Abstract
In contrast to the “Warburg effect” or aerobic glycolysis earlier generalized as a phenomenon in cancer cells, more and more recent evidence indicates that functional mitochondria are pivotal for ensuring the energy supply of cancer cells. Here, we report that cancer cells with reduced autophagy-related protein 12 (ATG12) expression undergo an oncotic cell death, a phenotype distinct from that seen in ATG5-deficient cells described before. In addition, using untargeted metabolomics with ATG12-deficient cancer cells, we observed a global reduction in cellular bioenergetic pathways, such as β-oxidation (FAO), glycolysis, and tricarboxylic acid cycle activity, as well as a decrease in mitochondrial respiration as monitored with Seahorse experiments. Analyzing the biogenesis of mitochondria by quantifying mitochondrial DNA content together with several mitochondrion-localizing proteins indicated a reduction in mitochondrial biogenesis in ATG12-deficient cancer cells, which also showed reduced hexokinase II expression and the upregulation of uncoupling protein 2. ATG12, which we observed in normal cells to be partially localized in mitochondria, is upregulated in multiple types of solid tumors in comparison with normal tissues. Strikingly, mouse xenografts of ATG12-deficient cells grew significantly slower as compared with vector control cells. Collectively, our work has revealed a previously unreported role for ATG12 in regulating mitochondrial biogenesis and cellular energy metabolism and points up an essential role for mitochondria as a failsafe mechanism in the growth and survival of glycolysis-dependent cancer cells. Inducing oncosis by imposing an ATG12 deficiency in solid tumors might represent an anticancer therapy preferable to conventional caspase-dependent apoptosis that often leads to undesirable consequences, such as incomplete cancer cell killing and a silencing of the host immune system.
Similar content being viewed by others
Introduction
Despite contradictory interpretations of autophagy as being either or both tumor-suppressive and oncogenic, the significance of this conserved cellular catabolic process in cancer cannot be ignored [1]. Through the activities of a set of autophagy-related proteins (ATGs), cellular elements, such as damaged organelles or long-lived proteins, can be sequestered in a typical double-membrane structure called the autophagosome, and finally, after fusion with lysosomes, degraded by lysosomal enzymes [2]. Interplay between autophagy and apoptosis, two essential cell fate determinants, define dual functions for several of the ATGs. For instance, ATG5 is cleaved by calpain and translocated to the mitochondria to induce apoptosis [3]. ATG5 is also able to translocate to the nucleus where it interacts with survivin and induces mitotic catastrophe after treatment of cells with DNA-damaging drugs [4].
Interestingly, the binding partner of ATG5, ATG12, has been reported to induce apoptosis by inactivating prosurvival BCL-2 family members, including BCL-2 and MCL-1, by directly binding to these proteins [5]. The downregulation of ATG12 inhibits BAX activation and cytochrome c release, thus protecting cells from apoptosis [5]. The mitochondrial proapoptotic effect of ATG12 has been linked to its relative instability. Free ATG12, which is short-lived, is rapidly degraded via proteasomes after its ubiquitination. Thus, the accumulation of free ATG12 is a factor in proteasome inhibitor-mediated cell death [6]. Besides BCL-2 and MCL-1, ATG12 has been reported to bind to ATG3; cells lacking an ATG12–ATG3 complex exhibit an increased mitochondrial mass and inhibited mitochondrial apoptosis [7].
A deficiency in autophagy has been linked to an impaired recycling of the tricarboxylic acid (TCA) cycle intermediates under starving conditions with Atg7−/− Kras-driven lung tumor cells [8]. In the present study, we newly report that ATG12-deficient cells undergo oncosis under nutrient-rich conditions owing to an insufficient level of cellular ATP. In contrast to apoptosis that is characterized by cell shrinkage, cells undergoing oncosis show cell and organelle swelling and cytoplasmic blebs owing to an impairment of ion pumps as a consequence of energy deficiency [9]. The downregulation of ATG12 was observed to correlate with a reduction in β-oxidation (FAO) of lipids, glycolysis, the pentose phosphate pathway (PPP), and TCA cycle activity as well as mitochondrial respiration, suggesting an overall decrease in metabolic activity of these cells. Furthermore, such cells contained reduced amounts of mitochondrial DNA (mtDNA), expressed diminished levels of mitochondrion-localizing proteins, and showed decreased expression of peroxisome proliferator-activated receptor gamma coactivator 1 alpha (PGC1-α), the master regulator of mitochondrial biogenesis. Subcellular localization experiments showed a mitochondrial localization for ATG12. Furthermore, ATG12 is significantly upregulated in diverse solid tumors as compared with their normal counterparts. Thus, our findings suggest a novel role for ATG12 in regulating mitochondrial biogenesis and cancer metabolism. In consequence, ATG12 may represent a new potential oncogene target for anticancer therapy.
Results
An ATG12 deficiency leads to decreased cell proliferation, and finally to oncosis, owing to reduced ATP availability in cancer cells
Since it has been reported that ATG12 binds to and inactivates antiapoptotic proteins, such as BCL-2 and MCL-1, thereby inducing apoptosis [5], one might expect a survival advantage for cells with reduced ATG12 expression. To test this hypothesis, we generated H1299 lung cancer and MDA-MB-231 breast cancer cells having deficient ATG12 expression using lentivirus-based gene transfer of the shATG12 construct as previously reported [10]. Successful knockdown of ATG12 was confirmed by western blotting (Fig. S1a). These cells showed a reduction in basal and starvation-induced autophagy as evidenced by a decrease in LC3 lipidation (Fig. S1a, b). However, surprisingly, ATG12-deficient cells formed significantly fewer colonies (Fig. 1a), thus contradicting findings in a previous publication [5].
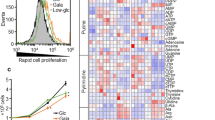
a A colony formation assay was performed and quantified in MDA-MB-231 and H1299 cells as described in the “Materials and methods” section. Data shown as bar diagrams are means ± SDs from three independent experiments. Successful knockdown of ATG12 in both cell lines is shown in Fig. S1a. b Cell proliferation was investigated by Ki-67 staining in vector control and shATG12 infected cancer cell lines. Ki-67 was stained red in the nucleus. The percentage of Ki-67-positive cells was calculated by counting more than 150 cells under the microscope. Scale bar: 50 µm. Data shown as bar diagrams are means ± SDs from three independent experiments. c Cell growth was followed by counting cell numbers daily beginning at the second until the sixth day after infection and the results are presented as a percentage of the vector control cells. d Cell viability was measured by the trypan blue exclusion assay daily beginning at the second until the sixth day after infection and the results are presented as the percentage of the vector control cells. e Morphological changes of vector control and shATG12 cells were observed 6 and 8 days after virus infection. Scale bar: 50 µm. Cells with blister formation were further quantified as percentage of the total cell number of three images (more than 230 cells). f Vector control and shATG12 cells were assessed for AV and PI staining via FACS. PI-positive cells after 6 days of virus infection are presented as bar diagrams. Data shown are means ± SDs from three independent experiments. g Cell morphology as well as AV and PI staining of MDA-MB-231 cells treated with 20 µM STS overnight are presented. The morphology of ATG12-deficient cells was compared with that of cells undergoing oncosis induced by 0.1% SA treatment for 24 h. Scale bars: 50 µm. h Vector control and shATG12 cells were stained with 50 μM JC-1 at 37 °C for 30 min to detect Δψm by FACS. Cells with polarized mitochondria (FITC and PE double positive) are presented as a bar diagram. Data shown are means ± SDs from three independent experiments. i The ATP level was measured and normalized to the protein amounts of cells as described in the “Materials and methods” section. The values for vector control cells were normalized to 100%. Data shown are means ± SDs from three independent experiments. j Cells were stained with 5 μM of Fluo-4 for 15 min to measure cytosolic calcium by FACS. After measuring the untreated samples, 1 μM of ionomycin was added directly to each sample to allow calcium release from ER, and then immediately assessed by FACS.
