Abstract
The carcinoembryonic antigen (CEA) is the main tumor-associated antigen of colorectal cancers. Previously, we developed a DNA vaccine using scFv6.C4, a CEA surrogate, against CEA-expressing tumors; 40% of the vaccinated mice were tumor-free after tumor challenge. In order to enhance vaccine efficacy, fragment C of Tetanus Toxin (FrC) was tested as adjuvant. C57BL/6J-CEA2682 mice were electroporated intramuscularly 4 times with uP-PS/scFv6.C4-FrC or uP-PS/scFv6.C4, challenged by s.c. injection of 1 × 105 MC38-CEA cells, and tumor growth was monitored over 100 days. The humoral and cellular immune responses were assessed by ELISA, immunocytochemistry, in-vitro lymphocyte proliferation, and CTL cytotoxicity assays. Immunization with uP-PS/scFv6.C4-FrC or uP-PS/scFv6.C4 induced similar anti-CEA antibody titers. However, immunocytochemistry analysis showed stronger staining with uP-PS/scFv6.C4-FrC-immunized mice sera. When challenged with MC38-CEA cells, 63% of the FrC-vaccinated mice did not develop tumors, half of the rest had a significant tumor growth delay, and the probability of being free of tumors was on average 40% higher than that of scFv6.C4-immunized mice. Addition of the adjuvant led to higher CD4+ and CD8+ proliferative responses and strong CD8+ CTL response against MC38-CEA cells. DNA immunization with scFv6.C4 and FrC increased antitumor effect via induction of high and specific humoral and cellular immune responses to CEA.
Similar content being viewed by others
Introduction
Colon and rectum cancers are highly prevalent among men and women, and prevention and treatment are yet major medical and scientific challenges [1, 2]. The carcinoembryonic antigen (CEA) is a 180-KDa membrane-bound highly glycosylated protein, which is the main tumor-associated antigen (TAA) of these cancers [3, 4]. The CEA is highly expressed during the fetal development, but its level is reduced to trace level in the normal adult colonic epithelium [3], which is a characteristic of a self-antigen. However, significant levels of autoantibodies against CEA were detected in some patients with CEA-expressing tumors [5,6,7], evidencing that the CEA self-antigen can activate the immune system.
Previously, our group developed a DNA vaccine against CEA-expressing tumors using scFv6.C4, a CEA surrogate, originated from the anti-idiotypic mAB 6.C4 variable heavy and light chain sequences [8,9,10,11]. Similar approaches with anti-idiotypic antibodies have been used in clinical trials with the production of consistent anti-CEA immune responses [12,13,14].
The scFv6.C4 DNA vaccine was able to both mimic CEA functionally and induce anti-CEA humoral and cellular immune responses [8]. The effectiveness of the preventive immunization with scFv6.C4, as evaluated in transgenic mice for CEA [15], was 40% tumor-free animals after challenge with B16F10-CEA cells. Despite the impressive result, for this vaccine to be translated into a clinical trial, the vaccine efficacy has to be improved.
Tetanus Toxin Fragment C (FrC) from Clostridium tetani has been used as an adjuvant in several DNA vaccine protocols [16,17,18,19,20,21,22]. This 50-kDa fragment has a promiscuous universal epitope for MHC class II in its sequence, which can induce a greater presentation and development of humoral and cellular immune responses [23,24,25]. In preclinical vaccines against lymphomas and melanomas, FrC fused to scFv of idiotypic antibodies increased humoral and cellular responses and caused tumor growth delay [16,17,18]. Additionally, phase-I and -II clinical trials with FrC fused to specific myeloma and prostatic cancer antigens showed safety and tolerance against the vaccines, inducing antigen specific CD8+ T cell responses [26, 27].
In this study, we evaluated the FrC adjuvant effect on DNA vaccination with scFv6.C4 in CEA2682 transgenic mice. Humoral and cellular responses were evaluated in immunized mice before and after challenge with MC38-CEA tumor cells.
Materials and methods
Research Ethics Committee approval
All animal procedures were performed in full compliance with the institutional guidelines, and were approved by the Institutional Research Ethics Committee (http://www.unifesp.br/reitoria/ceua; Approval number: CEUA 703012).
Construction of vectors
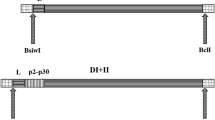
To construct the uP/PS-scFv6.C4-FrC plasmid vector, pFrC plasmid (kindly provided by Dr. Natalia Savelyeva from the Cancer Sciences Unit, Faculty of Medicine, University of Southampton, UK) [18] was used as a template for FrC amplification by PCR using the following primers: 5′-AAGCGGCCGCAAAAAACCTTGATTGTT-3′ and 5′-AATCTAGAGCGGCCGCTTAGTCGTTG-3′ (final concentration: 0.4 mM). The thermocycler was programmed for 95 °C (5 min) and 35 cycles of 95 °C (45 s), 60 °C (45 s), and 72 °C (2 min). After the final cycle, the reaction continued at 72 °C (7 min) and the reaction product was maintained at 4 °C. In the forward and reverse primers, the restriction sites for Not I and Xba I were respectively included, which are underlined. The 1364 bp FrC amplicon was digested with Not I and Xba I, and inserted into the uP-PS/scFv6.C4 plasmid vector [8], which was previously treated with the same enzymes and purified before cloning.
To construct the pT2-scFv6.C4 plasmid vector, uP-PS/scFv6.C4 was digested with Pvu II and Hind III to release the 1165-bp fragment with scFv6.C4 and bGH-PolyA. This fragment was treated with Klenow polymerase and inserted into the pT2 vector [28], which was previously digested with Ava I and treated with Klenow polymerase.
Cell culture, transfection and characterization
The human colorectal carcinoma cell line HCT-8 [29], free of mycoplasma, was maintained in RPMI-1640 (Thermo Fisher Scientific; Waltham, MA, USA), supplemented with 10% heat-inactivated fetal bovine serum (Invitrogen; Grand Island, NY, USA), L-glutamine (Gln; 2 mM), penicillin (100 units/ml), and streptomycin (100 μg/ml). The medium was buffered with sodium bicarbonate (24 mM), plus 2-[4-(2-hydroxyethyl)-1-piperazinyl] ethanesulfonic acid (HEPES) buffer (10 mM) in a humidified atmosphere (5% CO2; 37 °C). The supplemented medium was named RPMIc.
The murine colon adenocarcinoma cell line MC38 [30] was maintained in DMEM (Thermo Fisher Scientific) supplemented with 10% heat-inactivated fetal bovine serum (Invitrogen), Gln (2 mM), penicillin (100 units/ml), and streptomycin (100 μg/ml) in a humidified atmosphere (5% CO2; 37 °C). This medium was buffered with sodium bicarbonate (24 mM) plus HEPES buffer (10 mM). The supplemented medium was named DMEMc.
To promote permanent expression of CEA or scFv6.C4 in MC38 cells, 1 × 105 cells in SMEM (50 μl; Invitrogen) containing pINT (4 μg) [31], and uP-attb-CEA (4 μg) [8], or pSB100x (4 μg) [32] and pT2-scFv6.C4 (4 μg), were transferred into a 4-mm cuvette, and submitted to electroporation (500 V; 150-μs duration; 12 pulses; ECM830, BTX, Harvard Apparatus; Holliston, MA, USA). The transformed cells were named based on the transferred genes (MC38-CEA and MC38-scFv6.C4). The electroporated cells were cloned by limiting dilution, being maintained in DMEMc in a humidified incubator (5% CO2; 37 °C).
