Abstract
Three new 22-membered polyol macrolides, dactylides A−C (1−3), were isolated from Dactylosporangium aurantiacum ATCC 23491 employing repeated chromatographic separations, and their structures were established based on detailed analysis of NMR and MS data. The relative configurations at the stereocenters were established via vicinal 1H–1H coupling constants, NOE correlations, and by application of Kishi’s universal NMR database. In order to get insights into the biosynthetic pathway of 1−3, the genome sequence of the producer strain D. aurantiacum was obtained and the putative biosynthetic gene cluster encoding their biosynthesis was identified through bioinformatic analysis using antiSMASH. Compounds 1−3 showed significant in-vitro antimycobacterial and cytotoxic activity.

Similar content being viewed by others
Introduction
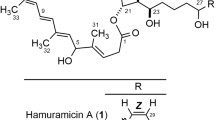
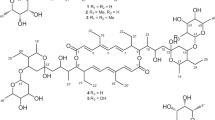
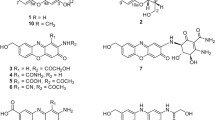
Naturally produced macrolides from actinomycetes are the paramount source of bioactive metabolites with profound structural diversities [1]. The structural diversity of macrolides ranges from 12- to 66-membered lactone rings, with unending scope for discovering new structural probabilities [2, 3]. Dactylosporangium aurantiacum, Gram-positive soil-based actinobacteria, is an exclusive source of lipiarmycins which are reported to exhibit potent antimicrobial properties against Gram-positive bacteria [4] and mycobacteria [5]. Lipiarmycin also known as fidaxomicin has been approved by the FDA to treat Clostridium difficile-associated diarrhea [6]. The pharmaceutical importance of lipiarmycins inspired us to further explore the D. aurantiacum for other bioactive metabolites. The actinomycete D. aurantiacum ATCC 23491 [7] remains unexplored in terms of secondary metabolites. With an aim for the isolation of interesting metabolites from D. aurantiacum, the strain was procured from ATCC and confirmed by 16 s rRNA sequencing (GenBank accession number OQ352622). Chemical investigation of ethyl acetate extract from the fermented broth using repeated chromatographic techniques led to the isolation of new 22-membered polyol macrolides named dactylides A−C (1−3) (Fig. 1). Employing various 1D- and 2D-NMR spectral analyses along with HR-ESI-MS data, structures of 1−3 were elucidated and their relative stereochemistry was determined using NOE correlations and Kishi’s universal NMR database [8]. Furthermore, the putative biosynthetic pathway for 1−3 was proposed on the basis of a genome sequence study using antiSMASH [9]. Further, 1−3 were evaluated for in vitro antimycobacterial and cytotoxic activities. Using the molecular docking, binding affinities of 1−3 were evaluated for pfDHFR-TS and pfLDH enzymes (See supporting information). Herein, the isolation, structural elucidation with stereochemical determination, biosynthetic pathway, and bioactivities of 1−3 are reported.
Material and methods
General experimental procedures
The NMR spectra were recorded on JEOL JNM-ECZR 600 MHz and Bruker Avance II 500 MHz spectrometers in DMSO-d6 and CD3OD. High-resolution electrospray ionization-mass spectrometry (HR-ESI-MS) data were acquired using Agilent 6545 LC/Q-TOF mass spectrometer. The FT-IR spectra were recorded on INTERSPEC 200-X FTIR spectrometer, CD spectra were recorded using JASCO J-815 CD spectrometer, and UV spectra were recorded on IGENE LABSERVE IG-UV302S (Microprocessor double beam xenon flash lamp spectrophotometer). Separation and purification of compounds was done using HPLC (Dionex Ultimate 3000, Thermo Scientific) coupled with a diode array detector using a semi-preparative column (YMC-Triart C18, 5 μm, 250 × 10 mm ID). HPLC grade solvents from Qualigens (Thermo Fisher Scientific) were used.
