Abstract
Plants and microbes communicate to collaborate to stop pests, scavenge nutrients, and react to environmental change. Microbiota consisting of thousands of species interact with each other and plants using a large chemical language that is interpreted by complex regulatory networks. In this work, we develop modular interkingdom communication channels, enabling bacteria to convey environmental stimuli to plants. We introduce a “sender device” in Pseudomonas putida and Klebsiella pneumoniae, that produces the small molecule p-coumaroyl-homoserine lactone (pC-HSL) when the output of a sensor or circuit turns on. This molecule triggers a “receiver device” in the plant to activate gene expression. We validate this system in Arabidopsis thaliana and Solanum tuberosum (potato) grown hydroponically and in soil, demonstrating its modularity by swapping bacteria that process different stimuli, including IPTG, aTc and arsenic. Programmable communication channels between bacteria and plants will enable microbial sentinels to transmit information to crops and provide the building blocks for designing artificial consortia.
Similar content being viewed by others
Introduction
Communication between plants and microbes consists of a rich language of chemical messages1,2,3,4. Plants release 100s of molecules from their roots, using up to 50% of the plant’s photosynthetic output to communicate with thousands of bacteria and fungi5,6,7. The exudate composition spans small volatile organic acids, flavonoids, lipids, oligosaccharides, peptides, and proteins that are received by bacteria that, in turn, can respond to the plant with their own chemical signals2,6,8,9,10. Manipulating plant-microbe communication has been used for crop improvement; however, system complexity limits predictability1,8,11,12,13,14,15. In engineering projects, defined communication channels would facilitate the distribution of functions across an agriculture system. For example, bacterial sentinels could survey the soil using genetically-encoded sensors and circuits and transmit the information to the plant.
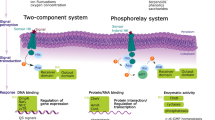
Synthetic biology projects often harness communication to coordinate cells in space and time16,17. A channel consists of a genetically-encoded “sender device” that produces a diffusible small molecule and “receiver device” that responds to it18,19,20. The term “device” refers to a transcriptional signal serving as the input (sender) or output (receiver), which simplifies the connection to other devices to build a larger system. Commonly, the chemical signals are acyl-homoserine lactones (acyl-HSLs) gleaned from bacterial quorum sensing systems. The acyl-HSL is produced by a single enzyme and binds to a regulatory protein. Specificity is determined by the length of the acyl chain, which has been exploited to build multiple non-interfering channels21,22,23,24. This language has been used for a plethora of projects, including stabilizing biofilm consortia, distributed computing, and timing metabolic flux in a bioreactor16,25,26,27,28,29,30,31,32,33,34. Communication channels between eukaryotes have been developed based on peptides and pheromones35,36,37.
Synthetic plant-to-microbe communication channels have been developed38. Plants can be engineered to excrete new chemicals from their roots, the receiver for which is put in a bacterium39,40. For example, tobacco was engineered to produce acyl-HSL and this could induce Escherichia coli carrying an acyl-HSL receiver41. Similarly, a sender device in barley was built by introducing two prokaryotic genes to make scyllo-inosamine (SI), the receiver for which is the SI-binding MocB regulator, placed in in the soil bacterium Azorhizobium caulinodans38,42.
Microbe-to-plant communication requires building a sensor in the plant with a low limit-of-detection. Constructing sensors in plants is difficult due to slow engineering cycles, tissue-specific expression, fewer genetic part (e.g., promoter) libraries, chromosome context effects and complex molecular transport43,44,45,46,47,48. Genetically-encoded plant sensors have been built that respond to ethanol, tetracycline, steroids, insecticides, trinitrotoluene (TNT), copper, fentanyl, and acetaldehyde44,49,50,51,52,53,54,55,56,57,58. However, these are not appropriate communication signals because of issues with specificity, diffusion, high limits-of-detection, or low production titers by bacteria.
Because of their role in establishing symbiotic relationships between bacteria and plants, acyl-HSLs have been proposed to be natural examples of interkingdom communication2,59,60. Biofilms on the root are abundant in acyl-HSLs, reflecting the volume of communication that occurs there59,61,62,63,64. Indeed, up to 12% of the species in soil make acyl-HSLs59,64,65. Plants have evolved means to eavesdrop on them to identify bacteria and respond appropriately1,2. Acyl-HSLs rapidly diffuse to the root surface from up to 30 µm away, are taken up and regulate hundreds of genes via poorly understood mechanisms1,66,67,68,69,70,71,72. The specific response depends on the plant, but short-chain acyl-HSL tend to change root morphology whereas long-chain (>C12) acyl-HSLs affect defense and immunity, and both can impact energy/metabolic process, hormone production and Ca2+ signaling70,71,73,74. Plants and microbes can interfere with acyl-AHSL signaling by producing degrading enzymes and chemical mimics66,70,74,75.
Rhodopseudomonas palustris is a plant-growth promoting bacterium isolated from rice paddies that produces only one quorum signal: p-coumaroyl-homoserine lactone (pC-HSL)76. In place of the acyl-group, it sources an aryl-group from p-coumarate secreted from plant roots65,77. No known soil bacterium has the complete pathway to pC-HSL65; however, a synthetic pathway (rpaI/4cl/tal) has been built in E. coli to make p-coumarate and incorporate it into pC-HSL78,79. The activator RpaR binds to pC-HSL and to the rpaO*A DNA operator65,78. Some other species make pC-HSL, but it is far less abundant than acyl-HSL59,64,65, plants do not respond except at high concentrations80,81, and there are far fewer deactivating enzymes and mimics in soil61,82.
Here, we demonstrate programmable microbe-to-plant communication from a pC-HSL sender in the soil bacteria Pseudomonas putida KT2440 and Klebsiella pneumoniae 342 to pC-HSL receivers in Arabidopsis thaliana and Solanum tuberosum (potato) (Fig. 1a). P. putida has been proposed to be used in agriculture to promote plant growth, is non-pathogenic and does not secrete any HSLs83,84,85. Klebsiella pneumoniae 342 is an endophytic nitrogen fixer first isolated from maize, colonizes roots of maize, wheat, rice, and Arabidopsis, and does not secrete any HSLs86,87,88,89. No plant pC-HSL receptor was known, so here we build a receiver device for plants by constitutively expressing RpaR and building a responsive promoter. This receiver is specific to pC-HSL and does not cross-react with acyl-HSL. The sender device can be connected to different sensors (IPTG, aTc, arsenic) and logic circuits and communicate the output to the plant root (Fig. 1b). This approach allows the plant to respond to different environmental signals by swapping the bacterium, rather than genetically modifying the plant. This work describes the division of labor by moving environmental sensing to bacterial sentinels at the roots, who relay the information to the plant.
a Bacteria receive a signal in the soil (grey diamonds) that induces the release of the communication signal (orange circles) to be sensed by regulatory proteins in the plant cell. b The communication channel is modular. To change the signals to which the plant responds, it simply can be grown with different bacteria engineered to connect different sensors (A or B) to the synthesis of the chemical used for communication. The bacterium can also integrate these signals using genetic circuits; an OR gate is shown. c The plant pC-HSL receiver device. The genetic part DNA sequences are provided in Supplementary Data 1. d The modifications to the prokaryotic RpaR regulator (orange) are shown to make it functional in plants. e Phenotypic comparison of A. thaliana wild-type to that carrying the pC-HSL receiver (A. thaliana 315_14_5). The plants were induced for 24 h in MS media in the hydroponic system. f Phenotypic comparison of wild-type A. thaliana with that carrying the pC-HSL receiver (A. thaliana 315_14_5_1) grown in soil. The data points represent replicates performed with different plants (n = 6 for height, fresh weight, dry weight, number of rosette leaves, number of primary shoots and primary root length; n = 13–14 for primary root length and lateral root density) and the bars represent the means of these points. Statistically significant differences were determined using two-tailed Student’s t test (ns, not significant P > 0.05). Source data are provided as a Source Data file.
Results
Plant HSL receiver devices
The plant receiver should detect a specific HSL and respond by generating a transcriptional output; in other words, activating a promoter (Fig. 1c). The receiver design was based on a plant promoter scaffold developed by Quatrano and co-workers that was shown to be functional in different plants, including A. thaliana90. This promoter was based on a minimal 35S motif (Pm35S) that is inactive unless an activator binds upstream. This scaffold had been used to create a 3-oxooctanyl-L-HSL (OC8-HSL) inducible promoter that cross-reacts with other HSLs90. Here, we changed this design to decrease the spacing between four operators from 10 bp to 2 bp, following our work in mammalian cells91, and added the TMVΩ translational enhancer92 downstream of the promoter to increase expression.
Four prokaryotic regulators were selected from quorum sensing systems: 1. LuxRAM (Vibrio fischeri), 2. CinRAM2 (Rhizobium leguminosarum), 3. LasRAM (Pseudomonas aeruginosa) and 4. RpaRAM (Rhodopseudomonas palustris). Each regulator and its cognate operator were gleaned from a set that we had previously evolved to improve their orthogonality and dynamic range (indicated by AM)21,93. To make the regulators functional in plants, their N-termini were fused to a SV40 nuclear localization signal (NLS), activation domain (VP16) and a 6 G flexible linker (Fig. 1d)94. The regulators were placed under the control of the strong 35S promoter, TMVΩ translation enhancer, and TOCS terminator.
The devices were constructed by combining the reporter expression cassette with the output promoter (Methods). The regulators are constitutively expressed as monomers and, in the presence of the HSL, they dimerize and bind to the output promoter, leading to green fluorescent protein (GFP) expression. The cassette included a phosphinothricin acetyltransferase gene to generate resistance to the herbicide phosphinothricin (PPT) as a selectable marker. The constructs were transformed into A. thaliana using the Agrobacterium floral dip method (Methods). Multiple homozygous A. thaliana lines for each of the receivers were identified after rounds of herbicide selection that segregated to all resistant progenies. No significant phenotypic differences were observed between wild-type A. thaliana and A. thaliana containing the pC-HSL receiver (315_14_5) (Fig. 1e, f).
The A. thaliana lines containing the receivers were then tested for their ability to respond to their cognate HSLs. Seeds were germinated and grown on agar plates for 7–12 days in a growth chamber before being transferred to a hydroponic system, following the protocol of Shank and co-workers (Methods) (Supplementary Fig. 1)95. This system allowed the plant roots to be exposed to a homogeneous inducer concentration in a 24-well plate format. We screened 4–9 plant lines for each HSL receiver. As an initial screen of activity, we added 100 µM of HSL inducer: N-3-oxohexanoyl-L-homoserine lactone (OC6-HSL) for the LuxR-expressing line, 3-hydroxytetradecanoyl-homoserine lactone (OHC14-HSL) for the CinR-expressing line, N-3-oxododecanoyl-L-homoserine lactone (OC12-HSL) for the LasR-expressing line or pC-HSL for the RpaR-expressing line. After 24 h of induction in the hydroponic plate, the fluorescence was visualized in the root tissue using confocal microscopy (Methods). To quantify fluorescence, we calculated the mean pixel intensity (MPI) across root tissue sections (Supplementary Fig. 2).
