Abstract
Transforming growth factor β (TGF-β) is a multifunctional polypeptide that plays critical roles in regulating a broad range of cellular functions and physiological processes. TGF-β signalling dysfunction contributes to many disorders, such as cardiovascular diseases, cancer and immunological diseases. The homoeostasis of negative feedback regulation is critical for signal robustness, duration and specificity, which precisely control physiological and pathophysiological processes. However, the underlying mechanism by which the negative regulation of TGF-β signalling is integrated and coordinated is still unclear. Here, we reveal that haematopoietic progenitor kinase-interacting protein of 55 kDa (HIP-55) was upregulated upon TGF-β stimulation, while the loss of HIP-55 caused TGF-β signalling overactivation and the abnormal accumulation of downstream extracellular matrix (ECM) genes. HIP-55 interacts with Smad7 and competes with Smad7/Axin complex formation to inhibit the Axin-mediated degradation of Smad7. HIP-55 further couples Smad7 to TβRI but not TβRII, driving TβRI degradation. Altogether, our findings demonstrate a new mechanism by which the effector and negative feedback functions of HIP-55 are coupled and may provide novel strategies for the treatment of TGF-β signalling-related human diseases.
Similar content being viewed by others
Introduction
Transforming growth factor β (TGF-β) is a secreted pleiotropic factor that plays critical roles in embryogenesis and adult tissue homoeostasis by regulating cell proliferation, differentiation, migration, and death and the immune response [1,2,3,4]. Impairment of TGF-β signalling has been implicated in the pathogenesis of many major diseases, including cancer, cardiovascular diseases, immunological diseases, metabolic disorders and tissue fibrosis [1, 5,6,7]. TGF-β ligands transmit downstream signals through type I receptors (TβRI) and type II receptors (TβRII) at the cell surface, which control the initiation, duration and termination of TGF-β signalling. Upon binding TGF-β ligands, TβRI and TβRII receptors assemble into heteromeric complexes that allow the TβRII phosphorylation, which are constitutively active to phosphorylate TβRI and activating TβRI kinases. Subsequently, receptor-regulated Smad family (R-Smads: Smad2 and Smad3) are directly phosphorylated by TβRI kinases to form a heteromeric complex with Co-Smad protein Smad4, which translocate to the nucleus and induce Smad-mediated transcriptional regulation of extracellular matrix (ECM) genes to further exert cellular responses [3, 5].
Because of its significant role in cell fate determination, TGF-β signalling is precisely regulated by various positive and negative feedback mechanisms that target ligands, receptors and intracellular mediators. The stability and activity of receptors are mainly determined by feedback, especially negative feedback regulation [8,9,10]. In this regard, Smad7 has been extensively reported to play a central role in negative feedback regulation. As a general inhibitor of TGF-β signalling, Smad7 can form a complex with TβRI upon ligand stimulation and therefore inhibit the recruitment and activation of R-Smads, subsequently recruiting E3 ubiquitin ligases to target TβRI for ubiquitination-mediated degradation in the proteasomal pathway [11,12,13]. Thus, the stability of Smad7 is also an important target in the negative feedback regulation of TGF-β signalling [10]. Axin-induced Smad7 degradation is the major way by which the stability and activity of Smad7 are modulated. Axin sequesters Smad7 in the cytoplasm, brining Smad7 in proximity to its E3 ligase, Arkadia (RNF111), facilitating Smad7 polyubiquitination and degradation [14, 15]. Smad7 that escapes the Axin/Arkadia degradation system can interact with TβRI by the MH2 domain, which is required to maximally inhibit TβRI activity [16,17,18]. However, how this integrated, multicomponent system functions in the negative feedback regulation of TGF-β remains a challenge that has not yet been explored.
Haematopoietic progenitor kinase 1 (HPK1)-interacting protein of 55 kDa (HIP-55) (also known as SH3P7 and mAbp1) is a multidomain adaptor protein that includes an N-terminal F-actin-binding (ADFH) domain and a C-terminal Src homology 3 (SH3) domain [19, 20]. HIP-55 is localised to the plasma membrane and to actin-rich sites in cells [21, 22]. HIP-55 can bind dynamin, a GTP-binding protein involved in vesicle fission, through its SH3 domain during endocytosis [21]. HIP-55 has been demonstrated to negatively regulate T cell receptor (TCR) and B cell receptor (BCR) signalling by recruiting inhibitory signalling molecules in BCR/TCR signalosomes [23, 24]. In addition, HIP-55 can bind the partner adaptor protein FHL2 to differentially regulate Rho GTPase signalling and impair cell invasion [25]. Although the presence of multiple domains in this adaptor protein implicates its functional diversity, few studies have examined the regulatory effects and mechanisms of HIP-55.
In the present study, we reveal that HIP-55 expression was increased in response to TGF-β stimulation and that the loss of HIP-55 damaged TGF-β signalling homoeostasis and exacerbated the accumulation of ECM genes. HIP-55 was found to maintain the stability of Smad7 by interacting with Smad7 to inhibit Smad7/Axin complex formation. HIP-55 Smad7 triggering the degradation of TβRI to further integrate components of the TGF-β negative feedback network. By harnessing HIP-55-mediated integration of the negative feedback regulation of TGF-β signalling, putative prophylactic and therapeutic strategies against TGF-β signalling-related human diseases can be developed.
Materials and methods
Isolation of primary mouse cardiac fibroblasts (CFs)
Primary mouse CFs were isolated from 8- to 12-week-old C57/BL6 wild-type mice and the mice were purchased from Weitonglihua Company (Beijing, China). The hearts were quickly removed from the mice under sterile conditions, washed gently in cold phosphate-buffered saline (PBS), minced as much as possible and then digested with 0.1% collagenase type II (Sigma-Aldrich, St. Louis, MO, USA) at 37 °C for 8 min with slow, constant shaking. The supernatants were collected in sterile tubes. This digestion cycle was repeated 6–10 times until all tissue was dissolved. Then, the digested products were suspended in equal volumes of Dulbecco’s Minimum Essential Medium (DMEM) containing 10% FBS and centrifuged at 1000 r/min for 5 min. Subsequently, the cells were resuspended in DMEM containing 10% fetal bovine serum (FBS) and 100 U/mL penicillin/streptomycin. After the cells were transferred to plates and incubated for ~4 h, floating cells and unattached or weakly adherent cells, including myocytes and endothelial cells, were removed by exchanging the medium for fresh medium. The culture medium was changed again after 24 h. Cells at passage 2 were used for immunoblotting (IB), and at passage 1 were used for qRT-PCR analysis. The animal study was approved by the Animal Care Committee, of Peking University.