Fermentation, extraction, and isolation
A seed broth was prepared by inoculating spores of the strain into ATCC medium 172 consisting of glucose 1.0%, starch 2.0%, yeast extract 0.5%, casein 0.5%, CaCO3 0.1% (pH 7.2), and the same medium was also used for production culture. After fermentation, the culture broth was centrifuged and the supernatant layer was extracted three times with equal volumes of ethyl acetate. These ethyl acetate layers were combined, dried over sodium sulfate, and concentrated in vacuo to afford 0.51 g of brown residue. The obtained crude extract was fractionated by flash chromatography on a C18 column with H2O – MeOH gradient elution and collected 32 fractions. All the factions were concentrated in vacuo and analyzed by HPLC. 1 mg ml−1 MeOH solution of the residue was subjected to HPLC-DAD analysis on analytical column (Thermo Fisher C18, 5 µm, 250 × 4.6 mm). Elution was performed at the flow rate of 1 ml min−1 with water (A) and acetonitrile (B) as the mobile phase. Fraction 27 was further purified by semi-preparative column (YMC-Triart C18, 5 µm, 250 × 10 mm) with an isocratic elution using acetonitrile/water solution (3:7) (6 ml min−1) to afford the major compound 1 (7.41 mg) along with two minor compounds 2 (2.2 mg) and 3 (2.9 mg).
Dactylide A (1): Amorphous off-white powder; UV (MeOH) λmax (log ε) 229 (3.80) nm; CD (c 7.6 × 10−4 M, MeOH) λmax (∆ε) 233 (−3.58), 226.6 (4.42), 222.6 (−3.32), 219.8 (3.63), 216 (3.89), 211 (4.04); IR vmax 3412, 2957, 2936, 1713, 1550, 1461, 1377, 1306, 1132, 1070, 994, 905, 826, 654, 590, 533 cm-1; 1H and 13C NMR data (see Table 1); (+)-HR-ESI-MS m/z 825.4994 [M + Na]+ (calcd for C42H74NaO14, 825.4970).
Dactylide B (2): Amorphous off-white powder; UV (MeOH) λmax (log ε) 229 (3.80) nm; CD (c 2.4 ×10−4 M, MeOH) λmax (∆ε) 247.6 (4.09), 237.4 (3.98), 232 (−4.32), 224.8 (4.69), 214.4 (4.25); IR vmax 3407, 2958, 1709, 1561, 1463, 1384, 1253, 1186, 1126, 1080, 995, 853, 655 cm−1; 1H and 13C NMR data (see Table 1); (+)-HR-ESI-MS m/z 681.4192 [M + Na]+ (calcd for C35H62NaO11, 681.4184).
Dactylide C (3): Amorphous off-white powder; UV (MeOH) λmax (log ε) 228 (3.83) nm; CD (c 5.7 ×10−4 M, MeOH) λmax (∆ε) 239 (4.15), −223 (4.57); IR vmax 3404, 2954, 2935, 1710, 1595, 1465, 1368, 1165, 1127, 1067, 996, 847, 668 cm−1; 1H and 13C NMR data (see Table 1); (+)-HR-ESI-MS m/z 811.4813 [M + Na]+ (calcd for C41H72NaO14, 811.4814).
Genome sequencing and bioinformatics analysis
Whole genome sequencing was done by Eurofins genomics India Pvt. Ltd. using an Illumina sequencer. Secondary metabolic genes were identified using the antiSMASH (antibiotics & Secondary Metabolite Analysis Shell) 6.0 program. Sequences of ketoreductase domains were used to validate the stereochemistry and structure of the compounds [10]. This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession JAQQGQ000000000.
