Abstract
Inhibitor of Growth 3 (ING3) is a candidate tumor suppressor gene whose expression is lost in tumors such as hepatocellular carcinoma, head and neck squamous cell carcinoma and melanoma. In the present study, we show that ING3-depleted human cells and yeast cells deleted for its ortholog YNG2 are sensitive to DNA damage suggesting a conserved role in response to such stress. In human cells, ING3 is recruited to DNA double strand breaks and is required for ATM activation. Remarkably, in response to doxorubicin, ATM activation is dependent on ING3 but not on TIP60, whose recruitment to DNA breaks also depends on ING3. These events lead to ATM-mediated phosphorylation of NBS1 and the subsequent recruitment of RNF8, RNF168, 53BP1, and BRCA1, which are major mediators of the DNA damage response. Accordingly, upon genotoxic stress, DNA repair by non-homologous end joining (NHEJ) or homologous recombination (HR) were impaired in absence of ING3. Finally, immunoglobulin class switch recombination (CSR), a physiological mechanism requiring NHEJ repair, was impaired in the absence of ING3. Since deregulation of DNA double strand break repair is associated with genomic instability, we propose a novel function of ING3 as a caretaker tumor suppressor involved in the DNA damage signaling and repair.
Similar content being viewed by others
Introduction
DNA double strand breaks (DSBs) represent the most severe form of damages exerted on DNA since it is the most difficult to repair. DNA DSBs are repaired by mechanisms achieved through the DNA damage response (DDR). The DDR comprises an integrated network of signaling pathways leading to DNA repair if successful or cell death if it fails. Abnormal repair of DNA DSBs can yield mutations and rearrangements thereby potentially giving rise to cell transformation [1]. DNA DSBs are detected by the proteins Mre11, Rad50, and NBS1, which constitute the MRN sensor complex. This allows the recruitment and activation of the ATM kinase which phosphorylates the histone variant H2AX (to form γH2AX), as well as a large number of DDR factors [2,3,4]. γH2AX clusters on the chromatin serve as docking sites for several DDR members therefore enhancing signaling. By binding γH2AX, MDC1 localizes to DSB and then recruits the ubiquitin ligases RNF8 and RNF168. These two ubiquitin ligases permit the accumulation of the downstream proteins 53BP1 and BRCA1 that balance DNA repair by either non-homologous end joining (NHEJ) or homologous recombination (HR) [5,6,7].
The expression of inhibitor of growth (ING) genes is frequently lost in human tumors [8]. The ING family contains five members (ING1–5) which are highly conserved through the evolution [9, 10]. Although ING1 and ING2 are proteins with characterized tumor suppressor functions [11], the biological functions of ING3, ING4, and ING5 are less clear. Previous studies showed the involvement of ING1 and ING2 in DNA replication and suggested their involvement in the DDR [12, 13]. Therefore, we investigated the role of ING genes in the DDR. In Saccharomyces cerevisiae, ING orthologs (Yng1, Yng2, and Pho23) belong to acetylation or deacetylation complexes [14, 15]. These features are conserved in human cells since ING1 and ING2 are part of the Sin3/HDAC1–2 complex, while ING3, ING4, and ING5 associate with NuA4/TIP60, HBO1, and MOZ/MORF complexes, respectively [16, 17]. Remarkably, in the yeast S. cerevisiae, Yng2, the ortholog of ING3, was shown to contribute to intra-S-phase DDR [18].
Herein, we found that ING3-depleted human cells and yeast cells deleted for its ortholog Yng2 are sensitive to DNA damage. In human cells, ING3 accumulates at DNA DSBs and is involved in DDR by promoting ATM phosphorylation and signaling. Moreover, the recruitment of ING3 is a prerequisite for the accumulation of the DNA DSBs signaling proteins RNF8, RNF168, 53BP1, and BRCA1 and allows proper repair by NHEJ or HR, and cell survival. Finally, we found that ING3 deficiency in B-cells results in decreased class switch recombination (CSR).
Results
Absence of ING3 sensitizes cells to DNA damage
We performed DNA damage sensitivity assays in S. cerevisiae deleted for PHO23 (the ING1/2 ortholog), YNG2 (ING3), YNG1 (ING4/5), or RAD9 (53BP1/MDC1/BRCA1). RAD9-deleted cells (rad9Δ) were used in the experiment as an internal control (Fig. 1a). Yeast cells were spotted on rich-media plates containing or exposed to various DNA damaging agents inducing different types of DNA damage. DNA DSBs are mostly induced by bleocin, hydroxyurea (HU) at high dose [19], and ionizing radiation (IR). On the other hand, low doses of HU depletes nucleotide pool and promotes replication stress [20]. Consistently with a previous report suggesting the involvement of Yng2 in cell growth regulation, yng1Δ, yng2Δ, and pho23Δ cells showed a slight growing delay compared with the wild type cells (WT) in the absence of DNA damage [21]. Remarkably, YNG2-deleted cells (yng2Δ) were the most sensitive to every DNA damaging agents tested. Altogether, these results indicate a role of the yeast INGs proteins in pathways regulating cell growth, DDR, and cell survival, with Yng2 possibly playing a key role in DNA damage resistance.
ING3 mediates resistance to DNA damage and cancer cell lines are dependent on ING3 for survival. (a) Wild type (WT) and mutant strains deleted for RAD9 (rad9Δ), YNG1 (yng1Δ), YNG2 (yng2Δ), or PHO23 (pho23Δ) were treated with various types of DNA damaging agents: 5 or 100 mM of hydroxyurea (HU), 0.5 μg/mL of bleocin (Bleo), or 10 Gy of ionizing radiation (IR). Plates were analyzed after 48 h. Summary table of the DNA damage sensitivity assay. (−) resistant, (+) to (+++) increased sensitivity, (++++) no survival. (b) Colony formation assay performed on U2OS cells transfected with siCT, siING3#1, siING3#2, or si53BP1 (53BP1 siRNA was used as a positive control) and treated with various type of DNA damaging agents: Cisplatin (CSP) (50 μM), CPT (10 nM), HU (2.5 mM), Mitomycin C (MMC) (100 ng/mL), MMS (0.5 μM), or Doxorubicin (Dox, 0.125 μM). Graph shows the number of colony formation; three independent experiment were performed. Bar graphs represent mean ± SEM (NSP > 0.05; *P < 0.05; **P < 0.01; ***P < 0.001). c Distribution of ING genes CERES score in cancer cell lines (v.2018Q1 of Avana 1.0 library). d Mean CERES score of ING genes in cancer cell lines (***P < 0.0001; n = 391 per group)
To extend these results to mammalian cells, we performed a DNA damage sensitivity assay in U2OS cells (Fig. 1b) with DNA damaging agents which generate mostly DNA DSBs, predominantly during S phase (CPT, Cisplatin, MMS, and HU). We also used MMC to induce DNA damages repaired by nucleotide excision repair, HR, and translesion synthesis [19, 22]. These results were confirmed with a second ING3 siRNA. Overall, ING3-depleted cells displayed increased sensitivity to all drugs tested. Furthermore, among the ING gene family, cancer cell lines are more dependent on ING3 for survival according to the Cancer Dependency Map database (Fig. 1c, d). Together, these results indicate an important role for ING3 in cell growth and its conserved involvement in the DDR in both yeast and human cells.
