Abstract
Endometriosis is characterized by the growth of endometrial-like tissue outside the uterus. It affects many women during their reproductive age, causing years of pelvic pain and potential infertility. Its pathophysiology remains largely unknown, which limits early diagnosis and treatment. We characterized peritoneal and ovarian lesions at single-cell transcriptome resolution and compared them to matched eutopic endometrium, unaffected endometrium and organoids derived from these tissues, generating data on over 122,000 cells across 14 individuals. We spatially localized many of the cell types using imaging mass cytometry. We identify a perivascular mural cell specific to the peritoneal lesions, with dual roles in angiogenesis promotion and immune cell trafficking. We define an immunotolerant peritoneal niche, fundamental differences in eutopic endometrium and between lesion microenvironments and an unreported progenitor-like epithelial cell subpopulation. Altogether, this study provides a holistic view of the endometriosis microenvironment that represents a comprehensive cell atlas of the disease in individuals undergoing hormonal treatment, providing essential information for future therapeutics and diagnostics.







Similar content being viewed by others
Data availability
The RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE179640. To further safeguard the genomic identities of participants, single-nucleotide variants (SNVs) relative to the reference genome are masked in all bam files (BAMboozle v.0.5.0)102. Moreover, we have made the final single-cell datasets available for download and interactive exploration at https://singlecell.jax.org/datasets/endometriosis-2022. For mapping of scRNA-seq and bulk RNA-seq data, GRCh38.p13 (Ensembl Release 93, https://www.ncbi.nlm.nih.gov/assembly/GCF_000001405.27) was used. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
All code developed for and utilized in this study are available at https://github.com/TheJacksonLaboratory/endometriosis-scrnaseq, including modified CellPhoneDB scripts developed to optimize data interpretation for this study.
Change history
29 September 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41556-022-01023-6
References
Zondervan, K. T. et al. Endometriosis. Nat. Rev. Dis. Prim. 4, 9 (2018).
Saunders, P. T. K. & Horne, A. W. Endometriosis: etiology, pathobiology, and therapeutic prospects. Cell 184, 2807–2824 (2021).
Nirgianakis, K., Ma, L., McKinnon, B. & Mueller, M. D. Recurrence patterns after surgery in patients with different endometriosis subtypes: a long-term hospital-based cohort study. J. Clin. Med. 9, 496 (2020).
Symons, L. K. et al. The immunopathophysiology of endometriosis. Trends Mol. Med. 24, 748–762 (2018).
Wang, W. et al. Single-cell transcriptomic atlas of the human endometrium during the menstrual cycle. Nat. Med. 26, 1644–1653 (2020).
Garcia-Alonso, L. et al. Mapping the temporal and spatial dynamics of the human endometrium in vivo and in vitro. Nat. Genet. 53, 1698–1711 (2021).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
Lv, H. et al. Deciphering the endometrial niche of human thin endometrium at single-cell resolution. Proc. Natl Acad. Sci. USA 119, e2115912119 (2022).
He, S. et al. Single-cell transcriptome profiling of an adult human cell atlas of 15 major organs. Genome Biol. 21, 294 (2020).
Voigt, A. P. et al. Bulk and single-cell gene expression analyses reveal aging human choriocapillaris has pro-inflammatory phenotype. Microvasc. Res. 131, 104031 (2020).
Goveia, J. et al. An integrated gene expression landscape profiling approach to identify lung tumor endothelial cell heterogeneity and angiogenic candidates. Cancer Cell 37, 21–36.e13 (2020).
Masuda, H., Anwar, S. S., Bühring, H. J., Rao, J. R. & Gargett, C. E. A novel marker of human endometrial mesenchymal stem-like cells. Cell Transplant. 21, 2201–2214 (2012).
Edwards, A. K., Ramesh, S., Singh, V. & Tayade, C. A peptide inhibitor of synuclein-γ reduces neovascularization of human endometriotic lesions. Mol. Hum. Reprod. 20, 1002–1008 (2014).
Huang, H., Bhat, A., Woodnutt, G. & Lappe, R. Targeting the ANGPT–TIE2 pathway in malignancy. Nat. Rev. Cancer 10, 575–585 (2010).
Alpdogan, Ö. et al. Keratinocyte growth factor (KGF) is required for postnatal thymic regeneration. Blood 107, 2453–2460 (2006).
