Abstract
Unidirectional fluid flow generated by motile cilia at the left–right organizer (LRO) breaks left–right (L–R) symmetry during early embryogenesis in mouse, frog and zebrafish. The chick embryo, however, does not require motile cilia for L–R symmetry breaking. The diversity of mechanisms for L–R symmetry breaking among vertebrates and the trigger for such symmetry breaking in non-mammalian amniotes have remained unknown. Here we examined how L–R asymmetry is established in two reptiles, Madagascar ground gecko and Chinese softshell turtle. Both of these reptiles appear to lack motile cilia at the LRO. The expression of the Nodal gene at the LRO in the reptilian embryos was found to be asymmetric, in contrast to that in vertebrates such as mouse that are dependent on cilia for L–R patterning. Two paralogues of the Nodal gene derived from an ancient gene duplication are retained and expressed differentially in cilia-dependent and cilia-independent vertebrates. The expression of these two Nodal paralogues is similarly controlled in the lateral plate mesoderm but regulated differently at the LRO. Our in-depth analysis of reptilian embryos thus suggests that mammals and non-mammalian amniotes deploy distinct strategies dependent on different Nodal paralogues for rendering Nodal activity asymmetric at the LRO.






Similar content being viewed by others
Data availability
All data are available in the main text, the Extended Data and the Supplementary Information. The molecular sequences identified in this study are included in Supplementary Dataset 1.
References
Blum, M., Feistel, K., Thumberger, T. & Schweickert, A. The evolution and conservation of left–right patterning mechanisms. Development 141, 1603–1613 (2014).
Shiratori, H. & Hamada, H. The left–right axis in the mouse: from origin to morphology. Development 133, 2095–2104 (2006).
Nonaka, S. et al. Randomization of left–right asymmetry due to loss of nodal cilia generating leftward flow of extraembryonic fluid in mice lacking KIF3B motor protein. Cell 95, 829–837 (1998).
Nakamura, T. et al. Fluid flow and interlinked feedback loops establish left–right asymmetric decay of Cerl2 mRNA. Nat. Commun. 3, 1322 (2012).
Montague, T. G., Gagnon, J. A. & Schier, A. F. Conserved regulation of Nodal-mediated left–right patterning in zebrafish and mouse. Development 145, dev171090 (2018).
Schweickert, A. et al. The nodal inhibitor Coco is a critical target of leftward flow in Xenopus. Curr. Biol. 20, 738–743 (2010).
Gros, J., Feistel, K., Viebahn, C., Blum, M. & Tabin, C. J. Cell movements at Hensen’s node establish left/right asymmetric gene expression in the chick. Science 324, 941–944 (2009).
Stephen, L. A. et al. The chicken left right organizer has nonmotile cilia which are lost in a stage-dependent manner in the talpid(3) ciliopathy. Genesis 52, 600–613 (2014).
Chang, C. F. et al. The cellular and molecular etiology of the craniofacial defects in the avian ciliopathic mutant talpid2. Development 141, 3003–3012 (2014).
Cui, C., Little, C. D. & Rongish, B. J. Rotation of organizer tissue contributes to left–right asymmetry. Anat. Rec. (Hoboken) 292, 557–561 (2009).
Kuraku, S. & Kuratani, S. Genome-wide detection of gene extinction in early mammalian evolution. Genome Biol. Evol. 3, 1449–1462 (2011).
Opazo, J. C., Kuraku, S., Zavala, K., Toloza-Villalobos, J. & Hoffmann, F. G. Evolution of nodal and nodal-related genes and the putative composition of the heterodimers that trigger the nodal pathway in vertebrates. Evol. Dev. 21, 205–217 (2019).
Hara, Y. et al. Madagascar ground gecko genome analysis characterizes asymmetric fates of duplicated genes. BMC Biol. 16, 40 (2018).
Yoshida, M. et al. Conserved and divergent expression patterns of markers of axial development in reptilian embryos: Chinese soft-shell turtle and Madagascar ground gecko. Dev. Biol. 415, 122–142 (2016).
Miller, J. D. Embryology of Marine Turtles (Wiley, 1985).
