Abstract
The early window of human embryogenesis is largely a black box for developmental biologists. Here we probed the cellular diversity of 4–6 week human embryos when essentially all organs are just laid out. On the basis of over 180,000 single-cell transcriptomes, we generated a comprehensive atlas of 313 clusters in 18 developmental systems, which were annotated with a collection of ontology and markers from 157 publications. Together with spatial transcriptome on embryonic sections, we characterized the molecule and spatial architecture of previously unappreciated cell types. Combined with data from other vertebrates, the rich information shed light on spatial patterning of axes, systemic temporal regulation of developmental progression and potential human-specific regulation. Our study provides a compendium of early progenitor cells of human organs, which can serve as the root of lineage analysis in organogenesis.






Similar content being viewed by others
Data availability
Sequencing data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE157329. An online depository for cell types and gene expression (developed with R package VisCello103 v1.1.2) is available at https://heoa.shinyapps.io/base/. Previously published data that were re-analysed here are from NCBI GSE119945 (single-nucleus RNA-seq in whole mouse embryos2), NCBI PRJNA637987 (scRNA-seq in mouse brain28), ArrayExpress E-MTAB-7320 (scRNA-seq in mouse neural tube26), ENCODE project available at https://cells.ucsc.edu/mouse-limb/10x/200120_10x.h5ad (scRNA-seq in mouse limb bud27), CNGB CNP0001543 (ST in mouse embryos29), NCBI GSE171892 (scRNA-seq in human neural tube63), NCBI GSE107618 (scRNA-seq in human retina9), NCBI GSE156793 (single-nucleus RNA-seq in human fetus7). For published datasets used in LIN28A analysis, see Supplementary Table 3. Mouse in situ database EMBRYS is available at https://www.embrys.jp/embrys/html/MainMenu.html. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
R scripts for analysis and figures are available at https://heoa.shinyapps.io/code/. All of the R packages used are available online. For the complete list of R packages and version numbers, see Supplementary Note.
References
Schoenwolf, G. C. & Larsen, W. J. (William J. Larsen’s Human Embryology (Churchill Livingstone/Elsevier, 2009).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Farrell, J. A. et al. Single-cell reconstruction of developmental trajectories during zebrafish embryogenesis. Science 360, eaar3131 (2018).
Wagner, D. E. et al. Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science 360, 981–987 (2018).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Han, X. et al. Construction of a human cell landscape at single-cell level. Nature 581, 303–309 (2020).
Cao, J. et al. A human cell atlas of fetal gene expression. Science 370, eaba7721 (2020).
Polioudakis, D. et al. A single-cell transcriptomic atlas of human neocortical development during mid-gestation. Neuron 103, 785–801.e8 (2019).
Hu, Y. et al. Dissecting the transcriptome landscape of the human fetal neural retina and retinal pigment epithelium by single-cell RNA-seq analysis. PLoS Biol. 17, e3000365 (2019).
Li, L. et al. Single-cell RNA-seq analysis maps development of human germline cells and gonadal niche interactions. Cell Stem Cell 20, 891–892 (2017).
Gao, S. et al. Tracing the temporal-spatial transcriptome landscapes of the human fetal digestive tract using single-cell RNA-sequencing. Nat. Cell Biol. 20, 721–734 (2018).
Popescu, D.-M. et al. Decoding human fetal liver haematopoiesis. Nature 574, 365–371 (2019).
Asp, M. et al. A spatiotemporal organ-wide gene expression and cell atlas of the developing human heart. Cell 179, 1647–1660.e19 (2019).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Young, M. D. et al. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. Science 361, 594–599 (2018).
Li, M. et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science 362, eaat7615 (2018).
Gao, R. et al. Delineating copy number and clonal substructure in human tumors from single-cell transcriptomes. Nat. Biotechnol. 39, 599–608 (2021).
Stefanovic, S., Etchevers, H. C. & Zaffran, S. Outflow tract formation—embryonic origins of conotruncal congenital heart disease. J. Cardiovasc. Dev. Dis. 8, 42 (2021).
