Abstract
In the past two decades, emerging studies have suggested that DExD/H box helicases belonging to helicase superfamily 2 (SF2) play essential roles in antiviral innate immunity. However, the antiviral functions of helicase SF1, which shares a conserved helicase core with SF2, are little understood. Here we demonstrate that zinc finger NFX1-type containing 1 (ZNFX1), a helicase SF1, is an interferon (IFN)-stimulated, mitochondrial-localised dsRNA sensor that specifically restricts the replication of RNA viruses. Upon virus infection, ZNFX1 immediately recognizes viral RNA through its Armadillo-type fold and P-loop domain and then interacts with mitochondrial antiviral signalling protein to initiate the type I IFN response without depending on retinoic acid-inducible gene I-like receptors (RLRs). In short, as is the case with interferon-stimulated genes (ISGs) alone, ZNFX1 can induce IFN and ISG expression at an early stage of RNA virus infection to form a positively regulated loop of the well-known RLR signalling. This provides another layer of understanding of the complexity of antiviral immunity.







Similar content being viewed by others
Data availability
RNA sequencing data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE132979. Source data for Figs. 1–7 and Extended Data Figs. 1–7 have been provided as Statistics Source Data. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
References
Ablasser, A. et al. cGAS produces a 2′–5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013).
Wu, J. et al. Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 5, 730–737 (2004).
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Goubau, D. et al. Antiviral immunity via RIG-I-mediated recognition of RNA bearing 5′-diphosphates. Nature 514, 372–375 (2014).
Ahmad, S. & Hur, S. Helicases in antiviral immunity: dual properties as sensors and effectors. Trends Biochem. Sci. 40, 576–585 (2015).
Oshiumi, H. et al. DDX60 is involved in RIG-I-dependent and independent antiviral responses, and its function is attenuated by virus-induced EGFR activation. Cell Rep. 11, 1193–1207 (2015).
Zhang, Z. et al. DDX1, DDX21 and DHX36 helicases form a complex with the adaptor molecule TRIF to sense dsRNA in dendritic cells. Immunity 34, 866–878 (2011).
Zhang, Z., Yuan, B., Lu, N., Facchinetti, V. & Liu, Y. J. DHX9 pairs with IPS-1 to sense double-stranded RNA in myeloid dendritic cells. J. Immunol. 187, 4501–4508 (2011).
Wang, P. H. et al. Nlrp6 regulates intestinal antiviral innate immunity. Science 350, 826–830 (2015).
Singleton, M. R., Dillingham, M. S. & Wigley, D. B. Structure and mechanism of helicases and nucleic acid translocases. Annu. Rev. Biochem. 76, 23–50 (2007).
Fairman-Williams, M. E., Guenther, U. P. & Jankowsky, E. SF1 and SF2 helicases: family matters. Curr. Opin. Struct. Biol. 20, 313–324 (2010).
Linder, P. & Jankowsky, E. From unwinding to clamping—the DEAD box RNA helicase family. Nat. Rev. Mol. Cell. Bio. 12, 505–516 (2011).
Seth, R. B., Sun, L., Ea, C. K. & Chen, Z. J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-κB and IRF 3. Cell 122, 669–682 (2005).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Chen, Q., Sun, L. J. & Chen, Z. J. Regulation and function of the cGAS-STING pathway of cytosolic DNA sensing. Nat. Immunol. 17, 1142–1149 (2016).
Munoz-Wolf, N. & Lavelle, E. C. Hippo interferes with antiviral defences. Nat. Cell Biol. 19, 267–269 (2017).
Zhang, Q. et al. Hippo signalling governs cytosolic nucleic acid sensing through YAP/TAZ-mediated TBK1 blockade. Nat. Cell Biol. 19, 362–374 (2017).
O’Shea, J. J. & Plenge, R. JAK and STAT signaling molecules in immunoregulation and immune-mediated disease. Immunity 36, 542–550 (2012).
Schoggins, J. W. et al. Pan-viral specificity of IFN-induced genes reveals new roles for cGAS in innate immunity. Nature 505, 691–695 (2014).
Chan, Y. K. & Gack, M. U. Viral evasion of intracellular DNA and RNA sensing. Nat. Rev. Microbiol. 14, 360–373 (2016).
