Abstract
Proper subcellular localization is crucial for the functioning of biomacromolecules, including proteins and RNAs. Nuclear transport is a fundamental cellular process that regulates the localization of many macromolecules within the nuclear or cytoplasmic compartments. In humans, approximately 60 proteins are involved in nuclear transport, including nucleoporins that form membrane-embedded nuclear pore complexes, karyopherins that transport cargoes through these complexes, and Ran system proteins that ensure directed and rapid transport. Many of these nuclear transport proteins play additional and essential roles in mitosis, biomolecular condensation, and gene transcription. Dysregulation of nuclear transport is linked to major human diseases such as cancer, neurodegenerative diseases, and viral infections. Selinexor (KPT-330), an inhibitor targeting the nuclear export factor XPO1 (also known as CRM1), was approved in 2019 to treat two types of blood cancers, and dozens of clinical trials of are ongoing. This review summarizes approximately three decades of research data in this field but focuses on the structure and function of individual nuclear transport proteins from recent studies, providing a cutting-edge and holistic view on the role of nuclear transport proteins in health and disease. In-depth knowledge of this rapidly evolving field has the potential to bring new insights into fundamental biology, pathogenic mechanisms, and therapeutic approaches.
Similar content being viewed by others
Introduction
Eukaryotic cells store genetic material in the nucleus and separate it from other cellular components using a double-layered membrane called the nuclear envelope (NE). This compartmentalization allows for complex and specialized cellular activities while simultaneously posing challenges for the exchange of materials between the nucleus and the cytoplasm. The vast majority of material exchange occurs through nuclear pore complexes (NPCs), which form channels in the NE. The transport of molecules into and out of the nucleus determines the subcellular localization of many macromolecules, e.g., transcription factor, and is fundamental in the regulation of gene expression, cell division, and other critical cellular functions.1
To facilitate nuclear transport (or nucleocytoplasmic transport), human cells dedicated approximately 60 different proteins to constitute the nuclear transport system (NTS).2 Each of these nuclear transport proteins (NTPs) has a defined function. One component of the NTS is the NPC, which is formed by nucleoporin proteins and presents a selective barrier to free diffusion of macromolecules into and out of the nucleus.3 The karyopherin family proteins, such as importins, exportins, and bidirectional transporters (biportins), act as molecular shuttles to transport macromolecules through NPCs.4 The small GTPase protein Ran and accessory factors regulate the transport direction and accelerate transport speed.5 In addition, proteins involved in nuclear transport have been demonstrated to have nontransport functions, including roles in mitosis, regulation of transcription, and regulation of biomolecular condensates.6,7 It is often possible to distinguish the contribution of canonical and noncanonical functions of NTPs in a certain cellular process. Due to these important cellular functions, dysregulation of the NTS is implicated in a range of human diseases, including cancer, neurodegenerative disorders, and viral infections.8,9
While different NTPs are often tightly linked in many cellular processes, most previous reviews have not included all three NTP classes: karyopherins, nucleoporins, and Ran system proteins. A holistic view of the NTS could facilitate the understanding of relevant phenomena and guide the development of therapies for diseases. In this review, we will explore the structure, function, and disease relevance of individual NTPs, with a focus on their interaction mechanism and networks, underlying principles, and potential therapeutic targets. We will draw on key foundations dating back decades as well as recent literature to summarize and discuss the vast body of knowledge acquired, and hopefully bring new perspectives to future research.
Retrospective summary of research milestones
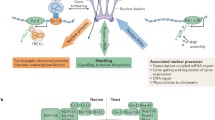
Due to their large size, cylindrical nuclear pore complex penetrating the nuclear envelope were discovered under electron microscopy as early as 1959 (Fig. 1).10,11. The first nuclear localization signal, which localizes yeast ribosomal protein L3 to the nucleus, was reported in 1985.12 Although the existence of nuclear import receptors was suspected at the time, first import receptor, p97 (now known as importin β1), was identified about a decade later.13,14 Shortly after, the first nuclear export signal and the first nuclear export receptor CRM1 were also identified.15,16,17 Ran-mediated regulation of nuclear cytoplasmic transport was discovered slightly earlier, but its role in nuclear transport was not well-understood until 1999.18 In the same year, the report of Ran-Importin β1 crystal structures marked that the field of nuclear transport entered the structural era.19,20 Using a collection of biophysical and proteomic techniques, the first molecular architecture of yeast NPC was built in 2007.21 Thereafter, with the development of cryo-EM and cryo-ET (electron tomography), the resolution of NPC structures has gradually increased to the current subatomic level.22,23 In the 1990s and 2000s, it was demonstrated that these NTS proteins also regulate mitosis, biomolecular condensates, and gene transcription, and are therefore implicated in various human diseases, including cancers, although many of the underlying mechanisms have not been revealed until recently.24,25,26,27 A drug targeting CRM1 was approved to treat two types of relapsed or refractory hematological cancers in 2019.28
Components of the nuclear transport system
The 60 NTPs can be classified into three groups: the nucleoporins that form the nuclear membrane-embedded NPC, the karyopherins that bind and ferry cargoes through NPCs, and the Ran system proteins that determine transport directionality and efficiency.2 In this section, we will discuss the structure and function of each NTP class.
Nucleoporins that form the nuclear pore complex
The NE consists of two lipid bilayer membranes—the inner and outer nuclear membranes - with NPCs embedded in NE pores where the inner and outer bilayers are curved and fused. A typical mammalian cell has approximately 2000–5000 NPCs.29 Each NPC can be visualized as a hollow cylinder with an outer diameter of ~1200 Å, a height of ~800 Å, and a total weight of ~120 MDa.30,31 The NPC can be divided into three parts: a central core that binds to the membrane and forms a diffusion barrier, eight thin filaments that bind to the central core and extend to the cytoplasm, and an additional eight thin filaments that form a basket-like structure on the nuclear side (Fig. 2a). All three parts of the NPC exhibit eightfold rotational symmetry along the channel axis, with all nucleoporins present as a multiple of eight in each NPC. The central core has an additional twofold symmetry between the cytoplasmic and nuclear halves.30 Therefore, each symmetric core nucleoporin (or symmetric nucleoporin) is present in at least a multiple of 16 in each NPC.32
Carton representation of the nuclear pore complex. a The nuclear pore complex can be divided into three parts: the central core, the cytoplasmic filaments, and the nuclear basket. The central core can be further divided into four rings surrounding the central channel: the inner ring, the cytoplasmic ring, the nuclear ring, and the luminal ring. b Inner ring viewed in the direction of transport. The inner ring consists of eight loosely associated subunits surrounding the central channel. The central channel is filled with disordered FG repeat polypeptides that inhibit free diffusion across the nuclear envelope. c Architecture of the outer ring. The two outer rings (the cytoplasmic and nuclear rings) are highly similar, and only one outer ring is drawn for clarity. Each outer ring contains 16 copies of the Y complex, arranged in two concentric rings and stabilized by linker nucleoporins. The Y complex consists of 10 nucleoporins, which can be divided into three regions: the stem, the short arm, and the long arm. d Architecture of the luminal ring. Thirty-two copies of Pom210 connected end-to-end surround the NPC and interact with the inner ring on the other side of the nuclear envelope. The parallelogram architectural features of Pom210 allows deformation that contract (left panel) or dilate (right panel) the central channel
Each NPC is constructed from approximately 1000 protein subunits, made up of multiple copies of approximately 34 unique nucleoporins encoded by the human genome. Approximately ten nucleoporins contain long stretches of FG repeats that are disordered and rich in FG dipeptides.33 These FG repeats are critical for passive diffusion barrier formation and karyopherin binding. The most prevalent domains are α-helical solenoids and β-propellers, which form the relatively rigid NPC scaffold. Nucleoporins have diverse functions, with some anchoring the NPC in the membrane (transmembrane nucleoporins), some forming the skeleton or scaffold of NPC (scaffold nucleoporins), some linking different scaffolds together (linker nucleoporins), some forming a diffusion barrier and/or interacting with different transport factors (FG nucleoporins), and some having mixed domains and functions. In this review, we introduce different nucleoporins according to their location within the NPC (Table 1). However, it should be noted that some nucleoporins are not restricted to a single location, especially those linking different parts of the NPC.
Symmetric core nucleoporins
The symmetric core can be further divided into four concentric rings: an inner ring which lines the central channel and forms the NPC diffusion barrier, two outer rings (nuclear ring and cytoplasmic ring) which dock the cytoplasmic filaments and the nuclear basket, and a luminal ring in the NE lumen surrounding the NPC (Fig. 2a).34 The inner ring and outer rings are connected by eight filaments on each side of the NPC. The filaments limit the movement of the inner ring towards outer rings but permit dilation or constriction in the NE plane. NPCs are conserved across diverse species from yeast to humans, but the degree of conservation for different parts are not the same: the inner ring, out rings, and other regions are in descending order of conservation. The inner ring thus represents the most critical part for NPC functions, especially nuclear transport.
Nucleoporins forming the inner ring
The inner ring is divided into eight subunits by eightfold symmetry, and each subunit is symmetrical on both the nuclear and cytoplasmic sides. When viewed from the direction of transport, each subunit resembles an eighth slice of pizza with the center portion cut away (Fig. 2b).35 The inner ring subunits are porous, plastic, and weakly connected to one another, allowing them to deform and change pore size in response to stimuli.36,37,38 The inner ring subunit can be further divided into three layers based on the distance to the transport axis: a middle layer of nucleoporins that form the central scaffold, an outer layer of coat nucleoporins that associate with the membrane, and an inner layer of nucleoporins that form the diffusion barriers.39,40
The central scaffold of each subunit is composed of two copies of Nup188, two copies of Nup205, and four copies of Nup93.41 These scaffold nucleoporins are mainly α-helical solenoids that intercalate extensively. The relatively rigid scaffold prevents excessive shrinkage of nuclear pores when subjected to compression force from the membrane.42 The coat nucleoporins include Nup155 (6 copies), NDC1 (2 copies), and ALADIN (2 copies). Nup155 contains a β-propeller domain as well as an α-helical solenoid. Four copies of Nup155 use α-helical solenoid domains to form a cushion for the central scaffold and use β-propeller domains to contact the membrane. The interaction between the inner ring and membrane is strengthened by ALADIN and NDC1. ALADIN is a β-propeller fold that interacts with the membrane and the pore domain of NDC1. NDC1 contains an additional transmembrane domain anchoring the inner ring to the NE. The two ALADIN-NDC1 heterodimers also interact with two other copies of Nup155 that contact the two outer rings.
The barrier nucleoporins, also known as the channel nucleoporin heterotrimer (CNT),43 include Nup54, Nup58, and Nup62 (four copies each) and are anchored by the N-terminal SLiMs of Nup93.44 Each of these nucleoporins contains a C-terminal coiled-coil domain bundled alongside the other coiled-coil domains of the heterotrimer, as well as an N-terminal FG repeat domain extending into the central transport channel to form the diffusion barrier. These FG repeats are depleted of charged amino acids and, at high concentrations, can self-assemble into a hydrogel-like condensate, which allows the diffusion and transport of FG-interacting karyopherins but prevents the passage of other macromolecules, biophysically similar to the NPC barrier.45,46 FG repeats in two disordered inner ring nucleoporins, Nup98 and Nup35 (also known as Nup53), can simultaneously bind several α-solenoid nucleoporins, which are structurally related to karyopherins, through interactions resembling those found in FG-karyopherins.47 In this way, these linker nucleoporins thread together all three layers, stabilize the NPC, and play a role in recruiting inner ring nucleoporins during NPC biogenesis.39,48
Nucleoporins forming the outer rings
Outer rings refer to the cytoplasmic outer ring (cytoplasmic ring) and the nuclear outer ring (nuclear ring). These two rings are largely identical, except for copy number differences of select components (ELYS, Nup205, and Nup93).49 Copy number differences for these proteins are also observed between species or even within a single cell, however, the functional difference remains poorly understood.50 Remarkably, the human outer rings contain twice as many Y-shaped structures (knowns as Y complexes or coat nucleoporin complexes, 32 vs. 16) as yeast.51 The outer rings bind and curve the membrane, connect the inner ring through Nup155, and form docking sites to recruit asymmetric nucleoporins (e.g., Nup358).52 Several asymmetric nucleoporin domains are firmly bound to outer rings and are sometimes regarded as a portion of the outer rings. For simplicity, we consider those domains to be part of the cytoplasmic filaments or nuclear baskets and will discuss asymmetric nucleoporins separately in later sections.