Golzio, C. et al. ISL1 directly regulates FGF10 transcription during human cardiac outflow formation. PLoS ONE 7, e30677 (2012).
Alquicira-Hernandez, J., Sathe, A., Ji, H. P., Nguyen, Q. & Powell, J. E. ScPred: accurate supervised method for cell-type classification from single-cell RNA-seq data. Genome Biol. 20, 264 (2019).
Cabello-Aguilar, S. et al. SingleCellSignalR: inference of intercellular networks from single-cell transcriptomics. Nucleic Acids Res. 48, e55 (2020).
Capdevila, J. & Izpisua Belmonte, J. C. Patterning mechanisms controlling vertebrate limb development. Annu. Rev. Cell Dev. Biol. 17, 87–132 (2001).
Echevarria, D., Vieira, C., Gimeno, L. & Martinez, S. Neuroepithelial secondary organizers and cell fate specification in the developing brain. Brain Res. Brain Res. Rev. 43, 179–191 (2003).
Ramilowski, J. A. et al. A draft network of ligand–receptor-mediated multicellular signalling in human. Nat. Commun. 6, 7866 (2015).
Delile, J. et al. Single cell transcriptomics reveals spatial and temporal dynamics of gene expression in the developing mouse spinal cord. Development 146, dev173807 (2019).
He, P. et al. The changing mouse embryo transcriptome at whole tissue and single-cell resolution. Nature 583, 760–767 (2020).
La Manno, G. et al. Molecular architecture of the developing mouse brain. Nature 596, 92–96 (2021).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792.e21 (2022).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Trainor, P. A., Tan, S. S. & Tam, P. P. L. Cranial paraxial mesoderm: regionalisation of cell fate and impact on craniofacial development in mouse embryos. Development 120, 2397–2408 (1994).
Bumcrot, D. A. & McMahon, A. P. Somite differentiation: sonic signals somites. Curr. Biol. 5, 612–614 (1995).
Chen, B., Kwan, K. Y., Rubenstein, J. L. R. & Rakic, P. Patterning and Cell Type Specification in the Developing CNS and PNS: Comprehensive Developmental Neuroscience (Academic Press, 2020).
van Wijk, B., Moorman, A. F. M. & van den Hoff, M. J. B. Role of bone morphogenetic proteins in cardiac differentiation. Cardiovasc. Res. 74, 244–255 (2007).
Kim, S. H., Turnbull, J. & Guimond, S. Extracellular matrix and cell signalling: the dynamic cooperation of integrin, proteoglycan and growth factor receptor. J. Endocrinol. 209, 139–151 (2011).
Chiovaro, F., Chiquet-Ehrismann, R. & Chiquet, M. Transcriptional regulation of tenascin genes. Cell Adh. Migr. 9, 34 (2015).
Benazet, J. D. & Zeller, R. Vertebrate limb development: moving from classical morphogen gradients to an integrated 4-dimensional patterning system. Cold Spring Harb. Perspect. Biol. 1, a001339 (2009).
Yokoyama, S. et al. A systems approach reveals that the myogenesis genome network is regulated by the transcriptional repressor RP58. Dev. Cell 17, 836–848 (2009).
Capellini, T. D. et al. Pbx1/Pbx2 requirement for distal limb patterning is mediated by the hierarchical control of Hox gene spatial distribution and Shh expression. Development 133, 2263–2273 (2006).
Coudert, A. E. et al. Expression and regulation of the Msx1 natural antisense transcript during development. Nucleic Acids Res. 33, 5208–5218 (2005).
Xu, B. et al. Hox5 interacts with Plzf to restrict Shh expression in the developing forelimb. Proc. Natl Acad. Sci. USA 110, 19438–19443 (2013).
Capellini, T. D. et al. Scapula development is governed by genetic interactions of Pbx1 with its family members and with Emx2 via their cooperative control of Alx1. Development 137, 2559–2569 (2010).
Kuijper, S. et al. Function and regulation of Alx4 in limb development: complex genetic interactions with Gli3 and Shh. Dev. Biol. 285, 533–544 (2005).
Cooper, K. L. et al. Patterning and post-patterning modes of evolutionary digit loss in mammals. Nature 511, 41–45 (2014).