Schneider, W. M., Chevillotte, M. D. & Rice, C. M. Interferon-stimulated genes: a complex web of host defenses. Annu. Rev. Immunol. 32, 513–545 (2014).
Schoggins, J. W. et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–485 (2011).
Pillai, P. S. et al. Mx1 reveals innate pathways to antiviral resistance and lethal influenza disease. Science 352, 463–466 (2016).
Bailey, C. C. et al. IFITM-family proteins: the cell’s first line of antiviral defense. Annu. Rev. Virol. 1, 261–283 (2014).
Munir, M. & Berg, M. The multiple faces of proteinkinase R in antiviral defense. Virulence 4, 85–89 (2013).
Neil, S. J., Zeng, T. & Bieniasz, P. D. Tetherin inhibits retrovirus release and is antagonized by HIV-1 Vpu. Nature 451, 425–430 (2008).
Yan, N. & Chen, Z. J. Intrinsic antiviral immunity. Nat. Immunol. 13, 214–222 (2012).
Yoh, S. M. et al. PQBP1 is a proximal sensor of the cGAS-dependent innate response to HIV-1. Cell 161, 1293–1305 (2015).
Song, G. et al. E3 ubiquitin ligase RNF128 promotes innate antiviral immunity through K63-linked ubiquitination of TBK1. Nat. Immunol. 17, 1342–1351 (2016).
Zhu, J. et al. Antiviral activity of human OASL protein is mediated by enhancing signaling of the RIG-I RNA sensor. Immunity 40, 936–948 (2014).
Jia, X. et al. The role of alternative polyadenylation in the antiviral innate immune response. Nat. Commun. 8, 14605 (2017).
Goodier, J. L., Cheung, L. E. & Kazazian, H. H. Jr. MOV10 RNA helicase is a potent inhibitor of retrotransposition in cells. PLoS Genet. 8, e1002941 (2012).
Serquina, A. K. et al. UPF1 is crucial for the infectivity of human immunodeficiency virus type 1 progeny virions. J. Virol. 87, 8853–8861 (2013).
Miller, M. S. et al. Senataxin suppresses the antiviral transcriptional response and controls viral biogenesis. Nat. Immunol. 16, 485–494 (2015).
López, D. G. et al. The Eutherian Armcx genes regulate mitochondrial trafficking in neurons and interact with Miro and Trak2. Nat. Commun. 3, 814 (2012).
Mukherjee, P. et al. Activation of the innate signaling molecule MAVS by bunyavirus infection upregulates the adaptor protein SARM1, leading to neuronal death. Immunity 38, 705–716 (2013).
West, A. P., Shadel, G. S. & Ghosh, S. Mitochondria in innate immune responses. Nat. Rev. Immunol. 11, 389–402 (2011).
Yoshizumi, T. et al. Influenza A virus protein PB1-F2 translocates into mitochondria via Tom40 channels and impairs innate immunity. Nat. Commun. 5, 4713 (2014).
Loo, Y. M. et al. Distinct RIG-I and MDA5 signaling by RNA viruses in innate immunity. J. Virol. 82, 335–345 (2008).
Lu, H. B. et al. DHX15 senses double-stranded RNA in myeloid dendritic cells. J. Immunol. 193, 1364–1372 (2014).
Lian, H. et al. The zinc-finger protein ZCCHC3 binds RNA and facilitates viral RNA sensing and activation of the RIG-I-like receptors. Immunity 18, 49 (2018).
Serrat, R. et al. The non-canonical Wnt/PKC pathway regulates mitochondrial dynamics through degradation of the arm-like domain-containing protein Alex3. PLoS One 8, e67773 (2013).
Panneerselvam, P. et al. T-cell death following immune activation is mediated by mitochondria-localized SARM. Cell Death Differ. 20, 478–489 (2013).
Yuan, S. et al. Amphioxus SARM involved in neural development may function as a suppressor of TLR signaling. J. Immunol. 184, 6874–6881 (2010).
Hou, F. et al. MAVS forms functional prion-like aggregates to activate and propagate antiviral innate immune response. Cell 146, 448–461 (2011).
Gack, M. U. et al. TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature 446, 916–920 (2007).
Jiang, X. et al. Ubiquitin-induced oligomerization of the RNA sensors RIG-I and MDA5 activates antiviral innate immune response. Immunity 36, 959–973 (2012).
