Abstract
Purpose
Bardet-Biedl syndrome (BBS) is a rare multisystem ciliopathy. The aim of this study was to describe the clinical and genetic features of a cohort of Chinese patients carrying biallelic BBS gene variants.
Methods
We recruited 34 patients from 31 unrelated pedigrees who carried biallelic pathogenic variants in BBS genes. All patients underwent ophthalmic and systematic evaluations, as well as comprehensive molecular genetic analyses. Ultimately, 14 patients were followed up over time.
Results
We identified 47 diseasing-causing variants in 10 BBS genes; 33 were novel. Diagnosis of BBS and non-syndromic retinitis pigmentosa (RP) were established in 28 patients from 27 pedigrees and 6 patients, respectively. The two most prevalent genes in patients with BBS were BBS2 and BBS4, accounting for 51.8% of the probands. The patients exhibited clinical heterogeneity, from patients with all six primary clinical components to patients suffering from non-syndromic RP. The common components were retinal dystrophy, polydactyly, and obesity, with frequencies of 78.6% to 100%, while renal anomaly frequencies were only 7.1%. Patients exhibited early and severe visual defects and retinal degeneration. Patients with biallelic missense variants in BBS2 suffered fewer clinical symptoms and mild visual impairment. Patients with BBS10 variants tended to have cone dystrophy.
Conclusions
Our study defined the mutated gene profiles and established the configuration of the variation frequencies for each BBS gene in Chinese patients. Overall, our patients showed early and severe visual defects and retinal degeneration. Genetic analysis is therefore crucial for diagnosis, genetic counseling, and future gene therapy in these patients.
Similar content being viewed by others
Introduction
Bardet-Biedl syndrome (BBS) is a rare multisystem ciliopathy with a variable prevalence between populations ranging from 1:100,000 in North America and Europe to 1:18,000 and 1: 13,500 in Newfoundland and Kuwait [1]. Patients with BBS usually have six primary clinical features: retinal dystrophy, postaxial polydactyly, obesity, intellectual disability, renal anomalies (RA), and genitourinary and reproductive abnormalities [1, 2]. Patients also show secondary features, such as speech disorder, brachydactyly/syndactyly, developmental delay, ataxia, diabetes, strabismus/cataracts/astigmatism, dental anomalies (dental crowding/hypodontia/high arched palate), and congenital heart defects [1, 2]. Patients display interfamilial and intrafamilial phenotypic variability in their clinical phenotypes. Clinically, the diagnosis of BBS is established by the presence of at least four primary features or three primary features in combination with two secondary features [3].
BBS is inherited in an autosomal recessive pattern and has high genetic heterogeneity. Currently, at least 21 BBS-causing genes (BBS1-21) have been identified [1, 2]. The BBS genes encode proteins whose functions are directly correlated with the primary cilium, such as the regulation of ciliary structure, biogenesis, and function; therefore, BBS pathogenesis is mainly associated with ciliary dysfunction [1, 2]. Of the BBS genes, eight (BBS1, 2, 4, 5, 7, 8, 9, and 18) encode a protein complex known as BBSome, which is essential for mediating ciliary trafficking activity [1, 2]. Three BBS genes (BBS6, 10, and 12) encode proteins that form a chaperone-like protein complex that is responsible for the BBSome assembly. The BBS3 gene encodes a small GTPase, which recruits the BBSome to membranes [4]. The remaining genes are involved in the maintenance of ciliary homeostasis and are referred to as non-canonical BBS genes [5]. The BBS genes show diverse mutational profiles, and variants in BBS1-12, 15, 17-19, 21, and 22 are mainly responsible for BBS, but variants of the genes BBS1, 2, 3, 8, 14, 21 can also sometimes caused non-syndromic retinal dystrophy (isolated retinal dystrophy) [6]. Variants of three genes (BBS13, 14, and 16) can also cause other syndromic ciliopathies, such as Joubert syndrome, Meckel-Gruber syndrome, and Senior-Løken syndrome, whose phenotypes partially genetic overlap with BBS [1].
