Abstract
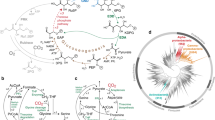
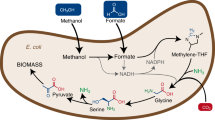
We engineered Escherichia coli to grow on CO2 and formic acid alone by introducing the synthetic CO2 and formic acid assimilation pathway, expressing two formate dehydrogenase genes, fine-tuning metabolic fluxes and optimizing the levels of cytochrome bo3 and bd-I ubiquinol oxidase. Our engineered strain can grow to an optical density at 600 nm of 7.38 in 450 h, and shows promise as a platform strain growing on CO2 and formic acid alone.


Similar content being viewed by others
Data availability
Data supporting our work are available in the paper, Extended Data Figs. 1–8 and Supplementary Information. Further information and materials related to the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Kumar, A. et al. Enhanced CO2 fixation and biofuel production via microalgae: recent developments and future directions. Trends Biotechnol. 28, 371–380 (2010).
Singh, A. K., Kishore, G. M. & Pakrasi, H. B. Emerging platforms for co-utilization of one-carbon substrates by photosynthetic organisms. Curr. Opin. Biotechnol. 53, 201–208 (2018).
Tashiro, Y., Hirano, S., Matson, M. M., Atsumi, S. & Kondo, A. Electrical-biological hybrid system for CO2 reduction. Metab. Eng. 47, 211–218 (2018).
Yishai, O., Bouzon, M., Döring, V. & Bar-Even, A. In vivo assimilation of one-carbon via a synthetic reductive glycine pathway in Escherichia coli. ACS Synth. Biol. 7, 2023–2028 (2018).
Döring, V., Darii, E., Yishai, O., Bar-Even, A. & Bouzon, M. Implementation of a reductive route of one-carbon assimilation in Escherichia coli through directed evolution. ACS Synth. Biol. 7, 2029–2036 (2018).
Innocent, B. et al. Electro-reduction of carbon dioxide to formate on lead electrode in aqueous medium. J. Appl. Electrochem. 39, 227 (2009).
Agarwal, A. S., Zhai, Y., Hill, D. & Sridhar, N. The electrochemical reduction of carbon dioxide to formate/formic acid: engineering and economic feasibility. ChemSusChem 4, 1301–1310 (2011).
Boddien, A. et al. CO2-“neutral” hydrogen storage based on bicarbonates and formates. Angew. Chem. Int. Ed. 50, 6411–6414 (2011).
Bang, J. & Lee, S. Y. Assimilation of formic acid and CO2 by engineered Escherichia coli equipped with reconstructed one-carbon assimilation pathways. Proc. Natl Acad. Sci. USA 115, E9271–E9279 (2018).
Gleizer, S. et al. Conversion of Escherichia coli to generate all biomass carbon from CO2. Cell 179, 1255–1263 (2019).
Kim, S. et al. Growth of E. coli on formate and methanol via the reductive glycine pathway. Nat. Chem. Biol. 16, 538–545 (2020).
Jeong, K. J. & Lee, S. Y. Enhanced production of recombinant proteins in Escherichia coli by filamentation suppression. Appl. Environ. Microbiol. 69, 1295–1298 (2003).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Song, C. W. & Lee, S. Y. Rapid one-step inactivation of single or multiple genes in Escherichia coli. Biotechnol. J. 8, 776–784 (2013).
Acknowledgements
We thank J. S. Cho at Systems Metabolic Engineering and Systems Healthcare Cross-Generation Collaborative Laboratory of KAIST for valuable advice during manuscript preparation. This work was supported by the C1 Gas Refinery Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT (NRF-2016M3D3A1A01913250).
Author information
Authors and Affiliations
Contributions
S.Y.L. conceived and coordinated the project; S.Y.L. and J.B. designed research; J.B. and C.H.H. performed experiments and analysed data; C.H.H., J.H.A. and J.A.L. performed microbial cultivations. J.B., C.H.H., J.H.A., J.A.L. and S.Y.L. wrote the manuscript and all authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have competing financial interests as the work described in this paper is covered by patents filed including, but not limited to, KR1020200086811, KR102000755, US16472876, DE112017006592.5, CN201780084543.8 and IL267579, and is of commercial interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Peer reviewer reports are available.