A. thaliana lines containing the pC-HSL, OC12-HSL and OHC14-HSL receivers showed 180-fold, 40-fold and 7-fold inductions, respectively (Fig. 2a, b and Supplementary Figs. 3–5). The OC6-HSL receiver yielded no functional lines, so it was not pursued further. Of the nine independent lines tested for pC-HSL induction, seven were active, of which we selected A. thaliana 315_14_5 for further characterization (Supplementary Fig. 6). We found that GFP was only expressed in mature tissues, with no GFP in the meristem (Fig. 2a, Supplementary Figs. 7–10). Using RT-qPCR, we found that GFP was mostly expressed in root tissues compared to leaf and stem tissues (Supplementary Fig. 11).
a Fluorescence microscopy images of the induction of the pC-HSL receiver expressing GFP (green) and stained with propidium iodide (PI, red). A. thaliana 315_14_5_1 was induced with 1 µM pC-HSL for 24 h in a hydroponic system (Methods). Images are representative of experiments performed on three different days with different plants (Supplementary Fig. 7). b Response function of the A. thaliana pC-HSL receiver. Each color represents experiments repeated on 6 different days with different plants (A. thaliana 315_14_5). All the data were fit to Eq. 1 (parameters in Supplementary Table 1). Raw images used to calculate the MPI are provided in Supplementary Fig. 12. c Orthogonality of the pC-HSL receiver. A. thaliana 315_14_5 was induced with 100 µM of each inducer (p-coumarate, OHC14-HSL, OC6-HSL, OC12-HSL and pC-HSL) for 24 h in a hydroponic system (Methods). The points represent replicates performed with different plants (A. thaliana 315_14_5) on different days (n = 6 for pC-HSL and uninduced, n = 3 for other HSLs, and n = 2 for p-coumarate) and the bars represent the means of these points. d Microscopy images of the induction of the pC-HSL receiver in soil. A. thaliana 315_14_5_1 was grown and induced by watering the plants with 100 µM pC-HSL in sterile soil (Methods). Images are representative of experiments performed on three different days with different plants. e Induction of the A. thaliana pC-HSL receiver in soil. The bars represent the mean fluorescence from three plants grown on different days (A. thaliana 315_14_5). Raw images used to calculate the MPI are provided in Supplementary Fig. 16. There is a 13-fold upregulation between induction with 0 µM of pC-HSL and 100 µM of pC-HSL. Statistical significance was determined using two-tailed Student’s t test (***P < 0.001; **P < 0.01; *P < 0.05; ns, not significant P > 0.05). Source data are provided as a Source Data file.
The full response functions were measured for the pC-HSL and OC12-HSL receivers (Fig. 2b and Supplementary Fig. 3). The minimum detection limit was 100 nM pC-HSL, which is an order of magnitude higher than the detection limit of RpaR in R. palustris65. In the absence of inducer, the background expression was 5-fold higher than wild-type A. thaliana (Supplementary Fig. 13). The whole root response functions for the pC-HSL and OC12-HSL receivers are shown in Supplementary Fig. 14.
Plants containing the pC-HSL receiver were then tested whether they respond to non-cognate HSLs (orthogonality) (Fig. 2c). Plants were grown and induced, as before, with 100 µM of each HSL for 24 h. There was no observed induction by the non-cognate HSLs, as the background fluorescence was indistinguishable from plants in the absence of inducer. This was consistent with the observation that these molecules do not bind the evolved RpaRAM21. The precursor p-coumarate does not induce the receiver, which is consistent with previous studies with RpaR96.
The inducibility of the pC-HSL receiver for plants grown in soil was tested (Fig. 2d, e). Seeds were sterilized and germinated on agar plates to achieve uniform germination. After the emergence of the first leaf, the plants were transferred into 1:3 vermiculite:soil non-sterile mix supplemented with fertilizers and grown for 10 days in a growth chamber before being induced in situ, in the soil (Methods). Plants were induced by pipetting 1 mL of water supplemented with 100 µM of pC-HSL directly on the plant-soil interface. We estimated the concentration of pC-HSL throughout the soil to be 260 nM, but it is expected to be higher near the surface as it is added through watering (Methods). Plants were then grown for an additional 24 h in the growth chamber before being prepared for imaging by washing the roots in water. The fluorescence from GFP was measured using confocal microscopy (Methods). As observed in the hydroponics experiments, GFP was only observed in the root tissue of plants containing the pC-HSL receiver in the presence of pC-HSL.
Bacterial pC-HSL sender device
Soil bacteria often produce acyl-HSLs. For example, P. putida IsoF and WCS358 produce 3OC12-HSL97,98,99. To confirm that our strains do not produce acyl-HSLs or pC-HSL, we performed a BLAST search on the genomes of P. putida KT2440 and K. pneumoniae (Methods). Neither species had any proteins with significant sequence similarity to RpaI, LuxI, LasI, CinI, or TraI. The inability of these species to produce pC-HSL was further validated experimentally by testing whether the wild-type strains could induce the A. thaliana pC-HSL receiver (Supplementary Figs. 17–18).
A sender device must convert the transcriptional output of a sensor or circuit into the production of the communication molecule to a concentration detectable by the receiver. A three-gene operon was designed to convert endogenous tyrosine to pC-HSL (Fig. 3a). The biosynthetic pathway was constructed using a pC-HSL synthase gene from Rhodopseudomonas palustris (rpaI), a tyrosine ammonia-lyase gene from Rhodobacter sphaeroides (tal) and a 4-coumarate coenzyme A ligase gene from Nicotiana tabacum (4cl)23,79. For experiments requiring the production of pC-HSL, p-coumarate was added to the media because it increases pC-HSL production79. A red fluorescent protein gene (mCherry) was included in the operon so that induction could be monitored.
a The constitutive production of pC-HSL by P. putida and K. pneumoniae was first used to induce the receiver in plants. b Phenotypic comparison of A. thaliana 315_14_5_1 (Supplementary Table 3) grown in hydroponics with and without wild-type P. putida or wild-type K. pneumoniae (Methods). c The induction of the pC-HSL receiver in A. thaliana (A. thaliana 315_14_5_1) by P. putida or by K. pneumoniae constitutively producing pC-HSL (pTT337, Supplementary Fig. 35) in hydroponics (Methods) is shown. The data were extracted from the images in Supplementary Fig. 18. The points were obtained for n = 3 plants on different days and the bars represent the means of these points. d Induction of the A. thaliana pC-HSL receiver in plant roots in hydroponics (Methods). The induction by wild-type P. putida (left) and K. pneumoniae (right) was compared to when P. putida and K. pneumoniae constitutively produce pC-HSL (pTT337). Images are representative of experiments performed on three different days with different plants (A. thaliana 315_14_5_1). e Induction of the A. thaliana pC-HSL receiver by P. putida constitutively producing pC-HSL (pTT337) in sterile and non-sterile soil (Methods). P. putida was introduced either by seed inoculation or through watering (Methods). The data were extracted from the images in Supplementary Fig. 23. The points were obtained for n = 3 plants (A. thaliana 315_14_5_1) on different days and the bars represent the means of these points. f Microscopy images of the induction of the A. thaliana pC-HSL receiver by P. putida from panel e. Statistical significance was determined using two-tailed Student’s t test (***P < 0.001; **P < 0.01; *P < 0.05; ns, not significant P > 0.05). Source data are provided as a Source Data file.
Initially, to test whether this pathway produced sufficient pC-HSL to theoretically turn on the plant receiver, the biosynthetic operon was placed under the control of the strong constitutive promoter BBa_J23100 on a pBBR1-ori plasmid. Before testing with plants, we built a surrogate receiver using E. coli engineered to respond to pC-HSL. The E. coli genome was modified to contain rpaRAM controlled by a constitutive promoter and PrpaR*A driving yellow fluorescent protein (YFP) expression21,100 (E. coli MG1655 sTT658, Supplementary Table 4). The fluorescence of YFP was measured using flow cytometry (Methods). This reporter strain showed pM sensitivity and was fully induced at 10 nM pC-HSL after 3 h (Supplementary Fig. 19) (Methods). To estimate the pC-HSL concentration produced by sender cells, we collected the supernatant after growth in MS medium supplemented with 100 µM of p-coumarate for 24 h and used it to induce the E. coli pC-HSL receiver strain. Using these data, we estimated the concentration produced by the P. putida in the presence of A. thaliana to be 1.5 ± 0.2 µM pC-HSL (Supplementary Fig. 19). When K. pneumoniae was cultured in MS medium for 24 h in the presence of A. thaliana, pC-HSL production was estimated to be 0.30 ± 0.07 µM.
Bacteria-to-plant signal relay
We then tested the ability for bacteria producing pC-HSL to induce the A. thaliana pC-HSL receiver (Fig. 3a). Initially, the strains constitutively producing pC-HSL were used (P. putida pTT337 and K. pneumoniae pTT337). The plants were germinated on solid agar and added to the 24-well plates. Separately, the bacteria were grown in LB medium overnight, then washed and diluted into MS medium into the wells containing the plants. The co-culture was grown for 24 h, and the roots imaged (Fig. 3b). The A. thaliana pC-HSL receiver was induced 30-fold when cultured with P. putida pTT337 and 50-fold with K. pneumoniae pTT337 producing pC-HSL but remained uninduced when grown with wild-type bacteria (Fig. 3c, d, Supplementary Figs. 17–18, 20). Induction in solid agar produced a similar result as the hydroponic system (Supplementary Fig. 21).
To test for non-specific interference in the communication between the plant and the bacteria, we then tested whether the receiver’s response to pC-HSL changed in the presence of wild-type P. putida, which could happen through a non-specific response to the bacteria. We did not observe a significant change in the response function when the bacteria were present (Supplementary Fig. 22).
The ability to transmit a signal in soil was then tested (Fig. 3e, f, Supplementary Fig. 23–24). Bacteria were introduced to the soil either by diluting them to OD600 = 0.1 into PBS supplemented with 100 µM p-coumarate and subsequently dispensing them into the soil, or by immersing the sterile A. thaliana pC-HSL receiver seeds into the P. putida overnight culture before sowing them into the soil (Methods). In sterile soil, we observed 32-fold and 39-fold inductions of the pC-HSL receiver when the system was inoculated with P. putida pTT337 compared to P. putida WT through seed inoculation and watering respectively (Fig. 3e). In non-sterile soil, a smaller 17-fold induction was observed. Whole-root imaging of the plants inoculated by watering in sterile and non-sterile soil show that GFP is expressed in the mature root tissues closer to the surface (Supplementary Fig. 24).
Induction of potato by bacteria producing pC-HSL
The pC-HSL receiver device was then moved to potato (S. tuberosum). The pC-HSL receiver construct (pTT315-Hyg, Supplementary Fig. 34) was used to construct S. tuberosum 315 (Methods). Carrying the pC-HSL receiver resulted in a small decrease in both the fresh and dry stem weight, and a small increase in the chlorophyll content index, but otherwise did not have a significant impact on the phenotype (Supplementary Fig. 25). The response of the S. tuberosum pC-HSL receiver was measured by adding pC-HSL to the plants in the hydroponic system. After 24 h, RNA was isolated from the root tissue and gfp transcription was quantified by qRT-PCR (Supplementary Fig. 26). qRT-PCR was used instead of confocal microscopy due to lower GFP expression. Still, inducing the S. tuberosum pC-HSL receiver with 100 µM pC-HSL resulted in the 10-fold upregulation of gfp transcripts. Inducing the S. tuberosum pC-HSL receiver with the bacterial sender (P. putida pTT337) resulted in the 6-fold upregulation of the gfp transcripts (Supplementary Fig. 26).
Modular sensing and signal processing by engineered bacteria
Once the pC-HSL receiver is put into a plant, the same plant can be made to sense different signals by changing the sensor contained by the bacterium. Multiple sensors can be integrated by genetic circuits performing logic operations that, in turn, control the pC-HSL sender, again relaying this signal to this same plant.