The CEA cDNA integration into the cell genome was assessed by PCR using the following primers: 5′- CTA CCT GTG GTG GGT AAA TGG 3′ and 5′- CTT GAC TAT GGA ATT ATT GCG GC -3′ (final concentration: 0.4 mM). The thermocycler was programmed for 95 °C (5 min) and 30 cycles of 95 °C (45 s), 51 °C (45 s), and 72 °C (45 s). After the final cycle, the reaction continued (72 °C; 7 min), and the reaction product was then maintained at 4 °C. The expected amplicon was 418 bp. The human colon cell lines HCT8 and CO112 were used as positive controls, which produce an amplicon of 1131 bp.
The scFv6.c4 integration was assessed by PCR, using the primers 5′-TGGGTGAGGCAGACGCCTGAA-3′ and 5′-GAGGTCCCAGACCCACTGCCA-3′ (final concentration: 0.4 mM). The thermocycler was programmed for 95 °C (5 min) and 30 cycles of 95 °C (45 s), 59 °C (45 s), and 72 °C (45 s). After the final cycle, the reaction continued at 72 °C (7 min) and the reaction product was maintained at 4 °C.
As an internal control for PCR, the glyceraldehyde-3-phosphate dehydrogenase housekeeping gene was used with the following primers: 5′-ACCACAGTCCATGCCATCAC-3′ and 5′-TCCACCACCCTGTTGCTGTA-3′. The PCR products were analyzed by 1% agarose gel electrophoresis.
Preventive DNA vaccination and tumor cell challenge
The CEA-expressing transgenic mice (CEA2682; kindly donated by Dr Wolfgang Zimmermann; University of Munich, Munich, Germany) [15] were immunized by injection of PBS (70 μl using insulin syringe) containing the uP-PS/scFv6.C4-FrC (90 μg) or uP-PS/scFv6.C4 (70 μg) plasmid (in order to maintain the same molar ratio of scFv6.C4) in each quadriceps muscle. Six electric pulses (100 V; duration: 40 ms per pulse; 1 s interval) were applied through 10-mm tweezer electrodes, which were placed around the DNA injection site (Electroporator ECM830). Three subsequent immunizations were performed at two-week intervals. Blood samples were collected 7 days after each immunization to quantify AB3 antibody production.
A week after the last immunization, mice were challenged by s.c. injection of 1 × 105 MC38-CEA cells into the left flank. The tumor size was periodically measured with a caliper for 100 days. Tumor volume (in mm3) was estimated using the following equation:
\({{V}} = 0.52 \times {{D}} \times {{d}}^2\) (D: major diameter; d: minor diameter; in mm)
Quantification of CEA‑specific antibody response
Sera from vaccinated mice were assessed for anti-CEA antibodies (AB3 = AB1′) by ELISA. Initially, CEA (50 µl) in PBS (1 μg/ml; Abcam; Cambridge, UK) was added to each well of 96-well plates and incubated at 37 °C (1 h). Plates were blocked with 1% bovine serum albumin (BSA; Sigma-Aldrich; St. Louis, MO, USA) and 2% non-fat milk in PBS (room temperature, RT; 1 h). The mouse sera diluted in PBS containing 0.1% BSA and 0.2% non-fat milk (1:50) were added and incubated (4 °C; overnight). The wells were washed three times with PBS containing 0.05% Tween-20 (PBS-Tween), and biotinylated rabbit anti-mouse IgG (Dako; Carpinteria, CA, USA) diluted in 0.1% PBS-BSA containing 0.2% non-fat milk (1:1000) was added and incubated (37 °C; 1 h). After washing, horseradish peroxidase-streptavidin (Dako) diluted in 0.1% PBS-BSA containing 0.2% non-fat milk (1:500) was added and incubated (dark chamber; RT; 30 min). The wells were washed before adding ortho-phenylenediamine (OPD; Sigma-Aldrich; 3 mg/ml) in 50 mM citrate-phosphate buffer (51.4 mM Na2HPO4; 24.3 mM acetic acid; pH 5.0) containing 0.03% H2O2. The reactions were stopped with H2SO4 (2 N; 25 mL). The optical density (I: 492 nm) was read in a microplate (Spectra Max M2e, Molecular Devices; Sunnyvale, CA, USA) spectrophotometer. Each sample was run in triplicate.
Immunocytochemistry of HCT‑8, MC38, and MC38-CEA cells
The immunocytochemistry was performed as we described previously [8]. Briefly, on day 0, cells were seeded onto 13-mm coverslips and incubated in a humidified chamber (5% CO2; 24 h). On day 1, media were aspirated, and the cells were fixed in 4% paraformaldehyde (1 h), incubated with NH4Cl (50 mM; 15 min), permeabilized with PBS containing 0.5% Triton X-100 (10 min), blocked with PBS containing 10% BSA (Sigma-Aldrich) and 8% non-fat milk (1 h), and incubated with 1:50 diluted sera from uP-PS/scFv6.C4-FrC immunized, uP-PS/scFv6.C4-immunized, or non-immunized mice in a humidified chamber (5% CO2; 4 °C; overnight). Monoclonal antibody 1F5H2 [10] (final concentration: 10 μg/ml) was used as a positive control. On day 2, the coverslips were incubated with biotinylated rabbit anti-mouse IgG (Dako; 1:100; 1 h), followed by Alexa 594-streptavidin (Invitrogen; 1:1000; 1 h) and 4′,6-diamidino-2-phenylindole (DAPI) for cell nuclei labeling (Invitrogen; 1:1000; 15 min). At the end, coverslips were rinsed with distilled water and mounted with Fluoromount (Sigma-Aldrich), and images were acquired using a fluorescent microscope (Olympus BX51, Tokyo, Japan).
Immunohistochemistry of MC38-CEA tumor
Subcutaneous tumors formed by injection of MC38-CEA cells were removed from the animals after euthanasia, fixed in formaldehyde (10%; 48 h) and dehydrated by sequential passage in 70% ethanol, 100% ethanol and xylol, and infiltrated with paraffin. Sections (4 μm; Leica RM2255 Microtome; Wetzlar, Germany) were dewaxed and hydrated by sequential passage in xylol, 100% ethanol, 70% ethanol, and distilled water. The sections were incubated with NH4Cl (50 mM; 15 min) and washed in distilled water (3 times; 5 min per washing). Permeabilization was done with PBS containing 0.5% Triton X-100 (15 min), followed by washing in PBS. Antigen retrieval was performed in citrate buffer (1.8 mM citric acid; 8.2 mM sodium citrate; pH 6.0) in a steamer (15 min), followed by distilled water washes. Endogenous peroxidase was blocked with a solution of methanol with hydrogen peroxide (30%; 15 min in the dark). The sections were washed in distilled water and blocked in PBS containing 10% BSA and 8% non-fat milk (1 h). After PBS washes, avidin and biotin blockage was performed with Avidin/Biotin Blocking Kit (Vector Laboratories; Burlingame, USA). Biotinylated mAB 1F5H2 [10] (final concentration: 50 μg/ml) was incubated in a humidified chamber (5% CO2; 4 °C; overnight). Next day, the sections were washed with PBS containing 0.25% Tween-20 and incubated with Streptavidin-HRP (Dako; 1:500; 1 h). Peroxidase activity was detected with DAB solution (Dako; Liquid DAB + Substrate Chromogen System) and counterstained with Mayer’s Hematoxylin. The slides were dehydrated by sequential passage in ethanol and xylol, and assembled with Entellan (Sigma-Aldrich). Images were acquired using the fluorescent microscope Olympus BX51.
