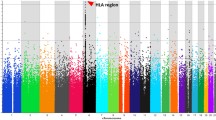
Abstract
Chronic hepatitis B (CHB) is a precursor to liver cirrhosis and hepatocellular carcinoma, caused by a Hepatitis B viral infection. Genome-wide association studies (GWASs) have been conducted to find genes associated with CHB risk. In previous GWAS, EHMT2 was identified as one of the susceptibility genes for CHB. To further characterize this association and discover possible causal variants, we conducted an additional association study. A total of 11 EHMT2 single-nucleotide polymorphisms (SNP) were selected and genotyped in 3902 subjects (1046 CHB patients and 2856 controls). An additional eight imputed SNPs were also included in further analysis. As a result, rs35875104 showed a strong association with the CHB, along with the previously reported genetic marker for CHB risk, rs652888 (odds ratio (OR) = 0.53, P = 2.20 × 10−8 at rs35875104 and OR = 1.58, P = 9.90 × 10−12 at rs652888). In addition, linkage disequilibrium and conditional analysis identified one SNP (rs35875104) as a novel genetic marker for CHB susceptibility. The GRSs (genetic risk scores) were calculated to visualize the combined genetic effects of all known CHB-associated loci, including EHMT2 rs35875104, which was additionally identified in this study. The findings from the present study may be useful for further understanding of the genetic etiology of CHB.
Similar content being viewed by others
Introduction
The hepatitis B virus (HBV) is a global public health problem that affects roughly 350 million people worldwide [1]. HBV infection often leads to chronic hepatitis B (CHB), which may eventually progress to liver failure, liver cirrhosis, or hepatocellular carcinoma (HCC) [2, 3]. The disease is of particular importance to certain Asian regions because the prevalence of HBV infection is higher there compared to other parts of the world [4]. A number of genome-wide association studies (GWASs) of CHB have been conducted recently to identify causal genes of the disease. One Japanese study reported that genes such as HLA-DPA1 and HLA-DPB1 are significantly associated with CHB risk [5]. Another study, also conducted in a Japanese population, found variants of HLA-DQ to be associated with CHB [6]. Moreover, genetic variants of GRIN2A showed association with the risk of CHB in Chinese populations [7]. Previously, our group conducted a GWAS of CHB in a Korean population [8]. In the study, we reported initial associations of genes such as euchromatic histone-lysine-methyltransferase 2 (EHMT2) and transcription factor 19 (TCF19) with the risk of CHB.
EHMT2 encodes a histone methyltransferase, which plays a critical role in catalyzing the transfer of methyl groups in the histone proteins. Histone methylation is an important component of epigenetic gene regulation, and methylated histones have been found to either repress or activate transcription of various genes involved in DNA replication and repair, as well as cell proliferation [9, 10]. Consequently, EHMT2 has been associated with various forms of cancer. A genetic variant of EHMT2 (rs535586) has shown a significant association with the risk of breast cancer and colorectal cancer [11, 12], and higher expressions of EHMT2 were observed in lung and gastric cancers [13, 14]. In addition to liver cancer, epigenetic change has been associated with other liver diseases [15]. On the basis of our previous GWAS, which reported that significant association exists between rs652888 of EHMT2 and CHB risk (P = 2.0 × 10−9), we conducted a fine-mapping association follow-up study to identify possible causal variants. Furthermore, GRSs (genetic risk scores) of all known genetic factors for CHB susceptibility were calculated to evaluate the combined genetic effects regarding CHB risk.
