Abstract
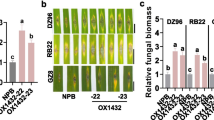
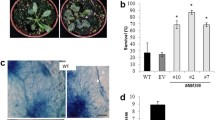
MicroRNA168 (miR168) is a key miRNA that targets Argonaute1 (AGO1), a major component of the RNA-induced silencing complex1,2. Previously, we reported that miR168 expression was responsive to infection by Magnaporthe oryzae, the causal agent of rice blast disease3. However, how miR168 regulates immunity to rice blast and whether it affects rice development remains unclear. Here, we report our discovery that the suppression of miR168 by a target mimic (MIM168) not only improves grain yield and shortens flowering time in rice but also enhances immunity to M. oryzae. These results were validated through repeated tests in rice fields in the absence and presence of rice blast pressure. We found that the miR168–AGO1 module regulates miR535 to improve yield by increasing panicle number, miR164 to reduce flowering time, and miR1320 and miR164 to enhance immunity. Our discovery demonstrates that changes in a single miRNA enhance the expression of multiple agronomically important traits.




Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this Article and in its Supplementary Information files. The data are available upon request. Source data are provided with this paper.
References
Li, Y. F. et al. Transcriptome-wide identification of microRNA targets in rice. Plant J. 62, 742–759 (2010).
Peters, L. & Meister, G. Argonaute proteins: mediators of RNA silencing. Mol. Cell 26, 611–623 (2007).
Li, Y. et al. Multiple rice microRNAs are involved in immunity against the blast fungus Magnaporthe oryzae. Plant Physiol. 164, 1077–1092 (2014).
Nelson, R., Wiesner-Hanks, T., Wisser, R. & Balint-Kurti, P. Navigating complexity to breed disease-resistant crops. Nat. Rev. Genet. 19, 21–33 (2018).
Bergelson, J. & Purrington, C. B. Surveying patterns in the cost of resistance in plants. Am. Nat. 148, 536–558 (1996).
Evans, J. R. Improving photosynthesis. Plant Physiol. 162, 1780–1793 (2013).
Rossi, M., Bermudez, L. & Carrari, F. Crop yield: challenges from a metabolic perspective. Curr. Opin. Plant Biol. 25, 79–89 (2015).
Xu, G. et al. uORF-mediated translation allows engineered plant disease resistance without fitness costs. Nature 545, 491–494 (2017).
Wang, J. et al. A single transcription factor promotes both yield and immunity in rice. Science 361, 1026–1028 (2018).
Fang, J. et al. Ef-cd locus shortens rice maturity duration without yield penalty. Proc. Natl Acad. Sci. USA 116, 18717–18722 (2019).
Tang, J. & Chu, C. MicroRNAs in crop improvement: fine-tuners for complex traits. Nat. Plants 3, 17077 (2017).
Chandran, V. et al. miR396-OsGRFs module balances growth and rice blast disease-resistance. Front. Plant Sci. 9, 1999 (2019).
Mallory, A. C., Elmayan, T. & Vaucheret, H. MicroRNA maturation and action—the expanding roles of ARGONAUTEs. Curr. Opin. Plant Biol. 11, 560–566 (2008).
Li, Y. et al. Identification of microRNAs involved in pathogen-associated molecular pattern-triggered plant innate immunity. Plant Physiol. 152, 2222–2231 (2010).
Sun, M. Z. et al. The multiple roles of OsmiR535 in modulating plant height, panicle branching and grain shape. Plant Sci. 283, 60–69 (2019).
Zhao, Y. F. et al. miR1432-OsACOT (acyl-CoA thioesterase) module determines grain yield via enhancing grain filling rate in rice. Plant Biotechnol. J. 17, 712–723 (2019).
Yang, R. et al. Fine-tuning of MiR528 accumulation modulates flowering time in rice. Mol Plant. 12, 1103–1113 (2019).
Salvador-Guirao, R., Hsing, Y. I. & San Segundo, B. The polycistronic miR166k-166h positively regulates rice immunity via post-transcriptional control of EIN2. Front. Plant Sci. 9, 337 (2018).
Zhao, Z. X. et al. Osa-miR167d facilitates infection of Magnaporthe oryzae in rice. J. Integr. Plant Biol. 62, 702–715 (2019).
Li, Y. et al. Osa-miR169 negatively regulates rice immunity against the blast fungus Magnaporthe oryzae. Front. Plant Sci. 8, 2 (2017).
Zhang, X. et al. Magnaporthe oryzae induces the expression of a microRNA to suppress the immune response in rice. Plant Physiol. 177, 352–368 (2018).
Li, Y. et al. Osa-miR398b boosts H2O2 production and rice blast disease-resistance via multiple superoxide dismutases. N. Phytol. 222, 1507–1522 (2019).
Xiao, Z. Y. et al. MiR444b.2 regulates resistance to Magnaporthe oryzae and tillering in rice. Acta Phytopathol. Sinica 47, 511–522 (2017).
