Abstract
An essential protein regulatory system in cells is the ubiquitin-proteasome pathway. The substrate is modified by the ubiquitin ligase system (E1-E2-E3) in this pathway, which is a dynamic protein bidirectional modification regulation system. Deubiquitinating enzymes (DUBs) are tasked with specifically hydrolyzing ubiquitin molecules from ubiquitin-linked proteins or precursor proteins and inversely regulating protein degradation, which in turn affects protein function. The ubiquitin-specific peptidase 32 (USP32) protein level is associated with cell cycle progression, proliferation, migration, invasion, and other cellular biological processes. It is an important member of the ubiquitin-specific protease family. It is thought that USP32, a unique enzyme that controls the ubiquitin process, is closely linked to the onset and progression of many cancers, including small cell lung cancer, gastric cancer, breast cancer, epithelial ovarian cancer, glioblastoma, gastrointestinal stromal tumor, acute myeloid leukemia, and pancreatic adenocarcinoma. In this review, we focus on the multiple mechanisms of USP32 in various tumor types and show that USP32 controls the stability of many distinct proteins. Therefore, USP32 is a key and promising therapeutic target for tumor therapy, which could provide important new insights and avenues for antitumor drug development. The therapeutic importance of USP32 in cancer treatment remains to be further proven. In conclusion, there are many options for the future direction of USP32 research.
Similar content being viewed by others
Facts
-
USP32 is involved in a variety of cell biological processes, such as cell cycle, proliferation, invasion, migration, and DNA damage repair.
-
USP32 acts as an oncogene in a variety of tumors. Therefore, USP32-specific inhibitors can be developed to study its role in tumors.
-
We summarize the structure and biological function of USP32 and discuss the mechanism of USP32 action in a variety of tumors.
Open Questions
-
The expression of USP32 is often out of balance, especially in cancer. Does USP32 have any other functions among many types of tumors?
-
How does USP32, a target for many cancer therapies, specifically regulate the content and function of relevant proteins through deubiquitination?
-
USP32 has a regulatory effect on a wide range of tumors, so it is possible to provide ideas for treating tumors by modulating USP32 proteins.
Introduction
Proteins are the most important performers of various cellular functions, and their proper function determines whether life activities can be carried out in an orderly and efficient manner, in which post-translational modification (PTM) plays a crucial role [1,2,3]. As a complex mechanism of biological function regulation, PTM is very important for many key events involving cellular response [4]. In general, intracellular proteins undergo several sorts of modifications upon translation, including phosphorylation, acetylation, methylation, and ubiquitination, each of which is associated with one or more distinct functions [5]. Ubiquitination is one of the post-translational modifications, which refers to the covalent binding of ubiquitin to target proteins catalyzed by a number of different enzymes. The series of enzymes refer to three enzymes that cooperate with each other in the process of ubiquitin cascade: E1 ubiquitin activating enzyme, E2 ubiquitin binding enzyme, and E3 ubiquitin ligase [6, 7]. Hundreds of ubiquitin ligases are found in mammals, suggesting that the diversity of E3 ligases provides an accurate basis for substrate selection [8]. The ubiquitin-proteasome pathway (UPP) is responsible for 80–90% of eukaryotic protein degradation [9]. Ubiquitin is precisely linked to the target protein or to the ubiquitin chain that has already been linked to the target protein under the sequential catalysis of the E1, E2, and E3 enzymes. The E3 ubiquitin ligase controls the target protein’s precise identification, and the human body has an E4 enzyme-ubiquitin chain extension factor that may lengthen the ubiquitin chain to produce a polyubiquitin chain [10] (Fig. 1). In addition, ubiquitin modification can also occur on the N-terminal of protein and some other amino acids (cysteine, serine, threonine) [11, 12].
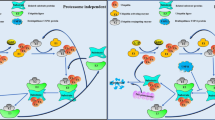
Multiple ongoing steps are required for the target protein’s ubiquitin breakdown. Ubiquitin is first activated by E1 enzymes when ATP (adenosine triphosphate) supplies a specific amount of energy. Ubiquitin activase E1 then sends the active ubiquitin molecules to E2 enzymes, and ubiquitin ligase E3 binds E2-binding ubiquitin to the target protein. The substrate protein’s ubiquitin molecule is extended by the E4 ubiquitin chain extension factor, and the tagged protein’s amino acid tail then forms a short ubiquitin molecular chain. Finally, the ubiquitin-tagged substrate protein is selectively recognized by the 26 S proteasome. The binding of 20 s catalytic core particles to 19 s regulatory complex, which binds the substrate protein tagged by ubiquitin chain and transfers the protein substrate to 20 s catalytic core under the energy of ATP, results in the formation of the structure of the 26 S proteasome. The substrate protein is degraded into small oligopeptides of less than 25 amino acids at the proteolytic β subunit of 20 s center, which will eventually be degraded into amino acids by protease in the cytoplasm. Ubiquitin molecules are recovered into the cytoplasmic pool. The deubiquitinating enzymes (DUBs) family hydrolyzes ubiquitin molecules from substrate proteins that include ubiquitin chains by hydrolyzing the ester, peptide, or isopeptide linkages at the carboxyl terminus of ubiquitin, and this process inversely controls protein deterioration. Ubiquitin molecules are also recycled into the cytoplasmic pool to exert their functions.
Ubiquitin is a strictly regulated and reversible process. DUBs can undo ubiquitin modification by hydrolyzing peptide or isopeptide links between ubiquitin molecules or between ubiquitin and substrate proteins [13]. Deubiquitinating enzymes not only inhibit ubiquitin processes, but also promote them by disassembling ubiquitin inhibitors, recycling ubiquitin molecules, and proofreading ubiquitin processes [12], which form a complex regulatory network with the ubiquitin system. Currently, there are seven families of ubiquitin-specific proteases (USPs), ubiquitin c-terminal hydrolases (UCHs), ovarian tumor proteases (OTUs), JAMMs (also known as MPN+), MJDs (also known as Josephins), MINDY family, and ZUP1 family, which make up human DUBs [14]. Cysteine proteases are categorized into six families (USPs, UCHs, OTUs, MJDs, MINDYs, and ZUP1), whereas zinc-dependent metalloproteinases make up the JAMM family [15]. Every family of DUBs is conformed from yeast to humans, with the exception of MJDs [16]. The conversion rate, activation, circulation, and localization of several proteins are all impacted by DUBs activity, which is crucial for intracellular homeostasis, protein stability, and a variety of signal pathways. Alterations in DUB function are simultaneously associated with many diseases, including cancer [17]. It is well known that USPs are the largest class of DUBs, with more than 60 members, and there is growing interest in its function and the important role that substrates play in the pathogenesis of many diseases, including cancer. As was already mentioned, USPs are a class of cysteine-dependent protein hydrolases whose catalytic structural domain (also known as the USP structural domain) is one of their most conserved structural domains. Other domains in USPs include those that can predict ubiquitin binding, such as the zinc finger ubiquitin-specific protease (ZnF-UBP) domain, the ubiquitin interaction motif (UIM) domain, and the ubiquitin associated (UBA) domain [18]. At the same time, although the additional domain around it does not bind to the substrate, the conformational change occurs during ubiquitin binding [18]. Dysfunction in the USPs family can lead to many diseases, including cancer [19], metabolic diseases [20], and neurodegenerative diseases [21], among others. Among them, the research on the relationship between USPs and malignant tumor has become a hot spot, and many studies have shown that targeted intervention of this pathway is expected to become an ideal anti-tumor therapy strategy. We list the function of USPs in carcinogenesis in Table 1.
The USPs family includes the ubiquitin-specific protease 32 (USP32), also known as NY-REN-60, which codes an ancient and unique gene. USP32 highly conserved homologous genes are evident in all intact post-animal genomes [22]. USP32 is a newly identified de-ubiquitin enzyme in recent years, which was first described in a research article in 2003, and then the related research on USP32 was gradually unveiled (Fig. 2). In recent years, the importance of USP32 continues to be revealed, especially in cancer, where USP32 is often disordered.
