Abstract
Background
Genetic correlations, causalities and pathways between large-scale complex exposures and ovarian and breast cancers need systematic exploration.
Methods
Mendelian randomisation (MR) and genetic correlation (GC) were used to identify causal biomarkers from 95 cancer-related exposures for risk of breast cancer [BC: oestrogen receptor-positive (ER + BC) and oestrogen receptor-negative (ER − BC) subtypes] and ovarian cancer [OC: high-grade serous (HGSOC), low-grade serous, invasive mucinous (IMOC), endometrioid (EOC) and clear cell (CCOC) subtypes].
Results
Of 31 identified robust risk factors, 16 were new causal biomarkers for BC and OC. Body mass index (BMI), body fat mass (BFM), comparative body size at age 10 (CBS-10), waist circumference (WC) and education attainment were shared risk factors for overall BC and OC. Childhood obesity, BMI, CBS-10, WC, schizophrenia and age at menopause were significantly associated with ER + BC and ER − BC. Omega-6:omega-3 fatty acids, body fat-free mass and basal metabolic rate were positively associated with CCOC and EOC; BFM, linoleic acid, omega-6 fatty acids, CBS-10 and birth weight were significantly associated with IMOC; and body fat percentage, BFM and adiponectin were significantly associated with HGSOC. Both GC and MR identified 13 shared factors. Factors were stratified into five priority levels, and visual causal networks were constructed for future interventions.
Conclusions
With analysis of large-scale exposures for breast and ovarian cancers, causalities, genetic correlations, shared or specific factors, risk factor priority and causal pathways and networks were identified.
Similar content being viewed by others
Background
Cancer is an important cause of worldwide morbidity and mortality. Breast cancer (BC) accounted for approximately one in four cancer cases among women in 2018 [1], and ovarian cancer (OC) is the second most common form and leading cause of death due to cancer in the female reproductive system [2]. Although hereditary factors can explain 5–10% of the risk for breast or ovarian cancer [1], non-hereditary factors remain the major drivers. One-third to two-fifths of new cancer cases could be avoided by eliminating or reducing exposure to known risk factors [1, 3,4,5]. Thus, primary preventive measures that can reduce risk of BC and OC by targeting intervention on complex causal biomarkers or pathways are of increasing interest.
Unobserved confounding and reverse causality limit the ability of epidemiological observational studies to identify causalities. Observational studies are also often limited in acquiring large-scale exposure measurements, whereas randomised clinical trials are not widely available because of ethical concerns, high cost and long duration [6]. Recently, thousands of summary-level statistics from genome-wide association studies (GWASs) have provided great opportunities to support wide-range causal findings. Mendelian randomisation (MR) uses genome-wide significant genetic variants as instrumental variables (IVs) that can accurately assess the causal effect and direction of one exposure on a specific outcome after ruling out unobserved confounders in theory (“causal” represents the causality in statistics). Genetic correlation (GC) analysis can identify the common genetic risk between two specific traits [7]. Identifying GCs can provide useful etiological insights and help prioritise likely causal relationships [7]. Combining GC with MR can be used to identify direct causal relations and shared genetic risks for an exposure–outcome pair.
To date, MR has been used to explore the effects of alcohol consumption and glycemic, lipids and obesity traits on ovarian and breast cancers [6, 8,9,10,11]. Although many causal biomarkers have been identified, focusing on a specific exposure can provide only limited evidence for primary prevention, compared with focusing on a wide range of potential biomarkers. In addition, genome-wide correlations between BC and OC and complex exposures remain unclear. There is intense interest identifying these associations because the shared genetic architecture and causal associations can provide valuable references for joint, precise and priority intervention targets. Although information has been obtained on many risk factors, prioritising these factors remains essential. Complex network pathways may also occur that lead from extensive exposures to the occurrence of cancer, which may involve mediators as potential targets. Si et al. used network-MR to study biomarkers and complex metabolic pathways [12]. With a network, upstream or downstream targets for a specific risk factor can be identified and also contribute to primary prevention.
In this study, large-scale genomic summary-level statistics were used in a comprehensive analysis to gain insight into the complex relations of 95 cancer-related factors and nine cancer types. The aim of the research was to screen for robust causal biomarkers of BC and OC and then identify shared or distinct risk factors, stratify risks by priority and construct visual causal networks to guide prevention measures.
Methods
Determination of exposures
Risk factors associated with human cancers were hypothesised to also have potential carcinogenic mechanisms in the occurrence of breast and ovarian cancers. To determine candidate exposures, all MR analyses (Supplementary Table S1) were reviewed for any cancer types, and only the factors with available summary-level datasets were used in this research. Details about the process are described in Supplementary Text 1. Ultimately, 95 complex traits were included in the study. Except for the established birth length (BIRL), birth weight (BIRW) and age at menarche (AAM), most factors could be modified. Data sources of the 95 exposures are listed in Supplementary Table S2.
Among the factors, literature review showed that only 32 factors had been explored for association with both BC and OC, only 18 for BC and only 1 for OC. The other 44 exposures have not been studied previously in BC and OC (Supplementary Table S2). Previous MR studies have not explored GCs, risk stratification and network pathways for these risk factors on BC or OC. Because of the lack of information, the exposures were examined as candidate factors for BC and OC in this research [13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48]. The 95 factors were further divided into the following categories: anthropometry traits (16), blood biochemistry traits (8), disease traits (5), lifestyle traits (14), lipids/glycemic traits (11), metabolites (13), nutrients (23) and sex-related traits (5) (see Supplementary Text 1 for details).
Data sources of breast and ovarian cancers
Summary statistics for 122,977 cases of breast cancer and 105,974 controls of European ancestry were acquired from a combined study including the Breast Cancer Association Consortium (BCAC), Discovery, Biology and Risk of Inherited Variants in Breast Cancer Consortium (DRIVE), Collaborative Oncological Gene-environment Study (iCOGS) and several other GWAS meta-analyses [49]. Summary statistics of two BC subtypes, oestrogen receptor-positive (ER + BC, 69,501 cases) and oestrogen receptor-negative (ER − BC, 21,468 cases), were also included.
Genetic associations with OC were obtained from the Ovarian Cancer Association Consortium using an Illumina Custom Infinium array (OncoArray) including 25,509 epithelial OC cases and 40,941 controls [50]. The OC cases were further divided into five major invasive histotypes: high-grade serous OC (HGSOC, 13,037 cases), low-grade serous OC (LGSOC, 1012 cases), invasive mucinous OC (IMOC, 1417 cases), endometrioid OC (EOC, 2810 cases) and clear cell OC (CCOC, 1366 cases). Detailed information about BC and OC research populations and sample sizes can be found in Supplementary Text 2 and Supplementary Table S3, as well as the original publications [49, 50].