Antimycobacterial assay
The MIC of the compounds was determined using the broth microdilution method. In brief, the concentrations 50 µg ml−1, 25 µg ml−1, 12.5 µg ml−1, 6.25 µg ml−1, 3.125 µg ml−1, and 1.56 µg ml−1 of the compounds were tested against Mycobacterium tuberculosis H37Rv (Mtb) in a 96 well plate. Middlebrook 7H9 media with growth supplement was used as growth media and for dilution of the compounds. Necessary controls were also included in the assay such as culture control (107 and 105 cells ml−1) and solvent control (DMSO). Plates were incubated at 37 °C for 5 days, and the growth of the Mtb was observed through an inverted microscope in the form of serpentine cords. All the tests were performed in triplicates and the minimum concentration required to completely inhibit the Mtb growth was considered as MIC [11].
Cytotoxicity assay
The cytotoxic activity of the compounds was tested using MTT cell proliferation assay. In brief, HCT116 cells were seeded in a 96-well plate with 5 × 103 cells per well and allowed to grow overnight. Cells were treated with different concentrations 10 µg ml−1, 7 µg ml−1, 5 µg ml−1, and 2 µg ml−1 of compounds 1 and 2. Cells were incubated with the compounds for 48 h and incubated with MTT for 4 h at 37 °C and 5% CO2. The produced formazan crystals were solubilized using the addition of DMSO and incubated in dark for 30 min. The absorbance was recorded at 570 nm using a microplate reader [12].
HCT116 colon cancer cell line was cultured in filter-sterilized Dulbecco’s Modified Eagle Medium (DMEM) (Sigma-Aldrich) supplemented with 1% penicillin, streptomycin, and amphotericin B (Sigma-Aldrich), and 10% Fetal Bovine Serum (FBS) (Gibco). 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide and DMSO were obtained from Sigma Aldrich for the MTT assay. The cells were maintained at 37 °C in a humidified atmosphere containing 5% CO2.
Results and discussion
Dactylide A (1) was obtained as an amorphous off-white powder. The molecular formula of 1 was determined to be C42H74O14 based on HR-ESI-MS analysis which showed m/z 825.4994 [M + Na]+ (calculated for C42H74NaO14+, 825.4970) indicating six degrees of unsaturation (Supplementary Fig. S1). The 13C NMR, DEPT, and HSQC spectral data (Table 1 and Supplementary Figs. S3, S4 and S6) of 1 revealed the presence of 42 carbon signals, including an ester carbonyl carbon (δC 175.3), four olefinic carbons (δC 135.3, 135.1, 132.5, and 132.1), a hemiketal carbon (δC 101.1), eighteen methines (δC 102.0, 81.9, 79.0, 77.6, 76.9, 75.6, 73.7, 73.3, 72.8, 71.3, 70.0, 69.6, 65.3, 48.2, 43.4, 42.5, 40.7, 25.4), ten methylenes (δC 44.4, 43.4, 43.0, 43.0, 42.4, 39.7, 39.1, 37.5, 36.2, 23.6), an oxymethyl (δC 57.4), and seven methyls (δC 24.6, 21.8, 18.4, 17.5, 12.2, 11.9, 6.6). In the 1H NMR spectrum of 1 (Supplementary Fig. S2), the presence of seven doublet signals at δH 1.25 (H3-6′), 1.10 (H3-32), 1.05 (H3-31), 0.94 (H3-30), 0.91 (H3-33), 0.91 (H3-35), and 0.76 (H3-34) indicated the presence of seven methyl groups.
COSY spectrum (Supplementary Fig. S5) data revealed the presence of three spin systems (H2-2 to H2-6, H-8 to H3-30, and H-1′ to H3-6′) (Fig. 2a). The spin system H-1′ to H3-6′ indicated a presence of 2,6-dideoxyhexosyl moiety on the basis of coupling from an anomeric proton δH 4.55 (H-1′) with methylene protons δH 2.32 (H-2′) and δH 1.34 (H-2′). These methylene protons δH 2.32 (H-2′) and δH 1.34 (H-2′) showed coupling with oxymethine proton δH 3.18 (H-3′) which was found coupling with another oxymethine proton δH 2.96 (H-4′) which showed coupling with oxymethine proton δH 3.21 (H-5′) which was further coupled with the methyl protons δH 1.25 (H3-6′) (Fig. 2a and Supplementary Fig. S5).