Ishidate, T. et al. ZNFX-1 functions within perinuclear nuage to balance epigenetic signals. Mol. Cell 70, 639–649 (2018).
Wan, G. et al. Spatiotemporal regulation of liquid-like condensates in epigenetic inheritance. Nature 557, 679–683 (2018).
Acknowledgements
This work was supported by the National Natural Science Foundation (NNSF) of China under grants 31770943, 81430099 and 31900661 and by the Natural Science Foundation of Guangdong Province of China under grants 2015A030306043, 2018A030313924 and 2018A030313051.
Author information
Authors and Affiliations
Contributions
Y.W., S.Y. and A.X. conceived the ideas and designed the experiments. Y.W., X.J., Y.G., T.L., M.N. and X.L. performed the experiments. Y.W., S.Y. and X.J. analysed the data. S.Y., Y.W., X.J. and S.C. contributed to editing the manuscript. S.Y. and A.X. supervised the research and wrote the paper. S.Y. and X.J. are joint co first authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Bioinformatics analysis of in vitro transcription-sequencing APA sites (IVT-SAPAS) data of VSV-infected macrophages and previously collected viral infection microarray data.
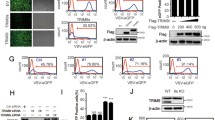
a, IVT-SAPAS revealed the genes with transcriptional changes in MDMs infected with VSV for 0, 2, 4, 8, 16, 24 hrs. b-c, The percentage of GFP+ cells of FACS analysis of A549 cells transfected with control siRNA or the indicated siRNAs followed by VSV-eGFP infection for another 6 hrs (n = 3 independent experiments). d, qRT-PCR revealed the mRNA expression of target gene in A549 cells transfected with indicated siRNAs for 48 hrs (n = 3 independent experiments). e-f, RNA levels of Znfx1 and Rig-I are significantly increased after different virial infection in different cell types. g, Schematic representation of ZNFX1 promoter containing core region bound by STAT1, STAT2, IRF1 and IRF9. For e, n = 3 independent experiments. Data in f, n = 4 wells for SeV infected epithelial cells and n = 6 wells for uninfected cells; n = 5 samples for IVA infected pDCs; n = 4 independent experiments for SeV infected monocytoid cells; n = 2 independent experiments for SeV or HIV infected Macrophages or mDCs. All data are shown as the mean ± s.d. Statistical differences were detected using two-tailed unpaired Student’s t-tests.
Extended Data Fig. 2 ZNFX1 deficiency impairs antiviral immune response in vitro and in vivo.
a, Quantitative RT-PCR (qRT-PCR) analysis of Znfx1 mRNA expression in A549 and L929 cells transfected with control siRNA or ZNFX1 siRNA 1#, 2# or 3# for 48 hrs (n = 3 independent experiments). b, Western blot analysis of ZNFX1 protein expression in A549 cells transfected with control siRNA or human ZNFX1 siRNA 1# for 48 hrs. c, ELISA of IFN-α or IFN-β production in cell supernatants from A549 cells with target gene knockdown for 48 hrs followed by VSV infection or poly I:C stimulation for another 12 hrs (n = 4 independent experiments). d, qRT-PCR analysis of VSV mRNA expression (left panel) and plaque assay analysis of VSV titer (right panel) of A549 cells transfected with RIG-I, ZNFX1 expressing plasmids or empty vector plasmid for 24 hrs and then infected with VSV at an MOI of 2 for 16 hrs (n = 3 independent experiments). e, FACS analysis of Znfx1+/+ and Znfx1-/- 293T cells followed by VSV-eGFP infection at 0.5 MOI for the indicated time points (n = 3 independent experiments). f, Znfx1-/- 293T and A549 clones were generated by the CRISPR-Cas9 method. Deficiency of target genes in the KO clones were confirmed by immunoblotting analysis. g, qRT-PCR analysis of viral mRNA transcripts in VSV, EMCV, H1N1 and HSV-1 infected A549 cells with control siRNA (si Control) or Znfx1-specific siRNA (si ZNFX1) (n = 3 independent experiments). h, ELISA of IFN-α in supernatants of BMDMs from WT and Znfx1-/- mice infected with VSV or HSV-1 for 16 hrs (n = 5 independent experiments). All data are shown as the mean ± s.d. P values were calculated using two-tailed unpaired Student’s t-test. For b, f, the experiments were repeated three times, independently, with similar results obtained.