After 3 days of infection with shATG12 lentivirus, a reduction in numbers of Ki-67-positive cells was observed (Fig. 1b). A time course follow-up of the proliferation and viability of these cells indicated that the reduced proliferation of ATG12-deficient cells occurred prior to a cell death that was obvious from day 6 of infection with shATG12 (Fig. 1c, d). After 6–9 days of in vitro culture of ATG12-deficient cells, we observed a high level of cell death in both MDA-MB-231 and H1299 cells with the latter exhibiting a faster and more massive effect (Fig. 1e). Interestingly, no obvious cell death was observed in primary human fibroblasts after shATG12 lentivirus infection (Fig. S2). Enlarged cell size, cytoplasmic vacuolization (Fig. 1e, red arrows), and blister formation (Fig. 1e, black arrows) were visible in the ATG12-deficient H1299 cells by 6 days after shRNA lentivirus infection. In MDA-MB-231 cells, blister formation began to appear only after 8 days of infection (Fig. 1e, black arrows). It should be noted that cells deficient in ATG5 and/or autophagy exhibit increased cell proliferation as demonstrated in multiple cell types, including cancer cells [11,12,13]. Interestingly, our data seem to suggest that ATG12 promotes proliferation rather than inducing apoptosis as previously reported [5].
However, the number of Annexin V (AV)-positive cells (<5% of MDA-MB-231 shATG12 cells and <12% of H1299 shATG12 cells, Fig. 1f) did not reflect the massive cell death shown in Fig. 1e, although ATG12-deficient cells did show significantly higher amounts of propidium iodide (PI)-positive cells (Fig. 1f). The fact that only a small number of cells were AV positive and the morphology of ATG12-deficient cells was distinct from that of cells undergoing apoptosis after treatment with 20 µM staurosporine (STS) overnight (Fig. 1g, left) raised the question whether these cells were in fact dying through apoptosis. Cell swelling, vacuolization, and blister formation of the ATG12-deficient cells (Fig. 1e) pointed to a different cell death mechanism, referred to as oncosis, in which cells enlarge due to an insufficient energy supply for maintaining the ion pumps in the plasma membrane. This then leads to an osmotic imbalance and swelling [9]. Morphological comparisons between ATG12-deficient cells and cells treated with 0.1% sodium azide (SA) to induce oncosis [14], revealed a close morphological similarity (Fig. 1g, right, blister formation indicated with black arrows). A typical oncotic morphology of the ATG12-deficient cells was further suggested by electron microscopic examination showing mitochondrial dilatation (Fig. S3a, black arrowheads) and cristae disintegration (Fig. S3a, black arrows). Applying the pan-caspase inhibitor, Q-VD-OPH, or the receptor-interacting serine/threonine-protein kinase 1 inhibitor, necrostatin-1, did not rescue the cell death caused by ATG12 deficiency (Fig. S3b). These data strongly suggested that cells deficient in ATG12 underwent oncotic cell death.
It has been reported that cells undergoing classical oncosis show reduced mitochondrial membrane potential (Δψm) and decreased ATP production [9, 15]. To ascertain if this applies also to ATG12-deficient cells, we stained the cells with the mitochondrial potential-dependent dye tetraethylbenzimidazolylcarbocyanine iodide (JC-1), which forms red aggregates in the energized mitochondria that can be detected in the PE channel [16, 17]. FACS analysis showed that cells with reduced ATG12 expression exhibited significantly decreased mitochondrial polarization relative to those with normal levels of ATG12 (Fig. 1h). In addition, the ATP level was reduced in ATG12-deficient cells (Fig. 1i). Moreover, levels of cytosolic calcium, which had been shown to be increased in oncotic cells [15], were elevated in ATG12-deficient cells (Fig. 1j). Ionomycin treatment, which induces calcium release from the endoplasmic reticulum (ER) [18], did not further increase the cytosolic calcium concentration in ATG12-deficient cells, though an increase in calcium was observed in vector control cells after treatment with ionomycin (Fig. 1j). Collectively, these data strongly support our hypothesis that ATG12-deficient cells die mainly through oncosis.
ATG12-deficient cells show reduced bioenergetic activity as compared with control cells
The insufficient energy supply in ATG12-deficient cells suggested a possible alteration in cellular metabolism. Hence, global metabolic profiles of the cells were characterized using ultrahigh performance liquid chromatography coupled with high-resolution mass spectrometry (UHPLC-HRMS). Principal component analysis (PCA) of the global untargeted metabolic profiles measured by reversed-phase (RP) UHPLC-HRMS indicated clear differences between the ATG12-deficient and vector control cells (Fig. S4). Similar PCA plots were obtained by hydrophilic interaction liquid chromatography (HILIC; data not shown). After isolation of the discriminative features and a review of potential identities (based on accurate mass, isotope similarity, and where available, adduct patterns, fragmentation patterns and retention time similarities compared against authentic standards), FAO, glycolysis, PPP, and TCA cycle activity were all found to be significantly altered in ATG12-deficient cells as compared with vector control cells (Fig. 2a–c and Table S1).
a Acylcarnitine profiles measured by UHPLC-HRMS. The acylcarnitine levels detected were normalized to the total level of carnitines and are presented as bar diagrams. Q: false discovery rate (FDR). Corresponding data are also presented in Table S1. b A simplified illustration of FAO. Cytoplasmic FAs need to be esterified with carnitines at the outer mitochondrial membrane in order to be transported into the mitochondria where they undergo transesterification to CoA esters for FAO. Carnitine esters are in equilibrium with the corresponding CoA esters, thus are used as markers for intramitochondrial β-oxidation. The four enzymatic reactions of FAO are chain length specific and produce the reducing equivalents, FADH2 and NADH for the respiratory chain, and acetyl-CoA, which enters into the TCA cycle. C carnitine, AC acylcarnitine, CPT1 carnitine O-palmitoyltransferase 1, CPT2 carnitine O-palmitoyltransferase 2, CACT carnitine acylcarnitine translocase, VLCAD very long-chain acyl-CoA dehydrogenase, LCAC long-chain acylcarnitine, MCAC medium chain acylcarnitine, SCAC short chain acylcarnitine, MOM mitochondrial outer membrane, MIM mitochondrial inner membrane. c Altered metabolites involved in glycolysis, PPP, and TCA cycle (indicated in the corresponding colored boxes) detected by untargeted metabolomics are presented. Q: FDR. The results suggest a decreased activity of these pathways in ATG12-deficient cells. Corresponding data are also shown in Table S1. d MDA-MB-231 and H1299 cells were applied to Seahorse analysis in order to detect OCR in the vector control and shATG12 cells as described in the “Materials and methods” section. The calculated basal, ATP-linked and maximal OCR are presented (see also Fig. S5a). Data shown are means ± SDs of five replicates from one out of three independent experiments. Representative OCR profiles are shown in Fig. S5a. Two-way ANOVA with Bonferroni posttests was performed for statistical analysis. MDA-MB-231 cells: pATG12 level < 0.0001, pOCR type = 0.0014. H1299 cells: pATG12 level < 0.0001, pOCR type = 0.0005. The individual compared p values are shown in the graph. e MDA-MB-231 and H1299 cells were subjected to Seahorse analysis in order to detect ECAR in the vector control and shATG12 cells as described in the “Materials and methods” section. The calculated basal, maximal ECAR, and glycolytic reserve are presented (see also Fig. S5b). Data shown are means ± SDs of five replicates from one out of three independent experiments. Representative ECAR profiles are shown in Fig. S5b. Two-way ANOVA with Bonferroni posttests was performed for statistical analysis. MDA-MB-231 cells: pATG12 level = 0.0095, p ECAR type < 0.0001. H1299 cells: pATG12 level = 0.0125, pECAR type < 0.0001. The individual compared p values are shown in the graph. f Lactate production in control and shATG12 cells was measured as described in the “Materials and methods” section. The calculated lactate levels per µg protein were normalized to those of the vector controls. Data shown are means ± SDs from three independent experiments. g Selected proteins were assessed by western blotting in total cell lysates from the vector control and shATG12 cells. h The OCRs and ECARs of H1299 and MDA-MB-231 cells were extracted from one representative out of three independent Seahorse experiments to characterize the metabotypes of these cells. The empty squares represent the utilization of mitochondrial respiration and glycolysis under normal/starting conditions. The filled squares represent the utilization of mitochondrial respiration and glycolysis when stressed. The metabolic potential is a parameter of the cells’ ability to meet energy demand under stressed conditions via mitochondrial respiration and glycolysis.
