Abstract
Tissue turnover requires activation and lineage commitment of tissue-resident stem cells (SCs). These processes are impacted by ageing, but the mechanisms remain unclear. Here, we addressed the mechanisms of ageing in murine hair follicle SCs (HFSCs) and observed a widespread reduction in chromatin accessibility in aged HFSCs, particularly at key self-renewal and differentiation genes, characterized by bivalent promoters occupied by active and repressive chromatin marks. Consistent with this, aged HFSCs showed reduced ability to activate bivalent genes for efficient self-renewal and differentiation. These defects were niche dependent as the transplantation of aged HFSCs into young recipients or synthetic niches restored SC functions. Mechanistically, the aged HFSC niche displayed widespread alterations in extracellular matrix composition and mechanics, resulting in mechanical stress and concomitant transcriptional repression to silence promoters. As a consequence, increasing basement membrane stiffness recapitulated age-related SC changes. These data identify niche mechanics as a central regulator of chromatin state, which, when altered, leads to age-dependent SC exhaustion.






Similar content being viewed by others
Data availability
Sequencing data that support the findings of this study have been deposited at the Gene Expression Omnibus (GEO) under the accession code GSE148619. Proteomics data have been deposited to the ProteomeXchange Consortium through the PRIDE partner repository51 under the dataset identifier PXD018352. Previously published sequencing data14 that were reanalysed here are available under the accession code GSE31239. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Lopez-Otin, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Signer, R. A. & Morrison, S. J. Mechanisms that regulate stem cell aging and life span. Cell Stem Cell 12, 152–165 (2013).
Voigt, P., Tee, W. W. & Reinberg, D. A double take on bivalent promoters. Genes Dev. 27, 1318–1338 (2013).
Harikumar, A. & Meshorer, E. Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep. 16, 1609–1619 (2015).
Hsu, Y. C., Li, L. & Fuchs, E. Emerging interactions between skin stem cells and their niches. Nat. Med. 20, 847–856 (2014).
Doles, J., Storer, M., Cozzuto, L., Roma, G. & Keyes, W. M. Age-associated inflammation inhibits epidermal stem cell function. Genes Dev. 26, 2144–2153 (2012).
Giangreco, A., Qin, M., Pintar, J. E. & Watt, F. M. Epidermal stem cells are retained in vivo throughout skin aging. Aging Cell 7, 250–259 (2008).
Keyes, B. E. et al. Nfatc1 orchestrates aging in hair follicle stem cells. Proc. Natl Acad. Sci. USA 110, E4950-9 (2013).
Matsumura, H. et al. Hair follicle aging is driven by transepidermal elimination of stem cells via COL17A1 proteolysis. Science 351, aad4395 (2016).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Saha, P. & Mishra, R. K. Heterochromatic hues of transcription—the diverse roles of noncoding transcripts from constitutive heterochromatin. FEBS J. 286, 4626–4641 (2019).
Brunet, A. & Rando, T. A. Interaction between epigenetic and metabolism in aging stem cells. Curr. Opin. Cell Biol. 45, 1–7 (2017).
Ge, Y. et al. The aging skin microenvironment dictates stem cell behavior. Proc. Natl Acad. Sci. USA 117, 5339–5350 (2020).
Lien, W. H. et al. Genome-wide maps of histone modifications unwind in vivo chromatin states of the hair follicle lineage. Cell Stem Cell 9, 219–232 (2011).
Ito, M., Kizawa, K., Hamada, K. & Cotsarelis, G. Hair follicle stem cells in the lower bulge form the secondary germ, a biochemically distinct but functionally equivalent progenitor cell population, at the termination of catagen. Differ. Res. Biol. diversity 72, 548–557 (2004).
Muller-Rover, S. et al. A comprehensive guide for the accurate classification of murine hair follicles in distinct hair cycle stages. J. Invest. Dermatol. 117, 3–15 (2001).
Chacon-Martinez, C. A., Klose, M., Niemann, C., Glauche, I. & Wickstrom, S. A. Hair follicle stem cell cultures reveal self-organizing plasticity of stem cells and their progeny. EMBO J. 36, 151–164 (2017).
Hsu, Y. C., Li, L. & Fuchs, E. Transit-amplifying cells orchestrate stem cell activity and tissue regeneration. Cell 157, 935–949 (2014).