Properties of USP32
USP32 chromosomal location, subcellular localization
The ancient and highly conserved human USP32 gene is found on the chromosomal band 17q23 [22]. According to the findings of the multi-tissue cDNA panel, USP32 was highly expressed in the leukocytes, thymus, spleen, testis, prostate, ovary, small intestine, and colon [23]. We queried USP32 in human tissues through the Human Protein Atlas website, the testis showed the greatest RNA expression of USP32 (Fig. 3). USP32 is upregulated in a variety of cancers, including small cell lung cancer [24], gastric cancer [25, 26], breast cancer [23, 27, 28], epithelial ovarian cancer [29], glioblastoma [30], gastrointestinal stromal tumor [31], pancreatic duct adenocarcinoma [32] and acute myeloid leukemia [33].
Available online: https://www.proteinatlas.org/ENSG00000170832-USP32/tissue (accessed on 26 March 2023).
Endogenous USP32 is found in the cytoplasm and membrane, according to the findings of a subcellular separation experiment [34] and a fluorescence protection experiment [23]. This is consistent with earlier findings that USP32 is an active membrane-bound ubiquitin protease [24]. In the investigation of drug resistance in tumor cells, USP32, a membrane protein, can result in resistance to the anticancer medication YM155 by interfering with the steady expression of SLC35F2 [28]. Subcellular localization studies also show that USP32 may be co-located with Golgi [23], and some studies have found that USP32 can regulate the participation of small GTPase Rab7 in Golgi endosome selection [35]. The absence of USP32 will also affect the structure and function of intracellular lysosomal vesicles and the transport of nuclear endosomes, thus affecting the occurrence of some diseases [35].
The structure and activities of USP32
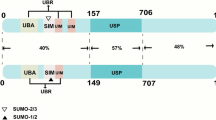
Human USP32 is composed of 1604 amino acids [23] and has an estimated molecular weight of 182KDa. The multi-domain protein includes calcium-binding EF-hand with signal transduction mechanism, DUSP (domain in USP) expected to mediate protein-protein interaction, a USP catalytic domain, and two ubiquitin-like (UBL) domains and c-terminal prenylation sites (CAAX box) (Fig. 4A) [36]. Among them, the USP catalytic structural domain is present in all member structures of the USPs and is the most important functional structural domain in the USPs [37], containing key cysteine and histidine residues [38]. The USP catalytic domain has strong homology in the regions around catalytic Cys box and His box [37]. All human DUBs have the two distinct structures of the N-terminal calcium-binding EF hand and the C-terminal prenylation site (CAAX box), the latter of which is present only in the USP32 structure and is associated with USP32-doped membrane structures [34]. Researchers discovered that the USP domain in USP32, which has substrates for both mono- and double-ubiquitin cleavage, does not often tend to break one of the eight ubiquitin connections, M1, K6, K11, K27, K29, K33, K48, and K63 [35]. In addition, a ubiquitin C-terminal hydrolase (UBP12) exists in the 518-1316 amino acid position of the USP32 structure, which can undergo post-translational modification and protein turnover, while the ubiquitin carboxyl terminal hydrolase has the 1231-1564 amino acid position of the USP32 structure (Fig. 4A). The structure of USP32 predicted by Alpha Fold [39, 40] is shown in Fig. 4B.
A schematic diagram of the composition of the USP32 domain; B the front and top view directions of the USP32 structure. Available online: https://alphafold.ebi.ac.uk/entry/Q8NFA0 (accessed on March 25th, 2023).
USP32 has a variety of molecular functions and characteristics. Firstly, USP32 is widely expressed in tissues and has the conservative peptidase characteristics of ubiquitin-specific proteases [23]. USP32 locates on the cell membrane by lipid anchoring and catalyzes the conversion of C-terminal thioesters to free ubiquitin and mercaptan, which may affect a variety of cell processes. Secondly, USP32 supports the role of Rab7 in the transport and circulation of MVB through two different mechanisms [35]. In a genome-wide sense, USP32 is linked to a higher incidence of Parkinson’s disease [41]. USP32 may also be the target gene of some miRNAs and participates in the ubiquitin proteolysis pathway [42]. USP32 gene also has sulfhydryl-dependent ubiquitin-specific protease activity, which can bind calcium ions [43]. The ability of USP32 to bind to proteins or protein complexes allows it to take part in the Golgi’s nuclear endosome selection process [44]. As a result, USP32 is engaged in a variety of biological activities. The following sections will explain USP32’s specific roles and mechanisms.
A homologue of USP32
USP32 and USP6
In human samples, the homologue of USP32 is USP6 (also known as TRE2). In actuality, phylogenetic study reveals that it is a chimera of two genes, USP32 (NY-REN-60) and TBC1D3, and shares 97% of its nucleotides with USP32 [22]. More than 20 years ago, USP6 was discovered to be a new oncogene [45]. It functions as a cysteine protease to control the degradation of ubiquitin and to take part in a number of cellular functions, such as intracellular transport, protein synthesis, inflammatory signaling, and cell transformation [46]. USP6 is a gene chaperone that has been discovered to have numerous partners. When these partners fuse with USP6, USP6 is transcriptionally activated and participates in the Wnt/β-catenin pathway, NF-κB pathway, and JAK1-STAT3 pathway, among other pathways involved in tumorigenesis [47]. Meanwhile, both USP6 and USP32 as ubiquitin protein hydrolases have conserved enzyme structural domains [22].
USP32 and PoUSP32
We know that USP32 is widely expressed in mammals and is involved in tumor activity, fragile X syndrome [48] and chronic nephropathy [42]. There is no previous record of USP32 in fish, and a recent study found that the sequence of Japanese flounder PoUSP32 is 73.6% and 74.5% identical to that of human and mouse USP32, respectively. The PoUSP32 of Japanese flounder consists of 1613 amino acid residues, which is expressed in 9 different tissues of Japanese flounder, but the expression level is significant in the intestine. PoUSP32 is similar to mammalian USP32 in that it has the conserved enzyme domain of USPs family and the conservative function of deubiquitin. In order to better understand the role of PoUSP32, scientists discovered through in vivo and in vitro investigations that pol-miR-363-3p48 interacts with the 3’UTR of PoUSP32 and decreases the expression of PoUSP32. Vitro experiments prevented dolphin infection with Streptococcus by down-regulation of PoUSP32 or overexpression of polmiR-363-3p in toothfish cells. Vivo research have shown that overexpression of USP32 and interfering with pol-miR-363-3p can increase Streptococcus dolphin infection in Paralichthys olivaceus tissue. These findings imply that pathogen infection substantially affects the regulation of PoUSP32 and pol-miR-363-3p expression [49].
The role of USP32 in several cancers
In recent years, it has been found that USP32 can regulate the growth and development of different tumors through the catalytic activity of deubiquitinating enzymes. We investigated the expression of USP32 in various malignant tumors by using the TCGA database and assessed the expression of USP32 in 33 tumor tissues and paraneoplastic tissues (Fig. 5). In terms of statistical significance, six of them, including cholangiocarcinoma (CHOL), esophageal carcinoma (ESCA), head and neck squamous cell carcinoma (HNSC), hepatocellular carcinoma (LIHC), and stomach adenocarcinoma (STAD), displayed increased levels of USP32 mRNA expression. However, in four additional tumor samples-glioblastoma multiforme (GBM), kidney suspicious cell (KICH), thyroid cancer (THCA), and endometrial cancer (UCEC)-USP32 gene expression were dramatically downregulated (Fig. 5A). Unexpectedly, glioblastoma has a low expression level of USP32 in the TCGA database. This outcome does not appear to be in line with those mentioned in the literature. We posit that this finding may be explained by post-translational alteration, tumor heterogeneity, or a small sample size. Thus, we need to learn more about USP32 expression in glioblastoma and its function. Additionally, we examined TCGA data sets to find USP32 expression in human cancer tissues that were linked with nearby normal tissues. We found that compared to normal tissue, USP32 expression was significantly elevated in 10 of 23 cancers, such as bladder thelial carcinoma (BLCA) and breast invasive carcinoma (BRCA), cholangiocarcinoma (CHOL), esophageal carcinoma (ESCA), head and neck squamous cell carcinoma (HNSC), kidney chromophobe (KICH), kidney renal papillary cell carcinoma (KIRP), liver hepatocellular carcinoma (LIHC), stomach adenocarcinoma (STAD) and thyroid carcinoma (THCA) (Fig. 5B). The increased expression of USP32 in different cancers is a necessary condition for cell proliferation and tumorigenesis. According to several studies, high levels of USP32 expression are associated with poor prognosis in various malignant tumors. In order to fully understand the mechanism of action of USP32 in different malignant tumors, researchers need to learn USP32 and its regulation of downstream targets.