Interestingly, we observed in ATG12-deficient cells an increase in long-chain acylcarnitines, substrates for FAO, together with a decrease in medium and short chain acylcarnitines as well as acetylcarnitine, the final product of FAO (Fig. 2a and Table S1). This profile is characteristic of an intramitochondrial block of long-chain FAO, leading to the assumption that the very long-chain acyl-CoA dehydrogenase, which processes long-chain acylcarnitines in the first step of FAO, was impaired in ATG12-deficient cells, thus resulting in a decreased FAO (Fig. 2a, b). Similarly, decreased levels of glycolysis, PPP, and TCA cycle activity were apparent in our global untargeted metabolic profiling measurements (Fig. 2c), indicating a reduced activity of all main energy producing pathways.
To inspect mitochondrial respiration and glycolysis in more detail, we performed extracellular flux analysis using Seahorse XF24 with vector control and shATG12 cells in both MDA-MB-231 and H1299 cells. As shown in Figs. 2d and S5a, there were significant reductions in basal, ATP-linked and maximal oxygen consumption rates (OCR) in ATG12-deficient MDA-MB-231 and H1299 cells. Regarding the extracellular acidification rate (ECAR), we observed a significant reduction in the glycolytic reserve in ATG12-deficient MDA-MB-231 and H1299 cells with no difference in basal or maximal ECAR (Figs. 2e and S5b). These data correspond to our untargeted metabolomics data obtained from H1299 cells, indicating again a downregulation of energy generation pathways, i.e., glycolysis, TCA cycle activity, and FAO.
Given the essential role of glycolysis in cancer cell metabolism and survival [19, 20], we decided to evaluate the level of glycolysis in these cells by measuring lactate production as previously described [21]. In line with our global metabolomics data (Fig. 2c and Table S1), we found reduced lactate levels after ATG12 knockdown in both cell lines (Fig. 2f), confirming a decrease in glycolysis. Taken together, the overall reduced energy metabolism in ATG12-deficient cells is likely the cause for the imbalances in ion distribution and in osmotic pressure and, finally, for the oncotic cell death observed in these cells (Fig. 1e, f).
The overall reduction in metabolites involved in global energetic pathways ranging from glycolysis, PPP to the TCA cycle indicated a likely decrease in glucose utilization. Western blot analysis showed that hexokinase II (HK II), which phosphorylates glucose to glucose-6-phosphate [22], was reduced in ATG12-deficient cells (Fig. 2g). This finding may also explain the elevated glucose in ATG12-deficient cells (Table S1). Interestingly, it is known that HK II, but not HK I, is often upregulated in tumor cells to ensure the glycolytic metabotype, thus promoting the growth and survival of cancer cells [23]. Therefore, the reduced HK II expression in ATG12-deficient cells could at least partially account for the overall decreased bioenergetics, leading to oncosis.
Dissecting the Seahorse data shown above (Fig. 2d, e as well as Fig. S5a, b), it seemed that an ATG12 deficiency impaired mitochondrial respiration more significantly than it inhibited glycolysis. A comparison of the metabolic phenotypes between the two investigated cancer cell lines revealed a clear difference in the preference of energy generation. H1299 cells were more dependent on mitochondrial respiration to obtain energy under normal conditions (Fig. 2h, dark green empty square) and exhibited a higher respiratory and glycolytic reserve once stressed, whereas MDA-MB-231 cells were less inclined towards mitochondrial respiration and had a lower bioenergetics reserve (Fig. 2h, light green empty square). Combining the results that ATG12-deficient H1299 cells showed an earlier and more severe oncosis than the ATG12-deficient MDA-MB-231 cells (Fig. 1e), these data strongly support our hypothesis that ATG12 plays an important role in mitochondrial respiration.
ATG12-deficient cells exhibit decreased mitochondrial biogenesis and ATG12 is upregulated upon mitochondrial damage
Given that an energy deficiency is the primary cause for oncosis [15], we asked whether ATG12-deficient cells are able to obtain energy from mitochondrial oxidative phosphorylation (OXPHOS), since cancer cells can switch their glycolysis-dependent metabotype to OXPHOS, if glycolysis is impaired [21]. Recalling our untargeted metabolomics data as well as our Seahorse data (Table S1, Figs. 2d, e and S5a, b), it seemed likely that these cells have an impaired mitochondrial metabolism. To investigate this possibility in detail, we examined the expression of complex IV/cytochrome C oxidase subunit II (CoxII), one of the components of the electron transport chain [24]. As hypothesized, we found a reduction in the expression of CoxII (Fig. 3a). In addition, the voltage-dependent anion-selective channel protein (VDAC) [24] was also seen to be reduced in ATG12-deficient cells (Fig. 3a). It seemed that there was a general reduction in the expression of mitochondrion-localizing proteins confirming our hypothesis that the biogenesis of mitochondria might be impaired in ATG12-deficient cells.
a Total cell lysates of vector control cells and ATG12-deficient cells were applied to western blot analysis. Multiple mitochondrion-localizing proteins were found to be reduced in ATG12-deficient cells. b mtDNA amounts in vector control cells and cells deficient for ATG12 were measured with quantitative real-time PCR using primers against the major and the minor arch of mtDNA and against the mitochondrion-encoded gene, 12S. The genomic region of β2M was used for normalization. Data shown are means ± SDs from three independent experiments. c The mRNA expression of PGC1α in vector control and ATG12-deficient cells was analyzed by real-time PCR. Values shown are means ± SDs from three independent experiments. d Western blotting analysis was performed using total cell lysates from vector control and shATG12 cells. The relative expression of UCP2 was evaluated by densitometry analysis and is presented as a bar diagram on the right. Data shown are means ± SDs from three independent experiments. Related data are also presented in Fig. S6. e H1299 cells were stained with JC-1 in the absence or presence of CCCP (50 µM) at 37 °C for 30 min in order to detect the Δψm by FACS. Total cell lysates from vehicle and CCCP-treated cells were applied to western blotting analysis.
To test this hypothesis, we compared the content of mtDNA as an additional approach to compare mitochondrial biogenesis [25] between control and ATG12-deficient cells. Indeed, the results of real-time PCR analysis of DNA isolated from control and shATG12-treated cells suggested a reduction in the amount of mtDNA in ATG12-deficient cells. Primers specific for the major or minor arches of mtDNA and primers for the mitochondrion-encoded gene, 12S, were employed. The genomic region of the β2M gene was chosen as reference [26] (Fig. 3b). In addition, we also examined the expression of the master regulator of mitochondrial biogenesis, PGC1-α [24], which was significantly reduced when ATG12 is downregulated (Fig. 3c), supporting the results shown in Fig. 3a, b that suggested the mitochondrial biogenesis is reduced in these cells.
Apart from the mitochondrial proteins decreased upon the downregulation of ATG12 (Fig. 3a) presumably owing to reduced mitochondrial biogenesis, we observed, to our surprise, that uncoupling protein 2 (UCP2) was significantly upregulated in ATG12-deficient cells (Fig. 3d). On the other hand, UCP2 was downregulated in cells overexpressing ATG12 (Fig. S6a), which we subsequently investigated [27]. Although not fully understood, UCP2 has been shown to be implicated in broad cancer-related pathological processes, such as cancer initiation, progression, and drug resistance [28]. It is maintained at relatively low levels to avoid proton leaks, because of its ability to transport protons from the intermembrane space of the mitochondria into the mitochondrial matrix, thereby dissipating the proton gradient which largely determines Δψm. Strikingly, the upregulation of UCP2 has been shown to be associated with oncosis, accounting for the loss of Δψm [29]. Since it is an unstable protein with a short half-life [30, 31], we asked whether ATG12-deficient cells were diminished in their ability to degrade UCP2, thus exhibiting an elevated level of UCP2 (Fig. 3d).