A very recent meta-analysis study revealed that more than half of the patients with BBS carried variants of one of the eight BBSome encoding genes, followed by patients (28%) who harbored causative variants of the three genes encoding chaperone-like proteins, patients (15%) who had non-canonical BBS genes, and patients (5%) with variants of other BBS genes [7].
Several previous studies revealed that the mutated gene profile varied in different ethnic origin patients [8,9,10,11,12,13]. For instance, BBS1 and BBS10 genes are the two prevalent genes for patients in North American and Northern Europe, accounting for 23% and 20% of the cases, respectively [8, 9, 11, 12]. By contrast, the most common genes for Pakistani patients are BBS10 and BBS5, accounting for 27% and 18%, respectively [10]. The prevalence of BBS in the Chinese population is not yet known, and studies on Chinese patients with BBS are rare.
In the current study, we described the clinical and genetic features of 34 patients from 31 unrelated families who carry biallelic variants in one of the BBS genes. We reported the gene profiles for Chinese patients with BBS and evaluated the probable genotype and phenotype correlations.
Methods
Subjects
This study was conducted strictly in accordance with the tenets of the Declaration of Helsinki and was approved by the Beijing Tongren Hospital Joint Committee on Clinical Investigation. In this study, 34 patients from 31 unrelated families were recruited at the Genetics Laboratory of Beijing Institute of Ophthalmology, Beijing Tongren Ophthalmic Center, during 2010 to 2021. The criteria for inclusion were that patients had retinal dystrophy and carried biallelic pathogenic or likely pathogenic variants in one of 18 BBS genes (BBS1-12, 15, and 17–21). We excluded three genes (BBS13, 14, and 16), as variants of these genes can also cause Joubert syndrome, Meckel-Gruber syndrome, and Senior-Løken syndrome, whose phenotypes overlap with BBS. We defined the recruited patients as having either BBS or isolated retinal dystrophy (non-syndromic retinal dystrophy). Diagnosis of BBS was based on the method described by Imhoff et al. [14]. whereby patients had at least two primary clinical features of BBS and harbored biallelic variants in one of the BBS genes.
All patients underwent standard ocular examinations, including best-corrected visual acuity (BCVA) using E decimal charts, slit-lamp biomicroscopy, and fundus photography (FP). Most patients were subjected to retinal optical coherence tomography (OCT, Heidelberg, Germany), fundus autofluorescence (FAF, Heidelberg, Germany), and full-field electroretinography (ERG), performed according to the ISCEV standard protocol.
We documented the general medical records and systemic examination results of all patients. The patients underwent laboratory investigations and ultrasonography of the kidney and liver, either in our hospital or in their local hospital. Obesity, polydactyly, reproductive abnormalities, and learning disabilities were established by physical examination or family testimony at their visit or by a telephone survey. Obesity was defined using the body mass index (BMI) according to the standards for Chinese children and adults [15, 16]. In this cohort, 14 patients were followed up.
Targeted Exon Sequencing (TES) and Whole Exome Sequencing (WES)
After informed consent was obtained, peripheral blood samples were collected from all participants for genetic analysis. Genomic DNA was extracted using a genomic DNA extraction kit (Vigorous, Beijing, China) following the manufacturer’s protocol. We performed TES analysis in all the probands, except for one proband (0191189) who underwent WES analysis, using a capture panel previously developed and evaluated by our group [17]. The capture panel included the 21 BBS genes. The Illumina library preparation and capture experiment were completed as previously described [17].
Bioinformatics analysis
Two databases, the HGMD database (http://www.hgmd.cf.ac.uk/ac/index.php) and the LOVD database (https://grenada.lumc.nl/LOVD2/eye/home.php), were used to explore any described pathogenic variants. The pathogenicity of the missense variants was predicted by in silico programs: SIFT (http://sift.jcvi.org/, in the public domain), Mutation Taster (http://www.mutationtaster.org, in the public domain), and PolyPhen2 (http://genetics.bwh.harvard.edu/pph, in the public domain). The pathogenicity of any variants involving splicing was analyzed by several programs, including Human Splice Finder (HSF, http://www.umd.be/HSF3/), NetGene2 Server (http://www.cbs.dtu.dk/services/NetGene2/), and the Berkeley Drosophila Genome Project (BDGP, http://www.fruitfly.org/seq_tools/splice.html). Co-segregation analysis was conducted when DNA of any family members was accessible.
