Abstract
Alkaptonuria is a rare autosomal recessive inherited disorder of tyrosine metabolism, which causes ochronosis, arthropathy, cardiac valvular calcification, and urolithiasis. The epidemiology of alkaptonuria in East Asia is not clear. In this study, patients diagnosed with alkaptonuria from January 2010 to June 2020 were reviewed. Their clinical and molecular features were further compared with those of patients from other countries. Three patients were found to have alkaptonuria. Mutation analyses of the homogentisate 1,2-dioxygenase gene (HGD) showed four novel variants c.16-2063 A > C, p.(Thr196Ile), p.(Gly344AspfsTer25), and p.(Gly362Arg) in six mutated alleles (83.3%). RNA sequencing revealed that c.16-2063 A > C activates a cryptic exon, causing protein truncation p.(Tyr5_Ile6insValTer17). A literature search identified another 6 patients with alkaptonuria in East Asia; including our cases, 13 of the 18 mutated alleles have not been reported elsewhere in the world. Alkaptonuria is rare in Taiwan and East Asia, with HGD variants being mostly novel and private.
Similar content being viewed by others
Introduction
Human homogentisate 1,2-dioxygenase, a homogentisic acid oxidase encoded by the HGD gene, catalyzes the conversion of homogentisic acid (HGA), a metabolite of tyrosine, to 4-maleylacetoacetate. Alkaptonuria (AKU; MIM # 203500) is a rare autosomal recessive disease caused by tissue accumulation and urinary excretion of HGA1. The hallmark of the disease is dark urine, which is caused by the oxidization of HGA. Associated morbidities include ochronosis, ochronotic arthropathy, cardiac valvular calcification, and urolithiasis in the 4th to 6th decades of life2. Although the life expectancy of patients with AKU is not reduced, their quality of life deteriorates mainly due to painful joint diseases3. The disease is not yet curable, and symptomatic relief management, such as physiotherapy, painkillers, and joint replacement therapy, is necessary4. Recently, nitisinone, a potent inhibitor of HGA production, was shown to decrease urinary HGA excretion and decelerate disease progression5.
The worldwide prevalence of alkaptonuria is estimated to be 1:100,000 to 1:1,000,0002,6, but the disease is more prevalent in Slovakia, Jordan, the Dominican Republic, and India4. HGD p.(Gly161Arg) is a common disease-causing variant in Slovakia and the Czech Republic; HGD p.(Cys120Trp) is common in the Dominican Republic7. The most prevalent variants in European countries excluding the abovementioned is p.(Met368Val), accounting for 11.2% of all HGD variants, followed by p.(Val300Gly), p.(Gly270Arg), and p.(Pro230Ser), which together account for ~9.5%7,8. Because AKU is rarely reported in Asia, we conducted this study to examine its characteristics in this region.
Results
Demographic data and clinical presentation of patients
Three Taiwanese (Han ethnicity) patients, two males, and one female, with a diagnosis of AKU were identified. Two of the patients had dark urine during infancy or early childhood, and one had dark urine at the age of 13 years. Their current ages are 10–20 years (Table 1). None of them showed a significant elevation in plasma tyrosine.
Patient 1 is a 20-year-old man. Dark urine was noted at 13 years of age. His porphyria test was negative, though urine organic acid analysis revealed a large amount of HGA. No arthralgia or hyperpigmentation was present currently.
Patient 2 is a 12-year-old female who was noted to have dark urine since early childhood. She was diagnosed at 8 years of age after a urine organic acid analysis, which revealed a large amount of HGA. Currently, she has ochronosis of the sclera (Fig. 1, arrow). No urolithiasis was detected by renal sonography, but she has frequent pain over the right hip and both knees.
Patient 3 is a 10-year-old boy. He was diagnosed at 5 years of age. When left at room temperature, his urine changed to purple-red in color in 1 hour and was brownish in half a day; these changes were more prominent after eating chocolate or seaweed. A urine organic acid analysis revealed a marked elevation in HGA. At present, he has no hyperpigmentation or urolithiasis; however, he did recently complain about intermittent bilateral knee joint pain.