Results
For genotyping of EHMT2 polymorphisms, 11 single-nucleotide polymorphisms (SNPs; one in the promoter region, five in the intron region, and five in the coding region) were selected and genotyped in 3902 study subjects composed of 1046 CHB patients and 2856 healthy controls. Ethnicity of all study subjects was Korean, and study samples did not have any ancestral diversity, which was confirmed by Principal components analysis (PCA; Supplementary Fig. S1). Detailed information about the 11 genotyped SNPs, such as allele, position, minor allele frequency (MAF), heterozygosity, and Hardy–Weinberg equilibrium (HWE), in this study is shown in Supplementary Table S1. Genetic composition of the 11 genotyped SNPs in this study was not significantly different from that of other Asian populations, specifically Chinese and Japanese (Supplementary Table S2). In order to maximize coverage, additional imputation analysis was conducted using the 1000 Genomes database as a reference, and eight EHMT2 SNPs (rs114386644, rs142338646, rs115485095, rs116027812, rs589428, rs570263, rs146903072, and rs605203) were included in further association analysis. Among the 19 investigated polymorphisms, 16 EHMT2 polymorphisms were used for linkage disequilibrium (LD) block construction. Three genetic variants (rs118097312, rs114386644, and rs146726232) were excluded from LD block construction due to its low frequency (MAF <5%). As a result, one LD block was constructed and it contains eight major haplotypes (frequency > 5%; Supplementary Fig. S2).
Logistic association analyses were performed between 19 EHMT2 SNPs and the risk of CHB (Table 1). As a result, four SNPs (rs7887, rs35875104, rs652888, and rs41267090) showed genetic effects on the CHB with a significance (P < 0.05). Among the four SNPs, two in the intron region, including the previously identified CHB-associated genetic marker rs652888, showed strong associations with CHB susceptibility (odds ratio (OR) = 0.53, P = 2.20 × 10−8 at rs35875104 and OR = 1.58, P = 9.90 × 10−12 at rs652888), although no SNPs were associated with HCC after multiple correction. Additional subgroup analyses were conducted to identify the statistical significances of CHB-related HCC progression, and rs35875104 was found to be marginally associated with the CHB-related HCC progression (OR = 0.56, P = 0.04; Table 1). In the haplotype analysis, six haplotypes (ht2, ht3, ht4, ht5, ht7, and ht8) showed genetic effects on the CHB with a significance (P < 0.05). Among the six haplotypes, ht7 and ht8 were tagged with rs652888 and rs35875104, respectively, and showed genetic effects on the CHB in the same direction as rs652888 and rs35875104, respectively (OR = 2.14, P = 7.84×10−15 at ht7 and OR = 0.51, P = 7.58×10−7 at ht8; Supplementary Table S3).
In order to confirm whether the significance of the newly identified genetic marker (rs35875104) was independent or influenced by known CHB risk loci, LD calculation and conditional analysis were conducted with rs35875104 and six known and nearby CHB-susceptible loci (rs9277535 and rs3077 of HLA-DP; rs2856716 and rs7453920 of HLA-DQ; rs1419881 of TCF19; rs652888 of EHMT2). The EHMT2 SNP rs35875104 did not show tight LDs with any known, nearby CHB-susceptible loci (pairwise r 2 ≤ 0.07; Supplementary Fig. S3). After adjusting for already known and nearby CHB-associated loci, rs35875104 still retained its significant association with CHB risk, suggesting an independent genetic effect of rs35875104 on CHB susceptibility (Table 2). Subsequent in silico analysis using SNP FuncPred (https://snpinfo.niehs.nih.gov/snpinfo/snpfunc.html) was conducted to predict the potential role of rs35875104. The SNP rs35875104 has the potential to affect the exon-splicing mechanism by regulating the binding of splicing factors, such as SRp55 and SF2ASFs (Supplementary Table S4).
To investigate the detailed genetic effects of all seven CHB loci (rs9277535 and rs3077 of HLA-DP; rs2856716 and rs7453920 of HLA-DQ; rs1419881 of TCF19; rs35875104 and rs652888 of EHMT2) in a Korean population, referent analysis based on the allele distribution of each SNP was performed. GRSs of genotypes were calculated using the OR from referent analysis (Table 3). To visualize the combined genetic effects of all seven CHB loci, including EHMT2 rs35875104, c ombined genetic effects in individuals were evaluated using the GRSs from Table 3 (Fig. 1 and Supplementary Table S5). The cumulative GRSs ranged from 3.44 (most protected group) to 11.39 (most susceptible group), and CHB patients showed significantly higher cumulative GRSs than healthy controls (Fig. 1a). Furthermore, individuals with higher cumulative GRSs showed significantly increased ORs. In particular, individuals with GRSs less than 4.8 showed an OR of 0.31 (log10 OR = −0.51), while individuals with GRSs over 8.8 showed an OR of 2.41 (log10 OR = 0.38; Fig. 1b).