Statistical analysis
Two-sample Mendelian randomization
The study graph model and flowchart are shown in Fig. 1. The univariate two-sample MR method was used to determine the causal effect of each exposure on a target outcome (Fig. 1a). In the MR analysis, genome-wide significant genetic variants were used as instrumental variables (IVs) to examine causal associations of exposures with OC or BC. The MR approach assumes that IVs (1) are associated with the candidate exposure, (2) are not associated with confounders (upper red cross) and (3) are associated with the outcome only through the candidate exposure but not through other pathways (lower red cross) [51].
Panel a was the causal graph of Mendelian randomization (MR), this method should satisfy three assumptions: (1) the IV is associated with the exposure, (2) the IV affects outcome only through the exposure (lower red cross), (3) the IV is not associated with the confounders (upper red cross). Panel b was a framework of multivariable MR, which used a pairwise design to measure exposure Xm–Xn pair on the risk of the outcome. The causal effects of candidate exposure Xn on outcome were calculated in turns. Panel c showed the framework of genetic correlation, the correlation between factors (Xn) and ovarian/breast cancer was measured. Panel d indicated a network-MR framework. This framework utilised the IVs of Xm to acquire the effect of Xm on Xn (potential mediator) and outcome, then, the IVs of Xn were utilised to acquire the effect of Xn on the outcome. Panel e was the flowchart of these analyses. LDSC trait linkage disequilibrium score regression, OC ovarian cancer, BC breast cancer, MR Mendelian randomization, IVW inverse variance weighted, WME weighted median estimator, MVMR multivariable Mendelian randomization.
Candidate IVs associated with a specific risk factor were determined at the standard threshold of genome-wide significance (P < 5 × 10−8). Furthermore, candidate IVs within the threshold of linkage disequilibrium (LD, r2 < 0.01) were pruned to keep nearly independent IVs. The proportion of variance in the exposure explained by the IVs (R2) was calculated as 2β2 × MAF × (1 − MAF), where β is the association of single-nucleotide polymorphisms (SNP) with the exposure and MAF is the minor allele frequency. The F statistic was calculated from the R2 statistic as F = [(N − K − 1) / K] × [R2 / (1 − R2)], where N is the sample size and K is the number of IVs [52]. Generally, an F value > 10 indicated a strong IV [12, 53].
To acquire the causal estimator, first, the Wald ratio (WRO) was used to estimate causal effects for each SNP. Then, the conventional inverse variance weighted (IVW) MR method was used to aggregate causal estimators of all SNPs for the principal analyses in this research [54]. Furthermore, the weighted median estimator (WME) method was applied simultaneously as one of the sensitivity analyses to assess the robustness of causal findings. The method produces robust estimates in the presence of some invalid genetic instruments (when the number of invalid IVs < 50%) [55]. For those exposures with one IV, only WRO results were reported. Additionally, testing for the intercept of the MR-Egger regression was used to assess horizontal pleiotropy. Significant results in both IVW and WME analysis were viewed as robust associations in this research.
Pairwise multivariate Mendelian randomization
The multivariable MR (MVMR) approach [56] is designed to assess the robustness of causality under the possible horizontal pleiotropy and acquire the direct effects of the interested exposure on the outcome. Currently, a consistent standard for covariates selection from large-scale exposures has not been determined. Because adjusting more covariates causes a sharp decline in the number of IVs, and to avoid unknown bias caused by overadjustment, the causal effect of a specific exposure was assessed by adjusting other exposures one by one (here, pairwise MVMR) (Fig. 1b). The exposures (or covariates) that were selected from the above univariate MR analysis were significant for BC, OC or their subtypes. Adjusted effects of the exposures were estimated by the standard IVW–MVMR method [56]. The pairwise MVMR adjusted for a wide range of covariates was also considered a sensitivity analysis in this research.
Genetic correlation
Cross-trait linkage disequilibrium score regression (LDSC) is a useful epidemiological tool to estimate the GC of two traits (Fig. 1c) [7, 57]. An LDSC analysis can rapidly screen for correlations among a diverse set of traits, without needing to measure multiple traits on the same individuals [7]. Genetic variants used in LDSC usually required whole genome-wide SNPs. To keep a consistent number of SNPs for traits from different consortiums, they were matched with a common SNP list that was used in previous work [58]. The SNP list was a file called “w_hm3.noMHC.snplist” that included ~1.2 million SNPs based on the HapMap 3 reference panel stored in “ldsc” software. It can also be download from the LD Hub website (http://ldsc.broadinstitute.org). In addition, LD Hub website also recommend using this SNP list to reduce the number of SNPs to improve computing performance. In this research, genetic variants of the candidate traits were extracted from the MR Base (https://www.mrbase.org/) or the corresponding consortiums based on the SNP list to retain the common 1.2 million recommended SNPs. Then, the direction of effect values (Z-values) of each SNP across all traits was adjusted to ensure they corresponded to consistent-effect alleles. In GC analysis, the genetic dataset of Europeans from 1000 Genomes was used as a reference to compute LD scores. The genetic correlation coefficient was termed rg, which ranged from −1 to 1. Details about this method are introduced elsewhere [7].
Network Mendelian randomization
Network-MR was used to investigate the intermediate phenotypes in causal pathways to help to construct causal networks from the detected risk factors to outcomes (Fig. 1d) [59]. Network-MR was based on the univariate MR approach to achieve point-by-point analyses for each component (exposure, mediator and outcome), which were robust biomarkers (also include the results of WRO for biomarkers with only one IV) in the principal MR analysis. The initial network-MR framework was composed of three separate two-sample MRs that included pairwise analysis of one trait on another trait, as described elsewhere [12]. The network was organised according to the following steps: (1) keep all significant factors (robust association) of BC and OC (or subtypes) from the univariate MR analysis, (2) perform MR analysis for these factors with one another, (3) select the robust associations (both IVW and WME, or WRO) among the factors and (4) connect the significant exposure–exposure pairs and the previous exposure–outcome pairs to construct the final network.
Study procedures
The study flowchart is shown in Fig. 1e. First, the univariable MR method (including IVW, WME and WRO methods) was used to determine the causal effect of each exposure on a target outcome. Simultaneously, cross-trait LDSC was used to measure GCs across OC, BC and large-scale exposures. Then, pairwise MVMR was used to detect potential pleiotropy. Furthermore, the shared and specific risk factors for the outcomes were summarised, and the priority and rank of risk factors were defined. Finally, according to the network-MR design, causal networks for different outcomes were constructed to guide prevention practice.
Risk factors were prioritised at five levels: (1) level 1, robust MR evidence for both BC and OC plus GC evidence; (2) level 2, only robust MR results for both BC and OC; (3) level 3, robust MR evidence plus GC evidence for either BC or OC; (4) level 4, factors only robustly associated with BC or OC but without GC and (5) level 5, remaining MR evidence (only significant in one method), which was suggestive. In addition, a score was defined for each factor to indicate its importance, and then, all significant factors were ranked according to their importance. The score was acquired by adding the number of times a specific exposure was significant for the nine outcomes in the three univariate MR methods and the GC evidence. The level of a factor and its rank indicated the degree of robustness and universality, respectively.