Firulli, B. A. et al. Altered Twist1 and Hand2 dimerization is associated with Saethre–Chotzen syndrome and limb abnormalities. Nat. Genet. 37, 373–381 (2005).
Akiyama, H. et al. Osteo-chondroprogenitor cells are derived from Sox9 expressing precursors. Proc. Natl Acad. Sci. USA 102, 14665–14670 (2005).
Abassah-Oppong, S. et al. A gene desert required for regulatory control of pleiotropic Shox2 expression and embryonic survival. Preprint at bioRxiv https://doi.org/10.1101/2020.11.22.393173 (2020).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Vargesson, N. et al. Cell fate in the chick limb bud and relationship to gene expression. Development 124, 1909–1918 (1997).
Zakany, J. & Duboule, D. The role of Hox genes during vertebrate limb development. Curr. Opin. Genet. Dev. 17, 359–366 (2007).
Zhang, B. et al. A human embryonic limb cell atlas resolved in space and time. Preprint at bioRxiv https://doi.org/10.1101/2022.04.27.489800 (2022).
Lefebvre, V. & Smits, P. Transcriptional control of chondrocyte fate and differentiation. Birth Defects Res. Part C 75, 200–212 (2005).
Kobayashi, T. et al. Indian hedgehog stimulates periarticular chondrocyte differentiation to regulate growth plate length independently of PTHrP. J. Clin. Invest. 115, 1734–1742 (2005).
Niedermaier, M. et al. An inversion involving the mouse Shh locus results in brachydactyly through dysregulation of Shh expression. J. Clin. Invest. 115, 900–909 (2005).
Duboule, D. The (unusual) heuristic value of Hox gene clusters; a matter of time?. Dev. Biol. 484, 75–87 (2002).
Philippidou, P. & Dasen, J. S. Hox genes: choreographers in neural development, architects of circuit organization. Neuron 80, 12–34 (2013).
Le Dreau, G. & Marti, E. Dorsal-ventral patterning of the neural tube: a tale of three signals. Dev. Neurobiol. 72, 1471–1481 (2012).
Uehara, M. et al. CYP26A1 and CYP26C1 cooperatively regulate anterior-posterior patterning of the developing brain and the production of migratory cranial neural crest cells in the mouse. Dev. Biol. 302, 399–411 (2007).
Sarropoulos, I., Marin, R., Cardoso-Moreira, M. & Kaessmann, H. Developmental dynamics of lncRNAs across mammalian organs and species. Nature 571, 510–514 (2019).
De Kumar, B. & Krumlauf, R. HOXs and lincRNAs: two sides of the same coin. Sci. Adv. 2, e1501402 (2016).
Zhang, X. et al. A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood 113, 2526–2534 (2009).
Wilson, L. & Maden, M. The mechanisms of dorsoventral patterning in the vertebrate neural tube. Dev. Biol. 282, 1–13 (2005).
Rayon, T., Maizels, R., Barrington, C. & Briscoe, J. Single-cell transcriptome profiling of the human developing spinal cord reveals a conserved genetic programme with human-specific features. Development 148, dev199711 (2021).
Moss, E. G. Heterochronic genes and the nature of developmental time. Curr. Biol. 17, R425–R434 (2007).
Ambros, V. & Horvitz, H. Heterochronic mutants of the nematode Caenorhabditis elegans. Science 226, 409–416 (1984).
Boroviak, T. et al. Lineage-specific profiling delineates the emergence and progression of naive pluripotency in mammalian embryogenesis. Dev. Cell 35, 366–382 (2015).
Tan, M. H. et al. RNA sequencing reveals a diverse and dynamic repertoire of the Xenopus tropicalis transcriptome over development. Genome Res. 23, 201–216 (2013).
Briggs, J. A. et al. The dynamics of gene expression in vertebrate embryogenesis at single-cell resolution. Science 360, eaar5780 (2018).
White, R. J. et al. A high-resolution mRNA expression time course of embryonic development in zebrafish. eLife 6, e30860 (2017).