In each outer ring, the Y complexes are arranged head-to-tail and form two concentric rings, each containing eight copies of Y complexes (Fig. 2c). The human Y complex is composed of 10 nucleoporin proteins that form a short arm (Nup75, Nup43, and Seh1), a long arm (Nup160, Nup37, and ELYS), and a stem (Sec13, Nup96, Nup107, and Nup133), together resembling the ‘Y’ shape (Fig. 2c).50,53 These nucleoporins contain either α-helical solenoid domains, β-propeller domains, or both, and membrane contact is primarily mediated by the β-propeller domains at the tips of the long arm and the stem. ELYS is not considered as a component of the Y complex by some groups because it is not uniformly present in all Y complexes. In X. laevis, the cytoplasmic ring has eight copies of ELYS, whereas the nuclear ring contains 16 copies.49 The extra copies of ELYS in the nuclear ring are well-exposed, functioning in chromatin binding, decondensation and gene transcription.50,54,55
The cytoplasmic ring can be divided into eight identical subunits, each containing two copies of Y complexes, one proximal and one peripheral (Fig. 2c). Except for the extensive interface between the two Y complexes, two linker nucleoporins, Nup205 and Nup93, connect and stabilize the two Y complexes. In addition, these two nucleoporins mediate inter-subunit interactions in a head-to-tail fashion, strengthening the outer ring scaffold. In Xenopus laevis, the cytoplasmic ring contains two copies of Nup205 and Nup93, while the nuclear ring subunit contains only one copy each of Nup205 and Nup93.49,56 Unlike the inner ring, the outer rings have extensive intersubunit interactions and rigid linkers, thus not allowing intersubunit movements. Stable outer rings are capable of restricting the inner ring movement through the Nup155 filament.38
Nucleoporins forming the luminal ring
The luminal ring (also known as the membrane ring) is within the perinuclear lumen of the NE and equatorially encircles the NPC.57 The luminal ring may sense membrane tension, set the NPC dilation limit, and buffer collisions with adjacent NPCs.34,37,58 The luminal ring appears as eight arches connected end to end and can be conceptualized as 16 parallelograms joined on their short sides (Fig. 2d).58,59 Since the luminal ring is connected to the inner ring via NDC1, the deformation of parallelograms can contract or dilate the associated inner ring. The deformation of parallelograms is likely passive, allowing the NPC to adapt to membrane tension and transport demands. The luminal ring contains Pom121 and Pom210 in vertebrates, both possessing a single-pass transmembrane region.42,57 Pom210, which contains 16 immunoglobulin-like domains, is responsible for the CryoEM density of the luminal ring, since each Pom121 contributes merely ~30 residues to the luminal ring.58 The pore side of Pom121 is largely unstructured and directly binds the β-propeller domains of Nup155 (the inter-ring filament) and Nup160 (Y complex component), thereby anchoring the cytoplasmic ring to the membrane.60,61 Whether and how Pom121 directly bind Pom210 are currently unclear.
The cytoplasmic filament nucleoporins
The cytoplasmic filaments are anchored to the cytoplasmic outer ring and possess long, flexible filamentous extensions into the cytoplasm. The exact architectural details of these extensions are not fully understood due to their conformational heterogeneity. These filaments are composed of less conserved accessory nucleoporins, being cell-type specific and modifiable by cellular processes.62,63 Nevertheless, cytoplasmic filaments play a crucial role in the recruitment of transport factors and the final step of protein and mRNA export.64 Nup214, Nup358, Nup98, and Nup42 are the main contributors to cytoplasmic FG repeats.33
Most of the molecular mass of the cytoplasmic filament is contributed by Nup358, which is large in size (358 kD) and high in copy number (five copies per filament).52 Five Nup358 molecules form a homopentameric complex using the coiled-coil domains and assemble onto the stems of two Y complexes using the N-terminal α-helical solenoid domains.52 Nup358 assembly in turn can stabilize the Y complex rings.65 The remaining domains of the five Nup358 molecules are entangled and flexibly extend into the cytoplasm, forming the observed 50 nm filamentous structures.66 The extended region of Nup358 contains four dispersed RanBP1-like Ran binding domains, a tandem array of eight zinc-finger RanGDP-binding domains, a binding site for the SUMO E2 ligase Ubc9 and RanGAP1, many FG repeats, and a catalytically active cyclophilin domain.67 These domains are involved in RanGTP hydrolysis, RanGDP recycling, and karyopherin docking.68,69,70,71
Alongside Nup358, the cytoplasmic ring is decorated with 16 copies of Nup214 complexes.50 This complex is constructed by eight nucleoporins including Nup214, Nup62, Nup88, Nup98, Nup42, Gle1, RAE1, and the ATP-dependent DEAD-box RNA helicase DDX19, although some of these proteins are not constitutively associated with NPCs.51 An earlier study showed that Nup358 assembly is dependent on the Nup214 complex, but the reverse is not true.72 Nup214, Nup88, and Nup62 uses the coiled-coil domains to form a heterotrimeric complex similar to the one observed in CNT. This complex is anchor to the short arm of the Y complexes and to the membrane, forming a multivalent interaction hub.50,52 Two other subcomplexes, Nup98/RAE1 and Nup42/Gle1/DDX19, are recruited to the vicinity using long linkers. The Nup214 complex thus localize critical factors to remove mRNA from its export factors in the final step of mRNA export.73 Unlike Nup358-mediated protein export termination, this process is independent of Ran and occurs closer to the central channel, but the biological significance is unclear.
Nuclear basket nucleoporins
In humans, the nuclear basket is made up of three nucleoporins: Nup50, Nup153, and Tpr. Tpr is the major structural component of the basket, as it has a large coiled-coil domain which allows for homo-oligomerization.74 Prior studies have demonstrated that Nup153 is responsible for tethering Nup50 to the nuclear pore and post-mitotic recruiting of Tpr to NPC, but not for stabilizing Tpr that is already anchored within the NPC.75,76,77 Unlike the cytoplasmic face, the nuclear face of NPC had minimal electron density beyond the symmetric core nucleoporins, indicating that the basket is anchored by short linear motifs.49,50,78 In agreement with this, depletion of multiple Y complex components, e.g., Nup75 (a Y short arm component) and Nup133 (a Y complex stem component), perturbed nuclear basket formation.59,79 Amphipathic helices from Nup1 (Nup153 orthologue) in yeast can induce membrane curvature and stabilize the nuclear ring.75,80 Nup50 and Nup153 forms a cargo disassembly station for nuclear import due to containing high affinity FG repeats interaction sites for importins.78 Besides nuclear transport, the nuclear basket is critical for cellular processes such as mRNA production and quality control, chromosome organization, and DNA damage repair,75,81,82 but how and why these processes occur at this location are largely unknown.
Karyopherins responsible for ferrying cargo across the nuclear pore complex
Karyopherins are molecules that ferry cargoes across NPCs either into or out of the karyo-compartment (the nucleus). These proteins are conserved from yeast to humans and are important in many cellular processes.83 Typical karyopherins are divided into three groups: importins, which import cargoes into the nucleus; exportins, which export cargoes to the cytoplasm; and biportins, which transport cargoes in either direction.4 These karyopherins rely on the GTPase RanGTP for cargo binding and dissociation.84 They are large in size (~ 1000 residues), forming alpha-helical solenoid structures.20 Generally, the highly acidic concave surface is used for interactions with RanGTP and cargo, and the convex surface presents hydrophobic pockets to interact with the FG repeats of the NPC. There are several small size transport factors that are not known as karyopherins, but similar to karyopherins, they can transport cargo through NPCs.85 On the other hand, a group of transport adaptor molecules (alpha karyopherins) are known as karyopherins, but they cannot independently transport cargo.86 In this section, each of these factors is explained in terms of the cargoes it recognizes, the mode in which cargoes are recognized, the cellular pathways in which it may specialize, and the associated diseases (Table 2).
The importins that import cargoes into the nucleus
Humans possess ten verified importins.87 The function of RanBP6 is unclear, but it is classified as an importin because it has high sequence homology (80% identity) to Importin 5. Importins bind to cargoes in the cytoplasm and release cargoes within the nucleus upon encountering the GTP-bound form of the GTPase Ran (Fig. 3). Generally, cargo binding and RanGTP binding are mutually exclusive, but RanGTP binds with a greater affinity and is thus able to dissociate cargoes.4 A proteomics study demonstrated that each importin recognizes a set of cargoes, although many of these interactions require further verification.88 Importins recognize cargoes in diverse ways, but all rely on positively charged amino acids in cargoes.4 This may explain why many cargoes are able to enter the nucleus using multiple importins.89,90
Model of protein nuclear import and export. Imported cargoes containing nuclear localization signals (NLSs) form complexes with importins in the cytoplasm, enter the nucleus through the NPC, and are dissociated from the importins with the aid of RanGTP. Nuclear export of cargoes starts with the formation of trimeric complexes consisting of exportin, nuclear export signal (NES)-containing cargo, and RanGTP. The trimeric complex transits through the NPC and is dissembled in the cytoplasm upon the hydrolysis of RanGTP. Certain species of RNA utilize protein adaptors to cross NPCs
Importin β1
The most widely studied importin is Importin β1 (Impβ1, also known as Importin β, karyopherin β1, or by its gene name KPNB1). Impβ1 acts as a transporter in classical nuclear import, wherein it recruits the adaptor protein Importin α (Impα, containing 7 isoforms) that directly binds to the cargo protein. The N-terminal Importin Beta Binding Domain (IBB) of Impα is basic and forms an α helix upon binding to Impβ1.91 Likewise, Impβ1 imports m3G-caped U snRNA by binding to the IBB of the adaptor snurportin.92 The use of adaptors enhances the diversity of cargoes recognized by Impβ1 and allows for fine regulation of nuclear import.93 Impβ1 may also directly recognize and import cargo without adaptors, for example, binding globular domains of the cholesterol metabolism transcription factor SREBP-2 to mediate its nuclear import.94 As a major import receptor, Impβ1 imports many cargoes, including the NF-κB subunit p65, autophagy transcription factor TFEB, and programmed cell death ligand 1 PD-L1.95,96,97 Most Impβ1 cargoes play a role in DNA synthesis and repair, as well as gene expression regulation.88 Impβ1 abnormalities are present in several diseases, such as upregulation in cancers and downregulation in neurodegenerative diseases.83,98
Transportin 1 and Transportin 2
Transportin 1 (TNPO1, also known as karyopherin β2) and Transportin 2 (TNPO2, also known as Importin 3) are highly homologous (85% sequence identity) and well-characterized importins that bind the PY (proline-tyrosine) nuclear localization signal (NLS) of cargo proteins.99 A typical PY NLS is disordered and contains two patches, an N-terminal positive/basic patch and a C-terminal [+]-X2–5-P-ϕ motif ([+], positively charged residue; ϕ, any hydrophobic residues including Y).99 In addition to the PY NLS, these importins bind arginine-glycine-glycine (RGG) domains in RNA binding proteins such as hnRNP A1, FUS, and the proline-arginine (PG) poly-dipeptides from C9orf72.100,101,102,103 As such, impairment of TNPO1 and TNPO2 causes those proteins to aggregate and condense in neuron cells, contributing to neurodegenerative diseases.83 Furthermore, TNPO1 imports the Wnt signaling effector β-catenin as well as the tumor suppressor BAP1, and many viruses exploit TNPO1 for nuclear entry and replication.104,105,106,107 Proteome analysis indicated that proteins related to nuclear division and tRNA ligases are preferentially cargoes of TNPO1, while proteins related to DNA repair and HMG proteins are preferentially imported by TNPO2.88 Interestingly, TNPO2 enhances export of a large proportion of mRNAs through the formation of a complex with RanGTP and the mRNA export factor NXF1,108 suggesting that it may be appropriate to classify TNPO2 as a biportin.