A The level of USP32 mRNA expression in tumor tissues and surrounding tissues according to the TCGA database. B USP32 expression levels in nearby normal tissues and matched malignant tissues from human cancer patients, according to the TCGA database. Box plots showing USP32 log2(TPM) expression in various malignancies and log2(TPM) transcript count per million. USP32 expression is denoted by the colors red and blue in normal and malignant tissues, respectively. The Wilcoxon test was used to determine the significant difference in USP32 expression between tumor and normal tissue. The significance level is indicated by the number of stars on top of the box plots (*p < 0.05, **p < 0.01, ***p < 0.001).
SCLC
About 15% of all lung cancers are small cell lung cancers (SCLC), which are distinguished by an unusually high rate of proliferation, significant early metastasis, and a poor prognosis [50, 51]. The preferred first- and second-line treatment for small cell lung cancer (SCLC) remains chemotherapy [52]. For small-cell lung cancer, USP32 is thought to be a potential therapeutic target. According to a study, USP32 is highly expressed in small-cell lung cancer. The association between high levels of USP32 and low survival rates was significant. In vitro experiments confirm that USP32 promotes tumor growth. It is interesting to note that USP32 can influence the development of small-cell lung cancer’s cell cycle. USP32 knockdown increased apoptosis in addition to causing cell cycle arrest in the G0/G1 phase, with concomitant increases in Caspase3, cleaved PARP and P53-related apoptotic proteins. The marker protein of epithelial–mesenchymal transformation (EMT) may be affected by the deletion of the USP32 gene, which can upregulate the expression of E-calcineurin and down-regulate the expression of N-calcineurin protein. Thus, the ability of small cell lung cancer cells to proliferate and generate clones, as well as the ability of cells to invade and migrate, could be inhibited by interfering with USP32 [24] (Fig. 6). These results will help to elucidate how small cell lung cancer tumors progress and guide the establishment of therapeutic targets for small cell lung cancer.
The expression of USP32 protein is negatively correlated with MicroRNAlet-7a in breast cancer, which can suppress USP32 expression and impede the growth of breast cancer cells. ETV1 controls the expression of USP32, which in turn stabilizes Rab35 by ubiquitinating Lys48 (K48) on Rab35 and increases exocrine secretion of imatinib-resistant gastrointestinal stromal tumors. USP32 supports the growth of acute myeloid leukemia (AML) through deubiquitinating and stabilizing Rap1b, while hsa_circ_0013880 controls USP32 expression in AML through miR-148a3p/miR-20a-5p. USP32 is extensively expressed in lung cancer tissues, where it dramatically speeds up cell division, encourages cell growth, and prevents cell death. USP32 contributes in the onset and progression of gastric cancer by controlling SHMT2, and USP32 enhances gastric carcinogenesis and cisplatin resistance in gastric cancer by eliminating ubiquitin and stabilizing SMAD2. USP32 increases resistance to YM155 in breast cancer cells through modulation of SLC35F2. By controlling the expression of FDFT1 protein, USP32 contributes to the development and progression of epithelial ovarian cancer. By controlling the cell cycle, DNA replication, basal excision repair, and mismatch repair in glioblastoma, USP32 promotes GBM.
Gastric cancer
Gastric cancer is characterized by high incidence, poor prognosis, and cellular and molecular heterogeneity, which makes it a major health problem in the world [53]. Effective treatments for gastric adenocarcinoma include systemic chemotherapy, radiation, surgery, immunotherapy, and targeted therapy [54]. Each type of gastric cancer has its own characteristics, and each gastric cancer patient can be effectively treated clinically through molecular targeting therapy [55]. Researchers have found a link between SMAD2 and USP32. SMAD2 is a key element of transforming growth factor signaling pathway in the occurrence and development of gastric cancer. USP32 has been discovered as an oncogene implicated in the development and metastasis of GC cells due to the considerable inhibition of GC cell proliferation and migration caused by USP32 gene knockout or deletion in both vivo and in vitro. It is more likely that USP32 will be used as a possible biomarker and therapeutic target for gastric cancer because of the high expression of USP32 in this disease and its relationship to the short overall survival rate and high T stage of gastric cancer patients. The scientists discovered that USP32 modulates SMAD2 expression in a ubiquitin protease-dependent manner. Gastric cancer cells’ cisplatin resistance can be decreased by down-regulating USP32 expression, indicating that USP32 is involved in the development of cisplatin resistance. Meanwhile, rescue experiments showed that overexpression of SMAD2 could reverse the effects of silenced USP32 on gastric cancer cell growth, metastasis and cisplatin resistance. These observations suggest that SMAD2 is located downstream of USP32 in GC, and USP32 could regulate the expression of SMAD2 and promote gastric carcinogenesis and cisplatin resistance by enhancing the stability of SMAD2 protein in gastric cancer cells, so targeting USP32 may be a potential therapeutic strategy for GC [25] (Fig. 6).
Another study found that early and late stages of gastric cancer had higher expression levels of the USP32 and SHMT2 proteins, and immunohistochemistry analysis supported these findings. The researchers also found that after the silencing of USP32 expression in gastric cancer, the expression of SHMT2 decreased [26] (Fig. 6). Therefore, targeting the USP32-SHMT2 axis may provide some ideas for the treatment of gastric cancer.
Breast cancer
According to the most recent report on cancer statistics, breast cancer continues to be the most common tumor among women [56]. Because the occurrence and development of tumors are regulated by complex molecular mechanisms, the choice of treatment options and disease prognosis of breast cancer are challenged at the molecular level [57]. USP32 was found to have an elevated copy number in estrogen receptor (ER) positive tumors, which is one of the 81 gene copy number traits that predict the metastatic capacity of breast cancer [58]. Meanwhile, it has been noted that USP32 is one of the transcripts that are up-regulated in malignant breast epithelial cells [59], indicating that USP32 may be a helpful biomarker in a subclass of breast cancer cells. In one work, USP32 transcript analysis was expanded to breast cancer cell lines and primary tumors in order to examine USP32 expression in breast cancer cells. USP32 was discovered to be overexpressed in both primary breast tumors and breast cancer cell lines. MCF7 cells, a typical cell line in which USP32 was overexpressed, were examined for mutations, but none were found, demonstrating that USP32 was overexpressed in wild-type transcripts [23] (Fig. 6).
In another study, we found that microRNA let-7a reduced the amount of USP32 protein in MCF-7 cells. USP32 was identified by bioinformatics study as a miR let-7a target gene. The 3’UTR location of USP32 mRNA was found to be the target of miR let-7a, and miR let-7a was able to reverse the reduction in ubiquitin levels caused by USP32. As a result, miR let-7a might suppress the expression of USP32 protein by directly binding to the USP32 3’UTR. As a result, miR let-7a can regulate USP32 in BCa, which can decrease proliferation and offer a new potential target for the treatment of breast cancer [27] (Fig. 6).
USP32 has reportedly been linked to the medication resistance mechanism in breast cancer. The uptake of the anti-cancer drug candidate YM155 by breast cancer cells is controlled by the expression of a solute carrier protein called SLC35F2. In contrast to SLC35F2, USP32 is highly expressed in breast, colon and lung malignancies. These cancer cells are resistant to YM155, a small molecule drug that targets a variety of cancers. Researchers further found that the direct binding of USP32 and SLC35F2 in the endoplasmic reticulum could negatively regulate the stability of SLC35F2 and promote drug resistance to YM155 in breast cancer cells [32] (Fig. 6). Therefore, inhibition of USP32 can increase the protein expression of SLC35F2 and improve the chemotherapeutic effect of YM155 on breast cancer, opening up a new pathway for clinical drug resistance treatment.
