Abstract
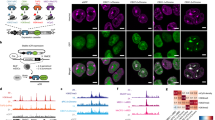
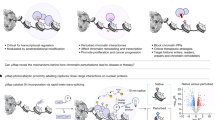
Elucidating the physiological binding partners of histone post-translational modifications (hPTMs) is key to understanding fundamental epigenetic regulatory pathways. Determining such interactomes will enable the study of how perturbations of these interactions affect disease. Here we use a synthetic biology approach to set a series of hPTM-controlled photo-affinity traps in native chromatin. Using quantitative proteomics, the local interactomes of these chemically customized chromatin landscapes are determined. We show that the approach captures transiently interacting factors such as methyltransferases and demethylases, as well as previously reported and novel hPTM reader proteins. We also apply this in situ proteomics approach to a recently disclosed cancer-associated histone mutation, H3K4M, revealing a number of perturbed interactions with the mutated tail. Collectively our studies demonstrate that modifying and interrogating native chromatin with chemical precision is a powerful tool for exploring epigenetic regulation and dysregulation at the molecular level.





Similar content being viewed by others
Data availability
All relevant data are included in the manuscript and supplementary information. Mass spectrometry data files have been uploaded to the PRIDE proteomics database (PXD017447).
References
Bannister, A. J. & Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381–395 (2011).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Dawson, M. A. & Kouzarides, T. Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012).
Weinberg, D. N., Allis, C. D. & Lu, C. Oncogenic mechanisms of histone H3 mutations. Cold Spring Harb. Perspect. Med. 7, a026443 (2017).
Bennett, R. L. et al. A mutation in histone H2B represents a new class of oncogenic driver. Cancer Discov. 9, 1438–1451 (2019).
Nacev, B. A. et al. The expanding landscape of ‘oncohistone’ mutations in human cancers. Nature 567, 473–478 (2019).
Vermeulen, M. et al. Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell 131, 58–69 (2007).
Vermeulen, M. et al. Quantitative interaction proteomics and genome-wide profiling of epigenetic histone marks and their readers. Cell 142, 967–980 (2010).
Li, X. et al. Quantitative chemical proteomics approach to identify post-translational modification-mediated protein-protein interactions. J. Am. Chem. Soc. 134, 1982–1985 (2012).
Machida, S. et al. Structural basis of heterochromatin formation by human HP1. Mol. Cell 69, 385–397 (2018).
Poepsel, S., Kasinath, V. & Nogales, E. Cryo-EM structures of PRC2 simultaneously engaged with two functionally distinct nucleosomes. Nat. Struct. Mol. Biol. 25, 154–162 (2018).
Bartke, T. et al. Nucleosome-interacting proteins regulated by DNA and histone methylation. Cell 143, 470–484 (2010).
Nikolov, M. et al. Chromatin affinity purification and quantitative mass spectrometry defining the interactome of histone modification patterns. Mol. Cell Proteomics 10, 1–16 (2011).
Local, A. et al. Identification of H3K4me1-associated proteins at mammalian enhancers. Nat. Genet. 50, 73–82 (2018).
Engelen, E. et al. Proteins that bind regulatory regions identified by histone modification chromatin immunoprecipitations and mass spectrometry. Nat. Commun. 6, 7155 (2015).
Ji, X. et al. Chromatin proteomic profiling reveals novel proteins associated with histone-marked genomic regions. Proc. Natl Acad. Sci. USA 112, 3841–3846 (2015).
Shah, R. N. et al. Examining the roles of H3K4 methylation states with systematically characterized antibodies. Mol. Cell 72, 162–177 (2018).
Suchanek, M., Radzikowska, A. & Thiele, C. Photo-leucine and photo-methionine allow identification of protein-protein interactions in living cells. Nat. Methods 2, 261–267 (2005).
Kleiner, R. E., Hang, L. E., Molloy, K. R., Chait, B. T. & Kapoor, T. M. A chemical proteomics approach to reveal direct protein-protein interactions in living cells. Cell Chem. Biol. 25, 110–120 (2018).
Zheng, Y., Gilgenast, M. J., Hauc, S. & Chatterjee, A. Capturing post-translational modification-triggered protein-protein interactions using dual noncanonical amino acid mutagenesis. ACS Chem. Biol. 13, 1137–1141 (2018).
Zheng, Y., Addy, P. S., Mukherjee, R. & Chatterjee, A. Defining the current scope and limitations of dual noncanonical amino acid mutagenesis in mammalian cells. Chem. Sci 8, 7211–7217 (2017).
David, Y., Vila-Perelló, M., Verma, S. & Muir, T. W. Chemical tagging and customizing of cellular chromatin states using ultrafast trans-splicing inteins. Nat. Chem 7, 394–402 (2015).
Stevens, A. J. et al. Design of a split intein with exceptional protein splicing activity. J. Am. Chem. Soc. 138, 2162–2165 (2016).
Stevens, A. J. et al. A promiscuous split intein with expanded protein engineering applications. Proc. Natl Acad. Sci. USA 114, 8538–8543 (2017).
Fischle, W. et al. Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438, 1116–1122 (2005).
Chang, Y., Horton, J. R., Bedford, M. T., Zhang, X. & Cheng, X. Structural insights for MPP8 chromodomain interaction with histone h3 lysine 9: Potential effect of phosphorylation on methyl-lysine binding. J. Mol. Biol. 408, 807–814 (2011).
