Abstract
Guanylate binding proteins (GBPs) are prominent regulators of immunity not known to be required for nuclear envelope formation and morphogenesis. Here we identify the Arabidopsis GBP orthologue AtGBPL3 as a lamina component with essential functions in mitotic nuclear envelope reformation, nuclear morphogenesis and transcriptional repression during interphase. AtGBPL3 is preferentially expressed in mitotically active root tips, accumulates at the nuclear envelope and interacts with centromeric chromatin as well as with lamina components transcriptionally repressing pericentromeric chromatin. Reduced expression of AtGBPL3 or associated lamina components similarly altered nuclear morphology and caused overlapping transcriptional deregulation. Investigating the dynamics of AtGBPL3–GFP and other nuclear markers during mitosis (1) revealed that AtGBPL3 accumulation on the surface of daughter nuclei precedes nuclear envelope reformation and (2) uncovered defects in this process in roots of AtGBPL3 mutants, which cause programmed cell death and impair growth. AtGBPL3 functions established by these observations are unique among dynamin-family large GTPases.








Similar content being viewed by others
Data availability
All data supporting the findings of this study are presented within the article (including Extended Data) and its Supplementary Information. Key reagents and resources are listed in Supplementary Table 1; unique materials contained in this table can be obtained from the authors. Oligonucleotide sequences are contained in Supplementary Table 2. RNA-seq dataset 1 (WT, gbpl3-5, gbpl3-5comp2 and mail1) is available at NCBI Sequence Read Archive (NCBI BioProject Accession PRJNA726378, BioSample Accessions SAMN18928784–SAMN18928795). RNA-seq dataset 2 (WT, gbpl3-5, gbpl3-5comp2, gbpl3-5compK83A_2, crwn1-1/4-1 and crwn4-1) as well as AtGBPL3–GFP and AtH2B–GFP ChIP–seq data have been deposited in NCBI Gene Expression Omnibus109 (NCBI GEO Series Accession GSE221669; RNA-seq GSE221661; ChIP–seq GSE221668). The following publicly accessible databases were used: TAIR (https://www.arabidopsis.org/index.jsp), Pfam (https://pfam.xfam.org), ReMap (https://remap2022.univ-amu.fr), NUP1 ReChIP–seq data (SRP079108) and CRWN1 ChIP–seq (PRJNA497671). Source data are provided with this paper.
References
Praefcke, G. J. K. & McMahon, H. T. The dynamin superfamily: universal membrane tubulation and fission molecules? Nat. Rev. Mol. Cell Biol. 5, 133–147 (2004).
Prakash, B., Praefcke, G. J. K., Renault, L., Wittinghofer, A. & Herrmann, C. Structure of human guanylate-binding protein 1 representing a unique class of GTP-binding proteins. Nature 403, 567–571 (2000).
Ferguson, S. M. & De Camilli, P. Dynamin, a membrane-remodelling GTPase. Nat. Rev. Mol. Cell Biol. 13, 75–88 (2012).
Hu, X., Wu, F., Sun, S., Yu, W. & Hu, J. Human atlastin GTPases mediate differentiated fusion of endoplasmic reticulum membranes. Protein Cell 6, 307–311 (2015).
McNew, J. A., Sondermann, H., Lee, T., Stern, M. & Brandizzi, F. GTP-dependent membrane fusion. Annu. Rev. Cell Dev. Biol. 29, 529–550 (2013).
Daumke, O. & Praefcke, G. J. K. Invited review: mechanisms of GTP hydrolysis and conformational transitions in the dynamin superfamily. Biopolymers 105, 580–593 (2016).
Vestal, D. J. The guanylate-binding proteins (GBPs): proinflammatory cytokine-induced members of the dynamin superfamily with unique GTPase activity. J. Interf. Cytokine Res. 25, 435–443 (2005).
Huang, S., Meng, Q., Maminska, A. & MacMicking, J. D. Cell-autonomous immunity by IFN-induced GBPs in animals and plants. Curr. Opin. Immunol. 60, 71–80 (2019).
Huang, S., Zhu, S., Kumar, P. & MacMicking, J. D. A phase-separated nuclear GBPL circuit controls immunity in plants. Nature 594, 424–429 (2021).
Kim, J. H. et al. Increasing the resilience of plant immunity to a warming climate. Nature 607, 339–344 (2022).
Krapp, C. et al. Guanylate binding protein (GBP) 5 is an interferon-inducible inhibitor of HIV-1 infectivity. Cell Host Microbe 19, 504–514 (2016).
Kim, B.-H. et al. A family of IFN-γ–inducible 65-kD GTPases protects against bacterial infection. Science 332, 717–721 (2011).
Kravets, E. et al. Guanylate binding proteins directly attack Toxoplasma gondii via supramolecular complexes. eLife 5, e11479 (2016).