To investigate this question, we performed a “UCP2 degradation assay” in which we applied the translation inhibitor cycloheximide (CHX) at 10 µg/ml [30] to the cells at different time points. We collected the cell lysates for western blotting to compare the degradation of UCP2 in vector control cells with that of ATG12-deficient cells. In contrast to the control cells in which UCP2 was rapidly degraded with a half-life of ~140 min, UCP2 remained undegraded in ATG12-deficient cells (Fig. S6b). Since it has been reported that the UCP2 degradation rate is regulated by mitochondrial bioenergetics, e.g., the level of ATP [30], the accumulation of UCP2 in ATG12-deficient cells could be a consequence of the reduced mitochondrial biogenesis that we have observed (Fig. 3a–c).
We next asked whether ATG12 might be increased following mitochondrial damage. Therefore, we applied the mitochondrial uncoupler, carbonyl cyanide m-chlorophenyl hydrazone (CCCP) [32] at 50 μM in H1299 cells for 30 min to induce mitochondrial depolarization as evidenced by JC-1 staining (Fig. 3e). Indeed, ATG12 was increased in cells treated with CCCP together with the other mitochondrion-localizing proteins we detected (Fig. 3e). These data suggest that ATG12 could be required for the biogenesis of mitochondria, e.g., under conditions of mitochondrial damage.
ATG12 is localized partially in mitochondria and is significantly upregulated in several solid tumors in vivo
The presence of a BAX-interacting domain in ATG12 [5] indicated a possible mitochondrial localization of ATG12. To test this hypothesis, we co-stained ATG12 with Mitotracker Orange (MTO), a mitochondrion-specific dye in MDA-MB-231 and H1299 cells and calculated the Pearson’s correlation coefficient (R) using Imaris software as described in the “Materials and methods” section. We found that ATG12 is co-localized with MTO in these cell lines as evidenced by the R-values (RMDA-MB-231 = 0.661 ± 0.014 and RH1299 = 0.848 ± 0.011) (Fig. 4a). The R-values obtained with these cells by co-staining of LC3 together with MTO are presented as negative controls (Figs. 4a and S7a). In contrast, the R-values of ATG12 with lysosome-associated membrane glycoprotein 1 (LAMP1), a lysosomal marker [33] or endoplasmic reticulum aminopeptidase 1 (ERAP1), an ER-localizing protein [34], were relatively low (Fig. S7b), suggesting that ATG12 is probably not localized in the lysosomes or ER.
a ATG12 was co-stained with MTO in MDA-MB-231 and H1299 cells. Scale bars: 20 µm. Images (n ≥ 3) were analyzed with Imaris software and the Pearson’s correlation coefficients plotted. Values are means ± SEMs. Related data are also presented in Fig. S7. b Mitochondrial fractions (M) from MDA-MB-231 and H1299 cells (16 × 106 cells) were separated from the cytosolic fractions (C) and were applied to western blotting. Conjugated ATG12 in cytosol and mitochondria is indicated with white and black stars, respectively. c Subcellular fractionation was performed, followed by western blotting with vector control and shATG12 H1299 cells (20 × 106 cells). C cytosol, M mitochondria. The mitochondrial ATG12 level is reduced in shATG12 cells (black stars) and the level of ATG5-ATG12 conjugate diminished, together with the appearance of free ATG5 in the cytosolic fraction (white rectangle). d ATG12 was co-stained with SSBP1 in freshly isolated human dermal fibroblasts and MDA-MB-231 cells. Scale bar: 20 µm. e ATG12 was co-stained with SSBP1 in tissue sections from paired colon and non-small cell lung cancer patients. Scale bars: 20 µm. f ATG12 was stained in a multicancer TMA. ATG12-positive cells from the normal and the corresponding tumor tissues were counted. Values are means ± SEMs. g H1299 cells were infected with either vector control virus or virus containing shATG12 plasmid. Three days post infection, cells were injected subcutaneously into NSG mice (four mice per group/cell type) and the tumor formation was assessed three times per week. The tumor sizes (mm3) of the vector control and the shATG12 cells were plotted. Values are means ± SDs from four mice per group. Two-way ANOVA with Bonferroni posttests was used to evaluate the statistical significance between the vector control cells and the cells deficient in ATG12 (pgenotype). The p values at different time points are also presented in the graph.
Following cell fractionation, we separated mitochondria out of the cytosolic fraction and in western blot could detect ATG12 in a band at ~56 kDa in the cytosol (Fig. 4b, white stars) as well as in the isolated mitochondrial fraction in a band together with other known mitochondrion-localizing proteins, such as VDAC and CoxII (Fig. 4b, black stars). The level of ATG12 in these two compartments seemed to vary between cell types: it was more dominant in the mitochondrial fraction of H1299 cells, whereas MDA-MB-231 cells showed more cytosolic ATG12 (Fig. 4b). ATG12 with the molecular weight of 15 kDa has been shown to be highly unstable and is rapidly degraded via the proteasome pathway [6]. The band at ~56 kDa detected in the mitochondrial fraction (Fig. 4b, black stars) strongly indicated the conjugated form of ATG12. Both ATG5 and ATG3 have been shown to form complexes with ATG12 with molecular weights of 32 and 36 kDa, respectively [7]. In contrast, an antibody against ATG5 detected the ATG5–ATG12 conjugate as well as free ATG5 in the cytosolic fraction (Fig. 4b, white rectangles). The same observation was made with an anti-ATG3 antibody (Fig. 4b, white rectangles). After a knockdown of ATG12, we observed a reduction in ATG12 protein in the mitochondrial fraction (Fig. 4c, black stars) together with other mitochondrial marker proteins, also accompanied by a decrease in the amount of ATG5–ATG12 conjugate and an increase in the ATG5 protein free in the cytosol [3] (Fig. 4c, white rectangle). The reduced expression of the mitochondrial CoxII caused by ATG12 downregulation (Fig. 4c) again suggested a potential role of ATG12 in mitochondrial biogenesis. Taken together, these data suggested a novel mitochondrial ATG12 complex, which performs mitochondrion-related functions, such as mitochondrial biogenesis and respiration, thus accounting for the oncotic phenotype observed in ATG12-deficient cells.
The mitochondrial localization of ATG12 was not only detected in cancer cell lines, but also in freshly isolated human dermal fibroblasts (Fig. 4d) as well as in the normal colon and lung tissues when we co-stained these cells/tissues with the mitochondrial marker protein, single stranded DNA-binding protein 1 (SSBP1) [35] (Fig. 4e). Interestingly, we observed a lower level of ATG12 in normal primary fibroblasts as well as in the normal tissues as compared with tumor cell lines or tumor tissues, which exhibited an upregulation of ATG12 (Fig. 4d, e), indicating a potential role of ATG12 in tumorigenesis.
Based on our in vitro data (Fig. 1), it seemed that ATG12, in contrast to its conjugation partner, ATG5, appears to behave as an oncogene product rather than a tumor suppressor [11]. Corresponding to the data shown in Fig. 4e, we observed a significant upregulation of ATG12 in tumor cells of several cancer types in our tumor tissue microarray (TMA), including colon cancer, lung cancer, prostate cancer, and stomach cancer (Fig. 4f). Furthermore, we have observed an overall significantly reduced tumor forming capacity of H1299 cells deficient in ATG12 compared with vector control cells when they were injected into NSG mice subcutaneously, despite the high variation of the tumor sizes of the vector control cells measured on day 25 after injection (Fig. 4g). Taken together, these data suggested a potential oncogenic role of ATG12 in cancer in vivo, most likely through regulating cellular metabolism to provide energy needed for cancer cell survival as shown in our in vitro data (Figs. 1–3).
Since the mitochondrion is a dynamic organelle and mitochondrial fusion and fission are essential for mitochondrial metabolism and cell survival [36], we then asked the question whether ATG12 executes its metabolism-related function by regulating mitochondrial fusion and/or fission. As could be demonstrated with subcellular fractionation experiments, vector control and ATG12-deficient cells showed no alteration in the expression of marker proteins regulating mitochondrial fusion or fission, such as mitofusin 2 (MFN2) or dynamin-like 120 kDa protein, mitochondrial (OPA1) (Fig. S8), suggesting that the role of ATG12 in regulating cellular energy metabolism is independent from mitochondrial dynamics.