In vitro cellular proliferation assay
After MC38-CEA injection (90–100 days), mice were euthanized, and splenocytes were collected and treated with ammonium chloride potassium buffer (0.15 M NH4Cl; 10 mM KHCO3; 0.1 mM Na2EDTA) to lyse the red blood cells. For the proliferation assay, 2 × 107 splenocytes suspended in PBS were labeled (37 °C; 5 min) with 2.0 μM carboxyfluorescein diacetate succinimidyl ester (CFSE) (Cell-Trace™; CFSE Cell Proliferation Kit; Invitrogen). Cells were centrifuged (1500 × g; 5 min) and washed three times with RPMIc. In a 96-well plate (U-bottom shape), 5 × 105 cells per well were plated in RPMIc (200 μl). The cells were stimulated with CEA (2 μg/ml; Abcam), mAB 6.C4 (10 μg/ml) [9], or concanavalin A (2.5 μg/ml; ConA; Sigma-Aldrich). Non-stimulated cells were used as a negative control. All experiments were carried out in triplicate. After incubation in a CO2 chamber (37 °C; 6 days), the cells were washed with MACS buffer (PBS containing 2 mM EDTA and 0.5% BSA; pH 7.2) and labeled in a dark chamber (4 °C; 45 min) with anti-CD4-APC (1:100) or anti-CD8-PE (1:100) antibodies (BD Biosciences; San Jose, CA, USA). The labeled cells were washed (three times) and suspended in MACS buffer (200 μl). The samples were quantified by flow cytometry (FACS Canto II, BD Biosciences), and analyzed by the FlowJo (Three Star; Ashland, OR, USA) software. The gate strategy was used to determine the lymphocyte population based on the SSC-A and FSC-A parameters, followed by determination of the CD8+ (SSC-A, PE) and CD4+ (SSC-A, APC) subpopulations. The proliferated cell gate (CFSE low) was established in non-stimulated samples and applied to the stimulated subpopulations. The percentage of CFSE-low cells was normalized by subtracting the percentage observed in non-stimulated samples of the same animal. The samples stimulated with ConA were used as experimental validation (not shown).
Cytotoxicity assay
Splenocytes from mice immunized and challenged with MC38-CEA were prepared as described above. For the cytotoxicity assay, 5–7 × 107 splenocytes were washed in MACS buffer and labelled with the anti-CD8-PE (BD Biosciences) antibody and sorted by flow cytometry (FACS Aria II, BD Biosciences).
The MC38, MC38-CEA, and MC38-scFv6.C4 cells were used as targets. Concomitantly, 2 × 104 tumor cells were plated in duplicates in 96-U-well plates with sorted T CD8+ cells in three different target:effect ratios: 1:5 (10 × 104 T CD8+), 1:2 (4 × 104 T CD8+), and 1:1 (2 × 104 T CD8+). The final volume was completed to 200 µl with RPMIc. The cells were incubated in a humidified chamber (5% CO2; 16 h). Wells containing only tumor cells were used as controls. The LDH activity assay (In Vitro Toxicology Assay Kit, Lactic Dehydrogenase based; Sigma-Aldrich) was performed in 50 µl medium following the manufacturer’s instructions. The lysis percentage was calculated using the following formula: % cell lysis = 100 × [(A − B) / (C − B)], where A: OD690nm tumor plus target, B: OD690nm target spontaneous release (background), and C: OD690nm target maximus lysis.
scFv6.C4 in silico analysis
The complementarity determining regions (CDRs) and structural regions (FR) of scFv6.C4 light and heavy chains were identified through the contact definition and with the IgBLAST algorithm [33, 34]. The forward and reverse sequences of these regions were individually aligned with the amino acid sequence of CEA (NP_004354.3) using the BLASTp and EMBOSS Needle algorithms [35, 36].
Prediction of CD8+ T-cell epitopes was performed in IEDB Analysis Resource (v2.19) using artificial neural network (ANN) method [37], considering the C57BL/6 MHC I alleles H-2-Db and H-2-Kb. Selection of predicted binders was based on the percentile rank of MHC binding affinity ≤ 1% for each combination (MHC allele, length) to cover most of the immune responses [38, 39]. Prediction of linear B-cell epitopes was performed in IEDB Analysis Resource (v2.19) using Bepipred Linear Epitope Prediction [40]. The predicted residues with scores above the threshold were considered as an epitope. The predicted T and B cells’ epitopes with scores above the threshold were individually aligned with the CEA amino acid sequence (NP_004354.3) using the BLASTp and EMBOSS Needle algorithms [35, 36] to assess similarities.
The results were presented as percentage of similarity between sequences, which represents the sum of the percentages of identical and chemically similar amino acids.
Statistical analysis
Statistical analyses were performed using the IBM SPSS Statistics (v. 21; New York, NY, USA) software, and graphs were plotted using the GraphPad Prism (v. 5.0; La Jolla, CA, USA) software. The anti-CEA antibody titers were analyzed by GEE (General Estimating Equations) using linear distribution, defined according to the QIC parameter, followed by Bonferroni. The significance level was set at 5%. The data are presented as mean and standard deviation. Survival analysis was performed by Cox Regression, in which the observed outcome was the time necessary to reach 0.5 cm3 tumor volume. The reported Hazard Ratio (HR) corresponds to the mean of all observed time. The effect of the different groups of animals on the proliferation and cytotoxicity assays were assessed by GLzM using linear distribution, defined according to the AIC parameter, followed by Bonferroni. Sample size was chosen based upon experiences from previous experiments and the numbers of samples per group were indicated in the legends. The significance was 5%. Data are presented as mean and standard deviation.
Results
In vivo tumor model with MC38 cells expressing CEA
To establish a murine colon adenocarcinoma cell line that permanently expresses human CEA, the murine colon adenocarcinoma cell line MC38 was electroporated with the integrative vector system composed of pINT(φC31) and uP-attb-CEA; the CEA-expressing cells were isolated by the limiting dilution technique. The CEA expression on transfected cells were detectable by immunocytochemistry with mAB 1F5H2 [10], even a year after clonal selection (Fig. 1a). Integration of the CEA gene was confirmed by PCR (Fig. 1b). Tumorigenesis of the selected MC38-CEA cells was validated after subcutaneous injection in CEA2682 mice. Visible tumors appeared about 20 days later and reached 1 cm3 40 days after injection, as shown by the standard tumor growth curve with a 95% prediction band (Fig. 1c). Tumors collected after 100 days maintained high CEA expression in most tumor cells (Fig. 1d).
