Abstract
The ribosome utilizes hydrogen bonding between mRNA codons and aminoacyl-tRNAs to ensure rapid and accurate protein production. Chemical modification of mRNA nucleobases can adjust the strength and pattern of this hydrogen bonding to alter protein synthesis. We investigate how the N1-methylpseudouridine (m1Ψ) modification, commonly incorporated into therapeutic and vaccine mRNA sequences, influences the speed and fidelity of translation. We find that m1Ψ does not substantially change the rate constants for amino acid addition by cognate tRNAs or termination by release factors. However, we also find that m1Ψ can subtly modulate the fidelity of amino acid incorporation in a codon-position and tRNA dependent manner in vitro and in human cells. Our computational modeling shows that altered energetics of mRNA:tRNA interactions largely account for the context dependence of the low levels of miscoding we observe on Ψ and m1Ψ containing codons. The outcome of translation on modified mRNA bases is thus governed by the sequence context in which they occur.
Similar content being viewed by others
Introduction
Chemically modified nucleosides are present in all organisms, often playing essential roles in key cellular processes including splicing and translation1,2,3. Defects in ribosomal RNA (rRNA) and transfer RNA (tRNA) modifying enzymes are linked to a host of deleterious human health outcomes, illustrating the importance of RNA modifications in protein synthesis4,5,6,7. There are over 150 unique modifications reported in RNAs that range in size and complexity from isomerized or saturated nucleosides (e.g. pseudouridine and dihydrouridine) to large, chemically diverse functional groups (e.g. NAD+, N6-threonylcarbamoyladenosine, glycan and farnesyl)8,9,10,11. RNA modifications have been widely studied for almost three quarters of a century and until recently were thought to be almost exclusively incorporated into non-coding RNA species (ncRNAs). However, the transcriptome wide mapping of 13 RNA modifications revealed that protein coding messenger RNAs (mRNAs) can also contain modifications at thousands of sites12,13. This discovery raised the possibility that mRNA modifications might play a previously underappreciated role in post-transcriptionally regulating gene expression.
The majority of enzymes that modify mRNAs also catalyze their incorporation into ncRNAs central to protein synthesis8. Like their protein post-translational counterparts, mRNA post-transcriptional modifications are generally present at sub-stoichiometric levels, with transcripts existing in a mixed population of modified and unmodified states14,15,16,17. Together these circumstances make ascertaining the impact of mRNA modifications on translation challenging. In cells, any changes to protein output observed when RNA modifying enzymes are removed cannot be directly attributed to alterations in a particular mRNA’s modification status. Therefore, studies using reconstituted translation systems, where it is possible to uniformly change the modification status of an mRNA without impacting that of ncRNA, have been particularly useful for assessing the consequences of mRNA modifications on translation elongation18. Although these reconstituted systems are typically bacterial in origin, the core mechanism of elongation phase of translation is fairly well conserved between bacteria and eukaryotes19,20,21. Initial studies reveal that modifications commonly slow the ribosome, though some only do so only in particular mRNA sequence contexts18. Additionally, a subset of mRNA modifications, including pseudouridine (Ψ) and inosine (I), also modulate codon decoding by the ribosome to varying degrees22,23,24,25. These findings suggest that there is a broad range of possible consequences when the ribosome encounters an mRNA modification. Developing a framework for understanding how individual modifications impact translation in differing sequence contexts will be crucial as researchers seek to uncover which of the thousands of chemically modified positions reported in mRNA codons are the most likely to have consequences for protein synthesis in cells.
In addition to being present in naturally occurring RNA molecules, modifications are also heavily incorporated in RNA-based therapeutics and mRNA vaccines26,27,28,29. Indeed, the mRNA transcripts that form the basis of many currently available COVID-19 mRNA vaccines substitute every uridine nucleoside with N1-methylpseudouridine (m1Ψ)30. The addition of m1Ψ limits the cellular innate immune response to dramatically stabilize the mRNA transcript, and ultimately increase the amount of protein synthesized31,32,33,34. Recent studies in a lysate-based translation system suggest that m1Ψ slows the ribosome in a manner that can be alleviated by the addition of membranes35. However, there is limited information available directly evaluating how m1Ψ can influence the rate and fidelity of amino acid addition, apart from studies establishing the translational accuracy of the carefully engineered COVID mRNA vaccine constructs36,37. This is an important question to ask because m1Ψ shares much of its structure with pseudouridine (Ψ) (Fig. 1A), a modification that has been shown to change translation speed and tRNA selection22,25,38,39,40. Furthermore, recent findings indicate that m1Ψ promote low levels of +1 ribosomal frameshifting on the COVID mRNA vaccine sequence, likely promoted by slowed-down translation32,36,37. Even subtle changes in translation rates or fidelity have the potential to impact protein folding or function41,42,43,44. Therefore, establishing if there are situations in which m1Ψ can alter translation will be critical for the continued design of mRNA-based therapeutics and vaccines in addition to understanding how different types of chemical moieties and contexts impact translation.
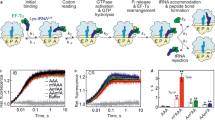
A The chemical structures of the nucleobases we investigated. B The formation of fMet-Phe (MF) dipeptide as a function of time by E. coli ribosomes containing 35S-fMet-tRNAfMet bound to an AUG start codon in the P site, and unmodified (black circles – UUU, n = 23 reactions divided between 2 experiments) or modified (blue squares - m1ΨUU (n = 36 reactions conducted in 3 experiments), green diamonds - Um1ΨU (n = 45 reactions over 4 experiments), red triangles - UUm1Ψ (n = 44 reactions/3 experiments)) codons in the A site. C The K½ curve for RF1. Fitted kobs values (n = 12 independent measurements of kobs) for RF1-catalyzed 35S-fMet release on UAA (circles) or m1ΨAA (squares) are displayed as a function of [RF1]. D The K½ curve for RF2. Fitted kobs values (n = 11 independent measurements of kobs) for RF2-catalyzed 35S-fMet release on an UAA (black circles) or m1ΨAA (blue squares) are displayed as a function of [RF2]. Error bars in (C) and (D) indicate the standard error of the fitted value of kobs. Source data are provided as Source Data file.
To ascertain the molecular level consequences of m1Ψ codon modifications on ribosome decoding, we compared the translation of unmodified and m1Ψ- modified codons in both a fully reconstituted bacterial in vitro translation system and HEK293 cells. These studies reveal that, in contrast to Ψ, the presence of a single m1Ψ does not substantially reduce the rate constant for cognate amino acid addition. However, m1Ψ does influence the accuracy of amino acid addition. We demonstrate that Ψ and m1Ψ can both impede and enhance alternative tRNA selection depending on the surrounding sequence context and identity of the tRNA. Comparison of how Ψ and m1Ψ modifications affect translation reveal that uridine base isomerization and methylation each contribute to the ability of m1Ψ to perturb mRNA decoding. Computational modeling of tRNAIle,UAG bound to modified and unmodified Phe (UUU) codons in the context of the A site suggests that changes in the energies of mRNA:tRNA interactions likely account for the context dependent effects of Ψ and m1Ψ we observe. These findings demonstrate that Ψ and m1Ψ modulate ribosome decoding and have the potential to impact the speed and accuracy of protein production from both native and therapeutic mRNA sequences.
Results
m1Ψ modestly impacts the rate constant for Phe addition in some sequence contexts
We used a fully reconstituted E. coli in vitro translation system to evaluate the consequences of incorporating m1Ψ into mRNA codons on translation elongation and termination. In contrast to reporter-based studies in cells and lysates, the in vitro system we implemented is not influenced by extra-translational factors that can change observed protein levels (e.g. RNases and proteases) and allows us to directly and quantitatively examine individual steps along the translation pathway45. This E. coli translation system has long been used to study translation elongation because tRNA binding sites and ribosome peptidyl-transfer center are highly conserved between eukaryotic and bacterial ribosomes19,20,21.