The connection between C-6 (δC 45.1) and C-8 (δC 48.4) via an oxygenated quaternary sp3 carbon C-7 (δC 101.1) was deduced on the basis of HMBC correlations (Fig. 2a and Supplementary Fig. S7) from protons H-8 (δH 1.25), H2-6 (δH 1.81, 1.52), H-5 (δH 4.06), and H3-31 (δH 1.05) to hemiketal carbon C-7. Moreover, an HMBC correlation between H-11 (δH 4.07) and C-7 confirmed the tetrahydropyran ring with an ether linkage between C-11 (δC 65.3) and C-7 (Fig. 2a and Supplementary Fig. S7). The ester linkage between C-1 (δC 175.3) and C-21 (δC 75.6) was established by the relatively downfield chemical shift for H-21 (δH 5.12) [13] and the HMBC correlations from H-21 (δH 5.12) to C-1 (Fig. 2a and Supplementary Fig. S7). The attachment of 2,6-dideoxyhexose sugar at C-13 was confirmed from the HMBC correlations from H-13 (δH 3.89) to C-1’ (δC 102.0) and from H-1’ (δH 4.55) to C-13 (δC 77.6) (Fig. 2a and Supplementary Fig. S7). The HMBC correlation from OCH3 (δH 3.44) to C-3’ (δC 81.9) confirmed the attachment of OCH3 group to C-3’ (Fig. 2a and Supplementary Fig. S7).
The trans geometry of double bonds was assigned from the large coupling constants 3JH-16/H-17 = 15.00 Hz and 3JH-18/H-19 = 15.00 Hz and COSY couplings from H-16 to H-17 to H-18 to H-19 established the presence of a conjugated diene group [14]. The putative glycosyltransferase enzymes in the present study belong to the GT1 family and thus invert the stereochemistry at the anomeric carbon of the activated sugar during its transfer to the aglycone which is clarified vide infra [15, 16]. The large coupling constants 3JH-1’/H-2’ = 9.6 Hz, 3JH-2’/H-3’ = 12.0 Hz, 3JH-3’/H-4’ = 9.0 Hz, and 3JH-4’/H-5’ = 9.0 Hz indicated that the H-1’, H-3’, H-4’, and H-5’ protons occupied axial orientations, suggesting the sugar to be β-glycosidically linked 3’-O-methyl-olivose (oleandrose). This was further confirmed through the NOE cross-peaks between H-1’ (δH 4.55) and H-5’ (δH 3.21), and H-1’ (δH 4.55) and H-3’ (δH 3.18) [15, 16] (Fig. 2b). Thus, the planar structure of 1 was elucidated to be a 22-membered macrolide glycoside possessing a conjugated diene, a tetrahydropyran ring, and an oleandrose sugar.