Discussion
The canonical role of ATG12 is in forming a complex with ATG5, participating in autophagosome formation, thus regulating autophagy [37]. However, it seems that ATG12-deficient cells show autophagy-unrelated phenotypes that have not been reported in ATG5-deficient cells. An example is the accelerated high-fat diet-induced obesity in mice lacking Atg12 in proopiomelanocortin-expressing neurons [38].
Compared with its conjugation partner, ATG5, the role of ATG12 in cancer has not been widely investigated. In contrast to a previous publication arguing that ATG12 promotes apoptosis by inhibiting BCL-2, we observed reduced proliferation, and then, later, oncotic death in ATG12-deficient H1299 and MDA-MB-231 cancer cells. The overall reduction in cellular bioenergetics in ATG12-deficient cells evidenced by our global metabolomics study suggested an essential role of ATG12 in cancer metabolism. A decrease in oxygen consumption in ATG12-deficient cells indicated a preference of ATG12 for acting on mitochondria. The reduction in the content of mtDNA, the expression levels of mitochondrion-localizing proteins as well as the master regulator of the mitochondrial biogenesis, PGC1-α, revealed that ATG12 probably regulates mitochondrial biogenesis. The insufficient energy supply caused by impaired mitochondrial biogenesis is the likely cause for oncosis of cells with insufficient ATG12 levels. Intriguingly, UCP2 was elevated and its degradation blocked in ATG12-deficient cells, suggesting a link between ATG12, UCP2, and normal mitochondrial function. Although it has been shown that the degradation of UCP2 is regulated by mitochondrial biogenesis [30], the possibility of a direct interaction between ATG12 and UCP2 cannot be excluded.
Although it is believed that cancer cells undergo aerobic glycolysis, the so-called Warburg effect [19], our data suggest an essential and indispensable role of mitochondria in tumor cell metabolism and survival. More and more evidence indicates that the Warburg effect cannot be explained just by the existence of dysfunctional mitochondria, although defective mitochondria have been found to accumulate in tumor cells [39]. Keeping the mitochondria functioning is also essential for glycolysis-dependent cancer cells. The two cancer cell lines studied in our work have different metabolic features as shown by us and others, i.e., MDA-MB-231 is highly glycolytic [22], whereas H1299 cells have a high mitochondrial respiration level and are capable of obtaining energy from OXPHOS in addition to glycolysis [21]. Interestingly, we observed a higher level of mitochondrial ATG12 in H1299 cells together with other mitochondrion-localizing proteins than we detected in MDA-MB-231 cells. It may be that H1299 cells need more ATG12 for mitochondrial biogenesis in order to perform an OXPHOS-dependent metabolism. Intriguingly, an ATG12 deficiency led to a faster and dramatic cell death phenotype in H1299 cells as compared with the more glycolytic MDA-MB-231 cells, implying that the reduced mitochondrial biogenesis was probably a leading cause for the cell death.
Changes in mitochondrial mass could be explained by an altered balance between mitochondrial biogenesis and mitophagy, a specific type of autophagy targeting damaged mitochondria [40,41,42]. The reduced mitochondrial mass in cells lacking ATG12 was unlikely to be owed just to an increased mitophagy, since these cells are autophagy-deficient. It is known that certain oncogenes, e.g., c-myc, induce mitochondrial biogenesis to increase energy generation through activating PGC1-β [43]. The question in our system is how ATG12 can promote mitochondrial biogenesis. This question still needs to be answered, despite the fact that the PGC1α transcript was reduced in ATG12-deficient cells.
In our system, no difference in the expression of proteins regulating mitochondrial dynamics was observed in ATG12-deficient compared with control cells, although it has been reported that cells lacking an ATG12–ATG3 complex exhibit reduced mitochondrial fusion with yet unknown mechanistic insights [7]. Furthermore, oncosis as well as the reduction in mitochondrial metabolic activities observed in ATG12-deficient cells seemed to be independent from ATG3 in our system. Future studies are required to identify the unknown mitochondrial binding partner of ATG12 that allows the translocation of ATG12 to the mitochondria.
The cell death phenotype caused by an ATG12 deficiency in tumor cells showed typical morphological features associated with oncosis, a type of caspase-independent cell death (CICD). In contrast to apoptosis that “silences” the host immune system, CICD promotes antitumor immunity in addition to elimination of cancer cells [44], pointing up the logic of engaging CICD, e.g., oncosis as means in anticancer cell killing. Collectively, our work points to an autophagy-independent function of ATG12 as an oncogene capable of promoting mitochondrial biogenesis. Targeting ATG12, that is often upregulated in tumors, may have anticancer therapeutic potential.
Materials and methods
Cells and cell culture
The H1299 human lung cancer cell line [45] and the breast cancer cell line MDA-MB-231 [46] were employed for this study and cultured as described previously. Briefly, H1299 cells were grown in RPMI 1640 and MDA-MB-231 in Dulbecco’s Modified Eagle’s medium supplemented with 10% fetal bovine serum and 100 U/ml penicillin and 100 µg/ml streptomycin (all media were from Thermo Fisher Scientific).
TMA
For this single-center, retrospective, cohort study, aimed at investigating the expression of ATG12 in diverse solid tumors, archived tissue samples obtained from patients at the Department of Pathology, University of Bern, Switzerland, were prepared in a TMA. The study was approved by the Ethics Commission of the Canton of Bern, Switzerland. To guarantee patient privacy, anonymization of data was performed immediately after export from the clinical database by deleting patient identifying fields, such as name, address, identification number, and histology number.
Lentiviral gene transfer
The reported shRNA construct of ATG12 [10] and its control plasmid were purchased from Sigma-Aldrich (sequence: CCG GCG AAC CAT CCA AGG ACT CAT TCT CGA GAA TGA GTC CTT GGA TGG TTC GTT TTT; TRCN0000007391, Sigma, Mission shRNA). Virus was produced by transfecting T293 cells with the plasmid of interest together with the viral envelope vector pMD2G and the packaging vector psPAX2 (provided by Dr D. Trono, EPFL, Lausanne, Switzerland) using a calcium phosphate transfection method [47]. The supernatants were collected after 24 h, filtered through a 0.22 µM filter (Millipore AG), and stored at −80 °C before use.
Colony formation assay
The colony formation assay was performed as previously described [11]. Briefly, cells deficient for ATG12 as well as vector control cells were seeded into a 24-well plate (1000 cells/well), grown for 1 week followed by crystal violet staining to visualize the colonies. A number of colonies were counted and further normalized to those of the controls.
AV and PI staining
Cells were incubated with AV in AV-binding buffer at room temperature for 15 min in the dark. After washing the cells with AV-binding buffer, cells were stained with PI and assessed for FACS analysis.
Immunofluorescence
Cells for immunofluorescence staining were first seeded on coverslips in a 24-well plate allowing adhesion. Cells were washed with PBS, fixed with 4% paraformaldehyde for 10 min, and permeabilized with 10% saponin for 5 min followed by acetone treatment at −20 °C for 10 min. After washing the slides with PBS, cells were blocked with normal goat sera at room temperature for 1 h. Cells were then incubated with primary antibody diluted in the blocking solution at 4 °C overnight. The primary antibodies were: anti-Ki-67 (1:200, Dako), anti-ATG12 (100 ng/ml; R&D Systems, MAB6807), anti-LC3 (1:100; Nanotools), anti-SSBP1 (1:100; Novus Biologicals), anti-LAMP1 (1:100; Abcam), and anti-ERAP1 (1:100; GeneTex). For those experiments where MTO was co-stained with ATG12, cells were first incubated with MTO (1:10,000; Thermo Fisher Scientific) at 37 °C for 20 min and processed as described above. After incubation with primary antibodies, cells were washed with PBS and incubated with secondary antibodies (goat anti-mouse 488, goat anti-mouse 555, or goat anti-rabbit 532; Thermo Fischer Scientific) at room temperature for 1 h protected from the light. Nuclei were stained with DAPI (Dako), slides were mounted with fluorescence mounting medium (Dako) and subsequently analyzed by confocal laser scanning microscopy (LSM 510; Carl Zeiss).