Teichert, M. et al. Pericyte-expressed Tie2 controls angiogenesis and vessel maturation. Nat. Commun. 8, 16106 (2017).
Carbone, C. et al. Angiopoietin-like proteins in angiogenesis, inflammation and cancer. Int. J. Mol. Sci. 19, 431 (2018).
Roca, C. & Adams, R. H. Regulation of vascular morphogenesis by Notch signaling. Genes Dev. 21, 2511–2524 (2007).
Pitulescu, M. E. et al. Dll4 and Notch signalling couples sprouting angiogenesis and artery formation. Nat. Cell Biol. 19, 915–927 (2017).
Mühleder, S., Fernández-Chacón, M., Garcia-Gonzalez, I. & Benedito, R. Endothelial sprouting, proliferation, or senescence: tipping the balance from physiology to pathology. Cell. Mol. Life Sci. 78, 1329–1354 (2020).
Wettschureck, N., Strilic, B. & Offermanns, S. Passing the vascular barrier: endothelial signaling processes controlling extravasation. Physiol. Rev. 99, 1467–1525 (2019).
Sharma, A. et al. Onco-fetal reprogramming of endothelial cells drives immunosuppressive macrophages in hepatocellular carcinoma. Cell 183, 377–394.e21 (2020).
Monzani, E., Bazzotti, R., Perego, C. & La Porta, C. A. M. AQP1 is not only a water channel: it contributes to cell migration through Lin7/β-catenin. PLoS ONE 4, e6167 (2009).
Chakarov, S. et al. Two distinct interstitial macrophage populations coexist across tissues in specific subtissular niches. Science 363, eaau0964 (2019).
Cheng, S. et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell 184, 792–809.e23 (2021).
Samaniego, R. et al. Folate receptor β (FRβ) expression in tissue-resident and tumor-associated macrophages associates with and depends on the expression of PU.1. Cells 9, 1445 (2020).
Kim, K.-W. et al. MHC II+ resident peritoneal and pleural macrophages rely on IRF4 for development from circulating monocytes. J. Exp. Med. 213, 1951–1959 (2016).
Gonzalez-Dominguez, E. et al. CD163L1 and CLEC5A discriminate subsets of human resident and inflammatory macrophages in vivo. J. Leukoc. Biol. 98, 453–466 (2015).
Hogg, C. et al. Macrophages inhibit and enhance endometriosis depending on their origin. Proc. Natl Acad. Sci. USA 118, e2013776118 (2021).
Lim, H. Y. et al. Hyaluronan receptor LYVE-1-expressing macrophages maintain arterial tone through hyaluronan-mediated regulation of smooth muscle cell collagen. Immunity 49, 326–341.e7 (2018).
Forster, R. et al. Macrophage‐derived insulin‐like growth factor‐1 is a key neurotrophic and nerve‐sensitizing factor in pain associated with endometriosis. FASEB J. 33, 11210–11222 (2019).
Lain, E. et al. A novel role for embigin to promote sprouting of motor nerve terminals at the neuromuscular junction. J. Biol. Chem. 284, 8930–8939 (2009).
Villar, J. & Segura, E. Decoding the heterogeneity of human dendritic cell subsets. Trends Immunol. 41, 1062–1071 (2020).
Maier, B. et al. A conserved dendritic-cell regulatory program limits antitumour immunity. Nature 580, 257–262 (2020).
Schulke, L. et al. Dendritic cell populations in the eutopic and ectopic endometrium of women with endometriosis. Hum. Reprod. 24, 1695–1703 (2009).
Hey-Cunningham, A. J. et al. Comprehensive analysis utilizing flow cytometry and immunohistochemistry reveals inflammatory changes in local endometrial and systemic dendritic cell populations in endometriosis. Hum. Reprod. 36, 415–428 (2021).
Cabeza-Cabrerizo, M. et al. Tissue clonality of dendritic cell subsets and emergency DCpoiesis revealed by multicolor fate mapping of DC progenitors. Sci. Immunol. 4, eaaw1941 (2019).
Karsunky, H., Merad, M., Cozzio, A., Weissman, I. L. & Manz, M. G. Flt3 ligand regulates dendritic cell development from Flt3+ lymphoid and myeloid-committed progenitors to Flt3+ dendritic cells in vivo. J. Exp. Med. 198, 305–313 (2003).