Cell culture and transfection
HEK293A cells, HEK293T cells and mouse CFs were maintained in DMEM supplemented with 10% FBS at 37 °C in a humidified, 5% CO2 incubator. A stable ShRNA-HIP-55-expressing HEK293A cell line was generated by transfecting HEK293A cells with ShRNA-HIP-55 (5′-CTTTGGCATGTTTCCTGCCAA-3′). SiRNA-mediated knockdown of Smad7 in CFs was achieved by transfection of predesigned ON-TARGET plus SMART pool mouse Smad7 siRNAs (Dharmacon, Lafayette, Co, USA) using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA). The cells were used for analyses at 48 h following transfection. In immunoprecipitation (IP) experiments, 48 h before harvesting, the cells were transiently transfected with the indicated plasmids using Lipofectamine 2000 (Invitrogen). To overexpress or knockdown HIP-55, CFs were transfected using one of the following vectors: adenovirus-Dsred as a control vector, adenovirus-Dsred-HIP-55, adenovirus-GFP as a control vector and adenovirus-GFP-HIP-55 knockdown. After 48 h, the infected CFs were collected for further research. The cells were treated with 5 ng/mL TGF-β (112.8668 nmol) for 3 h or 24 h, 50 μg/mL cycloheximide (CHX) (177.7146 μmol) for the indicated duration, and 5 μmol of MG132 for 6 h.
Plasmids and agents
GFP-Arkadia/RNF111 was a gift from Vasso Episkopou (Addgene plasmid #112228; http://n2t.net/addgene:112228; RRID:Addgene_112228) [26], Flag-Axin was a gift from Mariann Bienz (Addgene plasmid #109370; http://n2t.net/addgene:109370; RRID:Addgene_109370) [27], and Flag-Smad7 and Myc-TβRI were gifts from Kang-min He at the Chinese Academy of Sciences.
Human HIP-55 and ubiquitin cDNA were subcloned into pcDNA3.0 with the indicated tags. A short hairpin RNA (shRNA) construct used to knockdown mouse HIP-55 expression (5′-UCUUUGGCAUGUUUCCUGCCAACUCGAGUUGGCAGGAAACAUGCCAAAGUUUUUC-3′) was generated by PCR and subcloned into the pAV-GFP vector to target mouse HIP-55. A shRNA construct used to knockdown human HIP-55 expression (5′-CTTTGGCATGTTTCCTGCCAA-3′) was generated by PCR and subcloned into the pAV-GFP vector to target human HIP-55. The Dsred-HIP-55 RNA construct, used to overexpress HIP-55, was generated by subcloning HIP-55 cDNA into the pAV-IRES/Dsred vector.
Antibodies against Smad7 (mouse and rabbit, with the mouse antibody used for IB and the rabbit antibody used for IP) and HIP-55 (rabbit, used for both IP and IB) were purchased from Proteintech (Rosemont, RNP, USA). Antibodies against P-smad2, Smad2, P-smad3, Smad3, TβRII receptor, GAPDH, mouse IgG (light chain-specific), rabbit IgG (light chain-specific), IgG (mouse and rabbit) and TGF-β1 (mouse and human) were purchased from Cell Signalling Technology (Danvers, MA, USA). Antibodies against Smad7 (mouse, for IB), HIP-55 (mouse, for both IP and IB), and Axin were purchased from Santa Cruz Biotechnology (Delaware, CA, USA). Antibody against the TβRI receptor was purchased from Abcam (Cambridge, MA, USA), Arkadia was purchased from Abnova (Taipei, Taiwan), and HA was purchased from Huaxinbio (Beijing, China). MG132 was purchased from Selleck (Houston, TX, USA), and CHX was purchased from Sigma-Aldrich.
IP and Western blot analysis
Cells were lysed by resuspension in IP assay buffer (Beyotime, Shanghai, China) containing a protease inhibitor cocktail (Roche, Switzerland). After incubation for 30 min at 4 °C with shaking, the extracted proteins were recovered by centrifugation at 12,000 r/min for 15 min. For IP assays, cellular lysates were incubated with 2–5 μg of antibodies and a 30 mL bed volume of protein A/G-Sepharose beads (Thermo Fisher Scientific, Rockford, IL, USA) for 4–6 h at 4 °C, followed by 4 successive washes with IP buffer. Extracts and immunoprecipitated proteins were resolved by 8%, 10% or 12% SDS-PAGE, transferred to nitrocellulose membranes and analysed by Western blotting [28].
In vivo ubiquitination assay
HEK293T cells were transfected with His-HA-ubiquitin and other expression vectors as indicated. At 48 h after transfection, the cells were treated with 5 μmol of MG132 for 6 h and lysed with a lysis solution containing Triton X (Millipore, Darmstadt, Germany). The lysate was then treated with 5 ng/mL TGF-β (112.8668 nmol) for 3 h.
Transcriptional analysis
Total RNA was prepared with TRIzol reagent (Thermo Fisher Scientific), and cDNA was synthesised from 1 μg of RNA with ReverTra Ace. Quantitative reverse transcription-PCR (RT-PCR) was performed using the SYBR Green detection method and an Mx3000P quantitative PCR system (TransGen, Beijing, China). Gene expression was normalised against GAPDH mRNA expression and calculated using the 2−ΔΔCT method. Each sample was measured in triplicate experiments.