Statistical analysis
For statistical convenience, the Snellen ratios were converted into the logarithm of the minimum angle of resolution (logMAR) values. Snellen ratios of 1.0, 0.1, counting fingers, hand movements, and light perception match logMAR values of 0, 1.0, 2.1, 2.4, and 2.7, respectively. The Kolmogorov-Smirnov test was employed to assess whether the age and BCVA of a single group conformed to a normal distribution. Data with a normal distribution were described as means, and data with non-normal distribution were described as medians.
Results
BBS-related mutant genes and variants
We identified 47 distinct diseasing-causing variants related to 10 BBS genes in the 31 probands. The 10 BBS genes were detected in the following frequencies, from high to low: BBS2 in 11 probands (35.5%), BBS4, BBS8, and BBS10 each in four probands (12.9%), BBS12 and BBS21 each in 2 probands (6.5%), and BBS1, BBS3, BBS6, and BBS9 each in 1 proband (3.2%) (Fig. 1A). Variants in BBS2 and BBS8 cause both BBS and isolated retinal dystrophy, variants in BBS3 resulted in isolated retinal dystrophy, and variants in the remaining seven genes only caused BBS (Fig. 1A, B). Among the 47 variants, 33 were first detected in the current cohort. All the novel variants were either not recorded in any public database or were present at a very low frequency (between 0.0008% and 0.04%), and all were defined as pathogenic or likely pathogenic based on the ACMG guidelines and standards (Supplementary Table 1). The type of variants and allele numbers in each gene are summarized in Table 1. Of the 47 variants, only variants c.79A > C (p.T27P) and c.534 + 1G > T in BBS2 were identified three times or more, with a gene-specific allele frequency of 27.3% (6/22) and 13.6% (3/22), respectively.
A Proportion of all patients with variants of each involved gene. The shadow areas indicate patients with isolated retinal dystrophy. B Proportion in patients with BBS. C The frequencies of each primary clinical feature in BBS observed in the patients with variants of genes encoding BBSome and patients with variants of genes encoding CP.
Clinical findings
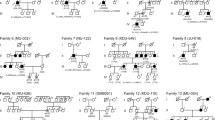
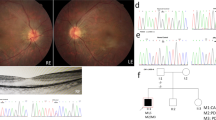
In the current cohort, 34 patients (24 males and 10 females) carried biallelic disease-causing variants in one of the BBS genes. These variants were confirmed by co-segregation analyses; the exceptions were two probands (010042 and 0191106) (Supplementary Fig. 1). Of the 34 patients, 28 from 27 unrelated families were diagnosed with BBS (Figs. 1B and 2), and the remaining six patients were diagnosed with isolated RP. The median age of the patients with BBS at last examination was 14 (range, 7–49) years and the mean age of the patients with isolated RP was 20 (range 5–43). As the patients were evaluated at an ophthalmic research institution, retinal dystrophy was observed in all the patients, followed by polydactyly in 24 patients (85.7%), obesity in 22 patients (78.6%), learning disabilities in 14 patients (50.0%), reproductive abnormalities in 11 patients (39.3%), and renal anomalies in two patients (7.1%). Only patient 019965 harboring BBS12 variants had developed all six primary components at their last examination. The remaining 27 patients with BBS included seven with five primary clinical signs, six with four components, eight presenting with three signs, and six displaying only two primary clinical features. Several secondary clinical features were noted in some patients, including brachydactyly in five patients, speech delay and dental abnormalities in four cases, ataxia in two cases, and congenital heart disease in one case. The main clinical characteristics of each patient are summarized in Supplementary Table 2. Some patients also experienced accessory spleen, fatty liver, hearing loss, hyperlipemia, hypertension, hypothyroidism, hyperuricemia, laryngeal cartilage dysplasia, midbrain dysplasia, ptosis, rachischisis, lateral ventricle cyst, and septum pellucidum cyst (Supplementary Table 2).