Co-localization was evaluated by calculating the Pearson’s correlation coefficient (R) using Imaris 7.5.0 (Bitplane).
Paraffin-embedded tissue sections were deparaffinized and rehydrated with graded ethanol dilutions. After antigen retrieval in sodium citrate buffer (10 mM, pH 6.0), immunofluorescence staining was performed by incubating the paraffin sections with anti-ATG12 antibody (100 ng/ml; R&D Systems) and anti-SSBP1 antibody (1:100; Novus Biologicals) or appropriate control antibodies in blocking buffer. Tissue sections were analyzed by confocal laser scanning microscopy (LSM 510).
Measurement of Δψm by FACS-based JC-1 staining
Cells either untreated or treated with 50 µM of CCCP were incubated with 2 µM of JC-1 (Thermo Fischer Scientific) at 37 °C for 45 min. After washing with PBS supplemented with 1% FCS, cells were analyzed by FACS (FACSverse, BD Biosciences).
Measurement of cellular ATP content
Cells, infected either with vector control or shATG12 virus, were seeded into a 96-well plate (10,000 cells/well) 2 days before the experiment to allow optimal attachment of cells. Cellular ATP was measured as previously described [21] with the CellTiter-Glo Luminescent Cell Viability Assay (Promega) according to the protocol provided. The relative luminescence units pro microgram protein in ATG12-deficient cells were normalized to those of vector control cells.
Untargeted metabolomics
Cell metabolites were extracted with an organic solvent for protein precipitation. Briefly, cells were washed twice with milli-Q water and the metabolites were subsequently extracted with cold methanol:acetonitrile (1:1, v/v). After centrifugation, the supernatant was evaporated until dryness and the protein pellets were used for measuring protein concentrations. One plate containing the cell culture medium and one with water were treated in the same way for use as blanks. The samples were then dissolved in the corresponding solvent and analyzed by RP chromatography and HILIC coupled to a Q-TOF mass spectrometer. RP analysis was performed as previously described [48, 49]. For HILIC chromatography, 2 µl of the extracted metabolites were separated on a ACQUITY UPLC BEH HILIC column (1.0 × 100 mm, 1.7 µm, Waters) at a flow rate of 0.2 ml/min. The mobile phase was composed of 10 mM ammonium acetate in 95:5 (v/v) acetontrile:H2O (A, pH 4.3 adjusted with acetic acid) and 1:1 (v/v) acetontrile:H2O (B, pH 4.3 adjusted with acetic acid). The following LC gradient was applied: 0 min: 100% mobile phase A, 1 min: 100% A, 8 min: 45% A, 8.5 min: 30% A, 8.6 min: 30% A (flow rate increased at 0.3 mL/min), 9.5 min: 30% A, 9.6 min: 100% A, 12.5 min: 100% A, 12.6 min: 100% A (flow rate decreased at 0.2 ml/min), 13 min: 100% A. The column temperature was set to 35 °C and the autosampler to 6 °C. For HILIC analysis, MS parameters were similar than the ones used for RP analysis. Metabolic features measured by UHPLC-HRMS were isolated with Progenesis QI (version 2.3, Nonlinear Dynamics, Newcastle, UK) and analyzed as previously described [48].
Seahorse (extracellular flux analysis)
The OCR and ECAR were measured simultaneously by Seahorse XF24 bioanalyzer (Agilent Technologies). Briefly, cells were seeded into a 24-well assay plate 1 day before the experiment. On the day of measurement, cells were washed with unbuffered culture medium and incubated at 37 °C without CO2 for 1 h, before the Mito Stress Test and Glycolysis Stress Test were performed. OCR and ECAR were measured under basal conditions followed by a sequential addition of oligomycin (1 µM), carbonyl cyanide-p-trifluoromethoxyphenylhydrazone (0.125 µM), rotenone (1 µM)/antimycin A (AA) (1 µM) and 2-deoxy-D-glucose (2DG) (50 mM) to estimate the contribution of individual parameters as described [50]. The optimal concentrations of these modulators were defined in pre-experiments. Three independent experiments with five replicates each were performed before protein concentrations in individual wells were measured. The calculated OCRs were corrected with nonmitochondrial respiration after addition of rotenone/AA and ECARs were corrected by nonglycolytic acidification after addition of 2DG. ATP-linked OCR represents the difference between basal and leak OCR after addition of oligomycin. The glycolytic reserve represents the difference between basal and maximal ECAR after addition of oligomycin.
Lactate production
Cells were seeded into a 24-well plate, replacing media after 24 h. On the day of measurement, cells were maintained either untreated or treated with oligomycin (100 ng/ml) for 6 h. Thereafter, cell culture media were collected and deproteinized by 1 M perchloric acid, followed by neutralization with 2 M potassium hydroxide until a pH of ~6.5–8 was reached. Lactate production was measured using the Lactate Assay Kit II according to the instructions provided (Sigma-Aldrich). Cells remaining in the wells were lysed and protein amounts were measured. The calculated lactate levels per µg protein were further normalized to those of the vector controls.
Quantitative real-time PCR
RNA was isolated using the ZR RNA MicroPrep Kit (Zymo Research) according to the protocol provided by the manufacturer. Real-time PCR was performed directly with the isolated RNA using the SYBR Green One Step qPCR Kit (Biotool). The following primers were used: Human PGC1α: 5′-TGA CAC AAC ACG GAC AGA AC-3′ and 5′-GCA TCA CAG GTA TAA CGG TAG G-3′ and human 18S: 5′-ATC CCT GAA AAG TTC CAG CA-3′ and 5′-CCC TCT TGG TGA GGT CAA TG-3′. The mtDNA content was quantified as previously described using primer pairs specific for the major or the minor arch of the mtDNA and normalized to the genomic region of the β2M gene [26, 51]. For the quantification of mtDNA, the following primers were used: major arch: 5′-CTG TTC CCC AAC CTT TTC CT-3′ and 5′-CCA TGA TTG TGA GGG GTA GG-3′; minor arch: 5′-CTA AAT AGC CCA CAC GTT CCC-3′ and 5′-AGA GCT CCC GTG AGT GGT TA-3′ and β2M: 5′-GCT GGG TAG CTC TAA ACA ATG TAT TCA-3′ and 5′-CCA TGT ACT AAC AAA TGT CTA AAA TGG T-3′. The qPCR reaction was performed with a CFX Connect detection system (Bio-Rad).
Immunoblotting
Cells were washed with cold PBS and lysed with lysis buffer containing 50 mM Tris pH 7.4, 150 mM NaCl, 10% glycerol, 1% TritonX 100, 2 mM EDTA, 10 mM sodium pyrophosphate, 50 mM sodium fluoride, and 200 µM sodium orthovanadate. Shortly before use, a protease inhibitor cocktail (Sigma-Aldrich) and 100 µM PMSF were freshly added to the lysis buffer. The cell pellet was incubated in lysis buffer on ice with frequent vortexing for 30 min. Thereafter, the supernatant was collected after centrifugation at 13,000 rpm for 10 min. The protein concentration was measured with the BCA Protein Assay kit (Thermo Fisher Scientific). Thirty micrograms of protein for each sample was loaded on NuPage gels (Thermo Fisher Scientific).
Subcellular fractionation was performed as previously described [52]. Briefly, ten million cells were collected and the cell pellet was washed with 1 ml of PBS containing 0.4% BSA. After removing the supernatant, 100 μl of cytosolic extraction buffer (CEB) (20 mM HEPES (pH 7.2), 250 mM sucrose, 10 mM KCl, 1.5 mM MgCl2, and 2 mM EDTA) with digitonin (0.625 mg/ml) were added to the pellet. It was resuspended and incubated for 20 min on ice to induce cell lysis. Lysed cells were centrifuged at 700 × g at 4 °C for 10 min. The supernatant was centrifuged again at 7000 × g for 30 min at 4 °C. The supernatant was then transferred into a new tube for ultracentrifugation at 21,000 × g at 4 °C for 60 min. The supernatant after ultracentrifugation contains the cytosolic fraction. The pellet obtained after the second centrifugation (7000 × g) contains the mitochondrial fraction and was washed two times with 500 μl of cold PBS followed each time by centrifugation at 7000 × g at 4 °C for 3 min. Thereafter, the cell pellet was lysed with 100 µl of CEB buffer containing 1% SDS by boiling at 95 °C for 5 min. The lysate was centrifuged at 14,000 × rpm at 4 °C for 3 min. The supernatant containing the mitochondrial fraction was collected for western blotting.