Britzen-Laurent, N., Herrmann, C., Naschberger, E., Croner, R. S. & Stürzl, M. Pathophysiological role of guanylate-binding proteins in gastrointestinal diseases. World J. Gastroenterol. 22, 6434–6443 (2016).
Tretina, K., Park, E.-S., Maminska, A. & MacMicking, J. D. Interferon-induced guanylate-binding proteins: guardians of host defense in health and disease. J. Exp. Med. 216, 482–500 (2019).
Tripal, P. et al. Unique features of different members of the human guanylate-binding protein family. J. Interf. Cytokine Res. 27, 44–52 (2007).
Tang, Y., Ho, M. I., Kang, B.-H. & Gu, Y. GBPL3 localizes to the nuclear pore complex and functionally connects the nuclear basket with the nucleoskeleton in plants. PLoS Biol. 20, e3001831 (2022).
Ciska, M. & de la Espina, S. M. D. The intriguing plant nuclear lamina. Front. Plant Sci. 5, e166 (2014).
Pradillo, M., Evans, D. & Graumann, K. The nuclear envelope in higher plant mitosis and meiosis. Nucleus 10, 55–66 (2019).
Graumann, K. Finding the missing piece of the puzzle: how NMCPs fit into the plant nuclear lamina. J. Exp. Bot. 72, 6077–6080 (2021).
Groves, N. R. et al. Recent advances in understanding the biological roles of the plant nuclear envelope. Nucleus 11, 330–346 (2020).
Barz, B., Loschwitz, J. & Strodel, B. Large-scale, dynamin-like motions of the human guanylate binding protein 1 revealed by multi-resolution simulations. PLoS Comput. Biol. 15, e1007193 (2019).
Vöpel, T. et al. Mechanism of GTPase-activity-induced self-assembly of human guanylate binding protein 1. J. Mol. Biol. 400, 63–70 (2010).
Wehner, M. & Herrmann, C. Biochemical properties of the human guanylate binding protein 5 and a tumor-specific truncated splice variant. FEBS J. 277, 1597–1605 (2010).
Britzen-Laurent, N. et al. Intracellular trafficking of guanylate-binding proteins is regulated by heterodimerization in a hierarchical manner. PLoS ONE 5, e14246 (2010).
Olszewski, M. A., Gray, J. & Vestal, D. J. In silico genomic analysis of the human and murine guanylate-binding protein (GBP) gene clusters. J. Interf. Cytokine Res. 26, 328–352 (2006).
Verbelen, J.-P., De Cnodder, T., Le, J., Vissenberg, K. & Baluška, F. The root apex of Arabidopsis thaliana consists of four distinct zones of growth activities. Plant Signal. Behav. 1, 296–304 (2006).
Dickman, M., Williams, B., Li, Y., de Figueiredo, P. & Wolpert, T. Reassessing apoptosis in plants. Nat. Plants 3, 773–779 (2017).
Ambastha, V., Friedmann, Y. & Leshem, Y. Laterals take it better—emerging and young lateral roots survive lethal salinity longer than the primary root in Arabidopsis. Sci. Rep. 10, 3291 (2020).
Graumann, K., Runions, J. & Evans, D. E. Characterization of SUN-domain proteins at the higher plant nuclear envelope. Plant J. 61, 134–144 (2010).
Boisnard-Lorig, C. et al. Dynamic analyses of the expression of the HISTONE::YFP fusion protein in Arabidopsis show that syncytial endosperm is divided in mitotic domains. Plant Cell 13, 495–509 (2001).
Goto, C., Tamura, K., Fukao, Y., Shimada, T. & Hara-Nishimura, I. The novel nuclear envelope protein KAKU4 modulates nuclear morphology in Arabidopsis. Plant Cell 26, 2143–2155 (2014).
Sakamoto, Y. & Takagi, S. LITTLE NUCLEI 1 and 4 regulate nuclear morphology in Arabidopsis thaliana. Plant Cell Physiol. 54, 622–633 (2013).
Wang, H., Dittmer, T. A. & Richards, E. J. Arabidopsis CROWDED NUCLEI (CRWN) proteins are required for nuclear size control and heterochromatin organization. BMC Plant Biol. 13, 200 (2013).
Tang, Y., Dong, Q., Wang, T., Gong, L. & Gu, Y. PNET2 is a component of the plant nuclear lamina and is required for proper genome organization and activity. Dev. Cell 57, 19–31 (2022).
Poulet, A. et al. The LINC complex contributes to heterochromatin organisation and transcriptional gene silencing in plants. J. Cell Sci. 130, 590–601 (2017).
Hu, B. et al. Plant lamin-like proteins mediate chromatin tethering at the nuclear periphery. Genome Biol. 20, 87 (2019).
