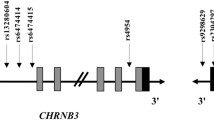
Abstract
Nicotine dependence (ND) is a worldwide health problem. Numerous genetic studies have demonstrated a significant association of variants in nicotinic acetylcholine receptors (nAChRs) with smoking behaviors. However, most of these studies enrolled only subjects of European or African ancestry. In addition, although an increasing body of evidence implies a causal connection of single-nucleotide polymorphisms (SNPs) and epigenetic regulation of gene expression, few studies of this issue have been reported. In this study, we performed both association and interaction analysis for 67 SNPs in CHRNA3-A5, CHRNA7, CHRNB2, and CHRNB4 with ND in a Chinese Han population (N = 5055). We further analyzed cis-mQTL for the three most significant SNPs and 5580 potential methylation loci within these target gene regions. Our results indicated that the SNPs rs1948 and rs7178270 in CHRNB4 and rs3743075 in CHRNA3 were significantly associated with the Fagerström Test for Nicotine Dependence (FTND) score (p = 6.6 × 10−5; p = 2.0 × 10−4, and p = 7.0 × 10−4, respectively). Haplotype-based association analysis revealed that two major haplotypes, T-G and C-A, formed by rs3743075–rs3743074 in CHRNA3, and other two major haplotypes, A-G-C and G-C-C, formed by rs1948–rs7178270–rs17487223 in CHRNB4, were significantly associated with the FTND score (p ≤ 8.0 × 10−4). Further, we found evidence for the presence of significant interaction among variants within CHRNA3/B4/A5, CHRNA4/B2/A5, and CHRNA7 in affecting ND, with corresponding p values of 5.8 × 10−6, 8.0 × 10−5, and 0.012, respectively. Finally, we identified two CpG sites (CpG_2975 and CpG_3007) in CHRNA3 that are significantly associated with three cis-mQTL SNPs (rs1948, rs7178270, rs3743075) in the CHRNA5/A3/B4 cluster (p ≤ 1.9 × 10−6), which formed four significant CpG–SNP pairs in our sample. Together, we revealed at least three novel SNPs in CHRNA3 and CHRNB4 to be significantly associated with the FTND score. Further, we showed that these significant variants contribute to ND via two methylated sites, and we demonstrated significant interaction affecting ND among variants in CHRNA5/A3/B4, CHRNA7, and CHRNA4/B2/A5. In sum, these findings provide robust evidence that SNPs in nAChR genes convey a risk of ND in the Chinese Han population.
Similar content being viewed by others
Introduction
Tobacco smoking is a major public health problem that causes nearly 6 million deaths worldwide every year.1 Because of the lack of effective treatment for smoking addiction, the addictive properties of nicotine in tobacco smoke, and lack of awareness of the consequences of smoking in many regions, the worldwide death toll caused by cigarette smoking might well reach 10 million annually by 2020.2 Although many developed countries have implemented regulations and laws against tobacco smoking, which have led to dramatic reductions in smoking during recent years, smoking remains a significant issue in many developing countries, especially in Asia.3,4 For example, the prevalence of smoking in Chinese men aged 15 or older is an estimated 52.1%,5 meaning that male Chinese smokers account for almost one of third of the total number of smokers in the world.6
Although environmental factors play an important role in nicotine dependence (ND),7 genetics is another important component, as ND has an average heritability of 0.56.7 Of the identified susceptibility genes for ND, the most investigated ones are members of the nicotinic acetylcholine receptor (nAChR) gene family, which encodes 12 subunits (i.e., α2–α7; β2–β4) that are widely expressed in many brain regions.8,9,10 Nicotine, a primary component of tobacco smoke, exerts its biological effects on these nAChRs, where it either stimulates them or inactivates them through desensitization.11
Numerous studies using approaches such as genome-wide linkage, candidate gene association, and genome-wide association (GWAS) analysis have greatly advanced our knowledge of the genetic architecture underlying ND.12 The most replicated susceptibility loci for ND are nAChR subunit genes in the CHRNA5/A3/B4 cluster on chromosome 15,12,13,14,15,16,17,18,19,20,21,22,23,24 CHRNB3/A6 on chromosome 8,12,25 and CYP2A6/A7 on chromosome 19.12,14,25 In addition, a significant association has been reported of variants in the CHRNA5/A3/B4 cluster with ND and lung cancer.18
Although the reported associations of variants in CHRNA7 with ND have not been replicated consistently in independent samples, pharmacological and molecular studies have strongly implicated α7 as an important target for ND and smoking cessation.26 Knockout α7 mice show a greater preference for oral nicotine than do their wild-type counterparts.27 In addition, local infusion of a highly selective antagonist (α-conotoxin ArIB) of α7 into the nucleus accumbens (NAc) shell or the anterior cingulate cortex increases nicotine self-administration.28
Numerous studies have shown that the β2 subunit is the most abundant of the nAChRs expressed in the brain and has high affinity for nicotine and acetylcholine (Ach). Further, many studies have demonstrated that the β2* (* indicates non-β2 subunits forming a functional nAChR by combining with β2) is required for nicotine reinforcement and reward.29 In the studies of therapeutics for smoking cessation, both β2-containing and α7-containing nicotine receptors have proved to be targets to curb tobacco addiction. However, α7 appears to act as modulator of nicotine reinforcement in opposition to α4β2* and α6β2*. Melis et al.30 showed a modulatory role for α7 in the ventral tegmental area to reduce β2* activation, especially for α4β2* and α6β2*, by stimulating the intracellular signaling cascades that inhibit β2*. These findings indicate that both α7 and β2* activation play an essential role in ND and smoking cessation.
