Abstract
Viruses manipulate cellular signalling by inducing the degradation of crucial signal transducers, usually via the ubiquitin–proteasome pathway. Here, we show that the murine cytomegalovirus (Murid herpesvirus 1) M45 protein induces the degradation of two cellular signalling proteins, the nuclear factor κ-light-chain-enhancer of activated B cells (NF-κB) essential modulator (NEMO) and the receptor-interacting protein kinase 1 (RIPK1), via a different mechanism: it induces their sequestration as insoluble protein aggregates and subsequently facilitates their degradation by autophagy. Aggregation of target proteins requires a distinct sequence motif in M45, which we termed ‘induced protein aggregation motif’. In a second step, M45 recruits the retromer component vacuolar protein sorting 26B (VPS26B) and the microtubule-associated protein light chain 3 (LC3)-interacting adaptor protein TBC1D5 to facilitate degradation of aggregates by selective autophagy. The induced protein aggregation motif is conserved in M45-homologous proteins of several human herpesviruses, including herpes simplex virus, Epstein–Barr virus and Kaposi’s sarcoma-associated herpesvirus, but is only partially conserved in the human cytomegalovirus UL45 protein. We further show that the HSV-1 ICP6 protein induces RIPK1 aggregation and degradation in a similar fashion to M45. These data suggest that induced protein aggregation combined with selective autophagy of aggregates (aggrephagy) represents a conserved viral immune-evasion mechanism.






Similar content being viewed by others
Data availability
The datasets generated and analysed during the course of this study are available from the corresponding author upon request without restrictions. Uncropped western blot images of all figures in the manuscript (Figs. 1–5, Extended Data Figs. 3, 4, 6 and 7) and numerical data with statistical analysis (Figs. 2, 3 and 5) are provided as supplementary source data.
References
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Orzalli, M. H. & Kagan, J. C. Apoptosis and necroptosis as host defense strategies to prevent viral infection. Trends Cell Biol. 27, 800–809 (2017).
Boutell, C. & Everett, R. D. Regulation of alphaherpesvirus infections by the ICP0 family of proteins. J. Gen. Virol. 94, 465–481 (2013).
Lanfranca, M. P., Mostafa, H. H. & Davido, D. J. HSV-1 ICP0: an E3 ubiquitin ligase that counteracts host intrinsic and innate immunity. Cells 3, 438–454 (2014).
Mizushima, N., Yoshimori, T. & Ohsumi, Y. The role of Atg proteins in autophagosome formation. Annu. Rev. Cell. Dev. Biol. 27, 107–132 (2011).
Birgisdottir, A. B., Lamark, T. & Johansen, T. The LIR motif—crucial for selective autophagy. J. Cell Sci. 126, 3237–3247 (2013).
Zaffagnini, G. & Martens, S. Mechanisms of selective autophagy. J. Mol. Biol. 428, 1714–1724 (2016).
Choi, Y., Bowman, J. W. & Jung, J. U. Autophagy during viral infection—a double-edged sword. Nat. Rev. Microbiol. 16, 341–354 (2018).
Kudchodkar, S. B. & Levine, B. Viruses and autophagy. Rev. Med. Virol. 19, 359–378 (2009).
Dong, X. & Levine, B. Autophagy and viruses: adversaries or allies? J. Innate Immun. 5, 480–493 (2013).
Pellet, P. E. & Roizman, B. in Fields Virology 6th edn (eds Knipe, D., M. & Howley, P. M.) 1802–1822 (Lippincott Williams & Wilkins, 2013).
Brune, W., Ménard, C., Heesemann, J. & Koszinowski, U. H. A ribonucleotide reductase homolog of cytomegalovirus and endothelial cell tropism. Science 291, 303–305 (2001).
Mack, C., Sickmann, A., Lembo, D. & Brune, W. Inhibition of proinflammatory and innate immune signaling pathways by a cytomegalovirus RIP1-interacting protein. Proc. Natl Acad. Sci. USA 105, 3094–3099 (2008).
Upton, J. W., Kaiser, W. J. & Mocarski, E. S. Cytomegalovirus M45 cell death suppression requires receptor-interacting protein (RIP) homotypic interaction motif (RHIM)-dependent interaction with RIP1. J. Biol. Chem. 283, 16966–16970 (2008).
Upton, J. W., Kaiser, W. J. & Mocarski, E. S. Virus inhibition of RIP3-dependent necrosis. Cell Host Microbe 7, 302–313 (2010).