Epithelial ovarian cancer
The most frequent type of gynecological cancer to cause death is epithelial ovarian cancer, which has a high mortality rate in part because of the patient’s late stage at the time of diagnosis [60]. The focus of immunotherapy in EOC clinical trials is similar to those of many other cancer subtypes, and programmed cell death 1 (PD-1) is its main inhibitor [61]. The standard of care for all female patients with epithelial ovarian cancer is the genetic identification of altered genes that affect treatment [62]. In one study, the researchers used the pertinent database search to discover that USP32 was substantially expressed in human ovarian cancer peritoneal tumors and was regulated in ovarian cancer. Epithelial ovarian cancer has a bad prognosis, which is directly associated with USP32. A peritoneal tumor’s ability to metastasize can be prevented by inhibiting USP32. Farnesyl-diphosphate farnesyltransferase 1 (FDFT1) in the valproic acid pathway is a new USP32 substrate regulated by the UPS, according to proteomic research. FDFT1, an oncogene and suppressor gene, has a direct connection to the growth of stem cells and cancerous cells. They showed that USP32 and FDFT1 expression was higher in tumor spheroids than in adnexal cells, and that the FDFT1 inhibitor ZA, blocking USP32, or interfering with the mevalonate pathway might inhibit the formation of tumor spheres in epithelial ovarian cancer (Fig. 6). These results suggest that USP32-FDFT1 axis promotes the migration of epithelial ovarian cancer and may become a new target for EOC therapy [29].
Glioblastoma
The most prevalent and aggressive high-grade primary malignant tumor of the adult central nervous system (CNS), glioblastoma has a very bad prognosis [63, 64]. Combination therapy may yield better results because the survival rate of currently approved GBM treatments is low and immunotherapy for GBM typically lacks sufficient clinical expertise to translate into meaningful benefits [65]. The elevated expression of USP32 protein in GBM tissues was validated by a study. Glioblastoma cell lines (Umur118 MG, Umur87 MG, A172, T98G, and Umur251 MG) had higher mRNA and protein levels of USP32 than normal brain cells SVG p12 [30]. In addition, they found that the higher the expression of USP32 in GBM, the worse the prognosis. Cancer cell proliferation and migration can be stopped in vitro by inhibiting USP32, and tumor growth can be stopped in vivo by down-regulating the USP32 gene. In their research, they discovered that USP32 can speed up the transition of the cell cycle from the G0 phase to the G1 phase [30], start DNA replication, and so enhance the growth of cancer cells. Then, RT-qPCR tests were used to confirm the effects of USP32 on a few functional molecules. The findings demonstrate that USP32 controls several genes’ expression and is a key oncogene in basal excision repair and mismatch repair [30] (Fig. 6). Therefore, USP32 is a promising new molecular target, from which researchers can study new treatments to control the development of glioblastoma.
Gastrointestinal stromal tumor
The gastrointestinal stromal tumor is a kind of malignant stromal tumor, which is treated separately because of its special histogenesis, clinical manifestation, and specific treatment [62]. Local GIST can be treated by surgery [66], while imatinib can be used for unresectable advanced and metastatic gastrointestinal stromal tumors [67]. However, about half of the patients will develop secondary imatinib resistance after two years [68]. Recent research has revealed that the USP32-Rab35 axis is crucial for controlling treatment resistance in gastrointestinal stromal tumors. The exocrine secreted by tumor cells has a great influence on the transmission of drug resistance in drug-sensitive cells. Previous studies have found that Rab35 regulates exocrine secretion through exosome recovery and early sorting [69]. A recent study discovered and concluded that Rab35 can influence the spread of drug resistance by controlling the exocrine secretion of gastrointestinal stromal tumors, and that gastrointestinal stromal tumors considerably overexpressed Rab35 [31]. In terms of mechanism, the expression of USP32 and transcription factor ETV1 have a positive correlation. In gastrointestinal stromal tumors that are imatinib-resistant, ETV1 increases the expression of USP32. USP32 promotes the exocrine secretion of imatinib-resistant gastrointestinal stromal tumors by decreasing the Lys48 (K48) ubiquitination of Rab35, protecting it from proteasome destruction (Fig. 6). Their research suggests that individuals with gastrointestinal stromal tumors resistant to imatinib may benefit from focusing on the USP32-Rab35 axis as a potential therapeutic target.
Acute myeloid leukemia
A lethal myeloid malignant tumor that develops in the blood system is acute myeloid leukemia (AML) [70]. According to WHO, the tumor is divided into four categories. It is mainly characterized by primordial and juvenile myeloid cells in the bone marrow and peripheral blood, but does not include lymphoid cell dysplasia, accompanied by infection, anemia and bleeding symptoms [71]. The disease can occur at any age, but mainly in the elderly [72]. At present, there are many options for the treatment of AML. The realization of complete response (CR) is the general method for the treatment of AML. Other treatment methods include induction therapy, induction therapy after remission, non-intensive therapy for newly diagnosed patients, recurrent refractory AML, etc [73]. A study reveals the regulatory mechanisms of USP32 and circRNAs in acute myeloid leukemia. In AML patients, the expression of USP32 and hsa_circ_0013880 is up-regulated and positively linked. Hsa_circ_0013880 overexpression can either positively regulate USP32 protein and increase AML cell proliferation. Through the analysis of bioinformatics and related experiments, we know that miR-148a3p/miR-20a-5p can regulate USP32, and hsa_circ_0013880 can combine with miR-148a3p/miR-20a-5p. In the BMNCs of patients with AML, the researchers found that USP32 acts as a well-intentioned deubiquitinase for Ras-related proteins (Rap1b). USP32 could interact with Rap1b and stabilize Rap1b. Knockdown of USP32 in HL-60 and U937 cells significantly reduced the expression of Rap1b. Overexpression of Rap1b largely reversed USP32 knockdown-induced apoptosis. As a result, the hsa_circ_0013880/USP32/-Rap1b axis plays a crucial regulatory role in the emergence and progression of AML (Fig. 6). Future studies of this axis may provide a new approach to the treatment of acute myeloid leukemia [33].
Conclusions
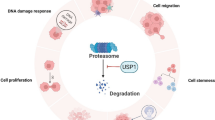
There are two main ways of protein degradation in human cells: one is in lysosome, which mainly degrades foreign proteins and has poor selectivity for proteins. The other is degraded in the proteasome, via an ATP-dependent pathway (energy-demanding), after ubiquitination modifications, which mainly degrades intracellular proteins with abnormal structure and short life span. The polyubiquitination of substrate proteins and proteasome degradation can affect or regulate a wide range of physiological processes, including gene transcription, cell cycle regulation, immune response, cell receptor function, tumor growth, inflammatory process, and others. As a member of the USPs family, USP32 controls DNA damage repair, cell cycle, cancer-related signaling, and protein stability.
This study analyzes the current findings on USP32, including its structure and biological functions, and discusses how it regulates the growth of various types of tumors. The presence of endogenous USP32 in both the cell membrane and cytoplasm determines that USP32 can be involved in the regulation of a number of small molecules. Meanwhile, USP32 is a multi-structural domain protein that possesses multiple protein properties and biological functions. Current researchers have found that the homologs of USP32 in human samples and in fish are USP6 and PoUSP32, respectively, which share the conserved enzyme structural domains of the USPs family and deubiquitination functions. USP32 is expressed in a wide range of tissues, but its expression is frequently dysregulated in some tumors. Therefore, a possible target for cancer therapy is USP32, which has been shown an important role in many malignant tumors and chemotherapy resistance. In light of the fact that USP32 is a target for the treatment of tumors and is dependent on ubiquitin-modified target genes as oncogenes, more research on USP32 as a means of regulating the efficacy of cisplatin and other DNA-damaging drugs should be done. This necessitates a comprehensive investigation, review, and evaluation of its therapeutic significance and application.
References
Zhang X-N, Cheng Q, Chen J, Lam AT, Lu Y, Dai Z, et al. A ribose-functionalized NAD+ with unexpected high activity and selectivity for protein poly-ADP-ribosylation. Nat Commun. 2019;10:4196.
Zhao P, Yao R, Zhang Z, Zhu S, Li Y, Ren C, et al. Eukaryotic ribosome quality control system: a potential therapeutic target for human diseases. Int J Biol Sci. 2022;18:2497–514.