Exogenous factors such as cigarette smoking can alter DNA methylation either locally or globally.31 Persistent exposure to smoke stimulates epigenetic reprogramming at the global level by affecting the methylation of repetitive elements.32 Although methylation is also under genetic influence, allele-specific DNA methylation is often correlated in related individuals.33 It is thus plausible to infer a correlation between these aberrant CpGs and risk variants for cigarette smoking.
A diversity of susceptibility variants for smoking has been identified, but the mechanisms by which these SNPs contribute to smoking-related traits are generally unclear. Benefits from high-throughput next-generation sequencing and high-density array platforms have allowed researchers to find regulatory variants by mapping expression and methylation quantitative trait loci (meQTLs).34,35 This approach provides a better way to reveal the mechanisms of significant variants from association studies. For example, considering the chromosome region of 15q25.1 as a well-established susceptibility region for smoking-related phenotypes, Hancock et al.36 assessed the number of methylation loci in the region based on the Illumina HumanMethylation27 array, which led to identification of a novel regulatory SNP, rs11636753, in CHRNA5 that modulates methylation and expression in ND-relevant brain regions in multiancestry groups. Because the Illumina array contains methylation loci only from promoter regions,37 it is necessary to perform a fine-mapping analysis of this region to detect more cis-meQTLs.
There were three objectives of this study: (1) to determine which individual SNPs or haplotypes in nAChR subunit genes are associated with ND in a Chinese Han smoker sample, a less commonly investigated population; (2) to detect significant interactive effects among these genes in exerting influence on the etiology of ND; and (3) to link risk variants for ND and differential DNA methylation loci by cis-mQTLs analysis to explore the underlying mechanisms involved in ND.
Materials and methods
Subjects
This study included 5055 unrelated subjects consisting of both smokers (N = 2616) and non-smokers (N = 2439), who were recruited from local hospitals in Jincheng and Taiyuan in Shanxi Province of China in 2013. Participants with clinically diagnosed psychiatric disorders such as schizophrenia, Alzheimer’s disease, and major depressive disorder were excluded. Because few Chinese women aged 15 years or older smoke (~2.7%),5 only male smokers were included.
A set of structured questionnaires on cigarette smoking; demographic information such as age, education, and annual income; drug or substance use history; and neighborhood environment were administered to each participant by trained researchers. The detailed demographic characteristics of this sample are shown in Table 1. The average age of the smokers was 40.4 ± 9.7 years and that of the non-smokers was 37.0 ± 10.9 years. Written informed consent was provided by each participant after receiving a detailed explanation of the project and process of this study. The study and all the questionnaires used in the study were approved by the Ethics Committee of First Affiliated Hospital of Zhejiang University School of Medicine.
Phenotype assessment
Non-smokers were defined as persons who had smoked fewer than 100 cigarettes in their lifetimes. Smoking dependence was assessed by the Fagerström Test for Nicotine Dependence (FTND) measure (0–10 scale).38,39 We defined the smokers with an FTND score of ≥6 as heavy smokers (N = 1243) and those with an FTND score of <6 as light smokers (N = 1373) (Supplementary Table 6).40
Selection of SNPs for genotyping analysis
Peripheral blood was collected from each participant. Genomic DNA was extracted by the Qiagen DNA purification kit according to the manufacturer’s protocol. A nanodrop was utilized to determine the DNA concentration of each sample based on the optical density (OD) at 260 nm, and the DNA quality of each sample was assessed by the OD 260/280 ratio.
Genotyping was performed with the Taqman OpenArray Genotyping Platform (Applied Biosystems, Inc.). For each sample, a mixture of 2 μl of DNA (ca. 100 ng) and 2 μl of 2× Taqman OpenArray Genotyping Master Mix was added to a 384-well plate. After the plate was sealed and centrifuged briefly, each plate was loaded into the QuantStudioTM 12 K Flex for PCR amplification. The amplified results were reviewed by Taqman Genotyper Software (v. 1.3.1) for SNP calling.
The SNPs examined in this study were three in CHRNB2, 14 in CHRNA3, 10 in CHRNB4, 16 in CHRNA5, 23 in CHNRA7, and 6 in CHRNA4. They were all selected with the SNP Browser software from Applied Biosystems by searching the dbSNP database and published papers. However, because of the calling rate of <95%, SNPs rs6495309 in CHRNA3, rs11637890 in CHRNB4, rs503464 and rs601079 in CHRNA5, and rs6494211 in CHRNA7 were excluded. After those quality control steps, a total of 67 SNPs remained for the association analysis. All of these SNPs have a minor allele frequency (MAF) of >1% and a p value of >1 × 10−4 in the Hardy–Weinberg equilibrium test.