The rate constants for amino acid addition were measured on unmodified (UUU) and m1Ψ modified (m1ΨUU, Um1ΨU, and UUm1Ψ) phenylalanine (Phe) codons (Fig. 1B). We chose to first evaluate amino acid incorporation rates on a UUU codon because the kinetics of Phe addition on UUU is well established, and UUU codons are present in the Pfizer/BioNTech mRNA COVID-19 vaccine sequence30. Our translation reactions were initiated by mixing E. coli 70 S ribosome initiation complexes (ICs; 35S-labeled formylmethionine-tRNAfMet [35S-fMet] bound to an AUG in the P site and Phe codon in the A site) with an excess of ternary complexes (TCs; Phe-tRNAPhe•EF-Tu•GTP). Reactions were quenched at select time points, and the unreacted 35S-fMet and 35S-fMet-Phe products visualized by electrophoretic TLC (eTLC) (Supplementary Fig. 2A). These studies reveal that cognate Phe incorporation on m1Ψ modified codons is largely unchanged, though we observe a very slight (2 ± 0.3 -fold) increase in the rate constant for Phe addition when m1Ψ is in the third position in the codon (Fig. 1B, Supplementary Fig. 2B, Supplementary Table 1). Similarly, the inclusion of m1Ψ across all codon positions (m1Ψm1Ψm1Ψ) results in no observable defect in Phe addition.
All three stop codons begin with uridine (UAA, UAG, UGA) ensuring that modified stop codons will be present in synthetic mRNA-based vaccines and therapeutics. We evaluated the ability of bacterial class I release factors (RF1 and RF2) to hydrolyze peptidyl-tRNA bonds and terminate translation on m1Ψ modified stop codons. To accomplish this, we reacted termination complexes (E. coli 70 S ribosomes with 35S-fMet bound to an AUG in the P site, and a universal stop codon positioned in the A site (UAA, m1ΨAA)) with varying concentrations of RF1 and RF2 (0.05-10 μM). The reactions were quenched at a range of time points and 35S-fMet hydrolyzed by RFs was detected on an eTLC (Supplementary Figs. 3 and 4). The observed rate constants for peptide release (kobs,max) on UAA and m1ΨAA codons are equivalent (~0.1 s−1) and comparable previously published termination rates on an unmodified UAA codon (Supplementary Tables 2 and 3)46,47. Effects on K1/2 were minimal (Fig. 1C, D, Supplementary Tables 2, 3) and not statistically significant. In sum, we do not expect m1Ψ to impede translation termination in cells unless the concentration of release factors becomes severely limited, or, the termination codon is in a particularly poor sequence context48,49. This supposition is supported by numerous observations that reporter peptides and therapeutic RNA sequences generated from fully m1Ψ-substituted mRNAs yield protein products of the expected length31,36,50.
m1Ψ influences aminoacyl-tRNA selection by the ribosome in a context dependent manner
Chemical modifications to mRNA nucleobases can change the propensity of the ribosome to incorporate alternative amino acids into a growing polypeptide chain18. In comparison to uridine, m1Ψ possesses a repositioned, methylated nitrogen in its pyrimidine ring (Fig. 1A). These two changes provide m1Ψ the opportunity to alter the conformational fit of an mRNA in the ribosome, and potentially change the strength and pattern of mRNA:tRNA interactions. Consistent with this idea, Ψ, which shares a repositioned nitrogen with m1Ψ, was previously shown to enhance the reaction of some non-cognate tRNAs on UUU codons in vitro and in HEK293 cells22. To begin examining if m1Ψ similarly influences aminoacyl-tRNA (aa-tRNA) selection, we qualitatively evaluated the impact of m1Ψ on the propensity of Phe codons to react with tRNAs beyond tRNAPhe. In these assays, 70 S E. coli initiation complexes were generated with unmodified (UUU) and modified (m1ΨUU, Um1ΨU, and UUm1Ψ) codons in the A site, and reacted with EF-Tu containing ternary complexes formed using a mixture of total tRNA aminoacylated either by reacting all 20 amino acids with S100 (total aa-tRNAaa), or with a single amino acid and aminoacyl tRNA synthetase (Phe-tRNAPhe, Ser-tRNASer, Leu-tRNALeu, Ile-tRNAIle and Val-tRNAVal)45. Relative to UUU, we observed that m1ΨUU reacts more robustly with total aa-tRNAaa, and promotes the production of higher-levels of miscoded Met-Ile (MI) and Met-Val (MV) peptides (Fig. 2A). Um1ΨU and UUm1Ψ codons also exhibit different levels of reactivity with multiple tRNAs than an unmodified UUU (Supplementary Fig. 5).
A Representative eTLC displaying dipeptide products from translation reactions performed with 70 S initiation complexes (ICs) containing an unmodified UUU or m1ΨUU codon in the A site and total E. coli tRNA aminoacylated with a single amino acid (aa-TC). Relative to ICs containing a UUU codon in the A site, higher levels of miscoded MI and MV dipeptide products and lower levels of MS were generated from m1ΨUU containing ICs. B Summary of amino acid substitutions observed by mass spectrometry in a luciferase peptide incorporated on m1Ψ-containing mRNAs translated in 293H cells (Supplementary Table 8). Source data are provided as Source Data file.
Using the information obtained from our qualitative screens, we selected to further study three tRNAs that either appeared to react more (tRNAIle(GAU)), less (tRNALeu(CAG)), or to the same extent (tRNASer(UGA)) on unmodified and m1Ψ modified codons (Fig. 2A). We performed multiple-turnover tRNA selection assays to quantitatively characterize differences in tRNA selection for these three tRNAs and the three modified codons. In these assays, saturating concentrations of ternary complex were reacted with initiation complex. Energy regeneration mix and EF-Ts were included to catalyze re-formation of ternary complex after rejection and maintain a saturating concentration of ternary complex51. The rate constants we observed for dipeptide formation reflects not only the rate constant for the rate-limiting step in peptide bond formation, but also the many other steps involved in formation and breakdown of the post-accommodation elongating ribosome complex. As such, these measurements should not directly be compared to the single-turnover rate constants measured for phenylalanine incorporation in Fig. 1. Our approach contrasts with that of a recent study which uses neither saturating aa-tRNA nor energy regeneration to maintain saturation, resulting in second-order kinetics36.
We find that tRNA identity and the position of m1Ψ within a codon influence the rate constants for amino acid substitution (Fig. 3, Supplementary Table 4). For example, m1Ψ substitution at the first position in the Phe codon (m1ΨUU) does not change the kobs values for Leu or Ser incorporation but increases the rate constant for Ile addition by 2.2 ± 0.4-fold (Figs. 3D and 4A, Supplementary Table 4). This differs markedly from what we observed on Um1ΨU-modified codons, which have a much larger effect on tRNA selection. The kobs values are significantly reduced for Ile (10 ± 2-fold) and Leu (4 ± 1-fold) addition, while the rate constant for Ser mis-incorporation is conversely increased by 3.5 ± 0.4-fold (Fig. 3D, Supplementary Table 4). Substitution at the wobble position (UUm1Ψ) generally had modest impacts on the rate constants for amino acid incorporation; decreasing the kobs for Leu addition (2 ± 0.3-fold), while not impacting the kobs values for Ile and Ser addition (Fig. 3D, Supplementary Table 4). The findings of our kinetic studies are generally consistent with our initial qualitative assays (Figs. 2, 3 and Supplementary Fig. 5), and together indicate that m1Ψ codon modifications can both increase and decrease the ability of Phe UUU codons to react with a variety of tRNAs in the A site.
Plots of dipeptide formation as a function of time (seconds). Miscoding reactions were performed with E. coli ICs containing 35S-fMet-tRNAfMet bound to an AUG start codon in the P site, and unmodified (black circles-UUU) or modified (blue squares-Ψ/m1ΨUU, green diamonds-UΨ/m1ΨU, red triangles-UUΨ/m1Ψ) codons in the A site. Purified ICs were reacted with TCs containing (A) Ile-tRNAIle(GAU), (B) Leu-tRNALeu(CAG)), or (C) Ser-tRNASer(UGA). At least 30 independent time points were collected for each IC in experiments conducted over three or more separate days. D The fitted rate constants (kobs) for isoleucine, leucine, and serine misincorporation on Ψ- and m1Ψ- modified codons relative to the fitted rate constants for isoleucine, leucine, and serine misincorporation on a UUU codon. Each ratio has an n of 1 (since each is computed by dividing kobs,modified/kobs,UUU) and error bars were calculated by propagating the 95% confidence intervals of each fitted rate constant. Source data are provided as Source Data file.