Ophthalmic features
All 34 patients had different extents of visual defects, and most (27 patients, 79.4%) experienced nyctalopia. In this cohort, 31 patients (91%) were diagnosed with RP, and only three patients (010042, 010179, and 0191189) were diagnosed with cone dystrophy (COD). Two patients with COD carried variants in BBS10. The median onset age for retinal dystrophy was 4 (range, 1–18) years. Among the patients suffering from severe visual impairments, more than 70% had reached the standard of low vision according to WHO criteria when they were under 20 years of age, and over 75% of the patients reached the standard of low vision or blindness when they were older than 20 years, regardless of the mutated genes (Fig. 3A, B). Nystagmus was observed in 14 patients, and five of them had isolated RP (three carrying BBS8 variants and two harboring BBS3 variants). The ERG examinations in 21 patients showed extinguished or almost extinguished recordings, except for two patients with COD, who displayed normal rod function and absent cone function.
And color fundus (CF), fundus autofluorescence (FAF), and macular optical coherence tomography (OCT) images of patients with variants in BBS genes. A The BCVA at the last visit for 34 patients, including 28 patients with BBS (25 RP presented as circles and 3 COD as squares) and 6 with isolated RP (triangles). B The stacked column chart of the number of patients with different clinical phenotypes in each visual impairment category. The first column shows the BCVA of all patients in each group, and the second and third columns show the BCVA of 12 patients at the first and last visit. VI, Visual impairment. C CF, OCT, and FAF images of the left eye of patient 0191229. During the follow-up period, CF and OCT images displayed more yellow dots in the macular area and thinning of the retina in the macular fovea. FAF image showed a hypo-AF zone in the macular region. D CF, FAF, and OCT images of patient 0191224 showed obvious tapetoretinal degeneration in the retina and macular atrophy. His fundus appearances showed macular atrophy enlargement and bone spicule during a 127month follow-up. E CF, OCT, and FAF images of the left eye of patient 019711 at age 37. FAF indicated a hyper-AF area surrounded by a mottled hypo-AF region at age 43. F CF, FAF, and OCT images of the left eye of patient 010042. At her first visit, her CF image showed small macular atrophy. Eleven years later, the CF image showed that the macular atrophy had enlarged, corresponding to the hypo-AF region in FAF. The OCT scan displayed a disruption and absence of the ellipsoid zone and RPE in the foveal region. G CF and OCT image of patient 0191189. CF image indicated diffused foveal reflection with a leopard fundus. OCT showed an almost normal macular structure.
Fundus features
Some patients with RP, aged approximately 10 years old, exhibited mild tapetoretinal degeneration (TD) in the mid-peripheral or peripheral retina and pigmentary changes in the macula (Fig. 3C). OCT scans revealed a relatively normal macular laminar architecture or mild macular atrophy. FAF showed a hypo-AF zone surrounded by a hyper-AF ring in the macular region. By contrast, some patients with disclosed obvious TD, macular dystrophy, and black pigment clumps scattered in the posterior pole and peripheral retina were under 10 years of age. Their OCT showed epiretinal membrane, macular dystrophy, and disruption and absence of the ellipsoid zone (EZ), photoreceptor inner/outer junction, and RPE in the central retina. The patients older than 20 years exhibited obvious retinal and retinal pigment epithelium (RPE) atrophy, with black bone spicule scattering in the peripheral retina (Fig. 3D) or even in the posterior pole. The exception was patient 019711, who carried biallelic missense variants in BBS2 and showed an almost normal macular structure at the age of 37 years (Fig. 3E).
Of the three patients with COD, two patients showed bull’s eye maculopathy with a smooth contour and a normal-appearing surrounding retina. FAF revealed a hypo-AF region in the macula. OCT scans displayed disruption and absence of the ellipsoid zone (EZ) and RPE in the foveal region (Fig. 3F). By contrast, patient 0191189 exhibited diffused fovea reflection and a leopard fundus. OCT showed an almost normal macular structure (Fig. 3G).