Population structure analysis
We used the Structure program (v. 2.3.4)41 to assess population stratification for our samples based on the genotyping data for a panel of 30 ancestry informative markers.42 Simulation parameters were set to 100,000 burn-ins and 100,000 iterations. To increase the accuracy when inferring admixture and taking account of the samples being recruited from two sites, we set K to 2. Our population structure analysis revealed no evidence of population admixture among the samples from the two recruitment sites, so we performed our association analysis on both sets of samples together with the goal of increasing our statistical power (Supplementary Figure 1).
DNA methylation
DNA methylation data used here were obtained from an ongoing whole-genome bisulfite sequencing project in this laboratory which consisted of 72 subjects selected from the same sample set as used for the abovementioned association analyses on the basis of their age, gender, and smoking status (36 non-smokers; 36 smokers). DNA methylation was identified using the Illumina HiSeq X Ten platform with an average of about 700 million (±75 million) 150-bp paired-end reads per sample. Clean reads were mapped to the hg19 reference genome using Bismark.43 We first combined two strands of information of CpG sites and then excluded those CpGs with <5 reads or that overlapped with common variants in the Chinese Han genome (MAF > 0.05). The MAF of each variant was determined by an unpublished Whole Genome Sequencing dataset of a Chinese Han sample (N = 1329) in our laboratory.
Individual SNP-based and haplotype-based association analysis
Individual SNP-based association analysis was performed for both smoking status and FTND score using PLINK (v. 1.07)44 under the logistic regression model. Adjusted covariates included age, site (Taiyuan or Jincheng), number of smoking family members, and income. In the haplotype-based association analysis, both pair-wise linkage disequilibrium (LD) and haplotypes were evaluated by Haploview (v. 4.2),45,46 and the analysis of those major haplotypes (with a frequency >5%) with each phenotype was performed by HaploStats (v. 1.7.7) in R and adjusted for same covariates under the additive model.47
Interaction analysis of variants in CHRNA3, CHRNA4, CHRNA5, CHRNA7, CHRNB2, and CHRNB4
To estimate the epistatic contribution of variants in six nAChR subunit genes to ND, we performed SNP-by-SNP interaction analysis using the GMDR-GPU program developed by our group,48,49 performing an exhaustive search of all combinations containing 2–5 SNPs each. The best interaction model was determined according to the following three parameters: (1) the cross-validation consistency (CVC) statistics for the selected variant combination; (2) the predictive accuracy, determined by 107 permutation tests for the selected SNP combinations; and (3) the significant p value.
On the basis of chromosomal location and known functional nAChR composition, we separated the genotype file into two: (1) all variants in CHRNA3, CHRNA5, CHRNB4, and CHRNA7, all of which are located on chromosome 15; and (2) all variants in CHRNA4, CHRNA5, and CHRNB2 because they can form a functional (α4β2)2α5 receptor in humans.
Determination of cis-mQTL
Our cis-mQTL analysis was restricted to 250 kb upstream and downstream of each SNP. The intervals for adjacent SNPs were combined if they overlapped. Taken together, a total of 8,915 CpG sites were revealed within the intervals of the target genes. Prior to analysis, we removed the low-quality DNA methylation sites with a calling rate of <80%, which left 5580 highly qualified methylated CpGs for cis-mQTL analysis.
We choose three significant SNPs for our cis-mQTL association analysis according to individual SNP-based analysis, namely rs1948, rs3743075, and rs7178270. They were all intronic polymorphisms associated with the extent of methylation. We used the Matrix eQTL (v. 2.1.1) R package50 to test association of the three SNPs, with linear regression under an additive model and adjusted for age, smoking status, and whether the subject was a coal miner. Bonferroni correction was used to define significant associations (i.e., p = 0.05/14,161 = 3.5 × 10−6), where 14,161 is the total number of associations tested for the abovementioned three significant SNPs (i.e., 4623 for rs3743075, 4743 for r1948, and 4795 for rs7178270).
Results
Individual SNP-based association analysis
As shown in Table 2, for smoking status, we found that eight variants in CHRNA5 and one variant in CHRNA4 showed significant associations (p = 8.0 × 10−3 to 4.4 × 10−2). However, they were no longer significant after Bonferroni correction for multiple testing.
For the FTND phenotype, we found that many variants showed significant associations prior to correction for multiple testing. However, only rs1948 and rs7178270 in CHRNB4 and rs3743075 in CHRNA3 remained significant after Bonferroni correction, with p values of 6.6 × 10−5, 2.0 × 10−4, and 7.0 × 10−4, the odds ratio (OR) under the additive model being 0.8, 0.7, and 0.8, respectively (Supplementary Table 1).
Haplotype-based association analysis
According to the haplotype block definition of Gabriel et al.46 there were seven blocks in the CHRNA5/A3/B4 cluster, four in CHRNA7, and one in CHRNA4 and CHRNB2 (D′ > 0.90) (see Supplementary Figure 2 and Supplementary Figure 3).