Bae J, Kim H, Kim G, Song J, Kim H. Dendrimer‐like supramolecular assembly of proteins with a tunable size and valency through stepwise iterative growth. Adv Sci (Weinh). 2021;8:2102991.
Olsen JV, Blagoev B, Gnad F, Macek B, Kumar C, Mortensen P, et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell. 2006;127:635–48.
Millar AH, Heazlewood JL, Giglione C, Holdsworth MJ, Bachmair A, Schulze WX. The scope, functions, and dynamics of posttranslational protein modifications. Annu Rev Plant Biol. 2019;70:119–51.
Swatek KN, Komander D. Ubiquitin modifications. Cell Res. 2016;26:399–422.
Kleiger G, Mayor T. Perilous journey: a tour of the ubiquitin-proteasome system. Trends Cell Biol. 2014;24:352–9.
Cheng J, Guo J, Wang Z, North BJ, Tao K, Dai X, et al. Functional analysis of Cullin 3 E3 ligases in tumorigenesis. Biochim Biophys Acta Rev Cancer. 2018;1869:11–28.
Tomko RJ, Hochstrasser M. Incorporation of the Rpn12 subunit couples completion of proteasome regulatory particle lid assembly to lid-base joining. Mol Cell. 2011;44:907–17.
Baranes-Bachar K, Levy-Barda A, Oehler J, Reid DA, Soria-Bretones I, Voss TC, et al. The ubiquitin E3/E4 ligase UBE4A adjusts protein ubiquitylation and accumulation at sites of dna damage, facilitating double-strand break repair. Mol Cell. 2018;69:866–878.e7.
McClellan AJ, Laugesen SH, Ellgaard L. Cellular functions and molecular mechanisms of non-lysine ubiquitination. Open Biol. 2019;9:190147.
Dittmar G, Winklhofer KF. Linear ubiquitin chains: cellular functions and strategies for detection and quantification. Front Chem. 2019;7:915.
Fennell LM, Rahighi S, Ikeda F. Linear ubiquitin chain-binding domains. FEBS J. 2018;285:2746–61.
Liu B, Ruan J, Chen M, Li Z, Manjengwa G, Schlüter D, et al. Deubiquitinating enzymes (DUBs): decipher underlying basis of neurodegenerative diseases. Mol Psychiatry. 2022;27:259–68.
Harhaj EW, Dixit VM. Deubiquitinases in the regulation of NF-κB signaling. Cell Res. 2011;21:22–39.
Clague MJ, Urbé S, Komander D. Breaking the chains: deubiquitylating enzyme specificity begets function. Nat Rev Mol Cell Biol. 2019;20:338–52.
Fraile JM, Quesada V, Rodríguez D, Freije JMP, López-Otín C. Deubiquitinases in cancer: new functions and therapeutic options. Oncogene. 2012;31:2373–88.
Komander D, Clague MJ, Urbé S. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol. 2009;10:550–63.
Young M-J, Hsu K-C, Lin TE, Chang W-C, Hung J-J. The role of ubiquitin-specific peptidases in cancer progression. J Biomed Sci. 2019;26:42.
Kitamura H. Ubiquitin-specific proteases (USPs) and metabolic disorders. Int J Mol Sci. 2023;24:3219.
Buneeva O, Medvedev A. Atypical ubiquitination and Parkinson’s disease. Int J Mol Sci. 2022;23:3705.
Paulding CA, Ruvolo M, Haber DA. The Tre2 (USP6) oncogene is a hominoid-specific gene. Proc Natl Acad Sci USA. 2003;100:2507–11.
Akhavantabasi S, Akman HB, Sapmaz A, Keller J, Petty EM, Erson AE. USP32 is an active, membrane-bound ubiquitin protease overexpressed in breast cancers. Mamm Genome. 2010;21:388–97.
Hu W, Wei H, Li K, Li P, Lin J, Feng R. Downregulation of USP32 inhibits cell proliferation, migration and invasion in human small cell lung cancer. Cell Prolif. 2017;50:e12343.
Dou N, Hu Q, Li L, Wu Q, Li Y, Gao Y. USP32 promotes tumorigenesis and chemoresistance in gastric carcinoma via upregulation of SMAD2. Int J Biol Sci. 2020;16:1648–57.
Li J, Bo Y, Ding B, Wang L. Understanding the regulatory role of USP32 and SHMT2 in the progression of gastric cancer. Cell J. 2023;25:222–8.
Liu C, Chen Z, Fang M, Qiao Y. MicroRNA let-7a inhibits proliferation of breast cancer cell by downregulating USP32 expression. Transl Cancer Res. 2019;8:1763–71.
Chandrasekaran AP, Kaushal K, Park C-H, Kim K-S, Ramakrishna S. USP32 confers cancer cell resistance to YM155 via promoting ER-associated degradation of solute carrier protein SLC35F2. Theranostics. 2021;11:9752–71.
Nakae A, Kodama M, Okamoto T, Tokunaga M, Shimura H, Hashimoto K, et al. Ubiquitin specific peptidase 32 acts as an oncogene in epithelial ovarian cancer by deubiquitylating farnesyl-diphosphate farnesyltransferase 1. Biochem Biophys Res Commun. 2021;552:120–7.
Chen S, Chen X, Li Z, Mao J, Jiang W, Zhu Z, et al. Identification of ubiquitin-specific protease 32 as an oncogene in glioblastoma and the underlying mechanisms. Sci Rep. 2022;12:6445.
Li C, Gao Z, Cui Z, Liu Z, Bian Y, Sun H, et al. Deubiquitylation of Rab35 by USP32 promotes the transmission of imatinib resistance by enhancing exosome secretion in gastrointestinal stromal tumours. Oncogene. 2023;42:894–910.
Wang Y, Zhou D, Kong Y, Yang Q, Ding Y, Wang W. USPs in pancreatic ductal adenocarcinoma: a comprehensive bioinformatic analysis of expression, prognostic significance, and immune infiltration. Biomed Res Int. 2022;2022:6109052.
Zhang H, Tao Y, Ding X, Wang Y, Wang X. Roles of the hsa_circ_0013880/USP32/Rap1b axis in the proliferation and apoptosis of acute myeloid leukemia cells. Acta Biochim Biophys Sin (Shanghai). 2023;55:382–93.
Hertel A, Alves LM, Dutz H, Tascher G, Bonn F, Kaulich M, et al. USP32-regulated LAMTOR1 ubiquitination impacts mTORC1 activation and autophagy induction. Cell Rep. 2022;41:111653.
Sapmaz A, Berlin I, Bos E, Wijdeven RH, Janssen H, Konietzny R, et al. USP32 regulates late endosomal transport and recycling through deubiquitylation of Rab7. Nat Commun. 2019;10:1454.
Xie Y, Li H, Luo X, Li H, Gao Q, Zhang L, et al. IBS 2.0: an upgraded illustrator for the visualization of biological sequences. Nucleic Acids Res. 2022;50:W420–W426.
Li Q, Ye C, Tian T, Jiang Q, Zhao P, Wang X, et al. The emerging role of ubiquitin-specific protease 20 in tumorigenesis and cancer therapeutics. Cell Death Dis. 2022;13:434.
Yue W, Chen Z, Liu H, Yan C, Chen M, Feng D, et al. A small natural molecule promotes mitochondrial fusion through inhibition of the deubiquitinase USP30. Cell Res. 2014;24:482–96.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9.
Varadi M, Anyango S, Deshpande M, Nair S, Natassia C, Yordanova G, et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 2022;50:D439–D444.
Pankratz N, Dumitriu A, Hetrick KN, Sun M, Latourelle JC, Wilk JB, et al. Copy number variation in familial Parkinson disease. PLoS ONE. 2011;6:e20988.
Jeong S, Oh JM, Oh K-H, Kim I-W. Differentially expressed miR-3680-5p is associated with parathyroid hormone regulation in peritoneal dialysis patients. PLoS ONE. 2017;12:e0170535.
Kour A, Deb SM, Nayee N, Niranjan SK, Raina VS, Mukherjee A, et al. Novel insights into genome-wide associations in Bos indicus reveal genetic linkages between fertility and growth. Anim Biotechnol. 2021;34:39–55.