Rothbart, S. B. et al. Association of UHRF1 with methylated H3K9 directs the maintenance of DNA methylation. Nat. Struct. Mol. Biol. 19, 1155–1160 (2012).
Lachner, M., O’Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001).
Bian, C., Chen, Q. & Yu, X. The zinc finger proteins ZNF644 and WIZ regulate the G9A/GLP complex for gene repression. eLife 4, e05606 (2015).
Schmitges, F. W. et al. Histone methylation by PRC2 is inhibited by active chromatin marks. Mol. Cell 42, 330–341 (2011).
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010).
Lewis, P. W. et al. Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340, 857–861 (2013).
Jang, Y. et al. H3.3K4M destabilizes enhancer H3K4 methyltransferases MLL3/MLL4 and impairs adipose tissue development. Nucleic Acids Res. 47, 607–620 (2019).
Wang, W. et al. Nucleolar protein Spindlin1 recognizes H3K4 methylation and stimulates the expression of rRNA genes. EMBO Rep. 12, 1160–1166 (2011).
Li, H. et al. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature 442, 91–95 (2006).
Wysocka, J. et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442, 86–90 (2006).
Guo, Y. et al. Methylation-state-specific recognition of histones by the MBT repeat protein L3MBTL2. Nucleic Acids Res 37, 2204–2210 (2009).
Chen, F. X. et al. PAF1 regulation of promoter-proximal pause release via enhancer activation. Science 357, 1294–1298 (2017).
Shi, X. et al. ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442, 96–99 (2006).
Peña, P. V. et al. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 442, 100–103 (2006).
Morgan, M. A. J. et al. A cryptic tudor domain links BRWD2/PHIP to COMPASS-mediated histone H3K4 methylation. Genes Dev. 31, 2003–2014 (2017).
Rotili, D. et al. A photoreactive small-molecule probe for 2-oxoglutarate oxygenases. Chem. Biol. 18, 642–654 (2011).
Sharifi-Zarchi, A. et al. DNA methylation regulates discrimination of enhancers from promoters through a H3K4me1-H3K4me3 seesaw mechanism. BMC Genomics 18, 964–985 (2017).
Zegerman, P., Canas, B., Pappin, D. & Kouzarides, T. Histone H3 lysine 4 methylation disrupts binding of nucleosome remodeling and deacetylase (NuRD) repressor complex. J. Biol. Chem. 277, 11621–11624 (2002).
Huang, Q. et al. Depletion of PHF14, a novel histone-binding protein gene, causes neonatal lethality in mice due to respiratory failure. Acta Biochim. Biophys. Sin 45, 622–633 (2013).
Saksouk, N. et al. HBO1 HAT complexes target chromatin throughout gene coding regions via multiple PHD finger with histone H3 tail. Mol. Cell 33, 257–265 (2009).
Gerace, M. et al. The scaffolding protein JADE1 physically links the acetyltransferase subunit HBO1 with its histone H3–H4 substrate. J. Biol. Chem. 293, 4498–4509 (2018).
Stevens, A. J., Sekar, G., Gramespacher, J. A., Cowburn, D. & Muir, T. W. An atypical mechanism of split intein molecular recognition and folding. J. Am. Chem. Soc. 140, 11791–11799 (2018).
Gramespacher, J. A., Burton, A. J., Guerra, L. F. & Muir, T. W. Proximity induced splicing utilizing caged split inteins. J. Am. Chem. Soc. 141, 13708–13712 (2019).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Acknowledgements
We thank F. Wojcik and R. Thompson and other members of the Allis and Muir laboratories for valuable discussions. We thank T. Srikumar, S. Kyin and H. Shwe from the Princeton Proteomics and Mass Spectrometry Core. A.J.B. is a Damon Runyon Fellow of the Damon Runyon Cancer Research Foundation (DRG-2283-17). J.D.B. was funded by a postdoctoral fellowship from the US National Institute of Health (GM123659). This work was supported by the US National Institutes of Health (NIH grants R37-GM086868 to T.W.M. and PO1-CA196539 to C.D.A. and T.W.M.).
Author information
Authors and Affiliations
Contributions
A.J.B. generated the modified histones in isolated nuclei and performed the cross-linking workflows. A.J.B. and M.H. generated the SILAC cell lines and performed fluorescence anisotropy experiments. A.J.B. and J.D.B. analysed the proteomics data. L.A.G. and C.D.A. contributed reagents and to data analysis. A.J.B. and T.W.M. conceived the project, analysed all data and wrote the manuscript with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures and Tables 1–28 (including uncropped gels displayed in Figs. 2–4).
Supplementary Data
Combined mass spectrometry datasets for the SILAC-based quantitative proteomics performed in this study.
Rights and permissions
About this article
Cite this article
Burton, A.J., Haugbro, M., Gates, L.A. et al. In situ chromatin interactomics using a chemical bait and trap approach. Nat. Chem. 12, 520–527 (2020). https://doi.org/10.1038/s41557-020-0474-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-020-0474-8
- Springer Nature Limited
This article is cited by
-
Spatiotemporal and direct capturing global substrates of lysine-modifying enzymes in living cells
Nature Communications (2024)
-
Single-electron transfer between sulfonium and tryptophan enables site-selective photo crosslinking of methyllysine reader proteins
Nature Chemistry (2024)
-
Interrogating epigenetic mechanisms with chemically customized chromatin
Nature Reviews Genetics (2024)
-
Tuning in to epigenetic cross-talk
Nature Methods (2023)
-
Tracking chromatin state changes using nanoscale photo-proximity labelling
Nature (2023)