Longitudinal observation of 14 follow-up patients
We followed 14 patients for their ocular longitudinal changes, including BCVA and fundus appearance by fundus photography, OCT scanning, and FAF examination (Table 2). Two patients did not have BCVA recordings for their initial visits. The mean follow-up time was 59.07 months (range, 7–127 months). We observed that visual acuity decreased in 8 patients, remained stable in 3 patients, and was slightly improved in one patient. The three patients with stable BCVA during the follow-up either had a severe visual impairment at their first visit or had short follow-up times. During follow-up, fundus examinations in some patients revealed enlargement of the macular atrophy and expanded retinal and RPE atrophy, as well as increases in bone spicules with aging (Fig. 3).
Discussion
This study provides the first description of mutated gene profiles identified in a relatively large cohort of Chinese patients with BBS or non-syndromic BBS-related retinal dystrophy and defines the variant spectrum for each BBS gene. We also depicted the clinical features in this patient cohort and preliminarily explored the correlation between genotype and phenotype.
In the current cohort, we detected pathogenic variants in the 10 BBS genes: seven caused BBS (BBS1, 4, 6, 9,10,12, and 21), two (BBS2 and BBS8) caused both BBS and isolated RP, and one (BBS3) caused isolated RP. The two most prevalent genes for BBS were BBS2 and BBS4, which accounted for 37.0% and 14.8% of the probands with BBS, respectively. Our findings were somewhat similar to those observed in a small Chinese cohort that included 10 probands with BBS: half of the probands carried BBS2 variants [18], but they differed distinctly from the observations made in other populations [8,9,10,11,12,13]. For example, the two most common BBS genes in Spanish patients were BBS1 and BBS10, comprising 54% and 22% of the patients, respectively [12]. These two genes were also the most frequent BBS genes in patients of North America and North Europe, at 23% and 20%, respectively [9, 10, 13]. By contrast, variants of BBS1 were only detected in one patient (3.6%, 1/28) in our cohort. Of the 47 variants identified in the current study, 33 were novel variants, and 38 were detected only once, indicating greater allelic heterogeneity. The two common variants in BBS2 (p.T27P and c.534 + 1 G > T), with a gene-specific allele frequency of 27.3% and 13.6%, respectively, were first identified in Chinese patients [18]. As expected, the most common BBS1 variant, p.Met390Arg, which had an 80% gene-specific allele frequency in Spanish patients and Caucasian patients [10, 12, 13], was not identified in the current cohort. All these findings collectively suggest that the variant spectrum for each BBS gene in Chinese patients also differed from the spectra reported in other populations.
The patients in the current cohort exhibited high variability in their phenotypes, ranging from patients with all six primary clinical components to patients with only isolated RP. Using the classic diagnosis criteria [3], only 15 patients (53.5%, 15/28) could be diagnosed with BBS; therefore, genetic analysis could define patients at early stages, especially atypical patients. We confirmed the results of previous studies that the most common clinical features were retinal dystrophy, polydactyly, and obesity [7,8,9,10,11,12,13,14], but the frequencies of polydactyly and obesity differed slightly from the previous findings. The slightly higher frequency of polydactyly (86%) might be related to the high proportion of patients carrying BBS2 variants, as a meta-analysis study revealed that patients with variants in BBS2 have a higher penetrance of polydactyly [7]. The penetrance of renal anomalies in BBS is highly variable, but it is usually around 50% [1]. By contrast, only two patients (7.1%) in our cohort suffered renal defects. In a recent study, which included 12 Chinese patients with BBS, renal anomalies were observed in one of the nine patients (11.1%) [18]. In another study that included two Chinese pedigrees, four patients carrying BBS2 variants from the same family did not display renal anomalies [19]. Currently, all published studies involving Chinese patients with BBS are either case reports or small cohort studies (fewer than 15 patients) [18,19,20,21,22]; therefore, we could not justify a conclusion that the incidence of renal anomalies was lower in Chinese patients with BBS. The pathophysiology of the renal defects caused by BBS protein deficiency remains unresolved. One previous study proposed that BBSome deficiency resulted in mislocalization and dysfunction of polycystin‐1 and polycystin‐2 [23]. Another study on animal models indicated that calorie restriction reversed the morphological and molecular changes in the kidney that occur in Bbs2−/− and Bbs4−/− mice and prevented renal defects [23]. Therefore, the low frequency of renal defects in Chinese patients might be related to dietary habits or other environmental and epigenetic factors.