Transportin 3
Transportin 3 (TNPO3, also known as Transportin-SR or Transportin-SR2) specifically binds cargoes containing an arginine/serine (RS) domain.109 Counter intuitively, serine phosphorylation of TNPO3 cargoes reduces net positive charges but enhances TNPO3 binding.110 Structural analysis has demonstrated that TNPO3 uses a unique arginine-rich helix for interaction with phosphorylated serine residues, in addition to the common acidic patches that interact with positively charged residues in importin cargoes.111 Examples of TNPO3 cargoes include the alternative splicing factor/splicing factor 2 (ASF/SF2), cold-inducible RNA-binding protein (CIRBP), and polyadenylation specificity factor 6 (CPSF6).101,111 Many TNPO3 cargoes are RS-rich splicing factors.88 The HIV virus utilizes TNPO3 to facilitate its nuclear import and replication, and a natural TNPO3 mutation that causes limb girdle muscular dystrophy also provides strong resistance against HIV-1 infection.112,113 Refer to the Supplemental File for reviews on other importins.
The exportins that export cargoes to the cytoplasm
The human genome encodes five exportins to conduct the export of cellular proteins and RNAs. Unlike importins, exportins display low affinities to either RanGTP or the cargo, typically in the micromolar range. However, exportins can cooperatively bind cargoes and RanGTP and form nanomolar affinity complexes in the nucleus. After translocating through NPCs to the cytoplasm, the complex is disassembled via RanGTP hydrolysis.114 Each exportin recognizes cargo by a different mechanism, and there is no common rule for cargo recognition.115
Exportin 1
Exportin 1 (XPO1, Exp1, also known as chromosomal region maintenance protein 1, CRM1) is the best characterized exportin.115 There are approximately 200 validated XPO1 protein cargoes, including p53, FOXO, Survivin, TFEB, and the cyclic GMP-AMP synthase cGAS.116,117,118 XPO1 cargoes are often involved in translation, cytoplasmic mRNA metabolism, vesicle coat complexes, and centrosome proteins.119,120. XPO1 interacts with leucine-rich nuclear export signals (NES), which are typically made up of four large hydrophobic residues separated by 1–3 linker residues (conforming to a Φ-X1–3-Φ-X2–3-Φ-X1–3-Φ motif).121 These hydrophobic residues are arranged linearly and bind in a long groove on the convex side of XPO1.122 The groove opens and closes dynamically, and RanGTP binding to the concave side stabilizes the open groove conformation.123 On the other hand, cargo binding displaces a loop on the concave side (H9 loop) and prepares XPO1 for RanGTP binding. Utilizing different protein adaptors, XPO1 can also export a variety of RNA molecules.124,125 XPO1 is frequently overexpressed in cancers and impairs the function of many tumor suppressors by exporting them to the cytoplasm.126
Exportin 2
Exportin 2 (XPO2, cellular apoptosis susceptibility, CAS, or chromosome segregation 1-like, Cse1, Cse1L) is a dedicated nuclear export factor for the classical nuclear import adaptor Impα, which is unable to traverse NPCs alone.127 By wrapping around RanGTP and Impα and folding the IBB in the NLS binding sites of Impα, XPO2 ensures cargo dissociation from Impα before export.128 XPO2 depletion alters the localization of multiple silencing factors and reactivates many repressed genes, due to its indispensable role in classical nuclear import.129 As Impβ1, XPO2 is overexpressed in many cancers.130,131
Exportin 5
Exportin 5 (XPO5) exports pre-miRNA, and this step is necessary for proper miRNA maturation.132,133 The crystal structure illustrates that Exp-5:RanGTP recognizes the 2-nucleotide 3’ overhang structure and the double-stranded stem of pre-miRNA.134 Likely through the same RNA interface, XPO5 exports tRNA and other double-stranded RNA molecules, as well as co-exports proteins bound to these RNAs.135,136,137,138 Impaired miRNA maturation due to XPO5 dysregulation such as genetic mutation and phosphorylation-mediated inhibition has been observed in several cancers.139,140 However, XPO5 was reported to be expressed in colorectal cancer that promotes the expression of oncogenic miRNA, but how this is selective for oncogenic miRNA but not tumor-suppressive miRNA is not clear.140 Exportin 6 and Exportin t are reviewed in the Supplementary File.
Bidirectional transporters
Biportins can function as importins to import cargoes or as exportins to export cargoes. There are three verified biportins in humans, and RanBP17 is classified as a biportin due to its high sequence homology to the biportin Exportin 7. The use of dedicated importins and exportins may allow for more diverse cargo recognition modes and more specific pathway control. On the other hand, using biportins in transport is likely more economical than using importins and exportins separately.
Importin 13
Importin 13 (IPO13) is a well-characterized biportin which imports glucocorticoid receptor GR, the exon junction complex components Mago-Y14, the E2 SUMO-conjugating enzyme Ubc9, programmed cell death 5 PDCD5, while exporting translation initiation factor 1 A eIF1A.141,142,143 A proteomic study demonstrated that IPO13 binds to many cargoes functioning in chromatin modification, chromatin remodeling, and transcription.88,144 Crystal structures of IPO13 in complex with Mago-Y14, Ubc9, and eIF1A have illustrated the mechanism by which this importin uses different surfaces to interact with different cargoes and how it plastically changes conformation upon binding to different cargoes.142,145 IPO13 overexpression plays a role in several cancers, and loss-of-function mutations cause defects in eye morphogenesis,146,147,148 but which cargo(es) mediate these pathological consequences are unknown.
Exportin 4
Exportin 4 (XPO4) mediates nuclear import of transcription factors Sox2 and SRY, the glycolytic enzyme PKM2, as well as mediates nuclear export of Smad proteins, the hypusine-containing translation factor eIF5A, and interestingly, a subset of circRNAs.149,150,151,152 Many XPO4-imported cargoes identified by mass spectrometry are RNAP II elongation factors and mRNA processing factors.88 The export cargo eIF5A is bound to the convex and concave surface of XPO4, with the hypusine bound in an acidic pocket.152 It is unclear how XPO4 recognizes other cargoes, but its plasticity may play a role in binding to different cargoes. Reduced expression of XPO4 due to copy number variation sustains nuclear Smad levels and TGFβ signaling, thereby enhancing the severity of fibrosis in patients with metabolic-associated fatty liver disease.153
Exportin 7 and RanBP17
Exportin 7 (XPO7) was initially identified as an exportin for RhoGAP1 and 14-3-3sigma.154 It was then demonstrated that XPO7 could also recognize positively charged folded domains and mediate the nuclear import of NFκB/p65.155 A recent proteomic study showed that XPO7 may import and export hundreds of cargoes with diverse structures and functions.156 How XPO7 recognize cargoes has not been reported. Depletion of XPO7 correlates with poor overall survival in several cancer types due to lack of oncogene-induced senescence caused by insufficient nuclear localization of p21 transcription factor TCF3.157 RanBP17 is 67% identical to XPO7 but little is known about this protein.
Smaller size transport factors
There are a few smaller-sized transport factors that do not form α-solenoid. However, they function as karyopherins and are capable of recognizing cargoes and translocating through NPCs.158 Unlike typical karyopherins, they are very specific in cargo recognition, do not rely on the RanGTP for cargo binding and dissociation, and contain fewer FG pockets.159
Nuclear Transport Factor 2
One ‘small karyopherin’ is Nuclear Transport Factor 2 (NTF2), which contains only 127 amino acids. NTF2 is a dedicated RanGDP nuclear import factor that recycles inactive RanGDP to the nucleus.160 NTF2 forms a homodimer and uses a distinct hydrophobic cavity for recognition of RanGDP.161,162 Two identical FxFG binding sites within the dimer are used for FG binding and NPC translocation.158 How NTF2 is dissociated from RanGDP in the nucleus is unclear, but NTF2 inhibits the guanine nucleotide exchange activity of RCC1 on Ran.163 Nuclear translocation of Ran may simultaneously import ankyrin repeat proteins and the filamentous actin capping protein CapG via a piggyback mechanism.162,164,165
NXF1 family
The nuclear RNA export factor 1 (NXF1, also known as TAP) family of proteins possesses an NTF2 domain capable of FG repeat binding and interacts with NTF2-like export factor 1 (NXT1) to form a heterodimer reminiscent of the NTF2 homodimer.166,167 This heterodimer binds to FG repeats but not RanGDP.166 NXF1 facilitates mRNA nuclear export since it also contains several other domains that interact with RNA and other mRNA processing factors, such as the TREX complex.168 Unspliced RNAs are generally not exported, but type-D retroviruses use a ∼130 nucleotide RNA called the constitutive transport element (CTE) to bind NXF1-NXT1 without protein adaptors (e.g., TREX) to export their unspliced genomic RNA.169 Structural analysis shows that this CTE-RNA forms a symmetrical stem-loop motif that binds to a symmetrical site formed by two copies of NXF1-NXT1 dimers.167 In humans, NXF1 is a major mRNA export factor, but there are a few other less-understood NXF family export factors, such as NXF2 and NXF3.170,171 NXF2 appears to be a tissue-specific mRNA export factor.171 Interestingly, NXF3 lacks FG binding pockets and instead relies on binding to XPO1 to translocate through NPCs, illustrating the diversity of RNA export.125,172
Hikeshi
The heat shock nuclear import factor Hikeshi contains 197 a.a. and is structurally unrelated to NTF2. Under conditions of heat shock, importins are globally downregulated and Hikeshi mediates nuclear import of molecular chaperone Hsp70 to counteract heat-shock damage and increase cell viability.173 Hikeshi contains an FG-binding N-terminal domain (NTD) and a C-terminal dimerization domain, and forms an asymmetric dimer that recognizes the full-length ATP-bound Hsp70.174 Interestingly, an loop in NTD contains a FG motif that can dock into its own FG pocket, thereby autoinhibits its interaction with FG nucleoporins and nuclear import function. How this autoinhibition is lifted under heat shock, how Hikeshi recognizes Hsp70, and whether Hsp70 is exported by Hikeshi after completing its nuclear function are unclear.175
The transport adaptor molecules
The transport adaptor itself does not have NPC translocation capabilities; however, it can bind karyopherin and cargo at the same time, thereby facilitating cargo transportation. They play important roles in nuclear transport, and in fact, alpha karyopherins are the first ‘karyopherins’ identified.176 Any protein that contains an NES or NLS and forms a tight complex with another protein/RNA is a potential nuclear transport adaptor. Because the list of adaptors is very long, except for the few examples shown above, two classes of well-studied adaptors with broad utility are reviewed here.