All MR results were reported as odds ratios (ORs) and 95% confidence intervals (CIs) for genetically predicted per standard deviation or per unit increment of each risk factor. Bonferroni-corrected P-values (two-tailed) were used to show the significance of multiple testing. Two Bonferroni thresholds were used. The “moderate” one was P < 5.26 × 10−4 (0.05/95), which considered only the number of candidate exposures, whereas the “strict” one was P < 5.85 × 10−5 (0.05/855), which considered both the number of exposures and cancer subtypes. We picked the strict one for standard multiply testing. P-values that exceeded the strict Bonferroni threshold but were less than 0.05 indicated a suggestive association. All statistical analyses for MR were performed in the R software v 3.6.2. R package “TwoSampleMR” was used for two-sample MR analysis. Large-scale LDSC analyses were performed using the ‘ldsc’ software with the Linux system.
Results
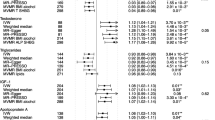
Figure 2 shows the associations of 95 genetically determined risk factors with BC in the IVW method or WRO analysis. Twenty-three exposures were significantly associated with overall BC. Positive associations [OR (95% CIs)] were with adult height (ADUH) [1.06 (1.02–1.10)], C-reactive protein (CRP) [1.09 (1.03–1.15)], platelet count (PLT) [1.04 (1.00–1.08)], schizophrenia [1.07 (1.03–1.10)], chronotype [1.19 (1.08–1.31)], high-density lipoprotein cholesterol (HDL-C) [1.10 (1.04–1.15)], apolipoprotein A1 (Apo A1) [1.07 (1.02–1.11)], insulin-like growth factor-1 (IGF-1) [1.08 (1.03–1.13)], omega-6:total fatty acids (O6:TFA) [1.06 (1.01–1.12)] and age at menopause (ANM) [1.05 (1.03–1.07)]. Negative associations were with body fat mass (BFM) [0.92 (0.87–0.98)], childhood obesity (COBE) [0.82 (0.76–0.88)], hip circumference (HC) [0.79 (0.65–0.94)], body mass index (BMI) [0.72 (0.65–0.80)], waist circumference (WC) [0.72 (0.60–0.86)], waist-to-hip ratio (WHR) [0.72 (0.55–0.94)], comparative body size at age 10 (CBS-10) [0.60 (0.54–0.67)], average acceleration (AVEA) [0.93 (0.89–0.98)], education attainment (EDUA) [0.91 (0.85–0.98)], age of smoking initiation (AOSI) [0.67 (0.51–0.89)], overall activity (OACT) [0.46 (0.24–0.91)], interleukin-6 receptor subunit alpha (IL-6 sRa) [0.98 (0.96–1.00)] and IGF-1 receptor (IGF-1R) [0.82 (0.76–0.88)]. Of the significant factors, COBE, BMI, CBS-10, IGF-1R and ANM passed the strict Bonferroni correction, whereas WC and schizophrenia also passed the correction when only considering the number of exposures. Supplementary Figures S1 and S2 show the results for ER + BC and ER − BC.
The odds ratio (OR) represented the effect of genetically predicted per unit increase in the risk factor. The majority of units were standard deviation (SD) and part of them were original units since the SD values were not available. Several R2 and F statistics were calculated by using the effect allele frequency in the outcome dataset since the missing allele frequency of exposure. For the number of SNP more than 1, the results were from the inverse variance weighted (IVW) Mendelian randomization; for the number of SNP equal to 1, the results were from the Wald ratio. * denotes P < 0.05, † denotes P < 5.26 × 10−4 (0.05/95), ‡ denotes P < 5.85 × 10−5 (0.05/855), # denotes potential pleiotropy in testing for the intercept of MR-Egger regression.
The relations between 95 traits and overall OC in the IVW method or WRO analysis are shown in Fig. 3. Fourteen exposures were significantly associated with overall OC. Positive associations [OR (95% CIs)] were with HC [1.24 (1.07–1.44)], WC [1.24 (1.05–1.46)], CBS-10 [1.20 (1.04–1.39)], BMI [1.19 (1.05–1.35)], body fat percentage (BFP) [1.18 (1.06–1.31)], BFM [1.14 (1.06–1.24)], basal metabolic rate (BMR) [1.13 (1.03–1.23)], body fat-free mass (BFFM) [1.12 (1.02–1.23)], schizophrenia [1.07 (1.02–1.12)] and omega-6:omega-3 fatty acids (O6/O3) [1.12 (1.02–1.24)]. Negative associations were with thyroid-stimulating hormone (TSH) [0.82 (0.67–0.99)], EDUA [0.83 (0.73–0.94)], adiponectin [0.87 (0.76–0.99)] and IGF-1R [0.64 (0.55–0.74)]. Of the factors, IGF-1R passed the Bonferroni correction. Supplementary Figures S3 to S7 show results for OC subtypes.
The odds ratio (OR) represented the effect of genetically predicted per unit increase in the risk factor. The majority of units were standard deviation (SD) and part of them were original units since the SD values were not available. Several R2 and F statistics were calculated by using the effect allele frequency in the outcome dataset since the missing allele frequency of exposure. For the number of SNP more than 1, the results were from the inverse variance weighted (IVW) Mendelian randomization; for the number of SNP equal to 1, the results were from the Wald ratio. * denotes P < 0.05, † denotes P < 5.26 × 10−4 (0.05/95), ‡ denotes P < 5.85 × 10−5 (0.05/855), # denotes potential pleiotropy in testing for the intercept of MR-Egger regression.
Forty-eight factors were significant for at least one cancer type in the IVW method or WRO analysis. Supplementary Figures S8 and S9 show the results of pairwise MVMR for BC and OC and the 48 factors. Overall, the associations were relatively robust after adjusting for other potential pleiotropy factors in turn. Effects of OACT, diet fat (D-Fat), folate and N3 docosapentaenoic acid (N3-DPA) could change to some extent when adjusted for other factors, which suggested potential horizontal pleiotropy.
Figure 4 summarises the robust associations (significant in both IVW and WME methods) for BC, OC and their subtypes. In general, 16 factors were identified as robust traits for overall BC, including chronotype, HDL-C, IGF-1, Apo A1, schizophrenia, O6:TFA, ANM, PLT, AVEA, BFM, EDUA, COBE, HC, BMI, WC and CBS-10 (Fig. 4a). In addition, CRP, O6:TFA, Apo A1, ADUH, IGF-1, schizophrenia, ANM, monounsaturated fatty acids (MUFAs), BFM, AVEA, COBE, WC, BMI, WHR and CBS-10 were robustly associated with ER + BC (Fig. 4b). Associated with ER − BC were HDL-C, schizophrenia, ANM, BMR, cognitive performance (CP), BFFM, COBE, EDUA, HC, BMI, WC and CBS-10 (Fig. 4c). For ER + BC, ANM, BFM, WC, BMI and CBS-10 passed the strict multiple testing, as did COBE, EDUA, BMI, WC and CBS-10 for ER − BC. For overall OC, WC, CBS-10, BMI, BFP, BFM, O6/O3 and EDUA remained significant after the sensitivity analysis by the WME method (Fig. 4d). For OC subtypes, CBS-10, linoleic acid (LA), O6FA, BFM, zinc and BIRW had robust associations with IMOC. Causally associated with EOC were HC, BMR, BFFM, O6/O3, BFM, ANM, BIRW and N3-DPA. The factors BMR and BFFM also passed multiple testing when considering the number of exposures. In addition, the BFP, BFM and adiponectin were robustly associated with HGSOC, and the BFFM, BMR, sex hormone-binding globulin (SHBG) and O6/O3 were also robust factors for CCOC.