Combined genetic impact of susceptible alleles from seven CHB genetic markers on the risk of CHB. a A comparison of GRS between CHB patients and PC. The ranges of GRS are shown below the bars. b Odds ratios of different GRS range in log10 scale. Median GRS in controls was used as the reference. GRS genetic risk score, CHB chronic hepatitis B, PC population control
Discussion
The role of histone methyltransferase in EHMT2 influences the transcription of several genes that are related to various types of cancer, such as pancreatic adenocarcinoma, leukemia, breast cancer, and liver cancer [16,17,18,19]. Histone methylation plays dynamic and crucial roles in regulating the chromatin structure. Precise coordination and organization of open and closed chromatin regions control normal cellular processes such as DNA replication, repair, and transcription [20]. A recent study reported that a change in histone methylation may affect mRNA expression of hepatocyte nuclear factor 4α (HNF4α) [21]. HNF4α is a transcription factor found in the liver and is essential for liver development and function. A previous study showed that the overexpression of HNF4α in a human liver cancer cell line transfected with the HBV genome led to a significant increase in viral DNA synthesis [22]. Considering this evidence, a decreased expression of EHMT2 may lead to a loose epigenetic modification and a more abundant expression of HNF4α, which consequently may lead to an increase in HBV synthesis and ultimately, CHB.
In this study, we conducted a follow-up to our previous GWAS by performing a fine-mapping association analysis of EHMT2 to identify possible causal variants. EHMT2 rs652888 was found to have a significant association with CHB risk in previous GWAS studies, and its genetic effect on the CHB was also validated [8, 23, 24]. However, we speculated that it might not be the only causal SNP. Accordingly, 11 EHMT2 SNPs, including rs652888, were selected for genotyping, and additional imputation analysis was conducted. To identify the novel causal variant, LD calculation and conditional analysis were conducted. As a result, one SNP in the intron region, rs35875104, showed an independent genetic effect on CHB, and subsequent in silico analysis was conducted to predict the function of rs35875104. The analysis showed that the rs35875104 variant was located in the exonic splicing enhancer region and promotes exon splicing that can induce exon skipping, and the production of a nonfunctional protein. The C allele of rs35875104 led to binding of splicing factors (SRp55, SF2ASF1, and SF2ASF2) to the motifs, including variant “C” allele of rs35875104 (Supplementary Table S4). Considering that genetic alterations of the splicing factor-binding site can be a critical pathogen in human diseases [25], rs35875104 may have a role in the mechanism related to histone methyltransfer during viral infection.
In addition to the association analyses, combined genetic effects in individuals were evaluated using seven already known CHB-susceptible loci (Supplementary Table S5). The results indicated that seven investigated SNPs in this study might have combined genetic effects on disease susceptibility, as anticipated. When the genetic effects of individual SNPs were considered together, a stronger and more consistent effect on disease could be observed. Consistent with a number of studies that suggest GRS as a significant predictor of disease susceptibility in individuals [26,27,28], the current study may provide prospecting ideas for genetic risk prediction in individuals.
HBV infection is a major factor for HCC development; more than 50% of HCC patients already have CHB [29]. Although the mechanisms of hepatocarcinogenesis in CHB patients are not fully understood, several studies have provided evidence that epigenetic modifications, including histone methylation, might affect HCC development [30, 31]. HBV infection can cause host epigenetic condition changes by affecting histone modification, and several genes, including tumor suppressor gene CDKN2A, have been found to be downregulated with high EHMT2 expression in HCC patients [32]. As a result of analyses conducted to investigate the genetic effects of EHMT2 SNPs on HBV infection and/or HCC progression, two SNPs, rs35875104 and rs652888, showed genetic effects on the HBV infection (OR = 0.54, P = 0.0006 and OR = 1.38, P = 0.001, respectively) and HCC (OR = 0.73, P = 0.02 and OR = 1.24, P = 0.004, respectively). Among them, rs35875104 was found to be marginally associated with the CHB-related HCC progression (OR = 0.56, P = 0.04; Table 1). Considering the importance of epigenetic modification during HCC development, the genetic effects induced by EHMT2 variants on HCC in this study suggest that EHMT2 polymorphisms may have a role in the mechanisms related to HCC progression from HBV infection by affecting epigenetic modification.