Molecular findings
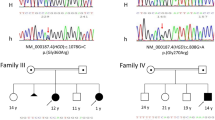
Molecular analyses revealed two HGD variants in each patient. Two of these variants have been previously reported: c.291 G > A p.(Trp97Ter) and c.473 C > T p.(Pro158Leu). p.(Trp97Ter), a nonsense variant, is reported as pathogenic in ClinVar, and other pathogenic variants at this amino acid p.(Trp97Arg) and p.(Trp97Gly) have also been reported, indicating that it is likely an important site for disease variants2,9. The variant p.(Pro158Leu), which occurs at CpG dinucleotides, is thought to be a mutation hot spot10,11. The other four variants have not been reported to date: c.16-2063 A > C, p.(Thr196Ile), p.(Gly344AspfsTer25), and p.(Gly362Arg) (Table 2). The frameshift variants p.(Gly344AspfsTer25) are predicted to be pathogenic. The two missense variants p.(Thr196Ile) and p.(Gly362Arg) are located in beta-strands of the HGD protein and have very low allele frequency in normal populations (maximal minor allele frequency of 0.0003 for p.(Thr196Ile); p.(Gly362Arg) was not previously described). Gly362 is proximal to the iron-binding site (residues 335, 341, and 371) of HGD protein. Both of them were predicted to be pathogenic by mCSM and HGDiscovery12,13.
DNA sequencing for Patient 3 revealed a heterozygous intronic variant c.16-2063 A > C (Fig. 2a, arrow) with a maximal minor allele frequency of 0.0006. To demonstrate the function of this variant, RNA-sequencing (RNA-Seq) from peripheral blood total RNA of Patient 3 and one control was performed. Sashimi plot analysis of the RNA-Seq data suggested the inclusion of a cryptic exon in the patient (Fig. 2b, arrow). Because the number of reads in RNA-Seq was small, we further verified aberrant splicing by reverse transcription PCR (RT-PCR). The results of RT-PCR of exons 1–3 of HGD revealed that, other than the normal product (arrow), several larger-than-expected fragments were found (Fig. 3a, star). Real-time PCR analysis reveals that the total amount of exons 1–3 HGD RNA was not decreased in the patient (Fig. 3b), and these PCR products revealed a shift in melting temperature (Fig. 3c). Next-generation sequencing (NGS) analysis of the PCR products revealed the inclusion of a previously described 126-bp cryptic exon in half of the reads in the patient (Fig. 3d). Some reads also contained other less-frequent cryptic exons (data not shown). A small portion of the reads in the control also contained the 126-bp cryptic exon. A zoom-in view of the cryptic exon revealed that the inclusion of the whole cryptic exon would cause frameshift and protein truncation (Fig. 3e). Therefore, variant c.16-2063 A > C is predicted as p.Tyr5_Ile6insValTer17. The c.16-2063 A > C variant is likely to disrupt an SRSF5-binding site (TATCAGG) and then activate the inclusion of the cryptic exon (Fig. 3f).
a DNA sequencing reveals a heterozygous variant, c.16-2063 A > C (arrow), in Patient 3 but not in the controls. b Results from RNA sequencing of Patient 3 and one control. Sashimi plot analysis suggests the inclusion of a cryptic exon (arrow) in the patient. The number of reads supporting the prediction of splicing is marked on the connecting lines.
a Agarose gel electrophoresis analysis reveals, other than the normal product (arrow), several larger-than-expected fragments in the patient (star). b Real-time PCR analysis reveals that the total amount of exons 1–3 HGD RNA was higher in Patient 3 than in the control. c High-resolution melting analysis reveals a shift in melting temperature in the patient. d NGS analysis reveals the inclusion of a 126-bp cryptic exon (283 reads) in half of the products (446 reads for exon 2) in the patient. A small portion of the reads in the control also contains this cryptic exon. There are other less-frequent cryptic exons included in the patient (not shown). e A zoom-in view of the cryptic exon reveals a major transcript (arrow) that causes protein truncation (Tyr5_Ile6insValTer17), and a minor transcript that also causes protein truncation (open arrow). f The c.16-2063 A > C variant is predicted to disrupt an SRSF5-binding site (TATCAGG) and then activate the inclusion of the cryptic exon (genomic coordinate 3:120,396,751–120,396,876).