To demonstrate this modularity, strains of P. putida were built containing sensors that respond to different chemicals. For proof-of-principle, two sensors were built for small molecules that are often used as inducers: isopropyl β-D-1-thiogalactopyranoside (IPTG, lacIAM) or anhydrotetracycline (aTc, tetR)93. The sensors controlling YFP expression were moved from E. coli to P. putida without making genetic modifications. The response functions of the sensors yielded dynamic ranges of 140-fold (IPTG) and 740-fold (aTc) (Supplementary Fig. 27). The small molecule sensors built for P. putida were connected to the pC-HSL sender (Fig. 4a). The bacteria were first grown without inducer and then added to the media containing the plant. They were then induced with 2 mM IPTG or 1 µM aTc, as appropriate. After 24 h, the induction of the pC-HSL receiver in plant roots was quantified (Fig. 4a, Supplementary Fig. 28). The ON and OFF states of the receiver induced by the bacteria were equivalent to the dynamic range observed by the induction with exogenous pC-HSL: 25-fold (IPTG) and 47-fold (aTc).
a.A. thaliana responds to P. putida engineered to relay pC-HSL upon sensing inducers IPTG or aTc (pTT409 and pTT410, Supplementary Fig. 35) in hydroponics (Methods). The data were extracted from the images in Supplementary Fig. 28. The points were obtained for n = 3 plants (A. thaliana 315_14_5_1) on different days and the bars represent the means. Microscopy images match the blue-circled replicate. b.P. putida and K. pneumoniae were engineered to detect arsenic (pTT417, Supplementary Fig. 37) and communicate the output to the A. thaliana pC-HSL receiver in hydroponics (Methods). The data were extracted from the images in Supplementary Fig. 29. The points were obtained for n = 3 plants (A. thaliana 315_14_5_1,) on different days and the bars represent the means. Microscopy images match the red-circled replicate. c A. thaliana 315_14_5_1 was co-cultured with P. putida sTT659 (Supplementary Table 4) engineered with an OR gate (pTT434, Supplementary Fig. 38), producing pC-HSL in response to either aTc or IPTG. The data were extracted from the images in Supplementary Fig. 30. Growth conditions and replicates were the same as in part a. P values for each of the induced state compared to the uninduced state are: +IPTG/-aTc: 0.02,-IPTG/+ aTc: 0.05, + IPTG/+ aTc: 0.03. Microscopy images match the blue-circled replicate. d A. thaliana 315 co-cultured with two strains of P. putida, each producing pC-HSL in response to a different signal. Strains, growth conditions, and replicates were the same as in part a. The data were extracted from the images in Supplementary Fig. 31. P values for each of the induced state compared to the uninduced state are: +IPTG/-aTc: 0.008,-IPTG/+ aTc: 0.010, + IPTG/+ aTc: 0.0007. Microscopy images match the yellow-circled replicate. Statistical significance was determined using two-tailed Student’s t test (***P < 0.001; **P < 0.01; *P < 0.05; ns, not significant P > 0.05). Source data are provided as a Source Data file.
Next, we demonstrated that P. putida and K. pneumoniae could act as bacterial sentinels for detecting arsenic. Arsenic is a prevalent and toxic heavy metal that is a global polluter of farmland101,102. We constructed an arsenic sensor by expressing the E. coli ArsR repressor103 under control of a strong constitutive promoter (PLacIQ)93 and synthetic RBS104 to ensure high levels of ArsR. An arsenic responsive promoter was constructed by overlapping the ArsR binding sequence103 with a strong constitutive promoter (BBa_J23100). The sensor was tested on a pBBR1-ori plasmid in both P. putida and K. pneumoniae. In P. putida, the sensor yielded a 52-fold response and a minimal detection limit of 10 ppb and in K. pneumoniae, it was 72-fold and 20 ppb (Supplementary Fig. 29). Note these detection ranges are close to the EPA’s limit for drinkable water105. The arsenic sensor was connected to the pC-HSL sender in P. putida and K. pneumoniae and GFP expression was present in the roots of the pC-HSL plant receiver: 15-fold and 16-fold induction with P. putida and K. pneumoniae respectively when induced with 1000 ppb arsenic (Fig. 4b, Supplementary Fig. 29).
Signal processing can modulate the response of sensors, integrate multiple sensors, or implement memory or dynamics106,107,108. While difficult to build in plants, many circuits have been constructed in bacteria, a process simplified with computer aided design (CAD) software43,47,109,110,111,112. Sensors that have promoters as outputs are easily connected to transcriptional genetic circuits113. Logic gates integrate multiple sensors, one of the simplest of which is an OR gate where the output is ON if either of two inputs is ON. Here, we implemented an OR gate in P. putida by placing the IPTG-inducible promoter (PTac) and the aTc-inducible promoter (PTet) in series in front of the pC-HSL sender device. The pC-HSL receiver in the Arabidopsis root turned on in the presence of either aTc or IPTG (Fig. 4c and Supplementary Fig. 30). This result demonstrates that the plant can be made to perform computational functions by placing the circuitry in root-associated bacteria.
Multiple bacterial sentinels could transmit information using the same communication channel. As a demonstration, the microbe-to-plant relay experiments were repeated, but with two strains of bacteria, each containing a different sensor connected to the pC-HSL sender (Fig. 4d). Separately, the bacteria were grown in LB medium overnight and then combined into the wells containing the plants. Combinations of the two inducers were added to the wells (1 µM aTc, 2 mM IPTG). Either inducer was sufficient to obtain the induction of the receiver in the root, noting that the magnitudes of the ON states varied for the different combinations of inducers (Fig. 4d, Supplementary Fig. 31). Therefore, the consortium is performing “fuzzy OR” logic.
Discussion
This work demonstrates a programmable channel of communication from a bacterium to a plant. The advantage of this approach is that the plant receiver only needs to be built and optimized once. Compared to bacteria, new plant sensors are much harder to build and optimize due to slow design-build-test cycles and few genetic parts. As a result, there are far fewer synthetic sensors available for plants and even simple inducible systems are notoriously unavailable. The few that are used are far worse in their performance and dynamic range than what is available for bacteria114,115. In contrast, there are hundreds of sensors built for bacteria any of which could be connected to the sender so long as its promoter output has an appropriate dynamic range. Such sensors have been built to respond to toxins, pollutants, nutrients, pathogens, agrochemicals, or any other stimuli18,106,116,117. In addition, optimization is much easier in bacteria because it is possible to implement complex selections and there are rapid generation times. The plant receiver allows one plant-based sensor to be developed and optimized, which can then be used with all the sensors and circuits available for bacteria. Changing what is being sensed and the signal processing involves changing the bacterium and not building a new system in the plant.
Complex circuitry could be moved to the bacterium, where it is easier to build93,108. Circuits can integrate information from multiple sensors or implements a dynamic response to an input signal (e.g., a pulse)108. The circuit’s output promoter could be connected to the pC-HSL sender to communicate the result to the plant. Similarly, the pC-HSL receiver was used here to control reporter expression, but it could be connected to metabolic pathways or transcription factors to control morphology118,119. Sense-and-respond systems could be distributed, where a bacterial sentinel receives information (e.g., toxin), transmits it to the plant, which then turns on a response (e.g., detoxification pathway).
Using a common communication signal addresses a second problem in that different molecules will have different uptake and transport properties in the plant. The principal constraint on selecting pC-HSL as a communication signal was that the molecule had to both be producible by a bacterium and able to be sensed by a plant. Transport was also an important consideration, as the molecule must be exported through the prokaryotic membrane and taken up by the root, ultimately entering the nucleus. Prior to selecting pC-HSL, we tested other candidates (data not shown). Lipo-chitooligosaccharides (nod factors) were considered, but we had difficulty both in producing high titers in bacteria and creating a sensor that could be moved to non-legumes. There is an indigo sensor for plants120 and we could make it at high titer in bacteria using published pathways121, but we could not export it from the bacterium without lysis. Another mode of interkingdom signaling is the production of plant hormones, such as auxins, by growth-promoting bacteria122. However, these have extensive effects on plant growth and gene expression and are ubiquitous across plant species. Using the same plant promoter scaffold described in this work, we also attempted to make sensors for DAPG (PhlF), OC6-HSL (LuxR) and OC8-HSL (TraR), but they failed.
More generally, rare HSLs may be the ideal molecules for interkingdom signaling. Homoserine lactones can transport across the walls of many cell types, can be produced at sufficient titer with a few enzymes, are non-toxic and bind to well-defined regulators. Note that while the HSL signal from the bacteria was detectable in non-sterile soil, it was weaker compared to sterile soil conditions, and factors such as the water source and composition, microbial content, soil treatment, and specifics to the system we use introduce variables that could attenuate signal strength and affect plant phenotype. These considerations are likely to lead to less predictability under field conditions. This reduced activity could be due to the microbes degrading or sequestering the signaling molecule. Thus, acyl-HSLs may be too common to enable specificity between an engineered plant and microbe and have too systematic of impact on plant gene expression. However, there are many other HSL structures from which to choose. This includes unusual branched-chain groups, other aroyl- groups (e.g., cinnamoyl-HSL) and structurally unrelated mimicking compounds9,123,124. Non-natural HSLs have been built with synthetic organic chemistry, including sulfonyl, aroyl and alkanoyl-HSLs81,125. All natural homoserine lactones are L-isomers and D-isomers are not biologically active in plants126,127. Synthetic pathways to the D-isomers could reduce crosstalk with plant signaling, but this would require engineering new regulators. Finally, retrosynthetic design software128 could create completely new chemistries that are not degraded by native bacteria.
Microbes beneficial to crops have long been used in agriculture. Typically, one species has been used or few have been combined into an artificial consortium, relying on the capabilities of unmodified species. Building such consortia has been ad hoc, and it has been noted that the positive effects of multiple species often do not combine additively62,129. Advances in genetic engineering make the design of multi-species consortia and the rational distribution of functions across the consortia possible. This work lays the foundation for using bacteria grown in proximity to a plant to perform tasks such as monitoring soil nutrient content, sensing pathogens, or detecting environmental contaminants.
In addition to agricultural applications, this approach could be applied to build plant sentinels. For example, it has been proposed to use engineered plants to detect landmines by sensing TNT130. Moving the sensing to root-associated bacteria allows the same plant to be used to detect different signals simply by swapping the engineered bacterium with which it is partnered that contains the new sensor. Further, when combined with the plant-to-microbe signal developed previously38, this completes two-way communication to coordinate interkingdom functions, such as the establishment of synthetic symbiosis131,132. By viewing the plant-microbe community holistically, we can select the organism best suited for a particular task and use orthogonal channels of communication to stabilize and coordinate the population. Desirable functions could be distributed amongst members of the consortium, where individual species are assigned duties such as detoxification, pathogen defense and nutrient scavenging.