Establishment of the MC38-CEA tumor cell lineage. The MC38 cells were electroporated with uP-attB-CEA and uP-INT (φC31) and cloned by the limiting dilution method. a CEA expression was detected by immunocytochemistry in three time periods: before, right after, and twelve months after clonal selection. CEA staining is shown in red, and nuclei staining in blue (DAPI). Bar = 50 µm. b CEA integration into the genome was analyzed by PCR. 1: 100 bp ladder plus, 2: MC38 (negative control), 3: MC38-CEA, 4: HCT-8 (positive control), 5: CO112 (positive control), 6: reaction without template. The expected amplicon size is 418 pb. c Tumor growth rate was determined after s.c. injection of 1 × 105 MC38-CEA cells into the left flank of C57bl/6-CEA2682 mice. Each point represents the temporal measure of one animal (n = 8 animals). The continuous red line represents the standard tumor growth curve and the dotted lines represent the 95% prediction band d) CEA expression detected by immunohistochemistry in MC38-CEA tumor. Cell nuclei were stained with hematoxylin. Bar = 100 µm
Humoral responses and protection against tumor challenge induced by uP-PS/scFv6.C4-FrC DNA vaccination
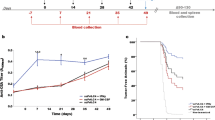
To evaluate the immune response after vaccination with the uP/PS-scFv6.C4-FrC vector, CEA2682 mice were immunized four times with 2-week intervals. The DNA vaccination with plasmid solution was performed by i.m. injection followed by electroporation based on our previous experience [8, 41]. Blood samples were collected 1 week after each immunization to determine the AB3 antibody titers (Fig. 2a). Mice immunized with the uP/PS-scFv6.C4 vector, which was used in a previous study [8], were used for comparison. Although animals immunized with uP/PS-scFv6.C4-FrC had a significant increase in AB3 antibody titer after the first (0.185 ± 0.025 vs 0.132 ± 0.010; P = 0.042), third (0.266 ± 0.028 vs 0.205 ± 0.025; P = 0.002), and fourth (0.322 ± 0.019 vs 0.266 ± 0.028; P = 0.003) immunizations in comparison with the immediately previous measure, no difference was observed in the AB3 titer between animals immunized with uP/PS-scFv6.C4-FrC and uP/PS-scFv6.C4 vectors (Fig. 2b).
Humoral response and protection against MC38CEA tumor challenge after uP-PS/scFv 6.C4-FrC DNA vaccination. a Immunization scheme. The arrows indicate electroporations performed with the uP-PS/scFv 6.C4-FrC vector every 14 days. Blood samples were collected 7 days after each immunization, as indicated by drops. The tumor challenge was performed 7 days after the last immunization, as indicated by cell drawings. The animals were euthanized between days 90 and 130 for cellular and humoral response evaluation. b CEA-specific AB3 antibodies in mice immunized with uP-PS/scFv6.C4 -FrC were detected by ELISA. Blood samples were diluted (1:50) for the assays. Sera from mice immunized with the uP-PS/scFv6.C4 vector were used as controls. Data are expressed as mean ± SD of OD490nm of each group (uP-PS/scFv6.C4-FrC (n = 14) and uP-PS/scFv6.C4 (n = 39)). Number 1: P = 0.042 compared to pre-immune sera; Number 2: P = 0.002 compared to the second immunization; Number 3: P = 0.003 compared to the third immunization. c Tumor-free animals estimated by Cox regression. Mice with tumors smaller than 500 mm3 were considered as survivors. Non-immunized and uP-PS/scFv6.C4-immunized animals were used as controls (uP-PS/scFv6.C4-FrC (n = 11) and uP-PS/scFv6.C4 (n = 17)). The animals were observed for 100 days; however, no change was observed on the survival rate since day 65
To evaluate the effectiveness of the preventive DNA uP-PS/scFv6.C4-FrC vaccine, 1 × 105 MC38-CEA cells were injected subcutaneously into the left flank of mice one week after the last immunization, and tumor growth was measured periodically. In the uP-PS/scFv6.C4-FrC-immunized mice, 63% remained free of tumor over 100 days of observation, and in the uP-PS/scFv6.C4-immunized mice only 47%. All mice without immunization developed tumor within 40 days after tumor injection. Survival analysis by Cox regression shows that the probability of uP-PS/scFv6.C4-FrC immunized mice to be free of tumor is 15.2 times higher than non-immunized mice (CI 95% HR 3.36–68.19; P < 0.001) and 1.4 times higher than the group immunized with uP-PS/scFv6.C4 (CI 95% HR: 0.434–4.58; P = 0.568) (Fig. 2c). These results show that the vaccine regimen can delay tumor growth and increase animal survival, or even can completely inhibit tumor growth. The summary of tumor cell challenge experiments is shown in the Table 1 to facilitate visualization of these results.
To assess the affinity of AB3 antibody on CEA, sera obtained after the last immunization and after tumor challenge were tested by immunocytochemistry on the CEA-expressing human colorectal cell line HCT-8 and murine cell line MC38-CEA. The MC38 cell line was used as a negative control. Pre-immune sera and sera from non-immunized mice were used as negative controls, while sera from the uP-scFv6.C4-immunized mice were used for comparison (Fig. 3). All the CEA-expressing cell lines were strongly marked with sera from uP/PS-scFv6.C4-FrC immunized and tumor-challenged mice, whereas the pre-immune sera or sera from non-immunized mice showed no reaction. Although fluorescent images are difficult to quantify, the staining of uP-scFv6.C4-FrC sera can be clearly observed, mainly in MC38-CEA cells. These results confirm the effectiveness of the scFv6.C4-FrC recombinant protein in mimicking CEA and inducing AB3 specific antibody.
Evaluation of humoral response by immunocytochemistry using sera from uP-PS/scFv6.C4-FrC immunized mice. Blood samples (diluted to 1:50) were collected before immunization, after immunization, and after tumor challenge. The MC38 and HCT-8 human colorectal cells were used as negative and positive controls, respectively. Samples obtained from non-immunized and uP-PS/scFv6.C4-immunized animals were used for comparison. CEA staining is shown in red, and nuclei staining in blue (DAPI). Images are representative of 10 animals per group. Bar = 100 µm
Cellular responses induced by uP‑PS/scFv6.C4-FrC DNA vaccination
To assess the specific T-cell proliferative activities, splenocytes from vaccinated and MC38-CEA-challenged mice were harvested and stimulated with CEA or mAB 6.C4 (Fig. 4). ConA (2.5 μg/ml) and no stimulation were used as positive and negative controls, respectively. As there were mice with and without tumor after vaccination and tumor-challenge, splenocytes from these mice were analyzed separately.
Cellular response of uP/PS-scFv6.C4-FrC immunized animals by proliferation assay. Splenocytes of mice challenged with MC38-CEA were harvested and labeled with 2.0 µM CSFE and stimulated with CEA or mAB 6.C4 for 6 days. CD4 +, and CD8 + CFSE-low cells were counted by flow cytometry. Non-immunized and uP/PS-scFv6.C4-immunized animals were used for comparison. Data are expressed as mean ± SD ((uP-PS/scFv6.C4-FrC (n = 4), uP-PS/scFv6.C4 (n = 8) and non-immunized (n = 5)). **P ≤ 0.01, ***P ≤ 0.001
Under CEA stimulation, a significantly higher proliferation of CD4+ T-cells was observed in scFv6.C4-FrC immunized animals that did not develop any tumors (4.59% ± 0.28%) as compared to animals immunized with scFv6.C4 only (2.06% ± 0.56%; P = 0.003) and non-immunized animals with tumor (1.93% ± 0.35%; P < 0.001). In the animals with tumor, no difference in CD4+ proliferation was observed. On the other hand, CD8+ proliferation was higher in the scFv6.C4-FrC immunized animals that were with tumor (12.50% ± 0.60%), as compared to animals immunized with scFv6.C4 only (1.29% ± 0.46%; P < 0.001) and non-immunized (8.59% ± 0.46%; P < 0.001). Considering the tumor-free animals, CD8+ proliferation in the scFv6.C4 group was higher than in the scFv6.C4-FrC group (8.52% ± 0.74% vs. scFv6.C4-FrC 3.42% ± 0.42%; P < 0.001).