Extended Data Fig. 3 ZNFX1 positively regulates IFN-β signaling.
a, Illustration of the CRISPR-Cas9 strategy to generate Znfx1-deficient mice and primer design used in (b). b, Genotyping of the ZNFX1 mutant pups. c, Immunoblot analysis of ZNFX1 protein levels in Znfx1+/+ and Znfx1-/- MEFs cells. d, qRT-PCR analysis of Ifnb1 and ISGs mRNA levels in A549 cells transfected with siControl (si Ctrl) or si ZNFX1 for 48 hrs and then infected with VSV for the indicated time points (n = 3 independent experiments). e, qRT-PCR analysis of Ifnb1 and ISGs mRNA expression in Znfx1+/+ and Znfx1-/- 293T cells followed by VSV infected with increasing MOI (0.5 and 1) for 0, 8 and 16 hrs (n = 3 independent experiments). For b, c, the experiments were repeated three times, independently, with similar results obtained. Data in d, e are the mean ± s.d. P values were calculated using two-tailed unpaired Student’s t-test.
Extended Data Fig. 4 ZNFX1 localizes to mitochondria and interacts with MAVS.
a, FACS analysis of Znfx1-/- A549 cells transfected with ZNFX1 and its mutants expressing plasmids or empty vector (EV) for 24 hrs followed by VSV-eGFP infection at an MOI of 2 for 6 hrs. b, Endogenous level of ZNFX1 protein in mitochondrial fractions in WT and Mavs-/- 293T with or without VSV infection at 1 MOI for 6 hrs. COX-IV was used as the loading control. Data are representative of at least three independent experiments.
Extended Data Fig. 5 The expression of Znfx1 in different tissues and cell types, and the phylogenetic tree of Znfx1.
a-c, The expression of Znfx1, Rig-I and Mda5 in different tissues and cell types as per BioGPS. d, Phylogenetic tree of ZNFX1 and RIG-I using an amino acid sequence alignment among different species.
Extended Data Fig. 6 Alignment of ZNFX1 amino acid sequences in human, mouse, rat and zebrafish.
Shading indicates sequence conservation, with darker gray indicating a higher degree of conservation.
Extended Data Fig. 7 Work model of mitochondria-localized ZNFX1 functions as a dsRNA sensor to initiate antiviral responses through MAVS.
Upon RNA virus infection, ZNFX1 induces type I interferon response by interacting with MAVS in the early stage, thus primes the expression of a number of ISGs, including RIG-I and MDA5. The induced sensors further enhance the antiviral immune response by amplifying ISGs expression.
Supplementary information
Supplementary Tables 1
Conservative analysis of ZNFX1 and DDX58 in different species.
Supplementary Tables 2
Information about the primers used in the study.
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 1
Unprocessed Western blots and/or gels
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 2
Unprocessed Western blots and/or gels
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 4
Unprocessed Western blots and/or gels
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 5
Unprocessed Western blots and/or gels
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 6
Unprocessed Western blots and/or gels
Source Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 2
Unprocessed Western blots and/or gels
Source Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 3
Unprocessed Western blots and/or gels
Source Data Extended Data Fig. 4
Unprocessed Western blots and/or gels
Rights and permissions
About this article
Cite this article
Wang, Y., Yuan, S., Jia, X. et al. Mitochondria-localised ZNFX1 functions as a dsRNA sensor to initiate antiviral responses through MAVS. Nat Cell Biol 21, 1346–1356 (2019). https://doi.org/10.1038/s41556-019-0416-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0416-0
- Springer Nature Limited
This article is cited by
-
Novel potent molecular glue degraders against broad range of hematological cancer cell lines via multiple neosubstrates degradation
Journal of Hematology & Oncology (2024)
-
Transcriptome Analysis of Heterosis in Survival in the Hybrid Progenies of ‘Haida No. 1’ and Orange-Shelled Lines of the Pacific Oyster Crassostrea gigas
Journal of Ocean University of China (2024)
-
Interferons are key cytokines acting on pancreatic islets in type 1 diabetes
Diabetologia (2024)
-
Cellular functions of eukaryotic RNA helicases and their links to human diseases
Nature Reviews Molecular Cell Biology (2023)
-
The MEF2A transcription factor interactome in cardiomyocytes
Cell Death & Disease (2023)