Methods
Strains, media and chemicals
All bacterial strains are listed in Supplementary Table 4. Escherichia coli NEB® 10-beta (New England BioLabs, C3019I) was used to clone all plasmids. Pseudomonas putida KT2440 (ATCC 47054) and Klebsiella pneumoniae 342 (ATCC BAA-2552) were used for co-culture experiments. Agrobacterium tumefaciens GV3101 (Gold Bio, GV3101 Electrocompetent) was used for the floral dip method. Bacterial cells were routinely grown in LB Miller broth (Difco, 244620) at 37 °C for E. coli and 30 °C for P. putida and A. tumefaciens. Plates were made using LB Miller broth with 1.5% Bacto Agar (Difco, 214010). Antibiotics were used to maintain plasmids during routine growth: kanamycin (GoldBio, K-120-10)-35 µg/mL for E. coli, 50 µg/mL for P. putida and A. tumefaciens; tetracycline (GoldBio, T-101-25)-10 µg/mL for E. coli, 25 µg/mL for P. putida; gentamycin (Enzo Lifesciences, 380-003-G001)-15 µg/mL for E. coli, 50 µg/mL for P. putida; chloramphenicol (Alfa Aesar, B20841)-25 µg/mL for P. putida; rifampicin (Santa Cruz Biotechnology, SC-200910)-50 µg/mL for A. tumefaciens. Stocks of 10 mM HSLs were solubilized in DMF and stored at -20 °C: N-3-oxohexanoyl-L-homoserine lactone (OC6-HSL; Sigma, K3007); 3-hydroxytetradecanoyl-homoserine lactone (OHC14-HSL; Sigma, 51481); N-3-oxododecanoyl-L-homoserine lactone (OC12-HSL; Sigma, O9139); p-coumaroyl-homoserine lactone (pC-HSL; Sigma, 07077). Bacterial cells were induced using the following chemical stocks: IPTG (isopropyl-ß-D-1-thiogalactopyranoside; Gold Biotechnology, I2481) in water; aTc (anhydrotetracycline; Sigma, 37919) in 50% (v/v) ethanol; arsenic (sodium meta-arsenite; Sigma, S7400) in water. For flow cytometry, cells were diluted in phosphate buffer saline (PBS; Sigma, 6505-4 L). MSVI vitamin solution (2 mg/mL glycine -Thermo Scientific, J16407-36; 0.5 mg/mL nicotinic acid - Sigma, N0761-100G; 0.5 mg/mL pyridoxine HCl - Sigma, P2680-25G; 0.4 mg/mL thiamine HCl - Sigma, T1270-25G), JHMS vitamin solution (0.4 mg/mL folic acid - Phytotech, F430; 0.05 mg/mL biotin - Phytotech, B140) and 3R vitamins solution (1 mg/mL thiamin HCl - Sigma, T1270-25G; 0.5 mg/mL nicotinic acid - Sigma, N0761-100G; 0.5 mg/mL pyridoxine HCl - Sigma, P2680-25G) were used for the preparation of CIM and 3C5ZR media.
A. thaliana growth and transformation
All plant lines are listed in Supplementary Table 3. A. thaliana Col-0 seeds (NACS, CS70000) were acquired from the Arabidopsis Biological Resource Center. When working with a small number of seeds, they were surface sterilized with 70% ethanol for 1–2 min, followed by 10% bleach for 10 min, and then rinsed 5 times with water. Larger numbers of seeds were sterilized using the chlorine gas method133. Sterilized seeds were sown on half strength Murashige and Skoog (Sigma, M5519) media with 1% sucrose (Fisher Scientific, S5-3) and adjusted to pH 5.7 with KOH (MS media). Plates were made with the addition of 0.8% agar (Sigma, A7921). Plates were sealed with MicroporeTM tape (3MTM, 1530-0) to allow for gas exchange. Seeds were stratified at 4 °C in the dark for 3 days before moving to a growth chamber (Percival Scientific, CU-36L5) where they were grown at 27 °C in 16/8 h light/dark cycles with a light intensity of 40 µmol/m2/s. For growth to seed, plants were also grown in soil in a greenhouse with 16/8 h light/dark cycles at 21 °C. Transgenic A. thaliana lines were generated by Agrobacterium tumefaciens-mediated floral dip. Briefly, A. tumefaciens strains containing plasmids of interest were cultured in 2 mL LB media containing appropriate antibiotics at 30 °C and 250 r.p.m. for 2 days (Brunswick Scientific, Innova 44). This culture was used to inoculate 500 mL LB media with appropriate antibiotics and cultured for an additional 24 h. Cultures were then pelleted by centrifugation at 4000 × g for 10 min at 4 °C. Pellets were resuspended by pipetting with a serological pipette in 5% (w/v) sucrose solution plus 0.02% (v/v) Silwet L-77 (Phytotech Labs, S7777). Arabidopsis inflorescences were submerged in the bacterial resuspension for 1 min with gentle agitation, removed and drip dried, and covered gently in plastic wrap before being transferred to the dark overnight. The next day transformed plants were returned to the greenhouse until they produced seeds. T1 seeds were sown on moistened soil and covered with clear plastic lid until cotyledons were visible at which point the lid was removed and the seedlings were sprayed with 50 µM of PPT (Phosphinothricin; GoldBio, P-165-1) twice per week for three weeks until only resistant lines remained. Transgenic T1 plants were grown to seed. T2 seeds were sown on agar plates containing 50 µM PPT. Resistant seedlings were transferred to soil at three weeks and grown to seed. T3 seeds were sown on agar plates containing 50 µM PPT and stable, homozygous lines were validated by segregation.
Arabidopsis phenotypic analysis
The pC-HSL receiver and wild-type Arabidopsis lines were transferred from tissue culture to 6 × 6 × 9 cm pots with Pro-Mix BK25 potting mix (Griffin Greenhouse Supplies, 94-1110). Plant height, number of rosette leaves, number of primary shoots, fresh weight was collected at time of flowering. Leaves and stems were dried in an oven at 60 °C for 1 week and weighed.
Arabidopsis root phenotypic analysis
The pC-HSL receiver and wild-type Arabidopsis seeds were germinated and grown on MS media with 1% sucrose. Seeds were stratified for 3 days at 4 °C, then petri plates were placed vertically in a growth chamber with light intensity at 90 μmol/m2/s, 16/8 h light/dark cycle, and a temperature of 24 °C. Eleven days after sowing seeds, plates were imaged on a ChemiDoc MP Imaging System (Bio-Rad) using a white tray. Images were taken with a 590/110 nm filter at 0.5 s exposure. Root length was measured using ImageJ segmented line tool. Lateral roots were counted from plates, and lateral root density is calculated by dividing lateral roots by total primary root length.
Chemical HSL induction of A. thaliana receivers in the hydroponic system
See Supplementary Fig. 1a. Seeds were surface sterilized with 70% ethanol for 1–2 min, followed by 10% bleach for 10 min, and then rinsed 5 times with water. They were then suspended in 300 µL of 0.1% agar (Sigma, A7921) and sown at 1 cm intervals onto square Petri dishes (Fisherbrand, FB0875711A) of half strength Murashige and Skoog (Sigma, M5519) medium with 1% sucrose, 0.8% agar (Sigma, A7921) and adjusted to pH 5.7 with KOH (MS medium). The plates were placed in the dark at 4 °C for a 3-day striation period before being moved to a growth chamber (Percival Scientific, CU-36L5) and grown for 7–12 days. In a tissue hood, individual wells of 24-well plates (Falcon, 353047) were filled with 1 mL MS medium. HSLs were added to appropriate wells. Finally, plants were carefully lifted from the agar plates with forceps and moved to individual wells such that the roots were entirely submerged. Each plate was covered with a lid, sealed with MicroporeTM tape, and returned to the growth chamber. After 24 h, plates were taken from the growth chamber for imaging.
Confocal microscopy
Microscopy experiments were performed using a Nikon A1R Ultra-Fast Spectral Scanning Confocal Microscope and Andor iXON EMCCD camera or the Leica SP8 Laser Scanning Confocal Microscope. A 4X/0.20 Plan Apo or a 10X/0.30 Plan Fluor air objective was used with the Nikon microscope and a Fluotar Visir 25X/0.95 water objective or a HC PL APO CS2 10x/0.40 air objective was used with the Leica SP8 microscope. For the Nikon microscope, the fluorescence signal was visualized with an excitation wavelength of 488 nm and emission wavelength of 525 nm. To enable comparisons between different days and plant lines, we used the same laser intensities and microscope settings for all experiments performed with the Nikon microscope: the 488 nm laser was used at 10% power, HV 100, and 0 offset. For the Leica SP8 microscope, GFP was visualized with the 488 nm laser at 2% power, 100% gain, emission was detected with the HyD 1 detector between 493 and 529 nm, with 2–4 times line average. Propidium iodide (PI) was visualized with the 534 nm laser at 5% power, 44.3% gain, emission was detected with the HyD 3 detector between 540 and 742 nm, with 2–4 times line average. Upon loading each sample, the entire root system of each plant was inspected, and images were captured only of the brightest portion. For spatiotemporal imaging with the Leica SP8, Arabidopsis roots were stained with 10 μg/ml propidium iodide (Invitrogen, P1304MP) in water. All images were analyzed using the FIJI package of ImageJ. Bright field and fluorescent images were aligned using the Landmark Correspondences plugin. For display purposes, the LUT of the fluorescent channels were inverted in composite images. To make the figures, the minimum and maximum brightness of the LUT were adjusted using the ImageJ default Brightness tool. Note that only the unaltered images were used for MPI quantification. A custom macro was used for fluorescence quantification (Supplementary Fig. 2). A Gaussian filter (σ = 2) was used for noise reduction followed by auto-thresholding using Otsu’s method to separate root tissue from background. The binary mask was then applied to the original image, selecting only the root sections. The mean pixel intensity of just the roots (MPIroots) was calculated using the Measure tool in ImageJ. Finally, the mean pixel intensity of the background (MPIbackground) was calculated and subtracted from the MPIroots to get the final MPI used in the paper.
Quantitative real time PCR analysis for Arabidopsis
Total RNAs were extracted from the seedling with the RNeasy Plant Mini Kit (Qiagen, 74904). cDNAs were generated by the High-Capacity cDNA Reverse Transcription Kit (ThermoFisher Scientific, 4368814). Transcript levels were amplified with the primers listed in Supplementary Table 1 and the reactions were set up by SYBR™ Select Master Mix (Thermo Fisher Scientific, A46109). ACTIN 2 was used as an internal control to normalize expression levels. Quantitative real-time PCRs were performed by the LightCycler 480 (Roche). The sequences of the target gene (primers) are: sfGFP (GAGGGTGAAGGTGACGCAACTAATG, GGACTTGAAGAAGTCATGCTGCTTC) and actin2 housekeeping gene (GTCGTACAACCGGTATTGTGCTG, CCTCTCTCTGTAAGGATCTTCATGAG).
Response functions
Data from multiple plants or multiple bacterial cultures were used to fit a Hill function:
where y is the output (au), x is the concentration of inducer, к is the threshold concentration, and n is the cooperativity (Supplementary Table 1).
Soil preparation
Soil was prepared by mixing three scoops (Grainger, REMCO 82 oz hand scoop, 3UE74) of soil (Lambert Peat Moss Inc, LM-2 Germination Mix, 664980-2325), 1 scoop (Grainger, REMCO 82 oz hand scoop, 3UE74) of vermiculite (Griffin, Whittemore D3 Fine, 65-3120) and one small scoop (30 mL) of Osmocote (The Scotts Miracle-Gro Company, Osmocote 14-14-14, 277960). Soil was sterilized by autoclaving.