Bi, X. et al. Nonrandom domain organization of the Arabidopsis genome at the nuclear periphery. Genome Res. 27, 1162–1173 (2017).
Mikulski, P. et al. The chromatin-associated protein PWO1 interacts with plant nuclear lamin-like components to regulate nuclear size. Plant Cell 31, 1141–1154 (2019).
Pontvianne, F. & Liu, C. Chromatin domains in space and their functional implications. Curr. Opin. Plant Biol. 54, 1–10 (2020).
Hohenstatt, M. L. et al. PWWP-DOMAIN INTERACTOR OF POLYCOMBS1 interacts with polycomb-group proteins and histones and regulates Arabidopsis flowering and development. Plant Cell 30, 117–133 (2018).
Ühlken, C., Horvath, B., Stadler, R., Sauer, N. & Weingartner, M. MAIN-LIKE1 is a crucial factor for correct cell division and differentiation in Arabidopsis thaliana. Plant J. 78, 107–120 (2014).
Ikeda, Y. et al. Arabidopsis proteins with a transposon-related domain act in gene silencing. Nat. Commun. 8, 15122 (2017).
Boltz, K. A., Jasti, M., Townley, J. M. & Shippen, D. E. Analysis of poly(ADP-ribose) polymerases in Arabidopsis telomere biology. PLoS ONE 9, e88872 (2014).
Olvera-Carrillo, Y. et al. A conserved core of PCD indicator genes discriminates developmentally and environmentally induced programmed cell death in plants. Plant Physiol. 169, 2684–2699 (2015).
Sequeira-Mendes, J. et al. The functional topography of the Arabidopsis genome is organized in a reduced number of linear motifs of chromatin states. Plant Cell 26, 2351–2366 (2014).
Dittmer, T. A., Stacey, N. J., Sugimoto-Shirasu, K. & Richards, E. J. LITTLE NUCLEI genes affecting nuclear morphology in Arabidopsis thaliana. Plant Cell 19, 2793–2803 (2007).
Graumann, K. Evidence for LINC1-SUN associations at the plant nuclear periphery. PLoS ONE 9, e93406 (2014).
Choi, J. & Richards, E. J. The role of CRWN nuclear proteins in chromatin-based regulation of stress response genes. Plant Signal. Behav. 15, 1694224 (2020).
Musseau, C. et al. The tomato guanylate-binding protein SlGBP1 enables fruit tissue differentiation by maintaining endopolyploid cells in a non-proliferative state. Plant Cell 32, 3188–3205 (2020).
Goldman, R. D., Gruenbaum, Y., Moir, R. D., Shumaker, D. K. & Spann, T. P. Nuclear lamins: building blocks of nuclear architecture. Genes Dev. 16, 533–547 (2002).
Choi, J., Strickler, S. R. & Richards, E. J. Loss of CRWN nuclear proteins induces cell death and salicylic acid defense signaling. Plant Physiol. 179, 1315–1329 (2019).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Rottmann, T. & Stadler, R. Measuring sucrose transporter activities using a protoplast-esculin assay. Methods Mol. Biol. 2014, 253–266 (2019).
Kleinboelting, N., Huep, G., Kloetgen, A., Viehoever, P. & Weisshaar, B. GABI-Kat SimpleSearch: new features of the Arabidopsis thaliana T-DNA mutant database. Nucleic Acids Res. 40, 1211–1215 (2012).
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Holsters, M. et al. The functional organization of the nopaline A. tumefaciens plasmid pTiC58. Plasmid 3, 212–230 (1980).
Lazo, G. R., Stein, P. A. & Ludwig, R. A. A DNA transformation–competent Arabidopsis genomic library in Agrobacterium. Biotechnology 9, 963–967 (1991).
Deblaere, R. et al. Efficient octopine Ti plasmid-derived vectors for Agrobacterium-mediated gene transfer to plants. Nucleic Acids Res. 13, 4777–4788 (1985).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 166, 557–580 (1983).
Bernard, P. & Couturier, M. Cell killing by the F plasmid CcdB protein involves poisoning of DNA-topoisomerase II complexes. J. Mol. Biol. 226, 735–745 (1992).
Villarejo, M. R. & Zabin, I. β-galactosidase from termination and deletion mutant strains. J. Bacteriol. 120, 466–474 (1974).
James, P., Halladay, J. & Craig, E. A. Genomic libraries and a host strain designed for highly efficient two-hybrid selection in yeast. Genetics 144, 1425–1436 (1996).
Schiml, S., Fauser, F. & Puchta, H. CRISPR/Cas-mediated site-specific mutagenesis in Arabidopsis thaliana using Cas9 nucleases and paired nickases. Methods Mol. Biol. 1469, 111–122 (2016).