When cells were stimulated with mAB 6.C4, a higher CD4+ proliferation was observed in both scFv6.C4-FrC animals that developed or did not develop tumors as compared to other groups. The main difference (almost 10-fold) was observed in the scFv6.C4-FrC animals that were with tumor (4.36% ± 0.46%) as compared to scFv6.C4 (0.49 ± 0.35%; P < 0.001) and non-immunized animals (0.59 ± 0.30%; P < 0.001). The CD8+ proliferation under mAB 6.C4 stimulus was also significantly higher in scFv6.C4-FrC animals with tumor (10.26 ± 0.60%) as compared to scFv6.C4 (5.65 ± 0.46%; P < 0.001) and non-immunized animals (1.02% ± 0.43%; P < 0.001).
To evaluate the CTL immune response, two target cells (MC38-CEA or MC38-scFv6.C4) were initially constructed to express CEA and scFv6.C4, respectively. Individually sorted CD8+ cells from tumor-challenged animals (which developed or not tumors) were used for cytotoxic assay in different targets: effector ratios, using MC38 cells as a negative control (Fig. 5).
CTL activity in CD8+ cells from uP/PS-scFv6.C4-FrC immunized animals by LDH activity assay. CEA and scFv6.C4-specific CTL lysis was carried out using CD8 + cells sorted by FACS from tumor-challenged animals that developed tumors or the animals that did not develop tumors. The cells were incubated for 16 h together with tumor target cells (target tumor: CD8+ ) in the ratios 1:5, 1:2, and 1:1 before LDH activity assessment. Non-immunized and uP/PS-scFv6.C4-immunized animals were used for comparison. Naive group represents non-immunized and non-challenged animals. MC38 cells were used as a negative control. Data are expressed as mean ± standard deviation, SD (uP-PS/scFv6.C4-FrC (n = 4), uP-PS/scFv6.C4 (n = 4), non-immunized (n = 6) and naïve (n = 4)). ***P ≤ 0.001
Analyzing 1:5 ratio, about 4% of MC38 cells were lysed by CD8+ from non-immunized mice, and this was considered as non-specific basal activity.
A strong CTL activity was observed in CD8+ cells derived from scFv6.C4-FrC-immunized animals that did not develop tumors, with specific lysis of MC38-CEA (77.83 ± 2.24%), which was 15-fold higher than that of scFv6.C4 (5.12 ± 2.24%; P < 0.001), and almost 7-fold higher than in non-immunized and non-challenged animals (naïve group) (12.39 ± 2.59; P < 0.001).
In the group of animals that developed tumors, specific lysis of MC38-CEA cells by CTL from scFv6.C4-FrC immunized animals (31.27 ± 0.91%) was almost 4-fold higher than that from uP/PS-scFv6.C4-immunized animals (8.11 ± 0.91; P < 0.001) and 2.5-fold higher than that from non-immunized animals (12.80 ± 0.65; P < 0.001).
The MC38-scFv6.C4 lysis (1:5 ratio) by cells of tumor-free scFv6.C4-FrC animals (52.45% ± 2.70%) was 5-fold higher than that of uP/PS-scFv6.C4 (10.03 ± 2.70%; P < 0.001). Regarding animals that developed tumors, MC38-scFv6.C4 lysis by CD8+ cells from uP/PS-scFv6.C4-FrC immunized animals (22.76 ± 0.47%) was about 3.6-fold higher than that of uP/PS-scFv6.C4 (6.23 ± 0.47%; P < 0.001) and 2-fold higher than that in non-immunized animals (12.26 ± 0.38%; P < 0.001).
A non-specific lysis of MC38 cells was observed in scFv6.C4-FrC animals that developed (19.06 ± 1.78%) or did not develop (15.82 ± 1.34%) tumors, but both were lower than the lysis observed in MC38-CEA and MC38-scFv6.C4 target cells. An overview of all outcomes after vaccination is schematized in the Fig. 6.
In silico analysis of scFv6.C4
Initially, the amino acid sequences of the complementarity determining regions (CDRs) and framework regions (FRs) of the light and heavy variable chains of scFv6.C4 were deduced based on the IgBLAST algorithm by contact definition (Fig. 7a). Using the BLASTp and EMBOSS Needle algorithms, scFv6.C4 and CEA sequences were aligned and the most significant alignment results are shown in Fig. 7b. Eight scFv6.C4 regions have more than 50% similarities to the CEA sequence; most of them are localized in the light chain. Only two heavy-chain regions show 50% similarity, and 6 light chain regions show similarities varying between 50.0–85.7%. Curiously, the reverse sequences showed more and higher similarity then forward sequences.
In silico analysis of scFv6.C4 regions. a Amino acid sequences of the complementarity determining (CDRs) and framework regions (FRs) of the light and heavy variable chains of scFv6.C4. b scFv6.C4 sequences with more than 50% similarity to CEA. c) scFv6.C4 B-cell predicted binders. Similarity was defined as the sum of the percentages of identical (│) and chemically similar amino acids (:)
Comparing the CEA antigenic epitopes [42,43,44], three scFv6.C4 regions (CDR-H1, CDR-L2-R, and CDR-L3-R) are similar to the CEA A1B1 sequence. Therefore, these regions can be similar to the GOLD IV antigenic region. The CDR-L2, CDR-L1-R, and FR-L1-R sequences have similarity to the CEA A3B3 region, where the GOLD I and III epitopes are present. In addition, scFv6.C4 CDR-H3 and FR-L1 sequences are similar to the CEA A2B2 sequence, which correspond to the GOLD II antigenic region. These results indicate a high collective similarity between the CEA and scFv6.C4 sequences.
The predicted epitopes for T and B cells were inferred from the scFv6.C4 amino acid sequence in IEBD Analysis Resource. Seven MHC-I and eleven B-cell predicted epitopes were selected based on the previously defined parameters. Four MHC-I and three B-cell predicted epitopes aligned with CEA showed similarities higher than 50%. The MHC-I-H2-Kb epitopes 1, 2 and 3 showed similarity to GOLD II antigenic region whereas the epitope 4 to GOLD V. B-cell epitopes 1 was similar to GOLD V, the epitope 2 to GOLD II, and the epitope 3 to GOLD I/III (Fig. 7c).