Immunofluorescence analysis
Cells were cultured on confocal dishes and grown to subconfluence. Then, they were fixed in 4% paraformaldehyde for 15 min. After being washed with PBS 3 times, the fixed cells were permeabilized with 0.1% Triton X-100 and blocked in PBS containing 4% BSA for 1 h at room temperature. The cells were then incubated with primary antibody overnight at 4 °C. After the cells were washed 3 times in PBS and incubated with secondary antibody conjugated to Alexa Fluor 488 (Invitrogen) or Alexa Fluor 594 (Invitrogen) for 1 h at room temperature, the nuclei were counterstained with Hoechst 33342 (Thermo Fisher Scientific). Then, images were captured on a Zeiss LSM 780 confocal microscope (Carl Zeiss, Thornwood, NY, USA) [29].
Statistical analyses
Data are reported as the mean values from the number of replicates indicated in the legends ± SEMs (shown as bars). Differences between two groups were analysed by the two-tailed Student’s t test. Differences between more than two groups were analysed by one-way analysis of variance (ANOVA) or two-way ANOVA with Tukey’s post hoc multiple comparison tests. Differences for which the P value < 0.05 were considered to be statistically significant.
Results
HIP-55 reduces TβRI protein levels and suppresses TGF-β signalling
To gain insight into the function of HIP-55 in TGF-β signalling, we detected the protein levels of TGF-β receptors and Smad signalling activation after HIP-55 was knocked down in primary CFs. Interestingly, the protein level of TβRI, but not that of TβRII, was upregulated after HIP-55 deletion. Moreover, the knockdown of endogenous HIP-55 elevated the levels of phosphorylated Smad2 and Smad3 (P-smad2 and P-smad3) (Fig. 1a). Consistent with the HIP-55 knockdown experiments, HIP-55 overexpression was accompanied by reductions in TβRI, P-smad2 and P-smad3, but no change in TβRII was detected (Fig. 1b). In addition, HIP-55 was upregulated in response to TGF-β exposure (Fig. 1a, b), suggesting that HIP-55 responds to TGF-β-induced pathological stimulation, as long-term TGF-β exposure (3 and 24 h) in this study can be regarded as equivalent to pathological conditions.
a, b Representative immunoblotting (IB) and quantification of TβRI, TβRII, P-smad2, T-smad2, P-smad3, T-smad3 and HIP-55 protein levels in cardiac fibroblasts (CFs) infected with Scrambled, ShRNA-HIP-55, Dsred or Dsred-HIP-55 and then treated with TGF-β (5 ng/mL) for 3 h. c, d qRT-PCR analysis of Collagen I, Collagen III, Fibronectin, α-SMA and HIP-55 mRNA expression in cultured CFs infected with Scrambled, ShRNA-HIP-55, Dsred or Dsred-HIP-55 and then treated with TGF-β (5 ng/mL) for 24 h. GAPDH was used as an internal control, and the values are the means ± SEMs of five independent experiments (n = 5). *P < 0.05, **P < 0.01, ***P < 0.001 vs. the Scrambled group or the Dsred group; #P < 0.05, ##P < 0.01, ###P < 0.001 vs. the Scrambled + TGF-β group or the Dsred + TGF-β group; &P < 0.05, &&P < 0.01, &&&P < 0.001 vs. the ShRNA-HIP-55 group or the Dsred-HIP-55 group; ns no significance (P > 0.05); two-way ANOVA with Tukey’s post hoc multiple comparisons for (a–d).
Since fibrotic ECM gene expression is an important downstream component of TGF-β signalling [30], we investigated the regulatory effects of HIP-55 on TGF-β-dependent ECM genes expression. As expected, mRNA levels of the fibrotic ECM genes Collagen I, Collagen III, Fibronectin and α-SMA were increased in the TGF-β treated CFs (Fig. 1c, d). Furthermore, TGF-β stimulation greatly augmented the expression of these ECM genes in HIP-55 knockdown CFs (Fig. 1c). And in contrast, the expression of these ECM genes was inhibited in HIP-55-overexpressing CFs upon TGF-β treatment (Fig. 1d). Consistent with the results of IB analysis, the qRT-PCR results indicated that HIP-55 sensed and responded to TGF-β stimulation in qRT-PCR analysis (Fig. 1c, d). Together, these data demonstrate that HIP-55 might modulate TβRI and TGF-β signalling to attenuate pathological effects in a receptor subtype-dependent manner.
HIP-55 represses TβRI protein levels and TGF-β signalling in a Smad7-dependent manner
As mentioned above, the negative regulatory effect of HIP-55 on TGF-β signalling may repress TβRI but not TβRII. This subtype-dependent form of regulation may rely on specific proteins, such as the inhibitory protein Smad7, that can repress the TβRI protein level without affecting TβRII [12, 31]. As expected, upon TGF-β treatment, HIP-55 had no inhibitory effect on the TβRI protein level or TGF-β signalling activation when Smad7 was silenced (Fig. 2a). Notably, Smad7 decreased P-smad2 and P-smad3 levels and slightly increased TβRI protein levels without HIP-55, in line with previous reports. Moreover, no significant changes in the expression of Collagen I, Collagen III, Fibronectin or α-SMA, fibrotic ECM genes linked to pathological effects, were observed when Smad7 was silenced (Fig. 2b). Together, these results indicate that Smad7 is required for the negative regulation of TβRI and TGF-β signalling by HIP-55.
a Representative immunoblotting (IB) and quantification of TβRI, P-Smad2, T-smad2, P-smad3, T-smad3, Smad7 and HIP-55 protein levels in cultured cardiac fibroblasts (CFs) infected with Scrambled, Smad7 siRNA, Dsred or Dsred-HIP-55 and then treated with TGF-β (5 ng/mL) for 3 h. b qRT-PCR analysis of Collagen I, Collagen III, Fibronectin, α-SMA, Smad7 and HIP-55 mRNA expression in cultured CFs infected with Scrambled, Smad7 siRNA, Dsred or Dsred-HIP-55 and then treated with TGF-β (5 ng/mL) for 24 h. GAPDH was used as an internal control, and the values are the means ± SEMs of five independent experiments (n = 5). *P < 0.05, **P < 0.01, ***P < 0.001 vs. the control group; #P < 0.05, ##P < 0.01, ###P < 0.001 vs. the TGF-β group; &P < 0.05, &&&P < 0.001 vs. the Dsred-HIP-55 group; $P < 0.05, $$P < 0.01, $$$P < 0.001 vs. the Dsred-HIP-55 + TGF-β group; aP < 0.05, aaaP < 0.001 vs. the Smad7 siRNA group; bbbP < 0.001 vs. the Smad7 siRNA + TGF-β group; ns no significance (P > 0.05); one-way ANOVA with Tukey’s post hoc multiple comparisons for (a, b).