By presenting a relatively large cohort of patients whose genotypes were confirmed, our study provided an opportunity to explore genotype and phenotype correlations. Consistent with a previous study [7], we did not detect any difference in the frequencies of any primary clinical features between the patients (n = 19) with variants in genes encoding BBSome and patients (n = 7) with variants of genes encoding chaperonin‐like proteins (Fig. 1C). In the largest genotype group (BBS2), the patients with truncating variants presented more clinical features of BBS, while patients with biallelic missense variants displayed fewer clinical features or isolated RP. All variants of BBS2 detected in isolated RP were missense variants, further suggesting that missense variants of BBS2 cause a mild phenotype and lower penetrance of the primary clinical features [6, 7]. The most common missense variant, p.T27P in BBS2, was detected in five pedigrees (six patients). The patients with variant p.T27P, one in the homozygous and the remaining five in the heterozygous state, tended to exhibit fewer primary components, as three patients suffered only RP and polydactyly. We speculated that variant p.T27P might be a mild or hypomorphic variant that causes only a partial loss of function of BBS2.
Apart from the patients with BBS2 variants, we also observed five other patients from three unrelated families who carried variants of BBS3 and BBS8, only suffered from isolated RP. Both BBS3 and BBS8 have retina-specific isoforms BBS3L and BBS8L [24, 25]. RNA rescue experiments revealed that variant p.A89V in BBS3, detected in patients with isolated RP, could rescue the transport delays induced by the loss of bbs3, but were unable to rescue vision impairment, indicating that this variant was critical and specific for the vision defect. One reported variant, c.115-2 A > G in BBS8, detected in four Pakistani patients with isolated RP, caused skipping of exon 2a of BBS8L, confirming a role of this isoform in retinal pathology [24]. The variants of BBS3 and BBS8 identified in the current study included missense, splicing, frameshift, and large deletion variants, but their mechanisms for causing isolated RP need comprehensive exploration in future studies.
Retinal dystrophy, the most highly penetrant feature in BBS, usually occurs early and is severe[7, 26, 27]. Most of the patients in our cohort exhibited more severe visual impairments than were reported in a cohort that included 67 patients carrying variants in BBS1 and BBS10 [28]. One previous study noted that VA was significantly worse in patients with BBS2 variants than in patients with BBS1 variants [26]. The severity of visual defects was related to early retinal degeneration involving the macular region. Similar to previous descriptions [18, 26], advanced retinal degeneration with profound macular atrophy was observed in all patients aged over 20 years, except for two patients, 010042 and 019711, who both had biallelic missense variants. Patient 010042 with COD with BBS10 variants had only small bull’s eye maculopathy and nearly normal BCVA (logMAR 0.1) at age 39. After over 10 years of follow-up, her maculopathy enlarged and her BCVA decreased to logMAR 0.6. Patient 019711 with BBS2 variants displayed relatively normal macular structures and mild visual defects (logMAR 0.2 OD and 0.05 OS) at age 37. After 6 years of follow-up, he had only a mild visual decrease in his left eye, from 0.05 to 0.2. Two of the four patients with BBS10 variants were diagnosed with COD, in agreement with the finding reported in a cohort that included 67 patients with variants in BBS1 and BBS10 [28].
The current study has some limitations. One is its retrospective design. Another is the small number of patients with variants in BBS genes other than BBS2; therefore, establishing a genotype-phenotype correlation is rather difficult. In addition, the frequency of renal anomalies and cognitive impairment might be underestimated as they were not properly assessed or were not displayed at the time of clinical evaluation.