The odds ratio (OR) represented the effect of genetically predicted per unit increase in the risk factor. Only factors both significant in IVW and WME method could be shown. The panel a-h showed the results of overall BC, ER+ BC, ER- BC, overall OC, IMOC, EOC, HGSOC, and CCOC, respectively. The results for low-grade serous ovarian cancer were not shown since no robust risk factors were detected. * denotes P < 0.05, † denotes P < 5.26 × 10−4 (0.05/95), ‡ denotes P < 5.85 × 10−5 (0.05/855). IVW inverse variance weighted, WME weighted median estimator.
Figure 5 summarises results for MR, GC and levels and rank of risk factors. Fifty-eight risk factors were significant for at least one type of cancer with at least one MR method (IVW, WRO or WME) (Fig. 5a). Thirty-one of the risk factors were robust factors for at least one BC/OC type (dark red). Moreover, many traits were shared risk factors across BC, OC and their subtypes. For example, BMI, BFM, CBS-10, HC, WC, schizophrenia, EDUA and IGF-1R were shared risk factors for overall BC and OC. In addition, 13 factors that shared genetic risk with these outcomes were also combined in the heat map, including OACT, COBE, BMI, BFFM, BMR, AVEA, CBS-10, schizophrenia, ANM, ADUA, CP, EDUA and insomnia (solid triangle). All significant results of GC analysis are shown in Fig. 5b. According to the results of MR and GC, five levels of risk factors were defined for BC and OC (Fig. 5c). Level 1 represented the robust MR plus the GC evidence for both BC and OC, which included only EDUA. Level 2 comprised only the robust MR results for both BC and OC, which included BMI, CBS-10, WC and BFM. In level 3, only COBE, AVEA, ANM, and schizophrenia were robustly associated with BC, and no factors were associated with OC. The above OACT, D-Fat, folate and N3-DPA, which showed potential horizontal pleiotropy in MVMR analysis, were mainly classified into level 5. Factors in level 5 were relatively unimportant compared with factors in other levels. Figure 5d shows the rank of the 58 significant factors according to their number of significant results (scores). The top 10 factors in order were CBS-10, EDUA, schizophrenia, BFM, BMI, ANM, WC, COBE, HC and O6/O3.
Panel a was the heat map of MR results. The coloured region represented the significant result in different methods including the IVW, WME and IVW + WME, while the white represented insignificant results. The direction of the triangle represents the direction of causal estimators where the upward triangle indicates positive association and the downward triangle indicates negative association. The solid triangle indicates a significant genetic correlation. Panel b was the result of genetic correlation (GC) analysis. The GC was quantified by the statistic of genetic correlation coefficient r, ranged from −1 to 1, and presented from blue to red colour. GC more than 0 represented a positive association and smaller than 0 represented a negative association. The asterisks (*) in the figure represent statistically significant results (P < 0.05). The results of low-grade serous ovarian cancer were not shown in GC analysis because the ldsc software failed to perform GC analysis on account of low h2 statistics or a small sample size. The insignificant factors for any outcomes were also not shown in this heat map. Panel c was the stratification of putative risk factors according to our criteria. The left one showed the factors for overall BC and OC only. The right one showed the factor for any BC and OC types. Panel d was the rank of these putative causal risk factors according to their scores defined by our criteria in the methods. The full annotation of the abbreviations in this Figure could be found in panel a or supplement Table S2.
Causal pathways and networks were developed from identified risk factors to BC (Fig. 6a) and to OC (Fig. 6b). In the networks, HC, WC, O6:TFA, IGF-1R, BFM, CBS-10, chronotype, ANM, IGF-1, Apo A1, PLT, AVEA, COBE, EDUA and BMI had both direct and indirect effects on BC, whereas BFM, CBS-10, BFP, O6/O3, EDUA, BMI, WC, IGF-1R and TSH affected the risk of OC through their respective pathways. For example, CBS-10 could affect the risk of OC by acting on BFM, WC, BMI and BFP (yellow pathways), suggesting that early-life body status could act on later-stage stature and lead to the risk of OC. In addition, the effect of EDUA on OC could also be mediated by obesity-related traits (grey pathways), indicating education could drive health-related behaviour to control obesity and ultimately modify the risk of OC. Causal pathways of BC and OC subtypes are shown in Supplementary Figs. S10 and S11. Causal estimators of exposure–outcome pairs between each node in the causal network diagrams are shown in Supplementary Table S4. The other independent factors were not shown in networks because they had no identifiable mediators that could be explained as having only direct causal effects on cancer.
The left panel was the network for breast cancer and the right one was for ovarian cancer. The arrows represented the direction of causal effect from one causal biomarker to another one, and finally, to ovarian/breast cancer (yellow highlight) from the network-MR analysis. This network only showed the factors with identifiable mediators. Each arrow represents a significant IVW-MR result.
Discussion
In this study, genetic statistical methods were used to identify the relations between large-scale cancer-related exposures and breast and ovarian cancers. Thirty-one exposures were robust risk factors for at least one type of BC or OC. Among them, BMI, BFM, CBS-10, WC and EDUA were shared robust risk factors for overall BC and OC, which implied potential joint intervention targets. In addition, 13 shared factors were detected in both GC and MR analyses, including OACT, COBE, BMI, BFFM, BMR, AVEA, CBS-10, schizophrenia, ANM, ADUA, CP, EDUA and insomnia. Furthermore, risk factors were stratified into five levels and ranked in order to prioritise future intervention measurements. Finally, visual causal networks were constructed that showed potential causal pathways from identified large-scale exposures to target outcomes to guide primary prevention practices.
Of the 31 putative robust factors from MR analysis, 16 factors were new causal biomarkers, including WC, BMR, BFP, BFFM, HC, COBE, BFM, PLT, CP, Apo A1, O6FA, O6:TFA, O6/O3, LA, MUFAs and N3-DPA. The other 15 factors, including BMI, WHR, ADUH, CBS-10, BIRW, CRP, schizophrenia, AVEA, EDUA, chronotype, HDL-C, adiponectin, IGF-1, ANM and SHBG, have been previously reported to be causal biomarkers. [8, 9, 11, 60,61,62,63,64,65,66,67,68,69,70,71,72,73] Consistent with previous studies, in this study, AUDH, schizophrenia, HDL-C, and IGF-1 were positively associated with BC and BMI and schizophrenia were positively associated with OC [8, 60, 63, 65, 67, 69]. In addition, BMI, CBS-10, AVEA and EDUA were negatively associated with BC, also consistent with previous studies [11, 62, 70, 73]. Same with previous studies [9, 61, 65, 68, 69], significant causal associations of BIRW and adiponectin with BC and WHR, ADUH, BIRW, CRP, adiponectin, IGF-1, ANM and SHBG with OC were not detected in this study. However, additional negative associations were detected, including WHR with BC and ER + BC, BIRW with IMOC and EOC and adiponectin with ER − BC, OC and HGSOC. Additional positive associations included ADUH with CCOC; CRP with BC and ER + BC; ANM with BC, ER + BC, ER − BC, and EOC and SHBG with CCOC, although some associations were only significant in the IVW method.