Rebsamen, M. et al. DAI/ZBP1 recruits RIP1 and RIP3 through RIP homotypic interaction motifs to activate NF-κB. EMBO Rep. 10, 916–922 (2009).
Upton, J. W., Kaiser, W. J. & Mocarski, E. S. DAI/ZBP1/DLM-1 complexes with RIP3 to mediate virus-induced programmed necrosis that is targeted by murine cytomegalovirus vIRA. Cell Host Microbe 11, 290–297 (2012).
Maelfait, J. et al. Sensing of viral and endogenous RNA by ZBP1/DAI induces necroptosis. EMBO J. 36, 2529–2543 (2017).
Fliss, P. M. et al. Viral mediated redirection of NEMO/IKKγ to autophagosomes curtails the inflammatory cascade. PLoS Pathog. 8, e1002517 (2012).
Krause, E., de Graaf, M., Fliss, P. M., Dölken, L. & Brune, W. Murine cytomegalovirus virion-associated protein M45 mediates rapid NF-κB activation after infection. J. Virol. 88, 9963–9975 (2014).
Hüttmann, J., Krause, E., Schommartz, T. & Brune, W. Functional comparison of molluscum contagiosum virus vFLIP MC159 with murine cytomegalovirus M36/vICA and M45/vIRA proteins. J. Virol. 90, 2895–2905 (2015).
Carisey, A., Stroud, M., Tsang, R. & Ballestrem, C. Fluorescence recovery after photobleaching. Methods Mol. Biol. 769, 387–402 (2011).
Brangwynne, C. P., Mitchison, T. J. & Hyman, A. A. Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc. Natl Acad. Sci. USA 108, 4334–4339 (2011).
Link, C. D. et al. Conversion of green fluorescent protein into a toxic, aggregation-prone protein by C-terminal addition of a short peptide. J. Biol. Chem. 281, 1808–1816 (2006).
Huang, Z. et al. RIP1/RIP3 binding to HSV-1 ICP6 initiates necroptosis to restrict virus propagation in mice. Cell Host Microbe 17, 229–242 (2015).
Lembo, D. et al. The ribonucleotide reductase R1 homolog of murine cytomegalovirus is not a functional enzyme subunit but is required for pathogenesis. J. Virol. 78, 4278–4288 (2004).
Mandal, P. et al. RIP3 induces apoptosis independent of pronecrotic kinase activity. Mol. Cell 56, 481–495 (2014).
Seaman, M. N., McCaffery, J. M. & Emr, S. D. A membrane coat complex essential for endosome-to-Golgi retrograde transport in yeast. J. Cell. Biol. 142, 665–681 (1998).
Swarbrick, J. D. et al. VPS29 is not an active metallo-phosphatase but is a rigid scaffold required for retromer interaction with accessory proteins. PLoS ONE 6, e20420 (2011).
Seaman, M. N. The retromer complex—endosomal protein recycling and beyond. J. Cell Sci. 125, 4693–470 (2012).
Collins, B. M. et al. Structure of Vps26B and mapping of its interaction with the retromer protein complex. Traffic 9, 366–379 (2008).
Bugarcic, A. et al. Vps26A and Vps26B subunits define distinct retromer complexes. Traffic 12, 1759–1773 (2011).
Popovic, D. et al. Rab GTPase-activating proteins in autophagy: regulation of endocytic and autophagy pathways by direct binding to human ATG8 modifiers. Mol. Cell Biol. 32, 1733–1744 (2012).
Johansen, T. & Lamark, T. Selective autophagy mediated by autophagic adapter proteins. Autophagy 7, 279–296 (2011).
Popovic, D. & Dikic, I. TBC1D5 and the AP2 complex regulate ATG9 trafficking and initiation of autophagy. EMBO Rep. 15, 392–401 (2014).
Lembo, D. & Brune, W. Tinkering with a viral ribonucleotide reductase. Trends Biochem. Sci. 34, 25–32 (2009).
Guo, H. et al. Herpes simplex virus suppresses necroptosis in human cells. Cell Host Microbe 17, 243–251 (2015).
Yu, X. et al. Herpes simplex virus 1 (HSV-1) and HSV-2 mediate species-specific modulations of programmed necrosis through the viral ribonucleotide reductase large subunit R1. J. Virol. 90, 1088–1095 (2016).
Alexander, D. E., Ward, S. L., Mizushima, N., Levine, B. & Leib, D. A. Analysis of the role of autophagy in replication of herpes simplex virus in cell culture. J. Virol. 81, 12128–12134 (2007).
Lussignol, M. et al. The herpes simplex virus 1 Us11 protein inhibits autophagy through its interaction with the protein kinase PKR. J. Virol. 87, 859–871 (2013).