Discussion
In our previous studies, we described the construction of scFv6.C4, a CEA epitope surrogate isolated from anti-idiotypic mAB 6.C4, and showed its capacity to raise specific humoral and cellular immune responses to CEA [8, 9, 11]. To evaluate the vaccine efficacy of scFv6.C4, CEA2682 transgenic mice were preventively immunized with uP-PS/scFv6.C4 and challenged with B16F10-CEA, a murine melanoma cell line expressing CEA (Denapoli et al., 2016). This vaccination induced sufficient antitumor responses to retard tumor growth and a population of immunized mice remained free of tumor. However, about 60% of them did not survive (Denapoli et al., 2016). The B16F10 tumor cell line is known to be a very aggressive and highly proliferative [45]. In mice, 5 × 105 B16F10-CEA cells injected subcutaneously reaches 200 mm3 in 12 days and 8 × 103 cells reaches the same size in 22 days (Denapoli et al., 2016). After reaching 200 mm3, cell growth rate changes to even faster and becomes very difficult to halt. So, we speculate that the window time for the immune system to fight against the tumor could be very narrow. To evaluate this speculation, here we used a murine colon adenocarcinoma cell lineage expressing CEA (MC38-CEA), which took about 40 days to reach 1 cm3 after injection of 1 × 105 MC38-CEA cells (Fig. 1c), whereas the same number of B16F10-CEA cells reached the same size in less than 15 days [8]. To generate MC38-CEA cell lineage, we used the phiC31 recombinase system because it allows CEA integration into the genome of MC38 cells (Fig. 1b) and promotes a long-term gene expression both in vitro and in vivo (Fig. 1a, d). The slower growth rate of MC38-CEA cells and their proximity to human colorectal tumor have made them the model of choice in many other studies [21, 46,47,48,49, 50,51,52,53,54,55,–56].
However, the challenge of uP/PS-scFv6.C4-immunized mice with MC38-CEA increased the survival rate from 40% (with B16F10-CEA cells) to 50%. In other words, only 10% increase was obtained compared to the challenge with B16F10 cells (Fig. 2c). This result showed the necessity to increase the immune response to CEA-expressing tumors irrespective of the tumor cell line for challenge because the extension of the window of time for the immune response to act on tumor growth affected the survival rate well less than expected.
The uP/PS-scFv6.C4-FrC-immunized mice raised an anti-CEA antibody response in CEA transgenic mice, but the antibody titer was not higher than that of the antibody response induced by scFv6.C4 alone (Fig. 2b). Nevertheless, the staining observed in CEA-expressing tumor cells with sera from the scFv6.C4-FrC-immunized mice was stronger than with sera from scFv6.C4-immunized mice (Fig. 3). The fluorescence intensity after immunocytochemistry staining is the sum of all individual antibody-antigen recognition. As the antibody titer of mice immunized with uP/PS-scFv6.C4-FrC or uP/PS-scFv6.C4 was similar, the stronger immunocytochemistry staining with the serum from uP/PS-scFv6.C4-FrC mice may indicate the presence of more antibodies that recognize other epitopes that can synergize in halting of tumor growth. Several of our results support this hypothesis: 63% of the vaccinated mice did not develop tumors and half of the rest had a significant delay in tumor growth. In addition, the probability of being free of tumors was on average 40% higher than that of mice immunized with scFv6.C4 alone (Fig. 2c).
The cellular immunity of uP/PS-scFv6.C4-FrC-immunized mice was assessed by the proliferation rate and CTL-specific tumor cell lysis assays. The proliferative activity of CD4+ and CD8+ cells in mice immunized with the FrC adjuvant showed to be higher than those immunized with scFv6.C4 alone (Fig. 4). Comparison between scFv6.C4-FrC-immunized mice with and without tumor after tumor challenge showed that there is an apparent increase in CD8+ cell proliferation in the group of mice with tumor. However, sorted CD8+ cells from mice without tumor showed higher CTL activity against the MC38-CEA (77.8%) and MC38-scFv6.C4 (52.5%) cells using the target:effector ratio 1:5 (Fig. 5), which could be achieved with the total splenocytes extract from scFv6.C4-immunized mice only at the 1:50 ratio [8], and this CTL activities were about 10 times higher than other studies using anti-idiotypic CEA vaccines and MC38-CEA tumor model [48]. Therefore, this high and specific CTL activities should be responsible for tumor elimination. The high lymphocyte proliferative activity seen in mice with tumor seems to occur to compensate the low specific lytic activity that was not able to halt tumor progression. Increased protective immunity was seen in the treatment of lymphomas and melanomas after the use of FrC as an adjuvant [17, 18], and cytotoxic response by another group using a plasmid vector carrying FrC fused with CEA for vaccination in wild-type animals [20], but it has never been reported in CEA-expressing transgenic mice yet.
Another important difference between mice with and without tumor after tumor challenge is that the group without tumor showed strong CD4+ and CD8+ proliferative responses with CEA, whereas the mice with tumor showed similar proliferative responses with CEA or mAB 6.C4. The immunization with scFv6.C4 should elicit immune responses against the sequences specific to scFv6.C4 and against the sequences common to both CEA and scFv6.C4 because of their similarities (Fig. 7), however, as the CEA2682 transgenic mice express CEA constitutively, we hypothesized that the endogenous CEA may have favored the selection of clones that are more reactive to CEA than scFv6.C4.
Generation of antibodies and lymphocytes able to recognize CEA after immunization with scFv6.C4 depends on amino acid sequence similarities between CEA and scFv6.C4 regions and epitopes (Fig. 7). In fact, we found eight scFv6.C4 regions, four MHC-I and three B-cell predicted epitopes with high (above 50%) similarities to the CEA epitopes (GOLD I to GOLD V). The mAB 3H1 epitope (surrogate of CEA) is a similar anti-idiotypic epitope that has already been tested clinically. It has only three regions whose similarities are higher than 50% to CEA [55], which is much lower than that found in scFv6.C4. More regions with high similarities between epitope surrogates and CEA should be advantageous in vaccination because it can broaden immune responses and, consequently, more CEA epitopes can be targeted later.
It is important to remind that although vaccination with scFv6.C4 induces the breakdown of tolerance to CEA, which can both cause autoimmune disease and attack their own cells and tissues. However, in our experimental conditions, the immunized mice apparently remained healthy during more than 100 days of follow up. Similar observations were made by others in such vaccine studies [8, 57,58,59].
In conclusion, preventive DNA vaccination of CEA-expressing transgenic mice with the uP-PS/scFv6.C4-FrC vector raised humoral and cellular responses, which were sufficient to maintain 63% of mice free of tumor over 100 days after challenge with MC38-CEA cells against 47% of mice vaccinated with uP-PS/scFv6.C4. The FrC adjuvant activity in delaying and preventing tumor growth is likely to be associated with the induction of high and specific humoral and cellular immune responses to CEA. FrC has proven to be safe and tolerable as a cancer vaccine adjuvant in phase I/II clinical trials [26, 27], as well as in clinical trials for neurological diseases [60]. Therefore, its use as adjuvant with scFv.6C4 for vaccination of patients with CEA-expressing cancer is promising.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62:10–29.
Stewart BW, Wild CP. World cancer report 2014. World Health Organganization. Lyon, France: 2014.
Hammarström S. The carcinoembryonic antigen (CEA) family: structures, suggested functions and expression in normal and malignant tissues. Semin Cancer Biol. 1999;9:67–81.
Thompson JA, Grunert F, Zimmermann W. Carcinoembryonic antigen gene family: molecular biology and clinical perspectives. J Clin Lab Anal U S. 1991;5:344–66.
Ura Y, Ochi Y, Hamazu M, Ishida M, Nakajima K, Watanabe T. Studies on circulating antibody against carcinoembryonic antigen (CEA) and CEA-like antigen in cancer patients. Cancer Lett Neth; Jan De. 1985;25:283–95.
Konstadoulakis MM, Syrigos KN, Albanopoulos C, Mayers G, Golematis B. The presence of anti-carcinoembryonic antigen (CEA) antibodies in the sera of patients with gastrointestinal malignancies. J Clin Immunol setembro De. 1994;14:310–3.