In order to confirm that the c.16-2063 A > C variant is in trans to another likely pathogenic variant (c.1084 G > A at exon 13), allele-specific PCR was designed for c.1084 G > A (Fig. 4a). The c.16-2063 A > C variant is supposed to be on the 1084 G (wildtype) allele. The results revealed that RT-PCR with the 1084 G primer gave 1157-bp products (Fig. 4b, arrow) in the controls, but gave a major higher molecular weight product (Fig. 4b, star) in the patient. There may be an 1157-bp band in the patient, but that band only constituted 18% of the total PCR product as estimated by densitometry.
a Allele-specific PCR was designed for the c.1084 G > A variant. The 1084 A allele is normal in splicing, while the 1084 G allele contains the cryptic exon. The positions of the primers are marked by blue arrows. b RT-PCR with the 1084 G wildtype primer gives the 1157-bp products (arrow) in the controls, but gives a major band with a higher molecular weight in the patient. There may be a small amount (18% by densitometry) of normal-size PCR products in the patient. RT-PCR with the 1084 A mutant primer gives the 1157-bp product (open arrow) in the patient but not in the controls.
Variant interpretation at the protein level for novel variants
The active form of HGD is a delicate hexamer, and amino-acid residues involving protomer folding, hexamer assembly, and substrate binding have been shown to play important roles in HGD disease variants4,12. We employed structure-based variant characterization tools to identify the molecular consequences of the novel variants in the current study, including mCSM, mCSM-lig, mCSM-PPI2, and HGDiscovery (Table 3)12,13,14,15,16,17,18,19.
Thr196 is located distal to the active site and hexamer interface (Fig. 5A). Therefore, its variant to Ile is unlikely to disrupt substrate binding or the formation of the active hexamer. Thr196 is, however, a buried and conserved residue that is intolerant to missense variant, and the introduction of an Ile leads to the loss of key hydrogen bonds to Val198 and mild steric clashes (Fig. 5B). Consistent with this, the variants of Thr196Ile are predicted to mildly destabilize the HGD structure, leading to it being predicted by HGDiscovery as pathogenic.
The sidechain of Thr196 (A) makes a series of hydrogen bonds to neighboring residues, which would be lost upon variant to Ile (B). A variant of the positive phi Gly362 (C) to Arg (D) would lead to large steric clashes within the structure of HGD. This highlights that both variants would lead to significant structural consequences and be quite deleterious. The wild-type residues are shown as green sticks, and the mutant residue is in magenta sticks. Hydrogen bonds are shown as red dashed lines, polar interactions as orange dashed lines, hydrophobic interactions as green dashed lines, and steric clashes as red disks.
Gly362 is a buried residue located 10 Å from the active site and within 15 Å of the hexamer interface (Fig. 5C). As a positive phi glycine, variant to arginine is likely to lead to significant disruption of the protein structure. The introduction of the larger arginine sidechain is also harder to accommodate within the tightly packed buried core and leads to significant steric clashes (Fig. 5D). This is consistent with G364 being identified as intolerant to missense variants and predicted to destabilize the protein, hexameric structure, and ligand binding. It was consequently identified as pathogenic by HGDiscovery.
Discussion
AKU has never been reported in Taiwan before, and we observed only three cases in a medical center that treats the largest number of rare disorders in Taiwan. Usually, symptoms of AKU initiate from ochronotic pigmentation in the 4th decade of life, which is followed by painful joint destruction requiring artificial joint replacement and cardiac valvular problems in the mid-50s and the formation of renal stones in the 60 s2. However, some of our three patients as young as 12 years already had ochronosis or arthralgia. Nevertheless, none of the 10- and 12-year-olds with arthralgia presented with joint destruction. Therefore, it is possible that the joint pain was psychological owing to the stress from having a chronic illness.
Biochemical diagnosis of AKU based on the detection of HGA in urine samples is accurate. Nevertheless, although HGA can be detected in routine urinary organic acid analysis, the disease is so rare that the operator may not recognize the compound unless they are reminded by a physician who suspects this diagnosis. Molecular diagnosis of AKU can also be challenging, especially in East Asia, where the disease is extremely rare and most of the variants are novel. For example, the deep intron variant c.16-2063 A > C detected in Taiwanese patients are not included in any of the human genome databases. Therefore, we needed to perform additional analyses to demonstrate the pathogenicity of this variant that affects splicing. It was fortunate that the cryptic exon has been described, thus, our capture probes included this exon; otherwise, we would not have detected this variant. RNA-Seq is certainly very helpful for revealing splicing aberrations. MetaDome software is also useful because this software provides good visualization of the tolerability of variants over the entire protein. With this tool, it is easier to evaluate novel variants by comparing them with other reported variants, which is especially helpful in ultrarare diseases such as AKU.