Following the elucidation of the planar structure, the relative stereochemistry of the aglycone part of 1 was determined on the basis of the 1H − 1H coupling constant values, NOE correlations, and by application of Kishi’s universal NMR database [17]. The large coupling constant 3JH-8/H-9 = 10.8 Hz, together with the key NOE correlations between H-11 (δH 4.07) and H-9 (δH 3.59), and H-9 (δH 3.59) and H-8 (δH 1.25) indicated the relative stereochemistry of the tetrahydropyran ring in 1 (Fig. 2b and Supplementary Fig. S8) [18, 19]. Finally, the stereochemistry of a contiguous dipropionate unit (C-23 to C-27) of 1 was predicted by the application of Kishi’s propionate universal NMR database [20]. By following the same procedure described previously, the 13C NMR characteristics of the C-23 to C-26 (first mode of correlation) and C-24 to C-27 (second mode of correlation) (Fig. 3, Supplementary Figs. S9, and S10) portions of the natural products were compared with those of each of the eight diastereomers. As seen from the structures (Fig. 3), peripheral carbons in standard molecules cannot be considered for similarity match due to the effect of functional groups at respective δC in 1 [21]. Hence, we chose to focus only on the 13C NMR characteristics of the C-24 and C-33 and of the C-26 and C-34 (Fig. 3, Supplementary Figs. S9 and S10) portions of the natural products. From these comparisons, 1d (βββα) and 1e (ββαβ) emerge from the first and second modes of correlation, respectively (The symbols, α and β, represent the direction of the bonds, outside the plane and inside the plane, respectively). Hence, the relative stereochemistry at C-23, C-24, C-25, C-26, and C-27 was predicted to be β, β, β, α, β, respectively. In addition, a comparison of the chemical shifts in the NMR database (Supplementary Fig. S11) for the dotted carbon atom in 1,3-diols with chemical shifts of the relevant portion of 1 immediately predicted the relative configuration of C-13 to C-15 as to be syn [22, 23]. Thus, the relative stereochemistry of compound 1 was proposed to be 5S*, 7S*, 8R*, 9S*, 11R*, 13R*, 15R*, 23S*, 24R*, 25S*, 26S*, 27S* based on coupling constant values, NOE correlations, Kishi’s universal NMR database, and reported data of similar compounds [18, 19]. During the revision stage of this manuscript, a recently published article disclosed dactylosporolide A, isolated from genetically engineered D. fulvum, having the same planar structure as of dactylide A (1) and therefore, the stereochemistry and configuration of the aglycone and sugar moieties was assumed to be the same as that of dactylosporolide A [18].
Compound 2 was isolated as an amorphous off-white powder and its molecular formula was determined to be C35H62O11 from the HR-ESI-MS peak at m/z 681.4192 ([M + Na]+, calculated for C35H62NaO11+, 681.4184) indicating five degrees of unsaturation (Supplementary Fig. S12). Comparison of 1H and 13C NMR data (Table 1 and Supplementary Figs. S13–S15) of 2 with that of 1 indicated that 2 lacks hexose sugar on the basis of the absence of signals which further aligned with the reduced number of degrees of saturation as suggested by HR-ESI-MS data. On the basis of COSY (Supplementary Fig. S16), HSQC (Supplementary Fig. S17), and HMBC (Supplementary Fig. S18) analyses, all 1H and 13C NMR signals of 2 were assigned. Thus, the structure of 2 was determined as an aglycone of 1. The absolute stereochemistry was assumed to be similar to that of 1 based on coupling constant values and NOE correlations (Supplementary Fig. S19).
The molecular formula of compound 3 was deduced as C41H72O14 based on a sodium adduct ion at m/z 811.4813 [M + Na]+ (calcd for C41H72NaO14+, 811.4814) in HR-ESI-MS spectrum (Supplementary Fig. S20), that differing by 14 Da from 1. The 1H- and 13C NMR spectra (Supplementary Figs. S21 and S22) of 3 showed considerable similarities to those of 1, except that the absence of a signal for methoxy group [as observed in 1 at δH 3.44 (H-5′) and δC 57.4 (H-5′)] which was replaced by a hydroxy group. On the basis of COSY (Supplementary Fig. S23), HSQC (Supplementary Fig. S24), and HMBC (Supplementary Fig. S25) analyses, all 1H and 13C NMR signals of 3 were assigned. NMR data of 3 in DMSO-d6 (Supplementary Table S1, Supplementary Figs. S27−S31), helped in further confirming the relative stereochemistry of the tetrahydropyran ring. The large coupling constant 3JH-8/H-9 = 10.3 Hz, together with the key NOE correlations between H-11 (δH 3.84) and 7-OH (δH 5.89), and H-9 (δH 3.87) and 7-OH (δH 5.89) indicated the relative stereochemistry of the tetrahydropyran ring (Fig. 2b and Supplementary Fig. S31) [18, 19]. Based on coupling constant values and NOE data (Supplementary Figs. S26 and S31), 3 was assumed to carry the same absolute configurations as that of 1.