ING3 is recruited to DNA DSBs
53BP1 is a marker of DNA damage and a mediator of DNA repair. To gain insight into the role ING3 could play in the DDR, 53BP1 foci formation was analyzed in siCt vs. siING3 U2OS cells (with the same genotoxic agents used above). A lack of increase of 53BP1 foci was observed in siING3 cells. It suggested that ING3 may be involved in the DDR by modulating 53BP1 foci formation (Fig. S1). Consequently, we investigated whether ING3 itself could be recruited at the site of DNA damage. Since no immunofluorescence-compliant ING3 antibody is currently available, we used Halo/GFP-tagged ING3 proteins. As observed for endogenous ING3, Halo-ING3 is mainly located into the nucleus (Fig. S2A), besides its functionality is conserved since it could interact with known partners, RuvBl2 and TIP60. Halo-ING3 or GFP-ING3 accumulated at DNA damage sites 30 min after the induction of DNA DSBs by laser micro-irradiation in U2OS and A549 cells (Fig. 2a and Fig. S2B). Then, we measured the kinetics of its assembly at DSBs in human U2OS cells. We observed that Halo-tagged ING3 accumulation peaked at 30 min (Fig. 2c and Fig. S2E). When comparing recruitment intensity of NBS1, Halo-ING3, and 53BP1, ING3 is recruited 5 min after irradiation, with a dynamic lagging NBS1 but it accumulates more quickly than 53BP1 (Fig. 2c and Fig. S2C). These results indicate that ING3 is recruited early to DNA DSBs and contributes to the recruitment of 53BP1.
ING3 is recruited at DNA damage sites and is required for 53BP1 and BRCA1 recruitment. a Immunofluorescence analysis of cells transfected with Halo-ING3 or GFP-ING3 and damaged by laser micro-irradiation. U2OS or A549 cells were probed, respectively, for NBS1 or 53BP1. Scale bars represent 9 µM. b Immunofluorescence analysis of U2OS cells transfected with Halo-ING3 or not and subjected to laser micro-irradiation. Cells were fixed 5 min after laser micro-irradiation and probed for γH2AX and for 53BP1 or NBS1. Scale bars represent 8 µM. c Immunofluorescence analysis of U2OS cells transfected with Halo-ING3 and subjected to laser micro-irradiation. Fluorescence intensity was assessed on 30 nuclei for each time and represented on a graph. Scale bars represent 11 µM. d Immunofluorescence analysis of 53BP1 or BRCA1 in U2OS cells damaged by laser micro-irradiation. Cells were probed for 53BP1 or HA-BRCA1. Graph shows the intensity of 53BP1 or HA-BRCA1 recruitment at DNA DSBs, at least 30 cells were analyzed. Bar graphs represent mean ± SEM (****P < 0.0001). Scale bars represent 9 µM. Western blot analysis of U2OS cells transfected with siCT or siING3#1 or siING3#2, treated with Dox for 3 h and probed with indicated antibody. All bar graphs represent mean ± SEM (***P < 0.001)
ING3 is necessary for the recruitment of RNF8, RNF168, 53BP1, and BRCA1 to DNA DSBs
To monitor the functional hierarchy of the DDR in cells depleted for ING3, we tested its role for the recruitment of RNF8 and RNF168 proteins. A slight decrease of GFP-RNF8 recruitment and a marked decrease of RNF168 accumulation at DNA DSBs was also observed in the absence of ING3 following laser micro-irradiation. ING3 silencing did not significantly affect the expression of either RNF8 or RNF168 proteins (Fig. S2F and G). The accumulation of RNF8 and RNF168 at DNA DSBs is considered as a critical event for the efficient recruitment of downstream signaling factors, such as 53BP1 and BRCA1, two major mediators of the DDR [23, 24]. Laser micro-irradiated or X-rays-irradiated cells silenced for ING3 exhibited a strong decrease of 53BP1 recruitment at DNA DSBs (Fig. 2d; Fig. S2D and S2H), corroborating with the 53BP1 defective accumulation in siING3 cells treated with DNA damaging drugs (Fig. S1). Similar results were confirmed in A549 cells treated with doxorubicin (Fig. S2I). Furthermore, time course experiments conducted in response to X-rays, showed a lack of recruitment of 53BP1 in the absence of ING3 (Fig. S2D). Importantly, ING3 siRNA had no effect on 53BP1 RNA or protein expression (Fig. 2d and Fig. S2J). Similarly, ING3 silencing also impaired the recruitment of HA-BRCA1 at DNA DSBs in laser micro-irradiated cells and had a slight effect on BRCA1 protein expression (Fig. 2d, Fig. S2H). In conclusion ING3 is required for the accumulation of RNF8, RNF168, 53BP1, and BRCA1.
ING3 recruitment to DSBs enhances ATM signaling
We then evaluated the impact of ING3 on ATM recruitment and activation, which represents an earlier event in the signaling cascade. Immunofluorescence analyses revealed an important decrease of phosphorylated ATM (p-ATM) at micro-irradiation sites and a reduced number of p-ATM foci after doxorubicin treatment in both U2OS and A549 cell lines (Fig. 3a and Fig. S3A). Similar results were observed in non-transformed cells (MRC5) treated with X-rays or laser micro-irradiation (Fig. S3B and Fig. S3E). Interestingly, NBS1 was recruited to laser generated DNA DSBs independently of ING3 (Fig. S3C). Furthermore, we identified two phases in p-ATM levels at DNA damage sites. Until 15 min after laser micro-irradiation p-ATM levels increase independently of ING3 expression. However, at later stages (>15 min post irradiation), p-ATM increase depends on ING3 since p-ATM decrease significantly in the absence of ING3 (Fig. 3b and S3D). This suggests that ING3 is required for ATM signaling maintenance and amplification. In MRC5 cells, when performing a longer time course, we also observed a lack of accumulation of p-ATM foci in the absence of ING3 in response to Dox (Fig S3F).