We retrieved 100 articles from PubMed but excluded 76 of them due to lack of information; we added both HGMD and ClinVar records. Not including our cohort, a total of 223 variants in 594 patients have been reported as disease-causing variants at the time of calculation (Dec 2020). However, only 6 patients from East Asia (China, South Korea, and Japan) are reported in PubMed20,21,22,23,24,25. Together with our three patients, 13 of the 18 East Asian mutated alleles have not been detected in other countries. The 13 mutated alleles comprise 12 variants p.(Gln33 Arg), p.(Glu42SerfsTer69), p.(Glu74Val), c.342 + 3A > C, p.(Gly152Ala), p.(Thr196Ile), p.(Glu329Cys), p.(Gly344AspfsTer25), p.(Gly362Arg), p.(Ser366_Thr367dup), c.16-2063 A > C, and c.469 + 1 G > C. Therefore, most of the AKU patients in East Asia harbor private variants in HGD. All HGD missense variants were analyzed by the pathogenicity analysis tool MetaDome, and the results revealed that most are located in intolerant parts of the protein; for the Asian variants, tolerant for p.(Glu74Val), slightly tolerant for p.(Gly362Arg), neutral for p.(Thr196Ile), slightly intolerant for p.(Gln33Arg) and p.(Phe329Cys), intolerant for p.(Pro158Leu) and p.(Glu168Lys), and highly intolerant for p.(Gly152Ala) (Fig. 6).
We only identified three cases in our institute, and there is no previous report of AKU in Taiwan. Although we were not able to calculate the prevalence of AKU in Taiwan, the disease must be very rare, as in all of East Asia. In addition, we were unable to fully elucidate the phenotype of the disease in our four Taiwanese patients because they were still young. Nonetheless, the suspicious ochronosis and arthralgia in a portion of them alert us to the potential for more serious symptoms in the future.
Alkaptonuria is rare in Taiwan and East Asia, with HGD variants being mostly novel. More efforts involving molecular analysis are expected to contribute to the diagnosis of new cases in this region.
Materials and methods
Patients
From Jan 2010 to Jun 2020, patients diagnosed with AKU at National Taiwan University Hospital were reviewed. Informed consent was obtained from all subjects or their guardians, and the guardian of patient 2 provided written consent for the publication of the photograph in Fig. 1. Clinical information, including the age of onset and clinical presentation, was collected. Diagnostic biochemical analyses performed for these patients included urine organic acid analysis by gas chromatography-mass spectrometry or thin-layer chromatography to reveal the presence of HGA in urine samples. The study was approved by the Institutional Review Board (IRB No. 201505135RIN) in our hospital.
Molecular analyses
Variant analyses of HGD (RefSeq: NM_000187.4) were performed by targeted panel sequencing using a SeqCap EZ probe (Roche Nimbelgen, Basel, Switzerland) and MiSeq sequencer (Illumina, San Diego, CA, USA) to produce 300-bp paired-end reads, with an average coverage >150 crossing exons and flanking intronic (50 bp) sequences. Sequence alignment to the human reference genome (GRCh37) was performed using Burrows-Wheeler Aligner, and variant calling was performed using Genome Analysis Tool Kit (GATK v4.0, Broad Institute)26. Variants were annotated by ANNOVAR (http://wannovar.wglab.org/)27. DNA and protein sequence variants were described as recommended by Human Genome Variation Society (HGVS, http://varnomen.hgvs.org/). For missense variants, we employed Sorting Intolerant From Tolerant (SIFT, https://sift.bii.a-star.edu.sg/), PolyPhen-2 (Polymorphism Phenotyping v2, http://genetics.bwh.harvard.edu/pph2/), mCSM (http://biosig.unimelb.edu.au/mcsm/)16, mCSM-PPI2 (http://biosig.unimelb.edu.au/mcsm_ppi2/)15,18 and mCSM-lig (http://biosig.unimelb.edu.au/mcsm_lig/)17 to predict effects on protein function and structure using the experimental crystal structure of HGD (PDB ID: 1EY2). Intramolecular interactions were calculated and visualized using Arpeggio28 and mutational tolerance was calculated using MTR-Viewer13,19. For intronic variants, we used Human Splicing Finder (http://umd.be/Redirect.html) and Splice AI to predict their splice effects. We also searched the HGD mutation database (http://hgddatabase.cvtisr.sk/home.php) and ClinVar to assess whether variants have been reported. The pathogenicity of variants was classified according to the American College of Medical Genetics and Genomics (ACMG) and the Association for Molecular Pathology guidelines29.