Chemical HSL induction of the A. thaliana pC-HSL receiver in soil
See Supplementary Fig. 1c. Seeds were surface sterilized with 70% ethanol for 1–2 min, followed by 10% bleach for 10 min, and then rinsed 5 times with water. They were then suspended in 300 µL of 0.1% agar (Sigma, A7921) and sown at 1 cm intervals onto square Petri dishes (Fisherbrand FB0875711A) of half-strength Murashige and Skoog (Sigma, M5519) medium with 1% sucrose, 0.8% agar (Sigma, A7921) and adjusted to pH 5.7 with KOH (MS media). The plates were placed in the dark at 4 °C for a 3-day striation period before being moved to a growth chamber under the conditions described above and grown for 5 days until the emergence of the first leaf. The soil mixture, prepared as described above, was thoroughly wet with sterile tap water, mixed with a spatula (Cole-Parmer, 17211), and used to fill 10 mL glass beakers (Pyrex, 1000-10). The soil mixture was lightly compressed to provide a firm bed for the seeds. Next, individual seedlings were transplanted into the beakers with forceps. The beakers were placed inside deep Petri dishes (Sigma, P5606-400EA), sealed with MicroporeTM tape, and incubated for 10 days in the growth chamber (Percival Scientific, CU-36L5). After 5 days in the growth chamber, the seedlings were watered by adding 1 mL of autoclaved MilliQ water to each beaker by pipetting and placed back into the growth chamber. After 10 days in the growth chamber, each plant was induced with autoclaved MilliQ water or pC-HSL by watering the plant by pipetting at the plant-soil interface with 1 mL of autoclaved MilliQ water and 1 mL of 100 µM pC-HSL respectively. We estimate the effective concentration of pC-HSL in the soil to be 260 nM (average molarity = (total moles of pC-HSL added)/(total water in soil) = (10−6 M x 10−3 L)/(((4.5 g wet soil-1.74 g dry soil)/1000 g/L) + 1 mL) = 260 nM). The plants were placed back into the growth chamber and incubated for 24 h. After 24 h, the plants roots were cleaned with tap water to remove the soil for imaging.
Genome analysis
To search for HSL producing genes in the P. putida KT2440 (Taxonomy ID: 160488) and K. pneumoniae 342 (Taxonomy ID: 507522) genomes were performed using the protein basic local alignment search tool (BLASTp; blast.ncvi.nlm.nih.gov) using the blastp (protein-protein BLAST) program and default parameters. The query genes were luxI (Vibrio fischeri, NCBI-Protein ID: AAW87994), cinI (Rhizobium leguminosarum, NCBI-Protein ID: WP_018242930), lasI (Pseudomonas aeruginosa, NCBI-Protein ID: QPV56976), traI (Agrobacterium tumefaciens, NCBI-Protein ID: WP_010974838), and rpaI (Rhodopseudomonas palustris, NCBI-Protein ID: WBU30219) which all resulted in no significant similarity found (no sequences were returned).
Potato growth and transformation
Solanum tuberosum (potato) var. ‘Desirée’ was grown in Magenta GA7 vessels (Thomas Scientific, 1190X31) with solid MS Reg medium (4.33 g/L MS basal salt mixture - Phytotech, M524; 25 g/L sucrose - Fisher Science Education, S25590B; 100 mg/L myo-inositol - Sigma, I7508-500G; 170 mg/L sodium phosphate monobasic monohydrate - Sigma, S9638-500G; 440 mg/L calcium chloride dihydrate - Sigma, C7902-500G; 0.9 mg/L thiamine-HCl - Sigma, T1270-25G; 2 mg/L glycine - Thermo Scientific, J16407-36; 0.5 mg/L nicotinic acid - Sigma, N0761-100G; 0.5 mg/L pyridoxine-HCl - Sigma, P2680-25G; 1 × MS vitamins - Phytotech, M553; 3 g/L PhytagelTM - Sigma, P8169-250G; pH 5.7 adjusted with KOH - Sigma, P2680-25G) under fluorescent lights (Sylvania, F34CW/SS/ECO-light intensity 70 m−2 s−1) at ambient temperature134. The pC-HSL receiver plasmid was modified to replace the phosphinothricin resistance cassette with one for hygromycin resistance for selection in potato. The pTT315-Hyg plasmid (Supplementary Fig. 34) was then transformed into A. tumefaciens LBA4404 with the freeze-thaw method135. For potato nuclear transformation, A. tumefaciens cultures were grown to OD600 = 0.6, centrifuged, and resuspended in liquid CIM medium (4.3 g/L MS salt - Polytech, M524; 1 mL/L MSVI vitamins; 1 mL/L JHMS vitamins; 0.1 g/L inositol - Sigma, I7508-500G; 30 g/L sucrose - Fisher Science Education, S25590B; 1 mg/L 6-benzylamino purine (BAP) - Sigma, B3408; 2 mg/L 1-naphthalene-acetic acid (NAA) - Phytotech, N605; 10 g/L agar - BD, 214010; pH 5.7 adjusted with KOH - Sigma, P2680)136. One-month old potato 1 cm internodes were placed on solid CIM media in petri plates with 20 mL of resuspended A. tumefaciens136. After 20 min, internodes were transferred to new solid CIM medium and placed in the dark136. After 48 h, internodes were transferred to 3C5ZR medium (4.3 g/L MS salt - Phytotech, M524; 1 mL/L 3R vitamins solution; 0.1 g/L inositol - Sigma, I7508-500G; 30 g/L sucrose - Fisher Science Education, S25590B; 10 g/L agar - BD, 214010; 0.5 mg/L 3-indoleacetic acid (IAA) - Phytotech, I364; 3 mg/L trans-zeatin-riboside - Phytotech, Z875; 500 mg/L timentin - Phytotech, T869 added after sterilization; pH 5.7 adjusted with KOH - Sigma, P2680-25G) until shoots formed, which were transferred to MS Reg media with 20 mg/L hygromycin (Phytotech, H397) and 200 mg/mL timentin (Phytotech, T869) in Magenta vessels. Five lines were regenerated and confirmed for transgene integration by PCR.
Quantitative real time PCR analysis for potato
Each line was propagated into new media until root formation, and then induced with either pC-HSL or P. putida as described for Arabidopsis. After 24 h, RNA was isolated from roots. cDNA was synthesized from 500 ng of RNA for each plant using ZymoScript RT PreMix Kit (Zymo Research, R3012). qRT-PCR was conducted with PowerUp SYBR Green Master Mix (Thermo Scientific, A25741) on a QuantStudio 3 (Thermo Fisher Scientific) using 1 µL of cDNA. The sequences of the target (primers) are: sfGFP (CTCCAATCGGTGATGGTCCT, GCAGAACCATATGATCGCGT), VP16 domain (TGGACATGTTGGGGGACGG, CTCGAAGTCGGCCATATCCAG), and the Ef1α housekeeping gene (GATGGTCAGACACGTGAACA, CCTTGGAGTACTTGGGGGTG).
Potato phenotypic analysis
The receiver and wild-type potato lines were transferred from tissue culture to 6 × 6 × 9 cm pots with Pro-Mix BK25 potting mix (Griffin Greenhouse Supplies, 94-1110). After 4 weeks, plants were transferred to 11.4 L pots and grown until bolting in a greenhouse. Plant height, fresh weight of leaves and stems, and chlorophyll content were collected at time of bolting. Chlorophyll content was measured with a CCM-200 plus Chlorophyll Content Meter (Opti-Sciences). Leaves and stems were dried in an oven at 60 °C for 1 week and weighed.
pC-HSL sensor characterization in E. coli
Single colonies of E. coli MG1655 sTT658 (Supplementary Table 4) were inoculated into 1 mL LB medium with antibiotics in 2 mL 96 deep-well plates (Thermo, AB-0788), sealed with AeraSeal film (Excel Scientific, BS-25), and grown at 30 °C at 900 rpm. (INFORS HT, Multitron Pro). A 0.5 µL aliquot of overnight culture was diluted into 150 µL LB medium with antibiotics and pC-HSL inducer in 96-well V-bottom plates (Thermo, 249952) and grown at 30 °C at 1000 rpm for 3 h in an EMLI shaker (ELMI, DTS-4). A 3 µL aliquot of the culture was diluted into 200 µL PBS in a round-bottom 96-well plate (Corning, 3797) and analyzed using cytometry.
Flow cytometry
A BD LSR Fortessa flow cytometer with High Throughput Sampler (HTS) attachment (BD Biosciences). At least 10,000 events were captured for each sample and gated by forward and side scatter. Measurements were made using a FITC channel voltage of 450 V, PE-TexasRed channel voltage of 600 V, an FSC voltage of 640 V, and SSC voltage of 289 V. FlowJo and Cytoflow (cytoflow.github.io) were used for analysis and gating. The median fluorescence values were reported.
Measurement of pC-HSL production from P. putida and K. pneumoniae
P. putida pTT337 or K. pneumoniae pTT337 was streaked onto LB agar plates with antibiotics and grown overnight at 30 °C or 37 °C for P. putida and K. pneumoniae respectively. Individual colonies were inoculated into 1 mL of LB medium with appropriate antibiotics in 2 mL 96 deep-well plates grown overnight at 30 °C at 900 rpm (INFORS HT, Multitron Pro). The next morning, the OD600 was measured in a 1 mL cuvette (VWR, 97000-586) in a spectrometer (Agilent Technologies, Cary 60 UV-Vis). The culture was diluted to OD600 = 0.01 into 1 mL of 100% MS medium supplemented with 100 µM of p-coumarate with or without A. thaliana pC-HSL receiver, and grown at 27 °C without shaking for 24 h in the growth chamber (Percival Scientific, CU-36L5). The culture was transferred to 2 mL microfuge tubes and centrifuged at 9600 × g for 5 min. The supernatant was moved to a 96-well filter plate (Whatman, Unifilter 800) and centrifuged at 2200 × g for 5 min. Dilutions of the filtered supernatant were used to induce E. coli MG1655 sTT658 (Supplementary Table 4) in 150 µL LB medium. The E. coli cultures were grown at 30 °C at 1000 r.p.m. for 3 h in an EMLI shaker (ELMI, DTS-4). The cells were diluted into PBS in a round-bottom 96-well plate (Corning, 3797) and analyzed using cytometry. Using the response function of the E. coli pC-HSL sensor strain (E. coli MG1655 sTT658, Supplementary Fig. 19) as a calibration curve, the YFP fluorescence from E. coli MG1655 sTT658 grown in the P. putida or K. pneumoniae supernatants was compared to the YFP fluorescence of the calibration curve (Supplementary Fig. 19). The pC-HSL concentrations of the diluted supernatants was determined by regression.
Bacterial induction of A. thaliana HSL receiver in the hydroponic system
See Supplementary Fig. 1a. Seeds were surface sterilized with 70% ethanol for 1–2 min, followed by 10% bleach for 10 min, and then rinsed five times with water. They were then suspended in 300 µL of 0.1% agar (Sigma, A7921) and sown at 1 cm intervals onto square Petri dishes (Fisherbrand, FB0875711A) of half strength Murashige and Skoog (Sigma, M5519) media with 1% sucrose, 0.8% agar (Sigma, A7921) and adjusted to pH 5.7 with KOH (MS media). The plates were placed in the dark at 4 °C for a 3-day striation period before being moved to a growth chamber under the conditions described above and grown for 7–12 days. Two days before plant inoculation, P. putida or K. pneumoniae strains were streaked from glycerol stocks onto LB agar plates with appropriate antibiotics and grown at 30 °C or 37 °C overnight for P. putida and K. pneumoniae respectively. The next day, individual colonies were selected and inoculated into 1 mL LB medium with appropriate antibiotics in 96 deep-well plates. These plates were grown overnight at 30 °C or 37 °C at 900 r.p.m. (INFORS HT Multitron Pro). The day of plant inoculation, the cultures were spun down for 3 min at 6100 × g, the supernatant was discarded, and the pellet was resuspended in 2 mL of MS medium. This rinsing step was repeated twice to ensure removal of pC-HSL produced overnight. The OD600 was measured in a 1 mL cuvette (VWR, 97000-586) in a spectrometer (Agilent Technologies, Cary 60 UV-Vis). In a tissue hood, individual wells of 24-well plates (Falcon, 353047) were filled with 1000 µL of MS media. P. putida or K. pneumoniae was inoculated in the wells to a starting OD600 of 0.01. For P. putida containing inducible control of HSL production from individual sensors, appropriate chemical inducers were added at the following concentrations: 2 mM IPTG; 1 µM aTc; 1000 ppb. For K. pneumoniae containing the arsenic sensor, a starting OD600 of 0.4 was used and arsenic was added at 1000 ppb. p-coumarate (Sigma, C9008) (100 µM) was also added. Finally, plants were lifted from the agar plates with forceps and moved to individual wells such that the roots were entirely submerged. Each plate was covered with a lid, sealed with MicroporeTM tape, and returned to the growth chamber. After 24 h, plants were removed from each well and placed on a microscope slide (VWR, Micro Slides 48300-026) under a 22x40mm No 1 cover slip (VWR, Cover glass 48393-048) for imaging.