HIP-55 interacts with Smad7 and inhibits Axin/Smad7 complex formation to maintain Smad7 stability
We further explored the underlying mechanistic relationship between HIP-55 and Smad7. The upregulation of HIP-55 elevated Smad7 protein levels both under basal conditions and upon TGF-β stimulation (Fig. 3a, b). However, Smad7 mRNA levels were not significantly changed in either the HIP-55-knockdown or HIP-55-overexpressing CFs (Fig. 3c, d), indicating that the relationship between HIP-55 and Smad7 does not occur at the transcriptional level. We further detected the degradation of Smad7 by ubiquitination and turnover experiments. HIP-55 showed the strong capacity to maintain the half-life of Smad7 in CFs (Fig. 3e, f). Moreover, the overexpression of HIP-55 inhibited Smad7 ubiquitination, especially upon TGF-β exposure (Fig. 3g). These results demonstrated that HIP-55 maintains Smad7 stability by inhibiting the proteasome-mediated degradation pathway.
a, b Representative immunoblotting (IB) and quantification of Smad7 and HIP-55 protein levels in cultured cardiac fibroblasts (CFs). c, d qRT-PCR analysis of Smad7 and HIP-55 mRNA expression in cultured CFs. e, f Representative IB of Smad7 and HIP-55 protein levels in each group and quantification of Smad7 protein levels in cultured CFs treated with CHX (50 μg/mL) for the indicated time periods. g Typical Western blot (WB) showing the levels of ubiquitinated Smad7. Lysates from HEK293T cells transfected with the indicated plasmids were treated with MG132 (5 μmol) for 3 h and then treated with TGF-β (5 ng/mL) for 3 h. GAPDH was used as an internal control, and the values are the means ± SEMs of five independent experiments (n = 5). *P < 0.05, **P < 0.01, ***P < 0.001 vs. the Scrambled group or the Dsred group; ##P < 0.01, ###P < 0.001 vs. the Scrambled + TGF-β group or the Dsred + TGF-β group; &P < 0.05, &&P < 0.01 vs. the ShRNA-HIP-55 group or the Dsred-HIP-55 group; ns no significance (P > 0.05); two-way ANOVA with Tukey’s post hoc multiple comparisons for (a–d). *P < 0.05, **P < 0.01, vs. the control group; two-way ANOVA with Tukey’s post hoc multiple comparisons for (e, f).
One of the major mechanisms underlying the degradation of Smad7 operates through binding with Axin, which recruits the ubiquitin E3 ligase Arkadia [14, 15]. We found by co-IP assays that the endogenous HIP-55 and Smad7 proteins could interact (Fig. 4a), and this interaction was dramatically enhanced by TGF-β treatment (Fig. 4b). Furthermore, when HIP-55 was knocked down, Axin/Smad7 complex formation was increased under basal conditions (Fig. 4c, lane 5 and lane 7 for IB: Axin) and even more so under TGF-β-treatment conditions (Fig. 4c, lane 6 and lane 8 for IB: Axin). The binding of HIP-55 and Smad7 was further confirmed in HEK293T cells overexpressing HIP-55 that were treated with MG132 (Fig. 4d). Remarkably, the interaction between HIP-55 and Smad7 was stronger than that between Axin and Smad7 upon TGF-β stimulation, implying the protective effect of HIP-55 against TGF-β-related diseases due to its high affinity as an adaptor protein, particularly upon pathological stimulation.
a Endogenous co-immunoprecipitation (co-IP) assay to detect Smad7 and HIP-55 in HEK293T cell lysates by Western blotting (WB) with the indicated antibodies. b Endogenous Smad7 and HIP-55 co-IP assay in HEK293T cell lysates after treatment with TGF-β (5 ng/mL) for 3 h before analysis by WB with the indicated antibodies. c Typical Western blot following the immunoprecipitation (IP) of Smad7 and Axin. Lysates from stable HEK293A cells were treated with MG132 (5 μmol) for 3 h and then treated with TGF-β (5 ng/mL) for 3 h before analysis by WB with the indicated antibodies. d Typical WB following a competitive IP assay to assess the relationship between HIP-55 and Axin to Smad7. Lysates from HEK293T cells transfected with the indicated plasmids, treated with MG132 (5 μmol) for 3 h and then treated with TGF-β (5 ng/mL) for 3 h were analysed by WB with anti-Axin or anti-HIP-55. e Typical WB showing the levels of ubiquitinated Smad7 upon competitive interaction. Lysates from HEK293T cells transfected with the indicated plasmids, treated with MG132 (5 μmol) for 3 h and then treated with TGF-β (5 ng/mL) for 3 h were analysed by WB with anti-HA (Ub).
To dissect the effect of this interaction, we performed a ubiquitination assay in which HIP-55 and Axin competed to interact with Smad7. The ubiquitination of Smad7 was ameliorated when Smad7 was bound to HIP-55 (Fig. 4e, lane 1 and lane 2 for IB:HA) but enhanced when Smad7 interacted with Axin (Fig. 4e, lane 1 and lane 3 for IB:HA) in cells were treated with MG132. This effect of HIP-55 was stronger upon TGF-β stimulation (Fig. 4e, lane 7 and lane 8 for IB:HA). Overall, our results suggest that HIP-55 can interact with Smad7 and compete with Smad7/Axin complex formation to inhibit the Axin-mediated ubiquitination and degradation of Smad7.