Importin α family
The importin α (Impα, or karyopherin α) family of adaptors functions in classical nuclear import, and it recognizes classical NLS signals. A classical NLS contains one or two stretches of polyK/R (2-4) sequences which bind to one or two acidic patches in the concave surface of Importin α.177 Impα contains an N-terminal Importin beta binding (IBB) domain that directly binds to Impβ1.178 This IBB can also bind to its own NLS binding sites, playing an autoinhibitory role so that cargo binding only occurs in the presence of Impβ1.179 In humans, there are seven Impα family members (Impα1 - Impα7) that are ~ 50%–80% identical in sequence and completely identical in the NLS interaction surface.180 These members are not entirely redundant, as they differ in cargo specificity and tissue- or developmental-stage-specific functions.181,182,183,184 Impα binds to a broad range of cargoes, including NFκB, STAT transcription factors, Ebola virus VP24 protein (eVP24), and influenza Polymerase PB2, thereby often involved in different cancers and viral infections.185,186,187,188
Snurportin
Snurportin (also known as Snurportin 1, SNUPN) is the nuclear import adaptor for m3G-capped U snRNPs, which participate in pre-mRNA splicing.92 Similar to Impα, Snurportin uses an IBB domain to interact with Impβ1.189 SNUPN contains an NES and is recycled to the cytoplasm via XPO1.190 Structural analysis has revealed that SNUPN binds to XPO1 in a manner incompatible with snRNP binding, thereby ensuring cargo unloading prior to nuclear export.191
Ran system proteins determining transport direction and speed
The transport directionality of importins, exportins, and biportins relies on an elaborate RanGTP system.5 This system generates the RanGTP gradient, strictly partitioning RanGTP in the nucleus and RanGDP in the cytoplasm.192 This RanGTP gradient is maintained by the nuclear-specific distribution of Ran guanine nucleotide exchange factor (GEF) RCC1 and the cytoplasm-specific localization of the GTPase-activating protein (GAP) RanGAP1 (Fig. 4).193 RanGDP, which is continuously generated throughout transport, is recycled to the nucleus by the aforementioned NTF2. In addition to these essential factors, four Ran binding proteins (RanBP1, RanBP2, RanBP3, and Nup50) regulate the interaction between RanGTP and karyopherins, accelerating transport speed.194
The RanGTP system and regulatory proteins. Ran is predominantly GTP-bound in the nucleus and GDP-bound in the cytoplasm. RanGTP is exported to the cytoplasm in complexes with karyopherins (either importins or export-cargo-bound exportins). In the cytoplasm, RanBP1 or RanBP2 (Nup358) promotes the dissociation of RanGTP from karyopherin complexes, allowing RanGAP1-mediated GTP hydrolysis. RanGDP is recycled back to the nucleus by NTF2, dissociated from NTF2, and reloaded with GTP by chromatin-bound RCC1 (GEF). RanBP3 enhances the recruitment of RanGTP to exportins
Ran
The Ras-related nuclear protein Ran contains a typical small GTPase domain and a C-terminal flexible tail that interacts with Ran-binding domains (RBD).195 Ran is active when it is GTP-bound and inactive when it is GDP-bound. RanGTP typically binds importins with nanomolar to picomolar affinities, and its binding either directly clashes with the cargo or induces an importin allosteric change to induce cargo dissociation.196,197 The binding affinity between an exportin and its cargo is usually higher than micromolar; however, RanGTP and the cargo cooperatively bind to the exportin at an affinity of tens to hundreds of nanomolar.198 In the cytoplasm, RanGTP in the export complex (either RanGTP-importin or RanGTP-exportin-cargo) is hydrolyzed to RanGDP through the cooperative action of RanGAP1 and RanBP1/2.67 In mitotic cells, RanGTP production is localized to chromosomes to promote local spindle assembly and at a later stage, local NE and NPC assembly.199,200,201
RCC1
Regulator of chromosome condensation (RCC1) contains an NLS that mediates its nuclear import and binds chromatin in the nucleus.202 RCC1 directly binds nucleosomal DNA via its N-terminal tail and a DNA binding loop, and it binds nucleosomal histones via a switchback loop.203,204 It collapses the P-loop of Ran to release bound nucleotides, and increases guanine nucleotide dissociation by over five orders of magnitude.205,206 GTP/GDP exchange catalyzed by RCC1 is indiscriminate, but due to the greater abundance of GTP compared to GDP in cells, nuclear Ran is eventually charged with GTP.205 Because NTF2 inhibits RCC1-mediated nucleotide exchange, an unknown ATP-dependent factor dissociates RanGDP from NTF2 to allow for RCC1-catalyzed guanine nucleotide exchange.163,207 Chromatin-bound RCC1 is responsible for local production of RanGTP in mitotic cells.
RanGAP1
RanGAP1 (RanGAP) is a cytoplasm-localized Ran-specific GAP recruited to the cytoplasmic filament protein Nup358 when SUMOylated.69,208,209 RanGAP1 does not use an arginine finger but positions Ran’s catalytic glutamine in the active conformation to trigger hydrolysis.210 RanGAP1 is anchored to the kinetochore and mediates chromatid segregation during mitosis, and depletion of RanGAP1 drives chromosome instability and tumorigenesis.211,212
RanBP1, RanBP2, RanBP3, and Nup50
RanGTP is tightly wrapped within karyopherins and is inaccessible to RanGAP1 when nuclear export complexes (RanGTP-importin or RanGTP-exportin-cargo) enter the cytoplasm.213,214 RanBP1 is a coactivator of RanGTP hydrolysis that increases the rate of RanGAP1-mediated RanGTP hydrolysis by an order of magnitude.215 This is achieved through its Ran-binding domain (RBD), which tightly binds to RanGTP and increases the rate of karyopherin-RanGTP dissociation.127,216 RanBP1 contains an NES and is located exclusively in the cytoplasm.217 RanBP2 (also known as Nup358) contains four RBDs functioning similarly to RanBP1, namely, in dissociating RanGTP from karyopherins and allowing RanGAP1-mediated GTP hydrolysis.218 In contrast, RanBP3 is a nuclear-localized RBD-containing protein that promotes nuclear export cargo assembly.219 RanBP3 contains several FG sequences that can form high-affinity anchors with exportins and an RBD domain, facilitating recruitment of RanGTP to exportins.220 This lowers the entropic barrier for RanGTP loading, as exportins typically have low affinity for RanGTP.221,222 Basket-localized Nup50 contains a high-affinity importin-binding FG domain and a C-terminal RBD that can recruit RanGTP to accelerate cargo dissociation from importins.223,224 These domain features of Nup50 enable it to increase the rate of nuclear import complex disassembly and, ultimately, nuclear import.
RanBP2, SUMOylated RanGAP1, and Ubc9 together form the NPC-localized SUMO E3 ligase, thus potentially linking SUMOylation and nuclear transport.225,226,227 SUMOylation of sites within or adjacent to the NES or NLS can disrupt karyopherin binding or alter the binding partner of modified proteins to render them inaccessible to karyopherins, thereby altering protein localization.228,229 SUMOylation of a protein may also enhance its nuclear import or export, but the mechanisms are largely unknown.230 On the other hand, nuclear transport also regulates protein SUMOylation. The nuclear import of many proteins, such as Sp100 (a component of the PML nucleosome), is critical for their SUMOylation.231,232 Although these proteins may undergo SUMOylation during nuclear entry, their SUMOylation may also occur inside the nucleus by other SUMO E3 ligases. In fact, most SUMO-modifying enzymes and SUMOylated proteins, including many kinetochore proteins, are found in the nucleus.233,234 For example, the kinase Aurora B, a key regulator of mitosis, is SUMOylated at the centromere in early mitosis by SUMO ligases including the RanBP2 complex.235 Interestingly, NPC also binds to deSUMOylase. The major de-SOMOylating enzyme SENP2 localizes to NPCs by binding to Nup153 and is critical for the de-SUMOylation of ribosomal precursors and their subsequent nuclear export.236,237 Removal of the highly hydrophilic SUMO groups may reduce energy required to penetrate the hydrophobic NPC barrier, which is especially important for translocation of large cargoes such as ribosomal precursors. Exported proteins could theoretically be SUMOylated by the RanBP2 complex, but reports on this are limited.67 Among many SUMO-regulated processes, gene expression, DNA damage response, and immune response can occur in the vicinity of NPCs, and future discoveries on how NPC-mediated SUMOylation participates in these processes to impact diseases such as tumors and infections are anticipated.238,239 It remains largely unclear what determines whether a translocating cargo is SUMOylated and how SUMOylation and nuclear transport cooperate in specific pathways.
Translocating across the NPC barrier
Nucleoporins, karyopherins, and Ran system proteins work together to transport cargo through the NPC. Each NPC can transport cargo at a staggering rate of ~ 1000 molecules per second, especially considering that it simultaneously prevents non-specific passive diffusion.240 The passive diffusion size limit is reported to be 40 kD, but few macromolecules employ passive diffusion to cross NPC due to low efficiency.241 Although much is known about the individual NTPs, we remain uncertain how NPC simultaneously achieve such a high level of transport efficiency and selectivity. Both the barrier and its interaction with karyopherin are highly dynamic and complex, making them difficult to study with most existing techniques.242 The variety of different models that have been proposed highlights our current lack of consensus in this regard.243,244,245,246,247
The nature of the barrier is highly debated. For example, whether the barrier is cohesive or non-cohesive, or in simpler terms, whether the barrier is formed of highly condensed “hydrogels” or more dynamic and loosely packed “polymer brushes”.248,249 GLFG repeats containing nucleoporins such as Nup98 are highly cohesive and form hydrogels in vitro at physiological concentration, but charged FG nucleoporins are less cohesive and do not naturally form hydrogels.250,251 The hydrogels formed in vitro exhibits many characteristics similar to the NPC barrier.252 It is not difficult to imagine that by anchoring in the relatively rigid NPC scaffold, the GLFG repeats are locally enriched,253 thereby forming a hydrogel barrier in NPC.244,254 However, this raises the question of whether and how karyopherins can rapidly melt and thus pass through cross-linked gels as rapidly as observed. Furthermore, high-speed atomic force microscopy revealed that the center of the barriers was entangled but ‘did not condense into a tightly cross-linked network’.255 Another model proposed that the highly dynamic FG repeats prevent the passage of non-interacting macromolecules by means of entropic exclusion, i.e., FG repeats exclude passive diffusion by forming a non-cohesive and highly entropic ‘virtual gate’.22,256 Invasion of inert macromolecules limits the entropy of the FG nucleoporins, thus being energetically unfavorable. Regardless of the debates, it is now known that the FG domains account for only ~ ¼ of the molecular mass of the NPC lumen, with the other ¾ being karyopherins and the cargoes they carry.257 A number of studies have highlighted a ‘karyopherin-centric’ model whereby karyopherins are integral constituents of the barrier and are critical for preventing NPC leakage.258,259,260 This model could complement both the hydrogel model (to reduce cohesiveness) and the virtual gating model (to outcompete non-specific diffusion).259,261 Although NPCs are somewhat heterogeneous in composition, it is unlikely that different gating mechanisms exist in different NPCs, but more likely that they coexist in all NPCs.262
Another controversy concerns the process of translocation, i.e. how karyopherins (with or without cargo) translocate from one side of NPC to the other.263 One of the earlier models proposed that certain FG nucleoporins bind karyopherins on one side of the NPC, escort them across the barrier, and release them on the other side.224,264 However, data generated later showed that the interaction between FG repeats and karyopherins is rather dynamic: FG pockets rapidly binds, dissociates, and rebinds other FGs in vicinity.265 Thus, rather than remaining tightly bound to FGs of one nucleoporin throughout transport, it is more plausible that karyopherins rapidly “slide” on the FG repeats of different nucleoporins to move forward.266 The ‘Brownian motion’ model suggest that the translocation process in the barrier is energy-independent and directionless.267,268,269 However, this model does not account for two important facts: (1) the distribution of different types of FGs, i.e., XXFG, GLFG, and FXFG, is asymmetrical in NPCs,270,271 and (2) cargo-loaded karyopherins always have a stronger affinity for the FG type on their destination side.272,273,274,275 Further, uncontrolled movement of karyopherins may cause traffic congestion and reduce transport efficiency, especially for large cargoes. Therefore, an ‘affinity gradient’ model demands that, besides RanGTP control of transport direction outside the barrier, the trafficking inside the barrier is constrained to a single direction, with the asymmetrically distributed FG types establishing an affinity gradient for karyopherins and luring it towards the high affinity end.273,274 Here, we further add that different FG types can be abstracted as hydrophobic balls of different sizes, with FXFG, GLFG, and XXFG representing large (2 F), medium (1.5 F), and small (1 F) balls, respectively (two Fs in FXFG or LF in GLFG are held together when inserted into FG pockets, as illustrated by different crystal structures, Fig. 5). RanGTP and cargo binding regulate the FG pocket size of karyopherin, shaping it selective for certain size balls, as observed in the ‘reversible collapse’ model where Importin β1 but not Importin β1-RanGTP can bind and collapse FXFG containing Nup153.249 Take importin as an example, after cargo binding, its FG pocket enlarges to bind 2 F balls and move along the 2 F gradient to the basket side. In the nucleus, RanGTP binding reshapes the importin FG pocket to select for 1 F balls and drive the importin to the cytoplasmic side. Free importins tend to stay in one compartment (cytoplasm for most importins) since it is energetically unfavorable to move against the affinity gradient. This prevents energy wasting, since GTP is consumed during transport (via RanGTP hydrolysis) even when there is no cargo. Furthermore, this model could explain why more karyopherins are required for large cargoes to cross the NPC:276 more karyopherins provide more energy (moving down the affinity gradient yields energy) or traction force to overcome the energy required for penetration of large cargoes. The affinity gradient can provide each karyopherin with energy equal to that generated by RanGTP hydrolysis in one round of import and export, if not considering any energy loss. Along this thread, the cytoplasmic filament and nuclear basket can use the affinity gradient to select export complexes and import complexes, and only at these exposed locations, these complexes are terminated by RanGTP hydrolysis and RanGTP binding, respectively. Without these cytoplasmic and nuclear extensions, these import and export complexes may spend a longer time in the transport channel where RanGTP and RanGAP are excluded, and ultimately ruduce transport efficiency. This model could better explain the observed high efficiency of NPC transport.