A Summary of data in Supplementary Tables 4 and 5 displaying how a Ψ and m1Ψ impact the rate constants for the reaction of near-cognate tRNAs. B, C Summary of MM data. Gray bars reflect the change in energy for interactions between a modified mRNA position (Ψ/m1Ψ) and the base paring (B) tRNAPhe(GAA) or (C) tRNAIle(GAU) residue (ΔEbp) relative to an unmodified mRNA U. Black bars reflect the total change in energy (ΔEΣΨ/m1Ψ:X-Y) for interactions between a modified mRNA position (Ψ/m1Ψ) and three (B) tRNAPhe(GAA) or (C) tRNAIle(GAU) residues (the base paired nucleotide, and nucleotides 5′ and 3′ the bp) relative to an unmodified mRNA U. D Molecular model of an unmodified sequence coding for a Phe UUU codon and a tRNAIle(GAU). The hypermodification t6A37 is also shown on the tRNA.
Uridine isomerization contributes to observed changes in amino acid substitution on m1Ψ containing codons
To determine how the C5-glycoside uridine isomerization and N1-methylation individually impact the ability of m1Ψ to modulate amino acid incorporation, we measured the rate constants for Leu, Ile and Ser mis-incorporation on Ψ modified Phe codons (ΨUU, UΨU and UUΨ). Ψ was selected for study because it contains the same isomerized uridine base as m1Ψ, but lacks the methylation at position N1 (Fig. 1A). The impact of Ψ on Ile and Leu insertion was similar to what we observed when m1Ψ is present in codons (Fig. 3, Supplementary Table 5). For example, the rate constant for Ile is significantly decreased (10 ± 2 -fold) when Ψ is incorporated at the second codon position (UΨU), while Leu is added more slowly when Ψ is at any position in the codon (Fig. 3, Supplementary Table 5). In contrast to m1Ψ, Ser incorporation occurs with a 4 ± 0.5-fold faster rate constants when Ψ is at the first and second positions in the codon and is not influenced by Ψ substitution at the wobble position (UUΨ). These observations indicate that uridine isomerization largely accounts for changes in how the ribosome decodes some tRNAs for m1Ψ−substituted mRNAs. However, comparing the reactivity profiles of Ψ and m1ΨUU reveals the N1 methyl group can suppress the effect of uridine isomerization on the rate constants for amino acid misincorporation by other tRNAs (e.g. ΨUU vs m1ΨUU reacting with tRNASer (UGA)) (Fig. 3D).
Amino acid substitution in HEK293 cells increases on some m1Ψ containing codons
Our in vitro translation data reveal that m1Ψ and Ψ affect tRNA selection by E. coli ribosomes in different ways depending on their sequence context. We next asked if m1Ψ has similar effects on amino acid selection in eukaryotic cells. To approach this question, we transfected luciferase encoding mRNAs transcribed in vitro with either UTP or m1ΨTP into HEK293 cells where they were translated. The base composition of the unmodified and modified mRNAs was assessed by liquid chromatography-mass spectrometry (LC-MS) and is consistent between the unmodified and modified mRNA species we generated (Supplementary Fig. 7). We observed increased levels of luciferase protein expression in the m1Ψ-substituted mRNAs, consistent with previous reports (Supplementary Figs. 8 and 9)22. The luciferase proteins generated from both unsubstituted and m1Ψ-substituted mRNAs were purified and analyzed by mass spectrometry to identify amino acid substitutions.
Our mass spectrometry data analyses focused on a specific luciferase peptide with favorable ionization characteristics (Fig. 2B)22. ~1% of the amino acids in this peptide were substituted. This is a >20-fold increase over the level of amino acid substitution we previously observed for peptides generated from an unmodified version of the same luciferase reporter (<0.05% of their amino acids substituted)22. Nonetheless, the levels of misincorporation are still quite low. m1Ψ-mediated substitutions were observed on multiple codons (e.g. UUU, UAU), though we did not observe amino acid substitutions above background for every U-containing codon (e.g. UGG) (Supplementary Table 6). The highest levels of substitution were observed on the two Phe codons (UUU and UUC). Similar to our in vitro observations, serine and isoleucine/leucine amino acid substitutions were detected on both Phe codons with a >6-fold increased frequency of substitution over peptide from unmodified mRNA (Fig. 2B, Supplementary Tables 6–8)22. Isoleucine and leucine have the same mass and therefore cannot be distinguished in this assay. We also noted that the likelihood of substitutions occurring was not uniform across m1Ψ containing codons. The levels of miscoding that we detect are consistent with what we would predict from our in vitro studies, as is the heterogeneity of amino acid substitution on m1Ψ-modified codons (Figs. 2, 3). Furthermore, the lack of uniformity in amino acid substitution was also seen in our previous findings indicating that Ψ also increases the levels of amino acid misincorporation in the same luciferase reporter peptide22. Our results collectively suggest that the extent of misincorporation on any codon containing a C5-glyocside uridine isomer is low and strongly depends on both the codon and sequence context in which the modification is present.
Impact of m1Ψ and Ψ on duplex melting temperatures is context dependent
Ψ and m1Ψ are known to affect the melting temperature of RNA duplexes31,52,53. We sought to better understand the role of sequence context in our kinetic data by assaying the melting temperature of short (7–8 basepairs) duplexes containing U, Ψ and m1Ψ, modeling our approach after that used in a recent study36. The Tms we observed (~23 °C) were far below those reported previously (~78 °C) on similar sequences36, and several mismatch-containing duplexes were too unstable to allow a Tm determination. The Tm values that we measure are in line with Tms predicted by programs for estimating the physical properties of RNAs such as OligoCalc54. The unusually high Tm values (>70 °C) previously reported likely reflect hydrolysis of RNA in the presence of magnesium at elevated temperatures, which also results in increased absorbance. Ultimately, we successfully assayed U, Ψ, and m1Ψ in the context of a perfect duplex, a duplex containing a mismatch, and in a duplex adjacent to a wobble base pair (Supplementary Figs. 10–12). Pseudouridine increased the melting temperature of all duplexes, as expected, though the extent to which Ψ substitutions increased Tm values varied with the surrounding sequence context (ΔTm = 1.5–4 °C). The effects of m1Ψ varied even more dramatically depending on the sequence context, and sometimes differed from that of Ψ. For example, in the CXU context, pseudouridine stabilized the duplex, while m1Ψ did not. In the UXU context, with a 3′ U:G wobble pair, Ψ and m1Ψ provided equal stabilization.
Ψ-derived modifications change the energetics of mRNA:tRNA nucleoside interactions in the ribosome A site
We sought to understand why Ψ and m1Ψ modifications alter the interactions between mRNAs and tRNAs during translation in a position dependent manner (Figs. 1–4A). Although our melting temperature studies were generally consistent with our translation assays and suggest that modification-induced alterations base pairings fluctuate with sequence context, the changes in Tm measured between oligonucleotides outside of the ribosome structural context did not satisfactorily explain alterations in tRNA selection on Ψ and m1Ψ-containing codons that we observed. Therefore, we turned to molecular modeling (MM) and quantum mechanical calculations to examine unmodified and Ψ-, m1Ψ- and 3-methylpseudouridine (m3Ψ) modified UUU mRNA codons interacting with a tRNAPhe(AAG) and tRNAIle(UAG) in a portion of the ribosome A site (Figs. 1A, 4). Although we did not investigate the translation of m3Ψ-containing codons, we included m3Ψ in our computational studies as a positive control for a modification that should severely perturb mRNA:tRNA interactions. Methylation at the uridine N3 position removes the ability of uridine to donate a hydrogen bond and will limit tRNA binding. Our MM studies were conducted using models developed based on previously published crystal structure of the 70 S Thermus thermophilus ribosome with tRNAPhe bound on a ΨUU codon22. The MM investigations were designed to examine how the location of a modification impacts the pairwise tRNA:mRNA interaction energies. Each modification was modeled in either the first, second, or third codon position, and the energetics of tRNAPhe/Ile:mRNA interactions on modified codons were compared to those on an unmodified UUU codon (ΔE) (Fig. 4, Supplementary Data 1 and 2). Only ΔE values with magnitudes ≳1 kcal/mol are considered large enough to potentially influence mRNA:tRNA interactions.