Steinert, J., Schiml, S., Fauser, F. & Puchta, H. Highly efficient heritable plant genome engineering using Cas9 orthologues from Streptococcus thermophilus and Staphylococcus aureus. Plant J. 84, 1295–1305 (2015).
Fischer, C., Kugler, A., Hoth, S. & Dietrich, P. An IQ domain mediates the interaction with calmodulin in a plant cyclic nucleotide-gated channel. Plant Cell Physiol. 54, 573–584 (2013).
Birdsell, D. N. et al. Melt analysis of mismatch amplification mutation assays (melt-MAMA): a functional study of a cost-effective SNP genotyping assay in bacterial models. PLoS ONE 7, e32866 (2012).
Rottmann, T. et al. Sugar transporter STP7 specificity for l-arabinose and d-xylose contrasts with the typical hexose transporters STP8 and STP12. Plant Physiol. 176, 2330–2350 (2018).
Johnson, M. A. & Kost, B. Pollen tube development. Methods Mol. Biol. 655, 155–176 (2010).
Bushnell, B. BBMap: A Fast, Accurate, Splice-Aware Aligner (Lawrence Berkeley National Laboratory, 2014).
Andrews, S., Krueger, F., Segonds-Pichon, A., Biggins, L. & Wingett, S. FastQC: A Quality Control Tool for High Throughput Sequence Data (Babraham Institute, 2010); https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Srivastava, A. et al. Alignment and mapping methodology influence transcript abundance estimation. Genome Biol. 21, 239 (2020).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2021).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Research 4, 1521 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer International, 2016).
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Softw. 4, 1686 (2019).
Lee, S., Cook, D. & Lawrence, M. plyranges: a grammar of genomic data transformation. Genome Biol. 20, 4 (2019).
Larsson, J. & Gustafsson, P. {eulerr}: area-proportional Euler and Venn diagrams with ellipses. Proc. Int. Work. Set. Vis. Reason. 2116, 84–91 (2018).
Wang, M., Zhao, Y. & Zhang, B. Efficient test and visualization of multi-set intersections. Sci. Rep. 5, 16923 (2015).
Gu, Z. Complex heatmap visualization. iMeta 1, e43 (2022).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Mi, H., Muruganujan, A., Ebert, D., Huang, X. & Thomas, P. D. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 47, D419–D426 (2019).
Gel, B. & Serra, E. karyoploteR: an R/Bioconductor package to plot customizable genomes displaying arbitrary data. Bioinformatics 33, 3088–3090 (2017).
Menetrier, Z. & Mestdagh, M. ReMapEnrich: Bioinformatics Tools to Compute Statistical Enrichment of Genomic Regions for ReMap Peaks (remap-cisreg, 2022); https://remap-cisreg.github.io/ReMapEnrich/
Karimi, M., Inzé, D. & Depicker, A. GATEWAYTM vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 7, 193–195 (2002).
Klahre, U., Becker, C., Schmitt, A. C. & Kost, B. Nt-RhoGDI2 regulates Rac/Rop signaling and polar cell growth in tobacco pollen tubes. Plant J. 46, 1018–1031 (2006).
Curtis, M. D. & Grossniklaus, U. A Gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133, 462–469 (2003).
Kost, B., Spielhofer, P. & Chua, N.-H. A GFP-mouse talin fusion protein labels plant actin filaments in vivo and visualizes the actin cytoskeleton in growing pollen tubes. Plant J. 16, 393–401 (1998).
Karasawa, S., Araki, T., Nagai, T., Mizuno, H. & Miyawaki, A. Cyan-emitting and orange-emitting fluorescent proteins as a donor/acceptor pair for fluorescence resonance energy transfer. Biochem. J. 381, 307–312 (2004).
Billou, I. et al. The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 433, 39–44 (2005).
Klahre, U. & Kost, B. Tobacco RhoGTPase ACTIVATING PROTEIN1 spatially restricts signaling of RAC/Rop to the apex of pollen tubes. Plant Cell 18, 3033–3046 (2006).
Soni, R., Carmichael, J. P. & Murray, J. A. H. Parameters affecting lithium acetate-mediated transformation of Saccharomyces cerevisiae and development of a rapid and simplified procedure. Curr. Genet. 24, 455–459 (1993).
Ueda, T., Yamaguchi, M., Uchimiya, H. & Nakano, A. Ara6, a plant-unique novel type Rab GTPase, functions in the endocytic pathway of Arabidopsis thaliana. EMBO J. 20, 4730–4741 (2001).
Camborde, L. et al. Detection of nucleic acid–protein interactions in plant leaves using fluorescence lifetime imaging microscopy. Nat. Protoc. 12, 1933–1950 (2017).
Escouboué, M., Camborde, L., Jauneau, A., Gaulin, E. & Deslandes, L. Preparation of plant material for analysis of protein–nucleic acid interactions by FRET-FLIM. Methods Mol. Biol. 1991, 69–77 (2019).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, 427–432 (2019).