Wang, Z. Y. et al. Osa-miR164a targets OsNAC60 and negatively regulates rice immunity against the blast fungus Magnaporthe oryzae. Plant J. 95, 584–597 (2018).
Jiao, Y. et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 42, 541–544 (2010).
Wang, L. et al. Coordinated regulation of vegetative and reproductive branching in rice. Proc. Natl Acad. Sci. USA 112, 15504–15509 (2015).
Mannai, Y. E., Akabane, K., Hiratsu, K., Satoh-Nagasawa, N. & Wabiko, W. The NAC transcription factor gene OsY37 (ONAC011) promotes leaf senescence and accelerates heading time in rice. Int. J. Mol. Sci. 18, 2165–2185 (2017).
Lin, Z. Z., Jiang, W. W., Wang, J. L. & Lei, C. L. Research and utilization of universally susceptible property of japonica rice variety Lijiangxintuanheigu. Sci. Agricultura Sinica 34, 116–117 (2001).
Tsumematsu, H. et al. Development of monogenic lines of rice for blast resistance. Breed. Sci. 50, 229–234 (2000).
Kanzaki, H. et al. Arms race co-evolution of Magnaporthe oryzae AVR-Pik and rice Pik genes driven by their physical interactions. Plant J. 72, 894–907 (2012).
Franco-Zorrilla, J. M. et al. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 39, 1033–1037 (2007).
Kankanala, P., Czymmek, K. & Valent, B. Roles for rice membrane dynamics and plasmodesmata during biotrophic invasion by the blast fungus. Plant Cell 19, 706–724 (2007).
Kong, L. A. et al. Different chitin synthase genes are required for various developmental and plant infection processes in the rice blast fungus Magnaporthe oryzae. PLoS Pathog. 8, e1002526 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Friedlander, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 40, 37–52 (2012).
Acknowledgements
We thank C.-L. Lei (Institute of Crop Science, Chinese Academy of Agricultural Sciences) for providing IRBLkm-Ts and J.-M. Zhou (Institute of Genetics and Developmental Biology, Chinese Academy of Sciences) for suggestions on writing the manuscript. This work was supported by grants from the Sichuan Applied Fundamental Research Foundation (no. 2020YJ0332) and National Natural Science Foundation of China (nos. U19A2033, 31672090 and 31430072) to W.-M.W. and by grants from the National Institutes of Health (no. GM59962), USDA NIFA (no. 2017-67013-26590) and the Joint BioEnergy Institute funded by the US DOE (no. DE-AC02-05CH11231) to P.C.R. and M.C.
Author information
Authors and Affiliations
Contributions
Yan Li and W.-M.W. conceived the project. H.W., Y.Z., L.-L.Z., J.-H.L., W.-Q.D., Z.-R.Y., S.-Z.Y. and Z.-X.Z. carried out the experiments. X.-P.L. and X.-C.M. performed the transgenic plant generation and analysis. J.-W.Z. and M.P. conducted the field trials. M.C., J.F. and X.-J.W. analysed the data. Yan Li, M.C., P.C.R. and W.-M.W. wrote the paper. X.-W.C., W.-T.L., J.W., M.H., Y.-Y.H., S.-G.L., P.L. and Yi Li discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Yong-Hwan Lee, Shaoqing Li and Guo-Liang Wang for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures and methods.
Supplementary Tables
Supplementary Tables 1–5.
Supplementary Data 1
Statistical source data.
Supplementary Data 2
Statistical source data.
Supplementary Data 3
Statistical source data.
Supplementary Data 4
Statistical source data.
Supplementary Data 5
Statistical source data.
Supplementary Data 6
Statistical source data.
Supplementary Data 7
Statistical source data.
Supplementary Data 8
Statistical source data.
Supplementary Data 9
Unprocessed western blots.
Supplementary Data 10
Unprocessed western blots.
Supplementary Data 11
Unprocessed northern blots.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Rights and permissions
About this article
Cite this article
Wang, H., Li, Y., Chern, M. et al. Suppression of rice miR168 improves yield, flowering time and immunity. Nat. Plants 7, 129–136 (2021). https://doi.org/10.1038/s41477-021-00852-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00852-x
- Springer Nature Limited
This article is cited by
-
miRNAs are involved in regulating the formation of recovery tissues in virus infected Nicotiana tabacum
Molecular Genetics and Genomics (2024)
-
The role of microRNAs in NBS-LRR gene expression and its implications for plant immunity and crop development
Transgenic Research (2024)
-
MicroRNAs Involved in Nutritional Regulation During Plant–Microbe Symbiotic and Pathogenic Interactions with Rice as a Model
Molecular Biotechnology (2024)
-
Fine-tuning of IPA1 transactivation activity by E3 ligase IPI7-mediated non-proteolytic K29-ubiquitination during Magnaporthe oryzae infection
Nature Communications (2024)
-
ARGONAUTE 1: a node coordinating plant disease resistance with growth and development
Phytopathology Research (2023)