As shown in Table 3, for the ND measured by smoking status, we found one haplotype, G-C-C, formed by rs1948, rs7178270, and rs17487223, to be marginally associated with smoking status (Hap-Score 2.13; p = 0.0331). For the FTND measure, we detected five major haplotypes in the CHRNA5/A3/B4 cluster that were significantly associated with ND. Of them, two major haplotypes, C-A and T-G, formed by rs3743075 and rs3743074, showed significant associations with the FTND score (Hap-Score 3.51; p = 4.0 × 10−4; Hap-Score −3.37; p = 8.0 × 10−4). Another haplotype, C-T, formed by rs12914385–rs2869546, also showed significance (Hap-Score −3.02; p = 2.6 × 10−3). Further, we found two other haplotypes, A-G-C and G-C-C, formed by SNPs rs1948, rs7178270, and rs17487223 with a frequency of 0.40 and 0.48, to be significantly associated with FTND (Hap-Score −3.57; p = 4.0 × 10−4; Hap-Score 3.79; p = 2.0 × 10−4, respectively). Importantly, the haplotype G-C-C, formed by rs1948, rs7178270, and rs17487223 in CHRNA5/A3/B4, was significantly associated with both smoking status and FTND. In addition, we found a haplotype in CHRNA4 to be marginally associated with FTND (Supplementary Table 2).
SNP-by-SNP interaction analysis
Combinations of different nAChR subunits can form various functional nicotinic receptors, which play various physiological roles in both the peripheral and the central nervous systems. To determine whether there exists any epistatic effect among these nAChR subunits, we performed interaction analysis using our own GMDR-GPU program, which revealed two significant interaction models for FTND. The first model consists of SNPs rs16969968 in CHRNA5 and rs7178270 in CHRNB4, with a CVC 7 of 10, prediction accuracy of 55.4%, and empirical p value of 8 × 10−5 based on 107 permutations. The second significant model consists of SNPs rs904951 and rs7178176 in CHRNA7, with a CVC 10 of 10, prediction accuracy of 56.0%, and empirical p value of 5.8 × 10−6 from 107 permutations (Supplementary Table S3). Although we also did interaction analysis on SNPs from CHRNA3, CHRNB4, and CHRNA5 with CHRNA7, we did not find any significant interaction for variants in CHRNA7 with those from the other nAChR subunit genes.
For smoking status, we detected a best interaction model, which consists of rs16969968 in CHRNA5, rs4845378 and rs3811450 in CHRNB2, and rs3787137 and rs1044393 in CHRNA4. This model has a CVC 7 of 10, prediction accuracy of 52.3%, and empirical p value of 0.012 (Supplementary Table S3).
Relations between genotype and methylation status
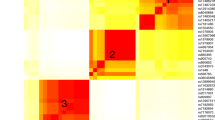
All association analysis results of top SNPs with cis-mQTLs are shown in Supplementary Table 4 and Supplementary Figure 4 (p < 5.0 × 10−4). Together, we detected four significant associations for SNP–CpG pairs (Fig. 1). These pairs were formed by three SNPs significant for ND (rs3743075 in CHRNA3 and rs1948 and rs7178270 in CHRNB4) and two distinct CpG sites (CpG_2975, CpG_3007). Significant cis-mQTL analysis results and the corresponding p values, with a range of 5.2 × 10−15 to 1.9 × 10−06 (Fig. 2).
Distribution of the values at the methylation sites is presented for individuals carrying zero, one, and two minor alleles. Only the most significant cis-mQTLs are shown for each variant. The cis-eQTL analysis associated three variants with CHRNA3 expression in human NAc basal ganglia. The cis-eQTL figures were downloaded from GTEX PORTAL (https://gtexportal.org/home/)
Given that the two highly methylated CpG sites are both located in CHRNA3, our association analyses of the three SNPs with RNA expression were conducted only for CHRNA3. According to the cis-eQTL association data from GTEX PORTAL (https://gtexportal.org/home/), we found that the three SNPs linked to ND (i.e., SNPs rs1948, rs7178270, and rs3743075) correlated significantly with two methylation sites that showed allele-specific mRNA expression of CHRNA3 and CHRNA5 in several ND-related brain regions of human post-mortem tissue (Supplementary Table 5), with p values ranging from 3.2 × 10−6 to 4.0 × 10−15.
In addition, we performed cis-mQTL analysis by adjusting with FTND score and age. Because the FTND score is a ND measure for smokers, we only included smokers in this analysis. We obtained consistent results (Supplementary Table 7). The significant CpG–SNP pairs were still formed by CpG_2975 with rs3743075, rs1948 and rs7178270, respectively (p = 2.2 × 10−6; p = 1.8 × 10−5; p = 2.4 × 10−5). These results further support that these three variants are risk variants that influence the extent of methylation in smokers.