Next, we compared the genetic composition of EHMT2 among ethnicities to investigate the ethnic differences in EHMT2 polymorphisms. Frequency analysis and Fisher’s exact test were conducted among various ethnic groups: Korean from the present study and other Asian populations (Chinese and Japanese), as well as African and Caucasian populations from the 1000 Genomes database. Frequencies of minor alleles were found to be more similar among Asian populations (Supplementary Table S3). In the additional PCA analysis, all the study subjects in this study were clustered in the Asian population (Supplementary Fig. S2A). These results suggest that genetic effects of CHB might be influenced by ethnic differences.
There are several limitations in this study, e.g., the use of population controls (PC) whose responses to an HBV infection are unknown, and no information for infection age in case of patients, by which we could not distinguish chronic carriers infected in infancy from those infected in adulthood. Although the population controls would not secure the best statistical power, it would be an alternative way to detect the genetic effects on diseases, when considering the difficulty of recruitment of large number of disease controls. Also, study subjects in this study were mostly the same as in our previous GWAS [8]. Although an independent sample of subjects was not used, the main purpose of this study was to identify the novel causal CHB risk loci that had not been identified in our previous GWAS. Moreover, in the PCA analysis, our study subjects were distinguished from the 1000 Genome project samples of other Asian populations, Chinese and Japanese (Supplementary Fig. S2B). On the basis of the PCA analysis, our study samples did not have any ancestral diversity, which increases the power and validity of the study results.
In conclusion, a follow-up study to a previous GWAS identified a novel genetic marker (rs35875104) for CHB susceptibility. In addition, it confirmed the genetic effect of rs652888 on CHB risk, this study only determined the potential functions for significantly associated variants using in silico analysis and further functional evaluations are required. In addition, the two SNPs were marginally associated with the risk of HCC. Furthermore, combined genetic effects were estimated with known CHB-susceptible loci, and these data might be used to help predict the CHB susceptibility.
Materials and methods
Study subjects
A total of 3902 subjects (1046 cases and 2856 controls) were recruited for the study. CHB patients (n = 1046) were obtained from the outpatient clinic of the Liver Unit and the Center for Health Promotion at Seoul National University Hospital, Ajou University Medical Center, and Ulsan University Hospital (Seoul, Korea). The PC samples (n = 2856) were provided by Korea BioBank, the Center for Genome Science, the National Institute of Health, and Korea Centers for Disease Control and Prevention. The seropositivity of the hepatitis B surface antigen (HBsAg; Enzygnost® HBsAg 5.0; Dade Behring, Marburg, Germany) over a 6-month period was used to diagnose chronic HBV infection (Supplementary Table S6). HBsAg detection is determined using the Enzygnost® HBsAg 5.0 Kit assay. A blood sample of 100 μl is added to a microplate well coated with sheep polyclonal antibodies against HBsAg, and the plate is then loaded into an assay processor, which performs all steps automatically, such as incubation, binding of the antigen, and the washing processes. Diagnosis of HCC was based on imaging findings of nodules that were larger than 1 cm, showing intense arterial uptake, followed by washout of contrast in the venous-delayed phases, in a four-phase multidetector CT scan or dynamic contrast-enhanced MRI and/or biopsy [33]. The ethnicity of all patients and healthy controls was Korean, and there was no ancestral diversity among the study subjects. The study protocol conformed to the Declaration of Helsinki. The study was approved by the institutional review board of Seoul National University Hospital, Ajou University Medical Center, and Ulsan University Hospital. All the subjects participating in the study provided written informed consent.
