Abstract
Cell migration depends on the dynamic organisation of the actin cytoskeleton and assembly and disassembly of focal adhesions (FAs). However, the precise mechanisms coordinating these processes remain poorly understood. We previously identified the oestrogen-related receptor α (ERRα) as a major regulator of cell migration. Here, we show that loss of ERRα leads to abnormal accumulation of actin filaments that is associated with an increased level of inactive form of the actin-depolymerising factor cofilin. We further show that ERRα depletion decreases cell adhesion and results in defective FA formation and turnover. Interestingly, specific inhibition of the RhoA-ROCK-LIMK-cofilin pathway rescues the actin polymerisation defects resulting from ERRα silencing, but not cell adhesion. Instead, we found that MAP4K4 is a direct target of ERRα and down-regulation of its activity rescues cell adhesion and FA formation in the ERRα-depleted cells. Altogether, our results highlight a crucial role of ERRα in coordinating the dynamic of actin network and FAs through the independent regulation of the RhoA and MAP4K4 pathways.
Similar content being viewed by others
Introduction
Cell migration is essential for embryonic development, wound healing and immune response [1]. Its dysregulation contributes to many pathologies including cancer cell dissemination [2].
The multistep process of cell movement requires highly coordinated changes in cell morphology and interactions with the extracellular matrix (ECM). Cell migration can be divided into four discrete steps: formation of protrusion, adhesion to the ECM, generation of traction forces at the adhesion sites and release of adhesion at the rear, which allow the cell to move forward [1, 3]. The growing actin network pushes the membrane and promotes lamellipodial protrusion at the leading edge. Protrusions are then stabilised by integrin-based protein complexes known as focal adhesions (FAs) connecting the actin cytoskeleton to the ECM. In addition, actomyosin fibres promote cell contraction and generate traction forces at the FA sites. It also induces cell retraction to allow the cells to move forward and migrate. Therefore this sequence of events involves a dynamic organisation of the actin cytoskeleton and a controlled assembly and disassembly of FAs that must be coordinated both in space and time [1, 4, 5].
The Rho GTPases plays a major role in regulating the actin cytoskeleton and cell migration [6,7,8]. Notably, RhoA initiates the process by inducing actin assembly at the cell front and mediates the interaction of contractile actin-myosin filaments that promotes FA maturation, cell body translocation and rear retraction [8,9,10,11]. RhoA promotes actin assembly through its effectors mammalian homologue of Drosophila diaphanous (mDIA) and Rho-associated protein kinase (ROCK) [6, 12]. mDIA initiates actin filament assembly by nucleation, whereas ROCK promotes actin polymerisation through the activation of the LIM kinase (LIMK) resulting in the inhibition of the actin-severing activity of cofilin [13, 14].
The mitogen-activated protein kinase kinase kinase kinase (MAP4K4) has also been shown to be involved in cell migration, in particular by specifically controlling FA dynamics [15, 16]. Interestingly, microtubules serve as tracks to deliver proteins that act locally at FAs to promote their turnover [11, 17]. In this way, MAP4K4 is delivered to FA sites to induce the activation of the GTPase Arf6, which promotes integrin internalisation from the cell surface and FA turnover [18]. MAP4K4 also regulates FA disassembly in migrating cells by phosphorylating moesin, which displaces talin from integrins and induces their inactivation [19]. Furthermore, the pro-migratory function of MAP4K4 relies on its ability to induce endocytosis and activation of the integrin β1 adhesion receptor [20]. Overall, cell migration is a complex and dynamic phenomenon, which involves crosstalks between actin and FAs. However, how these processes are coordinated to support cell migration is not clearly understood.
Our team and others have demonstrated that the oestrogen-related receptor α (ERRα) is an important factor promoting cell migration [21,22,23]. ERRα is an orphan member of the nuclear receptor superfamily and acts as a transcription factor [24, 25]. ERRα is strongly expressed in several types of cancers and its high expression correlates with poor prognosis [23, 25, 26]. In addition, accumulating evidence indicates that ERRα plays a major role in tumoral growth and progression via stimulation of cell proliferation [27, 28], angiogenesis [29,30,31,32], aerobic glycolysis [33, 34] and ECM invasion [35, 36]. In breast cancer cells, we previously showed that ERRα promotes directional cell migration by regulating RhoA stability and activity [22]. Consequently, the invalidation of ERRα leads to impaired cell migration, which is associated with cell disorientation, disorganised actin filaments and defective lamellipodium formation [22]. Yet the specific roles of ERRα in the regulation of the discrete processes involved in cancer cell migration, such as actin- and FA dynamics, remain unclear.
In the present study, we report that ERRα controls actin polymerisation and organisation by modulating cofilin activity through the RhoA-ROCK-LIMK pathway. We also found that ERRα promotes cell adhesion independently of its role on the actin cytoskeleton. Indeed, ERRα directly regulates the expression of MAP4K4, and thereby contributes to the modulation of FA formation and turnover. Together, our study identifies ERRα as a major actor involved in the coordination of actin and FA dynamics that may contribute to efficient cancer cell migration and metastasis.
Materials and methods
Cell lines
Mycoplasma-free MDA-MB231, HeLa and BT474 cells were cultured in 4.5 g/l glucose DMEM supplemented with 10% FCS (Gibco), 10 U/ml penicillin (Gibco) and 10 µg/ml streptomycin (Gibco). Cells were maintained in a 5% CO2 atmosphere at 37 °C. MDA-MB231 cells were transfected with pEGFP-paxillin plasmid (a generous gift from Sandrine Etienne-Manneville, Institut Curie, Paris, France), selected with 1 mg/ml G418 (Sigma-Aldrich) and maintained as cell populations.