ING3 regulates ATM activation and signaling in response to DNA DSBs. a Immunofluorescence analysis of U2OS cells transfected with siCT or siING3 and then subjected to laser micro-irradiation, treated with doxorubicin for 3 h or exposed to X-rays. Cells were fixed 15 min after X-rays exposure. Cells were probed for p-ATM. Scale bars represent 8 µM. b Immunofluorescence analysis of U2OS cells transfected with siCT or siING3 and subjected to laser micro-irradiation. Cells were probed for p-ATM. Scale bars represent 10 µM. Graph shows p-ATM signal intensity and fluorescence intensity was assessed on 50 nuclei for each time and represented on a graph. Bar graphs represent mean ± SEM (****P < 0.0001). c Western blot analysis of A549 cells transfected with siCT, siING3, or siATM, treated with Dox for 3 h and probed with indicated antibody. d Immunofluorescence analysis of U2OS cells transfected with siCT or siING3 and subjected to doxorubicin treatment for 3 h. Cells were probed with p-ATM and Cyclin A or PCNA antibodies. Graph shows the number of p-ATM foci per nucleus in PCNA or Cyclin A positive and negative cells, at least 50 cells were analyzed. Scale bars represent 9 µM. Bar graphs represent mean ± SEM (****P < 0.0001). e U2OS cells were transfected with Halo-ING3, treated with an ATM inhibitor (ATMi, 10 μM) for 6 h and submitted to laser micro-irradiation. Cells were then probed for 53BP1. Scale bars represent 9 µM. Graph shows the intensity of Halo-ING3 recruitment at DNA DSBs, at least 30 cells were analyzed. Bar graphs represent mean ± SEM (NSP > 0.05). Western blot analysis of U2OS cells treated ATMi and probed with indicated antibody
ATM is known to phosphorylate several DDR factors including NBS1 [25, 26]. Accordingly, we observed an impairment of both ATM and NBS1 phosphorylation after ING3 downregulation in A549 cells treated with Dox (Fig. 3c). There was no effect on NBS1 protein expression. These results were confirmed in U2OS cells (Fig. S3G). Similar results were observed with other known ATM substrates, CHK2 and BRCA1. Thus, ATM downstream signaling is impaired in siING3 cells, to the same extent observed in siATM cells (Fig. 3c).
Immunofluorescence analysis demonstrated that ING3 was involved in maintaining p-ATM levels throughout cell cycle progression (Fig. 3d and Fig. S3H). Reciprocally, inhibition of ATM activity (Fig. 3e and Fig. S3I) did not impair Halo-ING3 accumulation at DNA DSBs.
Finally, in colony formation experiments no synergy was observed when both ING3 and ATM are downregulated compared to ING3 or ATM downregulated alone. It suggests that the reduced survival observed in ING3-depleted cells is ATM-dependent (Fig S3J).
ING3 is required for TIP60 accumulation at DNA DSBs
In response to DNA damage several phosphatases are known to regulate the phosphorylation state of ATM and a deregulation in these processes can lead to a defective ATM phosphorylation. Indeed, PP2A is required to dephosphorylate ATM in undamaged cells [27] and WIP1 shuts down ATM-dependent phosphorylation and ATM auto-phosphorylation at the end of the DDR [28]. In U2OS cells treated with Dox, PP2A, or WIP1 expression was not affected by ING3 downregulation suggesting that it does not regulate ATM phosphorylation by modifying the expression of its canonical phosphatases (Fig. S4A).
Activation of ATM in response to DNA DSBs requires post-translational modifications. Among those, ATM acetylation by TIP60 has been described as essential for its activation and phosphorylation [29]. ING3 is known to be a stable component of the hNuA4/TIP60 complex [30, 31]. In addition to Tip60, some members of the hNuA4 complex are recruited to DNA DSBs suggesting that these proteins may be recruited together as components of the hNuA4 complex [27,28,29,30]. We tested whether TIP60 recruitment to DNA DSBs could depend on ING3. Interestingly, we observed an impairment of GFP-TIP60 accumulation at DNA DSBs in absence of ING3 (Fig. 4a and Fig. S4B). We then evaluated if TIP60 downregulation could reciprocally hinder ING3 recruitment at DNA DSBs (Fig. 4b and Fig. S4C). TIP60 regulates ING3 protein but not mRNA expression (Fig. S4D and S4E) and we also observed a decreased pool of Halo-tagged ING3 in the nucleus in siTIP60 cells. Thirty minutes after DNA damage induction, the recruitment of Halo-ING3 was reduced but still noticeable in TIP60 downregulated cells. We cannot exclude that TIP60 downregulation has some effect on ING3 recruitment to DNA DSBs, although this effect would be mild. Finally, Halo-ING3 pulldown analysis showed that Halo-ING3 and GFP-TIP60 co-existed in a complex with and without induction of DNA damages (Fig. 4c).
ING3 is required for TIP60 accumulation at DNA DSBs. a Immunofluorescence analysis of U2OS cells co-transfected with GFP-TIP60 and siCT or siING3#1 or siING3#2, and then submitted to laser micro-irradiations. Graph shows the intensity of GFP-TIP60 recruitment at DNA DSBs, at least 30 cells were analyzed. Bar graphs represent mean ± SEM (****P < 0.0001). Scale bars represent 9 µM. b Immunofluorescence analysis of U2OS cells transfected with Halo-ING3 and siCT or siTIP60 and subjected to laser micro-irradiation. Cells were probed for Halo-ING3 and 53BP1. Scale bars represent 13 µM. Bar graphs represent mean ± SEM (****P < 0.0001). c Western blot analysis of Halo pull down on U2OS cells transfected with an empty Halo or Halo-ING3 and treated or not with Doxorubicin for 3 h. d Western blot analysis of U2OS cells transfected with siCT or siTIP60 or siING3#1, treated with Dox for 3 h and probed with indicated antibody. e Immunofluorescence analysis of U2OS cells transfected with siCT, siTIP60, or siING3 and treated with Dox for 3 h. Cells were probed for p-ATM. Scale bars represent 13 µM. Graph shows the number of p-ATM foci per nucleus, at least 50 cells were analyzed. Bar graphs represent mean ± SEM (****P < 0.0001)
Then, we investigated whether ING3 could have an impact on TIP60 ability to acetylate its targets at the chromatin. TIP60 controls the acetylation of histone 2A on Lysine 5 (H2AK5), which can be considered as a marker of TIP60 activity [32, 33]. A decrease of H2AK5 acetylation was observed in siING3 cells similar to what was observed in siTIP60 cells treated with Dox (Fig. 4d and Fig. S4F). Interestingly, while we observed a loss of ATM phosphorylation in siING3 lysates it was not the case with siTIP60 lysates. This was confirmed by immunofluorescence in which pATM dots intensity decreased upon ING3 silencing but not in siTIP60 cells (Fig. 4e and Fig. S4G). Thus, ING3 is necessary for TIP60 recruitment at DNA damage sites but pATM foci accumulation in response to Dox is solely dependent on ING3.