Doherty LM, Mills CE, Boswell SA, Liu X, Hoyt CT, Gyori B, et al. Integrating multi-omics data reveals function and therapeutic potential of deubiquitinating enzymes. Elife. 2022;11:e72879.
Nakamura T, Hillova J, Mariage-Samson R, Onno M, Huebner K, Cannizzaro LA, et al. A novel transcriptional unit of the tre oncogene widely expressed in human cancer cells. Oncogene. 1992;7:733–41.
Legrand M, Jourdan M-L, de Pinieux G. Histopathogenesis of bone- and soft-tissue tumor spectrum with USP6 gene rearrangement: multiple partners involved in the tissue repair process. Histol Histopathol. 2023;38:247–60.
Hiemcke-Jiwa LS, van Gorp JM, Fisher C, Creytens D, van Diest PJ, Flucke U. USP6-associated neoplasms: a rapidly expanding family of lesions. Int J Surg Pathol. 2020;28:816–25.
Ma Y, Tian S, He S, Chen Q, Wang Z, Xiao X, et al. The mechanism of action of FXR1P-related miR-19b-3p in SH-SY5Y. Gene. 2016;588:62–68.
Sun Y-L, Guan X-L, Zhang P, Li M-F, Zhang J, Sun L. Pol-miR-363-3p plays a significant role in the immune defense of Japanese flounder Paralichthys olivaceus against bacterial and viral infection. Fish Shellfish Immunol. 2020;104:439–46.
Rudin CM, Brambilla E, Faivre-Finn C, Sage J. Small-cell lung cancer. Nat Rev Dis Prim. 2021;7:3.
Yin X, Li Y, Wang H, Jia T, Wang E, Luo Y, et al. Small cell lung cancer transformation: from pathogenesis to treatment. Semin Cancer Biol. 2022;86:595–606.
Yang S, Zhang Z, Wang Q. Emerging therapies for small cell lung cancer. J Hematol Oncol. 2019;12:47.
Seeneevassen L, Bessède E, Mégraud F, Lehours P, Dubus P, Varon C. Gastric cancer: advances in carcinogenesis research and new therapeutic strategies. Int J Mol Sci. 2021;22:3418.
Joshi SS, Badgwell BD. Current treatment and recent progress in gastric cancer. CA A Cancer J Clin. 2021;71:264–79.
Zeng Y, Jin RU. Molecular pathogenesis, targeted therapies, and future perspectives for gastric cancer. Semin Cancer Biol. 2022;86:566–82.
Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. Cancer J Clin. 2023;73:17–48.
Kawiak A. Molecular research and treatment of breast cancer. Int J Mol Sci. 2022;23:9617.
Zhang Y, Martens JWM, Yu JX, Jiang J, Sieuwerts AM, Smid M, et al. Copy number alterations that predict metastatic capability of human breast cancer. Cancer Res. 2009;69:3795–801.
Grigoriadis A, Mackay A, Reis-Filho JS, Steele D, Iseli C, Stevenson BJ, et al. Establishment of the epithelial-specific transcriptome of normal and malignant human breast cells based on MPSS and array expression data. Breast Cancer Res. 2006;8:R56.
Lheureux S, Gourley C, Vergote I, Oza AM. Epithelial ovarian cancer. Lancet. 2019;393:1240–53.
James NE, Woodman M, Ribeiro JR. Prognostic immunologic signatures in epithelial ovarian cancer. Oncogene. 2022;41:1389–96.
Kuroki L, Guntupalli SR. Treatment of epithelial ovarian cancer. BMJ. 2020;371:m3773.
Bikfalvi A, da Costa CA, Avril T, Barnier J-V, Bauchet L, Brisson L, et al. Challenges in glioblastoma research: focus on the tumor microenvironment. Trends Cancer. 2023;9:9–27.
Boccellato C, Rehm M. Glioblastoma, from disease understanding towards optimal cell-based in vitro models. Cell Oncol (Dordr). 2022;45:527–41.
Rong L, Li N, Zhang Z. Emerging therapies for glioblastoma: current state and future directions. J Exp Clin Cancer Res. 2022;41:142.
Blay J-Y, Kang Y-K, Nishida T, Von Mehren M. Gastrointestinal stromal tumours. Nat Rev Dis Prim. 2021;7:22.
Nguyen V, Banerjee S, Sicklick JK. Moving gastrointestinal stromal tumours towards truly personalised precision therapy. Lancet Oncol. 2020;21:865–7.
Søreide K, Sandvik OM, Søreide JA, Giljaca V, Jureckova A, Bulusu VR. Global epidemiology of gastrointestinal stromal tumours (GIST): A systematic review of population-based cohort studies. Cancer Epidemiol. 2016;40:39–46.
Hsu C, Morohashi Y, Yoshimura S-I, Manrique-Hoyos N, Jung S, Lauterbach MA, et al. Regulation of exosome secretion by Rab35 and its GTPase-activating proteins TBC1D10A-C. J Cell Biol. 2010;189:223–32.
Short NJ, Rytting ME, Cortes JE. Acute myeloid leukaemia. Lancet. 2018;392:593–606.
Newell LF, Cook RJ. Advances in acute myeloid leukemia. BMJ. 2021;375:n2026.
Pollyea DA, Bixby D, Perl A, Bhatt VR, Altman JK, Appelbaum FR, et al. NCCN guidelines insights: acute myeloid leukemia, version 2. 2021. J Natl Compr Canc Netw. 2021;19:16–27.
Shimony S, Stahl M, Stone RM. Acute myeloid leukemia: 2023 update on diagnosis, risk‐stratification, and management. Am J Hematol. 2023;98:502–26.
Liao Y, Shao Z, Liu Y, Xia X, Deng Y, Yu C, et al. USP1-dependent RPS16 protein stability drives growth and metastasis of human hepatocellular carcinoma cells. J Exp Clin Cancer Res. 2021;40:201.
Li J-T, Li K-Y, Su Y, Shen Y, Lei M-Z, Zhang F, et al. Diet high in branched-chain amino acid promotes PDAC development by USP1-mediated BCAT2 stabilization. Natl Sci Rev. 2022;9:nwab212.
Cui S-Z, Lei Z-Y, Guan T-P, Fan L-L, Li Y-Q, Geng X-Y, et al. Targeting USP1-dependent KDM4A protein stability as a potential prostate cancer therapy. Cancer Sci. 2020;111:1567–81.
Xu J, Li B, Song W, Cao L, Zhu C, Lin S. Tumor suppressor functions of miRNA-375 in nasopharyngeal carcinoma through inhibition of ubiquitin-specific protease 1 expression. Int J Biochem Cell Biol. 2021;141:106092.
Mussell A, Shen H, Chen Y, Mastri M, Eng KH, Bshara W, et al. USP1 regulates TAZ protein stability through ubiquitin modifications in breast cancer. Cancers (Basel). 2020;12:3090.
Tu Y, Xu L, Xu J, Bao Z, Tian W, Ye Y, et al. Loss of deubiquitylase USP2 triggers development of glioblastoma via TGF-β signaling. Oncogene. 2022;41:2597–608.
Xiao W, Wang J, Wang X, Cai S, Guo Y, Ye L, et al. Therapeutic targeting of the USP2-E2F4 axis inhibits autophagic machinery essential for zinc homeostasis in cancer progression. Autophagy. 2022;18:2615–35.
Zhu L, Chen Z, Guo T, Chen W, Zhao L, Guo L, et al. USP2 inhibits lung cancer pathogenesis by reducing ARID2 protein degradation via ubiquitination. Biomed Res Int. 2022;2022:1525216.
Shi K, Zhang JZ, Yang L, Li N-N, Yue Y, Du X-H, et al. Protein deubiquitylase USP3 stabilizes Aurora A to promote proliferation and metastasis of esophageal squamous cell carcinoma. BMC Cancer. 2021;21:1196.
Liang R-P, Zhang X-X, Zhao J, Zhu R-T, Wang W-J, Lu Q-W, et al. Ubiquitin-specific protease 3 facilitates cell proliferation by deubiquitinating pyruvate kinase L/R in gallbladder cancer. Lab Investig. 2022;102:1367–76.