Extended data
Extended Data Fig. 1 Recombinant plasmids and growth profiles of the FC1, 2, and 3 strains cultured using small amount of glucose.
a, Recombinant plasmids used to develop engineered E. coli strains. b, Growth profiles of the FC1, 2, and 3 strains when 0.1 g L−1 of glucose was supplemented every 24 h. The orange line with an open circle represents the cell growth profile of the FC1 strain. The black line with an open diamond represents the cell growth profile of the FC2 strain. The pink line with open square represents the cell growth profile of the FC3 strain. Data presented are shown in average values with error bars representing ± standard deviations obtained in triplicate experiments (n = 3).
Extended Data Fig. 2 Isotope analyses of the FC1 and FC2 strains.
a, Metabolic pathways of E. coli synthesizing amino acids from CO2, FA, and glucose. Dashed lines represent the multiple-step pathway. b, 12C ratios in proteinogenic alanine (Ala) and phenylalanine (Phe) of the FC1 and 2 strains. P values were calculated by two-tailed Student’s t-test (*P, 0.0002; **P, 0.003). c, M + 0, M + 1, and M + 2 proteinogenic tyrosine ratios of the FC1 and FC2 strains. P values were calculated by two-tailed Student’s t-test (*P, 0.0103; **P, 0.0493; ***P, 0.002). d, 12C ratios in proteinogenic valine (Val), threonine (Thr), aspartate (Asp), glutamate (Glu), glycine (Gly), and serine (Ser) of the FC1 and 2 strains. Abbreviations of metabolites are: FBP, fructose 1,6-bisphosphate; E4P, erythrose 4-phosphate PEP, phosphoenolpyruvate; PYR, pyruvate, Ac-CoA, acetyl-CoA; OXA, oxaloacetate; 2KG, α-ketoglutarate; PHE, phenylalanine; TYR, tyrosine; GLY, glycine; SER, serine; ALA, alanine; VAL, valine; ASP, aspartate; THR, threonine; GLU, glutamate. Data presented are shown in average values with error bars representing ± standard deviation obtained in triplicate experiments (n = 3). Open circles in graphs indicate individual data points.
Extended Data Fig. 3 Growth of the FC5 strain using CO2 and FA, FA only, CO2 only, and without CO2 and FA and colony forming unit profile cultured solely on CO2 and FA.
a, Images of culture broth of the FC5 strain at 0 and 100 h after cultivation. FA + and FA – represent with and without FA supplementation, respectively. CO2 + and CO2 – represent with and without CO2 supplementation, respectively. b, Colony forming unit profile of the FC5 strain cultured using CO2 and FA only. Data presented are shown in average values with error bars representing ± standard deviation obtained in triplicate experiments (n = 3). Black dots in graphs indicate individual data points.
Extended Data Fig. 4 Increase of 12C carbon ratios of the proteinogenic amino acids in the FC5 strain.
Increase of 12C ratios in proteinogenic alanine (Ala), glycine (Gly), valine (Val), serine (Ser), threonine (Thr), phenylalanine (Phe), aspartate (Asp) and glutamate (Glu) of the FC5 strain after 100 h of culture solely from CO2 and FA. Data presented are shown in average values with error bars representing ± standard deviation obtained in triplicate experiments (n = 3). Open circles in graph indicate individual data points.
Extended Data Fig. 5 Optimization of IPTG concentration and aeration condition.
a, Growth profiles of the FC5 strain cultured with different concentrations of isopropyl β-D-1-thiogalactopyranoside (IPTG) as an inducer. The light gray line with an open triangle represents cell growth profile of the FC5 strain when 1 mM of IPTG was supplemented. The gray line with an open square represents the cell growth profile of the FC5 strain when 0.1 mM of IPTG was supplemented. The black line with an open circle represents the cell growth profile of the FC5 strain when 0.05 mM of IPTG was supplemented. b, Growth profiles of the FC5 strain cultured in different aeration conditions. The FC5 strain was cultured using 300 mL baffled flask with a working volume of 50 mL (FC5 High aeration, dark blue line with open circle), 100 mL baffled flask with a working volume of 30 mL (FC5 Medium aeration, blue line with open square), and 100 mL Erlenmeyer flask with a working volume of 30 mL (FC5 Low aeration, light blue line with open triangle), respectively. c, Relative messenger RNA (mRNA) ratios of the ppsA and gcvT genes in the FC5 strain cultured supplementing FA, gas mixture and different concentrations of IPTG (1, 0.1, and 0.05 mM). Relative mRNA ratios were calculated by setting ppsA mRNA to rrsA mRNA and gcvT mRNA to rrsA mRNA ratios as 1, respectively, when cells were cultured supplementing 1 mM of IPTG. Data presented are shown in average values with error bars representing ± standard deviation obtained in triplicate experiments (n = 3). Open circles in graph indicate individual data points.