Of the robust risk factors, COBE, BMI, BFFM, BMR, AVEA, CBS-10, schizophrenia, ANM, ADUA, CP and EDUA also showed significant GCs, which suggested a higher level of causal evidence for which they could be stratified as priority intervention targets (all of these factors were classified into levels 1, 2 and 3). A factor that has a GC with cancer is more worthy of attention because it shares genetic risks with the outcome, and thus, an individual with such a trait would have a higher cancer risk than other people at the genetic level, even if it was not a causal risk factor. For a causal risk factor with GC, a comprehensive intervention should be implemented that intervenes not only with the particular factor but also with other risk factors (including those upstream, downstream and in other pathways in a network). For example, schizophrenia was a causal risk factor and had strong GC with BC (Fig. 5a), indicating that an individual with schizophrenia was at a genetically higher risk of BC. In this situation, intervention for only schizophrenia could not rule out additional genetic risk of BC, compared with other conventional risk factors. Therefore, a comprehensive intervention should target alternative pathways, such as BMI, AVEA, IGF-1, WC, HC and O6:TFA et al (Fig. 6). Moreover, although GC analysis showed significant results for coffee consumption (CC), childhood BMI (CBMI), alcohol consumption (AC), BIRL, Apo B, allergies, cigarettes per day (CPD), low-density lipoprotein cholesterol (LDL-C), sleep duration (SDU), triglycerides (TG) and depression with several outcomes, MR studies did not support a significant causal effect of those factors. Therefore, they were not included as intervention targets. Although GC could increase the priority of a causal risk factor, intervention could fail to decrease cancer risk because the factor did not have a causal effect.
The newly identified risk factors O6FA, O6:TFA, O6/O3 and MUFAs had a strong causal effect on OC, BC or several subtypes. These results suggested an adverse effect of O6FA and a potential protective effect of omega-3 fatty acids (O3FA). Causal evidence of fatty acids on cancer risk is limited, but in their review, Saini et al. [74] noted that O3FA and O6FA suppressed and induced inflammation, respectively. Diets enriched in O6FA are associated with inflammation, which provides an ideal tumour microenvironment and is linked to cancer risk and metastasis [74]. By contrast, O3FA help to resolve inflammation and alter the function of vascular and carcinogen biomarkers and thus reduce cancer risk [74, 75]. These differences may explain why different fatty acids can increase or decrease BC or OC risks. In addition, the results in study are consistent with the previous opinion that the ratio of O6FA to O3FA is more crucial than the absolute amounts [74].
Notably, there was dimorphism in obesity-related traits for OC and BC risks. This causal evidence suggests double-sided effects of obesity traits for different female cancers and indicates the importance of maintaining a moderate body shape. The factor CBS-10 reflects the early-life status of an individual and also had strong causality and GC with cancer (ranked No. 1 among the factors), indicating early childhood intervention is key. The protective effect of education on lung cancer has been previously verified [76], and the results of this study further extend the protective effect of education to OC and BC. A relatively high level of education is more likely to drive positive health-related behaviours and thus reduce the risk of cancer. The robust inverse associations combined with the extensive GCs with both BC and OC indicate education is one of the top factors to be addressed in intervention. The protective effect of IGF-1R may be due to negative feedback regulation of IGF-1, which has been reported as a risk factor for breast cancer [64]. The risk of Apo A1 for BC was consistent with the reported positive effect of HDL-C [8], likely because Apo A1 is a transporter of HDL-C.
Some important results that are not consistent with those of other studies should also be noted. For example, with AAM, whereas negative associations with ER + BC and IMOC were indicated, the association was not significant with BC. By contrast, Day et al. [77] reported a strong negative relation with BC after removing or adjusting BMI-related SNPs. To attempt to reproduce that result, additional analyses were conducted by using different AAM datasets (versions 2014 and 2017), different BC datasets (Oncoarray, iCOGS, GWAS and combined), different LD thresholds (0.01 and 0.001), and different methods (IVW, WME and BMI-adjusted and BMI-excluded SNPs) (Supplementary Fig. S12). Several results in only specific conditions were consistent the conclusion of Day et al. The possible difference in conclusions could be because the BMI GWAS dataset could not be acquired that, together with the reported AMM 61 BMI-related SNPs (AAM increasing/BMI decreasing) produced the strong positive association with BC. However, the results in this study are consistent with their unadjusted results and are also consistent with those of another MR study by Qi et al. [78]. Therefore, future research should focus on the mechanism by which AAM combined with BMI affects the risk of BC and distinguish the data sources in MR studies.
Finally, based on the above evidence, risk factors were stratified and ranked in order to provide a reference for future primary prevention targets. A complex network for BC, OC and each subtype was constructed based on network-MR analysis. This study was the first to summarise and stratify causal biomarkers and construct causal networks for BC and OC using biostatistics and data-driven evidence. When using a network, stratified risk factors can be referenced to identify more important targets. For each factor, primary prevention interventions can be directed not only at direct effects but also to interrupt its pathway in other related nodes. For example, CBS-10 could be altered in childhood, which would benefit downstream adult BMI and ultimately decrease the risk of OC (Fig. 6). Furthermore, when a risk factor cannot be easily modified, such as lower EDUA for risk of BC, alternative downstream targets (e.g. Apo A1 and obesity-related traits) can be identified to implement primary prevention interventions.
Compared with previous studies, this study had the advantage of examining large-scale factors (the largest to our knowledge) associated with OC and BC under a comprehensive framework in order to detect causalities, genetic correlations, and shared or distinct factors and to prioritise risk factors and develop causal networks. The study also had limitations that should be noted. First, some of the candidate factors did not have enough IVs and might suffer bias from that weakness. Second, pleiotropy is a dilemma in MR. Fortunately, the WME was more robust to IV assumptions, and the pairwise MVMR design found that few of the identified risk factors were affected by pleiotropy. Third, in large-scale exploratory research of observational datasets, there are natural limitations to detailed exploration of mechanisms of each factor, as in most similar studies. Further research is required to determine biological mechanisms of the risk factors newly identified in this research, as well as to explore the suggestive evidence. The practical value of risk stratification and a causal network needs to be verified in further public health intervention practices.