Lupas, A., Van Dyke, M. & Stock, J. Predicting coiled coils from protein sequences. Science 252, 1162–1164 (1991).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Marchler-Bauer, A. et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 45, D200–D203 (2017).
Kosugi, S., Hasebe, M., Tomita, M. & Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl Acad. Sci. USA 106, 10171–10176 (2009).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Barakat, A., Matassi, G. & Bernardi, G. Distribution of genes in the genome of Arabidopsis thaliana and its implications for the genome organization of plants. Proc. Natl Acad. Sci. USA 95, 10044–10049 (1998).
Hammal, F., De Langen, P., Bergon, A., Lopez, F. & Ballester, B. ReMap 2022: a database of human, mouse, Drosophila and Arabidopsis regulatory regions from an integrative analysis of DNA-binding sequencing experiments. Nucleic Acids Res. 50, D316–D325 (2022).
Houben, A. & Schubert, I. DNA and proteins of plant centromeres. Curr. Opin. Plant Biol. 6, 554–560 (2003).
Cheng, K. et al. Histone tales: lysine methylation, a protagonist in Arabidopsis development. J. Exp. Bot. 71, 793–807 (2020).
Nitsch, S., Shahidian, L. Z. & Schneider, R. Histone acylations and chromatin dynamics: concepts, challenges, and links to metabolism. EMBO Rep. 22, e52774 (2021).
Kumar, V., Thakur, J. K. & Prasad, M. Histone acetylation dynamics regulating plant development and stress responses. Cell. Mol. Life Sci. 78, 4467–4486 (2021).
Acknowledgements
We are grateful to S. Schulmeister and V. Schmidt for excellent technical support and to C. Fritz for helpful discussions (Cell Biology, University Erlangen-Nuremberg). We thank M. Weingartner (Molecular Plant Physiology, University of Hamburg) for mail1 seeds, E. Richards (Boyce Thompson Institute, Cornell University, Ithaca, New York) for crwn1-1/4-1 seeds, H. Puchta (Joseph Gottlieb Kölreuter Institute for Plant Sciences, Karlsruhe Institute of Technology) for plasmids enabling CRISPR–Cas9 and R. Stadler (Molecular Plant Physiology, University Erlangen-Nuremberg) for a binary plasmid containing a pWOX5:spGFP-HDEL expression cassette. Project funding was received from the German Research Foundation (DFG) (SFB796-52732026 start-up funding of collaborative project for M.S., H.S. and B.K.; SFB/TRR 241-375876048 subproject A06 for M.S.; STU 238/10-1-437201724 for M.S.; HE 2679/6-1 for C.H.), the Interdisciplinary Center for Clinical Research of the Clinical Center Erlangen (D34 for M.S.), the Ilse and Dr. Alexander Mayer Stiftung (2018-04-11 for B.K.) and the FAU UNIBUND (2017-12-12 for B.K.). Plant propagation and microscopy relied on DFG-sponsored major equipment (INST 90/1025-1 FUGG and INST 90/1074-1 FUGG for B.K.).
Author information
Authors and Affiliations
Contributions
The project was conceived by T.M.R., C.M., M.S. and B.K. Methodology was by T.M.R., C.S., C.M., H.S., C.H., M.S. and B.K. Formal analysis was conducted by T.M.R., C.S., C.M. and B.K. Investigation was undertaken by T.M.R., C.S., C.M., S.I. and H.S. Resources were obtained by T.M.R. and C.S. The original draft was written by T.M.R. and B.K. The final article was reviewed and edited by T.M.R., C.M., M.S., H.S., C.H. and B.K. Visualization was by T.M.R., C.S. and C.M. Supervision was carried out by C.H., M.S. and B.K. Project administration was by C.H., M.S. and B.K. Funding was acquired by H.S., C.H., M.S. and B.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Chang Liu, Ram Dixit and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
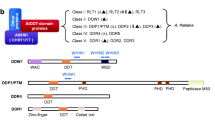
Extended Data Fig. 1 Phylogenetic relationships between GBP-related proteins identified in plants, animals, and protists.
The amino acid sequences of all plant (algae, mosses, ferns, seed plants) proteins containing adjacent GBP and GBP_C domains were retrieved from the Pfam database. Incomplete sequences apparently coding for truncated proteins were filtered out. The remaining sequences, together with the amino acid sequences of all human and of selected protist GBP-related proteins, were subjected to MUSCLE alignment using the Geneious 7.1.4 software package. A circular dendrogram was constructed using the Geneious tree builder function and the neighbour joining method. Human Atlastin2 (ATL2) was defined as outgroup.
Extended Data Fig. 2 AtGBPL3 expression pattern and intracellular distribution.