Merad, M., Ginhoux, F. & Collin, M. Origin, homeostasis and function of Langerhans cells and other langerin-expressing dendritic cells. Nat. Rev. Immunol. 8, 935–947 (2008).
Tewary, P. et al. Granulysin activates antigen-presenting cells through TLR4 and acts as an immune alarmin. Blood 116, 3465–3474 (2010).
Brown, C. C. et al. Transcriptional basis of mouse and human dendritic cell heterogeneity. Cell 179, 846–863.e24 (2019).
Durand, M. et al. Human lymphoid organ cDC2 and macrophages play complementary roles in T follicular helper responses. J. Exp. Med. 216, 1561–1581 (2019).
Yi, H. et al. Targeting the immunoregulator SRA/CD204 potentiates specific dendritic cell vaccine-induced T-cell response and antitumor immunity. Cancer Res. 71, 6611–6620 (2011).
Munawara, U. et al. Human dendritic cells express the complement receptor immunoglobulin which regulates T cell responses. Front. Immunol. 10, 2892 (2019).
Schmid, E. et al. Serum- and glucocorticoid-inducible kinase 1 sensitive NF-κB signaling in dendritic cells. Cell. Physiol. Biochem. 34, 943–954 (2014).
Halliday, N. et al. CD86 is a selective CD28 ligand supporting FoxP3+ regulatory T cell homeostasis in the presence of high levels of CTLA-4. Front. Immunol. 11, 3155 (2020).
Strunz, B. et al. Continuous human uterine NK cell differentiation in response to endometrial regeneration and pregnancy. Sci. Immunol. 6, eabb7800 (2021).
Aloisi, F. & Pujol-Borrell, R. Lymphoid neogenesis in chronic inflammatory diseases. Nat. Rev. Immunol. 6, 205–217 (2006).
Cabrita, R. et al. Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature 577, 561–565 (2020).
Ruffin, A. T. et al. B cell signatures and tertiary lymphoid structures contribute to outcome in head and neck squamous cell carcinoma. Nat. Commun. 12, 3349 (2021).
Hinrichs, B. H. et al. Serum amyloid A1 is an epithelial prorestitutive factor. Am. J. Pathol. 188, 937–949 (2018).
Taupin, D. & Podolsky, D. K. Trefoil factors: initiators of mucosal healing. Nat. Rev. Mol. Cell Biol. 4, 721–732 (2003).
Paulsen, F. P. et al. Intestinal Trefoil factor/TFF3 promotes re-epithelialization of corneal wounds. J. Biol. Chem. 283, 13418–13427 (2008).
Cocco, E. et al. Serum amyloid A (SAA): a novel biomarker for uterine serous papillary cancer. Br. J. Cancer 101, 335–341 (2009).
Badolato, R. et al. Serum amyloid a is a chemoattractant: induction migration, adhesion, and tissue infiltration of monocytes and polymorphonuclear leukocytes. J. Exp. Med. 180, 203–209 (1994).
Henze, D. et al. Endometriosis leads to an increased trefoil factor 3 concentration in the peritoneal cavity but does not alter systemic levels. Reprod. Sci. 24, 258–267 (2017).
Cindrova-Davies, T. et al. Menstrual flow as a non-invasive source of endometrial organoids. Commun. Biol. 4, 651 (2021).
Terakawa, J. et al. SIX1 cooperates with RUNX1 and SMAD4 in cell fate commitment of Müllerian duct epithelium. Cell Death Differ. 27, 3307–3320 (2020).
Maddern, J., Grundy, L., Castro, J. & Brierley, S. M. Pain in endometriosis. Front. Cell Neurosci. 14, 590823 (2020).
Izumi, G. et al. Mannose receptor is highly expressed by peritoneal dendritic cells in endometriosis. Fertil. Steril. 107, 167–173.e2 (2017).
Brech, D. et al. A mosaic renal myeloid subtype with T-cell inhibitory and protumoral features is linked to immune escape and survival in clear cell renal cell cancer. Preprint at bioRxiv https://doi.org/10.1101/2020.01.20.912865 (2020).
Gustafsson, C. et al. Gene expression profiling of human decidual macrophages: evidence for immunosuppressive phenotype. PLoS ONE 3, e2078 (2008).