Conclusions
Sixteen new factors associated with OC or BC were identified, including WC, BMR, BFP, BFFM, HC, COBE, BFM, PLT, CP, Apo A1, O6FA, O6:TFA, O6/O3, LA, MUFAs and N3-DPA. In addition, BMI, BFM, CBS-10, WC and EDUA were shared robust risk factors for overall BC and OC. Thirteen factors were significant in both GC and MR analyses, including OACT, COBE, BMI, BFFM, BMR, AVEA, CBS-10, schizophrenia, ANM, ADUA, CP, EDUA and insomnia. The risk factors were stratified into five levels and ranked to prioritise for future intervention measurements. Causal networks were developed that show pathways from putative factors to cancer in order to guide primary prevention.
Data availability
Researchers may have access to this data from the original researches shown in the supplement material.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Islami F, Goding Sauer A, Miller KD, Siegel RL, Fedewa SA, Jacobs EJ, et al. Proportion and number of cancer cases and deaths attributable to potentially modifiable risk factors in the United States. CA Cancer J Clin. 2018;68:31–54.
Wilson LF, Antonsson A, Green AC, Jordan SJ, Kendall BJ, Nagle CM, et al. How many cancer cases and deaths are potentially preventable? Estimates for Australia in 2013. Int J Cancer. 2018;142:691–701.
Brown KF, Rumgay H, Dunlop C, Ryan M, Quartly F, Cox A, et al. The fraction of cancer attributable to modifiable risk factors in England, Wales, Scotland, Northern Ireland, and the United Kingdom in 2015. Br J Cancer. 2018;118:1130–41.
Zhu J, Jiang X, Niu Z. Alcohol consumption and risk of breast and ovarian cancer: a Mendelian randomization study. Cancer Genet. 2020;245:35–41.
Bulik-Sullivan B, Finucane HK, Anttila V, Gusev A, Day FR, Loh P-R, et al. An atlas of genetic correlations across human diseases and traits. Nat Genet. 2015;47:1236–41.
Beeghly-Fadiel A, Khankari NK, Delahanty RJ, Shu X-O, Lu Y, Schmidt MK, et al. A Mendelian randomization analysis of circulating lipid traits and breast cancer risk. Int J Epidemiol. 2020;49:1117–31.
Yarmolinsky J, Relton CL, Lophatananon A, Muir K, Menon U, Gentry-Maharaj A, et al. Appraising the role of previously reported risk factors in epithelial ovarian cancer risk: a Mendelian randomization analysis. PLoS Med. 2019;16:e1002893.
Au Yeung SL, Schooling CM. Impact of glycemic traits, type 2 diabetes and metformin use on breast and prostate cancer risk: a Mendelian randomization study. BMJ Open Diabetes Res Care. 2019;7:e000872.
Ooi BNS, Loh H, Ho PJ, Milne RL, Giles G, Gao C, et al. The genetic interplay between body mass index, breast size and breast cancer risk: a Mendelian randomization analysis. Int J Epidemiol. 2019;48:781–94.
Si S, Tewara MA, Li Y, Li W, Chen X, Yuan T, et al. Causal pathways from body components and regional fat to extensive metabolic phenotypes: a Mendelian randomization study. Obesity. 2020;28:1536–49.
Manning AK, Hivert M-F, Scott RA, Grimsby JL, Bouatia-Naji N, Chen H, et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat Genet. 2012;44:659–69.
Bradfield JP, Taal HR, Timpson NJ, Scherag A, Lecoeur C, Warrington NM, et al. A genome-wide association meta-analysis identifies new childhood obesity loci. Nat Genet. 2012;44:526–31.
van der Valk RJP, Kreiner-Møller E, Kooijman MN, Guxens M, Stergiakouli E, Sääf A, et al. A novel common variant in DCST2 is associated with length in early life and height in adulthood. Hum Mol Genet. 2015;24:1155–68.
Liu M, Jiang Y, Wedow R, Li Y, Brazel DM, Chen F, et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet. 2019;51:237–44.
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Soranzo N, Sanna S, Wheeler E, Gieger C, Radke D, Dupuis J, et al. Common variants at 10 genomic loci influence hemoglobin A1(C) levels via glycemic and nonglycemic pathways. Diabetes. 2010;59:3229–39.
Wood AR, Esko T, Yang J, Vedantam S, Pers TH, Gustafsson S, et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat Genet. 2014;46:1173–86.
Willer CJ, Schmidt EM, Sengupta S, Peloso GM, Gustafsson S, Kanoni S, et al. Discovery and refinement of loci associated with lipid levels. Nat Genet. 2013;45:1274–83.
Lee JJ, Wedow R, Okbay A, Kong E, Maghzian O, Zacher M, et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat Genet. 2018;50:1112–21.
Lemaitre RN, Tanaka T, Tang W, Manichaikul A, Foy M, Kabagambe EK, et al. Genetic loci associated with plasma phospholipid n-3 fatty acids: a meta-analysis of genome-wide association studies from the CHARGE Consortium. PLoS Genet. 2011;7:e1002193.
International Multiple Sclerosis Genetics Consortium, Wellcome Trust Case Control Consortium 2, Sawcer S, Hellenthal G, Pirinen M, Spencer CCA, et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature. 2011;476:214–9.
Locke AE, Kahali B, Berndt SI, Justice AE, Pers TH, Day FR, et al. Genetic studies of body mass index yield new insights for obesity biology. Nature. 2015;518:197–206.
Köttgen A, Albrecht E, Teumer A, Vitart V, Krumsiek J, Hundertmark C, et al. Genome-wide association analyses identify 18 new loci associated with serum urate concentrations. Nat Genet. 2013;45:145–54.
Felix JF, Bradfield JP, Monnereau C, van der Valk RJP, Stergiakouli E, Chesi A, et al. Genome-wide association analysis identifies three new susceptibility loci for childhood body mass index. Hum Mol Genet. 2016;25:389–403.
Strawbridge RJ, Dupuis J, Prokopenko I, Barker A, Ahlqvist E, Rybin D, et al. Genome-wide association identifies nine common variants associated with fasting proinsulin levels and provides new insights into the pathophysiology of type 2 diabetes. Diabetes. 2011;60:2624–34.
Ahola-Olli AV, Würtz P, Havulinna AS, Aalto K, Pitkänen N, Lehtimäki T, et al. Genome-wide association study identifies 27 loci influencing concentrations of circulating cytokines and growth factors. Am J Hum Genet. 2017;100:40–50.
Evans DM, Zhu G, Dy V, Heath AC, Madden PAF, Kemp JP, et al. Genome-wide association study identifies loci affecting blood copper, selenium and zinc. Hum Mol Genet. 2013;22:3998–4006.
Jiang X, O’Reilly PF, Aschard H, Hsu Y-H, Richards JB, Dupuis J, et al. Genome-wide association study in 79,366 European-ancestry individuals informs the genetic architecture of 25-hydroxyvitamin D levels. Nat Commun. 2018;9:260.
Clarke T-K, Adams MJ, Davies G, Howard DM, Hall LS, Padmanabhan S, et al. Genome-wide association study of alcohol consumption and genetic overlap with other health-related traits in UK Biobank (N=112 117). Mol Psychiatry. 2017;22:1376–84.