Lamark, T. & Johansen, T. Aggrephagy: selective disposal of protein aggregates by macroautophagy. Int. J. Cell Biol. 2012, 736905 (2012).
Li, J. et al. The RIP1/RIP3 necrosome forms a functional amyloid signaling complex required for programmed necrosis. Cell 150, 339–350 (2012).
Ali, M., Roback, L. & Mocarski, E. S. Herpes simplex virus 1 ICP6 impedes TNF receptor 1-induced necrosome assembly during compartmentalization to detergent-resistant membrane vesicles. J. Biol. Chem. 294, 991–1004 (2019).
Cheng, A. Z. et al. Epstein–Barr virus BORF2 inhibits cellular APOBEC3B to preserve viral genome integrity. Nat. Microbiol. 4, 78–88 (2019).
Dufour, F. et al. The ribonucleotide reductase R1 subunits of herpes simplex virus types 1 and 2 protect cells against TNF-α and FasL-induced apoptosis by interacting with caspase-8. Apoptosis 16, 256–271 (2011).
Kwon, K. M., Oh, S. E., Kim, Y. E., Han, T. H. & Ahn, J. H. Cooperative inhibition of RIP1-mediated NF-κB signaling by cytomegalovirus-encoded deubiquitinase and inactive homolog of cellular ribonucleotide reductase large subunit. PLoS Pathog. 13, e1006423 (2017).
Carra, S., Seguin, S. J. & Landry, J. HspB8 and Bag3: a new chaperone complex targeting misfolded proteins to macroautophagy. Autophagy 4, 237–239 (2008).
Meriin, A. B. et al. Hsp70–Bag3 complex is a hub for proteotoxicity-induced signaling that controls protein aggregation. Proc. Natl Acad. Sci. USA 115, E7043–E7052 (2018).
Paul, P. & Münz, C. Autophagy and mammalian viruses: roles in immune response, viral replication, and beyond. Adv. Virus Res. 95, 149–195 (2016).
Kim, E. et al. Implication of mouse Vps26b–Vps29–Vps35 retromer complex in sortilin trafficking. Biochem. Biophys. Res. Commun. 403, 167–171 (2010).
Bresnahan, W. A., Hultman, G. E. & Shenk, T. Replication of wild-type and mutant human cytomegalovirus in life-extended human diploid fibroblasts. J. Virol. 74, 10816–10818 (2000).
Jordan, S. et al. Virus progeny of murine cytomegalovirus bacterial artificial chromosome pSM3fr show reduced growth in salivary glands due to a fixed mutation of MCK-2. J. Virol. 85, 10346–10353 (2011).
Tischer, B. K., Smith, G. A. & Osterrieder, N. En passant mutagenesis: a two step markerless red recombination system. Methods Mol. Biol. 634, 421–430 (2010).
Arase, H., Mocarski, E. S., Campbell, A. E., Hill, A. B. & Lanier, L. L. Direct recognition of cytomegalovirus by activating and inhibitory NK cell receptors. Science 296, 1323–1326 (2002).
Handke, W. et al. Viral inhibition of BAK promotes murine cytomegalovirus dissemination to salivary glands. J. Virol. 87, 3592–3596 (2013).
Brune, W., Hengel, H. & Koszinowski, U. H. A mouse model for cytomegalovirus infection. Curr. Protoc. Immunol. 43, 19.7.1–19.7.13 (2001).
Mahy, B. W. J. & Kangro, H. O. Virology Methods Manual (Academic Press, 1996).
Tanaka, M., Kagawa, H., Yamanashi, Y., Sata, T. & Kawaguchi, Y. Construction of an excisable bacterial artificial chromosome containing a full-length infectious clone of herpes simplex virus type 1: viruses reconstituted from the clone exhibit wild-type properties in vitro and in vivo. J. Virol. 77, 1382–1391 (2003).
Ashida, H. et al. A bacterial E3 ubiquitin ligase IpaH9.8 targets NEMO/IKKγ to dampen the host NF-κB-mediated inflammatory response. Nat. Cell Biol. 12, 66–73 (2010).
Ostermann, E. et al. Activation of E2F-dependent transcription by the mouse cytomegalovirus M117 protein affects the viral host range. PLoS Pathog. 14, e1007481 (2018).
van de Weijer, M. L. et al. A high-coverage shRNA screen identifies TMEM129 as an E3 ligase involved in ER-associated protein degradation. Nat. Commun. 5, 3832 (2014).