Albanopoulos K, Armakolas A, Konstadoulakis MM, Leandros E, Tsiompanou E, Katsaragakis S, et al. Prognostic significance of circulating antibodies against carcinoembryonic antigen (anti-CEA) in patients with colon cancer. Am J Gastroenterol abril De. 2000;95:1056–61.
Denapoli PMA, Zanetti BF, Dos Santos AA, de Moraes JZ, Han SW. Preventive DNA vaccination against CEA-expressing tumors with anti-idiotypic scFv6.C4 DNA in CEA-expressing transgenic mice. Cancer Immunol Immunother. 2016;66:333–42.
de Moraes JZ, Carneiro CR, Buchegger F, Mach JP, Lopes JD. Induction of an immune response through the idiotypic network with monoclonal anti-idiotype antibodies in the carcinoembryonic antigen system. J Cell Biochem novembro De. 1992;50:324–35.
de Moraes JZ, Gesztesi JL, Westermann P, Le Doussal JM, Lopes JD, Mach JP. Anti-idiotypic monoclonal antibody AB3, reacting with the primary antigen (CEA), can localize in human colon-carcinoma xenografts as efficiently as AB1. Int J Cancer maio De. 1994;57:586–91.
Pignatari GC, Takeshita D, Parise CB, Soares FA, de Moraes JZ, Han SW. Carcinoembryonic antigen (CEA) mimicry by an anti-idiotypic scFv isolated from anti-Id 6.C4 hybridoma. J Biotechnol Neth; Jan De. 2007;127:615–25.
Foon KA, John WJ, Chakraborty M, Das R, Teitelbaum A, Garrison J, et al. Clinical and immune responses in resected colon cancer patients treated with anti-idiotype monoclonal antibody vaccine that mimics the carcinoembryonic antigen. J Clin Oncol. 1999;17:2889–95.
Foon KA, Chakraborty M, John WJ, Sherratt A, Kohler H, Bhattacharyachatterjee M. Immune-response to the carcinoembryonic antigen in patients treated with an antiidiotype antibody vaccine. J Clin Invest. 1995;96:334–42.
Chong G, Bhatnagar A, Cunningham D, Cosgriff TM, Harper PG, Steward W, et al. Phase III trial of 5-fluorouracil and leucovorin plus either 3H1 anti-idiotype monoclonal antibody or placebo in patients with advanced colorectal cancer. Ann Oncol. 2006;17:437–42.
Eades-Perner aM, van der Putten H, Hirth a, Thompson J, Neumaier M, von Kleist S, et al. Mice transgenic for the human carcinoembryonic antigen gene maintain its spatiotemporal expression pattern. Cancer Res. 1994;54:4169–76.
Rice J, Elliott T, Buchan S, Stevenson FK. DNA fusion vaccine designed to induce cytotoxic T cell responses against defined peptide motifs: implications for cancer vaccines. J Immunol. 2001;167:1558–65.
Spellerberg MB, Zhu D, Thompsett A, King CA, Hamblin TJ, Stevenson FK. Promotion of anti-ldiotypic antibody responses induced by single chain Fv genes by fusion to tetanus toxin fragment C. J Immunol. 1997;159:1885–92
King CA, Spellerberg MB, Zhu D, Rice J, Sahota SS. Thompsett a R, et al. DNA vaccines with single-chain Fv fused to fragment C of tetanus toxin induce protective immunity against lymphoma and myeloma. Nat Med. 1998;4:1281–6.
Smahel M, Polakova I, Duskova M, Ludvikova V, Kastankova I. The effect of helper epitopes and cellular localization of an antigen on the outcome of gene gun DNA immunization. Gene Ther Nat Publ Group. 2014;21:225–32.
Lund LH, Andersson K, Zuber B, Karlsson A, Engström G, Hinkula J, et al. Signal sequence deletion and fusion to tetanus toxoid epitope augment antitumor immune responses to a human carcinoembryonic antigen (CEA) plasmid DNA vaccine in a murine test system. Cancer Gene Ther. 2003;10:365–76.
Facciabene A, Aurisicchio L, Elia L, Palombo F, Mennuni C, Ciliberto G, et al. DNA and adenoviral vectors encoding carcinoembryonic antigen fused to immunoenhancing sequences augment antigen-specific immune response and confer tumor protection. Hum Gene Ther. 2006;17:81–92.
Oosterhuis K, Hlschläger P, Van Den Berg JH, Toebes M, Gomez R, Schumacher TN, et al. Preclinical development of highly effective and safe DNA vaccines directed against HPV 16 E6 and E7. Int J Cancer. 2011;129:397–406.
Panina-Bordignon P, Tan A, Termijtelen A, Corradin G, Lanzavecchia A. Universally immunogenic T cell epitopes:promiscuous binding to human MHC class II and promiscuous recognition by T cells. Eur J Immunol. 1989;19:2237–42.
Demotz S, Lanzavecchia A, Eisel U, Niemann H, Widmann C, Corradin G. Delineation of several DR-restricted tetanus toxin T cell epitopes. J Immunol. 1989;142:394–402.
Helting TB, Zwisler O. Structure of Tetanus containing. J Biol Chem. 1977;252:194–8.
Chudley L, McCann K, Mander A, Tjelle T, Campos-Perez J, Godeseth R, et al. DNA fusion-gene vaccination in patients with prostate cancer induces high-frequency CD8+ T-cell responses and increases PSA doubling time. Cancer Immunol Immunother. 2012;61:2161–70.
McCann KJ, Godeseth R, Chudley L, Mander A, Di Genova G, Lloyd-Evans P, et al. Idiotypic DNA vaccination for the treatment of multiple myeloma: Safety and immunogenicity in a phase I clinical study. Cancer Immunol Immunother Springe Berl Heidelb. 2015;64:1021–32.
Aronovich EL, Bell JB, Belur LR, Gunther R, Koniar B, Erickson DCC, et al. Prolonged expression of a lysosomal enzyme in mouse liver after Sleeping Beauty transposon-mediated gene delivery: Implications for non-viral gene therapy of mucopolysaccharidoses. J Gene Med. 2007;9:403–15.
Tompkins WA, Watrach AM, Schmale JD, Schultz RM, Harris JA. Cultural and antigenic properties of newly established cell strains derived from adenocarcinomas of the human colon and rectum. J Natl Cancer Inst abril De. 1974;52:1101–10.
Corbett TH, Griswold DPJ, Roberts BJ, Peckham JC, Schabel FMJ. Tumor induction relationships in development of transplantable cancers of the colon in mice for chemotherapy assays, with a note on carcinogen structure. Cancer Res setembro De. 1975;35:2434–9.
Stilhano RS, Martin PKM, de Melo SM, Samoto VY, Peres GB, da Silva Michelacci YMC, et al. alpha- L-iduronidase gene-based therapy using the phiC31 system to treat mucopolysaccharidose type I mice. J Gene Med Engl. 2015;17:1–13.
Martin PKM, Stilhano RS, Samoto VY, Takiya CM, Peres GB, da Silva Michelacci YMC, et al. Mesenchymal stem cells do not prevent antibody responses against human alpha-L-iduronidase when used to treat mucopolysaccharidosis type I. PLoS One. 2014;9:e92420.
MacCallum RM, Martin AC, Thornton JM. Antibody-antigen interactions: contact analysis and binding site topography. J Mol Biol outubro De. 1996;262:732–45.