In conclusion, our study defined the mutation profiles of BBS genes and established the configuration of the variation frequencies for each BBS gene in a Chinese patient cohort. Our results revealed that Chinese patients showed early and severe visual defects and retinal degeneration. Our findings provide essential information for future genetic counseling and gene therapy in these patients.
Summary table
What was known before
-
Biallelic BBS gene variants cause Bardet Biedl syndrome. The two most common BBS genes in North America and North Europe patients were BBS1 and BBS10.
What this study adds
-
Genotype and phenotype of Chinese patients with biallelic BBS gene variants. Our findings were differed distinctly from the observations made in other populations.
Data availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Chandra B, Tung ML, Hsu Y, Scheetz T, Sheffield VC. Retinal ciliopathies through the lens of Bardet-Biedl Syndrome: Past, present and future. Prog Retin Eye Res. 2022;89:101035.
Florea L, Caba L, Gorduza EV. Bardet-Biedl Syndrome-Multiple Kaleidoscope Images: Insight into mechanisms of genotype-phenotype correlations. Genes. 2021;12:1–13.
Beales PL, Elcioglu N, Woolf AS, Parker D, Flinter FA. New criteria for improved diagnosis of Bardet-Biedl syndrome: results of a population survey. J Med Genet. 1999;36:437–46.
Riazuddin SA, Iqbal M, Wang Y, Masuda T, Chen Y, Bowne S, et al. A splice-site mutation in a retina-specific exon of BBS8 causes nonsyndromic retinitis pigmentosa. Am J Hum Genet. 2010;86:805–12.
Pretorius PR, Aldahmesh MA, Alkuraya FS, Sheffield VC, Slusarski DC. Functional analysis of BBS3 A89V that results in non-syndromic retinal degeneration. Hum Mol Genet. 2011;20:1625–32.
Shevach E, Ali M, Mizrahi-Meissonnier L, McKibbin M, El-Asrag M, Watson CM, et al. Association between missense mutations in the BBS2 gene and nonsyndromic retinitis pigmentosa. JAMA Ophthalmol. 2015;133:312–8.
Niederlova V, Modrak M, Tsyklauri O, Huranova M, Stepanek O. Meta-analysis of genotype-phenotype associations in Bardet-Biedl syndrome uncovers differences among causative genes. Hum Mutat. 2019;40:2068–87.
Guardiola GA, Ramos F, Izquierdo NJ, Oliver AL. A genotype-phenotype analysis of the Bardet-Biedl Syndrome in Puerto Rico. Clin Ophthalmol. 2021;15:3757–64.
Manara E, Paolacci S, D’Esposito F, Abeshi A, Ziccardi L, Falsini B, et al. Mutation profile of BBS genes in patients with Bardet-Biedl syndrome: an Italian study. Ital J Pediatr. 2019;45:72.
Esposito G, Testa F, Zacchia M, Crispo AA, Di Iorio V, Capolongo G, et al. Genetic characterization of Italian patients with Bardet-Biedl syndrome and correlation to ocular, renal and audio-vestibular phenotype: identification of eleven novel pathogenic sequence variants. BMC Med Genet. 2017;18:10.
Maria M, Lamers IJ, Schmidts M, Ajmal M, Jaffar S, Ullah E, et al. Genetic and clinical characterization of Pakistani families with Bardet-Biedl syndrome extends the genetic and phenotypic spectrum. Sci Rep. 2016;6:34764.
Castro-Sanchez S, Alvarez-Satta M, Corton M, Guillen E, Ayuso C, Valverde D. Exploring genotype-phenotype relationships in Bardet-Biedl syndrome families. J Med Genet. 2015;52:503–13.
Daniels AB, Sandberg MA, Chen J, Weigel-DiFranco C, Fielding Hejtmancic J, Berson EL. Genotype-phenotype correlations in Bardet-Biedl syndrome. Arch Ophthalmol. 2012;130:901–7.
Imhoff O, Marion V, Stoetzel C, Durand M, Holder M, Sigaudy S, et al. Bardet-Biedl syndrome: a study of the renal and cardiovascular phenotypes in a French cohort. Clin J Am Soc Nephrol. 2011;6:22–9.