Cell transfection
siRNA were transfected using INTERFERin (Polyplus Transfection) according to the manufacturer’s protocol. Briefly, 3 × 105 cells per ml were seeded in 6-well plates and transfected with 25 pmol/ml of control, ERRα or MAP4K4 siRNAs. Cells were harvested 48 or 72 h after transfection. Control siRNAs were from Thermo Fisher Scientific (medium GC Stealth RNA interference negative control duplexes). siRNAs were from Eurogentec: ERR#1(GGCAGAAACCUAUCUCAGGUU), ERR#2(GAAUGCACUGGUGUCACAUCUGCUG), MAP4K4#A (GUAGCACACUCCAGAAACA), MAP4K4#B (GCGAAGGAGAGAACAAGAA). Cells were harvested 48 or 72 h after transfection.
Biochemical reagents
Y27632 dihydrochloride monohydrate (Sigma-Aldrich, Y0503) was used at 2.5; 5 or 10 μM; Pyr1 (Lim K inhibitor) (a gift from Laurence Lafanechère, Institute for Advanced Biosciences, Grenoble, France) was used at 1; 5 or 10 μM; Blebbistatin (Sigma-Aldrich, B0560) was used at 5 μM; PF-06260933 dihydrochloride (Sigma-Aldrich, PZ0272) was used at 0.25 or 0.5 μM; and GNE495 (Clinisciences, HY-100343) was used at 1 μM. Cells were pre-treated (Western blot, xCELLigence) for 1h30 at 37 °C before cell lysis or cell adhesion assay, or incubated (micropatterns) with these inhibitors for 4 h at 37 °C before fixation.
Additional M&M can be found in Supplementary File.
Results
ERRα regulates actin dynamics
To investigate the potential role of ERRα in the regulation of actin polymerisation, we first performed differential sedimentation of actin filaments (F-actin) and actin monomers (G-actin) using ultracentrifugation. We observed that inactivation of ERRα with a specific siRNA induced a significant increase in F-actin level with stable G-actin level in both MDA-MB231 cells (Fig. 1A) and HeLa cells (Supplementary Fig. S1A). Immunofluorescence experiments showed that actin filaments are highly concentrated at the cell periphery in both the control and ERRα-depleted conditions. Interestingly, these actin filaments seemed to be particularly disorganised in the ERRα-depleted cells (Fig. 1B). In addition, quantitative image analysis revealed that the F-actin content significantly increased upon ERRα depletion (Fig. 1C).
A F-actin and G-actin from MDA-MB231 control and ERRα-depleted cells were segmented by ultra-speed centrifugation and analysed by western blot. Only one aliquot of each fraction was analysed by SDS-PAGE, which corresponded to 1% and 20% of the total volume of G-actin and F-actin fractions, respectively. Quantifications of F-actin and G-actin are relative to total actin level and control conditions and representative of three independent experiments. B F-actin was stained using phalloïdin (red) in control and ERRα-depleted cells. Nuclei are shown in blue. The lower panels show a high magnification of the boxed regions in the image above. Scale bars: 20 μm. C F-actin intensity was measured using ImageJ. D Triangle-shaped micropatterns were coated with 1,5 μg/cm2 of collagen I. Control or siERRα-transfected cells were then seeded onto micropatterns, F-actin was stained with SiR-actin (red) and nuclei were stained with Hoechst (blue). Scale bars: 10 μm. E F-actin intensity was measured using ImageJ. Box-and-whisker plots are representative of 4 (C) or 9 (E) independent experiments. Mann–Whitney test, ****p < 0.0001, n ≥ 25–30 cells per condition.
In order to further analyse defects in actin filaments associated with ERRα depletion, we used triangle-shaped micropatterns. When control cells adhered on these micropatterns, they spread and acquired a triangular shape. Actin filaments accumulated in these cells at the lateral edges of the triangle. By contrast, ERRα-depleted cells displayed a random localisation of the actin filaments and a strongly altered triangular morphology (Fig. 1D). Once again, depletion of ERRα induced an increase in the F-actin content as compared to the control condition (Fig. 1E). Taken together, these results demonstrate that ERRα regulates polymerisation of actin filaments.
ERRα acts on the RhoA-ROCK pathway to modulate actin polymerisation
The small GTPase protein RhoA plays a major role in regulating the organisation of the actin cytoskeleton through its effectors mDIA and ROCK [12, 14]. Downstream of ROCK, the activation of LIMK results in cofilin inactivation and consecutive increase in actin polymerisation [13, 14]. We previously showed that ERRα regulates cell migration by modulating RhoA protein expression and activation of the RhoA-ROCK pathway [22]. We, therefore, investigated if the effects of ERRα on the actin cytoskeleton could result from LIMK-dependent cofilin inhibition. Western blot experiments showed that depletion of ERRα strongly increased the level of the inactive phosphorylated form of cofilin, whereas total levels of cofilin remained unchanged (Fig. 2A). The level of RhoA also strongly increased in ERRα-depleted cells, as expected (Fig. 2B and Supplementary Fig. S1B). In addition, treatment with the selective ROCK inhibitor Y27632 decreased the phosphorylation level of cofilin in both control and ERRα-depleted cells (Fig. 2C). Interestingly, this treatment was able to put the phosphorylation level of cofilin in ERRα-depleted cells close to the one observed in control cells. Furthermore, the specific LIMK inhibitor Pyr1 similarly rescued the phosphorylation level of cofilin in these cells (Fig. 2D). These findings indicate that the RhoA-ROCK-LIMK pathway is involved in ERRα-mediated controls of cofilin activity. We next investigated whether the deregulation of this pathway may account for the defective actin regulation observed in ERRα-depleted cells. Treating micropatterned cells with 5 μM Y27632 rescued, at least partially, the increase in F-actin intensity induced by ERR depletion (Fig. 2E), indicating that the effect of the receptor on the RhoA-ROCK cascade is instrumental in regulating the F-actin content.