HIP-55 couples Smad7 to accelerate TβRI degradation
To elucidate the mechanism underlying the integrated, multicomponent system by which TGF-β signalling is negatively regulated, we found that endogenous HIP-55 and TβRI could also interact (Fig. 5a). Smad7 was diffusively distributed in the cytoplasm and nucleus and showed substantial colocalization with HIP-55 in the cytoplasm under basal conditions. After TGF-β exposure, a substantial amount of HIP-55 docked near the plasma membrane and then recruited Smad7 to the plasma membrane. Smad7 could not move to the plasma membrane upon TGF-β treatment when HIP-55 had been deleted (Fig. 5b). Next, we used an IP assay to unravel the effects of HIP-55-mediated regulation of the Smad7 subcellular localisation. As suggested by the results in Fig. 5c, d, HIP-55 stabilised the Smad7/TβRI complex, and this interaction was enhanced upon TGF-β stimulation when the cells were treated with MG132. Thus, HIP-55 can control Smad7 subcellular localisation and promote Smad7/TβRI complex formation.
a Endogenous TβRI and HIP-55 co-immunoprecipitation (co-IP) assay in HEK293T cell lysates, which were then analysed by Western blotting (WB) with the indicated antibodies. b Endogenous Smad7 and HIP-55 colocalization in stable HEK293A cells treated with TGF-β (5 ng/mL) for 3 h before fixation and staining. Staining with anti-HIP-55 (red) and anti-Smad7 (green) and nuclear staining with Hoechst were performed (scale bar, 5 μm). c Immunoprecipitation (IP) assay to detect Smad7 and TβRI in HEK293T cells. Lysates from HEK293T cells transfected with the indicated plasmids, treated with MG132 (5 μmol) for 3 h and then treated with TGF-β (5 ng/mL) for 3 h were analysed by WB with anti-TβRI. d Smad7 and TβRI IP assay in stable HEK293A cells. Lysates from stable HEK293 cells transfected with the indicated plasmids, treated with MG132 (5 μmol) for 3 h and then treated with TGF-β (5 ng/mL) for 3 h were analysed by WB with anti-TβRI.
As expected, HIP-55 showed a strong capacity to shorten the half-life of TβRI (Fig. 6a, b). Moreover, the overexpression of HIP-55 enhanced TβRI ubiquitination, especially upon TGF-β stimulation (Fig. 6c). No difference in TβRI half-life upon Smad7 silencing was observed between the Dsred group and Dsred-HIP-55 group (Fig. 6d). This finding was further supported by the effects of Smad7 overexpression in HEK293A cells with stable HIP-55 knockdown. The results showed that HIP-55 promotes TβRI ubiquitination and degradation in a Smad7-dependent manner (Fig. 6e). Altogether, the results suggest that HIP-55 binds Smad7, accelerating TβRI degradation.
a, b Representative immunoblotting (IB) to detect TβRI and HIP-55 protein levels in each group and quantification of TβRI protein levels in cultured cardiac fibroblasts (CFs) treated with CHX (50 μg/mL) for the indicated time periods. c Typical Western blot (WB) showing the levels of ubiquitinated TβRI. Lysates from HEK293T cells transfected with the indicated plasmids, treated with MG132 (5 μmol) for 3 h and then treated with TGF-β (5 ng/ml) for 3 h were analysed by WB with anti-HA (Ub). d Representative IB analysis of TβRI and HIP-55 protein levels in each group and quantification of TβRI protein levels in CFs treated with CHX (50 μg/mL) for the indicated time periods after Smad7 knockdown. e Typical WB showing the levels of ubiquitinated TβRI after Smad7 knockdown. Lysates from stable HEK293A cells transfected with the indicated plasmids, treated with MG132 (5 μmol) for 3 h and then treated with TGF-β (5 ng/mL) for 3 h were analysed by WB with anti-HA (Ub). GAPDH was used as an internal control, and the values are the means ± SEMs of five independent experiments (n = 5). *P < 0.05, **P < 0.01, vs. the control group; ns no significance (P > 0.05); two-way ANOVA with Tukey’s post hoc multiple comparisons for (a, b, d).
Discussion
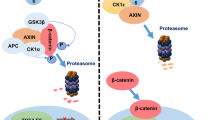
The TGF-β signalling system is extremely powerful, and an elaborate network of regulators maintain the homoeostasis of this system [1, 5]. Our work reveals a new mechanism by which a multifunctional adaptor protein may function as a hub to integrate and coordinate the negative feedback regulation of TGF-β signalling. As a multidomain adaptor protein, HIP-55 serves as a platform for a molecular assembly consisting of scaffold protein, receptors, and E3 ligases to stabilise Smad7, leading to receptor degradation. A schematic diagram of the roles of HIP-55 in the negative feedback regulation of TGF-β signalling is shown in Fig. 7. HIP-55 accumulates upon the pathological response to TGF-β1 and forms the HIP-55/Smad7 complex, which inhibits the Axin/Arkadia-mediated ubiquitin-dependent degradation of Smad7. Then, HIP-55 recruits Smad7 to the plasma membrane, triggering TβRI degradation and maintaining homoeostasis of the TGF-β signalling pathway. Because of its roles as an integrator and coordinator, HIP-55 may be a target for TGF-β signalling-related diseases.
TGF-β1 stimulation induces the accumulation of HIP-55, which interacts with Smad7 and blocks formation of the Smad7/Axin complex to maintain Smad7 stability. Surplus Smad7 is recruited by HIP-55 to the plasma membrane, triggering TβRI degradation, which maintains TGF-β signalling homoeostasis. Because of its roles as an integrator and coordinator, HIP-55 may be a target for TGF-β signalling-related diseases (Ub, ubiquitin).