We computed pairwise energy differences between a modified base and an unmodified base at the same position within a UUU codon. Both the change in base-pairing energy between the modified base and its partner in the tRNA were examined, as well as changes in energy derived from interactions between the modified base and neighboring tRNA bases. In general, the trends in modeled energy differences recapitulated trends in reaction free energy shown by changes in the observed rate constants for the reaction (Supplementary Fig. 13). In particular, the sum of changes in modeled energies between the modified base, its tRNA pair, and the upstream and downstream tRNA base (ΔEΣΨ/m1Ψ:tRNA-1,0,+1) were well correlated, while the individual pairwise interaction energies (Ebp) were less well correlated. The one exception was the summed energy change for m1Ψ in the second position, which predicted an increased kobs that was not borne out in the experimental data.
The in vitro translation, Tm, and modeling data all support the idea that pseudouridine-derived modifications affect the energetic landscape for codon interactions. As an additional check on our modeling, we calculated energy differences for codons containing m3Ψ, which should be strongly disruptive. Indeed, codon:anticodon pairings including this modification had significantly (+6 to +20 kcal/mol) increased interaction energies, strongly suggesting that m3Ψ-modified codons are not substrates for amino acid addition by tRNAPhe(GAA). In contrast to m3Ψ, Ψ and m1Ψ had changes in energy differences (ΔEΣΨ/m1Ψ:tRNA-1,0,+1) that ranged from significantly negative (−4.5 kcal/mol) to slightly positive (1.1 kcal/mol) depending on the modification, its position within the codon, and the anticodon. Pseudouridine substitution has small effects on both modeled energy differences (≲1 kcal/mol) and on changes in rate constant (≲2-fold) (Supplementary Fig. 13A) tRNAPhe(GAA). N1-methylpseudouridine substitution is predicted to increase codon:anticodon stability when in the second and third position (Supplementary Fig. 13A); this is observed experimentally in the third position but not in the second (Fig. 2, Supplementary Table S1).
Modeling studies were further performed with tRNAIle(GAU). The trends in calculated energy differences and changes in experimental rate constants correlated well for both Ψ and m1Ψ (Supplementary Fig. 13). This is likely attributable to differences in which steps in the kinetic mechanism the measured kobs values reflected, which were single-turnover with respect to amino acid addition (Fig. 1B) but multiple-turnover with respect to tRNA selection (Fig. 3). In the multiple turnover scenario, small energy differences in selection are effectively integrated over many cycles of tRNA selection, making the kobs in this experiment more sensitive to changes in tRNA selection than those in a true single-turnover experiment, such as those with tRNAPhe(GAA). Net energy differences (ΔEΣΨ/m1Ψ:tRNA-1,0,+1) were better correlated with kobs than individual mRNA:tRNA base pairs (ΔEbp). For m1Ψ in the first and third positions, this particularly reflects the contributions of compensatory interactions with tRNA bases adjacent to the modified base. The ΔEm1Ψ1:tRNAt6A37 interaction, for example, contributes −6.8 kcal/mol which is equal to the sum of contributions from all the other bases in the codon and anticodon. Our findings are consistent with previous studies which demonstrate that tRNA nucleotides adjacent to tRNA codon: anticodon base pairs are important determinants of mRNA decoding55. Our computational analyses generally support our experimental findings that Ψ-derived modifications differentially affect the interactions between codons and both tRNAs in context dependent manner to alter mRNA:tRNA interactions in the ribosome decoding center53,56,57,58,59,60 (Fig. 4, Supplementary Data 1, 2).
Discussion
During the selection of aminoacylated-tRNAs, the ribosome must compromise between the speed and accuracy of decoding. Chemical modifications of the RNAs involved in decoding (e.g. mRNA and tRNA) permit the fine tuning of this balancing act. m1Ψ modifications are heavily used in mRNA-based therapeutics and vaccines and we were interested in establishing how their inclusion in mRNA transcripts can impact translation elongation26,30. Our studies indicate that, depending on where it was located within a phenylalanine codon, a single m1Ψ substitution has little effect on the rate constant (kobs) for cognate amino acid incorporation (Fig. 1B). These findings are consistent with our melting temperature data (Supplementary Fig. 10), and the kobs values previously measured values for singly substituted Tyr codons (m1ΨAC), which were altered by <2-fold36. Similarly, the rate constants (khyd,max) for translation termination are not perturbed (Fig. 1C, D). The failure to detect defects in the rates of single amino acid incorporation on an exemplar codon does not preclude the possibility that overall translation elongation rate along an mRNA – which involves a variety of codons and ribosome translocation – is not perturbed by m1Ψ incorporation.
The small effect (if any) of m1Ψ on the ribosome reacting with cognate tRNAs depends largely on the position of the modification within a codon (Figs. 2, 3). Our computational studies reveal that the minor context-dependent effect of m1Ψ on Phe incorporation that we observe might be at least partially explained by changes in the energetics of base pairing interactions between m1Ψ-substituted UUU codons and their cognate tRNAPhe(GAA) (Fig. 4A, B), which vary depending on where m1Ψ is incorporated in the codon. Perturbations in base pairing energies can influence central steps in the translation elongation pathway including tRNA selection and accommodation51. Overall, our data support previous observations indicating that the increased protein yield observed from m1Ψ containing transcripts in cells (Supplementary Figs. 8 and 9) are largely due to m1Ψ-induced enhancements in mRNA stability and avoidance of the cell’s innate immune system33,34,50,61.
In our studies, we found that m1Ψ modestly alters tRNA interactions with the ribosome. Our kinetic investigations reveal that both m1Ψ and Ψ modifications can both enhance or limit amino acid substitution depending on aa-tRNA identity and the position of the modification within the codon (Figs. 3 and 4A). These in vitro observations are supported by cellular reporter studies indicating that m1Ψ can promote miscoding events when included in full-length transcripts expressed in human cells (Fig. 2B, Supplementary Tables 6, 8). The mass spectrometry assay does not have sufficient sensitivity to detect if there are any modified codons that exhibited lowered levels of amino acid substitution, as we might expect on Leu codons based on our kinetics. Our findings are consistent with the increase in miscoding we previously observed on Ψ-containing mRNAs in the same experimental system22. Comparison of miscoding rates on m1Ψ and Ψ-containing codons suggests that the addition of a methyl group and altered ring electronics resulting from the exchange of the nitrogen, play distinct positional and codon specific roles in the modulation of miscoding (Fig. 3). This is further supported by our MM calculations revealing that methylations (m1Ψ and m3Ψ) have larger impact on the energetics of tRNA:mRNA interactions than isomerization at the C5-position alone (Ψ) (Fig. 4).
Two studies published while this manuscript was under revision have also investigated the effects of m1Ψ in mRNA on translational fidelity in cells. Kim et al.36 expressed the SARS-CoV2 spike protein from mRNA constitutively substituted with Ψ or m1Ψ in HEK293 cells and utilized mass spectrometry to search for miscoded peptides. They achieved 39% coverage of the spike protein sequence and identified six peptide fragments with single amino acid substitutions detected in at least one sample. On average, any individual miscoded peptide was observed in 4 out of 9 samples (3 each of U, Ψ, and m1Ψ); the most-abundant/best-detected individual miscoded peptide was found in 7 out of 9 samples, though the two samples lacking the peptide were both m1Ψ samples, making reliable quantitation difficult. More recently, Mulroney et al.37 utilized multiple methods to determine that translation of m1Ψ-containing SARS-CoV2 spike protein mRNA yields +1 frameshifting products in cell culture and in mice, indicating a role for m1Ψ in frame maintenance. Although Mulroney et al.37 do not directly discuss amino acid substitutions, they observe that production of full-length spike protein from m1Ψ mRNA is increased when cells are treated with paromomycin, suggesting that m1Ψ does affect decoding by the ribosome. Although we are unable to say definitively why the results of Kim et al. differ from ours and those of Mulroney et al., we note that the constitutively-modified Pfizer mRNA vaccine sequence used in the Kim et al. study was likely experimentally optimized to minimize amino acid substitutions in human cells, which might make it a poor reporter of m1Ψ’s effects on translational fidelity.