ING3 promotes DNA repair and CSR
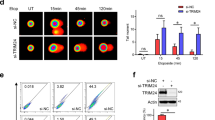
To better characterize the role of ING3 in DNA repair, we monitored the presence of γH2AX foci, a robust readout of DNA DSB repair, immediately after doxorubicin treatment (0 h) or 24 h after the end of this treatment. Interestingly, foci resolution was affected by the absence of ING3 as shown by the larger number of foci still present 24 h after the end of doxorubicin treatment (Fig. 5a and Fig. S5A). This result was confirmed in cells also probed for the S phase marker PCNA. Therefore, this defect in the resolution of γH2AX occurs in all the phases of the cell cycle including S phase (Fig. 5b and Fig. S5).
ING3 is required for DNA repair in response to DNA DSBs. a Immunofluorescence analysis of U2OS cells transfected with siCT or siING3, treated with Dox for 3 h. Cells were probed for γH2AX. Graph shows the number of γH2AX foci per nucleus, at least 50 cells were analyzed. Scale bars represent 12 µM. Bar graphs represent mean ± SEM (***P < 0.001; ****P < 0.0001). b Immunofluorescence analysis of U2OS cells transfected with siCT or siING3, treated with Dox for 3 h. Cells were probed for PCNA and γH2AX. Scale bars represent 12 µM. Graph shows the number of γH2AX foci per nucleus in PCNA positive and negative cells, at least 50 cells were analyzed. Bar graphs represent mean ± SEM (***P < 0.001; ****P < 0.0001)
Since NHEJ and HR are altered upon defective recruitment of 53BP1 and BRCA1 at DNA DSBs [32, 34] and since their recruitment at DNA DSBs was dampened in the absence of ING3, we evaluated if ING3 could impact on the NHEJ and/or HR. Using plasmid-based assays [35, 36] we found that ING3 silencing hindered DNA repair by NHEJ as in si53BP1 cells (Fig. 6a), as well as DNA repair by HR as in siBRCA1 cells (Fig. 6b).
ING3 is required for NHEJ, HR, and class switch recombination. a U2OS cells were transfected with siCT, siING3#1, siING3#2, or si53BP1 and assessed for DNA repair by non-homologous end joining (NHEJ). Bar graphs represent mean ± SEM (*P < 0.05; ***P < 0.001). b U2OS cells were transfected with siCT, siING3#1, siING3#2, or with siBRCA1 and assessed for DNA repair by homologous recombination (HR). Bar graphs represent mean ± SEM (NSP > 0.05; *P < 0.05). c qPCR analysis of CH12F3 cells transfected with GFP-miR-CT or four different GFP-miR-ING3. Bar graphs represent mean ± SEM (*P < 0.05; **P < 0.01). CSR to IgA in CH12F3 cells electroporated with GFP-miR-CT or GFP-miR-ING3#3 or GFP-miR-ING3#4 was measured by flow cytometry. The data are the means of three independent experiments. Bar graphs represent mean ± SEM (*P < 0.05; ***P < 0.001). d CH12F3 cells transfected with GFP-miR-CT or GFP-miR-ING3#4 were labeled with SNARF. Cells were harvested at the indicated time points following SNARF labeling and analyzed by flow cytometry. The data are representative of three independent experiments. e ATM recruitment and activation is a critical step in the DNA damage response and happens in the early stages of the pathway. Its auto-phosphorylation (on Ser 1981) is impaired when the cells are deficient for the ING3 protein in response to DNA damage, thus impacting the following recruitment of 53BP1 and BRCA1 at lesions sites and the DNA repair by NHEJ or HR
To further evaluate the importance of ING3 in the DDR, we measured its impact on the immunoglobulin CSR, which is a physiological pathway involving DNA DSBs repair by NHEJ. To this end, CH12F3 B cells silenced or not for ING3 were stimulated to undergo switching to IgA. These cells were also labeled with SNARF to track cell proliferation by flow cytometry. Interestingly, CH12F3 cells silenced for ING3 showed impaired CSR (Fig. 6c and Fig. S6) without defective cell proliferation (Fig. 6d). Importantly, the extent of CSR impairment correlated with the efficiency of ING3 silencing (Fig. 6c). Thus, ING3 is critical for DNA DSBs repair by NHEJ and HR and has an important role in CSR.
Discussion
ING3 expression is lost or reduced in human tumors [8] and a higher mortality has been associated with a decreased expression of ING3 [37, 38]. Two recent studies suggest that ING3 could act as an oncogene in prostate cancer [39, 40] but mainly, ING3 has been shown to exert tumor suppressor functions as a gatekeeper. Thus, ING3 modulates p53-mediated transcription and regulates apoptosis in melanoma cells in response to UV [41, 42]. ING3 is a stable component of the hNuA4 histone acetyltransferase (HAT) complex, known to be involved in transcriptional activation [16]. Therfore, it can modulate H4K12 acetylation on promoter of genes (mTOR, Cdc42, Rac1, and RhoA) [43]. In this study, we reveal a new role for ING3 in the DDR and repair. For the first time, we propose a new function of ING3 as a caretaker tumor suppressor involved in early DNA DSBs signaling and repair (Fig. 6e).
In accordance with previous reports [18, 44], we found that yeast cells deleted for the ING3 ortholog (Yng2) were highly sensitive to DNA damage. This yeast DNA damage sensitivity assay suggests that ING3 could also be involved in the DNA replication process [45]. Human cells depleted for ING3 are more sensitive to DNA damages and have a defective DDR by affecting 53BP1 foci formation. ING3 was specifically recruited at DNA DSBs in an ATM-independent manner. Interestingly, an ING3 knockdown led to decreased p-ATM at DNA DSBs that may be more dependent on its phosphorylation rather than the activities of its related phosphatases. ING3 has no impact on the expression of these phosphatases which are known to regulate ATM phosphorylation and its activation [27, 28]. However, we cannot exclude an impact of ING3 on the recruitment of these phosphatases at DNA damage sites. NBS1, a downstream target of ATM whose accumulation at DNA damage sites is ING3-independent, showed a defective phosphorylation status after doxorubicin treatment in siING3 cells. Thus, ING3 could be required for the subsequent maintenance and amplification of ATM signaling after the initial and rapid recruitment of ATM by MRN and therefore would take part in the positive feedback loop regulated by MDC1, which results in ATM accumulation on chromatin [46]. TIP60, via its chromodomain interacts specifically at DNA DSBs with H3K9me3 increasing its catalytic activity necessary for acetylation of ATM on Lysine 3016 [29, 30, 47]. Since the lack of ATM phosphorylation observed in siING3 cells is not observed in siTIP60 cells, our results suggest that ING3-dependent phosphorylation of ATM is independent on TIP60. One hypothesis would be that ING3 has an inhibitory effect on DNA-PKcs activity which itself negatively regulates ATM phosphorylation [48]. Or, the ATM-linked proliferation inhibition observed in siING3 cells could also be due to the activation of the tumor suppressor ARF, as it has been previously described [49].