MS agar induction of the A. thaliana pC-HSL receiver
See Supplementary Fig. 1b. Seeds were surface sterilized with 70% ethanol for 1–2 min, followed by 10% bleach for 10 min, and then rinsed 5 times with water. They were then suspended in 300 µL of 0.1% agar (Sigma, A7921) and sown at 1 cm intervals onto square Petri dishes (Fisherbrand, FB0875711A) of half strength Murashige and Skoog (Sigma, M5519) media with 1% sucrose, 0.8% agar (Sigma, A7921) and adjusted to pH 5.7 with KOH (MS media). The plates were placed in the dark at 4 °C for a 3-day striation period before being moved to a growth chamber (Percival Scientific, CU-36L5) and grown for 6 days. MS agar plates sectioned into quadrants (VWR, 25384-348) were prepared, such that the top quadrants would be filled with 1% MS agar without inducer and the bottom quadrants would be filled with MS agar supplemented with either 0 or 100 μM pC-HSL for chemical induction of the A. thaliana pC-HSL receiver. For induction with P. putida, the lower quadrants were filled with 1% MS agar supplemented with 100 μM p-coumarate and OD600 = 0.01 of P. putida WT or P. putida pTT337. Before measuring the OD600, the bacterial cultures were spun down for 3 min at 6100 × g, the supernatant was discarded, and the pellet was resuspended in 2 mL MS medium. This rinsing step was repeated twice to ensure removal of pC-HSL produced overnight. In a tissue hood, the 7-day old seedlings were transferred onto these new agar plates, carefully laying the leaves onto the top quadrants such that no leaf would be in direct contact with either pC-HSL or P. putida and laying the roots onto the bottom quadrants with or without pC-HSL inducer/P. putida. Plates were incubated in the growth chamber for another 24 h into stands to allow the plants to grow vertically. After 24 h, plates were taken from the growth chamber for imaging into the Leica SP8 Laser Scanning Confocal Microscope or from gfp transcript quantification using RT-qPCR.
Bacterial induction of the A. thaliana pC-HSL receiver by seed inoculation in soil
See Supplementary Fig. 1d. Seeds were surface sterilized with 70% ethanol for 1–2 min, followed by 10% bleach for 10 min, and then rinsed 5 times with water. They were then suspended in 300 µL of 0.1% agar (Sigma, A7921) and kept into the dark at 4 °C for a 3-day striation period. Sterile and non-sterile soil mixtures, prepared as described above, were thoroughly wet with sterile tap water, mixed with a spatula (Cole-Parmer, 17211), and used to fill 10 mL glass beakers (Pyrex, 1000-10). The soil mixture was lightly compressed to provide a firm bed for the seeds. After the striation period, the seeds were incubated for 1 h with P. putida WT or P. putida pTT337 grown overnight at 30 °C in LB medium at 900 rpm (INFORS HT, Multitron Pro). The seeds were then transferred to soil (either sterile or non-sterile). The glass beakers were covered with Saran wrap, held into place with elastic bands and 3 puncture holes were made with the tip of sterile forceps to allow for aeration in the beaker. Plants were then incubated for 20 days in the growth chamber (Percival Scientific, CU-36L5) and they were watered with autoclaved MilliQ water supplemented with 100 µM of p-coumarate at day 5 and day 15. After 20 days, the plants roots were cleaned with tap water to remove the soil for imaging.
Bacterial induction of the A. thaliana pC-HSL receiver by watering in soil
See Supplementary Fig. 1e. Seeds were surface sterilized with 70% ethanol for 1–2 min, followed by 10% bleach for 10 min, and then rinsed 5 times with water. They were then suspended in 300 µL of 0.1% agar (Sigma, A7921) and kept into the dark at 4 °C for a 3-day striation period. Sterile and non-sterile soil mixtures, prepared as described above, were thoroughly wet with sterile tap water, mixed with a spatula (Cole-Parmer, 17211), and used to fill 10 mL glass beakers (Pyrex, 1000-10). The soil mixture was lightly compressed to provide a firm bed for the seeds. After the striation period, the seeds were sowed into either sterile or non-sterile soil and incubated for 5 days in the growth chamber. After 4 days, P. putida WT or P. putida pTT337 were grown overnight in LB medium supplemented with antibiotics at 30 °C in LB medium at 900 rpm (INFORS HT, Multitron Pro). The next day, the cultures were spun down by centrifugation for 3 min at 6100 × g and resuspended in 2 mL of PBS. The wash step was repeated twice. The OD600 was measured in a 1 mL cuvette (VWR, 97000-586) in a spectrometer (Agilent Technologies, Cary 60 UV-Vis). The culture was diluted to OD600 = 1 into PBS for a final volume of 2 mL. The 2 mL of the diluted culture were supplemented with 100 µM of p-coumarate was added to the soil. At day 15, this was repeated. After 20 days, the plants roots were cleaned with tap water to remove the soil for imaging.
Characterization of bacterial genetically-encoded sensors
Overnight cultures from individual colonies were grown in 1 mL LB media with antibiotics in 96 deep-well plates (Thermo, AB-0788). A 3 µL aliquot of overnight culture was diluted into 150 µL LB medium with antibiotics and inducer in 96 well V-bottom plates (Thermo, 249952) and grown at 30 °C at 1000 rpm for 4.5 h in an EMLI shaker (ELMI DTS-4). A 3 µL aliquot of the culture was diluted into 200 µL PBS in a round-bottom 96-well plate (Corning, 3797) and analyzed using cytometry.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The P. putida KT2440 (Taxonomy ID: 160488) and K. pneumoniae 342 (Taxonomy ID: 507522) genomes were used to search for homologs of luxI (Vibrio fischeri, NCBI-Protein ID: AAW87994), cinI (Rhizobium leguminosarum, NCBI-Protein ID: WP_018242930), lasI (Pseudomonas aeruginosa, NCBI-Protein ID: QPV56976), traI (Agrobacterium tumefaciens, NCBI-Protein ID: WP_010974838), and rpaI (Rhodopseudomonas palustris, NCBI-Protein ID: WBU30219) query genes using the protein basic local alignment search tool BLASTp [https://blast.ncbi.nlm.nih.gov] with the blastp (protein-protein BLAST) program and default parameters. Plant genetic parts are available as part of Supplementary Data 1. Bacterial genetic parts are available as part of Supplementary Data 2. Plasmid maps are available as part of Supplementary Data 3. Movie files from Supplementary Fig. 9 are available as Supplementary Movies 1–4. All microscopy images used for fluorescence quantification are available in the Supplementary Information at a resolution of 600 dpi and the raw TIFF files are available at Zenodo [https://doi.org/10.5281/zenodo.10601326]. Source data are provided with this paper.
Code availability
The custom Fiji script used for the mean pixel intensity (MPI) calculation is available at Github [https://github.com/VoigtLab/plant-microbe-communication].
References
Mathesius, U. & Watt, M. in Progress in botany 72 125-161 (Springer, 2010).
Chagas, F. O., de Cassia Pessotti, R., Caraballo-Rodriguez, A. M. & Pupo, M. T. Chemical signaling involved in plant-microbe interactions. Chem. Soc. Rev. 47, 1652–1704 (2018).
Peters, N. & Verma, D. Phenolic compounds as regulators of gene expression in plant-microbe interactions. Mol. Plant-Microbe Interact. 3, 8 (1990).
Clear, M.R. & Hom, E.F. The evolution of symbiotic plant-microbe signaling. Annu. Plant Rev. Online 2, 1–52 (2019).
Strehmel, N., Böttcher, C., Schmidt, S. & Scheel, D. Profiling of secondary metabolites in root exudates of Arabidopsis thaliana. Phytochemistry 108, 35–46 (2014).
Badri, D. V., Weir, T. L., van der Lelie, D. & Vivanco, J. M. Rhizosphere chemical dialogues: plant-microbe interactions. Curr. Opin. Biotechnol. 20, 642–650 (2009).
van Dam, N. M. & Bouwmeester, H. J. Metabolomics in the rhizosphere: tapping into belowground chemical communication. Trends Plant Sci. 21, 256–265 (2016).
Hassan, S. & Mathesius, U. The role of flavonoids in root-rhizosphere signalling: opportunities and challenges for improving plant-microbe interactions. J. Exp. Bot. 63, 3429–3444 (2012).
González, O. et al. Non-ribosomal peptide synthases from Pseudomonas aeruginosa play a role in cyclodipeptide biosynthesis, quorum-sensing regulation, and root development in a plant host. Microb. Ecol. 73, 616–629 (2017).
Sharifi, R., Jeon, J.-S. & Ryu, C.-M. Belowground plant-microbe communications via volatile compounds. J. Exp. Bot. 73, 463–486 (2022).
Kumar, V., Baweja, M., Singh, P. K. & Shukla, P. Recent developments in systems biology and metabolic engineering of plant-microbe interactions. Front. Plant Sci. 7, 1421 (2016).
Haldar, S. & Sengupta, S. Plant-microbe cross-talk in the rhizosphere: insight and biotechnological potential. Open Microbiol. J. 9, 1 (2015).
Sanyal, D., Subhash, G. V., Dwivedi, V. & Dasgupta, S. in Climate Change and the Microbiome 695-715 (Springer, 2021).
Imam, J., Singh, P. K. & Shukla, P. Plant microbe interactions in post genomic era: perspectives and applications. Front. Microbiol. 7, 1488 (2016).
Farrar, K., Bryant, D. & Cope‐Selby, N. Understanding and engineering beneficial plant-microbe interactions: plant growth promotion in energy crops. Plant Biotechnol. J. 12, 1193–1206 (2014).
Boo, A., Amaro, R. L. & Stan, G.-B. Quorum sensing in synthetic biology: a review. Curr. Opin. Syst. Biol. 28, 100378 (2021).
Stephens, K. & Bentley, W. E. Synthetic biology for manipulating quorum sensing in microbial consortia. Trends Microbiol. 28, 633–643 (2020).
Voigt, C. A. Genetic parts to program bacteria. Curr. Opin. Biotechnol. 17, 548–557 (2006).
Canton, B., Labno, A. & Endy, D. Refinement and standardization of synthetic biological parts and devices. Nat. Biotechnol. 26, 787–793 (2008).