A Expression of phosphorylated cofilin (P-cofilin), total cofilin, and B RhoA was analysed by western blot after MDA-MB231 cell transfection with control or ERRα siRNAs. C, D Control or ERRα-depleted cells were treated either with Y27632 or Pyr1 as indicated, and subjected to western blot for analysis of P-cofilin and cofilin expression. A, C, D Quantifications indicate the ratio of P-cofilin/cofilin. A–D Hsp90 is used as a loading control and as the reference for quantification relative to control conditions. Quantifications are representative of three independent experiments. E Control or ERRα-depleted cells were seeded onto triangle-shaped micropatterns pre-coated with 1,5 μg/cm2 of collagen I and treated with 5 μM Y27632. F-actin was stained with SiR-actin (red), and intensity was quantified using ImageJ. The box-and-whisker plot is representative of 5 independent experiments. Kruskal–Wallis with Dunn’s multiple comparisons test, ns (not significant) for p > 0.05, *p < 0.05 and ****p < 0.0001, n ≥ 25–30 cells per condition.
ERRα regulates cell adhesion
Actin filament- and FA dynamics are tightly linked. Notably, Rho GTPases and actin dynamics play a crucial role in regulating FA maturation and turnover [39,40,41]. Therefore, we determined whether ERRα is able to regulate cell adhesion through its action on the RhoA pathway. Using the xCELLigence system, which allows real-time monitoring of cell adhesion [42], we first showed that depletion of endogenous ERRα resulted in a significant decrease in cell adhesion to collagen I compared to control condition (Fig. 3A). Similar effects were observed after depletion of ERRα in HeLa cells (Supplementary Fig. S1C). We also observed that cell adhesion to collagen IV or fibronectin decreased upon ERRα silencing (Fig. 3B). By contrast, control and ERRα-depleted cells were not able to adhere on the positively charged poly-L-lysine substrate, showing that the adhesion of MDA-MB231 cells does not rely on electrostatic interactions (Supplementary Fig. S1D). To investigate whether the upregulation of the RhoA pathway due to ERRα depletion could lead to defective cell adhesion, we then tested the effect of the ROCK inhibitor Y27632 on the adhesion of ERRα-depleted cells. Y27632 treatment exacerbated, rather than rescued, the adhesion defect of ERRα-depleted cells. It also induced a decrease, albeit moderate, of cell adhesion under control conditions (Fig. 3C). Treatment with the LIMK inhibitor Pyr1 also markedly reduced adhesion of the control and ERRα-depleted cells (Supplementary Fig. S1E). Notably, RhoA stimulates actomyosin contractility through ROCK that is required for cell adhesion [8]. We thus tested the possibility that abnormal actomyosin contractility resulting from ERRα depletion may impact this process. We found that the specific myosin II inhibitor blebbistatin did not change the adhesion ability of ERRα-depleted cells but slightly reduced adhesion of the control cells (Supplementary Fig. S1F). Taken together, these observations suggest that a correct level of activation of RhoA pathway and a precise control of the actomyosin contractility contribute to optimal cell adhesion in control conditions. Our result further indicate that the RhoA-ROCK pathway is not involved in ERRα-mediated regulation of cell adhesion.
Cell–matrix adhesion was analysed after siRNA inhibition of ERRα expression using the xCELLigence system, which measure electrical impedance induced by cells across microelectrodes integrated on the bottom of 16-well culture plates (E-plate). The impedance signal is proportional to the intensity of the interactions exerted by the cells on the substrate. A E-plates were coated with 1.5 μg/cm2 of collagen I. Control or ERRα-depleted cells were then seeded in E-plate for measurement of impedance, represented by the cell index (left panel) and the slope (right panel). Adhesion phase slopes, indicated as ‘Relative cell adhesion’, were calculated from the linear phase in a specific interval of time (blue and red arrowheads, left panel). B E-plates were coated with 1.5 μg/cm2 of fibronectin or collagen IV. Control or ERRα-depleted cells were then seeded in E-plate for measurement of impedance. C Control or ERRα-depleted cells were treated with 5 μM Y27632 and seeded in E-plates pre-coated with 1.5 μg/cm2 of collagen I for measurement of impedance. A–C Results are shown as mean ± SEM of three or four independent experiments performed in quadruplicate. 2-way ANOVA with Dunnett’s multiple comparisons, ns (not significant) for p > 0.05, *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
FAs represent the major sites of cell attachment to the ECM [1, 5]. Therefore, to determine how ERRα impacts on cell adhesion, we analysed FAs using vinculin as a marker. Immunofluorescence microscopy showed that FAs appeared smaller in ERRα-depleted cells as compared to control cells (Fig. 4A). Quantitative analysis of vinculin staining revealed indeed a significant decrease of FA area and length upon ERRα depletion (Fig. 4B, C). The distance of FAs to the cell periphery was also impaired in these cells, reflecting FA mislocalization (Fig. 4D). To determine the potential role of ERRα in FA dynamics, we next used MDA-MB231 cells stably expressing GFP-paxillin, a fluorescent FA marker protein. As for their wild type counterparts, adhesion of these cells to collagen I decreased upon ERRα depletion, demonstrating that the GFP tag did not compromise ERRα involvement in cell adhesion (Supplementary Fig. S3A). We performed live-cell imaging and we observed that FAs displayed more rapid phases of assembly and disassembly in ERRα-depleted cells as compared to control cells. Representative examples of these perturbations in FA dynamics are shown in montages in Fig. 4E (red arrows). Quantification of the kinetics of individual FA demonstrated that depletion of ERRα resulted in a significant increase in both the assembly and disassembly rates of FAs (Fig. 4F). Altogether, these data indicate that ERRα promotes cell adhesion by modulation of FA formation and turnover.