The finding of new dactylide class metabolites and their fascinating structures inspired us to investigate their biosynthesis. Structural similarity with macrolides suggested that a modular polyketide synthase (Type I PKS) and reverse synthesis prediction revealed that 13 modules are involved in the biosynthesis of dactylides core structure. The analysis of the draft genome sequence of D. aurantiacum ATCC 23491 using antiSMASH revealed the presence of 3 Type I PKS gene clusters and only 1 of them (designated as dac gene cluster) has all the necessary sequences required for the biosynthesis of dactylides. dac consists of 178.3 kb sequence having 60% similarity to catenulisporolides biosynthetic gene cluster from Catenulispora sp [19]. 74 open reading frames (ORFs) in dac putatively code for 5 genes of Type I PKS, 4 glycosyl transferases, 6 genes for biosynthesis of olivose and oleandrose sugar moieties, 5 genes for pathway regulation, 10 ORFs of non-ribosomal peptides, and remaining genes are either for hypothetical proteins or for transmembrane transportation. Out of 5 Type I PKS genes present in dac, 4 genes dacA1−dacA4 comprising 13 modules are proposed to synthesize the core structure of the dactylides. dacA2−dacA4 genes are present in continuation but dacA1 is situated 30 kb upstream of dacA2. antiSMASH analysis of the whole genome correlated with the structures as modules no. 1, 2, 4, and 10 are responsible for the addition of methylmalonyl CoA unit to the core structure which is equivalent to C-26, C-24, C-20, and C-8 of the dactylides (Fig. 4).
Proposed biosynthetic pathway of 1−3. 4 genes (dacA1-dacA4) involved in the synthesis of the core molecule with details of enzymatic modules and domains along with loading modules shown above. Stereochemistry prediction and types of ketoreductase domain identification based on amino acid sequences. Abbreviations: ACP Acyl Carrier Protein, AT Acyl transferase, KS Ketosynthase, KR Ketoreductase, DH Dehydratase, ER Enolreductase
The sequence of ketoreductase domains is utilized to predict the absolute stereochemistry of molecules. A comparison of the sequence with the literature resulted in the identification of types of conserved motifs. Therefore, the probable absolute stereochemistry of the 1−3 was proposed as shown in Fig. 4 based on the genomic analysis [24, 25]. The absolute stereochemistry of these compounds needs to be further confirmed via chemical experiments. The four putative glycosyltransferase genes predicted by antiSMASH were searched in BLAST and the best PDB (Protein database) match was found to be part of the GT1 family of glycosyltransferase listed in the CAZy database (Supplementary Table S2) [26].
Compound 2 was also found to possess antimycobacterial activity against the M. tuberculosis H37Rv strain. MIC of 2 was determined to be 25 µg ml−1 while 1 and 3 showed no inhibition up to 50 µg ml−1 against M. tuberculosis H37Rv. Cytotoxic activity of compounds 1 and 2 was assessed in vitro using an MTT cell proliferation assay on HCT116 colon cancer cells. At the highest concentrations of 10 µg ml−1, 1 and 2 exhibited a 34.60% and 23.95% reduction in cell proliferation, respectively when compared with the negative control.
Conclusion
In conclusion, Dactylosporangium aurantiacum ATCC 23491 was observed to produce three new 22-membered polyol macrolides (1–3) which were isolated and their structures were elucidated based on detailed analyses of NMR and MS data. The relative configurations at the stereocenters were established via vicinal 1H–1H coupling constants, NOE correlations, and by application of Kishi’s universal NMR database. Furthermore, the genome sequence of the producer strain D. aurantiacum was obtained and the putative biosynthetic gene cluster encoding their biosynthesis was identified through bioinformatic analysis using antiSMASH. Compounds 1–3 displayed significant in-vitro cytotoxicity and antimycobacterial activity, and found to show significant binding affinity for pfDHFR-TS and pfLDH enzymes in the in-silico studies (Supplementary Tables S4 and S5). In this way, we report an investigation of D. aurantiacum that led to the successful isolation and identification of three new compounds.