Yang, D.-H. & Moss, E. G. Temporally regulated expression of Lin-28 in diverse tissues of the developing mouse. Gene Expr. Patterns 3, 719–726 (2003).
Faas, L. et al. Lin28 proteins are required for germ layer specification in Xenopus. Development 140, 976–986 (2013).
Nieuwkoop, P. D. & Faber, J. Normal Table of Xenopus laevis (Daudin) (Taylor and Francis, 1994).
Carroll, E. J. & Hedrick, J. L. Hatching in the toad Xenopus laevis: morphological events and evidence for a hatching enzyme. Dev. Biol. 38, 1–13 (1974).
Irie, N. & Kuratani, S. Comparative transcriptome analysis reveals vertebrate phylotypic period during organogenesis. Nat. Commun. 2, 248 (2011).
Cardoso-Moreira, M. et al. Gene expression across mammalian organ development. Nature 571, 505–509 (2019).
Shyh-Chang, N. & Daley, G. Q. Lin28: primal regulator of growth and metabolism in stem cells. Cell Stem Cell 12, 395–406 (2013).
Wilbert, M. L. et al. LIN28 binds messenger RNAs at GGAGA motifs and regulates splicing factor abundance. Mol. Cell 48, 195–206 (2012).
Agarwal, V., Bell, G. W., Nam, J.-W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. eLife 4, e05005 (2015).
Shinoda, G. et al. Fetal deficiency of lin28 programs life-long aberrations in growth and glucose metabolism. Stem Cells 31, 1563–1573 (2013).
Robinton, D. A. et al. The Lin28/let-7 pathway regulates the mammalian caudal body axis elongation program. Dev. Cell 48, 396–405.e3 (2019).
Komarovsky Gulman, N., Armon, L., Shalit, T. & Urbach, A. Heterochronic regulation of lung development via the Lin28-Let-7 pathway. FASEB J. 33, 12008–12018 (2019).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Chu, Y. & Liu, T. On the shortest arborescence of a directed graph. Sci. Sin. 14, 1396–1400 (1965).
Marquardt, T. & Gruss, P. Generating neuronal diversity in the retina: one for nearly all. Trends Neurosci. 25, 32–38 (2002).
Fuhrmann, S. Eye morphogenesis and patterning of the optic vesicle. Curr. Top. Dev. Biol. 93, 61–84 (2010).
Theos, A. C., Truschel, S. T., Raposo, G. & Marks, M. S. The Silver locus product Pmel17/gp100/Silv/ME20: controversial in name and in function. Pigment Cell Res. 18, 322–336 (2005).
Ma, X. et al. The transcription factor MITF in RPE function and dysfunction. Prog. Retin. Eye Res. 73, 100766 (2019).
Regev, A. et al. The Human Cell Atlas. eLife 6, e27041 (2017).
Regev, A. et al. The Human Cell Atlas White Paper (2018).
Tyser, R. C. V. et al. Single-cell transcriptomic characterization of a gastrulating human embryo. Nature 600, 285–289 (2021).
Moris, N. et al. An in vitro model of early anteroposterior organization during human development. Nature 582, 410–415 (2020).
O’Rahilly, R. & Müller, F. Developmental stages in human embryos: revised and new measurements. Cells Tissues Organs 192, 73–84 (2010).
Kaufman, M. H. The Atlas of Mouse Development (Academic Press, 1992).
SL, W., R, L. & AM, K. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Zhang, T. et al. A single-cell analysis of the molecular lineage of chordate embryogenesis. Sci. Adv. 6, eabc4773 (2020).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Santos, A., Wernersson, R. & Jensen, L. J. Cyclebase 3.0: a multi-organism database on cell-cycle regulation and phenotypes. Nucleic Acids Res. 43, D1140–D1144 (2015).
Giotti, B. et al. Assembly of a parts list of the human mitotic cell cycle machinery. J. Mol. Cell. Biol. 11, 703–718 (2019).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014.e22 (2018).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Packer, J. S. et al. A lineage-resolved molecular atlas of C. elegans embryogenesis at single cell resolution. Science 365, eaax1971 (2019).