A MDA-MB231 control or ERRα-depleted cells were fixed and stained with vinculin for FAs (red) and Hoechst for nuclei (blue). Scale bars: 20 μm. B Area, C length of FAs and D relative distance to the cell periphery were visualised with vinculin staining, analysed using a Matlab code developed by R. De Mets and M. Balland, and represented by box-and-whisker plots. Data correspond to four (D) or six (B, C) independent experiments. E Representative time-lapse images (montages) of FA dynamic in control or ERRα-depleted cells. Red arrows point to the FAs of interest. Scale bars: 10 μm. F Box-and-whisker plots show the assembly and disassembly rates of FAs in ERRα-depleted cells relative to the control cells quantified with the Focal Adhesion Analysis (FAAS) method. Data are representative of three independent experiments. B–D and F Mann–Whitney test, ****p < 0.0001, n ≥ 25–30 cells and ≥ 470 FA per condition.
ERRα regulates FA dynamics via its transcriptional target MAP4K4
To investigate the molecular mechanisms through which ERRα controls cell adhesion, we examined its transcriptional targets. Transcriptomic and Gene Ontology (GO) analyses have been previously performed to identify ERRα target genes and associated biological functions [22]. Of particular interest, these analyses revealed MAP4K4, which encodes a Ser/Thr kinase involved in the regulation of FA dynamics and cell migration [15, 18, 19]. RT-qPCR experiments verified our finding, showing that silencing of ERRα led to an upregulation of MAP4K4 expression at the mRNA level (Fig. 5A). Examination of publicly available chromatin immunoprecipitation sequencing (ChIP-Seq) data performed on BT-474 cells [43] indicated the recruitment of ERRα on two distinct regions of intron 2 of the MAP4K4 gene, each displaying two putative ERRα response elements (ERREs) (Supplementary Fig. S2A). ChIP-qPCR experiments revealed that ERRα binds these regions in MDA-MB231 cells (Fig. 5B). Next, an upregulation of the MAP4K4 protein expression resulting from ERRα-depletion was confirmed by Western blot (Fig. 5C). Similar results were observed in HeLa and MDA-MB231 GFP-paxillin cells (Supplementary Fig. S2B). Consistently, an enhanced activity of MAP4K4 was observed in ERRα depleted cells, as indicated by an increased phosphorylation of moesin, a substrate of MAP4K4 involved in FA turnover (Fig. 5D) [19]. Together, these data demonstrate that ERRα directly reduces the expression of MAP4K4 and consequently influences its activity. Interestingly, we showed that loss of ERRα reduced the adhesion of BT474 cells on the collagen I substrate (Supplementary Fig. S3B), and increased the expression of the MAP4K4 protein in these cells (Supplementary Fig. S2B), suggesting that similar molecular mechanisms are involved in the adhesion of MDA-MB231, HeLa and BT474 cells.
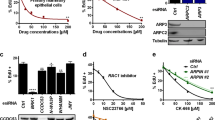
A The expression of MAP4K4 and ERRα genes was analysed by RT-qPCR after transfection with control or ERRα siRNA of MDA-MB231 cells. Data are mean ± SEM of three experiments performed in triplicate. 2-way ANOVA with Dunnett’s multiple comparisons, **p < 0.01 and ****p < 0.0001. B The position of the putative ERRα binding regions (red letters) was indicated relative to the transcriptional start site (TSS) (upper panel). Arrowheads: oligonucleotide primers used in qPCR. Note that the scheme is not to scale. ChIP experiments were performed using anti-ERRα antibody or IgG (lower panel). Percent enrichments relative to input were measured by qPCR, amplifying a region encompassing the putative ERREs for ERRα. Data represent mean ± SEM of two experiments, each in duplicate. Mann–Whitney test, *p < 0.05. nd not detected. Non specific (ns) downstream region was used as a negative binding control. C Expression of MAP4K4 was analysed by western blot in control or ERRα-depleted cells. Quantifications are relative to Hsp90 levels and control conditions and representative of three independent experiments. D Expression of phosphorylated moesin (P-moesin) and total moesin was analysed by western blot. Quantifications indicate the ratio of P-moesin/moesin relative to control conditions and represent four independent experiments.
MAP4K4 promotes FA disassembly by inducing integrin recycling [18] and inactivation [19]. These data raise the possibility that the overactivation of MAP4K4 observed in ERRα-depleted cells may account for the defects of FAs identified in these cells. To investigate this hypothesis, we suppressed endogenous expression of MAP4K4 in MDA-MB231 cells using siRNA targeting and we tested the ability of these cells to adhere to collagen I substrate. Unexpectedly, our results showed that the MAP4K4 knockdown induced a decrease in cell adhesion as compared to control conditions (Supplementary Fig. S3C). These data are contradictory with those of other teams, which demonstrated an increased cell adhesion resulting from siRNA-mediated depletion of MAP4K4 in human umbilical vein endothelial cells or knockout of MAP4K4 in mice keratinocytes [18]. However, it has been suggested that MAP4K4 could play a positive role in cell adhesion through protein interactions mediated by its C-terminal domain [15, 44]. This explains why deletion of MAP4K4 may impinge on the correct adhesion of cells as observed in our conditions. Therefore, since the depletion of MAP4K4 may lead to cell-type-specific effects, we choose to use specific MAP4K4 inhibitors to analyse the potential role of MAP4K4 in ERRα-mediated regulation of cell adhesion. First, cells were treated with the MAP4K4 inhibitor PF-06260933 [45]. We found that a low concentration of PF-06260933 rescued cell adhesion on the collagen I substrate (Fig. 6A) and restored FA area and length in ERRα-depleted cells (Fig. 6B, C). PF-06260933 also nearly completely rescued the relative distance of FAs to the cell periphery, which was impaired in these cells probably due to mislocalized MAP4K4 activity [19], but altered FA localisation in control cells (Fig. 6D). Second, we observed that treatment with another MAP4K4 inhibitor, GNE-495 [46], also rescued adhesion of the ERRα-depleted cells (Supplementary Fig. S3D). Together, these data demonstrate that ERRα regulates cell adhesion through MAP4K4.