Model of uni-directional translocation within NPCs. In NPC, XXFG (small ball), GLFG (medium ball), and FXFG (large ball) repeats are not uniformly distributed but exist in concentration gradients. NLS binding to importin in the cytoplasm rearranges importin HEAT repeats to generate large pockets with high affinity for FXFG repeats, thereby moving importin and the bound cargo toward the nucleus with the aid of the FG gradient. Nuclear RanGTP binding renders importin surface pockets small and selective for XXFG repeats, driving the export of the complex. Exportin can also take advantage of this FG concentration gradient. Concentration gradients of different FG repeats provides a traction force and restrains directionality, thereby contributing to the high transport efficiency of NPCs. This model also explains the biological significance of the cytoplasmic filaments and the nuclear basket in nuclear transport (see text)
Nuclear transport-independent functions of nuclear transport proteins
In interphase cells, NTPs play an important role in mediating the nuclear import and export of macromolecules. In mitotic cells, these proteins continue to regulate aspects of mitosis through fundamental principles of nuclear transport.277 Many NTPs act as molecular chaperones for highly basic cargoes to prevent aggregation and cellular degradation or form biomolecular condensates through phase separation.278 Some NTPs also interact extensively with chromatin, regulating its structure and transcription.279 Readers are redirected to these works for other atypical NTP functions such as cilia transport and nuclear sizing.280,281,282,283
Mitosis
In mitotic cells, the RanGTP system signals the location of chromosomes as a global positioning system (GPS). The RanGTP system cooperates with karyopherins to modulate key mitotic factors that are usually cargoes of karyopherins. Many nucleoporins are also critical players in mitosis and are highly involved in different stages of mitosis. These NTPs orchestrate many aspects of mitosis, and their defects may lead to genetic instability and tumorigenesis through different mechanisms. Although less studied, meiosis is also regulated by NTPs due to its similarity to mitosis.284,285
The role of RanGTP, Impβ1, and Impα1 in mitosis
In mitotic cells, RCC1 is constantly bound to chromosomes, continues to generate RanGTP surrounding the chromosome and promotes local mitotic spindle assembly and functioning.286 Defects in RCC1 localization or function perturb the RanGTP gradient, resulting in chromosomal misalignment, abnormal spindle pole number, abnormal chromosome segregation, and genome instability.287 RanBP1 can form a tight complex with RCC1/RanGTP and inhibit RCC1 function, thereby regulate spatial distribution and magnitude of mitotic Ran-GTP production at different stages.288 Many spindle assembly factors (SAFs), including NuMA, HURP, TPX2, and APC, are classical nuclear import cargoes, and a high concentration of RanGTP in the vicinity of the chromosome releases SAFs from Impβ1 and Impα1 (Fig. 6a).277 These released SAFs participate in microtubule nucleation, growth, stability, and organization.289 In the cortical region, Impβ1 and Impα1 inhibit the mitotic function of SAFs by binding to the NLS of these SAFs. NLS binding by Impβ1 and Impα1 often sterically masks the functioning region of an SAF, e.g., the microtubule-binding region of NuMA.290 This intricate system prevents spindle assembly at nonchromosomal locations. RanGTP regulation of spindle assembly is not a switch but rather a gradient from the chromosome to the cell cortex where importins and SAF activities are gradually tuned.291
Role of nuclear transport proteins in mitotic spindle assembly and chromatid segregation. a Chromosome-bound RCC1 catalyzes the production of RanGTP around chromosomes. RanGTP near the chromosomes dissociates spindle assembly factors (SAFs) from bound importins, and the released SAFs promote spindle assembly. In the cortical region of the cell, spindle assembly is inhibited by excess importins. b The Y complex nucleoporins are critical for the recruitment of γ-TuRC, which induces k-fiber formation (kinetochore-initiated spindle microtubules). XPO1 is recruited to the Y complex and strengthens the connection between the k-fiber and the kinetochore. RanBP2 and RanGAP1 are recruited by XPO1 and mediate chromatid segregation at anaphase
In addition to spindle assembly, the RanGTP-Impα1/β1 system regulates many other events in mitosis. For example, Impα directly competes with p115, a vesicle-tethering factor, for the interaction with the Golgi matrix protein GM130, inhibits p115/GM130-mediated vesicle fusion, and promotes Golgi disassembly in the early stages of mitosis.292 During anaphase, the TPX2 NLS is phosphorylated, and the bound importin α and β1 are dissociated, allowing Eg5 recruitment to promote centrosome separation.293 The formation of NE in late mitosis requires Lamin B-coated NE precursor vesicle-vesicle fusion at the vicinity of chromatin, a process possibly induced by local dissociation of Impβ1 from Lamin B receptor, which then simultaneously binds chromatin and NE vesicles.294 Likewise, nuclear pore assembly is also regulated by RanGTP and classical nuclear import karyopherins.295,296 These studies collectively highlight a model in which RanGTP regulates the activity of many mitotic factors through Impα1/β1 at different mitotic stages to ensure proper chromosome segregation.
The role of other karyopherins in mitosis
In addition to Impβ1/Impα1, TNPO1, TNPO2, and potentially other importins, are involved in mitosis.277 For example, TNPO1 is known to inhibit recruitment of the Y complex to kinetochores and suppress mitotic spindle assembly, a process that is counteracted by RanGTP.297 Furthermore, inhibiting TNPO1 induces the formation of microtubule asters in the mitotic cytosol, while cells depleted in TNPO1 display defects in spindle and cytokinesis.298 Other importins are rarely reported in mitosis regulation, but their importance should not be neglected because they can also bind many mitotic factors.88
As anticipated, exportins are involved in mitosis. In particular, exportin XPO1 is recruited to kinetochores via RanGTP- and NES-binding and is required for stabilizing the connection between kinetochores and k-fibers (kinetochore-initiated spindle microtubules).299 XPO1 is also present at centrosomes, recruiting pericentrin and γ-tubulin ring complex (γ-TuRC) to nucleate spindles from the centrosome.300 In addition, centrosome-localized XPO1 may also recruit other NES proteins such as NPM, p53, BRCA1, and cyclin B, to ensure the mitotic fidelity and prevent genomic instability.301 For example, XPO1-RanGTP controls the spatial/temporal recruitment of NES-containing NPM to prevent centrosome reduplication.302 Generation of the microtubule organizing center (MTOC) at the NPC in yeast requires Nup159 (human Nup214), XPO1, RanGTP, and the MTOC protein Mto1, where XPO1 simultaneously binds the Nup159 FG domain and the Mto1 NES to link them together.300 The E3 ligase Nup358/RanGAP1/Ubc9, which is recruited to kinetochores by XPO1, SUMOylates and recruits TopoIIσ to decatenate sister centromeres prior to anaphase onset.299,303 Nup358 haploid mice develop cells with anaphase bridges and severe aneuploidy, and are highly susceptible to tumor formation.257
The role of nucleoporins in mitosis
As shown in Fig. 6b, some of the dismantled NPC parts, especially the Y complex, are recruited to kinetochores and centrosomes, where they continue to function during mitosis.304 The recruitment of the Y complex to the kinetochore occurs through its component ELYS, and this process is inhibited by Impβ1 and TNPO1 that compete with kinetochores for ELYS binding.297,305 The kinetochore Y complex recruits γ-TuRC which induces k-fiber formation.285,306 The presence of the Y complex at kinetochores is required for the recruitment of XPO1 as well as chromosome passenger complex (CPC) proteins, a critical factor in chromosome alignment and segregation.307,308
Moreover, the proper expression levels of Nup98/Nup88/RAE1/Tpr/Nup153 are critical for spindle polarity, preventing aneuploidy and tumorigenesis.309 Mechanistically, Nup98 and RAE1 form a complex with the anaphase-promoting complex/cyclosome (APC/C) to inhibit premature progression into anaphase through inhibition of APC E3 ligase activity.310 Sequestration of Nup98/RAE1, by RAE1/Nup98 haploinsufficiency or overexpression of Nup88, can activate APC/C and induce degradation of the mitotic kinase PKL1, disrupt normal centrosome separation, and lead to aneuploidy.311 The spindle assembly checkpoint (SAC) proteins Mad1 and Mad2 associate with Tpr at NPC in interphase cells and are recruited to kinetochores in mitotic cells without Tpr, signaling to inhibit APC function until all kinetochores are attached to spindles.312 Cyclin B1-CDK1 is targeted to NPC at early mitosis and mediates Tpr phosphorylation to release Mad1 so that it can be recruited to kinetochore.313,314,315 In conclusion, studies have shown that NTS proteins play an indispensable role in mitosis, and that their dysfunction can distort mitosis and lead to genomic instability and cancers by different mechanisms. More studies are needed to reveal the complex spatiotemporal interactions and regulation mechanisms of NTPs in mitosis.
Disassembly and reassembly of NPC during mitosis
During mitotic entry, NPCs break down into subcomplexes within approximately 5 min and disperse to different regions of the cell.316 Phosphorylation of several nucleoporins is a decisive event for NPC disassembly and subsequent entry into mitosis.317,318 The responsible kinases include cyclin-dependent kinase 1 (CDK1), polo-like kinase 1 (PLK1), and NIMA-associated kinases (NEKs) NEK6/7.317,318 The CNT complex can recruit PLK1 to NPCs during mitosis in C. elegans.319 These kinases primarily hyperphosphorylate two nucleoporins, Nup98 and Nup35.318 Hyperphosphorylation of more than 15 sites in the Nup98 C-terminal domain dissociates Nup98 from NPCs and is the rate-limiting step in mitotic NPC disassembly.317 Nup98 and Nup53 are linker nucleoporins linking different NPC subcomplexes, and hyperphosphorylation of the intermolecular interaction sites disrupts their linker function, leading to their dissociation from NPC, dissociation of threaded subcomplexes such as the CNT complex, and exposure of more nucleoporins to be phosphorylated and disassembled.38,318 Y complexes are not further dissembled, but released into the mitotic cytoplasm, or recruited to kinetochores and assist in spindle assembly, or retained in the membrane with transmembrane nucleoporins to serve as templates for later reassembly.320 These studies highlight the role of hyperphosphorylation and inactivation of key linker nucleoporins in NPC disassembly.