Ye J, Ma N, Madden TL, Ostell JM. IgBLAST: an immunoglobulin variable domain sequence analysis tool. Nucleic Acids Res Engl; julho De. 2013;41:W34–40.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol outubro De. 1990;215:403–10.
McWilliam H, Li W, Uludag M, Squizzato S, Park YM, Buso N, et al. Analysis Tool Web Services from the EMBL-EBI. Nucleic Acids Res. 2013;41(Web Server issue):W597–600.
Nielsen M, Lundegaard C, Worning P, Lauemoller SL, Lamberth K, Buus S, et al. Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Protein Sci. 2003;12:1007–17.
Moutaftsi M, Peters B, Pasquetto V, Tscharke DC, Sidney J, Bui H-H, et al. A consensus epitope prediction approach identifies the breadth of murine T(CD8+ )-cell responses to vaccinia virus. Nat Biotechnol. 2006;24:817–9.
Kotturi MF, Peters B, Buendia-Laysa FJ, Sidney J, Oseroff C, Botten J, et al. The CD8+T-cell response to lymphocytic choriomeningitis virus involves the L antigen: uncovering new tricks for an old virus. J Virol. 2007;81:4928–40.
Larsen JEP, Lund O, Nielsen M. Improved method for predicting linear B-cell epitopes. Immunome Res. 2006;2:2.
Parise CB, Lisboa B, Takeshita D, Sacramento CB, de Moraes JZ, Han SW. Humoral immune response after genetic immunization is consistently improved by electroporation. Vaccines. 2008;26:3812–7.
Hammarstrom S, Shively JE, Paxton RJ, Beatty BG, Larsson a, Ghosh R, et al. Antigenic sites in carcinoembryonic antigen. Cancer Res. 1989;49:4852–8.
Bjerner J, Lebedin Y, Bellanger L, Kuroki M, Shively JE, Varaas T, et al. Protein epitopes in carcinoembryonic antigen. Tumor Biol. 2002;23:249–62.
Murakami M, Kuroki M, Arakawa F, Kuwahara M, Oikawa S, Nakazato H, et al. A reference of the GOLD classification of monoclonal antibodies against carcinoembryonic antigen to the domain structure of the carcinoembryonic antigen molecule. Hybridoma. 1995;14:19–28.
Sharma A, Kuzu OF, Nguyen FD, Sharma A, Noory M. Current state of animal (mouse) modeling in melanoma research. Cancer Growth Metastas-. 2015;8:81.
Pervin S, Chakraborty M, Bhattacharya-Chatterjee M, Zeytin H, Foon KA, Chatterjee SK. Induction of antitumor immunity by an anti-idiotype antibody mimicking carcinoembryonic antigen. Cancer Res. 1997;57:728–34.
Saha A, Chatterjee SK, Foon KA, Bhattacharya-Chatterjee M. Anti-idiotype antibody induced cellular immunity in mice transgenic for human carcinoembryonic antigen. Immunology. 2006;118:483–96.
Saha A, Chatterjee SK, Foon KA, Celis E, Bhattacharya-Chatterjee M. Therapy of established tumors in a novel murine model transgenic for human carcinoembryonic antigen and HLA-A2 with a combination of anti-idiotype vaccine and CTL peptides of carcinoembryonic antigen. Cancer Res. 2007;67:2881–92.
Schwegler C, Dorn-Beineke A, Nittka S, Stocking C, Neumaier M. Monoclonal anti-idiotype antibody 6G6.C4 fused to GM-CSF is capable of breaking tolerance to carcinoembryonic antigen (CEA) in CEA-transgenic mice. Cancer Res. 2005;65:1925–33.
Kass E, Schlom J, Thompson J, Guadagni F, Graziano P, Greiner JW. Induction of protective host immunity to carcinoembryonic antigen (CEA), a self-antigen in CEA transgenic mice, by immunizing with a recombinant vaccinia-CEA virus. Cancer Res. 1999;59:676–83.
Facciabene A, Aurisicchio L, Elia L, Palombo F, Mennuni C, Ciliberto G, et al. Vectors encoding carcinoembryonic antigen fused to the B subunit of heat-labile enterotoxin elicit antigen-specific immune responses and antitumor effects. Vaccine. 2007;26:47–58.
Kass E, Panicali DL, Mazzara G, Schlom J, Greiner JW. Granulocyte/macrophage-colony stimul factor prod recomb avian poxviruses enriches reg lymph nodes antigen-present cells acts immunoadjuvant. Cancer Res. 2001;27:206–14.
Lurquin C, Lethe B, De Plaen E, Corbiere V, Theate I, van Baren N, et al. Contrasting frequencies of antitumor and anti-vaccine T cells in metastases of a melanoma patient vaccinated with a MAGE tumor antigen. J Exp Med. 2005;201:249–57.
Disis ML, Shiota FM, Cheever MA. Human HER-2/neu protein immunization circumvents tolerance to rat neu: a vaccine strategy for “self” tumour antigens. Immunology. 1998;93:192–9.
Chatterjee SK, Tripathi PK, Chakraborty M, Yannelli J, Wang H, Foon KA, et al. Molecular mimicry of carcinoembryonic antigen by peptides derived from the structure of an anti-idiotype antibody. Cancer Res. 1998;1217–25.
Gaida FJ, Pieper D, Roder UW, Shively JE, Wagener C, Neumaier M. Molecular characterization of a cloned idiotypic cascade containing a network antigenic determinant specific for the human carcinoembryonic antigen. J Biol Chem. 1993;268:14138–45.
Greiner JW, Zeytin H, Anver MR, Schlom J. Vaccine-based therapy directed against carcinoembryonic antigen demonstrates antitumor activity on spontaneous intestinal tumors in the absence of autoimmunity. Cancer Res. 2002;62:6944–51.
Hodge JW, Poole DJ, Aarts WM, Yafal AG, Gritz L, Schlom J. Modified vaccinia virus ankara recombinants are as potent as vaccinia recombinants in diversified prime and boost vaccine regimens to elicit therapeutic antitumor responses. Cancer Res. 2003;63:7942–9.
Saha A, Chatterjee SK, Foon KA, Primus FJ, Sreedharan S, Mohanty K, et al. Dendritic cells pulse anti-idiot antib mimicking carcinoembryonic antigen (cea) can reverse immunol toler cea induce antitumor immun CEA transgenic mice. Cancer Res. 2004;7:4995–5003.
Toivonen JM, Oliván S, Osta R. Tetanus toxin c-fragment: The courier and the cure? Toxins (Basel). 2010;2:2622–44.
Acknowledgements
This work was supported by São Paulo Research Foundation (FAPESP; Grant Numbers: 2012/21861-1 and # 2013/17224-9). BFZ was a recipient of FAPESP scholarship (2012/21861-1).
Authors’ contributions
BFZ: conception and design of the study, acquisition of data, analysis and interpretation of the data, drafting of the manuscript; CPF: acquisition of data, analysis and interpretation of data; JRCV: analysis and interpretation of the data; SWH: conception and design of the study, drafting of the manuscript, critical review of the manuscript for important intellectual content, and final approval of the version to be submitted.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zanetti, B.F., Ferreira, C.P., de Vasconcelos, J.R.C. et al. scFv6.C4 DNA vaccine with fragment C of Tetanus toxin increases protective immunity against CEA-expressing tumor. Gene Ther 26, 441–454 (2019). https://doi.org/10.1038/s41434-019-0062-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41434-019-0062-y
- Springer Nature Limited