Data availability
Data will be made available on request.
References
Al-Fadhli AA, et al. Macrolides from rare actinomycetes: Structures and bioactivities. Int J Antimicrob Agents. 2022;59:106523. https://doi.org/10.1016/j.ijantimicag.2022.106523.
Elshahawi SI, Shaaban KA, Kharel MK, Thorson JS. A comprehensive review of glycosylated bacterial natural products. Chem Soc Rev. 2015;44:7591–697. https://doi.org/10.1039/C4CS00426D.
Onodera KI, Nakamura H, Oba Y, Ohizumi Y, Ojika M. Zooxanthellamide Cs: vasoconstrictive polyhydroxylated macrolides with the largest lactone ring size from a marine dinoflagellate of Symbiodinium sp. J Am Chem Soc. 2005;127:10406–11. https://doi.org/10.1021/ja050810g.
Cavalleri B, Arnone A, Di Modugno E, Nasini G, Goldstein BP. Structure and biological activity of lipiarmycin B. J Antibiot. 1988;41:308–15. https://doi.org/10.7164/antibiotics.41.308.
Kurabachew M, et al. Lipiarmycin targets RNA polymerase and has good activity against multidrug-resistant strains of Mycobacterium tuberculosis. J Antimicrob Chemother. 2008;62:713–9. https://doi.org/10.1093/jac/dkn269.
Zhanel GG, Walkty AJ, Karlowsky JA. Fidaxomicin: A novel agent for the treatment of Clostridium difficile infection. Can J Infect Dis Med Microbiol. 2015;26:305–12. https://doi.org/10.1155/2015/934594.
Thiemann JE. A new species of the genus Amorphosporangium isolated from Italian soil. Mycopathol Mycol Appl. 1967;33:233–40. https://doi.org/10.1007/BF02088915.
Kobayashi Y, Tan CH, Kishi Y. Toward creation of a universal NMR database for stereochemical assignment: Complete structure of the desertomycin/oasomycin class of natural products. J Am Chem Soc. 2001;123:2076–8. https://doi.org/10.1021/ja004154q.
Blin K, et al. AntiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021;49:W29–W35. https://doi.org/10.1093/nar/gkab335.
Keatinge-Clay AT. A tylosin ketoreductase reveals how chirality is determined in polyketides. Chem Biol. 2007;14:898–908. https://doi.org/10.1016/j.chembiol.2007.07.009c.
Heinrichs MT, et al. Mycobacterium tuberculosis Strains H37Ra and H37Rv have equivalent minimum inhibitory concentrations to most antituberculosis drugs. Int J Mycobacteriol. 2018;7:156–61. https://doi.org/10.4103/ijmy.ijmy_33_18.
Paramita P, et al. Evaluation of potential anti‐cancer activity of cationic liposomal nanoformulated Lycopodium clavatum in colon cancer cells. IET Nanobiotech. 2018;12:727–32. https://doi.org/10.1049/iet-nbt.2017.0106.
Shinde PB, et al. A non-immunosuppressive FK506 analogue with neuroregenerative activity produced from a genetically engineered Streptomyces strain. RSC Adv. 2015;5:6823–8. https://doi.org/10.1039/C4RA11907J.
Helaly SE, et al. Langkolide, a 32-membered macrolactone antibiotic produced by Streptomyces sp. acta 3062. J Nat Prod. 2012;75:1018–24.
Rix U, Fischer C, Remsing LL, Rohr J. Modification of post-PKS tailoring steps through combinatorial biosynthesis. Nat Prod Rep. 2002;19:542–80. https://doi.org/10.1039/B103920M.