It takes ~10 min to reassemble the NPC after anaphase onset.321 To start, the Y complex binds to chromatin via the C-terminal disordered region of ELYS.322 ELYS also recruits the phosphatase PP1, which dephosphorylates phosphorylated nucleoporins to allow their assembly.323,324 Chromatin-bound RCC1 and a high concentration of RanGTP in vicinity are critical in this process, since RanGTP relieves the inhibition of several importins on chromatin-Y complex interaction.277 Nup50 can bind and stimulate the activity of RCC1 and is also critical for NPC assembly.223 Mitotic NPC assembly and NE assembly are tightly coupled, possibly through the transmembrane nucleoporins.325 Membrane-embedded Pom121 can interact with the Y complex, allowing the nuclear membrane to form around newly formed (partial) NPCs.61,326,327 Another transmembrane nucleoporin, NDC1 is also critical for anchoring NPCs to membranes, since it interacts with (dephosphorylated) Nup35 which stabilize the inner ring subcomplexes.328 The recruitment of Nup98 to the inner ring and the Y complex may further stabilize the NPC scaffold.48 EM studies show that the cytoplasmic ring is assembled after the nuclear ring and inner ring.329 The mitotic reassembly of NPC described above is largely the reverse process of disassembly, but studies suggest that there may exist multiple reassembly pathways, e.g., cytoplasmic assembly of NPC precursors.330,331,332 More studies are needed to determine the proportion and detailed steps of different assembly pathways.
Regulation of biomolecular condensates
Owing to their acidic surface properties, karyopherins, especially importins, interact with many highly basic cargoes (HBCs).333 This interaction not only plays a role in the nuclear import of these HBCs but also prevents their binding to other cellular targets and sometimes their cellular degradation.278 A special group of HBCs is the RNA binding proteins (RBPs), including FUS, hnRNP A1, and TDP-43. These proteins contain intrinsically disordered regions that can phase separate within the cytoplasm to form membraneless liquid droplets or β-amyloid-like solid fibers.278 The chaperone activity of importins also disaggregates already oligomerized RBPs and may be exploited to halt or reverse neurodegeneration. In contrast, many FG nucleoporins can phase separate to form hydrogel-like permeable barriers or aggregate with other cellular condensates, playing key roles in physiological or pathological processes.
Ability to function as a molecular chaperone
Many highly abundant HBCs, such as histones and ribosomal proteins, readily aggregate with cytoplasmic polyanions such as RNAs.25 As early as 2002, Jakel et al. demonstrated that several importins, such as IPO4, IPO5, IPO7, IPO9, and Impβ1, can serve as chaperones for these HBCs. The chaperone activity of these importins requires their large acidic surfaces, which shield the basic patches in HBCs and thereby prevent the ionic aggregation of HBCs with cellular polyanions. This is conceptually similar to canonical chaperons that prevent hydrophobic aggregation of proteins with large hydrophobic surfaces. This chaperone activity not only prevents aggregation but also protects the HBCs from proteasome-mediated degradation, since aggregated proteins are prone to aggregation.334,335 Recent studies have confirmed and expanded upon this role.336,337 It has been shown that the disassembly of the IPO9-H2A-H2B complex requires the presence of DNA in addition to nuclear RanGTP.338 This stricter dissociation mechanism may also allow for the storage of unused histones.338 The chaperone function is not limited to importins, as it has shown that XPO4 can bind to the export cargo eIF5A and inhibit its undesired interactions before entering the cytoplasm.152
Ability to disaggregate RBPs
Some importin-chaperoned HBCs are the neurodegenerative disease-associated RBPs, including FUS, TAF15, hnRNP A1/A2, and TDP-43.278 These RBPs are typically larger and contain RNA recognition motif (RRM) domains, intrinsically disordered low complexity (LC) regions, and arginine-glycine-glycine rich (RGG) domains. These domains contain weak and multivalent interaction sites, predisposing these RBPs to undergo phase separation with or without RNA. Phase-separated RBPs can further form amyloid fibers under certain conditions.339 Importins, in contrast, inhibit their self-association and even dissolve aggregated RBPs (Fig. 7a).340 For example, TNPO1 inhibits and reverses fibrils formed by PY-NLS-containing FUS, TAF15, hnRNPA1, and hnRNPA2. Similarly, Impα and Impβ1 prevent and reverse TDP-43 fibrillation.341,342
Role of nuclear transport proteins in regulating biomolecular condensates. a Importins can act as molecular chaperons for highly positively charged cargoes, such as many RNA binding proteins. The interaction between importin and cargo inhibits cargo aggregation and disaggregates biomolecular condensates (including fibers) formed by cargo. Purple dots represent nuclear localization signal of cargo proteins. b Many FG nucleoporins can form biomolecular condensates either on their own and/or coaggregate with other biomolecular condensates, such as TDP-43 droplets and stress granules. Red dots represent FG dipeptides
The mechanism of importin-mediated RBP disaggregation is starting to be unveiled. First, an intact NLS in the cargo is required for the chaperone activity of importins. Equimolar importin are often required to fully dissolve the preformed hydrogels or fibrils.342 Second, binding to NLS alone is not sufficient to inhibit aggregation, since an antibody against FUS NLS did not inhibit FUS self-association.343 Similarly, XPO1 did not inhibit FUS aggregation when the NLS of FUS was replaced with an NES.344 This suggests that TNPO1 forms additional contacts with FUS in addition to its NLS interactions. However, these interactions are very weak, dynamic, and difficult to visualize using typical structural biology approaches.344 These interactions likely involve the acidic surfaces of importins and positively charged residues in the RRM/RGG domains of cargoes, as well as the FG pockets of importins and the FG dipeptides (or FG-like hydrophobic residues) of cargoes.278 For example, FUS contains two FG dipeptides and 19 FG-like YG dipeptides, and TDP-43 contains 8 FG dipeptides. Therefore, through NLS binding, importins may reduce the phase separation ability of RBPs by sequestering key elements that drive phase separation.345
Ability to form biomolecular condensates
FG domains are intrinsically disordered and contain weak multivalent interaction sites, such as positively charged residues and F/Y residues that can form π-cation interactions, and thus capable of phase separation like other intrinsically disordered domains. FG nucleoporins can be found in various cellular condensates, including amyloid-like fibers (Fig. 7b).346,347 Within the nuclear pore, the concentrated FG domains may aggregate into a condensed phase to form the selectively permeable barrier, although this is under debate.348,349 Karyopherins, which can form multivalent interaction with FG domain through FG pockets, coexist in these FG condensates and can fortify the NPC barrier to prevent NPC leakage. Additionally, many FG nucleoporins can phase-separate with other aggregation-prone proteins.248,350,351 Direct interaction between Nup98 and Tau is observed to promote phase separation of each other in vitro.352,353 Furthermore, fragments of Nup98 and Nup214, when fused with other DNA binding domains, can phase separate at chromosomal regions, a process that induces chromosomal looping and regulates transcription.354,355
Transcription regulation
As nuclear transport impacts the localization of different transcription factors, nuclear transport is naturally a critical step in transcription regulation.356 However, mounting evidence suggests that many nucleoporins can directly regulate transcription, independent of their function in nuclear transport.279,357 These nucleoporins can interact with transcription factors at promoters and enhancers.358 The end result may be either transcriptional activation or inhibition, and this regulation does not necessarily occur at the NPC locus.359 These activities are cell-type specific, and their dysregulation may drive the initiation and progression of different tumors.360,361
Transcription activation
More transcriptionally active genes are localized to NPCs from yeast to humans.362 Recruitment of active genes to NPCs may facilitate transcription factor binding immediately following nuclear import and coordinate transcription with subsequent nuclear export.363,364 Multiple studies have shown that nucleoporins can induce promoter-enhancer interactions, activating transcription.362,365 These nucleoporins are mainly from the nuclear outer ring and the nuclear basket.366 For example, Nup93 and Nup153 bind transcription factor-rich super-enhancers and drive the expression of key genes that specify cell identity.367 The nucleoporin Seh1 promotes transcription of proteins essential for oligodendrocyte differentiation through assembly of an Olig2-dependent transcription complex.368
Nucleoporins have been found to play a role in transcriptional activation not only at the nuclear pore complex but also in the nucleoplasm.359 Insides the nucleoplasm of Drosophila, several nucleoporins (Nup98, Nup62, and Nup50) interacts with development and cell cycle genes and activates their transcription.369 Similarly, Nup88 also binds to silent loci off-pore, and these nucleoporin-binding loci are often distinct from those NE contact sites.370 Nup98 can promote transcription by stimulating the ATPase activity of the DExH/D-box helicase DHX9.371 In leukemia, Nup98 is frequently fused to other DNA-binding homeodomain proteins, such as HOXA9, leading to the expression of oncogenes to drive leukemogenesis.372 The phase separation property of fusion nucleoporins seems critical for the transcription regulation activity. Interestingly, their condensation to chromatin depends on the chromatin-bound XPO1 that has formed a complex with RanGTP and a chromatin-bound NES-containing protein.373 Inhibition of XPO1 by leptomycin B disrupts the interaction of these nucleoporins with chromatin and reverses transcriptional activation mediated by these nucleoporins.373
Transcription repression
Less frequently transcribed heterochromatin is usually enriched at the nuclear periphery.374 Nup93 is associated with polycomb-silenced genes and physically interacts with a group of polycomb proteins, and polycomb repressive complexes containing Nup93 are more stable and localized to the nuclear periphery.375 Therefore, Nup93 may repress transcription by promoting heterochromatin formation.375 Similarly, Nup153 associates with the transcriptional start site of developmental genes and recruits polycomb-repressive complex 1, maintaining stem cell pluripotency in mammalian cells.376 Nup88 binds specifically to silenced genes; however, the regulatory mechanism is unclear.370
Genes near telomeres are less frequently transcribed due to the ‘positional effect’.377 Telomeres are localized at the nuclear periphery and bind silencing factors, such as Sir2, Sir3, and Sir4.378 In yeast, the Y complex component Nup170 (human Nup155), as well as the nuclear basket components Mlp1/2 (human Tpr), are critical for maintenance of the correct localization of telomeres.379,380 Furthermore, these nucleoporins can recruit silencing factors to telomeres.381 Depletion of these nucleoporins results in defective telomere silencing.382
Bimodal regulation
Actively transcribing genes are usually grouped into distinct topologically associated domains (TADs) with boundaries on both sides of the domain that insulate transcription within a TAD.383 The nuclear basket protein Nup153 interacts with key boundary proteins CTCF and cohesion to stabilize TADs.365 Therefore, Nup153 depletion leads to improper TAD boundaries as well as differential gene expression.365 Another study demonstrated that promoter binding by Nup153 increased gene expression, while transcriptional end site binding reduced gene expression.384 While it is conclusive that many nucleoporins can regulate transcription, whether phase-separation is involved in all these interactions and whether other NTPs regulate the process are largely unclear.
Diseases involving defects in nuclear transport proteins
Due to their high functional importance and relatively low gene redundancy, many NTPs are key players in different diseases. In particular, cancer cells often upregulate the expression of many karyopherins to alter the localization of cargoes or promote oncogenic transcription by creating nucleoporin fusion proteins.4,385,386 Defects in different NTPs downregulate nuclear transport and improperly localize key RNA-binding proteins such as TDP-43 in different neurodegenerative diseases.83 Many viruses exploit nuclear transport machinery to complete their life cycle in hosts and/or suppress host immune responses through impairment of nuclear transport.387,388 Inhibitors targeting various NTPs are being actively developed and clinically tested in relevant diseases.