Ansorge, H. L. et al. Type XIV collagen regulates fibrillogenesis: premature collagen fibril growth and tissue dysfunction in null mice. J. Biol. Chem. 284, 8427–8438 (2009).
Bradshaw, A. D. et al. SPARC-null mice display abnormalities in the dermis characterized by decreased collagen fibril diameter and reduced tensile strength. J. Invest. Dermatol. 120, 949–955 (2003).
Guimaraes, C. F., Gasperini, L., Marques, A. P. & Reis, R. L. The stiffness of living tissues and its implications for tissue engineering. Nat. Rev. Mater. 5, 351–370 (2020).
Langton, A. K., Graham, H. K., Griffiths, C. E. M. & Watson, R. E. B. Ageing significantly impacts the biomechanical function and structural composition of skin. Exp. Dermatol. 28, 981–984 (2019).
Mao, J. R. et al. Tenascin-X deficiency mimics Ehlers-Danlos syndrome in mice through alteration of collagen deposition. Nat. Genet. 30, 421–425 (2002).
Murrell, M., Oakes, P. W., Lenz, M. & Gardel, M. L. Forcing cells into shape: the mechanics of actomyosin contractility. Nat. Rev. Mol. Cell Biol. 16, 486–498 (2015).
Le, H. Q. et al. Mechanical regulation of transcription controls Polycomb-mediated gene silencing during lineage commitment. Nat. Cell Biol. 18, 864–875 (2016).
Vinogradova, M. et al. An inhibitor of KDM5 demethylases reduces survival of drug-tolerant cancer cells. Nat. Chem. Biol. 12, 531–538 (2016).
Conboy, I. M. et al. Rejuvenation of aged progenitor cells by exposure to a young systemic environment. Nature 433, 760–764 (2005).
Nishiguchi, M. A., Spencer, C. A., Leung, D. H. & Leung, T. H. Aging suppresses skin-derived circulating SDF1 to promote full-thickness tissue regeneration. Cell Rep. 24, 3383–3392 (2018).
Sarbacher, C. A. & Halper, J. T. Connective tissue and age-related diseases. Subcell. Biochem. 91, 281–310 (2019).
Segel, M. et al. Niche stiffness underlies the ageing of central nervous system progenitor cells. Nature 573, 130–134 (2019).
Marinkovich, M. P., Lunstrum, G. P. & Burgeson, R. E. The anchoring filament protein kalinin is synthesized and secreted as a high molecular weight precursor. J. Biol. Chem. 267, 17900–17906 (1992).
Sorokin, L. M. et al. Laminin α4 and integrin α6 are upregulated in regenerating dy/dy skeletal muscle: comparative expression of laminin and integrin isoforms in muscles regenerating after crush injury. Exp. Cell Res. 256, 500–514 (2000).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Liang, Y. et al. A cell-instructive hydrogel to regulate malignancy of 3D tumor spheroids with matrix rigidity. Biomaterials 32, 9308–9315 (2011).
Dodt, M., Roehr, J. T., Ahmed, R. & Dieterich, C. FLEXBAR-flexible barcode and adapter processing for next-generation sequencing platforms. Biology 1, 895–905 (2012).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Allhoff, M., Sere, K., J, F. P., Zenke, M. & I, G. C. Differential peak calling of ChIP-seq signals with replicates with THOR. Nucleic Acids Res. 44, e153 (2016).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank A. M. Luoto and N. Hachenberg for expert technical assistance; A. Gyenis for help with experiments; M. Loparic and D. Schneider for initial atomic force microscopy measurements; L. Sorokin for laminin antibodies; the staff at the FACS & Imaging Facility of MPI for Biology of Ageing, the Imaging Facility of CECAD Cologne and the Biomedicum Helsinki Imaging Unit for imaging support; and C. Becker and E. Kirstat at the Cologne Center for Genomics for help with sequencing. This work was supported by the Sigrid Juselius Foundation, Helsinki Institute of Life Science, Wihuri Research Institute, Max Planck Society, the Max Planck Förderstiftung and the European Research Council (ERC) under the EU Horizon 2020 research and innovation programme (grant agreement 770877—STEMpop) (all to S.A.W.) and the Deutsche Forschungsgemeinschaft (DFG; project number 73111208—SFB 829 (to S.A.W., C.M.N. and A.R.-I.), and FOR2722 to M.K. Y.A.M. is the recipient of the EMBO Long-Term fellowship ALTF 728-2017 and Human Frontier Science Program fellowship LT000861/2018.