RNA-Seq was performed on total RNA extracted from peripheral whole-blood samples. After the depletion of ribosomal RNA and globin mRNA, each sample was sequenced with a total output of 50 M reads. Data obtained from RNA-Seq were aligned using STAR and annotated for DNA analysis. RT-PCR was further applied to validate aberrant splicing. The region of exon 1–3 was amplified to demonstrate aberrant splicing. For quantitative PCR using the SYBR Master Mix (Applied Biosystems), GAPDH was used as an internal control. HRM was performed by Rotor-gene Q (Qiagen). Allele-specific PCR for RNA was designed for the c.1084 G > A variant, and the specific lower primers were 1084 G (wildtype) 5’-tcattgtgctgtgtagactccc and 1084 A (mutant) 5’-tcattgtgctgtgtagactcct. Densitometry was performed using ImageJ. All primer sequences are listed in Supplementary Table 1. All blots derived from the same experiment and were processed in parallel.
Computational analysis to evaluate possible enzyme inactivation of novel missense variants of HGD was used by webserver of HGDiscovery (http://biosig.unimelb.edu.au/hgdiscovery/submit_prediction)12,13,15. Asian HGD variants were illustrated by the tolerance landscape using Metadome Version 1.0.1 webserver (https://stuart.radboudumc.nl/metadome/dashboard)14.
Literature search
We searched PubMed publications using the criteria “(alkaptonuria) AND ((mutation) OR (variant) OR (gene diagnosis))”. We targeted studies published after 1996, and the time the human HGD gene was mapped to chromosome 3q. Exclusion criteria were as follows: 1. cases without ethnicity mentioned or a clear genotype, 2. literature without full-text availability, and 3. literature not written in English or Chinese. We then checked ClinVar, Human Gene Mutation Database (HGMD), and the HGD mutation database from Leiden Open Variation Database (LOVD) for the HGD mutations described in the publications retrieved.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Sequence data have been deposited at the NCBI (SRA), under accession number PRJNA765906. Variants are available in HGD mutation database (http://hgddatabase.cvtisr.sk/home.php) under number AKU_00248, AKU_00249, AKU_00245, and AKU_00246. All other data that support the findings of this study are available from the corresponding author upon request.
References
Ladu, B. N., Zannoni, V. G., Laster, L. & Seegmiller, J. E. Nature of the defect in Tyrosine metabolism in Alcaptonuria. J. Biol. Chem. 230, 251–260 (1958).
Phornphutkul, C. et al. Natural history of alkaptonuria. N. Engl. J. Med. 347, 2111–2121 (2002).
Mistry, J. B., Bukhari, M. & Taylor, A. M. Alkaptonuria. Rare Dis. 1, e27475 (2013).
Zatkova, A., Ranganath, L. & Kadasi, L. Alkaptonuria: current perspectives. Application Clin. Genet. 13, 37–47 (2020).
Ranganath, L. R. et al. Nitisinone arrests ochronosis and decreases rate of progression of Alkaptonuria: evaluation of the effect of nitisinone in the United Kingdom National Alkaptonuria Centre. Mol. Genet. Metab. 125, 127–134 (2018).
Ranganath, L. et al. Identification of alkaptonuria in the general population: a United Kingdom experience describing the challenges, possible solutions and persistent barriers. J. Inherit. Metab. Dis. 34, 723–730 (2011).
Zatkova, A. An update on molecular genetics of Alkaptonuria (AKU). J. Inherit. Metab. Dis. 34, 1127–1136 (2011).
Zatkova, A. et al. High frequency of alkaptonuria in Slovakia: evidence for the appearance of multiple mutations in HGO involving different mutational hot spots. Am. J. Hum. Genet. 67, 1333–1339 (2000).
Beltrán-Valero de Bernabé, D. et al. Mutation and polymorphism analysis of the human homogentisate 1, 2-dioxygenase gene in alkaptonuria patients. Am. J. Hum. Genet. 62, 776–784 (1998).
Gucev, Z. S. et al. Early-onset ocular ochronosis in a girl with alkaptonuria (AKU) and a novel mutation in homogentisate 1,2-dioxygenase (HGD). Prilozi 32, 305–311 (2011).
Vilboux, T. et al. Mutation spectrum of homogentisic acid oxidase (HGD) in alkaptonuria. Hum. Mutat. 30, 1611–1619 (2009).
Ascher, D. B. et al. Homogentisate 1,2-dioxygenase (HGD) gene variants, their analysis and genotype-phenotype correlations in the largest cohort of patients with AKU. Eur. J. Hum. Genet. 27, 888–902 (2019).
