Abstract
Many plant species monitor and respond to changes in day length (photoperiod) for aligning reproduction with a favourable season. Day length is measured in leaves and, when appropriate, leads to the production of floral stimuli called florigens that are transmitted to the shoot apical meristem to initiate inflorescence development1. Rice possesses two florigens encoded by HEADING DATE 3a (Hd3a) and RICE FLOWERING LOCUS T 1 (RFT1)2. Here we show that the arrival of Hd3a and RFT1 at the shoot apical meristem activates FLOWERING LOCUS T-LIKE 1 (FT-L1), encoding a florigen-like protein that shows features partially differentiating it from typical florigens. FT-L1 potentiates the effects of Hd3a and RFT1 during the conversion of the vegetative meristem into an inflorescence meristem and organizes panicle branching by imposing increasing determinacy to distal meristems. A module comprising Hd3a, RFT1 and FT-L1 thus enables the initiation and balanced progression of panicle development towards determinacy.




Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this Letter (and its supplementary files). The 3D protein structure of FT-L1 was modelled on the basis of available structural data on monomeric AtFT (PDB ID 6igh.1).
References
Andrés, F. & Coupland, G. The genetic basis of flowering responses to seasonal cues. Nat. Rev. Genet. 13, 627–639 (2012).
Komiya, R., Ikegami, A., Tamaki, S., Yokoi, S. & Shimamoto, K. Hd3a and RFT1 are essential for flowering in rice. Development 135, 767–774 (2008).
Tamaki, S., Matsuo, S., Wong, H. L., Yokoi, S. & Shimamoto, K. Hd3a protein is a mobile flowering signal in rice. Science 316, 1033–1036 (2007).
Komiya, R., Yokoi, S. & Shimamoto, K. A gene network for long-day flowering activates RFT1 encoding a mobile flowering signal in rice. Development 136, 3443–3450 (2009).
Wickland, D. & Hanzawa, Y. The FLOWERING LOCUS T/TERMINAL FLOWER1 gene family: functional evolution and molecular mechanisms. Mol. Plant 8, 983–997 (2015).
Taoka, K. et al. 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 476, 332–335 (2011).
Yano, M. et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12, 2473–2484 (2000).
Whipple, C. J. Grass inflorescence architecture and evolution: the origin of novel signaling centers. N. Phytol. 216, 367–372 (2017).
Gómez-Ariza, J. et al. A transcription factor coordinating internode elongation and photoperiodic signals in rice. Nat. Plants 5, 358–362 (2019).
Izawa, T. et al. Phytochrome mediates the external light signal to repress FT orthologs in photoperiodic flowering of rice. Genes Dev. 16, 2006–2020 (2002).
Brambilla, V. et al. Antagonistic transcription factor complexes modulate the floral transition in rice. Plant Cell 29, 2801–2816 (2017).
Goretti, D. et al. Transcriptional and post-transcriptional mechanisms limit Heading Date 1 (Hd1) function to adapt rice to high latitudes. PLoS Genet. 13, e1006530 (2017).
Tsuda, K., Ito, Y., Sato, Y. & Kurata, N. Positive autoregulation of a KNOX gene is essential for shoot apical meristem maintenance in rice. Plant Cell 23, 4368–4381 (2011).
Asano, T. et al. Rpp16 and Rpp17, from a common origin, have different protein characteristics but both genes are predominantly expressed in rice phloem tissues. Plant Cell Physiol. 43, 668–674 (2002).
Zhang, W. et al. Precise and heritable gene targeting in rice using a sequential transformation strategy. Cell Rep. Methods 3, 100389 (2023).
Gómez-Ariza, J. et al. Loss of floral repressor function adapts rice to higher latitudes in Europe. J. Exp. Bot. 66, 2027–2039 (2015).
Rigola, D. et al. High-throughput detection of induced mutations and natural variation using KeyPointTM technology. PLoS ONE 4, e4761 (2009).
Ho, W. W. H. & Weigel, D. Structural features determining flower-promoting activity of Arabidopsis FLOWERING LOCUS T. Plant Cell 26, 552–564 (2014).
Pires, D. E. V., Ascher, D. B. & Blundell, T. L. DUET: a server for predicting effects of mutations on protein stability using an integrated computational approach. Nucleic Acids Res. 42, W314-9 (2014).
Andrés, F. et al. SHORT VEGETATIVE PHASE reduces gibberellin biosynthesis at the Arabidopsis shoot apex to regulate the floral transition. Proc. Natl Acad. Sci. USA 111, E2760–E2769 (2014).
Zhu, W. et al. Rice SEPALLATA genes OsMADS5 and OsMADS34 cooperate to limit inflorescence branching by repressing the TERMINAL FLOWER1-like gene RCN4. N. Phytol. 233, 1682–1700 (2022).
Kobayashi, K. et al. Inflorescence meristem identity in rice is specified by overlapping functions of three AP1/FUL-like MADS box genes and PAP2, a SEPALLATA MADS box gene. Plant Cell 24, 1848–1859 (2012).
Prasad, K., Parameswaran, S. & Vijayraghavan, U. OsMADS1, a rice MADS-box factor, controls differentiation of specific cell types in the lemma and palea and is an early-acting regulator of inner floral organs. Plant J. 43, 915–928 (2005).