Discussion
To our knowledge, this is the first study exploring the underlying mechanisms of ND using multiple approaches of genetic associations, interaction, and cis-mQTLs in a Chinese Han population. Our results revealed that three SNPs and five major haplotypes in CHRNA5/A3/B4 were significantly associated with the FTND score. We also demonstrated that the significant ND-associated variants (rs1948, rs7178270, and rs3743075) are novel cis-meQTLs influencing both the extent of methylation and mRNA expression of CHRNA3. In addition, through SNP-by-SNP interaction study, we found several SNPs in CHRNA5/B4, CHRNA5/A4/B2, and CHRNA7 that interactively confer susceptibility to ND in our Han sample.
Two haplotypes formed by rs3743074 and rs3743075 were significantly associated with the FTND score. Of them, rs3743075 was significantly and rs3743074 was marginal associated with the score. Further, we showed that the minor allele of rs3743075 in CHRNA3 was significantly associated with the extent of methylation of CpG_2975 and CpG_3007 (p = 5.2 × 10−15 and p = 1.9 × 10−6, respectively) and marginally associated with CpG_3006 (p = 4.7 × 10−4). In concert with these findings, Hancock et al.36 reported that rs3743075 was negatively correlated with a methylation locus of cg22670733 in adult brain (p = 7.0 × 10−6). According to the annotation by GTEX PORTAL (https://gtexportal.org/home/), rs3743075 has strong evidence of affecting expression of CHRNA3 in the brain nucleus accumnems (NAc) (p = 4.9 × 10−11)51 and of CHRNA5 in the NAc, anterior cingulate cortex (BA 24), frontal cortex (BA 9), hippocampus and whole blood (p = 2.3 × 10−10; p = 4.9 × 10−8; p = 3.0 × 10−10; p = 1.6 × 10−5)52 as a cis-eQTL in humans. By using the Web-based tool SWISS-PROT (http://www.uniprot.org/), we found that rs3743075 represents a part of the conserved transcription factor-binding site for interferon regulatory factor 7 (IRF-7) in humans. Previously, we showed that the expression of IRF-7 mRNA was significantly suppressed by nicotine treatment in mouse RAW264.7 macrophages.53 Thus, it is highly likely that rs3743075 is a functional variant regulating the expression of CHRNA3 by altering the methylation contribution. In light of previous evidence demonstrating that smoking-associated abnormal methylation loci might convey a risk for lung cancer,54,55 this may indicate that rs3743075 is a biomarker for lung cancer.
SNPs rs1948 and rs7178270 in CHRNB4 showed the strongest association with the FTND score in our Chinese Han sample. In addition, we found that two haplotypes, A-G-C and G-C-C (formed by rs1948, rs7178270, and rs17487223), showed significant associations with the FTND score. Moreover, we demonstrated that the minor allele of rs1948 and rs7178270 significantly reduced the methylation of CpG_2975 (p = 4.4 × 10−13 and p = 1.2 × 10−12, respectively). Interestingly, these two SNPs increased the expression of CHRNA3 in the NAc.51 These results indicate that SNPs rs1948 and rs7178270 confer susceptibility to ND by suppressing the methylation that leads to increased expression of CHRNA3, which is consistent with previous documentation performed with Dutch persons born in The Netherlands.56 For CHRNA5, there were eight SNPs showing marginal associations with both smoking status and FTND (p < 0.05). Among them, two SNPs, rs16969968 and rs667282 in CHRNA5, have been widely associated with ND-related phenotypes and lung cancer in subjects of multiancestry, such as Europeans, Africans, and Asian.12,13,14,17,19,22,24,57,58 In addition, there exists one haplotype consisting of nine variants in the CHRNA5/A3/B4 cluster that is associated marginally with the FTND score and two haplotypes, formed by another nine SNPs in the CHRNA5/A3/B4 cluster, that are associated marginally with both the FTND and smoking status.
We performed gene–gene interaction analysis of all selected SNPs within CHRNA5/A3/B4 and CHRNA7. We detected the best interaction model between rs7178270 in CHRNB4 and rs16969968 in CHRNA5 (p = 8.0 × 10−5) with FTND, providing genetics-based evidence supporting the view that CHRNB4 interacts with CHRNA5 in contributing to ND. Previous molecular studies found that in mice, medial habenula overexpression of Chrnb4 strongly increases the aversive effect of nicotine, which can be reversed by lentiviral-mediated expression of Chrna5 D398N.59 Moreover, we observed that the interaction model of (α4β2)2α5 grouped by five SNPs (rs4845378, rs3811450, rs3787137, rs1044393, and rs16969968) was significantly associated with smoking status (p = 0.012). This interactive model was compatible with biological evidence that (α4β2)2α5 carrying the risk allele of rs16969968 has a twofold lower maximum response with a nicotinic agonist compared with the α5 carrying a protective allele.60 This is the first evidence of an epistatic effect of this gene cluster discovered via genetic methods in the Chinese Han population.