Acknowledgements
We thank R. Sheth, H. Etchevers, H. Zhang, S. Teichmann, C. Zhang, D. Huangfu, A. Joyner, L. Studer and G. Lin for discussions. We thank L. Xiao for website design. This work is supported by Funding Project of National Key Research and Development Program of China (2018YFD0900604), Natural Science Foundation of China (41676119 and 41476120) and Fundamental Research Funds for the Central Universities from the Ocean University of China to W.S. Z.B. is supported by funding from Memorial Sloan Kettering Cancer Center. T.Z. is a visiting student at Memorial Sloan Kettering Cancer Center sponsored by the China Scholarship Council.
Author information
Authors and Affiliations
Contributions
W.S. and Z.B. conceived the project. T.Z., M.H., Y.Q., L.W., Q.Z. and Y.X. collected the embryos and generated the atlas dataset. Y.X., T.Z. and Y.N. performed data processing, clustering and all the bioinformatics analysis. Y.X. created the atlas website. W.S., Z.B., Y.X. and T.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
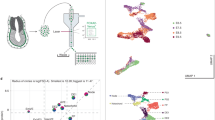
Extended Data Fig. 1 Embryos and data quality.
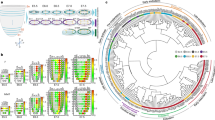
a, Images of embryos and the number of high-quality cells in each sample. Schematic diagrams (lower panel) showing the territory of embryo dissection parts. b, Histograms showing the distribution of total UMIs and gene numbers per cell. c, Distribution of XIST (female) versus RSP4Y1 (male) expression from randomly sampled 1,000 cells per embryo. d, UMIs of hemoglobin genes (HBA1, HBA2, HBE1, HBG1, HBG2, HBZ) against total UMIs. Each dot denotes a cell. Cells with high linear slope (above the green line) was identified as erythroid cells (red) (see Methods). Black dots denote cells that were not identified as erythroid cells. e, CopyKAT18, designed for 3’ or 5’ scRNA-seq with sparse coverage, was used to estimate copy number aberration in 7 embryos with default setting. 500 cells were randomly sampled in each embryo. Log2 ratios of copy number of segmentations (220 kb of size) were shown by embryo (left) and by genomic position (right).
Extended Data Fig. 2 Annotation of cell types.
Expression of selected known markers for cell types collapsed by unique terms in annotation (For all markers, see Supplementary Table 1). Cell types in limb and neural tube are detailed in Fig. 3 and Fig. 4, respectively. Dendrogram of cell types was defined according to the lineage relationship based on annotations. The third level of dendrogram shows 18 developmental systems and number of clusters in each system. Dot size represents the fraction of expressing cells (UMIs, unique molecular identifiers > 0) for a given gene (n = 7 embryos) and dot color shows developmental system. EP, epidermis; SN, sensory neuron; IM, intermediate mesoderm; soLPM, somatic LPM; spLPM, splanchic LPM; PGC, primordial germ cell; PA, pharyngeal arch; ZLI, zona limitans intrathalamica; AER, apical ectodermal ridge; NMP, neuro-mesodermal progenitor; a/pPSM, anterior/posterior presomitic mesoderm.
Extended Data Fig. 3 Iterative clustering and quality control.
a, The expression of markers in first round of clustering (solid lines) and second round of clustering (dashed lines) in brain. Markers that support second round of clustering were shown. Two color bars on top denote clustering results of first round and second round, respectively. b, The expression of markers relative to boundaries of one round of clustering and second round of clustering in endothelium. Convention follows panel a. c, The number of clusters resulted from a series of resolution ‘r’ and PCs in the clustering of spinal neuron. d, The pairwise ARI between clusters resulted from different resolution ‘r’ in spinal neuron. e, The mean ARI of clusters in testing resolution ‘r’ and PCs (Methods). Each dot denotes a super-cluster. f, Cross-validation on clustering by scPred21. The first column shows the AUROC (area under receiver operating curve) of testing the identity of developmental systems (each red dot is a system). Other columns show the AUROC of testing ‘Celltype_annotation’ and ‘Final_annotation’ (Supplementary Table 1) within each system (expect PGC) in red and blue dots, respective (Methods, n = 3~59 cell types). Testing of randomly shuffled identity was served as control in each column (grey). The center line denotes the median, while the box contains the 25th to 75th percentiles. The whiskers mark 1.5x interquartile range. g, Batch effects of embryos, technical replicates, cell cycle phase, and total UMIs estimated by the entropy of mixing100 (Methods, n = 1,700 randomly sampled cells). ‘+Ctrl’, cluster identities as batch. ‘-Ctrl’, randomly assigned batch. Boxplots are defined as in panel f. h, UMAP of all cells colored by embryos, technical replicates, cell cycle phase, total UMIs, and developmental systems in each embryo. The missing parts in embryos 03, 06, and 07 are due to 4 libraries that did not pass quality control (Methods).