Klimentidis YC, Raichlen DA, Bea J, Garcia DO, Wineinger NE, Mandarino LJ, et al. Genome-wide association study of habitual physical activity in over 377,000 UK Biobank participants identifies multiple variants including CADM2 and APOE. Int J Obes (Lond). 2018;42:1161–76.
Kilpeläinen TO, Carli JFM, Skowronski AA, Sun Q, Kriebel J, Feitosa MF, et al. Genome-wide meta-analysis uncovers novel loci influencing circulating leptin levels. Nat Commun. 2016;7:10494.
Kettunen J, Demirkan A, Würtz P, Draisma HHM, Haller T, Rawal R, et al. Genome-wide study for circulating metabolites identifies 62 loci and reveals novel systemic effects of LPA. Nat Commun. 2016;7:11122.
DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium, Asian Genetic Epidemiology Network Type 2 Diabetes (AGEN-T2D) Consortium, South Asian Type 2 Diabetes (SAT2D) Consortium, Mexican American Type 2 Diabetes (MAT2D) Consortium, Type 2 Diabetes Genetic Exploration by Nex-generation sequencing in muylti-Ethnic Samples (T2D-GENES) Consortium, Mahajan A, et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat Genet. 2014;46:234–44.
Meddens SFW, de Vlaming R, Bowers P, Burik CAP, Linnér RK, Lee C, et al. Genomic analysis of diet composition finds novel loci and associations with health and lifestyle. Mol Psychiatry. 2021;26:2056–69.
Sun BB, Maranville JC, Peters JE, Stacey D, Staley JR, Blackshaw J, et al. Genomic atlas of the human plasma proteome. Nature. 2018;558:73–9.
Doherty A, Smith-Byrne K, Ferreira T, Holmes MV, Holmes C, Pulit SL, et al. GWAS identifies 14 loci for device-measured physical activity and sleep duration. Nat Commun. 2018;9:5257.
Day FR, Ruth KS, Thompson DJ, Lunetta KL, Pervjakova N, Chasman DI, et al. Large-scale genomic analyses link reproductive aging to hypothalamic signaling, breast cancer susceptibility and BRCA1-mediated DNA repair. Nat Genet. 2015;47:1294–303.
Warrington NM, Beaumont RN, Horikoshi M, Day FR, Helgeland Ø, Laurin C, et al. Maternal and fetal genetic effects on birth weight and their relevance to cardio-metabolic risk factors. Nat Genet. 2019;51:804–14.
Demenais F, Margaritte-Jeannin P, Barnes KC, Cookson WOC, Altmüller J, Ang W, et al. Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nat Genet. 2018;50:42–53.
Dupuis J, Langenberg C, Prokopenko I, Saxena R, Soranzo N, Jackson AU, et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet. 2010;42:105–16.
Shungin D, Winkler TW, Croteau-Chonka DC, Ferreira T, Locke AE, Mägi R, et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature. 2015;518:187–96.
Benyamin B, Esko T, Ried JS, Radhakrishnan A, Vermeulen SH, Traglia M, et al. Novel loci affecting iron homeostasis and their effects in individuals at risk for hemochromatosis. Nat Commun. 2014;5:4926.
Dastani Z, Hivert M-F, Timpson N, Perry JRB, Yuan X, Scott RA, et al. Novel loci for adiponectin levels and their influence on type 2 diabetes and metabolic traits: a multi-ethnic meta-analysis of 45,891 individuals. PLoS Genet. 2012;8:e1002607.
Perry JR, Day F, Elks CE, Sulem P, Thompson DJ, Ferreira T, et al. Parent-of-origin-specific allelic associations among 106 genomic loci for age at menarche. Nature. 2014;514:92–7.
Ferreira MA, Vonk JM, Baurecht H, Marenholz I, Tian C, Hoffman JD, et al. Shared genetic origin of asthma, hay fever and eczema elucidates allergic disease biology. Nat Genet. 2017;49:1752–7.
Astle WJ, Elding H, Jiang T, Allen D, Ruklisa D, Mann AL, et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell. 2016;167:1415–1429.e19.
Michailidou K, Lindström S, Dennis J, Beesley J, Hui S, Kar S, et al. Association analysis identifies 65 new breast cancer risk loci. Nature. 2017;551:92–4.
Phelan CM, Kuchenbaecker KB, Tyrer JP, Kar SP, Lawrenson K, Winham SJ, et al. Identification of 12 new susceptibility loci for different histotypes of epithelial ovarian cancer. Nat Genet. 2017;49:680–91.
Burgess S, Thompson SG. Interpreting findings from Mendelian randomization using the MR-Egger method. Eur J Epidemiol. 2017;32:377–89.
Burgess S, Dudbridge F, Thompson SG. Combining information on multiple instrumental variables in Mendelian randomization: comparison of allele score and summarized data methods. Stat Med. 2016;35:1880–906.
Huang T, Afzal S, Yu C, Guo Y, Bian Z, Yang L, et al. Vitamin D and cause-specific vascular disease and mortality: a Mendelian randomisation study involving 99,012 Chinese and 106,911 European adults. BMC Med. 2019;17:160.
Burgess S, Butterworth A, Thompson SG. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 2013;37:658–65.
Bowden J, Davey, Smith G, Haycock PC, Burgess S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol. 2016;40:304–14.
Burgess S, Thompson SG. Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects. Am J Epidemiol. 2015;181:251–60.
Si S, Hou L, Chen X, Li W, Liu X, Liu C, et al. Exploring the causal roles of circulating remnant lipid profile on cardiovascular and cerebrovascular diseases: Mendelian randomization study. J Epidemiol. 2021 Jan 13. https://doi.org/10.2188/jea.JE20200305. Epub ahead of print. PMID: 33441507.
Si S, Li J, Tewara MA, Xue F. Genetically determined chronic low-grade inflammation and hundreds of health outcomes in the UK Biobank and the FinnGen population: a phenome-wide Mendelian randomization study. Front Immunol. 2021;12:720876.
Burgess S, Daniel RM, Butterworth AS, Thompson SG. EPIC-InterAct Consortium. Network Mendelian randomization: using genetic variants as instrumental variables to investigate mediation in causal pathways. Int J Epidemiol. 2015;44:484–95.
Adams CD, Neuhausen SL. Bi-directional Mendelian randomization of epithelial ovarian cancer and schizophrenia and uni-directional Mendelian randomization of schizophrenia on circulating 1- or 2-glycerophosphocholine metabolites. Mol Genet Metab Rep. 2019;21:100539.
Dimou NL, Papadimitriou N, Mariosa D, Johansson M, Brennan P, Peters U, et al. Circulating adipokine concentrations and risk of five obesity-related cancers: a Mendelian randomization study. Int J Cancer. 2021;148:1625–36.
Yuan S, Xiong Y, Michaëlsson M, Michaëlsson K, Larsson SC. Genetically predicted education attainment in relation to somatic and mental health. Sci Rep. 2021;11:4296.