A Control or ERRα-depleted cells were treated with 0.5 μM PF-06260933 and seeded in E-plate pre-coated with 1.5 μg/cm2 of collagen I. Then cell adhesion was measured using the xCELLigence system. Data are mean ± SEM of four experiments performed in quadruplicate. 2-way ANOVA with Dunnett’s multiple comparisons, ns (not significant) for p > 0.05, ***p < 0.001 and ****p < 0.0001. B Area C length of FA and D distance to the cell periphery were visualised with vinculin staining in control or ERRα-depleted cells treated with PF-06260933. FA analyses were performed using the Matlab algorithm developed by R. De Mets and M. Balland. The box-and-whisker plots represent data of three independent experiments. E Control or ERRα-depleted cells were seeded onto triangle-shaped micropatterns pre-coated with 1,5 μg/cm2 of collagen I and treated with 0.5 μM PF-06260933. F-actin was stained with SiR-actin (red), and intensity was quantified using ImageJ. Results are presented as a box-and-whisker plot and correspond to three independent experiments. Kruskal–Wallis with Dunn’s multiple comparisons test, ns (not significant) for p > 0.05, *p < 0.05, ***p < 0.001 and ****p < 0.0001, n ≥ 30 cells and ≥ 670 FAs per condition.
As shown above, impacting on the RhoA-ROCK axis in ERRα-depleted cells rescued the defects in actin polymerisation but not the reduced adhesion capacities. We thus examined the converse possibility, questioning whether impacting on the MAP4K4 axis could reduce the increased actin polymerisation observed upon ERRα inactivation. As shown in Fig. 6E, treatment with the MAP4K4 inhibitor PF-06260933 did not rescue the actin status in ERRα-depleted cells but rather increased actin polymerisation in control cells. Altogether our data show that ERRα regulates actin polymerisation and FA dynamics via two independent pathways.
Discussion
ERRα regulates many cellular processes contributing to tumour development and progression. Most of all, ERRα has been largely implicated in inducing migratory and invasive properties of cancer cells [21,22,23, 35, 36]. ERRα also contributes to cell migration under physiological conditions such as morphogenetic movements during gastrulation of the zebrafish embryo and chemotactic migration of activated macrophages [22, 47]. Although some molecular mechanisms through which ERRα promotes cell movements have been described, how these signalling actors are connected to the precise morphological changes required for cell migration per se is still unclear. In this report, we show that ERRα coordinates actin and FA dynamics, through the independent modulation of the RhoA-ROCK-LIMK-cofilin pathway and MAP4K4 activity, respectively.
A proper interaction between cells and ECM is an essential prerequisite for cell migration, and it needs to be precisely regulated. Nascent adhesion complexes recruit actin binding proteins to establish a link between ECM and the actin cytoskeleton [1]. RhoA contributes to FA maturation by controlling the growth of FA-associated actin filaments through the activation of the formin mDIA and inhibition of severing activity of cofilin [7, 13, 14]. RhoA also regulates actin binding to myosin II filaments via ROCK, which subsequently induces contractility required for FA maturation [8, 11]. We previously reported that ERRα depletion significantly increases RhoA expression and activation [22]. We show here that the upregulation of RhoA activity induces an increase in actin polymerisation resulting from an enhanced phosphorylation status of the ROCK downstream target cofilin. This suggests that an excess of actin filaments may impair their interaction with FAs and impact on FA maturation. Unexpectedly, our results reveal that the RhoA-ROCK pathway does not contribute to the regulation of cell adhesion downstream of ERRα. Therefore, global deregulation of RhoA activity due to ERRα depletion does not seem to have a strong influence on the precise control of actin polymerisation at the FA sites.
It has been reported that cofilin and myosin compete for binding to actin filaments [14]. We demonstrate here that upregulation of the RhoA-ROCK pathway leads to an increase in cofilin phosphorylation in the ERRα-depleted cells, which has been shown to inhibit its interaction with actin. As a consequence, depletion or inhibition of cofilin may promote actomyosin assembly [14]. In addition, we previously showed that the overactivation of ROCK resulting from ERRα depletion leads to an increased phosphorylation of MYPT1, one of the subunits of the myosin light chain phosphatase [22]. In addition, an overactivation of ROCK can increase the myosin II ATPase activity through phosphorylation of myosin light chain (MLC) [48]. Altogether these data suggest that ERRα functions as a regulator of actomyosin contractility by controlling the RhoA-ROCK pathway. Therefore, ERRα may modulate the formation of actin stress fibres not only through LIMK and cofilin, but also through myosin II. However, in accordance with the results obtained for the RhoA pathway, we demonstrate that ERRα does not promote cell adhesion through its potential effect on actomyosin filaments.
Upon ERRα silencing, cell adhesion decreases as a result from impaired FA formation and dynamics. These defects can be rescued by down-modulating the activity of MAP4K4. Interestingly, we show that ERRα depletion increases both FA assembly and FA disassembly. This is consistent with a report showing that loss of MAP4K4 exerts the inverse effect on FA dynamics [18]. MAP4K4 has been previously identified as a FA disassembly factor [18, 19]. Nevertheless it has been recently reported that MAP4K4 promotes the activation of β1-integrin and its downstream effector Focal Adhesion Kinase (FAK) [20], suggesting that it may also regulate FA assembly. Therefore, further investigations will be needed to determine the potential contribution of these MAP4K4-dependent mechanisms in the regulation of FA assembly and maturation by ERRα.