Our data reveal that the impact of m1Ψ and Ψ on mRNA decoding depends strongly on the sequence context of the modification (Figs. 3 and 4). These findings are in line with previous work demonstrating that naturally occurring mRNA modifications can differentially affect translation depending on their location within a codon or mRNA sequence22,23,62. In this work, we go beyond observing these differences to try and identify how pseudouridine-derived modifications change the fundamental interactions between mRNAs and tRNAs in a context dependent manner. Comparison of the rate constants for amino acid misincorporation on m1Ψ and Ψ containing codons, coupled with Tm measurements and MM calculations provide evidence that fundamental changes in the energetics of mRNA:tRNA base pairing contribute to the context dependent outcomes we observe. Indeed, we find that the strongest predicted interactions between the tRNAIle anticodon and Ψ and m1Ψ modified UUU Phe codons occurs when these modifications are in the first and third position of the codon (Fig. 4D, E). This is consistent with our tRNA selection assay indicating that Ile-tRNAIle reacts more rapidly with codons containing Ψ and m1Ψ in the first and third position of a codon than in the second position (Figs. 3, 4A).
tRNA anticodon step loop (ASL) modifications have long been known to influence ribosome decoding. Our data suggest that these modifications might also contribute to the context dependence decoding on modified Ψ and m1Ψ codons. We observe that the rate constants for non-cognate tRNAs possessing hyper-modifications (t6A37 in tRNAIle(GAU), ms2i6A37 and cmo5U34 in tRNASer(UGA)) were more sensitive to the position of mRNA modifications within a codon than the tRNALeu(CAG) that contains only a methylation at position 37 (m1G37) (Fig. 4A). Additionally, our MM calculations indicate that the hypermodifications adjacent to tRNAPhe(GAA) and tRNAIle(GAU) anticodons also influence the energetics of codon:anticodon interactions with Ψ and m1Ψ, and can make tRNA interactions with a non-cognate codon up to 7.6 kcal/mol more energetically favorable. This correlates with our kinetic studies demonstrating that tRNAIle(GAU) reacts more rapidly on m1ΨUU, and is further supported by previous studies demonstrating that modified tRNA A37 nucleosides (t6A, ms2t6A,ct6A) can improve the stability of the codon:anticodon duplex through enhanced base stacking30,41,56,57,63,64,65. Together, our biochemical amino acid misincorporation and modeling data suggest that tRNA ASL modifications may help to mediate tRNA discrimination on and modified codons during the decoding process (Fig. 4D, E and Supplementary Fig. 5).
The ability of m1Ψ and Ψ to modestly impact ribosome speed and decoding has several implications. Given the emerging evidence that Ψ is commonly included into mRNA at increased levels under cellular stress conditions, these findings support the possibility that Ψ−derived modifications can provide the cells with a way to transiently reshape the proteome under stress to increase fitness16,22,66,67. Furthermore, this could potentially be advantageous for mRNA vaccines relative to traditional vaccine platforms; greater antigen diversity might provide broader protection against circulating virus populations than single-strain vaccine formulations, similar to the increased efficacy of multivalent vaccines. Indeed, a recent report demonstrates that +1 ribosomal frameshift products generated from the translation of m1Ψ-substituted transcripts trigger an immune response37. While potentially beneficial in the context of mRNA vaccines, even small changes translational fidelity may need to be more carefully considered in the context of other classes of mRNA therapeutics. The complex rules which govern the translational outcome of mRNA modifications such as m1Ψ are only beginning to be elucidated, and may in some cases prove to be critical to designing effective mRNA therapeutics.
Methods
In vitro ribosome amino acid addition assays
E. coli MRE600 tight coupled 70 S ribosome were prepared by sedimentation and rate zonal ultracentrifugation45. Unmodified mRNAs were generated by run-off T7 transcription of DNA oligonucleotides. mRNAs containing modified nucleotides were synthesized and HPLC purified by Dharmacon. mRNA sequences were generally of the form GGUGUCUUGCGAGGAAUAAGUGCAUU AUG UUU UAA GCCCUUCUGUAGCCA; the coding sequence is underlined. Modified mRNA had either the first, second, third position of the Phe (UUU) codon modified with m1Ψ (Supplementary Data 3). Dharmacon verified quality control via ESI-MS data (Supplementary Fig. 14). E. coli translation factor and tRNA constructs were gifted from the laboratories of Dr. Rachel Green and Dr. Yury Polikanov unless otherwise noted. Recombinant translation factors and release factors were purified via sequential affinity, ion exchange, and gel filtration chromatography steps22,45. Natively modified tRNAPhe was expressed and purified from HB101 E. coli and Ser-, Ile-, and Leu- tRNAs were expressed and purified from BL21(DE3) E. coli utilizing ion exchange and reversed-phase chromatography45. The acceptor activity of the tRNA was validated via triplicate aminoacylation assays with both the appropriate synthetase and S100 lysate.
E. coli initiation complexes (ICs) were prepared in 1 × 219 – Tris Buffer (50 mM Tris pH 7.5, 70 mM NH4Cl, 30 mM KCl, 7 mM MgCl2, 5 mM β-ME) with 1 mM GTP22,45. Ternary complexes (TCs) were formed in the same buffer but with 10 mM GTP22. Amino acid addition reactions were conducted at final concentrations of 1 μM aminoacylated tRNA, 20 μM EF-Tu, 70 nM pre-formed ICs in buffer 1 × 219 at 37 °C. Reactions were quenched with 500 mM KOH (final) on a KinTek RF-3 quench-flow apparatus. eTLCs were visualized by phosophorimaging and then quantified with ImageQuant software (Cytiva). Data was fit to the following Eq. 1, where A is the amplitude of the signal.
In vitro translation termination assays
Pre-termination complexes (pre-TCs) were prepared by forming ICs on a mRNA containing the coding sequence AUG-UAA or AUG-m1ΨAA. Release assays were performed by mixing 70 nM pre-TCs with release factors (RF1 or RF2; 50 nM to 10 μM) at room temperature (~20 °C). Reaction time points were quenched in 5% formic acid. The fraction of released of f-[35S]-Met was fit to Equation 1 and K1/2 values were obtained by fitting to Eq. 2.
In vitro translation amino acid misincorporation
Assays performed with total aa-tRNAaa were conducted by reacting ICs (70 nM final) with TCs (1 μM total tRNA aminoacylated with either S100 enzymes or specific synthetases, 40 μM EF-Tu and 10 mM GTP) at 37 °C for 15 min. Reactions were quenched with 500 mM KOH (final). The reactants and products were separated by eTLC and visualized using ImageQuant software (Supplementary Figs. 5 and 6). For assays with high kinetic resolution (e.g. Fig. 3), ICs were reacted with Ternary complexes (40 μM EF-Tu:10 μM EF-Ts: 10 μM of aminoacylated tRNA (either Ile, Leu, Ser)). These reactions were prepared and conducted as previously published45.
Luciferase mRNA transfection and expression analyses
The template for in vitro synthesis of luciferase mRNA consists of: the T7 RNA polymerase promoter, followed by an N-terminal 3× Hemagglutinin (HA) tag fused in-frame with the firefly luciferase gene, in-frame C-terminal StrepII and FLAG tags. The open reading frame spans from the 3xHA tag to the FLAG tag enabling the purification of full-length luciferase protein and not translation truncated products. The luciferase mRNA was transcribed using T7 RNA polymerase (New England Biolabs) as previously published22.