Liao X-H, Wang Y, Zhong B, Zhu S-Y. USP3 promotes proliferation of non-small cell lung cancer through regulating RBM4. Eur Rev Med Pharm Sci. 2020;24:3143–51.
Geng N, Li Y, Zhang W, Wang F, Wang X, Jin Z, et al. A PAK5-DNPEP-USP4 axis dictates breast cancer growth and metastasis. Int J Cancer. 2020;146:1139–51.
Li F, Hu Q, He T, Xu J, Yi Y, Xie S, et al. The deubiquitinase USP4 stabilizes twist1 protein to promote lung cancer cell stemness. Cancers (Basel). 2020;12:1582.
Cheung BB, Kleynhans A, Mittra R, Kim PY, Holien JK, Nagy Z, et al. A novel combination therapy targeting ubiquitin-specific protease 5 in MYCN-driven neuroblastoma. Oncogene. 2021;40:2367–81.
Pan J, Qiao Y, Chen C, Zang H, Zhang X, Qi F, et al. USP5 facilitates non-small cell lung cancer progression through stabilization of PD-L1. Cell Death Dis. 2021;12:1051.
Huang W, Liu X, Zhang Y, Deng M, Li G, Chen G, et al. USP5 promotes breast cancer cell proliferation and metastasis by stabilizing HIF2α. J Cell Physiol. 2022;237:2211–9.
Li G, Yang T, Chen Y, Bao J, Wu D, Hu X, et al. USP5 Sustains the Proliferation of Glioblastoma Through Stabilization of CyclinD1. Front Pharm. 2021;12:720307.
Henrich IC, Jain K, Young R, Quick L, Lindsay JM, Park DH, et al. Ubiquitin-specific protease 6 functions as a tumor suppressor in ewing sarcoma through immune activation. Cancer Res. 2021;81:2171–83.
Park H-B, Hwang S, Baek K-H. USP7 regulates the ERK1/2 signaling pathway through deubiquitinating Raf-1 in lung adenocarcinoma. Cell Death Dis. 2022;13:698.
Wang Z, Kang W, Li O, Qi F, Wang J, You Y, et al. Abrogation of USP7 is an alternative strategy to downregulate PD-L1 and sensitize gastric cancer cells to T cells killing. Acta Pharm Sin B. 2021;11:694–707.
Saha G, Sarkar S, Mohanta PS, Kumar K, Chakrabarti S, Basu M, et al. USP7 targets XIAP for cancer progression: Establishment of a p53-independent therapeutic avenue for glioma. Oncogene. 2022;41:5061–75.
Li L, Liu Y, Zhao Y, Feng R, Li Y, Yu X, et al. Deubiquitinase USP8 increases ID1 stability and promotes esophageal squamous cell carcinoma tumorigenesis. Cancer Lett. 2022;542:215760.
Zhao Y, Peng D, Liu Y, Zhang Q, Liu B, Deng Y, et al. Usp8 promotes tumor cell migration through activating the JNK pathway. Cell Death Dis. 2022;13:286.
Huang Y, Xia L, Tan X, Zhang J, Zeng W, Tan B, et al. Molecular mechanism of lncRNA SNHG12 in immune escape of non-small cell lung cancer through the HuR/PD-L1/USP8 axis. Cell Mol Biol Lett. 2022;27:43.
Sulkshane P, Pawar SN, Waghole R, Pawar SS, Rajput P, Uthale A, et al. Elevated USP9X drives early-to-late-stage oral tumorigenesis via stabilisation of anti-apoptotic MCL-1 protein and impacts outcome in oral cancers. Br J Cancer. 2021;125:547–60.
Chen W, Song J, Liu S, Tang B, Shen L, Zhu J, et al. USP9X promotes apoptosis in cholangiocarcinoma by modulation expression of KIF1Bβ via deubiquitinating EGLN3. J Biomed Sci. 2021;28:44.
Yang R, Chen H, Xing L, Wang B, Hu M, Ou X, et al. Hypoxia-induced circWSB1 promotes breast cancer progression through destabilizing p53 by interacting with USP10. Mol Cancer. 2022;21:88.
Zhu H, Yan F, Yuan T, Qian M, Zhou T, Dai X, et al. USP10 Promotes Proliferation of Hepatocellular Carcinoma by Deubiquitinating and Stabilizing YAP/TAZ. Cancer Res. 2020;80:2204–16.
Qiu W, Xiao Z, Yang Y, Jiang L, Song S, Qi X, et al. USP10 deubiquitinates RUNX1 and promotes proneural-to-mesenchymal transition in glioblastoma. Cell Death Dis. 2023;14:207.
Li L, Deng T, Zhang L, Wang Y, Zhou Y, Liu Y, et al. ERK-mediated cytoplasmic retention of USP11 contributes to breast cancer cell proliferation by stabilizing cytoplasmic p21. Int J Biol Sci. 2022;18:2568–82.
Li H, Roy M, Liang L, Cao W, Hu B, Li Y, et al. Deubiquitylase USP12 induces pro-survival autophagy and bortezomib resistance in multiple myeloma by stabilizing HMGB1. Oncogene. 2022;41:1298–308.
Morgan EL, Patterson MR, Barba-Moreno D, Scarth JA, Wilson A, Macdonald A. The deubiquitinase (DUB) USP13 promotes Mcl-1 stabilisation in cervical cancer. Oncogene. 2021;40:2112–29.
Xie H, Zhou J, Liu X, Xu Y, Hepperla AJ, Simon JM, et al. USP13 promotes deubiquitination of ZHX2 and tumorigenesis in kidney cancer. Proc Natl Acad Sci USA. 2022;119:e2119854119.
Zhao C, Gong J, Bai Y, Yin T, Zhou M, Pan S, et al. A self-amplifying USP14-TAZ loop drives the progression and liver metastasis of pancreatic ductal adenocarcinoma. Cell Death Differ. 2023;30:1–15.
Du X-H, Ke S-B, Liang X-Y, Gao J, Xie X-X, Qi L-Z, et al. USP14 promotes colorectal cancer progression by targeting JNK for stabilization. Cell Death Dis. 2023;14:56.
Kim M-J, Min Y, Jeong S-K, Son J, Kim JY, Lee JS, et al. USP15 negatively regulates lung cancer progression through the TRAF6-BECN1 signaling axis for autophagy induction. Cell Death Dis. 2022;13:348.
Ge J, Yu W, Li J, Ma H, Wang P, Zhou Y, et al. USP16 regulates castration-resistant prostate cancer cell proliferation by deubiquitinating and stablizing c-Myc. J Exp Clin Cancer Res. 2021;40:59.
Li X, Liu Z, Xia C, Yan K, Fang Z, Fan Y. SETD8 stabilized by USP17 epigenetically activates SREBP1 pathway to drive lipogenesis and oncogenesis of ccRCC. Cancer Lett. 2022;527:150–63.
Feng L, Wang K, Tang P, Chen S, Liu T, Lei J, et al. Deubiquitinase USP18 promotes the progression of pancreatic cancer via enhancing the Notch1-c-Myc axis. Aging (Albany NY). 2020;12:19273–92.
Rossi FA, Enriqué Steinberg JH, Calvo Roitberg EH, Joshi MU, Pandey A, Abba MC, et al. USP19 modulates cancer cell migration and invasion and acts as a novel prognostic marker in patients with early breast cancer. Oncogenesis. 2021;10:28.
Li W, Shen M, Jiang Y-Z, Zhang R, Zheng H, Wei Y, et al. Deubiquitinase USP20 promotes breast cancer metastasis by stabilizing SNAI2. Genes Dev. 2020;34:1310–5.
Zhang Q, Chen Z, Tang Q, Wang Z, Lu J, You Y, et al. USP21 promotes self-renewal and tumorigenicity of mesenchymal glioblastoma stem cells by deubiquitinating and stabilizing FOXD1. Cell Death Dis. 2022;13:712.
Tian Y, Tang B, Wang C, Wang Y, Mao J, Yao Y, et al. Operative ubiquitin-specific protease 22 deubiquitination confers a more invasive phenotype to cholangiocarcinoma. Cell Death Dis. 2021;12:678.