Extended Data Fig. 4 Dissection parts, embryonic stages, and DEGs of cell types.
a, The origin of dissection parts of cells in each cluster ordered by system. See Supplementary Table 1 for the order of clusters. EP, epidermis; SN, sensory neuron; IM, intermediate mesoderm; LPM, lateral plate mesoderm; PGC, primordial germ cell; misc., miscellaneous. b, Stage distribution of cells in each cluster ordered by system. * denotes clusters missing cells from CS13-14, defined by total number of cells > 50, ratio of cells from CS12 embryo > 0.05, number of cells from CS12 embryo > 5, ratio of cells from CS15-16 embryo > 0.05, number of cells from CS15-16 embryo > 5, ratio of cells from CS13-14 embryo < 0.05, and number of cells from CS13-14 embryo < 5. c, Number of DEGs per cell type/cluster for each developmental system (see Fig. 1 for convention). The center line denotes the median, while the box contains the 25th to 75th percentiles. The whiskers mark 1.5x interquartile range. n = 3~59 cell types except PGC. d, The expression of ligands in DEGs of 9 signaling centers in human (top) and their expression in mouse cell types from published data26,27,28 (bottom) (Methods, n = 7 embryos). Asterisks denote ligands that are also expressed in the corresponding cell types in mouse. ANR, anterior neural ridge; MHB, mid-hindbrain boundary; ZLI, zona limitans intrathalamica; FP.b/s, floor plate (brain, spinal cord); RP.b/s, roof plate (brain, spinal cord); AER, apical ectodermal ridge; ZPA, zone of polarizing activity. e, The number of ligands that are expressed or not in mouse by signaling center. For each signaling center, left bar shows known ligands and right bar shows additional ligands.
Extended Data Fig. 5 Quality control of spatial transcriptome.
a, The distribution of total UMIs and gene numbers per spot. b, The distribution of total UMIs and gene numbers on each section. c, The co-clustering of 2 human sections and 2 published mouse sections29 on the spot level (Methods). Cluster identities of spots are indicated by color and number on each tissue section. d, The correlation of spatial neighborhood of each cluster between human and mouse. The spatial neighborhood for a cluster on a section was defined as the proportions of clusters in its neighbors. The neighbors of a cluster were defined as the union of nearest 6 spots on the section of each spot in this cluster, excluding spots from the same cluster. Human section 1 was compared to mouse E2S1 and human section 2 was compared to mouse E1S2. The color and number of each cluster correspond to panel c. e, The proportion of 5 recognizable structures on H&E staining of section 1 in deconvolution. f, The proportion of structures that are difficult to be recognized only by H&E staining on section 1 (first row) and section 2 (second row) in deconvolution. Five structures are shown in each section as examples.
Extended Data Fig. 6 Deconvolution, head mesoderm, and signaling interaction in spatial transcriptome.