TIP60 also regulates chromatin remodeling at DNA DSBs through its interaction with the hNuA4 complex [34]. We found that TIP60 accumulation at DNA damage sites is ING3-dependent. Moreover, we observed that TIP60 interaction with ING3 is independent on DNA damage. Since both TIP60 and ING3 interact in the hNuA4 complex we can hypothesize that this complex is relocalized or stabilized at DNA double strand breaks after a genotoxic stress. TIP60 has been shown to regulate ATM activation independently of the hNuA4 complex [30]. Thus, ING3 and TIP60 might be recruited at DSBs sites in the hNua4 complex and then are released to act in the DDR. The recruitment of this complex could depend on the interaction of ING3 with the histone mark H3K4me3. Indeed, ING3 is able to interact with H3K4me3 through its PHD domain and this association seems to be important for ING3 tumor suppressive activities [50]. But another mechanism could also be responsible. Under certain circumstances, a defective 53BP1 accumulation at DNA lesions could impact on TIP60 recruitment [51].
In the absence of TIP60, we observed a decrease in ING3 protein levels but no change in ING3 mRNA expression. These findings suggest that TIP60 might regulate the ING3 protein stability independently of DNA damage. Interestingly, previous studies reported that TIP60, by acetylating proteins improved their stabilization [44]. In agreement, Esa1, the yeast ortholog of TIP60 was shown to acetylate Yng2, the ING3 ortholog in order to prevent its proteasomal degradation [44]. Such a mechanism could also explain at least partly the accumulation of ING3 we observed following doxorubicin treatment. Taken together, these results suggest that ING3 could be acetylated by TIP60 to promote its stability.
Cells downregulated for ING3 failed to accumulate several proteins involved in the DDR, such as RNF8, RNF168, 53BP1, and BRCA1, suggesting ING3 might act earlier. ING3-depleted cells generated fewer colonies or displayed decreased NHEJ and HR activities, indicating a role of ING3 in these two repair pathways. The role of 53BP1 and ATM in a physiological context was highlighted in CSR, which involves NHEJ repair during the immunoglobulin switch in mature B cells [52, 53]. In accordance with our results showing that ING3 is necessary for ATM phosphorylation and recruitment of 53BP1, we have demonstrated that ING3 is also necessary for CSR, highlighting its crucial role in the DDR.
Our work is part of a study aiming at investigating the role of ING proteins in the DDR [54]. It indicates that ING3 may play an important role in maintaining genomic stability and preventing cancer initiation and progression. As a critical actor of the NHEJ signaling, the status of ING3 in tumors could be an important factor in predicting the response to chemotherapy and thus ING3 could be a target for synthetic lethality.
Material and methods
DNA damage sensitivity of yeast strains
Wild-type and RAD9, YNG1, YNG2, PHO23 deleted strains used in this study are in the BY4741 background and were obtained from the EUROSCARF collection (Frankfurt, Germany) (Accession numbers Y03576, Y01840, Y07786, Y07234, and Y00000 respectively). DNA damage sensitivity analysis was performed by drop test. Five-fold serial dilutions of exponentially growing cultures of the indicated strains were prepared in a sterile 96-well plate with the highest concentration being 5 × 106 cells/ml. Cells were then spotted on YPD media either containing DNA damaging agents (5 or 100 mM of hydroxyurea (Sigma-Aldrich) or 0.5 μg/ml of Bleocin (Calbiochem; San Diego, CA, USA) or irradiated with IR (10 Gy, CellRad, Faxitron (no filter) 130 kV, 5 mA). All plates were then incubated at 30 °C for 2 days and photographed.
Cell culture and drug treatments
U2OS osteosarcoma cell lines and MRC5 human lung fibroblast were cultured in McCoy medium (Thermo Scientific; Illkirch, France) supplemented with 10% of decomplemented fetal bovine serum and antibiotics (penicillin/streptomycin, Thermo Scientific). A549 adenocarcinoma cells were cultured in DMEM medium (Thermo Scientific) supplemented with 10% fetal bovine serum and antibiotics (penicillin/streptomycin). U2OS, MRC5, and A549 cells were treated with Doxorubicin (Dox), Camptothecin (CPT), Methyl methanesulfonate (MMS), Cisplatin (CSP), Mitomycin C (MMC), HU (Sigma-Aldrich; St. Louis, MO, USA) at indicated times. U2OS cells were irradiated with IR (2 Gy) (CellRad, Faxitron). CH12F3 mouse cell lines were cultured in RPMI medium (Thermo Scientific) supplemented with 10% of decomplemented fetal bovine serum, 10 mM Hepes (Thermo Scientific), 1 mM sodium pyruvate (Thermo Scientific), and 50 µM β-mercaptoethanol (Thermo Scientific). Cells were incubated at 37 °C in a humidified atmosphere, 5% CO2.
Plasmids construction, siRNAs, and transfection
ING3 cDNA was cloned using specific primers, 5′-CGAAGCGATCGCCATGGCGGACAGTGCGGAACTAAAG-3′ (sense) and 5′-GTCGGTTTAAACGTCCAATGAAATAATGTCTGGTATGATGCCAA-3′ (antisense) into Halo tag pFN21A vector according to the manufacturer’s instructions (Promega; Madison, WI, USA). A validated and a custom stealth siRNAs (Invitrogen; Carlsbad, CA, USA) were used for the ING3 downregulations, RNAi #HSS182564 for siING3#1 and for siING3#2 (5′-CCUAGAAGACUAUCUGGAAAUGAUU-3′). For ATM downregulation we used the validated stealth RNAi #HSS181472, #HSS181473, #HSS181474 (Invitrogen), for 53BP1 downregulation we used the validated stealth RNAi #HSS110908, #HSS110909, #HSS110910 (Invitrogen). As a control, the universal stealth RNAi negative control (#12935110, Invitrogen) was used. Stealth siRNA were transfected using Lipofectamine RNAimax (#13778-075, Invitrogen), according to the manufacturer’s instructions. Plasmids were transfected with Lipofectamine LTX in combination with Plus reagent (#15338-100, Invitrogen), according to the manufacturer’s instructions. To knockdown ING3 in CH12F3 mouse cells, we used the BLOCK-iT Pol II miR RNAi Expression Vector Kit (#K4936-00, Invitrogen). The miRNA duplex was inserted into the pcDNA 6.2-GW miR.