The TGF-β signalling pathway has great plasticity and adapts to particular physiological and pathological conditions by negative feedback, controlling finely tuned signal transduction and cellular responses [1, 5, 9, 10]. Smad7 is critical for the negative feedback regulation of TGF-β signalling as it targets TβRI for degradation [11, 31]. The stability and activity of Smad7 are also finely controlled by ubiquitination mediated by the E3 ubiquitin ligases Arkadia, RNF12 and Cbl-b [15, 32, 33]. Some posttranslational modifications also modulate Smad7 stability, including MPK38-induced phosphorylation, p300-mediated acetylation, HDAC1/Sirt1-triggered deacetylation and SET9-mediated methylation [34,35,36,37]. Among these modifications, Arkadia-triggered ubiquitination is the major way by which Smad7 stability is modulated. Axin interacts with Smad7, inducing the nuclear export of Smad7, which then constitutively recruits Arkadia to degrade Smad7 [14, 15].
In this study, we have revealed how HIP-55 maintains Smad7 stability by preventing Axin/Arkadia-mediated Smad7 degradation. We found that Smad7 can selectively bind HIP-55 or Axin, and this differential binding bestows different fates to Smad7, including its degradation and regulation of TGF-β receptors. The binding of Smad7 to HIP-55 may generate an optimal configuration for interaction with TβRI and drive receptor degradation. However, the binding of Smad7 to Axin may generate an optimal configuration for interaction with Arkadia, promoting Smad7 ubiquitination and degradation [14, 15]. The Smad7/HIP-55 complex is more refractory to TGF-β stimulation than the Smad7/Axin complex, which maintains the negative feedback regulation of TGF-β signalling. The competitive interaction between Smad7/HIP-55 and Smad7/Axin might involve similar domains in Smad7, or the dimensional structure of Smad7 might change after it interacts with HIP-55, impairing its capacity to bind Axin. Future studies will be required to obtain structural insights into the interaction between HIP-55 and Smad7 under different physiological and pathological circumstances, especially to determine the domains and sites involved in binding, which may preferentially promote Smad7/HIP-55 complex formation. However, Smad7 also appears to be a substrate of Smurf1/Smurf2, which are E3 ligases that act on TβRI [12, 13, 31]. Smad7 that escapes the Axin-mediated degradation system might be transmitted to the TβRI degradation system to ultimately degrade receptors via HIP-55. Another explanation is that the HIP-55/Smad7 complex recruits another E3 ligase, WW domain-containing protein 1 (WWP1), to degrade TβRI since WWP1 can promote TβRI degradation without affecting the stability of Smad7 [38, 39].
Several regulators have been reported to negatively regulate TGF-β signalling by modulating TβRI activity and stability. BMP and activin membrane-bound inhibitor (BAMBI) and the E3 ligase AIP4/Itch can form a ternary complex with TβRI and Smad7 upon TGF-β stimulation, impeding the recruitment of R-Smad to receptors and further inhibiting TGF-β signalling [40, 41]. The adaptor protein Toll interacting protein (Tollip), Salt-inducible kinase (SIK) and TGF-β-stimulated clone 22 (TSC-22) can regulate TβRI degradation through Smad7/Smurfs [42,43,44]. In contrast to these regulators, the multifunctional adaptor protein HIP-55 utilises a new regulatory mechanism in which it integrates multiple components for the negative feedback regulation of TGF-β signalling. HIP-55 damaged the Axin-mediated degradation of Smad7, sustaining Smad7-mediated degradation of TβRI, showing the diverse and flexible functions of HIP-55 in two TGF-β-related degradation systems. HIP-55 is simultaneously connected to several components along the same route during negative feedback regulation of TGF-β signalling. The subcellular localisation of Smad7 is responsive to TGF-β stimulation, which may involve microtubules and associated motor proteins in the cytoskeleton [45, 46]. HIP-55 can move to different cellular compartments, which may engage other proteins. As an actin-binding protein, HIP-55 may serve as a structural platform distributed along the cytoskeleton that also anchors Smad7 in the plasma membrane [47].
We propose that HIP-55 is a promising target for TGF-β signalling-related diseases in precise medicine. Endogenous TGF-β signalling is essential for the development of pathological conditions and regulates matrix deposition in many organs [6]. TGF-β enables cancer cell invasion and metastasis, stem cell properties and tumour therapeutic resistance [48]. TGF-β also controls adaptive immunity and the innate immune system by regulating the complex behaviour of regulatory T (Treg) cells, T cells, natural killer cells, macrophages and neutrophils, thus forming a negative immunoregulatory network [2]. Because of the broad effects of TβRI inhibitors on TGF-β signalling and the important role of TβRI in maintaining tissue homoeostasis, TβRI inhibitors may not be a successful strategy to protect against TGF-β signalling-related diseases [49, 50]. Negative feedback is an important mechanism that inhibits the overactivation of TGF-β signalling in the early stage of disease [51, 52]. Thus, a suitable therapeutic strategy targeting negative TGF-β signalling regulation is promising. In our work, we found that HIP-55 could stabilise Smad7 and couple Smad7 to TβRI, but not TβRII, accelerating TβRI degradation. Selective targeting with this subtype-specific mechanism may bring more effective and less toxic treatments. Targeting HIP-55 may provide novel strategies for the treatment of TGF-β signalling-related diseases.
References
Budi EH, Duan D, Derynck R. Transforming growth factor-β receptors and smads: regulatory complexity and functional versatility. Trends Cell Biol. 2017;27:658–72.
Batlle E, Massagué J. Transforming growth factor-β signaling in immunity and cancer. Immunity. 2019;50:924–40.
Hata A, Chen YG. TGF-β signaling from receptors to smads. Cold Spring Harb Perspect Biol. 2016;8:a022061.
Gu L, Zhu YJ, Yang X, Guo ZJ, Xu WB, Tian XL. Effect of TGF-beta/Smad signaling pathway on lung myofibroblast differentiation. Acta Pharmacol Sin. 2007;28:382–91.