SNP genotyping
Candidate SNPs of EHMT2 were selected from genotype data from Japanese and Han Chinese populations in the 1000 Genomes database (http://browser.1000genomes.org/index.html) based on the following conditions: (1) MAF > 5%, (2) LD status (LD coefficient (r 2) > 0.98), and (3) low-frequency (MAF ≤ 5%) promoter region and non-synonymous SNPs. A total of 11 EHMT2 SNPs were selected from 105 reported SNPs in the 1000 Genomes Asian population database and were genotyped in 1046 CHB patients and 2856 healthy controls at the multiplex level using the Golden Gate genotyping system (Illumina Inc., San Diego, CA, USA). Approximately 250 ng of genomic materials were used to genotype each sample that had undergone DNA activation, binding to paramagnetic particles, hybridization to oligonucleotides, washing, extension, ligation, amplification by polymerase chain reaction, and hybridization to the Bead plate in an appropriate hybridization buffer. Image intensities were scanned using the BeadXpress® Reader and they were genotyped using the GenomeStudio® software (Illumina Inc.). The genotype quality score for retaining data was set to 0.25. The locations of the genotyped SNPs are shown in Supplementary Fig. S2.
Statistical analysis
LD was obtained using Haploview v4.2 software downloaded from the Broad Institute (http://www.broadinstitute.org/mpg/haploview), with examination of Lewontin’s D’ (|D’|) and the LD coefficient r 2 between all pairs of bi-allelic loci [34]. In order to increase coverage, imputation analysis was performed using IMPUTE2 software [35], with the genotypes from the 1000 Genomes database as a reference. The following criteria were applied to remove improper results from the imputation analysis: (1) call rate > 95%, (2) MAF > 5%, and (3) HWE P value > 0.05. Haplotypes of investigated SNPs were estimated using PHASE software [36]. Logistic regression models were used to compare genotype distributions, including MAF and HWE, among CHB patients and controls, and to calculate ORs, 95% confidence intervals, and corresponding P values adjusted for age (continuous value) and sex (male = 0, female = 1) as covariates using SAS, version 9.4 (SAS Inc., Cary, NC, USA). In corrections for multiple comparisons, Bonferroni correction for multiple testing was applied to the P values based on the number of independent SNPs and the analysis model. Additional subgroup analyses were also performed. Along with the age and sex, liver cirrhosis was also used as a covariate for the association analysis between HCC (+) and HCC (−) cases. In order to investigate whether the association signals of EHMT2 SNPs are independent or affected by known CHB-susceptible loci, conditional logistic regression analyses were performed. Referent model analysis based on the allele distribution of each SNP was also performed to assess the detailed genetic effects. Six known CHB-susceptible loci from our previous study (rs9277535 and rs3077 of HLA-DP; rs2856718 and rs7453920 of HLA-DQ; rs1419881 of TCF19; rs652888 of EHMT2) [8] were used for the conditional analysis and referent analysis. Based on the results from referent analysis, GRSs were calculated by multiplying the number of minor alleles by effect size (OR) of the SNP. Then, the combined genetic effects for each individual were calculated as the sum of the GRSs. We also examined the distribution of cumulative GRSs, ORs, 95% confidence intervals, and corresponding P values between patients and controls. In addition, Fisher’s exact test was conducted using SAS, version 9.4 (SAS Inc.). PCA was conducted to check for possible ethnic diversity using the genotype data from our previous GWAS [8] and the 1000 Genomes database.
References
Kane MA. [World-wide epidemiology of hepatitis B]. Soz Praventivmed. 1998;43(Suppl 1):S98–100 S24-6.
Chu CM, Karayiannis P, Fowler MJ, Monjardino J, Liaw YF, Thomas HC. Natural history of chronic hepatitis B virus infection in Taiwan: studies of hepatitis B virus DNA in serum. Hepatology. 1985;5:431–4.
Hadziyannis SJ, Vassilopoulos D, Hepatitis B e antigen-negative chronic hepatitis B. Hepatology.2001;34:617–24.