Author information
Authors and Affiliations
Contributions
J.K. designed and performed most of the experiments and analysed data. Y.A.M. performed atomic force microscopy, immunostainings, organoid experiments, designed experiments and analysed data. S.G. performed the HFSC quantification and BrdU feeding experiments. C.A.C.-M. performed cell sorting and depilation experiments with J.K. J.M. performed immunostainings. X.L. and I.A. performed proteomics experiments. W.B. performed electron microscopy. J.A. performed ChIP sequencing. D.E.B. and M.K. provided collagen XIV and Tnx mutant mice and other key reagents. M.B. and A.R.-I. supported the ATAC-seq experiments and analysis of sequencing data. C.M.N. and A.R.-I. designed experiments and analysed data. S.A.W. conceived and supervised the study, designed experiments, analysed data and wrote the paper. All of the authors commented and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Karl Lenhard Rudolph and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Analyses of young and aged HFSCs using flow cytometry, ATACseq and RNAseq.
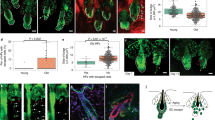
a, Representative images of young (6 month) and aged (24 month) old mice show no visible differences in haircoats. (b,c) Immunofluorescence imaging (b) and quantification (c) of hair follicle density from young and old mice (mean ±SD; n = 5 mice/group). Scale bars 100 µm. d, Quantification of CD34 + /α6 integrin+ HFSCs by flow cytometry from young and aged mice cohorts from two different animal facilities (facilities #2 and #3, facility #1 shown in main Fig. 1b), where the respective young and aged mice were housed in the same room. Note decreased HFSC levels in mice from both facilities (mean, n = 7 mice/group; *p = 0.0293 Student’s t-test (facility #2), n = 4 mice/group; *p = 0.0283, Mann-Whitney (facility #3)). e, Plotting relative abundance of CD34 + HFSCs as a function of anagen-to-telogen ratio shows no correlation between the two parameters in young or aged mice (n = 15 young/14 aged mice; R2 = 0.0001168, Pearson’s correlation). f, Fragment frequency distribution of the ATACseq reads shows expected distribution both in young and aged biological replicates (n = 4 mice/group). g, Motif enrichment analyses of differentially accessible ATACseq peaks shows enrichment of CpG islands and promoter regions in regions more accessible in young HFSCs, whereas more accessible peaks in aged HFSCs are enriched in transposable elements. (-log(p) values: CpG islands – yHFSCs: 670.48/ aHFSCs: 2.03; promoters - yHFSCs: 289.13, aHFSCs: 1.22; 5’ utr - yHFSCs: 203.36, aHFSCs: 2.69; IAPEz-int|LTR | ERVK - young HFSCs: -40.18, aged HFSCs: 83.66; LTR | ERVK - young HFSCs: -151.31, aged HFSCs: 12.56) (h) Volcano plot of significantly altered transcripts in young and aged HFSCs. Genes involved in differentiation (eg. Mef2c, Gata6, Sox21, Tcf24), proliferation and signaling (Ptch1, Fgfr2, Tgfa, Nptx1), actin cytoskeleton (Espn, Fhod3, Smoc2), and stemness (Peg3) are among the most regulated (n = 3 mice/group; padj<0.05, Wald test/Benjamini-Hochberg). (i) GO-term analyses of genes up- and down-regulated in aged HFSCs.