To date, there are far fewer loci in CHRNA7 reported to be significantly associated with smoking-related phenotypes, in contrast to the numerous reports that the gene is essential in ND in pharmacological studies. In the present work, we did not detect any significant association by either individual SNP-based or haplotype-based association analysis. One possible explanation for the inconsistent observations is that the variants in CHRNA7 contribute to ND risk through SNP-by-SNP interactions. Concordant with this assumption, we found a significant interaction model formed by rs904951 and rs7178176 associated with FTND. rs7178176 was previously reported in association with an increasing probability of dizziness at first smoke inhalation by adolescents in a Canada sample with a mixed ethnical origins.61 For the first time, we provide evidence that an epistatic effect of CHRNA7 is implicated in ND, which is in accord with the biological fact that (α7)5 forms a homomeric pentamer in humans.
There are a few limitations of this study. First, the number of subjects in the methylation cohort might be too small to detect weak signals of risk polymorphisms and mapping trans-mQTLs. Considering that trans-meQTL is less common and contributes only minor effects to phenotypes,62 we concentrated our analysis on identification of cis-meQTLs. In spite of the small samples, because we adopted stringent criteria to select high-quality CpG sites, we believe that our cis-meQTL analysis results are trustworthy and deserve to be replicated by others if possible. Second, we chose 67 variants within the six genes to perform association analysis for our sample, and the majority of them are common SNPs. Thus, we could not assess the effects of rare variants, which have been thought to exert more influence on traits of interest. Further next-generation sequencing-based studies are warranted to identify more rare variants within these regions for ND in the Chinese Han population.
To sum up, this is the first study to demonstrate the significant effects of CHRNA3–CHRNB4 variants on ND in the Chinese Han population. Having succeeded in performing integrative data analysis for genetic polymorphisms and DNA methylation, we relate SNP-based association, cis-mQTL analysis, and mRNA expression using public data to explore the biological mechanisms of ND. Further, we provide novel evidence of a significant genetic interactive model for CHRNA7 in affecting ND, which extends our knowledge of the potential biological mechanism for this gene’s actions affecting ND. A complete understanding of the genetic variants will help us find pharmacologic targets to account for the addictive properties of nicotine in Chinese smokers.
References
Centers for Disease Control and Prevention. Current cigarette smoking among adults - United States, 2011. MMWR Morb. Mortal. Wkly. Rep. 61, 889–894 (2012).
Warren, C. W., Jones, N. R., Eriksen, M. P. & Asma, S., Global Tobacco Surveillance System collaborative group. Patterns of global tobacco use in young people and implications for future chronic disease burden in adults. Lancet 367, 749–753 (2006).
WHO Media centre. Tobacco Fact sheet N 339. http://www.who.int/mediacentre/factsheets/fs339/en/ (2016).
Benowitz, N. L. Clinical pharmacology of nicotine: implications for understanding, preventing, and treating tobacco addiction. Clin. Pharmacol. Ther. 83, 531–541 (2008).
Center for Disease Control. Major finding of 2015 Chinese adults tobacco survery (Chinese Center for Disease Control and Prevention, Beijing, 2015).
Liu, B. Q. et al. Emerging tobacco hazards in China: 1. Retrospective proportional mortality study of one million deaths. BMJ 317, 1411–1422 (1998).
Li, M. D., Cheng, R., Ma, J. Z. & Swan, G. E. A meta-analysis of estimated genetic and environmental effects on smoking behavior in male and female adult twins. Addiction 98, 23–31 (2003).
Le Novere, N., Corringer, P. J. & Changeux, J. P. The diversity of subunit composition in nAChRs: evolutionary origins, physiologic and pharmacologic consequences. J. Neurobiol. 53, 447–456 (2002).
Elgoyhen, A. B. et al. alpha10: a determinant of nicotinic cholinergic receptor function in mammalian vestibular and cochlear mechanosensory hair cells. Proc. Natl Acad Sci USA 98, 3501–3506 (2001).
Elgoyhen, A. B., Johnson, D. S., Boulter, J., Vetter, D. E. & Heinemann, S. Alpha 9: an acetylcholine receptor with novel pharmacological properties expressed in rat cochlear hair cells. Cell 79, 705–715 (1994).
Kuryatov, A., Berrettini, W. & Lindstrom, J. Acetylcholine receptor (AChR) alpha5 subunit variant associated with risk for nicotine dependence and lung cancer reduces (alpha4beta2)(2)alpha5 AChR function. Mol. Pharmacol. 79, 119–125 (2011).
Yang, J. & Li, M. D. Converging findings from linkage and association analyses on susceptibility genes for smoking and other addictions. Mol. Psychiatry 21, 992–1008 (2016).
Saccone, N. L. et al. The CHRNA5-CHRNA3-CHRNB4 nicotinic receptor subunit gene cluster affects risk for nicotine dependence in African-Americans and in European-Americans. Cancer Res. 69, 6848–6856 (2009).
Tobacco, Genetics C. Genome-wide meta-analyses identify multiple loci associated with smoking behavior. Nat. Genet. 42, 441–447 (2010).
Li, M. D. et al. Association and interaction analysis of variants in CHRNA5/CHRNA3/CHRNB4 gene cluster with nicotine dependence in African and European Americans. Am. J. Med. Genet. B Neuropsychiatr. Genet. 153B, 745–756 (2010).