Synthesized, purified mRNAs were transfected into 293H cells using TransIT-mRNA transfection kit as recommended by the manufacturer (Mirus). Tandem purification of the luciferase translation products was performed using the FLAG tag followed by selection for the N-terminal HA tag as described previously22,68,69 Three independent transfections were performed for uridine/ N1-methylpseudouridine -containing mRNAs. The tandem-affinity purified products were analyzed on 8% SDS-PAGE, gels were then silver stained (ProteoSilver, Sigma) and processed for mass spectrometry. For western blot analyses of the luciferase protein, proteins were separated by SDS- PAGE and blotted as previously described22.
In-gel digestion and LC-MS/MS analysis
In-gel digestion and LC-MS/MS analysis were performed as previously published22. Three independent transfections were performed for each RNA (modified and unmodified) and each sample was harvested and processed independently. UPLC was performed on a Waters NanoAcquity instrument using water +0.1% formic acid as mobile phase A and acetonitrile + 0.1% formic acid as mobile phase B. Peptides were loaded at 5% mobile phase B and separated using a 5–35% B gradient over 45 min at a flow rate of 4 uL/min. Positive mode electrospray ionization with a liquid junction potential of 1.4 kV was used to introduce ions into a Thermo Scientific Q Exactive hybrid mass spectrometer. An m/z range of 300–1750 at 70,000 resolution and an AGC target of 1e6 were used. Data-dependent acquisition selected the ten most abundant precursor ions for HCD fragmentation using an isolation width of 1.6 Da, fill time of 110 ms, and an AGC target of 1e5. Peptides were fragmented using a normalized collision energy of 27, and fragment spectra were acquired with a resolution of 17,500 at m/z = 200.
Raw data files were peak-picked by Proteome Discoverer (version 2.1), and preliminary searches were performed using the MASCOT search engine (version 2.4) against the SwissProt Human FASTA file (downloaded 05/2018) modified to include the luciferase protein sequence. Search parameters included Trypsin/P specificity, up to 2 missed cleavages, a fixed modification of carbamidomethyl cysteine, and variable modifications of oxidized methionine, pyroglutamic acid for Q, and N-terminal acetylation. The processed peak list was then re-searched in MASCOT against luciferase only as described previously22, using an error-tolerant search allowing for all possible substitutions. Substitutions with a greater than 90% probability in Scaffold (v4.8.8) were added back to the original search in Proteome Discoverer and re-run using MASCOT with these included as variable modifications22. This final search was loaded into Skyline-daily (University of Washington, v4.1), extracted ion chromatograms were generated for each peptide of interest, and peaks were manually inspected for proper peak picking, isotope dot product >0.8, good fragment ion coverage, and elution times consistent with the time of MS/MS detection. The sum of the top 3 isotopes were then exported for each modification for further analysis.
Molecular modeling
Fragment molecular orbital (FMO) calculations were used to quantify the pairwise Phe UUU codon anti-codon interaction energies70,71,72. First, the initial coordinates of the Phe UUU codon:phe tRNA complex were taken from the X-ray crystal of the Thermus thermophilus 70 S ribosome in complex with mRNA (PDB ID: 6UO1). Starting from these coordinates, we generated six additional codon:tRNA complexes with the UUU codon changed to m1ΨUU, Um1Ψ U, Uum1Ψ, ΨUU, UΨ U, and UUΨ, respectively. The codon:tRNA complexes were then processed through CHARMM-GUI webserver add hydrogens, patch the terminal 5′ and 3′ residues, and prepare the input files for energy minimization73,74. Each complex was energy minimized with 50 steps of steepest descent (SD) and 200 steps of adopted basis Newton-Raphson (ABNR) method with a gradient tolerance of 0.001. During energy minimization, the non-bonded list was generated at a cutoff of 15.0 Å and updated heuristically; and the Lennard-Jones and electrostatic interactions were treated with the switching function. Energy minimization was carried out using the CHARMM36 nucleic force field for the RNAs and solvation effects were models using the Generalized Born using Molecular Volume (GBMV) implicit solvent75,76,77. The energy minimized coordinates of the complexes were used as the starting geometries for FMO calculations.
All calculations were carried out at the Møller-Plesset perturbation theory (MP2)/6-31 G* level of theory78,79,80,81. Solvation effects were modeled using the polarizable continuum model (PCM) interfaced with the FMO method82. For the FMO-MP2/6-31 G*/PCM calculations, each RNA residue was treated as a single fragment. Input files for FMO calculations we generated from the energy minimized coordinates of the codon:tRNA complexes using an in-house fragment script (https://github.com/atfrank/RNAFMO). Briefly, fragmentation was performed at the C5′-O5′ bond of the RNA residues following the approach used in the computational chemistry software Facio (version 22.1.1.32)83,84. All FMO calculations were carried out using the ab initio quantum chemistry package, general atomic and molecular electronic structure system (GAMESS) (September 2018, R3)85. The Pair interaction energy decomposition analysis (PIEDA) facility in GAMESS was used to compute and decompose the pairwise interaction energies between individual fragments (nucleotides) in the mRNA:tRNA complexes86.
Melting temperature determination
Annealing and melting were performed in a Beckman-Coulter DU-6000 equipped with the 6-position Tm cell changer and Peltier temperature control. RNA oligos were obtained from Horizon Discovery and from IDT. Stoichiometric quantities of RNA oligos were mixed in a cuvette in 20 mM sodium cacodylate, pH 7, and 100 mM NaCl at a sufficient concentration to give an A254 reading of approximately 0.5 at 8 °C. RNA oligos were annealed by heating from 10 °C to 65 °C at 5 °C/min, holding at 65 °C for 5 min, and then cooling to 8 °C at 5 °C/min. Annealing was validated by observation of the A254 trace during the annealing procedure. Following annealing, cuvettes were removed from the instrument, kept on ice, and ice-cold MgCl2 was added to each cuvette to a final concentration of 10 mM. Cuvettes were returned to the instrument and the melting experiment was performed according to the following protocol. Cuvettes were held at 8 °C for 10 min, then heated at 0.1 °C per minute to 45 °C with readings every 0.1 °C. Subsequently, samples were heated to 65 °C with readings every 5 °C. Each experiment contained one blank cuvette containing only buffer, and one cuvette for each duplex containing U, Ψ, or m1Ψ. Three independent experiments were performed on different days for each set of duplexes.
Absorbance data at 254 nm from independent experiments were normalized using Eq. 3:
Normalized absorbance data were overlaid on the same plot and fitted using Eq. 4:
This equation assumes a two-state model for melting and allows us to explicitly fit Tm as a parameter without performing material-intensive determination of heat capacities. It further assumes that the duplex and single-stranded RNAs display the same dependence of absorbance on temperature; however, due to the limited amount of data in the fully-annealed region below 10 °C, we cannot prove or disprove this assumption. The key feature of this equation is that the maximum value of its first derivative occurs at Tm, which is a characteristic it shares with the theoretically complete treatment (Turner Methods Enzymology 2009). GraphPad Prism was used for curve fitting. Reported Tms are the values determined by fitting curves to absorbance data collected in three separate experiments (~500 data points per RNA duplex) and the reported error values are the 95% confidence intervals of the fitted parameter calculated by Prism.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw mass spectrometry data have been deposited to the ProteomeXchange Consortion (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository (https://www.ebi.ac.uk/pride/) with the dataset identifiers PXD053919. Structural data from 6UO1 was used for molecular modeling. Source data are provided with this paper.
Code availability
The in-house script used in the molecular modeling is available on the GitHub repository at https://github.com/atfrank/RNAFMO.
References
Frye, M., Jaffrey, S. R., Pan, T., Rechavi, G. & Suzuki, T. RNA modifications: what have we learned and where are we headed? Nat. Rev. Genet. 17, 365–372 (2016).
Nachtergaele, S. & He, C. Chemical modifications in the life of an mRNA transcript. Annu Rev. Genet. 52, 349–372 (2018).