Svensson, J. et al. Macrophages at the fetal–maternal interface express markers of alternative activation and are induced by M-CSF and IL-10. J. Immunol. 187, 3671–3682 (2011).
Ścieżyńska, Komorowski, Soszyńska & Malejczyk NK cells as potential targets for immunotherapy in endometriosis. J. Clin. Med. 8, 1468 (2019).
Nothnick, W. B. Treating endometriosis as an autoimmune disease. Fertil. Steril. 76, 223–231 (2001).
Malhotra, D. et al. Transcriptional profiling of stroma from inflamed and resting lymph nodes defines immunological hallmarks. Nat. Immunol. 13, 499–510 (2012).
Rodda, L. B. et al. Single-cell RNA sequencing of lymph node stromal cells reveals niche-associated heterogeneity. Immunity 48, 1014–1028.e6 (2018).
Csibi, N. et al. Gamma-synuclein levels are elevated in peritoneal fluid of patients with endometriosis. Med. Sci. Monit. 26, e922137 (2020).
Kang, T.-Y. et al. Pericytes enable effective angiogenesis in the presence of proinflammatory signals. Proc. Natl Acad. Sci. USA 116, 23551–23561 (2019).
Egorova, A. et al. Anti-angiogenic treatment of endometriosis via anti-VEGFA siRNA delivery by means of peptide-based carrier in a rat subcutaneous model. Gene Ther. 25, 548–555 (2018).
Becker, C. M. & D’Amato, R. J. Angiogenesis and antiangiogenic therapy in endometriosis. Microvasc. Res. 74, 121–130 (2007).
Liu, Z. et al. Single-cell transcriptomic analysis of eutopic endometrium and ectopic lesions of adenomyosis. Cell Biosci. 11, 51 (2021).
Ma, J. et al. Single-cell transcriptomic analysis of endometriosis provides insights into fibroblast fates and immune cell heterogeneity. Cell Biosci. 11, 125 (2021).
Zou, G. et al. Cell subtypes and immune dysfunction in peritoneal fluid of endometriosis revealed by single-cell RNA-sequencing. Cell Biosci. 11, 98 (2021).
Adam, M., Potter, A. S. & Potter, S. S. Psychrophilic proteases dramatically reduce single-cell RNA-seq artifacts: a molecular atlas of kidney development. Development 144, 3625–3632 (2017).
Tan, Y., Luo, D., Bozal, S., Robson, P. & Courtois, E. Human endometrium and endometriosis tissue dissociation for single-cell RNA sequencing. protocols.io https://doi.org/10.17504/protocols.io.bvy8n7zw (2022).
Boretto, M. et al. Patient-derived organoids from endometrial disease capture clinical heterogeneity and are amenable to drug screening. Nat. Cell Biol. 21, 1041–1051 (2019).
Turco, M. Y. et al. Long-term, hormone-responsive organoid cultures of human endometrium in a chemically defined medium. Nat. Cell Biol. 19, 568–577 (2017).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Giotti, B. et al. Assembly of a parts list of the human mitotic cell cycle machinery. J. Mol. Cell. Biol. 11, 703–718 (2019).
O’Flanagan, C. H. et al. Dissociation of solid tumor tissues with cold active protease for single-cell RNA-seq minimizes conserved collagenase-associated stress responses. Genome Biol. 20, 210 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Lun, A. T. L., Chen, Y. & Smyth, G. K. in Statistical Genomics. Methods in Molecular Biology Vol. 1418 (eds Mathé, E. & Davis, S.) 391–416 (Humana Press, 2016).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Bankhead, P. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Windhager, J., Bodenmiller, B. & Eling, N. An end-to-end workflow for multiplexed image processing and analysis. Preprint at bioRxiv https://doi.org/10.1101/2021.11.12.468357 (2021).
Schapiro, D. et al. HistoCAT: analysis of cell phenotypes and interactions in multiplex image cytometry data. Nat. Methods 14, 873–876 (2017).
Ziegenhain, C. & Sandberg, R. BAMboozle removes genetic variation from human sequence data for open data sharing. Nat. Commun. 12, 6216 (2021).