MAP4K4 regulates FA dynamics by promoting internalisation and inactivation of β1-integrin [18, 19]. In migrating cells, MAP4K4 is delivered to FA sites through its association with the microtubule end-binding protein EB2 (ending binding 2). MAP4K4 subsequently activates IQSEC1 (IQ motif and SEC7 domain-containing protein 1) and Arf6 to induce FA disassembly and cell migration [18]. Furthermore, MAP4K4 is involved in phosphorylation of moesin, which competes with talin for binding to β1-integrin. This leads to β1-integrin inactivation and FA disassembly [19]. Since we observed the activation of moesin in the ERRα-depleted cells, our results strongly suggest that ERRα reduces FA disassembly through the MAP4K4-moesin pathway. We cannot completely exclude the possibility that ERRα also regulates FA disassembly via the regulation of IQSEC1 and Arf6 activation. However, ERRα is more probably involved in the regulation of integrin inactivation rather than their recycling because surface expression of β1-integrin (and other tested integrins) is not modified upon ERRα depletion (Tribollet and Forcet, unpublished). Furthermore, ERRα promotes cell adhesion to different ECM proteins, suggesting the involvement of the MAP4K4-moesin pathway in the regulation of distinct types of integrins. In line with that observation, the role of talin in activation of multiple integrins been reported [49, 50]. Therefore, it is plausible that ERRα, by inducing moesin competition with talin via MAP4K4, has a more general role in the regulation of integrin activation and FA turnover.
We previously performed a transcriptomic analysis of breast cancer cells that led us to identify MAP4K4 as an ERRα-target gene [22]. This analysis showed that the expression of MAP4K4 increased by about 80% upon ERRα depletion, and also revealed that its expression was decreased but not fully suppressed in the control condition. Thus, we postulate that a correct protein level of MAP4K4 is critical for cell migration. It has been shown that the knockout or the knockdown of MAP4K4 leads to FA stabilisation and impaired or reduced cell migration probably due to a strong attachment of the cells to the substrate [18]. On the contrary, a marked increase in expression of MAP4K4 resulting from ERRα depletion leads to FA disassembly and may result in a reduced capacity of cells to respond to extracellular matrix cues. Consequently, the directionality of cell migration may be impaired rather than the migration process per se [22, 51]. We, therefore, propose that ERRα is important in ensuring a correct level of MAP4K4 necessary for proper oriented cell migration.
A role of MAP4K4 in regulation of cortical actin dynamics has been previously shown [52,53,54]. Notably, these data show that MAP4K4 silencing decreases the accumulation of actin filaments in cell protrusions. Thus, it suggests that the upregulation of MAP4K4 resulting from ERRα depletion may promote aberrant actin polymerisation. However, we demonstrate here that MAP4K4 is not involved in the regulation of the actin network downstream of ERRα. Therefore, we speculate that ERRα-mediated regulation of MAP4K4 expression plays a major role in cell adhesion, but that another upstream signal may be needed to activate MAP4K4-dependent actin polymerisation. In this way, it has been shown that MAP4K4, or its mouse homologue NIK, phosphorylates ARP2 and the ERM proteins to induce actin polymerisation in response to growth factor stimulation [44, 52, 53]. Altogether, our data firmly demonstrate that ERRα coordinates actin polymerisation and adhesion via two independent pathways.
Both ERRα and MAP4K4 have been shown to be highly expressed in cancers [15, 23]. Our data demonstrate here that ERRα is a key transcriptional regulator of the MAP4K4 gene. However, epigenetic modifications and other signalling pathways also appear to be involved in the modulation of MAP4K4 expression [15]. In these conditions, compensation mechanisms may bypass the MAP4K4-induced cell adhesion defects to facilitate oriented migration of cancer cells. Thus, this study revealed that the modulation of cell adhesion exerted by ERRα through MAP4K4 is one specific way to control breast cancer cell migration.
In conclusion, we report that ERRα modulates actin polymerisation through the RhoA-ROCK axis and FA formation and turnover through the MAP4K4 pathway. As a consequence, deregulation of ERRα expression deeply impacts cell adhesion and cell morphology, pointing to a critical role played by ERRα in cancer cell migration.
Data availability
All relevant data can be found within the published article and its supplementary files. FA quantification was done using a Matlab (MathWorks, Natick, MA) algorithm developed by M. Balland and R. De Mets, and the corresponding source code can be found in GitHub: https://github.com/rdemets/FA_Quantif_Matlab.
Code availability
https://github.com/rdemets/FA_Quantif_MatlabThe source code of the Focal Adhesion Analysis Server developed by M Berginski [37, 38] is also available in GitHub: https://github.com/mbergins/.
References
Gardel ML, Schneider IC, Aratyn-Schaus Y, Waterman CM. Mechanical integration of actin and adhesion dynamics in cell migration. Annu Rev Cell Dev Biol. 2010;26:315–33.
Bravo-Cordero JJ, Hodgson L, Condeelis J. Directed cell invasion and migration during metastasis. Curr Opin Cell Biol. 2012;24:277–83.
Ridley AJ. Cell migration: integrating signals from front to back. Science. 2003;302:1704–9.
Blanchoin L, Boujemaa-Paterski R, Sykes C, Plastino J. Actin dynamics, architecture, and mechanics in cell motility. Physiol Rev. 2014;94:235–63.
De Pascalis C, Etienne-Manneville S. Single and collective cell migration: the mechanics of adhesions. Mol Biol Cell. 2017;28:1833–46.