Lu, Z. et al. Genome-wide binding analysis of the transcription activator ideal plant architecture1 reveals a complex network regulating rice plant architecture. Plant Cell 25, 3743–3759 (2013).
Wang, L. et al. Coordinated regulation of vegetative and reproductive branching in rice. Proc. Natl Acad. Sci. USA 112, 15504–15509 (2015).
Jiao, Y. et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 42, 541–544 (2010).
Miura, K. et al. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 42, 545–549 (2010).
Wang, L. et al. Bract suppression regulated by the miR156/529–SPLs–NL1–PLA1 module is required for the transition from vegetative to reproductive branching in rice. Mol. Plant 14, 1168–1184 (2021).
Cerise, M. et al. OsFD4 promotes the rice floral transition via florigen activation complex formation in the shoot apical meristem. N. Phytol. 229, 429–443 (2021).
Kaur, A., Nijhawan, A., Yadav, M. & Khurana, J. P. OsbZIP62/OsFD7, a functional ortholog of FLOWERING LOCUS D, regulates floral transition and panicle development in rice. J. Exp. Bot. 72, 7826–7845 (2021).
Miki, D., Zhang, W., Zeng, W., Feng, Z. & Zhu, J. K. CRISPR/Cas9-mediated gene targeting in Arabidopsis using sequential transformation. Nat. Commun. 9, 1967 (2018).
Miki, D. et al. CRISPR/Cas9-based genome editing toolbox for Arabidopsis thaliana. Methods Mol. Biol. 2200, 121–146 (2021).
AL-Tam, F. et al. P-TRAP: a panicle trait phenotyping tool. BMC Plant Biol. 13, 122 (2013).
Lowder, L. G. et al. A CRISPR/Cas9 toolbox for multiplexed plant genome editing and transcriptional regulation. Plant Physiol. 169, 971–985 (2015).
Miao, J. et al. Targeted mutagenesis in rice using CRISPR–Cas system. Cell Res. 23, 1233–1236 (2013).
Sahoo, K. K., Tripathi, A. K., Pareek, A., Sopory, S. K. & Singla-Pareek, S. L. An improved protocol for efficient transformation and regeneration of diverse indica rice cultivars. Plant Methods 7, 49 (2011).
Toriba, T. et al. Suppression of leaf blade development by BLADE-ON-PETIOLE orthologs is a common strategy for underground rhizome growth. Curr. Biol. 30, 509–516.e3 (2020).
Betti, C. et al. Sequence-specific protein aggregation generates defined protein knockdowns in plants. Plant Physiol. 171, 773–787 (2016).
Abbruscato, P. et al. OsWRKY22, a monocot wrky gene, plays a role in the resistance response to blast. Mol. Plant Pathol. 13, 828–841 (2012).
Acknowledgements
We thank G. Valè, CREA-RIS and the staff of the Botanical Garden ‘Città Studi’ for technical support with the field experiments and the NO LIMITS facility for technical support with confocal microscopy. We thank P. E. Colleoni for support with the molecular studies. This work was supported by the Shanghai Science and Technology Innovation Plan (grant no. 20ZR1467000) to D. Miki, by an ERC Starting Grant (no. 260963) to F.F. and by the Italy–Japan bilateral collaboration programme funded by the Italian Ministry of Foreign Affairs and International Cooperation (Grande Rilevanza, grant no. PGR10047) to F.F.
Author information
Authors and Affiliations
Contributions
F.G., G.A.B., D. Martignago, Y.M., G.V., T.T., M.C., D.C., M.B., B.K., P.M., D.G. and V.B. performed the expression analyses, generated the transgenic plants, recorded the phenotypes and analysed the data. R.W., D.K. and D. Miki designed and produced the GFP knock-in plants. W.T. and M.K. produced the Volano mutant collection. M.M. performed the field experiments. G.C., V.B., D. Miki and J.K. conceived the experiments. F.F. and F.G. conceived the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Richard Immink and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9.
Supplementary Table 1
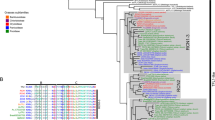
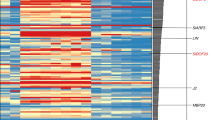
Pearson correlation coefficients among gene transcripts (FT-L1 versus 728 Affymetrix rice microarrays). Genes indicated in red have been demonstrated to regulate aspects of reproductive (mostly floral) development.
Supplementary Data 1
Statistical source data.
Supplementary Data 2
Unprocessed western blots.
Supplementary Data 3
Primers used in this study.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Giaume, F., Bono, G.A., Martignago, D. et al. Two florigens and a florigen-like protein form a triple regulatory module at the shoot apical meristem to promote reproductive transitions in rice. Nat. Plants 9, 525–534 (2023). https://doi.org/10.1038/s41477-023-01383-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01383-3
- Springer Nature Limited
This article is cited by
-
Functional divergence of FTL9 and FTL10 in flowering control in rice
BMC Genomics (2024)
-
QTL detection and candidate gene identification of qCTB1 for cold tolerance in the Yunnan plateau landrace rice
Molecular Breeding (2024)