Tsao, C.-Y., Quan, D. N. & Bentley, W. E. Development of the quorum sensing biotechnological toolbox. Curr. Opin. Chem. Eng. 1, 396–402 (2012).
Vaiana, C. A. et al. Characterizing chemical signaling between engineered “microbial sentinels” in porous microplates. Mol. Syst. Biol. 18, e10785 (2022).
Collins, C. H., Leadbetter, J. R. & Arnold, F. H. Dual selection enhances the signaling specificity of a variant of the quorum-sensing transcriptional activator LuxR. Nat. Biotechnol. 24, 708–712 (2006).
Du, P. et al. De novo design of an intercellular signaling toolbox for multi-channel cell-cell communication and biological computation. Nat. Commun. 11, 4226 (2020).
Scott, S. R. & Hasty, J. Quorum sensing communication modules for microbial consortia. ACS Synth. Biol. 5, 969–977 (2016).
Chen, T., Al-Radhawi, M. A., Voigt, C. A. & Sontag, E. D. A synthetic distributed genetic multi-bit counter. IScience 24, 103526 (2021).
Choudhary, S. & Schmidt-Dannert, C. Applications of quorum sensing in biotechnology. Appl. Microbiol. Biotechnol. 86, 1267–1279 (2010).
Dinh, C. V., Chen, X. & Prather, K. L. Development of a quorum-sensing based circuit for control of coculture population composition in a naringenin production system. ACS Synth. Biol. 9, 590–597 (2020).
Basu, S., Gerchman, Y., Collins, C. H., Arnold, F. H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
Tamsir, A., Tabor, J. J. & Voigt, C. A. Robust multicellular computing using genetically encoded NOR gates and chemical ‘wires’. Nature 469, 212–215 (2011).
Tabor, J. J. et al. A synthetic genetic edge detection program. Cell 137, 1272–1281 (2009).
Danino, T., Mondragón-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010).
Gupta, A., Reizman, I. M. B., Reisch, C. R. & Prather, K. L. Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat. Biotechnol. 35, 273–279 (2017).
Brenner, K., Karig, D. K., Weiss, R. & Arnold, F. H. Engineered bidirectional communication mediates a consensus in a microbial biofilm consortium. Proc. Natl. Acad. Sci. USA 104, 17300–17304 (2007).
Kobayashi, H. et al. Programmable cells: interfacing natural and engineered gene networks. Proc. Natl. Acad. Sci. 101, 8414–8419 (2004).
Billerbeck, S. et al. A scalable peptide-GPCR language for engineering multicellular communication. Nat. Commun. 9, 5057 (2018).
Martin, S. G. Quorum sensing with pheromones. Nat. Microbiol. 4, 1430–1431 (2019).
Khakhar, A., Bolten, N. J., Nemhauser, J. & Klavins, E. Cell-cell communication in yeast using auxin biosynthesis and auxin responsive CRISPR transcription factors. ACS Synth. Biol. 5, 279–286 (2016).
Haskett, T. L. et al. Engineered plant control of associative nitrogen fixation. Proc. Natl. Acad. Sci. USA 119, e2117465119 (2022).
Mondy, S. et al. An increasing opine carbon bias in artificial exudation systems and genetically modified plant rhizospheres leads to an increasing reshaping of bacterial populations. Mol. Ecol. 23, 4846–4861 (2014).
Abdel-Ghany, S. E., Day, I., Heuberger, A. L., Broeckling, C. D. & Reddy, A. S. Production of Phloroglucinol, a Platform Chemical, in Arabidopsis using a Bacterial Gene. Sci. Rep. 6, 38483 (2016).
Fray, R. G. et al. Plants genetically modified to produce N-acylhomoserine lactones communicate with bacteria. Nat. Biotechnol. 17, 1017–1020 (1999).
Geddes, B. A. et al. Engineering transkingdom signalling in plants to control gene expression in rhizosphere bacteria. Nat. Commun. 10, 1–11 (2019).
Kassaw, T. K., Donayre-Torres, A. J., Antunes, M. S., Morey, K. J. & Medford, J. I. Engineering synthetic regulatory circuits in plants. Plant Sci. 273, 13–22 (2018).
Zhang, Z. et al. Inducible Gene Expression Systems in Plants. Plant Gene Trait 11, 1–14 (2020).
Liu, W. & Stewart, C. N. Plant synthetic biology. Trends Plant Sci. 20, 309–317 (2015).
Nemhauser, J. L. & Torii, K. U. Plant synthetic biology for molecular engineering of signalling and development. Nat. Plants 2, 1–7 (2016).
Schaumberg, K. A. et al. Quantitative characterization of genetic parts and circuits for plant synthetic biology. Nat. Methods 13, 94–100 (2016).
Medford, J. I. & Prasad, A. Towards programmable plant genetic circuits. Plant J. 87, 139–148 (2016).
Aoyama, T. & Chua, N. H. A glucocorticoid‐mediated transcriptional induction system in transgenic plants. Plant J. 11, 605–612 (1997).
Beerli, R. R., Schopfer, U., Dreier, B. & Barbas, C. F. Chemically regulated zinc finger transcription factors. J. Biol. Chem. 275, 32617–32627 (2000).
Roslan, H. A. et al. Characterization of the ethanol‐inducible alc gene‐expression system in Arabidopsis thaliana. Plant J. 28, 225–235 (2001).
Zuo, J., Niu, Q.-W. & Chua, N.-H. An estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J. 24, 265–273 (2000).
Bick, M. J. et al. Computational design of environmental sensors for the potent opioid fentanyl. eLife 6, e28909 (2017).
Zuo, J. & Chua, N.-H. Chemical-inducible systems for regulated expression of plant genes. Curr. Opin. Biotechnol. 11, 146–151 (2000).
Corrado, G. & Karali, M. Inducible gene expression systems and plant biotechnology. Biotechnol. Adv. 27, 733–743 (2009).
Morey, K. J. et al. Crosstalk between endogenous and synthetic components-synthetic signaling meets endogenous components. Biotechnol. J. 7, 846–855 (2012).
Verhounig, A., Karcher, D. & Bock, R. Inducible gene expression from the plastid genome by a synthetic riboswitch. Proc. Natl. Acad. Sci. USA 107, 6204–6209 (2010).
Weber, W., Daoud-El Baba, M. & Fussenegger, M. Synthetic ecosystems based on airborne inter-and intrakingdom communication. Proc. Natl. Acad. Sci. 104, 10435–10440 (2007).
Ferluga, S., Steindler, L. & Venturi, V. in Secondary metabolites in soil ecology 69–90 (Springer, 2008).
Calatrava-Morales, N., McIntosh, M. & Soto, M. J. Regulation mediated by N-acyl homoserine lactone quorum sensing signals in the rhizobium-legume symbiosis. Genes 9, 263 (2018).
Chong, T.-M. et al. Characterization of quorum sensing and quorum quenching soil bacteria isolated from Malaysian tropical montane forest. Sensors 12, 4846–4859 (2012).
Rosier, A., Bishnoi, U., Lakshmanan, V., Sherrier, D. J. & Bais, H. P. A perspective on inter-kingdom signaling in plant-beneficial microbe interactions. Plant Mol. Biol. 90, 537–548 (2016).
Begum, J. F., Tamilarasi, M., Pushpakanth, P. & Balachandar, D. A simple method for direct isolation of N-acyl-L-homoserine lactone mediated biofilm-forming rhizobacteria from roots. J. Microbiol. Methods 156, 34–39 (2019).
Steidle, A. et al. Visualization of N-acylhomoserine lactone-mediated cell-cell communication between bacteria colonizing the tomato rhizosphere. Appl. Environ. Microbiol. 67, 5761–5770 (2001).
Schaefer, A. L. et al. A new class of homoserine lactone quorum-sensing signals. Nature 454, 595–599 (2008).
Delalande, L. et al. N-hexanoyl-L-homoserine lactone, a mediator of bacterial quorum-sensing regulation, exhibits plant-dependent stability and may be inactivated by germinating Lotus corniculatus seedlings. FEMS Microbiol. Ecol. 52, 13–20 (2005).
Gantner, S. et al. In situ quantitation of the spatial scale of calling distances and population density-independent N-acylhomoserine lactone-mediated communication by rhizobacteria colonized on plant roots. FEMS Microbiol. Ecol. 56, 188–194 (2006).
Dulla, G. & Lindow, S. E. Quorum size of Pseudomonas syringae is small and dictated by water availability on the leaf surface. Proc. Natl. Acad. Sci. 105, 3082–3087 (2008).
von Rad, U. et al. Response of Arabidopsis thaliana to N-hexanoyl-DL-homoserine-lactone, a bacterial quorum sensing molecule produced in the rhizosphere. Planta 229, 73–85 (2008).
Babenko, L. M., Kosakivska, I. V. & Romanenko, КО Molecular mechanisms of N‐acyl homoserine lactone signals perception by plants. Cell Biol. Int. 46, 523–534 (2022).
Mathesius, U. et al. Extensive and specific responses of a eukaryote to bacterial quorum-sensing signals. Proc. Natl. Acad. Sci. USA 100, 1444–1449 (2003).
Sieper, T. et al. N-acyl-homoserine lactone uptake and systemic transport in barley rest upon active parts of the plant. N. Phytol. 201, 545–555 (2014).
Song, S., Jia, Z., Xu, J., Zhang, Z. & Bian, Z. N-butyryl-homoserine lactone, a bacterial quorum-sensing signaling molecule, induces intracellular calcium elevation in Arabidopsis root cells. Biochem. Biophys. Res. Commun. 414, 355–360 (2011).
Teplitski, M., Mathesius, U. & Rumbaugh, K. P. Perception and degradation of N-acyl homoserine lactone quorum sensing signals by mammalian and plant cells. Chem. Rev. 111, 100–116 (2011).
Uroz, S. & Heinonsalo, J. Degradation of N-acyl homoserine lactone quorum sensing signal molecules by forest root-associated fungi. FEMS Microbiol. Ecol. 65, 271–278 (2008).
Lo, K. J., Lin, S. S., Lu, C. W., Kuo, C. H. & Liu, C. T. Whole-genome sequencing and comparative analysis of two plant-associated strains of Rhodopseudomonas palustris (PS3 and YSC3). Sci. Rep. 8, 12769 (2018).
Palmer, A. G. & Blackwell, H. E. Deciphering a protolanguage for bacteria-host communication. Nat. Chem. Biol. 4, 452–454 (2008).
Hirakawa, H. et al. Activity of the Rhodopseudomonas palustris p-coumaroyl-homoserine lactone-responsive transcription factor RpaR. J. Bacteriol. 193, 2598–2607 (2011).
Kang, S. Y., Lee, J. K., Jang, J. H., Hwang, B. Y. & Hong, Y. S. Production of phenylacetyl-homoserine lactone analogs by artificial biosynthetic pathway in Escherichia coli. Micro. Cell Fact. 14, 191 (2015).
Du, X. et al. Rhodopseudomonas palustris Quorum Sensing Molecule pC-HSL Induces Systemic Resistance to TMV Infection via Upregulation of NbSIPK/NbWIPK Expressions in Nicotiana benthamiana. Phytopathology 111, 500–508 (2021).
Cooley, M., Chhabra, S. R. & Williams, P. N-Acylhomoserine lactone-mediated quorum sensing: a twist in the tail and a blow for host immunity. Chem. Biol. 15, 1141–1147 (2008).
Momb, J., Yoon, D. W. & Fast, W. Enzymic disruption of N-aroyl-L-homoserine lactone-based quorum sensing. Chembiochem 11, 1535–1537 (2010).