Extended Data Fig. 6 CO2 and FA assimilation and FA consumption by the FC8 strain.
a, Increase of 12C ratios in proteinogenic alanine (Ala), glycine (Gly), valine (Val), serine (Ser), threonine (Thr), phenylalanine (Phe), aspartate (Asp), and glutamate (Glu) of the FC8 strain after 100 h of culture solely from CO2 and FA. b, Consumed FA by the FC5 and 8 strains after 100 h of cultivation. Red bars represent the consumed FA of the FC5 and FC8 strains. Data presented are shown in average values with error bars representing ± standard deviation obtained in triplicate experiments (n = 3). Open circles indicate individual data points.
Extended Data Fig. 7 Growth of the FC8 strain solely on CO2 and FA at lower culture temperatures.
a, Cell growth profiles of the FC8 strain cultured solely from FA and CO2 at 30 and 33 °C. The light blue line with an open circle represents the cell growth profile of the FC8 strain at 30 °C. The purple line with an open diamond represents the cell growth profile of the FC8 strain at 33 °C. The dashed line is the growth profile of the FC8 strain at 32 °C (Fig. 2d) and was provided for better comparison. b, Images of culture broths of the FC8 strain at 0 and 200 h after cultivation under different conditions. Cells were cultured at 32 °C. FA+ and FA– indicate with and without FA supplementation, respectively. CO2+ and CO2– indicate with and without CO2 supplementation, respectively. c, 12C ratios in proteinogenic alanine (Ala), glycine (Gly), valine (Val), serine (Ser), threonine (Thr), phenylalanine (Phe), aspartate (Asp), and glutamate (Glu) of the FC8 strain at 0, 140, and 260 h. Cells were first cultured using 13C labeled FA and U-13C-glucose. Afterward, cells were cultured at 32 °C using CO2 and FA only. d, Relative messenger RNA (mRNA) ratios of cyoA, cyoB, cydA, and cydB in the FC8 strain cultured at 32 °C compared with those in the FC8 strain cultured at 37 °C. Relative mRNA ratios were calculated by setting those four mRNA to rrsA ratios as 1, when cells were cultured at 37 °C. Data presented are shown in average values with error bars representing ± standard deviation obtained in triplicate experiments (n = 3). Black dots and open circles in graph indicate individual data points.
Extended Data Fig. 8 Growth properties of the FC8 strain in flask and bioreactor cultures.
a, Doubling time and specific growth rate profiles of the FC8 strain cultured at low initial inoculum (OD600 of 0.018). Data presented are shown in average values obtained in duplicate experiments (n = 2). b, Cell growth, FA concentration, and consumed FA profiles of a single bioreactor cultivation of the FC8 strain using agitation speed of 500 rpm. c, Doubling time and specific growth rate profiles of the Fig. 2g bioreactor culture. d–f, Reproduced bioreactor culture profiles of the FC8 strain solely on CO2 and FA. Data shown in d, e, and f are obtained from the second, third, and fourth bioreactor culture trials of the FC8 strain solely on CO2 and FA, respectively.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, Notes 1–10 and discussion.
Source data
Source Data Fig. 1
Data presented in Fig. 1.
Source Data Fig. 2
Data presented in Fig. 2.
Source Data Extended Data Fig. 1
Data presented in Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Data presented in Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Data presented in Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Data presented in Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Data presented in Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Data presented in Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Data presented in Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Data presented in Extended Data Fig. 8.
Rights and permissions
About this article
Cite this article
Bang, J., Hwang, C.H., Ahn, J.H. et al. Escherichia coli is engineered to grow on CO2 and formic acid. Nat Microbiol 5, 1459–1463 (2020). https://doi.org/10.1038/s41564-020-00793-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00793-9
- Springer Nature Limited
This article is cited by
-
Structure of recombinant formate dehydrogenase from Methylobacterium extorquens (MeFDH1)
Scientific Reports (2024)
-
Artificial synthesis of polyesters at ambient condition via consecutive CO2 electrolysis and fermentation
Nano Research (2024)
-
Turn air-captured CO2 with methanol into amino acid and pyruvate in an ATP/NAD(P)H-free chemoenzymatic system
Nature Communications (2023)
-
Discovery and remodeling of Vibrio natriegens as a microbial platform for efficient formic acid biorefinery
Nature Communications (2023)
-
Engineering a new-to-nature cascade for phosphate-dependent formate to formaldehyde conversion in vitro and in vivo
Nature Communications (2023)