Western blot
Whole cell protein extracts were prepared for immunoblotting by cell lysis with RIPA buffer (#9806, Cell Signaling; Danvers, MA, USA) in combination with a protease inhibitor cocktail (#5871, Cell Signaling). Protein samples were subjected to electrophoresis using the NuPAGE® Novex 4–12% Bis–Tris Gels Electrophoresis system (# NP0329BOX, Invitrogen). The antibodies used in this study were 53BP1, MDC1, WIP1, Cyclin A, RNF8 from Santa Cruz Biotechnology, p-ATM, p-p53, p-Chk2, PP2A, PP2AC, PP2AB, γH2AX, p-NBS1, and p-BRCA1 from Cell Signaling Technology, ATM and RNF168 from Millipore, H2AK5ac and RuvBl2 from Abcam, NBS1 from Novus Biological or from GeneTex, PCNA from BD Pharmigen and β-actin from Sigma Aldrich. ING3 antibody was a kind gift of K. Riabowol (University of Calgary, Canada).
Halo pull down
U2OS cells were lysed in mammalian lysis buffer (Promega) and subjected to pull down using the kit HaloTag® Protein Purification System (Promega) following manufacturer’s instructions.
Laser micro-irradiation and imaging of live and fixed cells
Cells were grown on chambered coverglasses (Lab-Tek). Cells transfected with Halo-tag plasmids were treated overnight with Halo-tag TMR Direct Ligand (#G299A) according to the manufacturer’s instructions (Promega). Then, cells were sensitized with 10 µM of Hoechst (Hoechst 33258, Sigma-Aldrich) for 15 min at 37 °C. Localized DNA DSBs were generated by an exposure to ultraviolet-A laser beam [55] (Kong X 2009). Laser micro-irradiation was performed by using a Spinning-Disk confocal microscope NIKON Ti-E equipped with a 37 °C heating stage and a 405 nm laser diode (55 mW) focused through a ×60 Plan Apo/1,4 oil. The time of cell exposure to the laser beam was around 500 ms. Laser settings (0.40 mW output, 50 scans) were chosen to generate a detectable damage response restricted to the laser path in a pre-sensitization-dependent manner without noticeable cytotoxicity.
Kinetic of recruitment to DNA DSBs and fluorescence analysis
Fluorescence intensity was assessed on 10 nuclei at different time points (0, 5, 15, 30, 60, 240 min) by using ImageJ software. Quantification of Halo-ING3, γH2AX, NBS1, 53BP1, and p-ATM signal intensities was measured on the laser track path and normalized with the background fluorescence intensity.
Immunofluorescence staining
Cells were grown on coverslips in 12-well plates and probed as described previously [56]. Briefly, cells were subjected to a pre-lysis treatment for removing protein’s soluble fraction. Then, cells were fixed in a 100% cold ethanol solution for 4 min at 4 °C or HaloTag TMRDirect Ligand-treated cells were fixated in a 2% PFA solution for 20 min at room temperature. Cells were washed in 1X PBS and probed. Images were acquired with a confocal SP8 TCS Leica microscope.
Cell sensitivity assay
U2OS cells were transfected with siRNA and 48 h later exposed to doxorubicin (0.125 µM) for 3 h or exposed to IR (2 Gy). Then, fresh media was added and cells were incubated at 37 °C in a humidified atmosphere of 5% CO2 for 11 days to allow colony formation. Plates were probed with 1% methylene blue in Tris buffer and colonies with 50 cells or more were counted. Results were normalized with the corresponding non-treated condition. Cells were seeded to take into account the relative plating efficiencies and treatment effect.
NHEJ and HR assay
NHEJ assay were performed as previously described [57]. HR assay were performed as previously described using a DR-GFP reporter in U2OS cells [58, 59]
CSR assay, flow cytometry analysis, and SNARF labeling
For the CSR assay, CH12F3 cells downregulated for ING3 were stimulated with 250 ng/ml of functional grade purified anti-mouse CD40 clone HM40-3 (eBioscience), 10 ng/ml recombinant murine IL-4 (Preprotech; London, UK), and 1 ng/ml recombinant human TGF-β1 (R&D Systems) for 24 h and then analyzed by flow cytometry.
CH12F3 cells were probed with PE-conjugated anti-murine IgA (Southern Biotech) and APC-conjugated anti-murine IgM (Southern Biotech). Cells were then acquired on Gallios flow cytometer (Beckman Coulter) and the data were analyzed using Kaluza software. For SNARF labeling, cells were incubated with 9 μM SNARF (carboxylic acid acetate, succinimidyl ester) in 5% FBS for 10 min at 37 °C. Then, the reaction was quenched with FBS and cells were washed with 2.5% FBS. Cells were analyzed on Gallios flow cytometer at indicated time points.
Evaluation of dependency in cancer cell lines
CERES score for each ING gene was extracted from the latest (2018Q1) release of 391 cell lines screened with the CRISPR Avana 1.0 library (May 2018; https://depmap.org/portal). Two-sided paired T-test were performed to compare mean of CERES score of all IING1, ING2, ING4, and ING5 genes to ING3. Data were plotted and p-values were calculated in GraphPad Prism®.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was isolated from U2OS cells by NucleoSpin® RNA (Macherey-Nagel, UK) and then reversely transcribed into cDNA using High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, USA). RT-qPCR was carried out with the QuantBio5 system (Applied Biosystems, USA) under the following conditions: 95 °C for 10 min, 40 cycles of 95 °C for 10 s, 55 °C for 10 s, 72 °C for 20 s, and 72 °C for 10 min. The primers used U6-Fwd: CTCGCTTCGGCAGCACA; U6-Rev: AACGCTTCACGAATTTGCGT; TP53BP1-Fwd: GGGATCGATCTGGAGGGACT; TP53BP1-Rev: TCCAGTAGGGTCCATCTGCT; LIG4-Fwd: TGTAGACTGCGCCGCATTAG; LIG4-Rev: TCCAGGGAGTCAAAAACGGG; DCLRE1C-Fwd: TGGAGTGCAGCTTGAAGGTTTAT; DCLRE1C-Rev: TCTCTTCCTTCTCTCCTGATG; XRCC5-Fwd: GTCGGCGTGGCTTTTCCTCAT; XRCC5-Rev: AAGCTCTGTGCAGCAGACAC; RAD51-Fwd: ATACTGTGGAGGCTGTTGCC; RAD51-Rev: GCTTTGGCTTCACTAATTCCCT; RBBP8-Fwd: AGGGCGAAAGAGAAAAGCGA; RBBP8-Rev: TGGACAGGTCAAATACCGCC; BRCA1-Fwd: TTGCGGGAGGAAAATGGGTA; and BRCA1-Rev: TACCATCCATTCCAGTTGATC. Relative quantification in RT-PCR was performed using 2−ΔΔCT (threshold cycle value) for calculation of fold change with U6 as control.
References
Ciccia A, Elledge SJ. The DNA damage response: making it safe to play with knives. Mol Cell. 2010;40:179–204.
You Z, Chahwan C, Bailis J, Hunter T, Russell P. ATM activation and its recruitment to damaged DNA require binding to the C terminus of Nbs1. Mol Cell Biol. 2005;25:5363–79.
Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J Biol Chem. 1998;273:5858–68.
Lavin MF, Kozlov S, Gatei M, Kijas AW. ATM-dependent phosphorylation of all three members of the MRN complex: From sensor to adaptor. Biomolecules. 2015;5:2877–902.
Nakada S. Opposing roles of RNF8/RNF168 and deubiquitinating enzymes in ubiquitination-dependent DNA double-strand break response signaling and DNA-repair pathway choice. J Radiat Res. 2016;57:i33–i40.
Doil C, Mailand N, Bekker-Jensen S, Menard P, Larsen DH, Pepperkok R, et al. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell. 2009;136:435–46.
Mailand N, Bekker-Jensen S, Faustrup H, Melander F, Bartek J, Lukas C, et al. RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell. 2007;131:887–900.
Guérillon C, Bigot N, Pedeux R. The ING tumor suppressor genes: Status in human tumors. Cancer Lett. 2014;345:1–16.
He GHY, Helbing CC, Wagner MJ, Sensen CW, Riabowol K. Phylogenetic analysis of the ING family of PHD finger proteins. Mol Biol Evol. 2005;22:104–16.
Ythier D, Larrieu D, Brambilla C, Brambilla E, Pedeux R. The new tumor suppressor genes ING: genomic structure and status in cancer. Int J Cancer. 2008;123:1483–90.
Guérillon C, Larrieu D. Pedeux RING1 and ING2: multifaceted tumor suppressor genes. Cell Mol Life Sci. 2013;70. https://doi.org/10.1007/s00018-013-1270-z.
Larrieu D, Ythier D, Binet R, Brambilla C, Brambilla E, Sengupta S, et al. ING2 controls the progression of DNA replication forks to maintain genome stability. EMBO Rep. 2009;10:1168–74.
Wong RPC, Lin H, Khosravi S, Piche B, Jafarnejad SM, Chen DWC, et al. Tumour suppressor ING1b maintains genomic stability upon replication stress. Nucleic Acids Res. 2011;39:3632–42.
Loewith R, Smith JS, Meijer M, Williams TJ, Bachman N, Boeke JD, et al. Pho23 is associated with the Rpd3 histone deacetylase and is required for its normal function in regulation of gene expression and silencing in Saccharomyces cerevisiae. J Biol Chem. 2001;276:24068–74.
Woo H, Dam HaS, Lee SB, Buratowski S, Kim T. Modulation of gene expression dynamics by co-transcriptional histone methylations. Exp Mol Med. 2017;49:e326.
Doyon Y, Cayrou C, Ullah M, Landry A-J, Côté V, Selleck W, et al. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol Cell. 2006;21:51–64.
Avvakumov N, Lalonde M-E, Saksouk N, Paquet E, Glass KC, Landry A-J, et al. Conserved molecular interactions within the HBO1 acetyltransferase complexes regulate cell proliferation. Mol Cell Biol. 2012;32:689–703.
Choy JS, Kron SJ. NuA4 subunit Yng2 function in intra-S-phase DNA damage response. Mol Cell Biol. 2002;22:8215–25.
Wyatt HDM, Sarbajna S, Matos J, West SC. Coordinated actions of SLX1-SLX4 and MUS81-EME1 for Holliday junction resolution in human cells. Mol Cell. 2013;52:234–47.
Merrill BJ, Holm C. A requirement for recombinational repair in Saccharomyces cerevisiae is caused by DNA replication defects of mec1 mutants. Genetics. 1999;153:595–605.
Loewith R, Meijer M, Lees-Miller SP, Riabowol K, Young D. Three yeast proteins related to the human candidate tumor suppressor p33ING1 are associated with histone acetyltransferase activities. Mol Cell Biol. 2000;20:3807–16.
Lee Y-J, Park S-J, Ciccone SLM, Kim C-R, Lee S-H. An in vivo analysis of MMC-induced DNA damage and its repair. Carcinogenesis. 2006;27:446–53.
Panier S, Durocher D. Regulatory ubiquitylation in response to DNA double-strand breaks. DNA Repair. 2009;8:436–43.
Jackson SP, Durocher D. Regulation of DNA damage responses by ubiquitin and SUMO. Mol Cell. 2013;49:795–807.
Zhao S, Weng YC, Yuan SS, Lin YT, Hsu HC, Lin SC, et al. Functional link between ataxia-telangiectasia and Nijmegen breakage syndrome gene products. Nature. 2000;405:473–7.
Lavin MF. ATM and the Mre11 complex combine to recognize and signal DNA double-strand breaks. Oncogene. 2007;26:7749–58.
Goodarzi AA, Jonnalagadda JC, Douglas P, Young D, Ye R, Moorhead GBG, et al. Autophosphorylation of ataxia-telangiectasia mutated is regulated by protein phosphatase 2A. EMBO J. 2004;23:4451–61.
Shreeram S, Demidov ON, Hee WK, Yamaguchi H, Onishi N, Kek C, et al. Wip1 phosphatase modulates ATM-dependent signaling pathways. Mol Cell. 2006;23:757–64.
Sun Y, Xu Y, Roy K, Price BD. DNA damage-induced acetylation of lysine 3016 of ATM activates ATM kinase activity. Mol Cell Biol. 2007;27:8502–9.
Sun Y, Jiang X, Chen S, Fernandes N, Price BD. A role for the Tip60 histone acetyltransferase in the acetylation and activation of ATM. Proc Natl Acad Sci USA. 2005;102:13182–7.
Bhoumik A, Singha N, O’Connell MJ, Ronai ZA. Regulation of TIP60 by ATF2 modulates ATM activation. J Biol Chem. 2008;283:17605–14.
Kimura A, Horikoshi M. Tip60 acetylates six lysines of a specific class in core histones in vitro. Genes Cells. 1998;3:789–800.
Bhaumik SR, Smith E, Shilatifard A. Covalent modifications of histones during development and disease pathogenesis. Nat Struct Mol Biol. 2007;14:1008–16.
Murr R, Loizou JI, Yang Y-G, Cuenin C, Li H, Wang Z-Q, et al. Histone acetylation by Trrap-Tip60 modulates loading of repair proteins and repair of DNA double-strand breaks. Nat Cell Biol. 2006;8:91–9.
Stucki M, Clapperton JA, Mohammad D, Yaffe MB, Smerdon SJ, Jackson SP. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell. 2005;123:1213–26.
Sartori AA, Lukas C, Coates J, Mistrik M, Fu S, Bartek J, et al. Human CtIP promotes DNA end resection. Nature. 2007;450:509–14.
Gunduz M, Beder LB, Gunduz E, Nagatsuka H, Fukushima K, Pehlivan D, et al. Downregulation of ING3 mRNA expression predicts poor prognosis in head and neck cancer. Cancer Sci. 2008;99:531–8.