Analysis of transgenic Arabidopsis reporter lines, which expressed genomic AtGBPL3-GUS or AtGBPL3–GFP fusion constructs under the control of the AtGBPL3 promoter in the WT background. 7 or 8 independent AtGBPL3-GUS or AtGBPL3–GFP reporter lines were investigated, respectively, which all displayed identical patterns of GUS activity or GFP distribution. a-f, Histochemical visualization of GUS activity (indicated by blue staining) in AtGBPL3-GUS reporter plants. a, 6-day-old seedling incubated with GUS substrate for 24 h. Arrows: cotyledons; arrowhead: vasculature. b, Mature rosette leaf. Arrow: vasculature. c, Inflorescence with flowers at different developmental stages. Arrow: inflorescence stem. d, Young un-pollinated flower. Arrow: pistil; arrowhead: sepal. e, Pollinated flower. Arrow: petal; closed arrowhead: stamen filament; open arrowhead: nectaries. f, Base of a young silique (fruit). Arrow: nectaries. g-t, Confocal imaging of AtGBPL3–GFP reporter plants emitting green AtGBPL3–GFP fluorescence along with red chlorophyll or cell wall (r, pollen) autofluorescence indicated in magenta (j, k, maximum projections of serial confocal sections; t, image shown in s overlaid with a transmitted light reference image; all other images, single confocal sections). g–k, Low-magnification images of intact tissues showing AtGBPL3–GFP fluorescence confined to small puncta representing nuclei. l-q, High-magnification images showing AtGBPL3–GFP association with the nuclear envelope in each of the tissues depicted in g–k. g, m, Rosette leaf. h, n, Inflorescence stem. i, o, Petal of a mature flower. j-q, Anther (j (arrowhead), p) and stamen filament (j (arrow), q) of a mature flower. k, l, Seed coat enclosing mature embryo (outlined by dashed line). r–t, High-magnification images showing AtGBPL3–GFP accumulation in the vegetative nucleus (arrow) and in the two generative cells (arrowheads) contained in a pollen grain (r) or a pollen tube (s, t). Interestingly, AtGBPL3–GFP association with the nuclear envelope was not observed in these male reproductive cell types. Scale bars: a, c, 2 mm; b, h-k, 100 µm; d–g, 250 µm; l-r, 10 µm; s, t, 8 µm.
Extended Data Fig. 3 AtGBPL1 and AtGBPL2 are nuclear or tonoplast proteins, respectively.
a–c, e, Confocal imaging of AtGBPL1–GFP (a, b) or AtGBPL2–GFP (c, e) fluorescence (green) in roots of transgenic Arabidopsis reporter lines (WT background) containing genomic constructs (pAtGBPL1:AtGBPL1genomic-GFP or pAtGBPL2:AtGBPL2genomic-GFP, respectively), which confer expression of these fusion proteins under the control of the corresponding endogenous promoter (pAtGBPL1 or pAtGBPL2). a, c, Maximum projections of serial confocal optical sections showing the tips of main roots. Scale bars: 50 µm. b, e, Single confocal optical sections showing epidermal root tip cells at higher magnification. d, Single confocal optical section showing FM4-64 labelling (magenta, 60 minutes after the application of this membrane dye to the imaged root) of the tonoplast (membrane enclosing the vacuole) in the AtGBPL2–GFP expressing cells displayed in e. f, Overlay of the images shown in d and e. Scale bars: b, 5 µm; d–f, 10 µm. g, h, Confocal imaging of AtGBPL2–GFP fluorescence (green) and of chlorophyll autofluorescence (magenta) in Arabidopsis mesophyll protoplast transiently transformed with a construct conferring AtGBPL2–GFP expression under the control of the CaMV (cauliflower mosaic virus) 35 S promoter. g, Single confocal optical section. h, Maximum projection of serial confocal optical sections. n = 17 (a, b), 18 (c), 5 (d, e, f), 10 (g, h). Scale bars: 5 µm.
Extended Data Fig. 4 Molecular and genetic characterization of CRISPR/Cas9 AtGBPL3 knockout mutants.