Derynck R, Budi EH. Specificity, versatility, and control of TGF-β family signaling. Sci Signal. 2019;12:eaav5183.
Hu HH, Chen DQ, Wang YN, Feng YL, Cao G, Vaziri ND, et al. New insights into TGF-β/Smad signaling in tissue fibrosis. Chem Biol Interact. 2018;292:76–83.
Mauviel A. Transforming growth factor-beta: a key mediator of fibrosis. Methods Mol Med. 2005;117:69–80.
Kang JS, Liu C, Derynck R. New regulatory mechanisms of TGF-beta receptor function. Trends Cell Biol. 2009;19:385–94.
Miyazono K. Positive and negative regulation of TGF-beta signaling. J Cell Sci. 2000;113:1101–9.
Yan X, Xiong X, Chen YG. Feedback regulation of TGF-β signaling. Acta Biochim Biophys Sin. 2018;50:37–50.
Nakao A, Afrakhte M, Morén A, Nakayama T, Christian JL, Heuchel R, et al. Identification of Smad7, a TGFbeta-inducible antagonist of TGF-beta signalling. Nature. 1997;389:631–5.
Ebisawa T, Fukuchi M, Murakami G, Chiba T, Tanaka K, Imamura T, et al. Smurf1 interacts with transforming growth factor-beta type I receptor through Smad7 and induces receptor degradation. J Biol Chem. 2001;276:12477–80.
Kavsak P, Rasmussen RK, Causing CG, Bonni S, Zhu H, Thomsen GH, et al. Smad7 binds to Smurf2 to form an E3 ubiquitin ligase that targets the TGF beta receptor for degradation. Mol Cell. 2000;6:1365–75.
Liu W, Rui H, Wang J, Lin S, He Y, Chen M, et al. Axin is a scaffold protein in TGF-beta signaling that promotes degradation of Smad7 by Arkadia. EMBO J. 2006;25:1646–58.
Koinuma D, Shinozaki M, Komuro A, Goto K, Saitoh M, Hanyu A, et al. Arkadia amplifies TGF-beta superfamily signalling through degradation of Smad7. EMBO J. 2003;22:6458–70.
Hayashi H, Abdollah S, Qiu Y, Cai J, Xu YY, Grinnell BW, et al. The MAD-related protein Smad7 associates with the TGFbeta receptor and functions as an antagonist of TGFbeta signaling. Cell. 1997;89:1165–73.
Hanyu A, Ishidou Y, Ebisawa T, Shimanuki T, Imamura T, Miyazono K. The N domain of Smad7 is essential for specific inhibition of transforming growth factor-beta signaling. J Cell Biol. 2001;155:1017–27.
Mochizuki T, Miyazaki H, Hara T, Furuya T, Imamura T, Watabe T, et al. Roles for the MH2 domain of Smad7 in the specific inhibition of transforming growth factor-beta superfamily signaling. J Biol Chem. 2004;279:31568–74.
Ensenat D, Yao Z, Wang XS, Kori R, Zhou G, Lee SC, et al. A novel src homology 3 domain-containing adaptor protein, HIP-55, that interacts with hematopoietic progenitor kinase 1. J Biol Chem. 1999;274:33945–50.
Kessels MM, Engqvist-Goldstein AE, Drubin DG. Association of mouse actin-binding protein 1 (mAbp1/SH3P7), an Src kinase target, with dynamic regions of the cortical actin cytoskeleton in response to Rac1 activation. Mol Biol Cell. 2000;11:393–412.
Kessels MM, Engqvist-Goldstein AE, Drubin DG, Qualmann B. Mammalian Abp1, a signal-responsive F-actin-binding protein, links the actin cytoskeleton to endocytosis via the GTPase dynamin. J Cell Biol. 2001;153:351–66.
Xu W, Stamnes M. The actin-depolymerizing factor homology and charged/helical domains of drebrin and mAbp1 direct membrane binding and localization via distinct interactions with actin. J Biol Chem. 2006;281:11826–33.
Seeley-Fallen MK, Liu LJ, Shapiro MR, Onabajo OO, Palaniyandi S, Zhu X, et al. Actin-binding protein 1 links B-cell antigen receptors to negative signaling pathways. Proc Natl Acad Sci USA. 2014;111:9881–6.
Le Bras S, Foucault I, Foussat A, Brignone C, Acuto O, Deckert M. Recruitment of the actin-binding protein HIP-55 to the immunological synapse regulates T cell receptor signaling and endocytosis. J Biol Chem. 2004;279:15550–60.
Boateng LR, Bennin D, De Oliveira S, Huttenlocher A. Mammalian Actin-binding Protein-1/Hip-55 Interacts with FHL2 and Negatively Regulates Cell Invasion. J Biol Chem. 2016;291:13987–98.
Mavrakis KJ, Andrew RL, Lee KL, Petropoulou C, Dixon JE, Navaratnam N, et al. Arkadia enhances Nodal/TGF-beta signaling by coupling phospho-Smad2/3 activity and turnover. PLoS Biol. 2007;5:e67.
Fiedler M, Mendoza-Topaz C, Rutherford TJ, Mieszczanek J, Bienz M. Dishevelled interacts with the DIX domain polymerization interface of Axin to interfere with its function in down-regulating β-catenin. Proc Natl Acad Sci USA. 2011;108:1937–42.
Tang M, Li Z, Zhang C, Lu X, Tu B, Cao Z, et al. SIRT7-mediated ATM deacetylation is essential for its deactivation and DNA damage repair. Sci Adv. 2019;5:eaav1118.
Wang W, Li W, Liu K, Niu X, Guan K, Jiang Y, et al. Src mediates β-adrenergic receptor induced YAP tyrosine phosphorylation. Sci China Life Sci. 2020;63:697–705.
Dobaczewski M, Chen W, Frangogiannis NG. Transforming growth factor (TGF)-β signaling in cardiac remodeling. J Mol Cell Cardiol. 2011;51:600–6.