Acknowledgements
The authors would like to thank all participants for their tissue donations and valuable participation in this study. We thank the following Jackson Laboratory (JAX) Scientific Services cores, partially supported through the JAX Cancer Center Support Grant (CCSG) P30CA034196-30, for expert technical assistance: Single Cell Biology, Flow Cytometry and A. Carcio and T. Prosio, Genome Technologies and R. Maurya, Histology, and Microscopy. We also thank the JAX Cyberinfrastructure team for computational resources, L. Perpetua and the UConn Health Research Biorepository, and the UConn Health Surgery Center Personnel for assistance in collection of biopsy samples; the UCHC Pathology and Laboratory Medicine and M. Yu for assistance with histological examination of biopsies; the Clinical and Translational Research Support group, the Sponsored Research Administration service, and the Research Program Development service and A. L. Lucido for administrative assistance. All schematic panels were created using Biorender.com. This study was supported by the Assistance Secretary of Defense for Health Affairs endorsed by the Department of Defense, through the Peer Reviewed Medical Research Program under Award No. W81XWH-19-1-0130 (to E.T.C.), JAX Institutional startup funds (to P.R.) and UCHC/JAX Training Program in Genomic Science T32HG010463 (to M.D.). Opinions, interpretations, conclusions and recommendations are those of the authors and are not necessarily endorsed by the Department of Defense. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
E.T.C., P.R. and D.E.L. conceived and designed the study. Y.T. and E.T.C. performed and supervised the experiments. Y.T. and E.T.C. optimized the tissue-dissociation protocol. Y.T., D.L., S.B.B. and E.T.C. performed tissue dissociation, FACS and single-cell experiments. D.E.L. and A.A.L. collected consent from participants and clinical samples. S.S. performed IMC experiments. M.D performed organoid immunostaining. Y.T., W.F.F. and E.T.C. performed data analyses. Y.T., W.F.F. and E.T.C. wrote the manuscript and generated figures and schematics. Y.T., W.F.F., A.A.L., P.R., D.E.L. and E.T.C. contributed crucial data interpretation. All authors read or provided comments on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Junyue Cao, Chandrakant Tayade and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
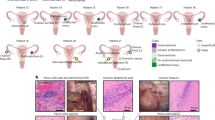
Extended Data Fig. 1 Overview of experiment design and comparison of bulk RNA-seq and scRNA-seq transcriptomic profiles from Ctrl and endometriosis tissues.
a, Experimental workflow. b, UMAP showing distribution of cell based on tissue types, PID, and endometriosis stage, before and after batch correction with Harmony. c, Box plot showing Spearman’s correlation rank (ρ) between bulkRNA-seq and pseudobulk from scRNA-seq in Eutopic (Ctrl & EuE, n = 144), Peritoneal (EcP & EcPA, n = 90), or Ovary (EcO, n = 24). Each dot represents a sample pair. The box represents the interquartile range with median and minimum/maximum represented by box centerline and whiskers, respectively. d, Scatterplot showing distribution of average gene expression between bulk RNA-seq and scRNA-seq (Spearman ρ). Each dot represents a gene. e, Volcano plots representing DEGs between scRNA-seq pseudo bulk (red) and bulk RNA-seq from undissociated tissue (blue) (edgeR, FDR < 0.001, LogFC > 3). The genes highlighted are exclusively expressed in bulk RNA-seq and associated with erythrocytes (orange), neuronal projections (green), adipocytes (brown), and muscle cells (purple). Related to Fig. 1.
Extended Data Fig. 2 Proportion of major cell types in each replicate and IMC panel for spatial profiling of Ctrl and endometriosis tissues.
a, Major cell types were determined based on UMAP. The mean distribution for all 5 major cell populations is represented for each tissue type Ctrl, EuE, EcP, EcPA, and EcO (left of the line). Each pie chart represents major cell type proportions for each replicate (right of the line). b, Each antibody was selected according to the cell types identified by the scRNA-seq data analysis. Representative images show single channels for each metal-conjugated antibody in a EuE biopsy. A total of 26 antibodies was used to identify cellular heterogeneity within stromal, endothelial, epithelial, lymphocyte, and myeloid major cell types. Additional antibodies (in ‘Others’) were used to identify cell proliferation (Ki67), active metabolism (pS6), extracellular matrix (Collagen1), and nuclei (DNA). A complete list of cell subpopulations identified through this panel of markers is listed on Supplementary Table 8b. Related to Fig. 1.