Mutants were generated by transforming Arabidopsis plants with genes conferring expression of CAS9 and of a sgRNA targeting either the first (CRISPR 1) or the second (CRISPR 2) AtGBPL3 exon (see Fig. 2a). a, Application of genomic PCR to identify lines, which contained the CAS9 gene in the T1 generation, but had lost this gene in the T2 generation due to segregation. Negative control (WT): WT genomic DNA; positive control (Pos): plasmid containing the CAS9 gene. b, Sequencing the AtGBPL3 locus in independent lines (selected as described above: (a)) established that each of them was heterozygous for a 1-bp frameshift insertion in the first (CRISPR 1) or the second (CRISPR 2) exon. c, Analysis of the segregation of 1-bp insertions within in the AtGBPL3 gene among the offspring of self-fertilized heterozygous CRISPR 1 or CRISPR 2 plants based on modified Melt-MAMA-PCR. Heterozygous mutant offspring (he) was detected at a reduced rate (ratio he/WT: 0.6 (CRISPR 2) to 0.75 (CRISPR 1) instead of 2). No homozygous (ho) mutants could be identified (ho/WT: 0 instead of 1). d, e, Seed set of self-fertilized WT and heterozygous CRISPR 1 or CRISPR 2 plants. d, Mature siliques collected prior to dehiscence and cleared in 70% ethanol for several days to reveal seeds (arrows). Scale bar: 5 mm. e, Seed occupancy: number of seeds per silique normalized based on the average WT seed count (46.2 ± 4.0). n ≥ 18 siliques from ≥ 3 plants for each genotype. Boxplot: median (centerline), upper/lower quartiles (box), minimum/maximum (whiskers), datapoints (dots). Statistical analysis: Kruskal–Wallis with post hoc Wilcoxon test (two-sided). Letters indicate significant differences (p ≤ 0.05). f, Segregation of 1-bp insertions within the AtGBPL3 gene among offspring obtained after reciprocal backcrossing of heterozygous CRISPR 1 or CRISPR 2 plants to WT as determined by Melt-MAMA-PCR. Transmission of these insertions through the female (he x WT) and the male (WT x he) gametophyte was reduced to 10.2% (CRISPR 1: 8.7%, CRISPR 2: 11.5%) and 34.6% (CRISPR 1: 33.8%, CRISPR 2: 36.2%) instead of 50%, respectively.
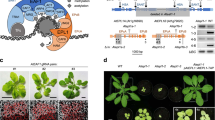
Extended Data Fig. 5 Unlike wild-type AtGBPL3, GTPase-dead AtGBPL3K83A does not complement PCD induction or reduced root growth displayed by gbpl3-5 seedlings.
a, Medial confocal optical sections through tips of main roots of 7-day-old seedlings with the indicated genotype (n = 30 (gbpl3-5), 27 (gbpl3-5comp2), 25 (gbpl3-5comp1) or 14 (gbpl3-5compK83A_1)), which show GFP fluorescence (green) along with propidium iodide (PI) staining of cell walls and of dead cells (magenta). PCD induction in gbpl3-5 roots (dead cells brightly labelled by PI) is complemented by AtGBPL3–GFP expression at endogenous (gbpl3-5comp2) or about 7x higher (gbpl3-5comp1) levels (see Fig. 2d). By contrast, tips of gbpl3-5 roots transformed with a genomic AtGBPL3K83A-GFP fusion construct (pAtGBPL3:AtGBPL3K83Agenomic-GFP), which expressed an AtGBPL3K83A-GFP fusion protein under the control of the AtGBPL3 promotor (pAtGBPL3) at endogenous level (gbpl3-5compK83A_1, see Fig. 4h), contained dead cells in outer cell layers and displayed a typical gbpl3-5 phenotype. A total of 10 independent lines expressing AtGBPL3K83A-GFP in the gbpl3-5 background were investigated, which all failed to display complementation. Scale bars: 50 µm. b, Quantitative analysis of the length of roots of seedlings with the indicated genotype at the different time points after germination (compare Fig. 3a). n ≥ 22 measurements/genotype and time point, 6 independent experiments. Boxplot: median (centerline), upper/lower quartiles (box), minimum/maximum (whiskers). Statistical analysis: ANOVA with post hoc Tukey´s tests (two-sided). Letters indicate significant differences (p ≤ 0.05).
Extended Data Fig. 6 AtGBPL3–GFP speckles formed during mitosis do not colocalize with markers for centromeres, the nucleolus, or peripheral nuclear pore components.
a–c, Confocal optical sections through cells co-expressing AtGBPL3–GFP (green) together with the indicated markers for different nuclear structures (magenta). Epidermal, cortical, or endodermal cells of gbpl3-5comp2 roots stably expressing AtGBPL3–GFP at endogenous level together with a centromere-associated histone (a, CENH3; n = 25 (Interphase), 22 (Mitosis)), a nucleolar protein (b, FIB2; n = 23 (Interphase), 11 (Mitosis)), or a peripheral nuclear pore component (c, NUP1; n = 29 (Interphase), 10 (Mitosis)) tagged with mCherry were imaged during interphase and mitosis (metaphase to late anaphase). Scale bars: 4 µm.
Extended Data Fig. 7 AtGBPL3 dynamics from late anaphase to cytokinesis revealed by time-lapse imaging of an individual mitotic root cell.