Chen, D. & Huang, S. Nucleolar components involved in ribosome biogenesis cycle between the nucleolus and nucleoplasm in interphase cells. J. Cell Biol. 153, 169–176 (2001).
Montespan, C. et al. Multi-layered control of Galectin-8 mediated autophagy during adenovirus cell entry through a conserved PPxY motif in the viral capsid. PLoS Pathog. 13, e1006217 (2017).
Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 16, 10881–10890 (1988).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Acknowledgements
We thank J. Connor, J. Bertin and E. Mocarski for RIPK3 kinase-dead mice, S. Jonjic, Y. Kawaguchi and R. Teasdale for reagents, F. Giraudo for technical assistance and T. Potgieter for critical readings of the manuscript. This study was supported by funding from Deutsche Forschungsgemeinschaft (BR 1730/3-2 to WB). The Heinrich Pette Institute is supported by the Free and Hanseatic City of Hamburg and the Federal Ministry of Health. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
E.M. designed, performed and analysed most experiments. R.S. performed a few biochemical experiments. E.K. and S.L. performed SILAC and AP–MS analyses. A.S. supervised the mass spectrometry analyses. E.C. performed and analysed FRAP experiments. M.R. generated and analysed TBC1D5-deficient cell clones. C.S. and R.R. performed correlative light and electron microscopy analyses. Y.-H.K. provided mice and biochemical reagents. V.J.L. provided biochemical reagents. E.O. performed in vivo experiments. W.B. designed and supervised the study, acquired funding and provided resources. E.M. and W.B. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 M45 induced aggregates.
(a) Atg5-/- MEFs were transfected with an M45-mCherry plasmid. 24 h post transfection cells were fixed and analyzed by fluorescence microscopy. Nuclei were stained with Hoechst 33342. Scale bar, 10 µm. (b) NIH-3T3 cells transfected with plasmids expressing HA-tagged full-length M45 or M45-Ct3. HA-tagged proteins were detected by immunofluorescence (green), protein aggregates by using the ProteoStat dye (red). (c) Another two ultrathin sections of the same WT MEF cell shown in Fig. 1f (ix – xi). The arrow points to a different aggregate than the one in Fig. 1f. Magnification of the same aggregate (A) close to an autophagosome (*) in the two different sections (x – xii). Scale bars, 5 µm (ix, xi) and 200 nm (x, xii). (d) Maximum intensity projection of NIH-3T3 cell transfected with a plasmid encoding M45-mCherry and observed 6 h post transfection by live cell imaging for 30 h. Scale bar, 10 µm. These data are representative of three (a-b) or two (d) biologically independent experiments. This data (c) is representative of three independent infected cells per group where at least 10 different sections (50 nm) per cells were analyzed.
Extended Data Fig. 2 Proteins co-purifying with M45 detected by AP-MS.
NIH-3T3 cells were labelled by SILAC and infected with MCMV WT or MCMV-M45HA. Cell lysates were harvested 15 hpi and subjected to anti-HA affinity-purification. Purified were analyzed by mass spectrometry.
Extended Data Fig. 3 M45 interacts with VPS26B, independent of VPS26B’s interaction with VPS35, and TBC1D5.
(a) Schematic of WT and mutant VPS26B. (b) HEK-293A cells were co-transfected with VPS26B-myc and VPS35-Flag plasmids. Immunoprecipitations were done as indicated. (c) HEK-293A cells co-transfected with plasmids expressing M45- HA and myc-tagged VPS26B or VPS26A (negative control). M45-HA was immunoprecipitated. (d) NIH-3T3-Vps26B-myc cells infected with MCMV-M45HA, MCMV-M45mut2HA or HA- tagged M45 C-terminal truncation mutants (MOI 3). Immunoprecipitation was done with an anti-HA antibody. (e) MEFs were infected with MCMV-M45HA or MCMVΔM45 (MOI 3). (f) VPS35, VPS26A, VPS29, and viral protein levels were determined by immunoblot at different times post infection. HEK-293A cells co-transfected with plasmids expressing M45-HA and Flag-TBC1D5 or Flag-IFI16 (negative control) or Flag-NEMO (positive control). HA was immunoprecipitated. (g) HEK-293A cells co-transfected with plasmids expressing M45- HA full-length or C-terminus truncation mutants (HA tagged) and Flag-TBC1D5. Immunoprecipitation was done with an anti-HA antibody. (h) WT and Vps26b-/- MEFs were infected with MCMV-M45HA (MOI 3). Immunoprecipitation was done with an anti-HA antibody. Immunoblot labels are in kDa. These data are representative of two (b-e-h) or three (c-d-f-g) biologically independent experiments.