Extended Data Fig. 3 Stromal cell analysis across sample types.
a, Bar plot representing the proportion of stromal cell types in control endometrium and endometriosis lesions. Endometrial fibroblasts were found in all lesions. Fibroblast C7 is the predominant fibroblast type in EcO. b, Density plot showing distribution of mural cells for each tissue. Arrows points to Prv-CCL19. c, Heatmap of markers genes for mural cell subtypes. d, Track plot representing gene expression pattern for selected DEG in Prv-CCL19 subpopulations. GGT5 and ABCC9 are pan-markers for this cell subtype. e, Box plot showing the proportion of CCL19-expressing cells in Prv-CCL19 subpopulation within each tissue type. Each dot represents the percentage of CCL19+ cells in a tissue biopsy (Ctrl n = 3, EuE n = 9, EcP n = 8, EcPA n = 6, EcO n = 4). The box represents the interquartile range with median and minimum/maximum represented by box centerline and whiskers, respectively. Related to Fig. 2.
Extended Data Fig. 4 Characterization of endothelial cells (EC) across sample types.
a, Unique cell-to-cell interaction counts obtained from a modified CellPhoneDB procedure. To recover meaningful interactions, we analyzed ligand-receptor interaction in each sample independently. Unique interactions in each tissue type are counted as follows; each ligand-receptor pair observed in a specific cell type pair is counted as one interaction; this is tabulated for all possible pairwise cell type combinations (up to 58 subpopulations in this study) within a sample (n). The total count (Σ, n_celltype_pairs) represents the commonality of the ligand-receptor interaction of interest. The more common interactions (observed in multiple cell type pairs and in all individual samples) will have higher counts while restricted interactions (observed in specific cell type pairs) will have lower counts. We arbitrarily restricted our analysis to interactions observed fewer than 150 times to narrow the scope of analysis and focus on potentially uncovering unique cell-to-cell interactions. b, Box plot showing the proportion of DLL4-expressing cells in EC-tip subpopulation within each tissue type. c, Density plot showing distribution of endothelial cells for each tissue. d, EC proportions by sample type. e, AQP1+ cell abundance is substantially increased in peritoneal lesions (EcP and EcPA). f, (top) Proportion of aPCV among ECs across tissue types. (bottom) Swarm plot showing AQP1 expression per cell. Horizontal lines represent the median value. For box plots, each dot represents percentage of DLL4+ cells in EC-tip cluster (b) or AQP1+ cells in EC-aPCV cluster (e), in a tissue biopsy (Ctrl n = 3, EuE n = 9, EcP n = 8, EcPA n = 6, EcO n = 4). The box represents the interquartile range with median and minimum/maximum represented by box centerline and whiskers, respectively. Related to Fig. 3.
Extended Data Fig. 5 Myeloid cell diversity in control and endometriosis.
a, Heatmap representing marker genes for each myeloid subpopulation. b, Dendrogram showing the hierarchical clustering (Pearson correlation) for the myeloid cell clusters. c, Bar plot showing the representation of each myeloid subtype across tissue types. Related to Fig. 4.
Extended Data Fig. 6 DC subpopulations.
a, Bar plot represents the proportion of DCs among all myeloid cells for each patient (Ctrl n = 3, EuE n = 9, EcP n = 8, EcPA n = 6, EcO n = 4). Patient-to-patient variability was observed in DC proportions within the myeloid population and across different tissue types. The box represents the interquartile range with median and minimum/maximum represented by box centerline and whiskers, respectively. b, PAGA and RNA velocity trajectory analyses suggest that pre-cDC2 differentiate towards cDC2 and DC3 in Ctrl and EuE. Red arrows indicate that some cDC2 and DC3 cells derive from a smaller intermediate cell population. c, Cell cycle analysis for pre-cDC2, cDC2 and DC3 populations. d, Expression of DC progenitor markers FLT3, AXL, and SIGLEC6. e, Phagocytosis pathway is enriched in cDC2 subpopulations of peritoneal lesions. Bar plot shows the Normalized Enrichment Score (NES) for the top 10- GSEA pathways in cDC2 cells of EuE and EcPA (FDR < 0.1). Related to Fig. 5.