Extended Data Fig. 2 Histone methylation analyses of young and aged HFSCs.
a, Heatmap of differentially accessible intergenic regions clustered according to H3K4me1, H3K27ac, H3K27me3 as well as the transcripts of these regions quantified by RNAseq from young and aged HFSCs. Note that most regions have high H3K4me1, but low H3K27ac and no detectible transcripts, indicative of a primed enhancer state. b, GO term analyses of the regions in (a) implicate enhancers for genes involved processes such as cell-cell adhesion, tissue development, cell differentiation, and proliferation to be less accessible (upper panel), whereas enhancers for genes involved in actin cytoskeleton organization and regulation of cell adhesion are more accessible (lower panel) in aged HFSCs. c, Heatmap of H3K27me3 ChIP sequencing peaks from young (H3K27me3-Y) and aged (H3K27me3-A) HFSCs shows slightly increased peak intensity in aged HFSCs (n = 2 mice/group). d, Box plot of -log2 fold changes of H3K4me3 occupancy at promoters of aged HFSC compared to young (top, middle and bottom delimiters are the 75th, 50th, 25th percentiles; top and bottom whiskers are the 90th and 10th percentiles; n = 2 mice/group). e, Representative genes from cluster-4 with functions in stem cell (SC) fate, differentiation and self-renewal include key genes required for HFSC activation and differentiation.
Extended Data Fig. 3 Niche-dependent regulation of HFSC potency and epigenetic state.
a, Representative images and quantification of BrdU incorporation in bulge (Bu) HFSCs from mice fed with BrdU in drinking water for 4 weeks. Note reduced BrdU incorporation in aged HFSCs (n = 3 mice/group). Scale bars 25 µm. b, Quantitative RT-PCR of RNAi knockdowns for selected genes with bivalent promoter state (n = 3 independent experiments). (c-e) Representative images (c) and quantification of colony number (d) and size (e) from colony forming assays with HFSCs depleted of the indicated genes (n = 3 independent experiments; *p = 0.0263, RM-ANOVA/Fischer’s, **p = 0.0045, Friedman/Dunn’s). f, Representative images and quantification of H3K4me3 in hair follicles generated in young nude mice by transplanting young or aged HFSCs (n = 3 mice/group). Scale bars 25 µm. g, Quantitative RT-PCR of selected genes with bivalent promoter state and reduced accessibility in vivo from H3K4me3 ChIP of young and aged HFSCs in 24 h 3 C organoid cultures. No significant differences in expression are observed (n = 4 mice/group). h, Quantitative RT-PCR of selected genes with bivalent promoter state and reduced accessibility in vivo from young and aged HFSCs in 3 C organoid cultures. No significant differences in expression are observed (n = 4 mice/group). Bar graphs in g, h show mean ±SEM, others mean ±SD.
Extended Data Fig. 4 Basement membranes in young and aged skin and decellularized scaffolds.
a, Representative immunofluorescence images and quantification of Laminin 332 staining show increased levels in aged BMs (mean; n = 4 mice/group, *p = 0.0286, Mann-Whitney; Scale bars 50 µm). b, Representative immunofluorescence images and quantification of Laminin 511 staining show increased levels in aged BMs (mean; n = 4 mice/group, *p = 0.0286, Mann-Whitney; Scale bars 50 µm). c, Schematic illustration of decellularized basement membrane scaffold preparation from young and aged mice. d, Immunofluorescence images of decellularized basement membrane scaffolds showing absence of epidermal cell nuclei but intact basement membranes as marked by Collagen IV and Laminin 332 staining. A representative of 3 mice is shown. Scale bars 100 µm. e, Box and whiskers plot of atomic force microscopy force indentation measurements of 3 C organoid hydrogel stiffness (top, middle and bottom delimiters are the 75th, 50th, 25th percentiles; top and bottom whiskers are the maximum and minimum; n = 13 force curves pooled across 4 independent experiments; **p = 0.0012, Kolmogorov-Smirnov).
Extended Data Fig. 5 Analyses of Collagen-XIV- and Tenascin-X-deficient mice.