Hubert, J. Embryology of the Squamata (Wiley, 1985).
Stower, M. J. et al. Bi-modal strategy of gastrulation in reptiles. Dev. Dyn. 244, 1144–1157 (2015).
Bertocchini, F., Alev, C., Nakaya, Y. & Sheng, G. A little winning streak: the reptilian-eye view of gastrulation in birds. Dev. Growth Differ. 55, 52–59 (2013).
Coolen, M. et al. Molecular characterization of the gastrula in the turtle Emys orbicularis: an evolutionary perspective on gastrulation. PLoS ONE 3, e2676 (2008).
Collignon, J., Varlet, I. & Robertson, E. J. Relationship between asymmetric nodal expression and the direction of embryonic turning. Nature 381, 155–158 (1996).
Levin, M., Johnson, R. L., Stern, C. D., Kuehn, M. & Tabin, C. A molecular pathway determining left–right asymmetry in chick embryogenesis. Cell 82, 803–814 (1995).
Loges, N. T. et al. DNAI2 mutations cause primary ciliary dyskinesia with defects in the outer dynein arm. Am. J. Hum. Genet. 83, 547–558 (2008).
Shinohara, K. et al. Two rotating cilia in the node cavity are sufficient to break left–right symmetry in the mouse embryo. Nat. Commun. 3, 622 (2012).
Meno, C. et al. Left–right asymmetric expression of the TGF beta-family member lefty in mouse embryos. Nature 381, 151–155 (1996).
Meno, C. et al. Two closely-related left–right asymmetrically expressed genes, lefty-1 and lefty-2: their distinct expression domains, chromosomal linkage and direct neuralizing activity in Xenopus embryos. Genes Cells 2, 513–524 (1997).
Boettger, T., Wittler, L. & Kessel, M. FGF8 functions in the specification of the right body side of the chick. Curr. Biol. 9, 277–280 (1999).
Tanaka, C., Sakuma, R., Nakamura, T., Hamada, H. & Saijoh, Y. Long-range action of Nodal requires interaction with GDF1. Genes Dev. 21, 3272–3282 (2007).
Kawasumi, A. et al. Left–right asymmetry in the level of active Nodal protein produced in the node is translated into left–right asymmetry in the lateral plate of mouse embryos. Dev. Biol. 353, 321–330 (2011).
Adachi, H. et al. Determination of left/right asymmetric expression of nodal by a left side-specific enhancer with sequence similarity to a lefty-2 enhancer. Genes Dev. 13, 1589–1600 (1999).
Norris, D. P. & Robertson, E. J. Asymmetric and node-specific nodal expression patterns are controlled by two distinct cis-acting regulatory elements. Genes Dev. 13, 1575–1588 (1999).
Saijoh, Y. et al. Left–right asymmetric expression of lefty2 and nodal is induced by a signaling pathway that includes the transcription factor FAST2. Mol. Cell 5, 35–47 (2000).
Krebs, L. T. et al. Notch signaling regulates left–right asymmetry determination by inducing Nodal expression. Genes Dev. 17, 1207–1212 (2003).
Noro, M., Uejima, A., Abe, G., Manabe, M. & Tamura, K. Normal developmental stages of the Madagascar ground gecko Paroedura pictus with special reference to limb morphogenesis. Dev. Dyn. 238, 100–109 (2009).
Hara, Y. et al. Optimizing and benchmarking de novo transcriptome sequencing: from library preparation to assembly evaluation. BMC Genomics 16, 977 (2015).
Copeland, N. G., Jenkins, N. A. & Court, D. L. Recombineering: a powerful new tool for mouse functional genomics. Nat. Rev. Genet. 2, 769–779 (2001).
Gong, S. et al. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature 425, 917–925 (2003).
Takaoka, K. et al. The mouse embryo autonomously acquires anterior-posterior polarity at implantation. Dev. Cell 10, 451–459 (2006).
Saijoh, Y. et al. Distinct transcriptional regulatory mechanisms underlie left–right asymmetric expression of lefty-1 and lefty-2. Genes Dev. 13, 259–269 (1999).