Ridley AJ. Rho GTPase signalling in cell migration. Curr Opin Cell Biol. 2015;36:103–12.
Lawson CD, Ridley AJ. Rho GTPase signaling complexes in cell migration and invasion. J Cell Biol. 2018;217:447–57.
Guan X, Guan X, Dong C, Jiao Z. Rho GTPases and related signaling complexes in cell migration and invasion. Exp Cell Res. 2020;388:111824.
Machacek M, Hodgson L, Welch C, Elliott H, Pertz O, Nalbant P, et al. Coordination of Rho GTPase activities during cell protrusion. Nature. 2009;461:99–103.
Martin K, Reimann A, Fritz RD, Ryu H, Jeon NL, Pertz O. Spatio-temporal co-ordination of RhoA, Rac1 and Cdc42 activation during prototypical edge protrusion and retraction dynamics. Sci Rep. 2016;6:1–14.
Seetharaman S, Etienne-Manneville S. Microtubules at focal adhesions – a double-edged sword. J Cell Sci. 2019;132:jcs232843.
Spiering D, Hodgson L. Dynamics of the Rho-family small GTPases in actin regulation and motility. Cell Adhes Migr. 2011;5:170–80.
Mizuno K. Signaling mechanisms and functional roles of cofilin phosphorylation and dephosphorylation. Cell Signal. 2013;25:457–69.
Kanellos G, Frame MC. Cellular functions of the ADF/cofilin family at a glance. J Cell Sci. 2016;129:3211–8.
Gao X, Gao C, Liu G, Hu J. MAP4K4: an emerging therapeutic target in cancer. Cell Biosci. 2016;6:56.
Tripolitsioti D, Grotzer MA, Baumgartner M. The Ser/Thr kinase MAP4K4 controls pro-metastatic cell functions. J Carcinog Mutagen. 2017;8:2.
Stehbens S, Wittmann T. Targeting and transport: how microtubules control focal adhesion dynamics. J Cell Biol. 2012;198:481–9.
Yue J, Xie M, Gou X, Lee P, Schneider MD, Wu X. Microtubules regulate focal adhesion dynamics through MAP4K4. Dev Cell. 2014;31:572–85.
Vitorino P, Yeung S, Crow A, Bakke J, Smyczek T, West K, et al. MAP4K4 regulates integrin-FERM binding to control endothelial cell motility. Nature. 2015;519:425–30.
Tripolitsioti D, Kumar KS, Neve A, Migliavacca J, Capdeville C, Rushing EJ, et al. MAP4K4 controlled integrin β1 activation and c-Met endocytosis are associated with invasive behavior of medulloblastoma cells. Oncotarget. 2018;9:23220–36.
Dwyer MA, Joseph JD, Wade HE, Eaton ML, Kunder RS, Kazmin D, et al. WNT11 expression is induced by estrogen-related receptor alpha and beta-catenin and acts in an autocrine manner to increase cancer cell migration. Cancer Res. 2010;70:9298–308.
Sailland J, Tribollet V, Forcet C, Billon C, Barenton B, Carnesecchi J, et al. Estrogen-related receptor α decreases RHOA stability to induce orientated cell migration. Proc Natl Acad Sci. 2014;111:15108–13.
Tam IS, Giguère V. There and back again: The journey of the estrogen-related receptors in the cancer realm. J Steroid Biochem Mol Biol. 2016;157:13–9.
Horard B, Vanacker J-M. Estrogen receptor-related receptors: orphan receptors desperately seeking a ligand. J Mol Endocrinol. 2003;31:349–57.
Ranhotra HS. The estrogen-related receptors in metabolism and cancer: newer insights. J Recept Signal Transduct. 2018;38:95–100.
Ranhotra HS. Estrogen-related receptor alpha and cancer: axis of evil. J Recept Signal Transduct. 2015;35:505–8.
Chang C, Kazmin D, Jasper JS, Kunder R, Zuercher WJ, McDonnell DP. The metabolic regulator ERRα, a downstream target of HER2/IGF-1, as a therapeutic target in breast cancer. Cancer Cell. 2011;20:500–10.
Bianco S, Sailland J, Vanacker J-M. ERRs and cancers: Effects on metabolism and on proliferation and migration capacities. J Steroid Biochem Mol Biol. 2012;130:180–5.
Ao A, Wang H, Kamarajugadda S, Lu J. Involvement of estrogen-related receptors in transcriptional response to hypoxia and growth of solid tumors. Proc Natl Acad Sci. 2008;105:7821–6.
Zou C, Yu S, Xu Z, Wu D, Ng C-F, Yao X, et al. ERRα augments HIF-1 signalling by directly interacting with HIF-1α in normoxic and hypoxic prostate cancer cells. J Pathol. 2014;233:61–73.
Stein RA, Gaillard S, McDonnell DP. Estrogen-related receptor alpha induces the expression of vascular endothelial growth factor in breast cancer cells. J Steroid Biochem Mol Biol. 2009;114:106–12.
Zhang K, Lu J, Mori T, Smith-Powell L, Synold TW, Chen S, et al. Baicalin increases VEGF expression and angiogenesis by activating the ERR{alpha}/PGC-1{alpha} pathway. Cardiovasc Res. 2011;89:426–35.
Tennessen JM, Baker KD, Lam G, Evans J, Thummel CS. The Drosophila estrogen-related receptor directs a metabolic switch that supports developmental growth. Cell Metab. 2011;13:139–48.
Cai Q, Lin T, Kamarajugadda S, Lu J. Regulation of glycolysis and the Warburg effect by estrogen-related receptors. Oncogene. 2013;32:2079–86.
Carnesecchi J, Forcet C, Zhang L, Tribollet V, Barenton B, Boudra R, et al. ERRα induces H3K9 demethylation by LSD1 to promote cell invasion. Proc Natl Acad Sci. 2017;114:3909–14.