Extended Data Fig. 7 Lymphocyte subpopulations in control and endometriosis tissues.
a, Density plot showing distribution of lymphocyte cells for each tissue. b, Dot plot representing marker genes for each lymphocyte subpopulation, including four natural killer cell (NK) clusters, innate lymphoid cells (ILCs), effector memory T-cells (TEM), cytotoxic T-lymphocytes (CTL), naïve/central memory T-cells (TN/TCM), T regulatory cells (TReg), CD4- and CD8- tissue resident T cell (CD4-TRM and CD8-TRM, respectively), CD8 mucosal-associated invariant T cells (CD8-MAIT), plasma cells, and B cells. c, Representative IMC images showing the presence and proximity of myeloid cells labelled with CD68 (yellow) with T cells labelled with CD3 (cyan), and TReg labelled with FOXP3 (magenta) in EcO (n = 5); nuclei are marked with DNA intercalation (blue). Scale bar = 100 μm. d, Proportion bar plot of CTLA4 expressing cells from the total TReg subpopulation. e, Proportion box plot of BCL6, SEMA4A, CXCR5 expressing cells from the total B cells within each sample type. For box plots, each dot represents a unique patient (Ctrl n = 3, EuE n = 9, EcP n = 8, EcPA n = 6, EcO n = 4). The box represents the interquartile range with median and minimum/maximum represented by box centerline and whiskers, respectively. Related to Fig. 6.
Extended Data Fig. 8 Characterization of in vivo epithelial and in vitro endometrial epithelial organoid (EEO) cells.
a, Proportions of epithelial subpopulations per sample type. b, Representative IMC images of MUC5B+ epithelial cells in eutopic endometrium (Ctrl: C07, EuE: E12, E06) from multiple tissues. Epithelial cells are marked with PanCK, EpCAM, E-cadherin (green); MUC5B (magenta); nuclei (white). Scale bar = 100 μm. c, Proportion box plot of SAA1 expressing cells from the total MUC5B+ cells within each sample type. Each dot represents a unique patient (Ctrl n = 3, EuE n = 9, EcP n = 8, EcPA n = 6, EcO n = 4). The box represents the interquartile range with median and minimum/maximum represented by box centerline and whiskers, respectively. d, Sequencing metrics from EEO scRNA-seq; UMIs and unique genes counts are shown for Control (C) and endometriosis (E) patients and across tissue type. Undetermined (UD) group represents single cells which could not be assigned due to the lack of multiplexing hashtag but otherwise passed QC. e, Density plot showing distribution of EEO cells derived from Ctrl, EuE, EcP, and EcPA (UD cells were not included). f, UMAP showing the co-expression of MUC5B, SAA1, TFF3, and RUNX3 in the MUC5B+ population comprising in vivo epithelial cells and EEO. Related to Fig. 7.
Extended Data Fig. 9 MUC5B+ cells display a progenitor-like capacity in in vitro organoid culture.
a, Schematic and FACS sorting gating strategy to isolate MUC5B+ and MUC5B− epithelial cells from eutopic tissue for organoid generation. b, Representative brightfield images showing the progression of organoid generation from sorted single cells at day 2, 6 and 10. MUC5B+ cells formed EEO faster than MUC5B− cells. Each panel shows a whole Matrigel dome and magnified organoids are shown in the inset. Inset scale bar = 100 μm. c, Line graph showing area (top) and number (bottom) of EEO generated from MUC5B+ (dark blue, n = 1) and MUC5B− (sky blue, n = 1) cells over time. Area and Count of EEO is significantly higher in MUC5B+ compared to MUC5B− (paired t-test, two-tailed p < 0.0001). d, IF staining of EEO generated from MUC5B+(n = 1) and MUC5B- (n = 1) sorted cells showing the co-localization of endometrial epithelial (E-Cadherin, in green) and MUC5B+ (magenta) staining. Nuclei were counterstained with DAPI (gray). Scale bar = 100 µm. Related to Fig. 7.
Extended Data Fig. 10 Schematic illustrating the proposed microenvironment alterations for ectopic peritoneal and ovary lesions.