Liaw YF, Leung N, Kao JH, Piratvisuth T, Gane E, Han KH, et al Asian-Pacific consensus statement on the management of chronic hepatitis B: a 2008 update. Hepatol Int. 2008;2:263–83.
Kamatani Y, Wattanapokayakit S, Ochi H, Kawaguchi T, Takahashi A, Hosono N, et al A genome-wide association study identifies variants in the HLA-DP locus associated with chronic hepatitis B in Asians. Nat Genet. 2009;41:591–5.
Mbarek H, Ochi H, Urabe Y, Kumar V, Kubo M, Hosono N, et al A genome-wide association study of chronic hepatitis B identified novel risk locus in a Japanese population. Hum Mol Genet. 2011;20:3884–92.
Liu L, Li J, Yao J, Yu J, Zhang J, Ning Q, et al A genome-wide association study with DNA pooling identifies the variant rs11866328 in the GRIN2A gene that affects disease progression of chronic HBV infection. Viral Immunol. 2011;24:397–402.
Kim YJ, Kim HY, Lee JH, Yu SJ, Yoon JH, Lee HS, et al. A genome-wide association study identified new variants associated with the risk of chronic hepatitis B. Hum Mol Genet. 2013;22:4233–8.
Jones PL, Veenstra GJ, Wade PA, Vermaak D, Kass SU, Landsberger N, et al. Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat Genet. 1998;19:187–91.
Jaenisch R, Bird A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet. 2003;33:245–54.
Cebrian A, Pharoah PD, Ahmed S, Ropero S, Fraga MF, Smith PL, et al. Genetic variants in epigenetic genes and breast cancer risk. Carcinogenesis. 2006;27:1661–9.
de Vogel S, Wouters KA, Gottschalk RW, van Schooten FJ, de Goeij AF, de Bruine AP, et al. Genetic variants of methyl metabolizing enzymes and epigenetic regulators: associations with promoter CpG island hypermethylation in colorectal cancer. Cancer Epidemiol Biomarkers Prev. 2009;18:3086–96.
Lee SH, Kim J, Kim WH, Lee YM. Hypoxic silencing of tumor suppressor RUNX3 by histone modification in gastric cancer cells. Oncogene. 2009;28:184–94.
Chen MW, Hua KT, Kao HJ, Chi CC, Wei LH, Johansson G, et al. H3K9 histone methyltransferase G9a promotes lung cancer invasion and metastasis by silencing the cell adhesion molecule Ep-CAM. Cancer Res. 2010;70:7830–40.
Mann DA. Epigenetics in liver disease. Hepatology. 2014;60:1418–25.
Son HJ, Kim JY, Hahn Y, Seo SB. Negative regulation of JAK2 by H3K9 methyltransferase G9a in leukemia. Mol Cell Biol. 2012;32:3681–94.
Dong C, Wu Y, Yao J, Wang Y, Yu Y, Rychahou PG, et al. G9a interacts with Snail and is critical for Snail-mediated E-cadherin repression in human breast cancer. J Clin Invest. 2012;122:1469–86.
Yuan Y, Tang AJ, Castoreno AB, Kuo SY, Wang Q, Kuballa P, et al. Gossypol and an HMT G9a inhibitor act in synergy to induce cell death in pancreatic cancer cells. Cell Death Dis. 2013;4:e690.
Hung SY, Lin HH, Yeh KT, Chang JG. Histone-modifying genes as biomarkers in hepatocellular carcinoma. Int J Clin Exp Pathol. 2014;7:2496–507.
Kouzarides T. Histone methylation in transcriptional control. Curr Opin Genet Dev. 2002;12:198–209.
Zhang Q, Lei X, Lu H. Alterations of epigenetic signatures in hepatocyte nuclear factor 4alpha deficient mouse liver determined by improved ChIP-qPCR and (h)MeDIP-qPCR assays. PLoS ONE. 2014;9:e84925.
Yu X, Mertz JE. Distinct modes of regulation of transcription of hepatitis B virus by the nuclear receptors HNF4alpha and COUP-TF1. J Virol. 2003;77:2489–99.