Miyazawa K, Miyazono K. Regulation of TGF-β Family Signaling by Inhibitory Smads. Cold Spring Harb Perspect Biol. 2017;9:a022095.
Zhang L, Huang H, Zhou F, Schimmel J, Pardo CG, Zhang T, et al. RNF12 controls embryonic stem cell fate and morphogenesis in zebrafish embryos by targeting Smad7 for degradation. Mol Cell. 2012;46:650–61.
Gruber T, Hinterleitner R, Hermann-Kleiter N, Meisel M, Kleiter I, Wang CM, et al. Cbl-b mediates TGFβ sensitivity by downregulating inhibitory SMAD7 in primary T cells. J Mol Cell Biol. 2013;5:358–68.
Seong HA, Jung H, Ha H. Murine protein serine/threonine kinase 38 stimulates TGF-beta signaling in a kinase-dependent manner via direct phosphorylation of Smad proteins. J Biol Chem. 2010;285:30959–70.
Grönroos E, Hellman U, Heldin CH, Ericsson J. Control of Smad7 stability by competition between acetylation and ubiquitination. Mol Cell. 2002;10:483–93.
Kume S, Haneda M, Kanasaki K, Sugimoto T, Araki S, Isshiki K, et al. SIRT1 inhibits transforming growth factor beta-induced apoptosis in glomerular mesangial cells via Smad7 deacetylation. J Biol Chem. 2007;282:151–8.
Elkouris M, Kontaki H, Stavropoulos A, Antonoglou A, Nikolaou KC, Samiotaki M, et al. SET9-mediated regulation of TGF-β signaling links protein methylation to pulmonary fibrosis. Cell Rep. 2016;15:2733–44.
Morén A, Imamura T, Miyazono K, Heldin CH, Moustakas A. Degradation of the tumor suppressor Smad4 by WW and HECT domain ubiquitin ligases. J Biol Chem. 2005;280:22115–23.
Komuro A, Imamura T, Saitoh M, Yoshida Y, Yamori T, Miyazono K, et al. Negative regulation of transforming growth factor-beta (TGF-beta) signaling by WW domain-containing protein 1 (WWP1). Oncogene. 2004;23:6914–23.
Yan X, Lin Z, Chen F, Zhao X, Chen H, Ning Y, et al. Human BAMBI cooperates with Smad7 to inhibit transforming growth factor-beta signaling. J Biol Chem. 2009;284:30097–104.
Lallemand F, Seo SR, Ferrand N, Pessah M, L’Hoste S, Rawadi G, et al. AIP4 restricts transforming growth factor-beta signaling through a ubiquitination-independent mechanism. J Biol Chem. 2005;280:27645–53.
Zhu L, Wang L, Luo X, Zhang Y, Ding Q, Jiang X, et al. Tollip, an intracellular trafficking protein, is a novel modulator of the transforming growth factor-β signaling pathway. J Biol Chem. 2012;287:39653–63.
Kowanetz M, Lönn P, Vanlandewijck M, Kowanetz K, Heldin CH, Moustakas A. TGFbeta induces SIK to negatively regulate type I receptor kinase signaling. J Cell Biol. 2008;182:655–62.
Yan X, Zhang J, Pan L, Wang P, Xue H, Zhang L, et al. TSC-22 promotes transforming growth factor β-mediated cardiac myofibroblast differentiation by antagonizing Smad7 activity. Mol Cell Biol. 2011;31:3700–9.
Ekman M, Mu Y, Lee SY, Edlund S, Kozakai T, Thakur N, et al. APC and Smad7 link TGFβ type I receptors to the microtubule system to promote cell migration. Mol Biol Cell. 2012;23:2109–21.
Zhu S, Goldschmidt-Clermont PJ, Dong C. Transforming growth factor-beta-induced inhibition of myogenesis is mediated through Smad pathway and is modulated by microtubule dynamic stability. Circ Res. 2004;94:617–25.
He K, Xing R, Yan X, Tian A, Zhang M, Yuan J, et al. Mammalian actin-binding protein 1/HIP-55 is essential for the scission of clathrin-coated pits by regulating dynamin-actin interaction. FASEB J. 2015;29:2495–503.
Derynck R, Turley SJ, Akhurst RJ. TGFβ biology in cancer progression and immunotherapy. Nat Rev Clin Oncol. 2021;18:9–34.
Petersen M, Thorikay M, Deckers M, van Dinther M, Grygielko ET, Gellibert F, et al. Oral administration of GW788388, an inhibitor of TGF-beta type I and II receptor kinases, decreases renal fibrosis. Kidney Int. 2008;73:705–15.
Schiemann WP, Rotzer D, Pfeifer WM, Levi E, Rai KR, Knaus P, et al. Transforming growth factor-beta (TGF-beta)-resistant B cells from chronic lymphocytic leukemia patients contain recurrent mutations in the signal sequence of the type I TGF-beta receptor. Cancer Detect Prev. 2004;28:57–64.
Novák B, Tyson JJ. Design principles of biochemical oscillators. Nat Rev Mol Cell Biol. 2008;9:981–91.
Oshimori N, Fuchs E. The harmonies played by TGF-β in stem cell biology. Cell Stem Cell. 2012;11:751–64.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (91939301, 81820108031, 91539123 and 81471893) and the Beijing Municipal Natural Science Foundation (7172235). The authors also thank Dr. Kang-min He at the Chinese Academy of Sciences for providing the pcDNA3.1-Flag-Smad7 and pcDNA3.1-myc-TβRI plasmids.
Author information
Authors and Affiliations
Contributions
YS: planned and performed the experiments, analysed the data and drafted the paper. ZJL: the principal investigator, conceived the scientific ideas, oversaw the project, designed the experiments and refined the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Sun, Y., Li, Zj. The multifunctional adaptor protein HIP-55 couples Smad7 to accelerate TGF-β type I receptor degradation. Acta Pharmacol Sin 43, 634–644 (2022). https://doi.org/10.1038/s41401-021-00741-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-021-00741-1
- Springer Nature Singapore Pte Ltd.