After electrophoresis, proteins were transferred onto PVDF membranes (Immobilion-P; Millipore). Membranes were incubated with a blocking buffer consisting of Tris-buffered saline (pH 7.4) with 0.1% Tween 20 (TBST) and 5% nonfat dry milk at room temperature for 1 h, and then incubated with primary antibodies at 4 °C overnight. The primary antibodies were: anti-ATG12 (100 ng/ml; R&D Systems, MAB6807), anti-Complex IV subunit II (1:1000; Thermo Fisher Scientific), anti-VDAC (1:1000; Millipore), anti-ATG5 (1:1000; Nanotools), anti-LC3 (1:1000; Nanotools), anti-HK II (1:1000, Cell Signaling), anti-HK I (1:1000, Novus), anti-UCP2 (Cell Signaling), anti-OPA1 (1:1000, Sigma), anti-ATG3 (1:1000, Abgent), anti-MFN2 (1:1000, Cell Signaling), anti-OPA1 (1:1000, Cell Signaling), and anti-GAPDH (1:10000; Chemicon International, Inc.). Membranes were then incubated with the corresponding HRP-conjugated secondary antibody (Amersham Pharmacia Biotech) in TBST with 5% nonfat dry milk at room temperature for 1 h and developed using an ECL approach (ECL-kit, Amersham).
UCP2 degradation assay
Cells were treated with CHX (10 µg/ml) for different times as indicated. Cells were counted and equal numbers of cells were collected for cell lysates, which were then applied for western blotting. UCP2 levels relative to GAPDH at different time points were normalized to that at the time point zero.
Immunohistology
Paraffin-embedded tissue sections were deparaffinized and rehydrated with graded ethanol dilutions, after which antigen retrieval was carried out. Immunohistologic staining was performed using the Dako REAL Detection System, Alkaline Phosphatase/RED Kit according to the manufacturer's instructions and as described [11]. Staining was performed with the anti-ATG12 antibody (100 ng/ml; R&D Systems, MAB6807). The fraction of ATG12-positive cells exhibiting a membranous staining pattern was compared to those of their normal counterparts.
Transmission electron microscopy
Cells were first fixed in 2.5% glutaraldehyde in 0.1 M sodium cacodylate (pH 7.4, 340 mOsm) before fixing in OsO4 in the same buffer used in the first fixation step. After dehydration in graded ethanol and embedding in EPON (Sigma), blocks were cut with diamond knife (Diatome, Switzerland) and stained with uranyl acetate and lead citrate. Sections were observed under TEM (Philips, Eindhoven, Holland).
Mouse xenografts
NOD-scid L2Rgnull (NSG) mice [53] from Charles River Laboratories were maintained under pathogen-free condition in individually ventilated cages with food and water ad libitum. After 3 days of infection, vector control and shATG12 H1299 cells were suspended in 100 µl of PBS (40 × 106 cells/ml) and injected in both flanks of NSG mice. Tumor growth was monitored using caliper at different time points and tumors were explanted 4 weeks after tumor cell infection. Animal experiments were approved by the animal committee of the Canton of Bern (KEK 76/17) and performed according to the Swiss laws for animal protection.
Statistics
Data are presented as means ± S.D.s and were analyzed by two-tailed Student's t test, if not indicated. Differences between means were considered significant at the p < 0.05 level.
References
Fulda S. Targeting autophagy for the treatment of cancer. Biol Chem. 2018;399:673–7.
Koyama-Honda I, Itakura E, Fujiwara TK, Mizushima N. Temporal analysis of recruitment of mammalian ATG proteins to the autophagosome formation site. Autophagy. 2013;9:1491–9.
Yousefi S, Perozzo R, Schmid I, Ziemiecki A, Schaffner T, Scapozza L, et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat Cell Biol. 2006;8:1124–32.
Maskey D, Yousefi S, Schmid I, Zlobec I, Perren A, Friis R, et al. ATG5 is induced by DNA-damaging agents and promotes mitotic catastrophe independent of autophagy. Nat Commun. 2013;4:2130.
Rubinstein AD, Eisenstein M, Ber Y, Bialik S, Kimchi A. The autophagy protein Atg12 associates with antiapoptotic Bcl-2 family members to promote mitochondrial apoptosis. Mol Cell. 2011;44:698–709.
Haller M, Hock AK, Giampazolias E, Oberst A, Green DR, Debnath J, et al. Ubiquitination and proteasomal degradation of ATG12 regulates its proapoptotic activity. Autophagy. 2014;10:2269–78.
Radoshevich L, Murrow L, Chen N, Fernandez E, Roy S, Fung C, et al. ATG12 conjugation to ATG3 regulates mitochondrial homeostasis and cell death. Cell. 2010;142:590–600.
Guo JY, Teng X, Laddha SV, Ma S, Van Nostrand SC, Yang Y, et al. Autophagy provides metabolic substrates to maintain energy charge and nucleotide pools in Ras-driven lung cancer cells. Genes Dev. 2016;30:1704–17.
Shubin AV, Demidyuk IV, Komissarov AA, Rafieva LM, Kostrov SV. Cytoplasmic vacuolization in cell death and survival. Oncotarget. 2016;7:55863–89.
Lin JF, Lin YC, Tsai TF, Chen HE, Chou KY, Hwang TI. Cisplatin induces protective autophagy through activation of BECN1 in human bladder cancer cells. Drug Des Devel Ther. 2017;11:1517–33.
Liu H, He Z, von Rutte T, Yousefi S, Hunger RE, Simon HU. Down-regulation of autophagy-related protein 5 (ATG5) contributes to the pathogenesis of early-stage cutaneous melanoma. Sci Transl Med. 2013;5:202ra123.
Hwang SH, Han BI, Lee M. Knockout of ATG5 leads to malignant cell transformation and resistance to Src family kinase inhibitor PP2. J Cell Physiol. 2018;233:506–15.
Rozman S, Yousefi S, Oberson K, Kaufmann T, Benarafa C, Simon HU. The generation of neutrophils in the bone marrow is controlled by autophagy. Cell Death Differ. 2015;22:445–56.
Del Nagro C, Xiao Y, Rangell L, Reichelt M, O'Brien T. Depletion of the central metabolite NAD leads to oncosis-mediated cell death. J Biol Chem. 2014;289:35182–92.
Weerasinghe P, Buja LM. Oncosis: an important non-apoptotic mode of cell death. Exp Mol Pathol. 2012;93:302–8.
St John JC, Amaral A, Bowles E, Oliveira JF, Lloyd R, Freitas M, et al. The analysis of mitochondria and mitochondrial DNA in human embryonic stem cells. Methods Mol Biol. 2006;331:347–74.
Liu H, He Z, April SL, Trefny MP, Rougier JS, Salemi S, et al. Biochemical re-programming of human dermal stem cells to neurons by increasing mitochondrial membrane potential. Cell Death Differ. 2019;26:1048–1061.
Morgan AJ, Jacob R. Ionomycin enhances Ca2+ influx by stimulating store-regulated cation entry and not by a direct action at the plasma membrane. Biochem J. 1994;300:665–72.
Warburg O, Wind F, Negelein E. The metabolism of tumors in the body. J Gen Physiol. 1927;8:519–30.
Potter M, Newport E, Morten KJ. The Warburg effect: 80 years on. Biochem Soc Trans. 2016;44:1499–505.
Hao W, Chang CP, Tsao CC, Xu J. Oligomycin-induced bioenergetic adaptation in cancer cells with heterogeneous bioenergetic organization. J Biol Chem. 2010;285:12647–54.
Gatenby RA, Gillies RJ. Why do cancers have high aerobic glycolysis? Nat Rev Cancer. 2004;4:891–9.