Cancer
In the mitosis section, we showed that NTP dysfunction can result in improper mitosis, genetic instability and cancers. Cancer cells also frequently overexpress karyopherins or employ mutations to manipulate the localization of key proteins and RNAs to promote proliferation and evade tumor suppression. The most prominent example is overexpression or mutation of XPO1, which localizes many tumor suppressors to the cytoplasm to render them inactive. In addition, the transcriptional regulation function of several NTPs is also implicated in cancers. In leukemia patients, fragments of Nup98 and Nup214 are frequently fused to other proteins, resulting in fusion proteins that promote oncogenic transcription.
Overexpression of NTPs in cancer
Cancer cells often exhibit increased nuclear translocation velocity and capacity in response to faster signaling and metabolic stress, and many NTP proteins are overexpressed in cancer.98,389 XPO1 overexpression in many types of cancer correlates with disease severity and prognostic outcome in various studies.390 Mislocalization and inactivation of tumor suppressor proteins, such as P53, P21, and Rb, in the cytoplasm have been linked to XPO1 overexpression (Fig. 8a).391 Since overexpression of XPO1 is required to sustain multiple hallmark features of cancer,392 genetic or pharmacological inhibition of XPO1 is effective in a broad spectrum of cancer cells.131 XPO1 frequently mediates drug resistance, and XPO1 inhibitors were reported to enhance the efficacy of many clinically used drugs.115,393 However, the first-generation XPO1 inhibitor leptomycin B failed clinical trials due to high toxicity.394 Leptomycin B covalently binds to XPO1 and permanently inhibits its nuclear export function, but XPO1 is essential for the survival of all eukaryotic cells.16,395
Role of nuclear transport proteins in cancer. a Overexpression of XPO1 leads to mislocalization and subsequent inactivation of many tumor suppressors, such as P53. Inhibition of XPO1, for example, by the FDA-approved drug selinexor, correctly repositions the tumor suppressors in the nucleus and inhibits cancer cell growth. b In several hematopoietic cancers, nucleoporin fusion proteins cocondensate with other transcription factors around chromatin, induce aberrant chromatin looping and activate HOXA cluster oncogenes
Several reversible second-generation XPO1 inhibitors have been subsequently developed with significantly reduced toxicity.396,397,398 Among them, selinexor (KPT-330) was approved by the FDA in 2019 for the treatment of relapsed and refractory diffuse large B-cell lymphoma as well as multiple myeloma.115 Dozens of selinexor clinical trials are underway, either as a single agent or in combination with other therapies (Table 3).399,400,401 Eltanexor (KPT-8602), a next-generation inhibitor that is fast reversible and less permeable to the blood-brain barrier, is also being investigated in several clinical trials.402 These studies clearly demonstrate the efficacy of XPO1 inhibitors in patients with advanced and refractory human cancers, especially hematological cancers, either alone or in combination with pre-existing therapies. The adverse effects are mostly gastrointestinal and hematological, such as nausea, vomiting, anemia, and thrombocytopenia. In elderly patients with acute myeloid leukemia, reduced survival with selinexor was observed, but the reasons were unclear.403 See these reviews for earlier clinical trials.115,126,131
Canonical nuclear import factors Impβ1 and Impα1 are overexpressed in multiple cancers, albeit less frequently than XPO1.87 Overexpression of these proteins may lead to nuclear entry of many oncogenic transcription factors (such as PDL1 and β-catenin) to promote tumorigenesis.98,404 The overexpression of other Impα isoforms and other karyopherins, such as XPO5, XPO6, and Imp8, has been reported in a few specific cancer types.140,184,405,406,407 In addition to karyopherins, other NTPs, such as Ran, Nup93, and POM121, have also been reported to be overexpressed in cancers, playing a role in cancer initiation and/or progression.356,408,409 Mechanistically, overexpression of nucleoporins and Ran is unlikely to cause cancer via promoting nuclear transport, but rather through their other functions such as mitotic and transcriptional regulation functions.
Mutations that change protein localization
Mutation also plays a role in altering cellular localization of key proteins in cancers. Notably, the XPO1 E571K mutation is present in a quarter of patients with Hodgkin lymphoma and primary mediastinal B-cell lymphoma.410 Moreover, E571K accelerated leukemogenesis in a mouse model of chronic lymphocytic leukemia.389 This mutation altered XPO1 localization and the interactome of XPO1 in B-cell lymphoma.411 Structurally, E571 is located proximal to the NES groove, and the E571K mutation can increase the affinity for XPO1 cargoes that have more acidic residues in the NES sequence.412
Many studies have identified pathogenic mutations occurring within cargoes rather than karyopherins.87 For example, the tumor potential of cyclin D1 depends on its nuclear retention.413 T286 mutations, which specifically disrupts cyclin D1 phosphorylation and XPO1-mediated nuclear export, have been found in primary esophageal carcinoma samples.413 Moreover, nucleophosmin (NPM), which is localized in the nucleus of normal cells, is cytoplasmic in approximately one-third of acute myeloid leukemia samples and plays a key role in leukemogenesis.414 This cytoplasmic localization of NPM occurred because of a frameshift in its last exon generated a new NES sequence that promoted its nuclear export.414 Understanding these different pathogenic mechanisms can help guide the development of precise medicines which specifically correct the localization of a particular cargo.
RNA export dysregulation and cancer
Similar to protein nuclear transport, RNA export is altered in many cancers. Human mRNA can be exported via XPO1/RanGTP-dependent pathways and NXT1/NXF1-dependent bulk export pathway.415 Multiple factors within these pathways are dysregulated across various cancers, such as XPO1, THO1, HuR, and eIF4E.415,416,417 In particular, ~ 30% of human cancers upregulate eIF4E levels, exporting a set of mRNAs containing eIF4E-sensitive elements through the XPO1 pathway.418 Many of these mRNAs encode oncogenes, including cyclin D1, NBS1, cMyc, and MDM2, leading to activation of proliferation pathways such as Akt.419 There are over a dozen clinical trials using the eIF4E inhibitor ribavirin on different cancers.420 The bulk mRNA export factor NXT1 was also identified as a genetic dependency in neuroblastoma and several pediatric cancers.421
Defects in pre-miRNA export are also observed across many cancers. Mature miRNAs are often downregulated in cancer, and several mechanisms are known to reduce miRNA levels.422 In a subset of human tumors with microsatellite instability, several XPO5-inactivating mutations trapped pre-miRNAs within the nucleus and reduced miRNA-target inhibition.423 In another study, ERK phosphorylation of XPO5 induces a conformational change in XPO5, making it unable to load pre-miRNA appropriately.424 XPO5 phosphorylation is associated with poor prognosis in liver cancer patients.424 Epigenetic change and abnormal XPO5 expression levels also impact miRNA expression and have profound effects on tumorigenesis.132
Nucleoporin fusions that alter transcription
In a wide array of hematopoietic malignancies, chromosome translocations often result in Nup98 oncogenic fusion proteins associated with poor prognosis.425,426 Fusion proteins typically include the N-terminal FG domain of Nup98 and the C-terminal domain of a partner protein such as HOXA9.427 Many fusion proteins physically interact with mixed lineage leukemia 1 (MLL1) and nonspecific lethal (NSL) histone-modifying complexes, an interaction that is critical for its leukemogenesis ability.428,429 These fusion proteins can upregulate the HOXA cluster gene and inhibit hematopoietic precursor differentiation (Fig. 8b).385,425 In acute myeloid leukemia, the loss of the direct transcriptional target CDK6 severely attenuated fusion-driven leukemogenesis.430 In addition to Nup98 fusions, fusions containing fragments of Nup214 or Tpr have also been observed in several cancers and can similarly drive cancer progression.431,432,433
The oncogenic property of fusions depends not only on fused domains that bind DNA or modify histones but also on the FG domain of Nup98.385,434 The FG domain is capable of liquid-liquid phase separation and is critical for puncta formation on chromatin.354,372 This property promotes binding between the fused domain and chromatin, generating a broad superenhancer-like binding pattern that potentiates transcriptional activation of proto-oncogenes.354
Neurodegenerative diseases
Unlike cancers, nuclear transport is often impaired in neurodegenerative diseases (NDDs), including amyotrophic lateral sclerosis, frontotemporal dementia, Alzheimer’s disease, and Huntington’s disease.435 Cytoplasmic aggregation of RBP proteins such as TDP-43 and FUS is a hallmark of NDD.436 The microtubule-associated protein Tau is not known as an RBP but it is also capable of binding RNA and other RBPs, especially in disease.437,438 These RBP proteins are normally nuclear but are predominantly aggregated in the cytoplasm of diseased neurons.439 For example, mislocalization and aggregation of TAR-DNA binding protein 43 (TDP-43) is observed in ~98% of ALS cases.440 Similarly, cytoplasmic FUS aggregates are a pathological hallmark in a subset of patients with FTD or ALS. The loss of RBP nuclear function and gain of RBP cytoplasmic function are critical for the pathogenesis of NDD.439 Many studies indicate that impaired nuclear transport is responsible for RBP pathology and is a common factor in many NDDs,350 highlighting a promising area of research that could lead to the discovery of new therapies for NDDs.
The vicious cycle between nuclear transport impairment and RBP pathology
Impaired nuclear transport of RBP proteins is a major cause of RBP mislocalization. Some RBP cargo mutations that occur in neurodegenerative diseases promote accumulation and aggregation in the cytoplasm.278 For example, many FUS-NLS mutations impair TNPO1 binding, promoting cytoplasmic phase separation and stress granule partitioning of FUS.343 Phosphorylation or mutation of the NLS of TDP-43 disrupts the nuclear import and chaperone activity of Impα1/β1.441 Furthermore, many NTPs, including karyopherins and nucleoporins, are downregulated in NDD cells. Therefore, impaired nuclear transport is increasingly recognized as a pathogenic driver of neurodegeneration.442
The cytoplasmic aggregation of RBPs is not only a consequence of impaired nuclear transport but may in turn lead to defective nuclear transport. For example, cytoplasmic TDP-43 droplets may recruit and mislocalize importin-α, Nup62, RanGAP1, Ran, and Nup107, leading to inhibition of nuclear transport and eventual neuronal cell death.443 Likewise, pathogenic Tau can lead to NPC dysfunction by directly interacting with NPC components, causing their mislocalization.444 Therefore, current studies support the existence of a vicious cycle in NDD, i.e., the progressive deterioration of RBP localization and nuclear transport (Fig. 9).
Vicious cycle between cytoplasmic condensation of RNA binding proteins (RBPs) and defective nuclear transport. Cytoplasmic condensation of RBPs (such as FUS and TDP-43) recruits many nuclear transport proteins (such as importins, nucleoporins, and Ran system proteins) into the condensates and disrupts normal nuclear transport (especially import). Impaired nuclear transport further leads to cytoplasmic accumulation of RBPs and excessive condensation. The vicious cycle can be triggered by genetic factors such as mutations or external factors such as chronic stress
Nuclear transport protein abnormalities in NDD
Karyopherins, especially importins, play an important role in the nuclear import of RBPs. For example, upregulation of Impα3 reduces the cytoplasmic accumulation of TDP-43 and mitigates behavioral deficits in mice.445 Alternatively, inhibition of protein nuclear export also suppresses neurodegeneration.446 In addition to their canonical role in nuclear import, importins sculpt cytoplasmic membraneless organelles and dissolve the ‘irreversible’ precipitates or beta-amyloid-like fibers formed by RBPs, as explained in section 4.2.2.103 Therefore, karyopherins exhibit two distinct roles in NDD, as mediators of RBP nuclear localization and as molecular chaperones that inhibit RBP aggregation, and karyopherin abnormalities can trigger the onset and progression of NDD.