Confocal optical sections showing an individual dividing cortical cell of a gbpl3-5comp2 root stably expressing AtGBPL3–GFP at endogenous level (green) together with SUN1-mCherry (magenta), a marker for the nuclear envelope (n = 8). Images were recorded at the indicated time points during late anaphase (0 s), telophase (80 s, 160 s), and cytokinesis (235 s, 290 s). Arrowheads: AtGBPL3–GFP accumulation on the surface of emerging daughter nuclei in regions into which the SUN1-mCherry labelled reforming nuclear envelope is about to extend. Scale bar: 5 µm.
Extended Data Fig. 8 Validation of RNA-seq data and complementation of deregulated gene expression in gbpl3-5 seedlings by wild-type or GTPase-dead AtGBPL3.
a, b, Expression levels as determined by RNA-seq (FPKM: average fragments per kilo base per million mapped reads, 3 biological replicates) of AtGBPL3 (GBPL3), CRWN1, and CRWN4 (a), or of a selection of the 10 genes statistically most significantly up- or downregulated in gppl3-5 compared to WT (gppl3-5 DEGs) (b), in 4-day-old seedlings with the indicated genotype. Boxplots: median (centerline), upper/lower quartiles (box), minimum/maximum (whiskers), datapoints (dots). c, qRT-PCR analysis of relative expression levels of the gbpl3-5 DEGs displayed in b in 5-day-old seedlings with the indicated genotype. Data were generated according to the 2-ΔΔCT (Livak) method and normalized using UBI10 as reference gene (3 biological replicates, each technically replicated twice). Boxplots: median (centerline), upper/lower quartiles (box), minimum/maximum (whiskers). d, Heat map displaying relative expression levels (Z-score) of all gbpl3-5 DEGs (differentially expressed between gbpl3-5 and WT) in 4-day-old seedlings with die indicated genotype (three biological replicates/genotype). gbpl3-5comp2 and gbpl3-5compK83A_2 seedlings expressed at endogenous levels GFP fused to AtGBPL3 or to GTPase-dead AtGBPL3K83A, respectively. DEGs are sorted from most strongly upregulated (top) to most strongly downregulated (bottom) in gbpl3-5. e, Quantitative analysis of the complementation of deregulated gene expression in gbpl3-5 seedlings by AtGBPL3 (gbpl3-5comp2) or AtGBPL3K83A (gbpl3-5compK83A_2) expressed at endogenous levels. The percentage of all up- or downregulated gbpl3-5 DEGs is indicated, which fall into the following categories: 1) partially complemented (not differentially expressed between gbpl3-5compX (gbpl3-5comp2 or gbpl3-5compK83A_2) and WT & not differentially expressed between gbpl3-5compX and gbpl3-5), or 2) fully complemented (not differentially expressed between gbpl3-5compX and WT & differentially expressed between gbpl3-5compX and gbpl3-5).
Extended Data Fig. 9 Marker genes for developmental PCD or DNA repair are strongly deregulated in mail1, but not in gbpl3-5 seedlings.
Expression values as determined based on RNA-seq (FPKM) of Mail1 and of selected PCD or DNA repair marker genes in 4-day-old seedlings with the indicated genotype. ANAC046: At3g04060, EXI1: At2g14095, DMP-4: At4g18425, PARP2: At4g02390, RNS3: At1g26820, SMR1: At5g02420. Statistical analysis: pairwise comparison with WT (RNA-seq dataset 2; NCBI BioProject Accession PRJNA726378). * p ≤ 0.5, ** p ≤ 0.01, *** p ≤ 0.001.
Extended Data Fig. 10 Root tips of homozygous kaku4-2, crwn1-2, and crwn4-1 knockout mutants do not display local PCD or defects in tissue structure.
Medial confocal optical sections through tips of main roots of 7-day-old WT (n = 12), kaku4-2 (n = 10), crwn1-2 (n = 13), and crwn4-1 (n = 12) seedlings. Cell walls are stained with propidium iodide (magenta). Scale bars: 50 µm.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary Table 1
Key reagents and resources.
Supplementary Table 2
Oligonucleotides and Arabidopsis gene accession numbers.
Supplementary Video 1
Direct comparison of ER structure in WT and gbpl3-5 root cells. Three-dimensional (3D) rendering of stacks of at least 80 confocal optical sections through WT and gbpl3-5 root cells in 4-day-old seedlings stably expressing ER targeted spGFP-HDEL under the control of the pWOX5 promoter. The stacks used for 3D rendering are shown at the end of the video. Three-dimensional rendering was performed using the LAS X 3D software module (Leica). Scale bars, 25 µm (3D rendering), 20 µm (stacks).
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Reimann, T.M., Müdsam, C., Schachtler, C. et al. The large GTPase AtGBPL3 links nuclear envelope formation and morphogenesis to transcriptional repression. Nat. Plants 9, 766–784 (2023). https://doi.org/10.1038/s41477-023-01400-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01400-5
- Springer Nature Limited