Zhang L, Carnesecchi J, Cerutti C, Tribollet V, Périan S, Forcet C, et al. LSD1-ERRα complex requires NRF1 to positively regulate transcription and cell invasion. Sci Rep. 2018;8:10041.
Berginski ME, Vitriol EA, Hahn KM, Gomez SM. High-resolution quantification of focal adhesion spatiotemporal dynamics in living cells. PLoS ONE. 2011;6:e22025.
Berginski ME, Gomez SM. The Focal Adhesion Analysis Server: a web tool for analyzing focal adhesion dynamics. F1000Research. 2013;2:68.
Vicente-Manzanares M, Choi CK, Horwitz AR. Integrins in cell migration – the actin connection. J Cell Sci. 2009;122:199–206.
Juanes MA, Bouguenina H, Eskin JA, Jaiswal R, Badache A, Goode BL. Adenomatous polyposis coli nucleates actin assembly to drive cell migration and microtubule-induced focal adhesion turnover. J Cell Biol. 2017;216:2859–75.
Romero S, Le Clainche C, Gautreau AM. Actin polymerization downstream of integrins: signaling pathways and mechanotransduction. Biochem J. 2020;477:1–21.
Agaësse G, Barbollat-Boutrand L, Sulpice E, Bhajun R, Kharbili ME, Berthier-Vergnes O, et al. A large-scale RNAi screen identifies LCMR1 as a critical regulator of Tspan8-mediated melanoma invasion. Oncogene. 2017;36:446–57.
Deblois G, Smith HW, Tam IS, Gravel S-P, Caron M, Savage P, et al. ERRα mediates metabolic adaptations driving lapatinib resistance in breast cancer. Nat Commun. 2016;7:12156.
Baumgartner M, Sillman AL, Blackwood EM, Srivastava J, Madson N, Schilling JW, et al. The Nck-interacting kinase phosphorylates ERM proteins for formation of lamellipodium by growth factors. Proc Natl Acad Sci. 2006;103:13391–6.
Ammirati M, Bagley SW, Bhattacharya SK, Buckbinder L, Carlo AA, Conrad R, et al. Discovery of an in vivo tool to establish proof-of-concept for MAP4K4-based antidiabetic treatment. ACS Med Chem Lett. 2015;6:1128–33.
Ndubaku CO, Crawford TD, Chen H, Boggs JW, Drobnick J, Harris SF, et al. Structure-based design of GNE-495, a potent and selective MAP4K4 inhibitor with efficacy in retinal angiogenesis. ACS Med Chem Lett. 2015;6:913–8.
Bardet P-L, Horard B, Laudet V, Vanacker J-M. The ERRα orphan nuclear receptor controls morphogenetic movements during zebrafish gastrulation. Dev Biol. 2005;281:102–11.
Amano M, Nakayama M, Kaibuchi K. Rho-Kinase/ROCK: a key regulator of the cytoskeleton and cell polarity. Cytoskelet Hoboken Nj. 2010;67:545–54.
Klapholz B, Brown NH. Talin – the master of integrin adhesions. J Cell Sci. 2017;130:2435–46.
Sun Z, Costell M, Fässler R. Integrin activation by talin, kindlin and mechanical forces. Nat Cell Biol. 2019;21:25–31.
Petrie RJ, Doyle AD, Yamada KM. Random versus directionally persistent cell migration. Nat Rev Mol Cell Biol. 2009;10:538–49.
Ma M, Baumgartner M. Intracellular theileria annulata promote invasive cell motility through kinase regulation of the host actin cytoskeleton. PLoS Pathog. 2014;10:e1004003.
LeClaire LL, Rana M, Baumgartner M, Barber DL. The Nck-interacting kinase NIK increases Arp2/3 complex activity by phosphorylating the Arp2 subunit. J Cell Biol. 2015;208:161–70.
Santhana Kumar K, Tripolitsioti D, Ma M, Grählert J, Egli KB, Fiaschetti G, et al. The Ser/Thr kinase MAP4K4 drives c-Met-induced motility and invasiveness in a cell-based model of SHH medulloblastoma. SpringerPlus. 2015;4:19.
Acknowledgements
The authors warmly thank members of the Vanacker lab for support and discussion, as well as Séverine Périan for technical assistance. We are grateful to Sandrine Etienne-Manneville (Institut Pasteur, Paris) and Laurence Lafanechère (Institute for Advanced Biosciences, Grenoble) for reagents. We also thank the staff of PLATIM (UMS3444/CNRS, US8/INSERM, ENS de Lyon, UCBL) and IGFL microscopy facilities for their precious help with microscopy studies.
Funding
Work in our laboratory is funded by Ligue contre le Cancer (comité Rhône), Région Auvergne Rhône Alpes (grant SCUSI OPE2017_004), ANSES (grant EST15-076), and ENS Lyon (programme JoRISS).
Author information
Authors and Affiliations
Contributions
VT, JMV and CF were responsible for designing research; VT, CC, EDB, JC and CF performed research; RDM, MB, and AG contributed to analytic tools; VT, CC, JMV and CF extracted, analysed data and interpreted results; and CF wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tribollet, V., Cerutti, C., Géloën, A. et al. ERRα coordinates actin and focal adhesion dynamics. Cancer Gene Ther 29, 1429–1438 (2022). https://doi.org/10.1038/s41417-022-00461-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-022-00461-6
- Springer Nature America, Inc.
This article is cited by
-
ERRα: unraveling its role as a key player in cell migration
Oncogene (2024)
-
Mdm2 requires Sprouty4 to regulate focal adhesion formation and metastasis independent of p53
Nature Communications (2024)