Qian F, Wang S, Mitchell J, McGuffog L, Barrowdale D, Leslie G, et al. Height and body mass index as modifiers of breast cancer risk in BRCA1/2 mutation carriers: a Mendelian randomization study. J Natl Cancer Inst. 2019;111:350–64.
Murphy N, Knuppel A, Papadimitriou N, Martin RM, Tsilidis KK, Smith-Byrne K, et al. Insulin-like growth factor-1, insulin-like growth factor-binding protein-3, and breast cancer risk: observational and Mendelian randomization analyses with ∼430 000 women. Ann Oncol. 2020;31:641–9.
Larsson SC, Carter P, Vithayathil M, Kar S, Mason AM, Burgess S. Insulin-like growth factor-1 and site-specific cancers: a Mendelian randomization study. Cancer Med. 2020;9:6836–42.
Richmond RC, Anderson EL, Dashti HS, Jones SE, Lane JM, Strand LB, et al. Investigating causal relations between sleep traits and risk of breast cancer in women: mendelian randomisation study. BMJ. 2019;365:l2327.
Byrne EM, Ferreira MAR, Xue A, Lindström S, Jiang X, Yang J, et al. Is schizophrenia a risk factor for breast cancer? Evidence from genetic data. Schizophr Bull. 2019;45:1251–6.
Robinson T, Martin RM, Yarmolinsky J. Mendelian randomisation analysis of circulating adipokines and C-reactive protein on breast cancer risk. Int J Cancer. 2020;147:1597–603.
Gao C, Patel CJ, Michailidou K, Peters U, Gong J, Schildkraut J, et al. Mendelian randomization study of adiposity-related traits and risk of breast, ovarian, prostate, lung and colorectal cancer. Int J Epidemiol. 2016;45:896–908.
Papadimitriou N, Dimou N, Tsilidis KK, Banbury B, Martin RM, Lewis SJ, et al. Physical activity and risks of breast and colorectal cancer: a Mendelian randomisation analysis. Nat Commun. 2020;11:597.
Dimou NL, Papadimitriou N, Gill D, Christakoudi S, Murphy N, Gunter MJ, et al. Sex hormone binding globulin and risk of breast cancer: a Mendelian randomization study. Int J Epidemiol. 2019;48:807–16.
Kar SP, Andrulis IL, Brenner H, Burgess S, Chang-Claude J, Considine D, et al. The association between weight at birth and breast cancer risk revisited using Mendelian randomisation. Eur J Epidemiol. 2019;34:591–600.
Richardson TG, Sanderson E, Elsworth B, Tilling K, Davey Smith G. Use of genetic variation to separate the effects of early and later life adiposity on disease risk: mendelian randomisation study. BMJ. 2020;369:m1203.
Saini RK, Keum Y-S. Omega-3 and omega-6 polyunsaturated fatty acids: dietary sources, metabolism, and significance—a review. Life Sci. 2018;203:255–67.
Adkins Y, Kelley DS. Mechanisms underlying the cardioprotective effects of omega-3 polyunsaturated fatty acids. J Nutr Biochem. 2010;21:781–92.
Zhou H, Zhang Y, Liu J, Yang Y, Fang W, Hong S, et al. Education and lung cancer: a Mendelian randomization study. Int J Epidemiol. 2019;48:743–50.
Day FR, Thompson DJ, Helgason H, Chasman DI, Finucane H, Sulem P, et al. Genomic analyses identify hundreds of variants associated with age at menarche and support a role for puberty timing in cancer risk. Nat Genet. 2017;49:834–41.
Qi G, Chatterjee N. Mendelian randomization analysis using mixture models for robust and efficient estimation of causal effects. Nat Commun. 2019;10:1941.
Acknowledgements
We thank to the MR Base platform, Integrative Epidemiology Unit (IEU) project, Neale Lab (Neale Lab analysis of UK Biobank phenotypes, round 2), MRC-IEU (MRC-IEU UK Biobank GWAS pipeline v 2), Borges et al. (metabolic biomarkers in the UK Biobank measured by Nightingale Health 2020), all related consortiums including BCAC, OCAC, GIANT, EGG, MRC-IEU, GUGC, Neale Lab, DIAGRAM, IMSGC, PGC, SSGAC, UK Biobank, GSCAN, GLGC, MAGIC, ADIPOGen, GIS, CHARGE and ReproGen, and original research by Astle et al., Jiang et al., Sun et al., Kettunen et al., Clarke et al., Doherty et al., Ferreira et al., Demenais et al., Klimentidis et al., Strawbridge et al., Manning et al., Dupuis et al., Evans et al. and Borges et al. for providing open access summary-level data resources. For a full description of acknowledgements concerning these studies, cohorts and consortiums, as well as funding information and other details, see Supplementary Text 3. We also thank the Charlesworth Group (https://www.cwauthors.com.cn/) for its linguistic assistance during the round 2 revision.
Funding
This research was funded by the National Key Research and Development Program (2020YFC2003500), the National Natural Science Foundation of China (81773547) and the Natural Science Foundation of Shandong Province (ZR2019ZD02). The corresponding author FX obtained the funding. The funders had no role in this work.
Author information
Authors and Affiliations
Contributions
SS and FX have the conception. SS did the statistical analyses and drafted the initial manuscript. MAT completed the revision of English grammar. All authors participated in the interpretation of the results, edited and reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Our research is only based on publicly available summarised data and no individual data were involved. These genome-wide association studies (including the GIANT, EGG, MRC-IEU, GUGC, Neale Lab, DIAGRAM, IMSGC, PGC, SSGAC, UK Biobank, GSCAN, GLGC, MAGIC, ADIPOGen, GIS, CHARGE and ReproGen consortium, and the individual research by Astle WJ et al, Jiang X et al, Sun BB et al, Kettunen et al, T-K Clarke et al, Aiden Doherty et al, Ferreira MA et al, Demenais F et al, Klimentidis YC et al, Strawbridge RJ et al, Manning AK et al, Dupuis J et al, Evans et al and Borges CM et al) had declared the ethic approval from the relevant institutional review board and in accordance with the declaration of Helsinki, or other (see reference [15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50]). This secondary analysis is based on summary-level statistics that are not applicable for the clauses of ethics, the Ethics Committee of the School of Public Health of Shandong University ruled that ethics approval was not required in this particular case.
Consent for publication
Not applicable
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Si, S., Li, J., Tewara, M.A. et al. Identifying causality, genetic correlation, priority and pathways of large-scale complex exposures of breast and ovarian cancers. Br J Cancer 125, 1570–1581 (2021). https://doi.org/10.1038/s41416-021-01576-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01576-7
- Springer Nature Limited
This article is cited by
-
Relationship between oily fish intake and breast cancer based on estrogen receptor status: a Mendelian randomization study
Breast Cancer Research and Treatment (2024)
-
Ovarian cancer risk of Chinese women with BRCA1/2 germline pathogenic variants
Journal of Human Genetics (2022)