a, The deconvolution result of selected cell types on section 1. From left to right, second heart field (SHF) along with pharynx arches (PA) and heart, sclerotome, arterial and vascular endothelium. Arrows in the panel of SHF denote the detection of SHF in PA 1/2, PA 3/4, and heart. b, The UMAP of cells in head mesoderm in scRNA-seq colored by cell type/cluster. c, The average z-score of DEGs of the 5 undefined cell types in each spot of section 1. The window of ST is the same with that in Fig. 2e. Only top DEGs from scRNA-seq shown in Fig. 2e were used. d, The comparison of detection of cell types between scRNA-seq and ST. Each dot is a cell type in head mesoderm and red dots are the 5 undefined cell types. Pearson’s correlation was calculated on the detection of 5 undefined cell types between scRNA-seq and section 1 (mainly mesodermal tissues), between scRNA-seq and section 2 (mainly ectodermal tissues). To be consistent on developmental stage between scRNA-seq and ST, only cells from CS13-CS14 were counted in scRNA-seq. e, The pipeline of inferring signaling interaction by scRNA-seq and spatial transcriptome (Methods). f, Examples of known signaling interactions identified by integrating scRNA-seq and ST. For all significant interactions, see Supplementary Table 1. Top left, the -log10 adjusted p value of selected signaling interactons on section 2. Each row is a pair of cell types and each column is a pair of ligand-receptor. Bottom left, the proportion of cell types involved in these interactions in ST. Right panel, the expression of ligands and receptors in these interaction by pathway. g, The expression of genes in ST involved in signaling interaction shown in Fig. 2g. First row is ligand and second row is receptor.
Extended Data Fig. 7 Spatial domains in limb mesenchymal cells.
a, UMAP and RNA velocity analysis of human forelimb at CS12 and CS15-16 (see Fig. 3 for conventions). b, Batch effect of embryo estimated by entropy of mixing at CS13-14 and CS15-16 (Methods, Mann-Whitney U test p < 2.2 × 10−16, n = 100 randomly sampled cells). The center line denotes the median, while the box contains the 25th to 75th percentiles. The whiskers mark 1.5x interquartile range. c, Normalized UMIs of top DEGs of domain m in domain m by embryo. d, UMAP of mouse forelimb at E10.5 and E12 by reclustering a published dataset27 (Supplementary Table 2). e, The expression of marker genes in each domain from human scRNA-seq (red), mouse scRNA-seq (blue), and mouse in situ data (black) at three stages. Mouse in situ data in each domain were manually classified to strong (large dot), weak (small dot) and no expression (no dot) based on in situ images. Domain m was not identified in E12 data of mouse. HOX genes (black boxes) show developmental stages are aligned between human and mouse datasets. f, The expression of specific markers of domain m on UMAP in human and mouse. Dashed circles, domain m. g, Left panel, UMAP co-embedding of CS15-16 of human and E12 of mouse colored by species and domains (see panel a for conventions). Right panel, the average percentage of mouse cells in k-nearest neighbors (KNN) of human cells in each domain in co-embedding space (Methods, mean +/− SEM, number of cells indicated above each bar). The first bar (background) denotes the percentage of mouse cells in total cells of human and mouse. Asterisks denote significant p values by Mann-Whitney U test (domain a, 3 × 10−7; domain b, 5 × 10−7; domain m, 9 × 10−9). In all panels, n = 1, 3, 3 embryos at CS12, CS13-14, and CS15-16, respectively.
Extended Data Fig. 8 Neural tube patterning.
a, LncRNAs in AP patterning. Upper panel, Monocle analysis shown in pseudo value and cell type (see Fig. 4d for conventions). Middle panel, schematic diagram of the genomic location of the 5 lncRNAs identified as AP related genes in neural tube. Lower-left panel, the expression pattern of 5 lncRNAs (solid lines, see left bar of the heatmap for color legend) and nearby HOX genes (dashed lines) along the AP axis on section 2 of ST. Lower-right panel, the division between hindbrain and spinal cord on H&E staining of section 2, and the correlation on gene expression between lncRNAs and HOX genes along AP axis of section 2 (Methods). b, The comparison of expression of canonical markers for cell types in neural progenitors (upper panel) and neurons (lower panel) between human and mouse (n = 7 and 6 embryos, respectively). Box denotes the difference of MSX2 between human and mouse in neural progenitors. c, The comparison of expression of PAX7 and NKX6-2 between human and mouse in neural progenitors (n = 7, 3, and 6 embryos, respectively). Boxes denote human specific expression that are consistent between two datasets of human. d, The expression of markers (dp1 marker MSX1 and dp4 marker ASCL1, log2 scaled) that distinguish dp1 and dp4 cells in individual cells in human (our dataset and Rayon’s dataset) and mouse datasets. Dot size shows MSX2 expression. Dashed boxes denote most confident dp4 cells (ASCL1 > 0 and MSX1 = 0), which are shown in Fig. 4f. e, The expression of MSX2 in dp2-4 in each human embryo in our data (mean + /- SEM). Each dot denotes a cell (n = 34~209 cells). The number on top is mean of normalized UMIs in each embryo. Note the trunk sample of Emb.07 did not pass quality control so that no cell is from Emb.07.