Li H, Ji CY, Zong XN, Zhang YQ. Body mass index growth curves for Chinese children and adolescents aged 0 to 18 years. Chin J Pediatr. 2009;47:493–8.
Zhou B. Predictive values of body mass index and waist circumference to risk factors of related diseases in Chinese adult population. Zhonghua Liuxingbingxue Zazhi 2002;23:5–10.
Sun T, Xu K, Ren Y, Xie Y, Zhang X, Tian L, et al. Comprehensive molecular screening in Chinese usher syndrome patients. Invest Ophthalmol Vis Sci. 2018;59:1229–37.
Meng X, Long Y, Ren J, Wang G, Yin X, Li S. Ocular characteristics of patients with bardet-biedl syndrome caused by pathogenic BBS gene variation in a Chinese cohort. Front Cell Dev Biol. 2021;9:635216.
Huang L, Sun L, Wang Z, Li S, Chen C, Luo X, et al. Novel compound heterozygous BBS2 and homozygous MKKS variants detected in Chinese families with Bardet-Biedl Syndrome. J Ophthalmol. 2021;2021:6751857.
Tang HY, Xie F, Dai RC, Shi XL. Novel homozygous protein-truncating mutation of BBS9 identified in a Chinese consanguineous family with Bardet-Biedl syndrome. Mol Genet Genom Med. 2021;9:e1731.
Tao T, Wang L, Chong W, Yang L, Li G. Characteristics of genotype and phenotype in Chinese patients with Bardet-Biedl syndrome. Int Ophthalmol. 2020;40:2325–43.
Xing DJ, Zhang HX, Huang N, Wu KC, Huang XF, Huang F, et al. Comprehensive molecular diagnosis of Bardet-Biedl syndrome by high-throughput targeted exome sequencing. PLoS One. 2014;9:e90599.
Guo DF, Beyer AM, Yang B, Nishimura DY, Sheffield VC, Rahmouni K. Inactivation of Bardet-Biedl syndrome genes causes kidney defects. Am J Physiol Ren Physiol. 2011;300:F574–80.
Murphy D, Singh R, Kolandaivelu S, Ramamurthy V, Stoilov P. Alternative splicing shapes the phenotype of a mutation in BBS8 to cause nonsyndromic Retinitis Pigmentosa. Mol Cell Biol. 2015;35:1860–70.
Pretorius PR, Baye LM, Nishimura DY, Searby CC, Bugge K, Yang B, et al. Identification and functional analysis of the vision-specific BBS3 (ARL6) long isoform. PLoS Genet. 2010;6:e1000884.
Denniston AK, Beales PL, Tomlins PJ, Good P, Langford M, Foggensteiner L, et al. Evaluation of visual function and needs in adult patients with bardet-biedl syndrome. Retina 2014;34:2282–9.
Berezovsky A, Rocha DM, Sacai PY, Watanabe SS, Cavascan NN, Salomão SR. Visual acuity and retinal function in patients with Bardet-Biedl syndrome. Clinics 2012;67:145–9.
Grudzinska Pechhacker MK, Jacobson SG, Drack AV, Scipio MD, Strubbe I, Pfeifer W, et al. Comparative natural history of visual function from patients with Biallelic variants in BBS1 and BBS10. Invest Ophthalmol Vis Sci. 2021;62:26.
Acknowledgements
This study was supported by the National Key R&D Program of China (2016YFC0905200). The funding organizations had no role in designing or conducting this research.
Funding
This work was supported by the National Key R&D Program of China, 2016YFC0905200. The funding organization had no role in the design or conduct of this research.
Author information
Authors and Affiliations
Contributions
JZ and YX participated in the study design, acquired the data, and drafted the manuscript. HY, CC, TS, KX, and XZ analyzed the data and aided in interpreting the results. YL designed the current study and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhong, J., Xie, Y., Ye, H. et al. Phenotypic diversity observed in a Chinese patient cohort with biallelic variants in Bardet-Biedl syndrome genes. Eye 37, 3398–3405 (2023). https://doi.org/10.1038/s41433-023-02516-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41433-023-02516-w
- Springer Nature Limited