Wang Y, Dai DL, Martinka M, Li G. Prognostic significance of nuclear ING3 expression in human cutaneous melanoma. Clin Cancer Res. 2007;13:4111–6.
Nabbi A, McClurg UL, Thalappilly S, Almami A, Mobahat M, Bismar TA, et al. ING3 promotes prostate cancer growth by activating the androgen receptor. BMC Med. 2017;15:103.
McClurg UL, Nabbi A, Ricordel C, Korolchuk S, McCracken S, Heer R, et al. Human ex vivo prostate tissue model system identifies ING3 as an oncoprotein. Br J Cancer. 2018;118:713–26.
Nagashima M, Shiseki M, Pedeux RM, Okamura S, Kitahama-Shiseki M, Miura K, et al. A novel PHD-finger motif protein, p47ING3, modulates p53-mediated transcription, cell cycle control, and apoptosis. Oncogene. 2003;22:343–50.
Wang Y, Li G. ING3 promotes UV-induced apoptosis via Fas/caspase-8 pathway in melanoma cells. J Biol Chem. 2006;281:11887–93.
Suzuki S, Nozawa Y, Tsukamoto S, Kaneko T, Imai H, Minami N. ING3 is essential for asymmetric cell division during mouse oocyte maturation. PLoS One. 2013;8:e74749.
Lin Y, Qi Y, Lu J, Pan X, Yuan DS, Zhao Y, et al. A comprehensive synthetic genetic interaction network governing yeast histone acetylation and deacetylation. Genes Dev. 2008;22:2062–74.
Larrieu D, Pedeux R. SharING out the roles in replicating DNA. Cell Cycle 2009;8. https://doi.org/10.4161/cc.8.22.9947.
Lou Z, Minter-Dykhouse K, Franco S, Gostissa M, Rivera MA, Celeste A. et al. MDC1 maintains genomic stability by participating in the amplification of ATM-dependent DNA damage signals. Mol Cell. 2006;21:187–200.
Sun Y, Jiang X, Xu Y, Ayrapetov MK, Moreau LA, Whetstine JR, et al. Histone H3 methylation links DNA damage detection to activation of the tumour suppressor Tip60. Nat Cell Biol. 2009;11:1376–82.
Zhou Y, Lee J-H, Jiang W, Crowe JL, Zha S, Paull TT. Regulation of the DNA damage response by DNA-PKcs inhibitory phosphorylation of ATM. Mol Cell. 2017;65:91–104.
Velimezi G, Liontos M, Vougas K, Roumeliotis T, Bartkova J, Sideridou M, et al. Functional interplay between the DNA-damage-response kinase ATM and ARF tumour suppressor protein in human cancer. Nat Cell Biol. 2013;15:967–77.
Kim S, Natesan S, Cornilescu G, Carlson S, Tonelli M, McClurg UL, et al. Mechanism of histone H3K4me3 recognition by the plant homeodomain of inhibitor of growth 3. J Biol Chem. 2016;291:18326–41.
Jacquet K, Fradet-Turcotte A, Avvakumov N, Lambert J-P, Roques C, Pandita RK, et al. The TIP60 complex regulates bivalent chromatin recognition by 53BP1 through direct H4K20me binding and H2AK15 acetylation. Mol Cell. 2016;62:409–21.
Bothmer A, Robbiani DF, Di Virgilio M, Bunting SF, Klein Ia, Feldhahn N, et al. Regulation of DNA end joining, resection, and immunoglobulin class switch recombination by 53BP1. Mol Cell. 2011;42:319–29.
Lumsden JM, McCarty T, Petiniot LK, Shen R, Barlow C, Wynn TA, et al. Immunoglobulin class switch recombination is impaired in Atm-deficient mice. J Exp Med. 2004;200:1111–21.
Guérillon C. Les protéines suppressives de tumeurs ING1, ING2 et ING3: régulation par sumoylation et implication dans la réponse aux dommages à l’ADN. http://www.theses.fr 2014.http://www.theses.fr/s175004. Accessed 3 Jul 2018.
Kong X, Mohanty SK, Stephens J, Heale JT, Gomez-Godinez V, Shi LZ, et al. Comparative analysis of different laser systems to study cellular responses to DNA damage in mammalian cells. Nucleic Acids Res. 2009;37:e68.
Pedeux R, Sengupta S, Shen JC, Demidov ON, Saito S, Onogi H, et al. ING2 regulates the onset of replicative senescence by induction of p300-dependent p53 acetylation. Mol Cell Biol. 2005;25:6639–48.
Galanty Y, Belotserkovskaya R, Coates J, Polo S, Miller KM, Jackson SP. Mammalian SUMO E3-ligases PIAS1 and PIAS4 promote responses to DNA double-strand breaks. Nature. 2009;462:935–9.
Pierce AJ, Johnson RD, Thompson LH, Jasin M. XRCC3 promotes homology-directed repair of DNA damage in mammalian cells. Genes Dev. 1999;13:2633–8.
Pierce AJ, Jasin M. Measuring recombination proficiency in mouse embryonic stem cells. Methods Mol Biol. 2005;291:373–84.
Acknowledgements
We thank E. Curran, A. Breton, D. Jacquet, S. Huet, and T. Lebeaupin for their participation and the technical support. We thank K. Riabowol for the ING3 antibody. We thank E. Chevet and O. Binda for discussion and helping with the manuscript. We thank the photonic facility of the Microscopy Rennes Imaging Center (MRic-Photonics) of Biosit, Université de Rennes 1. AM was a recipient of a doctoral fellowship from La Ligue Contre le Cancer and Region Bretagne. RP and MG were recipients of Ulysses “The France–Ireland Exchange Scheme”. NB was a recipient of ANR program (SAFE 2012) (ANR-11-RPIB-0012). CR is a recipient of FHU Camin (CHU Rennes) and Nuovo-Soldati Fundation (CHU Genève) fellowships. RP is supported by Institut National de la Santé et de la Recherche Médicale (INSERM). The work was supported by La Ligue Contre le Cancer (Grand Ouest), Association pour la Recherche sur le Cancer (ARC), AIS Rennes Métropole, Biosit and Action Incitative Université de Rennes 1.
Author information
Authors and Affiliations
Contributions
AM, JA, MG, and RP conceived and designed the experiments. AM, JA, NB, CR, LC, and RP performed the experiments. AM, NB, TG, MG, and RP analyzed the data. RP and TG approved the manuscript and contributed to the financial support. AM, JA, CR, and RP wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Oren
Supplementary information
Rights and permissions
About this article
Cite this article
Mouche, A., Archambeau, J., Ricordel, C. et al. ING3 is required for ATM signaling and DNA repair in response to DNA double strand breaks. Cell Death Differ 26, 2344–2357 (2019). https://doi.org/10.1038/s41418-019-0305-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0305-x
- Springer Nature Limited