Jackman, J. E. & Alfonzo, J. D. Transfer RNA modifications: nature’s combinatorial chemistry playground: transfer RNA modifications. WIREs RNA 4, 35–48 (2013).
Jonkhout, N. et al. The RNA modification landscape in human disease. RNA 23, 1754–1769 (2017).
Sloan, K. E. et al. Tuning the ribosome: the influence of rRNA modification on eukaryotic ribosome biogenesis and function. RNA Biol. 14, 1138–1152 (2017).
Pan, T. Modifications and functional genomics of human transfer RNA. Cell Res. 28, 395–404 (2018).
Haruehanroengra, P., Zheng, Y. Y., Zhou, Y., Huang, Y. & Sheng, J. RNA modifications and cancer. RNA Biol. 17, 1560–1575 (2020).
Boccaletto, P. et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 46, D303–D307 (2018).
Dumelin, C. E., Chen, Y., Leconte, A. M., Chen, Y. G. & Liu, D. R. Discovery and biological characterization of geranylated RNA in bacteria. Nat. Chem. Biol. 8, 913–919 (2012).
Chen, Y. G., Kowtoniuk, W. E., Agarwal, I., Shen, Y. & Liu, D. R. LC/MS analysis of cellular RNA reveals NAD-linked RNA. Nat. Chem. Biol. 5, 879–881 (2009).
Flynn, R. A. et al. Small RNAs are modified with N-glycans and displayed on the surface of living cells. Cell 184, 3109–3124.e22 (2021).
Jones, J. D., Monroe, J., & Koutmou, K. S. A molecular‐level perspective on the frequency, distribution, and consequences of messenger RNA modifications. WIREs RNA https://doi.org/10.1002/wrna.1586 (2020).
Moshitch-Moshkovitz, S., Dominissini, D. & Rechavi, G. The epitranscriptome toolbox. Cell 185, 764–776 (2022).
Li, X., Xiong, X. & Yi, C. Epitranscriptome sequencing technologies: decoding RNA modifications. Nat. Methods 14, 23–31 (2017).
Arango, D. et al. Acetylation of cytidine in mRNA promotes translation efficiency. Cell 175, 1872–1886.e24 (2018).
Schwartz, S. et al. Transcriptome-wide mapping reveals widespread dynamic-regulated pseudouridylation of ncRNA and mRNA. Cell 159, 148–162 (2014).
Xu, L. et al. Three distinct 3-methylcytidine (m3C) methyltransferases modify tRNA and mRNA in mice and humans. J. Biol. Chem. 292, 14695–14703 (2017).
Franco, M. K. & Koutmou, K. S. Chemical modifications to mRNA nucleobases impact translation elongation and termination. Biophys. Chem. 285, 106780 (2022).
Melnikov, S. et al. One core, two shells: bacterial and eukaryotic ribosomes. Nat. Struct. Mol. Biol. 19, 560–567 (2012).
Ramakrishnan, V. Ribosome structure and the mechanism of translation. Cell 108, 557–572 (2002).
Jobe, A., Liu, Z., Gutierrez-Vargas, C. & Frank, J. New insights into ribosome structure and function. Cold Spring Harb. Perspect. Biol. 11, a032615 (2019).
Eyler, D. E. et al. Pseudouridinylation of mRNA coding sequences alters translation. PNAS 116, 23068–23074 (2019).
Licht, K. et al. Inosine induces context-dependent recoding and translational stalling. Nucleic Acids Res. 47, 3–14 (2019).
You, C., Dai, X. & Wang, Y. Position-dependent effects of regioisomeric methylated adenine and guanine ribonucleosides on translation. Nucleic Acids Res. 45, 9059–9067 (2017).
Dai, Q. et al. Quantitative sequencing using BID-seq uncovers abundant pseudouridines in mammalian mRNA at base resolution. Nat. Biotechnol. 41, 344–354 (2023).
Agrawal, S. RNA therapeutics are stepping out of the maze. Trends Mol. Med. 26, 1061–1064 (2020).
Zhou, L.-Y., Qin, Z., Zhu, Y.-H., He, Z.-Y. & Xu, T. Current RNA-based therapeutics in clinical trials. Curr. Gene Ther. 19, 172–196 (2019).
Sergeeva, O. V., Koteliansky, V. E. & Zatsepin, T. S. mRNA-based therapeutics–advances and perspectives. Biochem. Mosc. 81, 709–722 (2016).
Gao, M., Zhang, Q., Feng, X.-H. & Liu, J. Synthetic modified messenger RNA for therapeutic applications. Acta Biomater. 131, 1–15 (2021).
Nance, K. D. & Meier, J. L. Modifications in an emergency: the role of N1-methylpseudouridine in COVID-19 vaccines. ACS Cent. Sci. https://doi.org/10.1021/acscentsci.1c00197 (2021).
Parr, C. J. C. et al. N 1-Methylpseudouridine substitution enhances the performance of synthetic mRNA switches in cells. Nucleic Acids Res. 48, e35–e35 (2020).
Svitkin, Y. V. et al. N1-methyl-pseudouridine in mRNA enhances translation through eIF2α-dependent and independent mechanisms by increasing ribosome density. Nucleic Acids Res. 45, 6023–6036 (2017).
Karikó, K. et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol. Ther. 16, 1833–1840 (2008).
Karikó, K., Buckstein, M., Ni, H. & Weissman, D. Suppression of RNA recognition by toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175 (2005).
Svitkin, Y. V., Gingras, A.-C., & Sonenberg, N. Membrane-dependent relief of translation elongation arrest on pseudouridine- and N1-methyl-pseudouridine-modified mRNAs. Nucleic Acids Res. https://doi.org/10.1093/nar/gkab1241 (2021).
Kim, K. Q. et al. N1-methylpseudouridine found within COVID-19 mRNA vaccines produces faithful protein products. Cell Rep. 40, 111300 (2022).
Mulroney, T. E., et al. N1-methylpseudouridylation of mRNA causes +1 ribosomal frameshifting. Nature https://doi.org/10.1038/s41586-023-06800-3 (2023).
Hoernes, T. P. et al. Nucleotide modifications within bacterial messenger RNAs regulate their translation and are able to rewire the genetic code. Nucleic Acids Res. 44, 852–862 (2016).
Levi, O. & Arava, Y. S. Pseudouridine-mediated translation control of mRNA by methionine aminoacyl tRNA synthetase. Nucleic Acids Res. https://doi.org/10.1093/nar/gkaa1178 (2020).
Adachi, H. et al. Targeted pseudouridylation: an approach for suppressing nonsense mutations in disease genes. Mol. Cell 83, 637–651.e9 (2023).
Kurland, C. G. Translational accuracy and the fitness of bacteria. Annu. Rev. Genet. 26, 29–50 (1992).
Sitron, C. S. & Brandman, O. Detection and degradation of stalled nascent chains via ribosome-associated quality control. Annu. Rev. Biochem 89, 417–442 (2020).
Kim, S. J. et al. Translational tuning optimizes nascent protein folding in cells. Science 348, 444–448 (2015).
Yu, C.-H. et al. Codon usage influences the local rate of translation elongation to regulate co-translational protein folding. Mol. Cell 59, 744–754 (2015).
Monroe, J. G., Smith, T. J., & Koutmou, K. S. Investigating the consequences of mRNA modifications on protein synthesis using in vitro translation assays. in Methods in Enzymology (Academic Press, 2021). https://doi.org/10.1016/bs.mie.2021.06.011.
Zaher, H. S. & Green, R. Quality control by the ribosome following peptide bond formation. Nature 457, 161–166 (2009).
Hetrick, B., Lee, K. & Joseph, S. Kinetics of stop codon recognition by release factor 1. Biochemistry 48, 11178–11184 (2009).
Dabrowski, M., Bukowy-Bieryllo, Z. & Zietkiewicz, E. Translational readthrough potential of natural termination codons in eucaryotes – The impact of RNA sequence. RNA Biol. 12, 950–958 (2015).
Wangen, J. R. & Green, R. Stop codon context influences genome-wide stimulation of termination codon readthrough by aminoglycosides. eLife 9, e52611 (2020).