Costa-Gutierrez, S. B. et al. Plant growth promotion by Pseudomonas putida KT2440 under saline stress: role of eptA. Appl. Microbiol. Biotechnol. 104, 4577–4592 (2020).
Bernal, P., Allsopp, L. P., Filloux, A. & Llamas, M. A. The Pseudomonas putida T6SS is a plant warden against phytopathogens. ISME J. 11, 972–987 (2017).
Subramoni, S. & Venturi, V. PpoR is a conserved unpaired LuxR solo of Pseudomonas putida which binds N-acyl homoserine lactones. BMC Microbiol. 9, 125 (2009).
Chelius, M. K. & Triplett, E. W. Immunolocalization of Dinitrogenase Reductase Produced by Klebsiella pneumoniae in Association with Zea mays L. Appl. Environ. Microbiol. 66, 783–787 (2000).
Dong, Y., Glasner, J. D., Blattner, F. R. & Triplett, E. W. Genomic interspecies microarray hybridization: rapid discovery of three thousand genes in the Maize Endophyte, Klebsiella pneumoniae 342, by Microarray Hybridization with Escherichia coli K-12 Open Reading Frames. Appl. Environ. Microbiol. 67, 1911–1921 (2001).
Fouts, D. E. et al. Complete Genome Sequence of the N2-Fixing Broad Host Range Endophyte Klebsiella pneumoniae 342 and Virulence Predictions Verified in Mice. PLOS Genet. 4, e1000141 (2008).
Balestrino, D., Haagensen, J. A. J., Rich, C. & Forestier, C. Characterization of Type 2 Quorum Sensing in Klebsiella pneumoniae and Relationship with Biofilm Formation. J. Bacteriol. 187, 2870–2880 (2005).
You, Y.-S. et al. Use of bacterial quorum-sensing components to regulate gene expression in plants. Plant Physiol. 140, 1205–1212 (2006).
Stanton, B. C. et al. Systematic transfer of prokaryotic sensors and circuits to mammalian cells. ACS Synth. Biol. 3, 880–891 (2014).
Gallie, D. R. The 5′‐leader of tobacco mosaic virus promotes translation through enhanced recruitment of eIF4F. Nucleic Acids Res. 30, 3401–3411 (2002).
Meyer, A. J., Segall-Shapiro, T. H., Glassey, E., Zhang, J. & Voigt, C. A. Escherichia coli “Marionette” strains with 12 highly optimized small-molecule sensors. Nat. Chem. Biol. 15, 196–204 (2019).
Dingwall, C. & Laskey, R. A. Nuclear targeting sequences—a consensus? Trends Biochem. Sci. 16, 478–481 (1991).
Harris, S. L., Pelaez, C. A. & Shank, E. A. Monitoring bacterial colonization and maintenance on Arabidopsis thaliana roots in a floating hydroponic system. J. Visual. Exp. 147, e59517 (2019).
Miano, A., Liao, M. J. & Hasty, J. Inducible cell-to-cell signaling for tunable dynamics in microbial communities. Nat. Commun. 11, 1–8 (2020).
Steidle, A. et al. Identification and characterization of an N-acylhomoserine lactone-dependent quorum-sensing system in Pseudomonas putida strain IsoF. Appl. Environ. Microbiol. 68, 6371–6382 (2002).
Bertani, I. & Venturi, V. Regulation of the N-acyl homoserine lactone-dependent quorum-sensing system in rhizosphere Pseudomonas putida WCS358 and cross-talk with the stationary-phase RpoS sigma factor and the global regulator GacA. Appl. Environ. Microbiol. 70, 5493–5502 (2004).
Rampioni, G. et al. The Pseudomonas quorum-sensing regulator RsaL belongs to the tetrahelical superclass of HTH proteins. J. Bacteriol. 189, 1922–1930 (2007).
Park, Y., Espah Borujeni, A., Gorochowski, T. E., Shin, J. & Voigt, C. A. Precision design of stable genetic circuits carried in highly-insulated E. coli genomic landing pads. Mol. Syst. Biol. 16, e9584 (2020).
Sandhi, A., Yu, C., Rahman, M. M. & Amin, M. N. Arsenic in the water and agricultural crop production system: Bangladesh perspectives. Environ. Sci. Pollut. Res. 29, 51354–51366 (2022).
Kayode, O., Aizebeokhai, A. & Odukoya, A. in IOP Conference Series: Earth and Environmental Science, 012081 (IOP Publishing, 2021).
Xu, C., Shi, W. & Rosen, B. P. The chromosomal arsR gene of Escherichia coli encodes a trans-acting metalloregulatory protein. J. Biol. Chem. 271, 2427–2432 (1996).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
EPA. Fact Sheet: Drinking Water Standard for Arsenic. EPA 815-F-00-015. Washington, DC:Office of Water, U.S. Environmental Protection Agency (2001).
Slusarczyk, A. L., Lin, A. & Weiss, R. Foundations for the design and implementation of synthetic genetic circuits. Nat. Rev. Genet. 13, 406–420 (2012).
Lu, T. K., Khalil, A. S. & Collins, J. J. Next-generation synthetic gene networks. Nat. Biotechnol. 27, 1139–1150 (2009).
Brophy, J. A. & Voigt, C. A. Principles of genetic circuit design. Nat. methods 11, 508–520 (2014).
Nielsen, A. A. et al. Genetic circuit design automation. Science 352, aac7341 (2016).
Jones, T. S., Oliveira, S. M., Myers, C. J., Voigt, C. A. & Densmore, D. Genetic circuit design automation with Cello 2.0. Nat. Protoc. 17, 1097–1113 (2022).
Brophy, J. A. et al. Synthetic genetic circuits as a means of reprogramming plant roots. Science 377, 747–751 (2022).
Zhong, V., Archibald, B. N. & Brophy, J. A. Transcriptional and post-transcriptional controls for tuning gene expression in plants. Curr. Opin. Plant Biol. 71, 102315 (2023).
Anderson, J. C., Voigt, C. A. & Arkin, A. P. Environmental signal integration by a modular AND gate. Mol. Syst. Biol. 3, 133 (2007).
Borghi, L. Inducible gene expression systems for plants. Plant Dev. Biol.: Methods Protoc. 655, 65–75 (2010).
Omelina, E. S. et al. Optogenetic and chemical induction systems for regulation of transgene expression in plants: Use in basic and applied research. Int. J. Mol. Sci. 23, 1737 (2022).
Ford, T. J. & Silver, P. A. Synthetic biology expands chemical control of microorganisms. Curr. Opin. Chem. Biol. 28, 20–28 (2015).
Salis, H., Tamsir, A. & Voigt, C. Engineering bacterial signals and sensors. Bact. Sens. Signal. 16, 194–225 (2009).
Brophy, J. A. Toward synthetic plant development. Plant Physiol. 188, 738–748 (2022).
Lau, W., Fischbach, M. A., Osbourn, A. & Sattely, E. S. Key applications of plant metabolic engineering. PLoS Biol. 12, e1001879 (2014).
Kodama, S., Okada, K., Inui, H. & Ohkawa, H. Aryl hydrocarbon receptor (AhR)-mediated reporter gene expression systems in transgenic tobacco plants. Planta 227, 37–45 (2007).
Hsu, T. M. et al. Employing a biochemical protecting group for a sustainable indigo dyeing strategy. Nat. Chem. Biol. 14, 256–261 (2018).
Fahad, S. et al. Phytohormones and plant responses to salinity stress: a review. Plant Growth Regul. 75, 391–404 (2015).
Bauer, W. D. & Robinson, J. B. Disruption of bacterial quorum sensing by other organisms. Curr. Opin. Biotechnol. 13, 234–237 (2002).
Corral-Lugo, A., Daddaoua, A., Ortega, A., Espinosa-Urgel, M. & Krell, T. Rosmarinic acid is a homoserine lactone mimic produced by plants that activates a bacterial quorum-sensing regulator. Sci. Signal 9, ra1 (2016).
Geske, G. D. et al. Comparative analyses of N‐acylated homoserine lactones reveal unique structural features that dictate their ability to activate or inhibit quorum sensing. ChemBioChem 9, 389–400 (2008).
Li, S.-Z. et al. Influence of the d/l configuration of N-acyl-homoserine lactones (AHLs) and analogues on their Lux-R dependent quorum sensing activity. Bioorg. Chem. 77, 215–222 (2018).
Shrestha, A. & Schikora, A. AHL-priming for enhanced resistance as a tool in sustainable agriculture. FEMS Microbiol. Ecol. 96, fiaa226 (2020).
Lin, G.-M., Warden-Rothman, R. & Voigt, C. A. Retrosynthetic design of metabolic pathways to chemicals not found in nature. Curr. Opin. Syst. Biol. 14, 82–107 (2019).
Johns, N. I., Blazejewski, T., Gomes, A. L. & Wang, H. H. Principles for designing synthetic microbial communities. Curr. Opin. Microbiol. 31, 146–153 (2016).
Antunes, M. S. et al. Programmable ligand detection system in plants through a synthetic signal transduction pathway. PLoS One 6, e16292 (2011).
Mus, F. et al. Symbiotic nitrogen fixation and the challenges to its extension to nonlegumes. Appl. Environ. Microbiol. 82, 3698–3710 (2016).
Geddes, B. A. et al. Use of plant colonizing bacteria as chassis for transfer of N 2-fixation to cereals. Curr. Opin. Biotechnol. 32, 216–222 (2015).
Lindsey, B. E., Rivero, L., Calhoun, C. S., Grotewold, E. & Brkljacic, J. Standardized Method for High-throughput Sterilization of Arabidopsis Seeds. J. Vis. Exp. 128, e56587 (2017).
Occhialini, A. et al. Generation, analysis, and transformation of macro-chloroplast Potato (Solanum tuberosum) lines for chloroplast biotechnology. Sci. Rep. 10, 1–13 (2020).
Weigel, D. & Glazebrook, J. Transformation of agrobacterium using the freeze-thaw method. CSH protocols. 7, pdb-prot4666 (2006).
Chronis, D. et al. Potato transformation. Bio-Protoc. 4, e1017–e1017 (2014).
Acknowledgements
We thank the Whitehead Institute and Jing-Ke Weng for access to the greenhouse facility and technical support. We also thank the Microscopy Core at the Koch Institute’s Robert A. Swanson (1969) Biotechnology Center, the Barbara K. Ostrom (1978) Bioinformatics Facility, Jeffrey Kuhn, Jeff Wyckoff and Duanduan Ma for technical support in acquiring and analyzing confocal microscopy images. This material is based upon work supported by the National Science Foundation Graduate Research Fellowship grant #1745302 (T.T.), the Defense Advanced Research Projects Agency-Advanced Plant Technologies grant HR0011-18-2-0049 (S.C.L., C.N.S., C.A.V.), and the MIT Climate Grand Challenges (CGC) program (C.A.V.). USDA Hatch funding to C.N.S. and S.C.L. is acknowledged.
Author information
Authors and Affiliations
Contributions
TT, AB, SCL, CNS and CAV conceived the study and designed the experiments. TT, AB, and AP performed the experiments and analyzed the data. QY provided technical assistance with plant genetic engineering. BF developed the arsenic sensor. TT, AB, and CAV wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks José Dinneny and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boo, A., Toth, T., Yu, Q. et al. Synthetic microbe-to-plant communication channels. Nat Commun 15, 1817 (2024). https://doi.org/10.1038/s41467-024-45897-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-45897-6
- Springer Nature Limited