Wen, L., Jiang, K., Yuan, W., Cui, W. & Li, M. D. Contribution of variants in CHRNA5/A3/B4 gene cluster on chromosome 15 to tobacco smoking: from genetic association to mechanism. Mol. Neurobiol. 53, 472–484 (2016).
Berrettini, W. et al. Alpha-5/alpha-3 nicotinic receptor subunit alleles increase risk for heavy smoking. Mol. Psychiatry 13, 368–373 (2008).
Thorgeirsson, T. E. et al. A variant associated with nicotine dependence, lung cancer and peripheral arterial disease. Nature 452, 638–642 (2008).
Saccone, N. L. et al. Multiple distinct risk loci for nicotine dependence identified by dense coverage of the complete family of nicotinic receptor subunit (CHRN) genes. Am. J. Med. Genet. B Neuropsychiatr. Genet. 150B, 453–466 (2009).
Saccone, S. F. et al. Cholinergic nicotinic receptor genes implicated in a nicotine dependence association study targeting 348 candidate genes with 3713 SNPs. Hum. Mol. Genet. 16, 36–49 (2007).
Lassi, G. et al. The CHRNA5-A3-B4 gene cluster and smoking: from discovery to therapeutics. Trends Neurosci. 39, 851–861 (2016).
Li, M. D. et al. Associations of variants in CHRNA5/A3/B4 gene cluster with smoking behaviors in a Korean population. PLoS ONE 5, e12183 (2010).
Liu, J. Z. et al. Meta-analysis and imputation refines the association of 15q25 with smoking quantity. Nat. Genet. 42, 436–440 (2010).
David, S. P. et al. Genome-wide meta-analyses of smoking behaviors in African Americans. Transl. Psychiatry 2, e119 (2012).
Thorgeirsson, T. E. et al. Sequence variants at CHRNB3-CHRNA6 and CYP2A6 affect smoking behavior. Nat. Genet. 42, 448–453 (2010).
Ghasemzadeh-Hasankolaei, M., Batavani, R., Eslaminejad, M. B. & Sayahpour, F. Transplantation of autologous bone marrow mesenchymal stem cells into the testes of infertile male rats and new germ cell formation. Int. J. Stem Cells 9, 250–263 (2016).
Levin, E. D. et al. Nicotinic alpha7- or beta2-containing receptor knockout: effects on radial-arm maze learning and long-term nicotine consumption in mice. Behav. Brain Res. 196, 207–213 (2009).
Brunzell, D. H. & McIntosh, J. M. Alpha7 nicotinic acetylcholine receptors modulate motivation to self-administer nicotine: implications for smoking and schizophrenia. Neuropsychopharmacology 37, 1134–1143 (2012).
Brunzell, D. H., Boschen, K. E., Hendrick, E. S., Beardsley, P. M. & McIntosh, J. M. Alpha-conotoxin MII-sensitive nicotinic acetylcholine receptors in the nucleus accumbens shell regulate progressive ratio responding maintained by nicotine. Neuropsychopharmacology 35, 665–673 (2010).
Melis, M. et al. PPARalpha regulates cholinergic-driven activity of midbrain dopamine neurons via a novel mechanism involving alpha7 nicotinic acetylcholine receptors. J. Neurosci. 33, 6203–6211 (2013).
Breitling, L. P., Yang, R., Korn, B., Burwinkel, B. & Brenner, H. Tobacco-smoking-related differential DNA methylation: 27K discovery and replication. Am. J. Hum. Genet. 88, 450–457 (2011).
Furniss, C. S., Marsit, C. J., Houseman, E. A., Eddy, K. & Kelsey, K. T. Line region hypomethylation is associated with lifestyle and differs by human papillomavirus status in head and neck squamous cell carcinomas. Cancer Epidemiol. Biomark. Prev. 17, 966–971 (2008).
Bell, J. T. & Spector, T. D. DNA methylation studies using twins: what are they telling us? Genome Biol. 13, 172 (2012).
Liu, Y. et al. Epigenome-wide association data implicate DNA methylation as an intermediary of genetic risk in rheumatoid arthritis. Nat. Biotechnol. 31, 142–147 (2013).
Jaffe, A. E. et al. Mapping DNA methylation across development, genotype and schizophrenia in the human frontal cortex. Nat. Neurosci. 19, 40–47 (2016).
Hancock, D. B. et al. A multiancestry study identifies novel genetic associations with CHRNA5 methylation in human brain and risk of nicotine dependence. Hum. Mol. Genet. 24, 5940–5954 (2015).
Roadmap Epigenomics, C. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Fagerstrom, K. O. Measuring degree of physical dependence to tobacco smoking with reference to individualization of treatment. Addict. Behav. 3, 235–241 (1978).
Heatherton, T. F., Kozlowski, L. T., Frecker, R. C. & Fagerstrom, K. O. The fagerstrom test for nicotine dependence: a revision of the Fagerstrom Tolerance Questionnaire. Br. J. Addict. 86, 1119–1127 (1991).