Kuraku, S., Zmasek, C. M., Nishimura, O. & Katoh, K. aLeaves facilitates on-demand exploration of metazoan gene family trees on MAFFT sequence alignment server with enhanced interactivity. Nucleic Acids Res. 41, W22–W28 (2013).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Cunningham, F. et al. Ensembl 2019. Nucleic Acids Res. 47, D745–D751 (2019).
Nguyen, N. T. T., Vincens, P., Roest Crollius, H. & Louis, A. Genomicus 2018: karyotype evolutionary trees and on-the-fly synteny computing. Nucleic Acids Res. 46, D816–D822 (2018).
Acknowledgements
We thank S. Aizawa for sharing reptile cDNA clones and for advice on reptile embryos, A. Kawasumi for advice on the immunostaining of pSmad2/3 and K. Yamaguchi for technical discussion. This study was supported by grants from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT) of Japan (17H01435) and by Core Research for Evolutional Science and Technology (CREST) of the Japan Science and Technology Agency (JST) (JPMJCR13W5). U.H. was supported by the Global Science Campus of JST.
Author information
Authors and Affiliations
Contributions
H.H., E.K., U.H. and S.K. designed the project. E.K. performed most experiments with reptile embryos. U.H. performed the initial expression studies. T.I., K.Mizumo and K.Minegishi performed the SEM, TEM and ciliary motility analysis. Y.I. and H.N. generated transgenic mice harbouring various BAC clones. Y.H. and S.K. performed the genomic and phylogenetic analyses. H.K. maintained and provided the gecko animals. M.U. examined Nodal2 expression in chick embryos. H.H. and S.K. wrote the manuscript. All authors reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
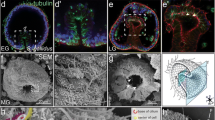
Extended Data Fig. 1 Detection of structures with features typical of immotile cilia in gecko embryos.
a–c, SEM of gecko embryos at stage 9. The anteroposterior (A-P) and dorsoventral (D-V) axes are indicated. The blastopore (BP) region indicated by the rectangle in a is shown at higher magnification in b. The boxed region in b is shown at higher magnification in c. Cilium-like and microvillus-like structures are indicated by red and white arrowheads, respectively. Scale bars: 100 µm (a), 50 µm (b), or 5 µm (c). d–h, TEM of gecko embryos at stage 9. The boxed regions near the blastopore (BP) in d are shown at higher magnification in e and g. The boxed regions in e and g are shown at higher magnification in f and h, respectively. Both f′ and f′′ are adjacent distal sections of f. The box in f′′ shows one of the doublet microtubules (black arrowhead) at higher magnification. Note that primary cilia with a typical 9+0 microtubule configuration were detected, but that outer dynein arms characteristic of motile cilia were not. Scale bars: 50 µm (d) or 1 µm (e, g).
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Dataset 1.
Supplementary Video 1
Movie of the region of the gecko embryo shown in Supplementary Fig. 7b. Note the absence of motile cilia. The images were captured at 150 frames per second for 4 s. Time is shown as minutes:seconds.centiseconds.
Supplementary Video 2
Movie of the region of the gecko embryo shown in Supplementary Fig. 7c. Note that a cilium (arrowhead) is present in this region but does not move. The images were captured at 30 frames per second for 10 s. Time is shown as minutes:seconds.centiseconds.
Supplementary Video 3
Movie of the region of the gecko embryo shown in Supplementary Fig. 7d. Note that a cilium (arrowhead) is present in this region but does not move. The images were captured at 30 frames per second for 10 s. Time is shown as minutes:seconds.centiseconds.
Rights and permissions
About this article
Cite this article
Kajikawa, E., Horo, U., Ide, T. et al. Nodal paralogues underlie distinct mechanisms for visceral left–right asymmetry in reptiles and mammals. Nat Ecol Evol 4, 261–269 (2020). https://doi.org/10.1038/s41559-019-1072-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-1072-2
- Springer Nature Limited
This article is cited by
-
Myosin1G promotes Nodal signaling to control zebrafish left-right asymmetry
Nature Communications (2024)
-
Discovery of a genetic module essential for assigning left–right asymmetry in humans and ancestral vertebrates
Nature Genetics (2022)