In peritoneal lesion (left), the proportion of myeloid and endothelial is increased, and endometrial-like epithelial population is reduced. CCL19-expressing perivascular cells mediate immune cell recruitment, such as macrophages and T cells, which contributes to the immunomodulatory microenvironment. We observe the presence of MSR1-expressing dendritic cells contributing to immunomodulation. TLS is also observed in some lesions. In addition, Mø1-LYVE1 and perivascular cells contribute to angiogenesis by regulating endothelial tip proliferation. In contrast, ovarian ectopic lesions (right) show a striking increase in the proportion of stromal cell and a reduced endometrial-like-epithelial cell presence. The immunomodulatory microenvironment is mainly driven by Mø1-LYVE1 expressing IL10. In ovary lesions, the regulation of angiogenesis is marked by endothelial cell arrest, resulting in mature vasculature. Created with Biorender.com.
Supplementary information
Supplementary Tables
Supplementary Table 1: Demographic and endometrium histology of participants. Related to Figs. 1 and 6, Extended Data Figs 1 and 2, and Supplementary Fig. 1. Supplementary Table 2: Experiment performed on each tissue. Oral contraceptive status is decoded as treated (Y) and non-treated (N). Supplementary Table 3: Marker genes expressed by 5 major cell types and 57 subpopulations found in this study. Significance was determined by two-sided Wilcoxon rank-sum test, P values were adjusting using Benjamini–Hochberg correction. The ‘in_group_fraction’ denotes the fraction of cells expressing the marker gene within the intended cluster, and ‘out_group_fraction’ denotes the fraction of cells expressing the marker gene outside the intended cluster. Related to Fig. 1d. Supplementary Table 4: DEGs between transcriptome obtained from scRNA-seq and bulk RNA-seq. Significance was determined with edgeR exact test (two-sided) and P values were adjusted using Benjamini–Hochberg correction (FDR < 0.001). Related to Extended Data Fig. 1e. Supplementary Table 5: Top 1,000 DEGs between tissue type within each subpopulation, in order from left to right: dS2, Prv-CCL19, EC tip, EC-aPCV, EC-PCV, Mɸ1-LYVE1, Mɸ4-infiltrated, cDC2, TReg, B cell. Significance was determined with edgeR quasi-likelihood F-test (two-sided), and P values were adjusted using Benjamini–Hochberg correction. Related to Figs. 2–5 and Extended Data Fig. 2. Correction (FDR < 0.01). A comprehensive list of the 46 cell subpopulation that this analysis was performed on is available upon request. Related to Figs. 2–6. Supplementary Table 6: List of 802 unique ligand–receptor interactions in each tissue type. A comprehensive list of ligand-–receptor interactions (including common interactions) is available on GitHub (Methods). Related to Fig. 3, Figs. 6 and 7, and Extended Data 4a. Supplementary Table 7: Enriched gene ontology in each biopsy type in comparison to control endometrium, in Mɸ1-LYVE1 (top) and cDC2 (bottom). Significance was determined with GSEA Fisher’s exact test (one-sided), and P values were adjusted using Benjamini–Hochberg correction (FDR < 0.1). ES: enrichment score; NES: normalized enrichment score. A comprehensive list of the 46 cell subpopulation that this analysis was performed on is available upon request. Related to Fig. 5 and Extended Data Fig. 6e. Supplementary Table 8: Additional information about cell hashtaging reagents used in this study. Related to Fig. 7 and Extended Data Fig. 8. Supplementary Table 9: Detailed information about antibodies used for IMC in this study. Related to Figs. 1–4, Figs. 6 and 7, and Extended Data Figs. 2, 7 and 8. Supplementary Table 10: Additional information about antibodies used for FACS in this study. Related to Extended Data Fig. 9.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tan, Y., Flynn, W.F., Sivajothi, S. et al. Single-cell analysis of endometriosis reveals a coordinated transcriptional programme driving immunotolerance and angiogenesis across eutopic and ectopic tissues. Nat Cell Biol 24, 1306–1318 (2022). https://doi.org/10.1038/s41556-022-00961-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-022-00961-5
- Springer Nature Limited
This article is cited by
-
Vitamin D and reproductive disorders: a comprehensive review with a focus on endometriosis
Reproductive Health (2024)
-
Unveiling heterogeneity and prognostic markers in ductal breast cancer through single-cell RNA-seq
Cancer Cell International (2024)
-
Endometriotic lesions exhibit distinct metabolic signature compared to paired eutopic endometrium at the single-cell level
Communications Biology (2024)
-
Immune micro-environment analysis and drug screening for ovarian endometriosis
Genes & Genomics (2024)
-
An integrated single-cell reference atlas of the human endometrium
Nature Genetics (2024)