Lis P, Dylag M, Niedzwiecka K, Ko YH, Pedersen PL, Goffeau A, et al. The HK2 dependent "Warburg effect" and mitochondrial oxidative phosphorylation in cancer: targets for effective therapy with 3-bromopyruvate. Molecules. 2016;21:1730.
Ngamsiri P, Watcharasit P, Satayavivad J. Glycogen synthase kinase-3 (GSK3) controls deoxyglucose-induced mitochondrial biogenesis in human neuroblastoma SH-SY5Y cells. Mitochondrion. 2014;14:54–63.
Xu W, Zhu H, Gu M, Luo Q, Ding J, Yao Y, et al. DHTKD1 is essential for mitochondrial biogenesis and function maintenance. FEBS Lett. 2013;587:3587–92.
Malik AN, Shahni R, Rodriguez-de-Ledesma A, Laftah A, Cunningham P. Mitochondrial DNA as a non-invasive biomarker: accurate quantification using real time quantitative PCR without co-amplification of pseudogenes and dilution bias. Biochem Biophys Res Commun. 2011;412:1–7.
Mizushima N, Sugita H, Yoshimori T, Ohsumi Y. A new protein conjugation system in human. The counterpart of the yeast Apg12p conjugation system essential for autophagy. J Biol Chem. 1998;273:33889–92.
Valle A, Oliver J, Roca P. Role of uncoupling proteins in cancer. Cancers. 2010;2:567–91.
Mills EM, Xu D, Fergusson MM, Combs CA, Xu Y, Finkel T. Regulation of cellular oncosis by uncoupling protein 2. J Biol Chem. 2002;277:27385–92.
Azzu V, Brand MD. Degradation of an intramitochondrial protein by the cytosolic proteasome. J Cell Sci. 2010;123(Pt 4):578–85.
Rousset S, Mozo J, Dujardin G, Emre Y, Masscheleyn S, Ricquier D, et al. UCP2 is a mitochondrial transporter with an unusual very short half-life. FEBS Lett. 2007;581:479–82.
Itami N, Shiratsuki S, Shirasuna K, Kuwayama T, Iwata H. Mitochondrial biogenesis and degradation are induced by CCCP treatment of porcine oocytes. Reproduction. 2015;150:97–104.
Schroder B, Wrocklage C, Pan C, Jager R, Kosters B, Schafer H, et al. Integral and associated lysosomal membrane proteins. Traffic. 2007;8:1676–86.
Chang SC, Momburg F, Bhutani N, Goldberg AL. The ER aminopeptidase, ERAP1, trims precursors to lengths of MHC class I peptides by a "molecular ruler" mechanism. Proc Natl Acad Sci USA. 2005;102:17107–12.
Bogenhagen DF, Rousseau D, Burke S. The layered structure of human mitochondrial DNA nucleoids. J Biol Chem. 2008;283:3665–75.
Chan DC. Mitochondria: dynamic organelles in disease, aging, and development. Cell. 2006;125:1241–52.
Liu H, He Z, Simon HU. Targeting autophagy as a potential therapeutic approach for melanoma therapy. Semin Cancer Biol. 2013;23:352–60.
Malhotra R, Warne JP, Salas E, Xu AW, Debnath J. Loss of Atg12, but not Atg5, in pro-opiomelanocortin neurons exacerbates diet-induced obesity. Autophagy. 2015;11:145–54.
Galluzzi L, Morselli E, Kepp O, Vitale I, Rigoni A, Vacchelli E, et al. Mitochondrial gateways to cancer. Mol Asp Med. 2010;31:1–20.
Kelly DP, Scarpulla RC. Transcriptional regulatory circuits controlling mitochondrial biogenesis and function. Genes Dev. 2004;18:357–68.
Twig G, Hyde B, Shirihai OS. Mitochondrial fusion, fission and autophagy as a quality control axis: the bioenergetic view. Biochim Biophys Acta. 2008;1777:1092–7.
Youle RJ, Narendra DP. Mechanisms of mitophagy. Nat Rev Mol Cell Biol. 2011;12:9–14.
Zhang H, Gao P, Fukuda R, Kumar G, Krishnamachary B, Zeller KI, et al. HIF-1 inhibits mitochondrial biogenesis and cellular respiration in VHL-deficient renal cell carcinoma by repression of C-MYC activity. Cancer Cell. 2007;11:407–20.
Ichim G, Lopez J, Ahmed SU, Muthalagu N, Giampazolias E, Delgado ME, et al. Limited mitochondrial permeabilization causes DNA damage and genomic instability in the absence of cell death. Mol Cell. 2015;57:860–72.
Law BY, Mok SW, Chan WK, Xu SW, Wu AG, Yao XJ, et al. Hernandezine, a novel AMPK activator induces autophagic cell death in drug-resistant cancers. Oncotarget. 2016;7:8090–104.
Guan F, Ding Y, Zhang Y, Zhou Y, Li M, Wang C. Curcumin suppresses proliferation and migration of MDA-MB-231 breast cancer cells through autophagy-dependent Akt degradation. PLoS ONE. 2016;11:e0146553.
Okayama H, Chen C. Calcium phosphate mediated gene transfer into established cell lines. Methods Mol Biol. 1991;7:15–21.
Rindlisbacher B, Schmid C, Geiser T, Bovet C, Funke-Chambour M. Serum metabolic profiling identified a distinct metabolic signature in patients with idiopathic pulmonary fibrosis—a potential biomarker role for LysoPC. Respir Res. 2018;19:7.
Li S, Park Y, Duraisingham S, Strobel FH, Khan N, Soltow QA, et al. Predicting network activity from high throughput metabolomics. PLoS Comput Biol. 2013;9:e1003123.
Tan B, Xiao H, Li F, Zeng L, Yin Y. The profiles of mitochondrial respiration and glycolysis using extracellular flux analysis in porcine enterocyte IPEC-J2. Anim Nutr. 2015;1:239–43.
Phillips NR, Sprouse ML, Roby RK. Simultaneous quantification of mitochondrial DNA copy number and deletion ratio: a multiplex real-time PCR assay. Sci Rep. 2014;4:3887.
Conus S, Perozzo R, Reinheckel T, Peters C, Scapozza L, Yousefi S, et al. Caspase-8 is activated by cathepsin D initiating neutrophil apoptosis during the resolution of inflammation. J Exp Med. 2008;205:685–98.
Shultz LD, Lyons BL, Burzenski LM, Gott B, Chen X, Chaleff S, et al. Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2R gamma null mice engrafted with mobilized human hemopoietic stem cells. J Immunol. 2005;174:6477–89.
Acknowledgements
We thank the Clinical Metabolomics Facility, Center for Laboratory Medicine from the Bern University Hospital (Inselspital Bern) for performing UHPLC-HRMS analysis. We also thank Dr José Alberto Galvάn Hernάndez (Institute of Pathology, University of Bern) for his help in evaluating the pathological images, Jeannine Wagner (Institute of Anatomy, University of Bern) for her assistance in electron microscopy and Michaela Römmele (Department for BioMedical Research, University of Bern) for helping us with the mouse xenograft model. NG and ZF are PhD students of the Graduate School of Cellular and Biomedical Sciences of the University of Bern. Images were acquired on equipment supported by the Microscopy Imaging Centre of the University of Bern. This work was supported by grants from the Swiss National Science Foundation (grant no. 310030_166473 and 310030_184816), Swiss Cancer League (grant no. KFS-3703-08-2015), Stiftung zur Krebsbekämpfung (Zurich) and European Union’s Horizon 2020 research and innovation program (Marie Sklodowska-Curie grant no. 642295; MEL-PLEX).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Piacentini
Supplementary information
Rights and permissions
About this article
Cite this article
Liu, H., He, Z., Germič, N. et al. ATG12 deficiency leads to tumor cell oncosis owing to diminished mitochondrial biogenesis and reduced cellular bioenergetics. Cell Death Differ 27, 1965–1980 (2020). https://doi.org/10.1038/s41418-019-0476-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0476-5
- Springer Nature Limited
This article is cited by
-
Serine and one-carbon metabolisms bring new therapeutic venues in prostate cancer
Discover Oncology (2021)
-
The role of neutrophil death in chronic inflammation and cancer
Cell Death Discovery (2020)