Karmakar, M. et al. HGDiscovery: an online tool providing functional and phenotypic information on novel variants of homogentisate 1,2- dioxigenase. bioRxiv, Preprint at: https://doi.org/10.1101/2021.04.26.441386 (2021).
Wiel, L. et al. MetaDome: pathogenicity analysis of genetic variants through aggregation of homologous human protein domains. Hum. Mutat. 40, 1030–1038 (2019).
Rodrigues, C. H. M., Myung, Y., Pires, D. E. V. & Ascher, D. B. mCSM-PPI2: predicting the effects of mutations on protein-protein interactions. Nucleic Acids Res. 47, W338–W344 (2019).
Pires, D. E., Ascher, D. B. & Blundell, T. L. mCSM: predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics 30, 335–342 (2014).
Nemethova, M. et al. Twelve novel HGD gene variants identified in 99 alkaptonuria patients: focus on ‘black bone disease’ in Italy. Eur. J. Hum. Genet. 24, 66–72 (2016).
Rodrigues, C. H. M., Pires, D. E. V. & Ascher, D. B. mmCSM-PPI: predicting the effects of multiple point mutations on protein-protein interactions. Nucleic Acids Res 49, W417–W424 (2021).
Silk, M., Petrovski, S. & Ascher, D. B. MTR-Viewer: identifying regions within genes under purifying selection. Nucleic Acids Res. 47, W121–W126 (2019).
Higashino, K. et al. A novel point mutation associated with alkaptonuria. Clin. Genet 53, 228–229 (1998).
Yang, Y. J. et al. First report of HGD mutations in a Chinese with alkaptonuria. Gene 518, 467–469 (2013).
Li, H. et al. Two novel mutations in the homogentisate-1,2-dioxygenase gene identified in Chinese Han child with alkaptonuria. J. Pediatr. Endocrinol. Metab. 28, 453–456 (2015).
Cho, S.-Y. & Kim, J. H. Identification of HGD mutations in an alkaptonuria patient: using the Internet to seek rare diseases. J. Genet. Med. 15, 17–19 (2018).
Hu, M. et al. Gene diagnosis of alkaptonuria in an infant. Zhongguo Dang Dai Er Ke Za Zhi 14, 796–797 (2012).
Zatkova, A. et al. Identification of 11 novel homogentisate 1,2 dioxygenase variants in alkaptonuria patients and establishment of a novel LOVD-based HGD mutation database. JIMD Rep. 4, 55–65 (2012).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303 (2010).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Jubb, H. C. et al. Arpeggio: a web server for calculating and visualising interatomic interactions in protein structures. J. Mol. Biol. 429, 365–371 (2017).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Acknowledgements
This work was funded by a grant from the Ministry of Science and Technology (107-2314-B-002 -164 -MY3) of Taiwan and National Taiwan University Hospital (NTUH 105-002959). We thank all the parents and their children who participated in the study. D.B.A. was supported by an Investigator Grant from the National Health and Medical Research Council (NHMRC) of Australia (GNT1174405) and in part by the Victorian Government’s Operational Infrastructure Support Program. We would also thank Dr. Andrea Zatkova for helping in the submission of variants to HGD mutation database.
Author information
Authors and Affiliations
Contributions
N.C.L, Y.H.C., D.B.A., and W.L.H. conceptualized the study and provided the methodology; C.Y.L., I.J.T., P.C.C., Y.H.H., and Y.L.L analyzed the clinical data; C.Y.L., Y.H.H., and Y.L.L analyzed and interpreted the next-generation sequencing and RT-PCR sequencing; D.B.A. analyzed the computational prediction for novel missense variants; C.Y.L., N.C.L, Y.H.C., and W.L.H. drafted the first version of the manuscript; all authors participated to the writing, reviewing, and editing of the manuscript and agreed on the final version for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lai, CY., Tsai, IJ., Chiu, PC. et al. A novel deep intronic variant strongly associates with Alkaptonuria. npj Genom. Med. 6, 89 (2021). https://doi.org/10.1038/s41525-021-00252-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41525-021-00252-2
- Springer Nature Limited
This article is cited by
-
Alkaptonuria
Nature Reviews Disease Primers (2024)
-
Clinical development innovation in rare diseases: overcoming barriers to successful delivery of a randomised clinical trial in alkaptonuria—a mini-review
Orphanet Journal of Rare Diseases (2023)