Extended Data Fig. 9 LIN28A in vertebrate embryogenesis.
a, Expression pattern of 4 groups of TFs in zebrafish and frog. Each group of TFs were identified as TFs that are highly expressed in the corresponding stage. Numbers above curves indicate number of TFs in each group. The width of each curve represents 2 standard deviations within each group of TFs. The black lines show the expression of LIN28A in each species. b, Pairwise correlation of timepoints between species by homologous TFs. Correlation is scaled by row (Methods). Black line denotes the alignment of timepoints between species by dynamic time warping. The yellow asterisks denote the match of timepoints from a previous study74. c, Numbers of systemically up- (red) and down-regulated (blue) genes from stage 1 to stage 2 (labeled as 1→2) and from stage 3 to stage 4 (labeled as 3→4). No transcriptome data is available for stages 1 and 2 in human. d, Expression dynamics of genes in cell cycle, mRNA splicing and translation pathways that are positively correlated to LIN28A in zebrafish and frog.
Extended Data Fig. 10 Match of cell types between embryo and fetus.
The mutual best match (red edge), best match (black edge) and 2nd-best match (grey edge) in our dataset for each cell type in Cao’s dataset by Slingshot (also see Supplementary Table 6). Cell types are colored by developmental system (see Fig. 1 for conventions). The thickness of lines is inversely proportional to z-score across distances between a cell type in Cao’s dataset and all cell types in our dataset. The lineage relationship between cell types within our dataset (orange edge) was determined by annotation.
Supplementary information
Supplementary Information
Supplementary Note.
Supplementary Tables
Supplementary Table 1. Cell types and DEGs. Supplementary Table 2. The comparison between human and mouse scRNA-seq (whole embryo and limb). Supplementary Table 3. LIN28A in vertebrate embryogenesis. Supplementary Table 4. Systemically changing genes in vertebrate embryogenesis. Supplementary Table 5. Enriched pathways in systemically changing genes. Supplementary Table 6. Data integration in human embryos.
Source data
Source Data Fig. 1
The colours and coordinates of UMAP.
Source Data Fig. 2
The proportions of cell types at each spot in ST (pie chart).
Source Data Fig. 3
Mean of normalized UMIs and fraction of cells with UMI >0 (dot plot).
Source Data Fig. 4
The value of each dot in Fig. 4f.
Source Data Fig. 5
The value of each dot in Fig. 5a.
Source Data Fig. 6
The meta-data of cells in Fig. 6.
Source Data Extended Data Fig./Table 1
The copy number ratio at each genomic location (e).
Source Data Extended Data Fig./Table 5
The co-clustering of ST in human and mouse.
Source Data Extended Data Fig./Table 8
The value of each dot in e.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, Y., Zhang, T., Zhou, Q. et al. A single-cell transcriptome atlas profiles early organogenesis in human embryos. Nat Cell Biol 25, 604–615 (2023). https://doi.org/10.1038/s41556-023-01108-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-023-01108-w
- Springer Nature Limited
This article is cited by
-
Single‑cell RNA sequencing analysis of human embryos from the late Carnegie to fetal development
Cell & Bioscience (2024)
-
Single-cell analysis reveals the spatial-temporal expression of genes associated with esophageal malformations
Scientific Reports (2024)
-
In vitro culture of cynomolgus monkey embryos from blastocyst to early organogenesis
Nature Protocols (2024)
-
Retinoic acid induces human gastruloids with posterior embryo-like structures
Nature Cell Biology (2024)
-
A patterned human neural tube model using microfluidic gradients
Nature (2024)