Lairson LL, Henrissat B, Davies GJ, Withers SG. Glycosyltransferases: structures, functions, and mechanisms. Annu Rev Biochem. 2008;77:521–55. https://doi.org/10.1146/annurev.biochem.76.061005.092322.
Kobayashi Y, Lee J, Tezuka K, Kishi Y. Toward creation of a universal NMR database for the stereochemical assignment of acyclic compounds: the case of two contiguous propionate units. Org Lett. 1999;1:2177–80. https://doi.org/10.1021/ol9903786.
Caradec T, et al. Dactylosporolides: Glycosylated macrolides from Dactylosporangium fulvum. J Nat Prod. 2022;85:2714–22. https://doi.org/10.1021/acs.jnatprod.2c00484.
Son S, et al. Catenulisporolides, glycosylated triene macrolides from the chemically underexploited actinomycete Catenulispora species. Org Lett. 2018;20:7234–8. https://doi.org/10.1021/acs.orglett.8b03160.
Fleury E, et al. Advances in the universal NMR database: toward the determination of the relative configurations of large polypropionates. Eur J Org Chem. 2009;2009:4992–5001. https://doi.org/10.1002/ejoc.200900616.
Kobayashi Y, Tan CH, Kishi Y. Stereochemical assignment of the C21–C38 portion of the desertomycin/oasomycin class of natural products by using universal NMR databases: prediction. Angew Chem. 2000;112:4449–51. https://doi.org/10.1002/1521-3757(20001201)112:23<4449::AID-ANGE4449>3.0.CO;2-0.
Kobayashi Y, Czechtizky W, Kishi Y. Complete stereochemistry of Tetrafibricin. Org Lett. 2003;5:93–96. https://doi.org/10.1021/ol0272895.
Seike H, Ghosh I, Kishi Y. Attempts to assemble a universal NMR database without synthesis of NMR database compounds. Org Lett. 2006;8:3861–4. https://doi.org/10.1021/ol061580t.
Zheng J, Taylor CA, Piasecki SK, Keatinge-Clay AT. Structural and functional analysis of A-type ketoreductases from the amphotericin modular polyketide synthase. Structure. 2010;18:913–22. https://doi.org/10.1016/j.str.2010.04.015.
Kwan DH, Schulz F. The stereochemistry of complex polyketide biosynthesis by modular polyketide synthases. Molecules. 2011;16:6092–115. https://doi.org/10.3390/molecules16076092.
Zhang P, Zhang Z, Zhang L, Wang J, Wu C. Glycosyltransferase GT1 family: Phylogenetic distribution, substrates coverage, and representative structural features. Comput Struct Biotechnol J. 2020;18:1383–90. https://doi.org/10.1016/j.csbj.2020.06.003.
Acknowledgements
This work was supported by Scientific and Engineering Research Board (SERB), Department of Science and Technology [ECRA/2016/000788 and EEQ/2016/000268]; and Council of Scientific and Industrial Research (CSIR), New Delhi [MLP/0027]. Yedukondalu Nalli acknowledges Scientific and Engineering Research Board (SERB) for providing National Postdoctoral Fellowship (N-PDF). Sanju Singh acknowledges Council of Scientific and Industrial Research (CSIR) for JRF and SRF fellowship. Padmaja D. Wakchaure acknowledges CSIR, New Delhi, India, for the GATE-SRF fellowship. Pankaj Kumar acknowledges Department of Biotechnology (DBT) for providing JRF and SRF fellowship. Central Instrumentation facility CSIR-CSMCRI is acknowledged for acquiring MS, FT-IR, CD, and NMR spectral data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumar, P., Nalli, Y., Singh, S. et al. Dactylides A−C, three new bioactive 22-membered macrolides produced by Dactylosporangium aurantiacum. J Antibiot 76, 503–510 (2023). https://doi.org/10.1038/s41429-023-00632-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-023-00632-z
- Springer Japan KK