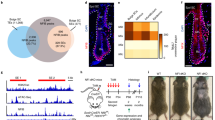
a, Hematoxylin/Eosin stainings from 1 year old Collagen-XIV-deficient (Col XIV -/-) and wild type (ColXIV + /+) control mice reveal no overt skin pathology. Scale bars 100 µm. Representative images from 3 mice/genotype are shown. b, Immunofluorescence images of Cd34 staining to mark the bulge (bu) niche from 1 year old Col XIV + / + and -/- mice show comparable hair follicle morphology. Representative images from 3 mice/genotype are shown. Scale bars 25 µm. c, Quantification of CD34 + /α6 integrin+ HFSCs by flow cytometry from 6 months old Col XIV + / + and -/- mice shows no major reduction in levels of HFSCs (n = 5 Col XIV + / + and n = 4 Col XIV -/- mice). d, Quantification of basement membrane (BM) thickness from skin electron micrographs of Tenascin-X-deficient (TnX -/-) and wild type (TnX + /+) control mice. Note increased BM thickness in TnX -/- mice (n = 4 mice/group; *p = 0.0286, Mann-Whitney). e, mHematoxylin/Eosin stainings from 1 year old TnX + /+ and TnX -/- mice show no overt skin pathology. Representative images from 3 mice/genotype are shown. Scale bars 100 µm. f, Immunofluorescence images of Keratin-15 staining to mark the bulge (bu) niche from 1 year old TnX + /+ and -/- mice show comparable hair follicle morphology. Representative images from 3 mice/genotype are shown. Scale bars 25 µm. g, Quantification of CD34 + /α6 integrin+ HFSCs by flow cytometry from 6 months old TnX + /+ and -/- mice shows slightly reduced levels of HFSCs in TnX -/- mice (n = 7 TnX + /+ and n = 8 TnX -/- mice; *p = 0.0301, Student’s t-test). Bar graphs show mean ±SD.
Extended Data Fig. 6 Analyses of contractility and transcription in hair follicles and organoid cultures.
a, Representative immunofluorescence images and quantification of phosphorylated myosin light chain 2 (pMLC2). Note increased pMLC2 indicating increased contractility in aged bulge (bu) HFSCs (n = 5 young and n = 4 aged mice; *p = 0.0451, Student’s t-test; Scale bar 50 µm). b, Quantification of the nuclear aspect ratio (major axis/minor axis) shows increased nuclear elongation of aged HFSCs (n = 5 mice/group with >100 nuclei per mouse; **p = 0.0031, Student’s t-test). c, Representative images and quantification of H3K4me3 intensity from HFSC organoids treated with Kdm5 inhibitor (Kdm5i) (n = 6 independent experiments; p = 0.0515, Student’s t-test; Scale bar 50 µm). d, Quantitative RT-PCR of selected genes with bivalent promoter state from HFSC organoids treated with Kdm5i (n = 4 independent experiments). Bar graphs show mean ±SD.
Supplementary information
Supplementary Information
Supplementary Fig. 1: the gating strategy for quantification and purification of integrin α6+CD34+ HFSCs by FACS.
Supplementary Tables 1–7
Supplementary Table 1: differential peak analysis of ATAC-seq data. Analysis of differentially accessible genomic regions in young and aged HFSCs using ATAC-seq. Supplementary Table 2: differential gene expression analysis of RNA-seq data. Analysis of differentially expressed genes in young and aged HFSCs using RNA-seq. Supplementary Table 3: differential peak analysis of H3K4me3 ChIP–seq data. Analysis of differential H3K4me3 peaks in young and aged HFSCs. Supplementary Table 4: differential peak analysis of H3K27me3 ChIP–seq data. Analysis of differential H3K27me3 peaks in young and aged HFSCs. Supplementary Table 5: differential protein abundance analysis of proteomics data. Analyses of differences in proteome in young and aged skin. Supplementary Table 6: differential peak analysis of RNAPII ChIP–seq data. Analysis of differential RNAPII peaks in young and aged HFSCs. Supplementary Table 7: primers used in this study. A list of primers used for ChIP PCR and PCR in this study.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
About this article
Cite this article
Koester, J., Miroshnikova, Y.A., Ghatak, S. et al. Niche stiffening compromises hair follicle stem cell potential during ageing by reducing bivalent promoter accessibility. Nat Cell Biol 23, 771–781 (2021). https://doi.org/10.1038/s41556-021-00705-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00705-x
- Springer Nature Limited
This article is cited by
-
Biomaterial-based mechanical regulation facilitates scarless wound healing with functional skin appendage regeneration
Military Medical Research (2024)
-
Epidermal stem cells: skin surveillance and clinical perspective
Journal of Translational Medicine (2024)
-
Profiling native pulmonary basement membrane stiffness using atomic force microscopy
Nature Protocols (2024)
-
Mechanical state transitions in the regulation of tissue form and function
Nature Reviews Molecular Cell Biology (2024)
-
Deciphering the molecular mechanisms of stem cell dynamics in hair follicle regeneration
Experimental & Molecular Medicine (2024)