Wei, J. et al. Association study of 45 candidate genes in nicotine dependence in Han Chinese. Addict. Behav. 37, 622–626 (2012).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Yang, B. Z., Zhao, H., Kranzler, H. R. & Gelernter, J. Practical population group assignment with selected informative markers: characteristics and properties of Bayesian clustering via STRUCTURE. Genet. Epidemiol. 28, 302–312 (2005).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Gabriel, S. B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
Schaid, D. J., Rowland, C. M., Tines, D. E., Jacobson, R. M. & Poland, G. A. Score tests for association between traits and haplotypes when linkage phase is ambiguous. Am. J. Hum. Genet. 70, 425–434 (2002).
Zhu, Z. et al. Development of GMDR-GPU for gene-gene interaction analysis and its application to WTCCC GWAS data for type 2 diabetes. PLoS ONE 8, e61943 (2013).
Lou, X. Y. et al. A generalized combinatorial approach for detecting gene-by-gene and gene-by-environment interactions with application to nicotine dependence. Am. J. Hum. Genet. 80, 1125–1137 (2007).
Shabalin, A. A. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Consortium, G. T. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Westra, H. J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Cui, W. Y. et al. Identification and characterization of poly(I:C)-induced molecular responses attenuated by nicotine in mouse macrophages. Mol. Pharmacol. 83, 61–72 (2013).
Alexandrov, L. B. et al. Mutational signatures associated with tobacco smoking in human cancer. Science 354, 618–622 (2016).
Ma, Y. & Li, M. D. Establishment of a strong link between smoking and cancer pathogenesis through DNA methylation analysis. Sci. Rep. 7, 1811 (2017).
van Eijk, K. R. et al. Genetic analysis of DNA methylation and gene expression levels in whole blood of healthy human subjects. BMC Genomics 13, 636 (2012).
Wang, Y., Peng, X., Zhu, L., Hu, L. & Song, Y. Genetic variants of CHRNA5-A3 and CHRNB3-A6 predict survival of patients with advanced non-small cell lung cancer. Oncotarget 7, 26436–26443 (2016).
Wu, C. et al. Genetic variants on chromosome 15q25 associated with lung cancer risk in Chinese populations. Cancer Res. 69, 5065–5072 (2009).
Frahm, S. et al. Aversion to nicotine is regulated by the balanced activity of beta4 and alpha5 nicotinic receptor subunits in the medial habenula. Neuron 70, 522–535 (2011).
Bierut, L. J. et al. Variants in nicotinic receptors and risk for nicotine dependence. Am. J. Psychiatry 165, 1163–1171 (2008).
Pedneault, M. et al. The association between CHRN genetic variants and dizziness at first inhalation of cigarette smoke. Addict. Behav. 39, 316–320 (2014).
McClay, J. L. et al. High density methylation QTL analysis in human blood via next-generation sequencing of the methylated genomic DNA fraction. Genome Biol. 16, 291 (2015).
Acknowledgements
This study was supported in part by the China Precision Medicine Initiative (2016YFC0906300), Research Center for Air Pollution and Health of Zhejiang University, the State Key Laboratory for Diagnosis and Treatment of Infectious Diseases of the First Affiliated Hospital of Zhejiang University, and Shanxi Key Laboratory of Ecological Animal Science and Environmental Veterinary Medicine of Shanxi Agricultural University. We thank Dr. David L. Bronson for excellent editing of this manuscript.
Author information
Authors and Affiliations
Contributions
H.H., L.W., K.J., Y.M., K.S., Z.Y., W.C., W.Y., and X.J. participated clinical data collection. Q.L., H.H,. M.W., R.F., J.C., and X.J. performed the laboratory experiments. Q.L., L.W., and Y.Y. participated data analysis. Q.L., T.J.P., and J.W. participated in writing of the paper. M.D.L. conceived the study and was involved in every step of the research. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, Q., Han, H., Wang, M. et al. Association and cis-mQTL analysis of variants in CHRNA3-A5, CHRNA7, CHRNB2, and CHRNB4 in relation to nicotine dependence in a Chinese Han population. Transl Psychiatry 8, 83 (2018). https://doi.org/10.1038/s41398-018-0130-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-018-0130-x
- Springer Nature Limited
This article is cited by
-
Integrative analysis of genetics, epigenetics and RNA expression data reveal three susceptibility loci for smoking behavior in Chinese Han population
Molecular Psychiatry (2024)
-
Genetic variations in the bitter taste receptor gene TAS2R38 are related to cigarette smoking behavior in Han Chinese smokers
Genes & Genomics (2022)
-
Genome-wide methylation and expression analyses reveal the epigenetic landscape of immune-related diseases for tobacco smoking
Clinical Epigenetics (2021)
-
Methylation quantitative trait locus rs5326 is associated with susceptibility and effective dosage of methadone maintenance treatment for heroin use disorder
Psychopharmacology (2021)
-
Association and cis-mQTL analysis of variants in serotonergic genes associated with nicotine dependence in Chinese Han smokers
Translational Psychiatry (2018)