Jiang DK, Ma XP, Yu H, Cao G, Ding DL, Chen H, et al. Genetic variants in five novel loci including CFB and CD40 predispose to chronic hepatitis B. Hepatology. 2015;62:118–28.
Posuwan N, Payungporn S, Tangkijvanich P, Ogawa S, Murakami S, Iijima S, et al. Genetic association of human leukocyte antigens with chronicity or resolution of hepatitis B infection in thai population. PLoS ONE. 2014;9:e86007.
Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, Yamamoto R, et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature. 2011;478:64–9.
Cheong HS, Lee JH, Yu SJ, Yoon JH, Lee HS, Cheong JY, et al. Association of VARS2-SFTA2 polymorphisms with the risk of chronic hepatitis B in a Korean population. Liver Int. 2015;35:1934–40.
Fava C, Sjogren M, Olsson S, Lovkvist H, Jood K, Engstrom G, et al. A genetic risk score for hypertension associates with the risk of ischemic stroke in a Swedish case-control study. Eur J Hum Genet. 2015;23:969–74.
Zhou XJ, Qi YY, Hou P, Lv JC, Shi SF, Liu LJ, et al. Cumulative effects of variants identified by genome-wide association studies in IgA nephropathy. Sci Rep. 2014;4:4904.
El-Serag HB, Rudolph KL. Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology. 2007;132:2557–76.
Guerrieri F, Belloni L, Pediconi N, Levrero M. Molecular mechanisms of HBV-associated hepatocarcinogenesis. Semin Liver Dis. 2013;33:147–56.
Herath NI, Leggett BA, MacDonald GA. Review of genetic and epigenetic alterations in hepatocarcinogenesis. J Gastroenterol Hepatol. 2006;21:15–21.
Kondo Y, Shen L, Suzuki S, Kurokawa T, Masuko K, Tanaka Y, et al. Alterations of DNA methylation and histone modifications contribute to gene silencing in hepatocellular carcinomas. Hepatol Res. 2007;37:974–83.
Bruix J, Sherman M. Management of hepatocellular carcinoma: an update. Hepatology. 2011;53:1020–2.
Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21:263–5.
Howie BN, Donnelly P, Marchini J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009;5(6):e1000529.
Stephens M, Smith NJ, Donnelly P. A new statistical method for haplotype reconstruction from population data. Am J Hum Genet. 2001;68(4):978–89.
Acknowledgements
We thank the patients and their parents without whom this study would not have been possible. This research was supported by a grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (No. HI16C1074, No. A101023); Korea Science and Engineering Foundation (KOSEF), funded by the Korean government (MEST; No. 2011-0004453); a Sogang University Research Grant for 2011 (SRF-201114006.01); and the SNUH Research Fund (04-2016-0300).
Author contributions
Conceived and designed the experiments: Y.J.K. and H.D.S. Performed the experiments: J.G.S., H.S.C., J.Y.K., S.N., and L.H.K. Analyzed the data: J.G.S., H.S.C., and H.D.S. Provided and cared for study patients: J.H.L., S.J.Y., J.H.Y., J.Y.C., S.W.C., N.H.P., and Y.J.K. Contributed to the writing of the manuscript: J.G.S., Y.J.K., and H.D.S.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Shin, JG., Cheong, H.S., Kim, J.Y. et al. Identification of additional EHMT2 variant associated with the risk of chronic hepatitis B by GWAS follow-up study. Genes Immun 20, 1–9 (2019). https://doi.org/10.1038/s41435-017-0004-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41435-017-0004-x
- Springer Nature Limited
This article is cited by
-
HLA-B*39:01:01 is a novel risk factor for antithyroid drug-induced agranulocytosis in Japanese population
The Pharmacogenomics Journal (2021)
-
Cross-cancer evaluation of polygenic risk scores for 16 cancer types in two large cohorts
Nature Communications (2021)
-
A non-synonymous variant rs12614 of complement factor B associated with risk of chronic hepatitis B in a Korean population
BMC Medical Genetics (2020)