Zeng K, Xie W, Wang C, Wang S, Liu W, Su Y, et al. USP22 upregulates ZEB1-mediated VEGFA transcription in hepatocellular carcinoma. Cell Death Dis. 2023;14:194.
Wang S, Zhong X, Wang C, Luo H, Lin L, Sun H, et al. USP22 positively modulates ERα action via its deubiquitinase activity in breast cancer. Cell Death Differ. 2020;27:3131–45.
He H, Yi L, Zhang B, Yan B, Xiao M, Ren J, et al. USP24-GSDMB complex promotes bladder cancer proliferation via activation of the STAT3 pathway. Int J Biol Sci. 2021;17:2417–29.
Nelson JK, Thin MZ, Evan T, Howell S, Wu M, Almeida B, et al. USP25 promotes pathological HIF-1-driven metabolic reprogramming and is a potential therapeutic target in pancreatic cancer. Nat Commun. 2022;13:2070.
Tang J, Luo Y, Xiao L. USP26 promotes anaplastic thyroid cancer progression by stabilizing TAZ. Cell Death Dis. 2022;13:326.
Zou T, Wang Y, Dong L, Che T, Zhao H, Yan X, et al. Stabilization of SETD3 by deubiquitinase USP27 enhances cell proliferation and hepatocellular carcinoma progression. Cell Mol Life Sci. 2022;79:70.
Chen L, Xu Z, Li Q, Feng Q, Zheng C, Du Y, et al. USP28 facilitates pancreatic cancer progression through activation of Wnt/β-catenin pathway via stabilising FOXM1. Cell Death Dis. 2021;12:887.
Gao R, Buechel D, Kalathur RKR, Morini MF, Coto-Llerena M, Ercan C, et al. USP29-mediated HIF1α stabilization is associated with Sorafenib resistance of hepatocellular carcinoma cells by upregulating glycolysis. Oncogenesis. 2021;10:52.
Zhang X, Han Y, Liu S, Guo B, Xu S, He Y, et al. MF-094 nanodelivery inhibits oral squamous cell carcinoma by targeting USP30. Cell Mol Biol Lett. 2022;27:107.
Guo F, Zhang C, Wang F, Zhang W, Shi X, Zhu Y, et al. Deubiquitinating enzyme USP33 restrains docetaxel-induced apoptosis via stabilising the phosphatase DUSP1 in prostate cancer. Cell Death Differ. 2020;27:1938–51.
Lin C, Xia J, Gu Z, Meng Y, Gao D, Wei S. Downregulation of USP34 Inhibits the Growth and Migration of Pancreatic Cancer Cells via Inhibiting the PRR11. Onco Targets Ther. 2020;13:1471–80.
Liu C, Chen Z, Ding X, Qiao Y, Li B. Ubiquitin-specific protease 35 (USP35) mediates cisplatin-induced apoptosis by stabilizing BIRC3 in non-small cell lung cancer. Lab Investig. 2022;102:524–33.
Zhang W, Luo J, Xiao Z, Zang Y, Li X, Zhou Y, et al. USP36 facilitates esophageal squamous carcinoma progression via stabilizing YAP. Cell Death Dis. 2022;13:1021.
Wu L, Zhao N, Zhou Z, Chen J, Han S, Zhang X, et al. PLAGL2 promotes the proliferation and migration of gastric cancer cells via USP37-mediated deubiquitination of Snail1. Theranostics. 2021;11:700–14.
Zhan W, Liao X, Liu J, Tian T, Yu L, Li R. USP38 regulates the stemness and chemoresistance of human colorectal cancer via regulation of HDAC3. Oncogenesis. 2020;9:48.
Li X, Yuan J, Song C, Lei Y, Xu J, Zhang G, et al. Deubiquitinase USP39 and E3 ligase TRIM26 balance the level of ZEB1 ubiquitination and thereby determine the progression of hepatocellular carcinoma. Cell Death Differ. 2021;28:2315–32.
Yoon J-Y, Seo S-U, Woo S-M, Kwon T-K. USP41 enhances epithelial-mesenchymal transition of breast cancer cells through snail stabilization. Int J Mol Sci. 2023;24:1693.
Xue Y, Li M, Hu J, Song Y, Guo W, Miao C, et al. Cav2.2-NFAT2-USP43 axis promotes invadopodia formation and breast cancer metastasis through cortactin stabilization. Cell Death Dis. 2022;13:812.
Chen Y, Zhao Y, Yang X, Ren X, Huang S, Gong S, et al. USP44 regulates irradiation-induced DNA double-strand break repair and suppresses tumorigenesis in nasopharyngeal carcinoma. Nat Commun. 2022;13:501.
Liu L, Chen C, Liu P, Li J, Pang Z, Zhu J, et al. MYH10 combines with MYH9 to recruit USP45 by deubiquitinating snail and promotes serous ovarian cancer carcinogenesis, progression, and cisplatin resistance. Adv Sci. 2023;10:e2203423.
Tian M, Zhu R, Ding F, Liu Z. Ubiquitin-specific peptidase 46 promotes tumor metastasis through stabilizing ENO1 in human esophageal squamous cell carcinoma. Exp Cell Res. 2020;395:112188.
Pan B, Yang Y, Li J, Wang Y, Fang C, Yu F-X, et al. USP47-mediated deubiquitination and stabilization of YAP contributes to the progression of colorectal cancer. Protein Cell. 2020;11:138–43.
Du L, Li Y, Kang M, Feng M, Ren Y, Dai H, et al. USP48 Is upregulated by Mettl14 to attenuate hepatocellular carcinoma via regulating SIRT6 Stabilization. Cancer Res. 2021;81:3822–34.
Liu Z, Li J, Ding Y, Ma M, Chen J, Lei W, et al. USP49 mediates tumor progression and poor prognosis through a YAP1-dependent feedback loop in gastric cancer. Oncogene. 2022;41:2555–70.
Li J, Xiao X, Wang H, Wang W, Ou Y, Wang Z, et al. CDK4/6-USP51 axis regulates lung adenocarcinoma metastasis through ZEB1. Cancer Gene Ther. 2022;29:1181–92.
Yao Y, Ma W, Guo Y, Liu Y, Xia P, Wu X, et al. USP53 plays an antitumor role in hepatocellular carcinoma through deubiquitination of cytochrome c. Oncogenesis. 2022;11:31.
Zhang C, Ma X, Wei G, Zhu X, Hu P, Chen X, et al. Centrosomal protein 120 promotes centrosome amplification and gastric cancer progression via USP54-mediated deubiquitination of PLK4. iScience. 2023;26:105745.
Wang L, Lin Y, Zhou X, Chen Y, Li X, Luo W, et al. CYLD deficiency enhances metabolic reprogramming and tumor progression in nasopharyngeal carcinoma via PFKFB3. Cancer Lett. 2022;532:215586.
Haq S, Sarodaya N, Karapurkar JK, Suresh B, Jo JK, Singh V, et al. CYLD destabilizes NoxO1 protein by promoting ubiquitination and regulates prostate cancer progression. Cancer Lett. 2022;525:146–57.
Funding
This research was funded by the National Natural Science Foundation of China (Nos. 81871231, 81471546, 81001346, and 81802759), Taishan Scholars Young Experts Program, Youth Innovation and Technology Plan of Shandong Province (2019KJK016), Science and Technology Project of Qingdao (No. 18-6-1-91-nsh), the China Postdoctoral Science Foundation Funded Project (No. 2019M652312), and the Qingdao Postdoctoral Application Research Project.
Author information
Authors and Affiliations
Contributions
SL wrote the manuscript, YS, KXW, and GXL drew the figures, XLD, FHY, GC, and CC reviewed relevant materials and organized the tables, HHZ, MJW, YL, and TZ collected relevant references and participated in the discussion. BL and CYL designed this review and revised the manuscript. All authors contributed to this manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, S., Song, Y., Wang, K. et al. USP32 deubiquitinase: cellular functions, regulatory mechanisms, and potential as a cancer therapy target. Cell Death Discov. 9, 338 (2023). https://doi.org/10.1038/s41420-023-01629-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-023-01629-1
- Springer Nature Limited