Pardi, N. et al. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J. Control Release 217, 345–351 (2015).
Rodnina, M. V. & Wintermeyer, W. Fidelity of Aminoacyl-tRNA selection on the ribosome: kinetic and structural mechanisms. Annu. Rev. Biochem. 70, 415–435 (2001).
Meroueh, M. Unique structural and stabilizing roles for the individual pseudouridine residues in the 1920 region of Escherichia coli 23S rRNA. Nucleic Acids Res. 28, 2075–2083 (2000).
Kierzek, E. et al. The contribution of pseudouridine to stabilities and structure of RNAs. Nucleic Acids Res. 42, 3492–3501 (2014).
Kibbe, W. A. OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Res. 35, W43–W46 (2007).
Zhang, W., Foo, M., Eren, A. M. & Pan, T. tRNA modification dynamics from individual organisms to metaepitranscriptomics of microbiomes. Mol. Cell 82, 891–906 (2022).
Charette, M. & Gray, M. W. Pseudouridine in RNA: what, where, how, and why. IUBMB Life 49, 341–351 (2000).
Davis, D. R., Veltri, C. A. & Nielsen, L. An RNA model system for investigation of pseudouridine stabilization of the codon-anticodon interaction in tRNALys, tRNAHis and tRNATyr. J. Biomol. Struct. Dyn. 15, 1121–1132 (1998).
Davis, D. R. Stabilization of RNA stacking by pseudouridine. Nucleic Acids Res. 23, 5020–5026 (1995).
Lucas, X., Bauzá, A., Frontera, A. & Quiñonero, D. A thorough anion–π interaction study in biomolecules: on the importance of cooperativity effects. Chem. Sci. 7, 1038–1050 (2016).
Westhof, E. Pseudouridines or how to draw on weak energy differences. Biochem. Biophys. Res. Commun. 520, 702–704 (2019).
Andries, O. et al. N1-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice. J. Controlled Release 217, 337–344 (2015).
Choi, J. et al. N6-methyladenosine in mRNA disrupts tRNA selection and translation elongation dynamics. Nat. Struct. Mol. Biol. 23, 110–115 (2016).
Durant, P. C., Bajji, A. C., Sundaram, M., Kumar, R. K. & Davis, D. R. Structural effects of hypermodified nucleosides in the Escherichia coli and human tRNALys anticodon loop: the effect of nucleosides s2U, mcm5U, mcm5s2U, mnm5s2U, t6A, and ms2t6A. Biochemistry 44, 8078–8089 (2005).
Murphy, F. V., Ramakrishnan, V., Malkiewicz, A. & Agris, P. F. The role of modifications in codon discrimination by tRNA(Lys)UUU. Nat. Struct. Mol. Biol. 11, 1186–1191 (2004).
Satpati, P., Bauer, P. & Åqvist, J. Energetic tuning by tRNA modifications ensures correct decoding of isoleucine and methionine on the ribosome. Chem. Eur. J. 20, 10271–10275 (2014).
Garcia, D. M. et al. A prion accelerates proliferation at the expense of lifespan. Elife 10, e60917 (2021).
Purchal, M. K. et al. Pseudouridine synthase 7 is an opportunistic enzyme that binds and modifies substrates with diverse sequences and structures. Proc. Natl. Acad. Sci. USA 119, e2109708119 (2022).
Roy, B., Leszyk, J. D., Mangus, D. A. & Jacobson, A. Nonsense suppression by near-cognate tRNAs employs alternative base pairing at codon positions 1 and 3. Proc. Natl. Acad. Sci. USA 112, 3038–3043 (2015).
Roy, B. et al. Ataluren stimulates ribosomal selection of near-cognate tRNAs to promote nonsense suppression. Proc. Natl. Acad. Sci. USA 113, 12508–12513 (2016).
Kitaura, K., Ikeo, E., Asada, T., Nakano, T. & Uebayasi, M. Fragment molecular orbital method: an approximate computational method for large molecules. Chem. Phys. Lett. 313, 701–706 (1999).
Nakano, T. et al. Fragment molecular orbital method: application to polypeptides. Chem. Phys. Lett. 318, 614–618 (2000).
Nakano, T. et al. Fragment molecular orbital method: use of approximate electrostatic potential. Chem. Phys. Lett. 351, 475–480 (2002).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput Chem. 29, 1859–1865 (2008).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016).
Foloppe, N. & MacKerell, A. D. Jr. All-atom empirical force field for nucleic acids: I. Parameter optimization based on small molecule and condensed phase macromolecular target data. J. Comput. Chem. 21, 86–104 (2000).
Denning, E. J., Priyakumar, U. D., Nilsson, L. & MacKerell, A. D. Impact of 2′-hydroxyl sampling on the conformational properties of RNA: Update of the CHARMM all-atom additive force field for RNA. J. Comput Chem. 32, 1929–1943 (2011).
Lee, M. S., Feig, M., Salsbury, F. R. & Brooks, C. L. New analytic approximation to the standard molecular volume definition and its application to generalized Born calculations. J. Comput Chem. 24, 1348–1356 (2003).
Takematsu, K. et al. Possibility of mutation prediction of influenza hemagglutinin by combination of hemadsorption experiment and quantum chemical calculation for antibody binding. J. Phys. Chem. B. 113, 4991–4994 (2009).
Kurisaki, I. et al. Fragment molecular orbital (FMO) study on stabilization mechanism of neuro-oncological ventral antigen (NOVA)–RNA complex system. J. Mol. Structure THEOCHEM 962, 45–55 (2010).
Mazanetz, M. P., Ichihara, O., Law, R. J. & Whittaker, M. Prediction of cyclin-dependent kinase 2 inhibitor potency using the fragment molecular orbital method. J. Cheminform 3, 2 (2011).
Okimoto, N., Otsuka, T., Hirano, Y. & Taiji, M. Use of the multilayer fragment molecular orbital method to predict the rank order of protein–ligand binding affinities: a case study using tankyrase 2 inhibitors. ACS Omega 3, 4475–4485 (2018).
Fedorov, D. G., Kitaura, K., Li, H., Jensen, J. H. & Gordon, M. S. The polarizable continuum model (PCM) interfaced with the fragment molecular orbital method (FMO). J. Comput. Chem. 27, 976–985 (2006).
Suenaga, M. Facio: new computational chemistry environment for PC GAMESS. J. Comput. Chem. Jpn. 4, 25–32 (2005).
Suenaga, M. Development of GUI for GAMESS / FMO calculation. J. Comput. Chem., Jpn. 7, 33–54 (2008).
Barca, G. M. J. et al. Recent developments in the general atomic and molecular electronic structure system. J. Chem. Phys. 152, 154102 (2020).
Fedorov, D. G. & Kitaura, K. Pair interaction energy decomposition analysis. J. Comput Chem. 28, 222–237 (2007).
Acknowledgements
We would like to acknowledge Dr. Tyler Smith for his thoughtful discussions and careful reading of the manuscript. We thank the following funding sources for their support: National Institutes of Health (NIGMS R35 GM128836 (K.S.K.); NIGMS R01GM093278 (C.M.D.); T32-GM008602 (P.S.)), National Science Foundation (NSF CAREER 2045562 to K.S.K.), Research Corporation for Science Advancement (Cottrell Scholar Award to K.S.K.) and New England Biolabs (NEB) Inc. (L.M. and B.R. were New England Biolabs, Inc. employees).
Author information
Authors and Affiliations
Contributions
J.M., D.E., L.M., B.R., and A.B. performed key experiments. P.S. and C.M.D. prepared and provided critical reagents. J.M., D.E., L.M., and K.S.K. analyzed data. J.M., I.D., and A.T.F. performed computational work. J.M., D.E., A.T.F. and K.S.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hashim Al-Hashimi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Monroe, J., Eyler, D.E., Mitchell, L. et al. N1-Methylpseudouridine and pseudouridine modifications modulate mRNA decoding during translation. Nat Commun 15, 8119 (2024). https://doi.org/10.1038/s41467-024-51301-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-51301-0
- Springer Nature Limited