Extended Data Fig. 4 HSV-1 ICP6 induces aggregate formation.
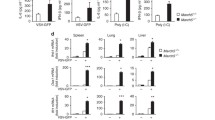
(a) Ripk3-/- fibroblasts were transfected with plasmids expressing WT or mutant ICP6. 24 h post transfection protein aggregates were detected by using the ProteoStat dye (red) and HA-tagged ICP6 by immunofluorescence (green). Scale bar, 10 µm. (b) HFF were infected with WT or ICP6mut HSV-1 (MOI 1). 24 hpi cells were fixed and stained for LC3BII (green). Nuclei were stained with Hoechst 33342. Scale bar, 10 µm. (c) HFF infected as in b. LC3BII was detected by immunoblot at 8 and 24 hpi. (d) HFF were infected with HSV-1 ICP6HA or ICP6mutHA (MOI 1). 24 hpi cells were fixed and stained for gamma-tubulin (green) and HA (red). Nuclei were stained with Hoechst 33342. Scale bar, 10 µm. Immunoblot labels are in kDa. These data are representative of two (a-b-c) or three (d) biologically independent experiments.
Extended Data Fig. 5 M45 aggregates co-localize with LC3BII.
(a, b) NIH-3T3 infected with MCMV-M45HA or MCMV- M45mut2HA (MOI 3). 24 hpi cells were fixed and stained for HA (red) and either LC3BII (a) or Caveolin-1 (b) (green). Nuclei were stained with Hoechst 33342. Scale bar, 10 µm. These data (a-b) are representative of three biologically independent experiments.
Extended Data Fig. 6 M45-interacting regions in NEMO and RIPK1 do not share a common motif.
(a) HEK-293A cells were co-transfected with plasmids expressing M45-HA and Flag-tagged RIPK1. Full-length (FL) and deletion mutants lacking the N-terminus, the C-terminus, or the death domain (DD) were used. M45-HA was immuno- precipitated and co-precipitating proteins were detected by immunoblot. (b) HEK-293A cells were co-transfected with plasmids expressing M45-HA and Flag-tagged NEMO (FL or N- terminal truncation mutants). M45-HA was immuno- precipitated and co-precipitating proteins were detected by immunoblot. (c) Sequence alignment of M45-interacting regions in NEMO and RIPK1 with APOBEC3B. Immunoblot labels (a-b) are in kDa. These data are representative of two (a) or three (b) biologically independent experiments.
Extended Data Fig. 7 M45 aggregates do not co-localize with HSP70.
(a) NIH-3T3 were infected with MCMV-M45HA or MCMV- M45mut2HA (MOI 3). 24 hpi cells were fixed and immunostained for HA (red) and HSP70 (green). Nuclei were stained with Hoechst 33342. Scale bar, 10 µm. (b) Immunoblot analysis of the soluble and insoluble fractions of MCMV- M45HA and MCMV-M45mut2 infected Atg5-/- MEFs (MOI 5). These data (a-b) are representative of three biologically independent experiments.
Supplementary information
Source data
Source Data Fig. 1
Unprocessed western blot.
Source Data Fig. 2
Unprocessed western blot.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Unprocessed western blot.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Unprocessed western blot.
Source Data Fig. 5
Unprocessed western blot.
Source Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 3
Unprocessed western blot.
Source Data Extended Data Fig. 4
Unprocessed western blot.
Source Data Extended Data Fig. 6
Unprocessed western blot.
Source Data Extended Data Fig. 7
Unprocessed western blot.
Rights and permissions
About this article
Cite this article
Muscolino, E., Schmitz, R., Loroch, S. et al. Herpesviruses induce aggregation and selective autophagy of host signalling proteins NEMO and RIPK1 as an immune-evasion mechanism. Nat Microbiol 5, 331–342 (2020). https://doi.org/10.1038/s41564-019-0624-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0624-1
- Springer Nature Limited
This article is cited by
-
The role of autophagy in viral infections
Journal of Biomedical Science (2023)
-
Master mitotic kinases regulate viral genome delivery during papillomavirus cell entry
Nature Communications (2023)
-
Targeting autophagy in prostate cancer: preclinical and clinical evidence for therapeutic response
Journal of Experimental & Clinical Cancer Research (2022)
-
Isoforms of autophagy-related proteins: role in glioma progression and therapy resistance
Molecular and Cellular Biochemistry (2022)
-
Protein clearance strategies for disease intervention
Journal of Neural Transmission (2022)