Karyopherin abnormalities are widely observed in NDD.83 For example, protein levels of Impα1 and XPO2 are reduced in the frontal cortex of FTD patients, and Impβ1 is reduced in the spinal cord of ALS patients.447,448 Furthermore, karyopherins are often sequestered into stress granules and TDP-43 droplets in diseased neuron cells.351,449 Several patient-derived TNPO2 variants that impair RanGTP or cargo binding have been shown to be responsible for neurodevelopmental abnormalities.450 These observations are consistent with genetic perturbation studies, suggesting that karyopherin abnormalities may mediate NDD pathogenesis. Upregulating the expression levels of several aforementioned importins in neuronal cells of patients is a promising strategy for the treatment of those NDDs without RBP NLS mutations.
In addition to karyopherin abnormalities, loss of nuclear pores, nucleoporin aggregation, and altered nuclear morphology are some of the most prominent features across a variety of NDD cells and animal models.451,452 Through phase separation, aggregated TDP-43, FUS, and Tau mislocalize FG nucleoporins and trigger the structural and functional impairment of NPCs.444,453 For example, the nucleoporin Nup62 colocalizes with TDP-43 granules in diseased brain cells.351,454 Although importins can reduce these cytoplasmic condensates formed by RBPs and nucleoporins, they may be overwhelmed and trapped by these excessive biocondensates in diseased cells.455 Impaired mRNA export is often observed in NDD cells, as many FG nucleoporins are critical for mRNA export.452,456 Furthermore, specific nucleoporin mutations may lead to cell type-specific neurological disorders.350 For example, a homozygous splicing mutation in Nup133 causes Galloway-Mowat syndrome, highlighting the importance of nuclear transport in NDD.457 Nucleoporin alterations and the consequential loss of NPC function may lie upstream of TDP-43 mislocalization in NDD.458 Abnormalities in nucleoporins (such as Nup62, Nup93, Tpr, and Nup153) also impact non-neuron cells, contribute to aging and premature aging at the organism level.459
Other factors that may initiate the vicious cycle
Many studies have demonstrated that C9ORF72 hexanucleotide (GGGGCC) repeat expansion (HRE, either familial or sporadic) may act as an initiator of the vicious cycle. HRE, the most common genetic cause of ALS and FD, encodes proteins containing extra intrinsically disordered regions. One of the encoded poly-GR proteins tightly binds to Impα1, directly disrupting the nuclear import of endogenous cargoes.460 A mutant C9ORF72 can induce proteasome-mediated degradation of select nucleoporins.461 Mutations that generate extra intrinsically disordered regions in proteins such as huntingtin and ataxin1 can similarly initiate the vicious cycle.462,463 In addition to the protein, HRE RNA initiates a decrease in POM121, which may further lead to downregulation of seven additional nucleoporins.464 The HRE RNA can also sequester RanGAP1 and distort the RanGTP gradient, disrupting nuclear integrity and transport.465 Several components of nuclear transport can also effectively combat the toxicity of C9ORF72 HRE by means of nuclear import and anti-aggregation.466
In addition to genetic factors, external factors may initiate the vicious cycle. The formation of stress granules is a typical response of cells to a broad range of stresses. However, stress granule formation may sequester critical NTPs, like RBP biocondensates, thereby inhibiting nuclear transport.467 Therefore, constitutive oxidative stress throughout aging may persistently impair nuclear transport and lead to irreversible NDD. Likewise, a recent study using Drosophila demonstrated that traumatic injury leads to NPC defects, impairing the RanGTP gradient, and leading to cytoplasmic aggregation of Nup62 and TDP-43.454 This may explain why traumatic brain injury is a predisposing factor for several neurodegenerative diseases. Since cellular localization of a protein is determined by both its nuclear import and nuclear export, nuclear export inhibitors may be applied after a traumatic injury to prevent NDD. Based on two encouraging preclinical studies,468,469 it is worth further testing whether XPO1 inhibitors can clinically slow down or even reverse some of the discussed NDDs.
Viral infection
Many viruses replicate in the nucleus of host cells and rely on the nuclear transport system for their nuclear entry. In addition, nuclear transport plays a role in other stages of the viral life cycle, such as uncoating and viral RNA export. Viruses have developed specific strategies to suppress host immune responses by targeting karyopherins, thereby avoiding clearance by the host. In addition to targeting karyopherins, viruses may inhibit or distort host nuclear transport by altering NPC integrity.
Exploitation of nuclear transport proteins to complete the viral infection cycle
Several RNA viruses and nearly all DNA viruses require access to the host cell nucleoplasm for replication.470 Following virus-cell fusion, a core consisting of capsid proteins (CA) surrounding the viral genomic DNA/RNA enters the cytoplasm of host cells.471 The NPC filament proteins Nup214 and Nup358 can bind to capsid proteins and are critical for NPC docking.472 Prior to genome import and replication in the nucleus, the core of most viruses must be uncoated by cytosolic host proteins such as karyopherins.473 TNPO1, which promotes the removal of M1 from the core by binding to a PY-NLS sequence in the matrix protein M1, is a common uncoating factor for influenza A virus (IAV) and human immunodeficiency virus type 1 (HIV-1).107
Most of these viruses utilize cellular nuclear import machinery for their nuclear entry.474 For example, IAV vRNP uses Impα7 for its nuclear import.475 Although viral mechanisms for crossing NPCs are complex and diverse, interaction with Impα/β1 is absolutely critical for nuclear entry of many viruses.476 The macrocyclic lactone ivermectin, which is reported to target IMPα/β1, has broad-spectrum activity against a variety of viruses, including HIV-1, DENV, ZIKV, West Nile virus (WNV), and SARS-CoV-2 (COVID-19).387 Our group, however, have observed no direct binding between ivermectin to IMPα1 or β1, nor inhibition of classical nuclear import using physiological relevant concentrations of ivermectin (unpublished). Through inhibition of the binding of NS5 and Impα1, N-(4-hydroxyphenyl) retinamide (4-HPR) has anti-ZIKV activity at low μM concentrations.387
Human retroviruses such as HIV and human T-cell leukemia virus type 1 (HTLV-1) require export of their intron-containing RNAs from the nucleus to the cytoplasm for translation and packaging. Two viral proteins, Rev and Rex, act as export adaptors to facilitate export of viral RNA through simultaneous binding of the viral RNA and the export factor XPO1.477 Inhibition of XPO1 thus results in sequestration of key viral accessory proteins and genomic materials in the host cell nucleus, thereby reducing the replication of viruses such as influenza, respiratory syncytial virus (RSV), and SARS-CoV.478 Clinical studies using the XPO1 inhibitor selinexor are currently in progress.388 Since XPO1 suppress inflammation and immune activation through the NFκB pathway, the application of XPO1 inhibitors in different cancers may also result in bacterial infection, a major cause of poor clinical outcomes.478,479 Therefore, combination with antibiotics should probably be considered in the treatment of cancers or viral infections.
Evasion of immune clearance by inhibiting protein nuclear import or mRNA nuclear export
As a defense mechanism, human cells upregulate the interferon response to combat viral infections. Typically, upon viral stimulation, IRF and STAT transcription factors are imported into the nucleus, subsequently transcribing and exporting an array of mRNAs encoding immune factors.480 Viruses have evolved different strategies to inhibit this process. Open reading frame 6 (ORF6) of SARS-CoV-2 binds to Impα1 to inhibit IRF3 nuclear import and the type I interferon response.481 Similarly, Ebola virus VP24 binds importin alpha proteins and inhibits STAT1 nuclear import, rendering cells refractory to IFNs.187 Zika virus NS2A protein induces degradation of Impα1 through chaperone-mediated autophagy.482 In contrast, the influenza virus NS1 protein blocks host mRNA nuclear export by directly interacting with the NXF1-NXT1 export machinery.483
Viruses can also disrupt the host immune response by targeting nucleoporins. For example, the 2Apro protease of poliovirus and rhinovirus cleaves Nup62, Nup98, and Nup153.484,485 Alternatively, mengovirus and cardioviruses can inhibit nuclear transport through induction of hyperphosphorylation of nucleoporins such as Nup162, Nup35, and Nup214.486,487 Cytoplasmic mislocalization of NPC parts, such as Nup214, Nup358, and Nup62, is frequently observed in cells infected with different viruses.472,488,489 Moreover, ORF10 and ORF6 from several viruses repress host mRNA export by interacting with RAE1 and Nup98.490,491 These actions inhibit antiviral responses and may also prevent cell death to allow viral replication or induce NE leakage to permit viral genome entry into the nucleus.472 While the above findings are well documented, actual infections are often more complex and dependent on the specific virus and the infection stage.492
Conclusion and perspectives
Due to the complexity and structural dynamics of NPCs, a complete atomic model has not been achieved thus far, despite enormous efforts. It is expected that with continuous improvements in resolution power and artificial intelligence, an almost complete atomic NPC structure will soon be obtained. Different conformations of NPCs may be obtained to illustrate how NPCs transit from one to another. While atomic models cannot be established for disordered regions, it should be possible to correctly understand their function in the broader context of structured regions. Such structural information is crucial for understanding the canonical function of NPCs, interpreting the pathogenic mechanisms of disease mutations, and developing targeted drugs for related diseases.
A few karyopherins, including XPO1 and classical nuclear import factors, are well studied; however, little is known about many other karyopherins. While some karyopherins (especially importins) are redundant in transporting certain cargoes, they do vary widely in terms of function and spatiotemporal expression. Furthermore, often selected karyopherins are reported to play key roles in different diseases. Therefore, it is important to study the structures, cargo recognition mechanisms, and affiliated pathways of each karyopherin. Furthermore, the development of potent and selective inhibitors has been limited to only a few karyopherins.493 The development of specific inhibitors is not only beneficial to basic research but may also provide new therapies for imminently threatening diseases.
In addition to their role in nuclear transport, nuclear transport proteins play roles in other cellular processes, such as mitosis, biomolecular condensate regulation, and transcription regulation. These functions share common fundamental principles with the nuclear transport function, including the interaction network and assembly principle, but can be completely distinguished from nuclear transport. For example, nuclear transport, mitosis, and gene transcriptional regulation of NTPs are important for tumorigenesis, while the nuclear transport and regulation of biomolecular condensate functions of NTPs are clearly involved in NDDs. Future studies involving NTPs should try to clarify which specific functions of an NTP are important for the phenotype or disease in question.
The nuclear transport system is a double-edged sword that keeps cells functioning properly. Generally, upregulation of nuclear transport may lead to cancer, and downregulation may lead to NDD. Viruses may either use nuclear transport to facilitate their replication or inhibit nuclear transport to evade immune surveillance. Therefore, while treating one disease, care must be taken to avoid causing another. For example, when treating tumors through inhibition of highly expressed NTPs, it is important to avoid triggering NDD development. Likewise, when upregulating importins to treat different NDDs, the risk of carcinogenesis should be considered. Tissue-specific targeting or delivery may be helpful in this regard.
References
Macara, I. G. Transport into and out of the nucleus. Microbiol Mol. Biol. Rev. 65, 570–594 (2001). table of contents.
Cautain, B., Hill, R., de Pedro, N. & Link, W. Components and regulation of nuclear transport processes. FEBS J. 282, 445–462 (2015).
Beck, M. & Hurt, E. The nuclear pore complex: understanding its function through structural insight. Nat. Rev. Mol. Cell Biol. 18, 73–89 (2017).
Wing, C. E., Fung, H. Y. J. & Chook, Y. M. Karyopherin-mediated nucleocytoplasmic transport. Nat. Rev. Mol. Cell Biol. 23, 307–328 (2022).
Izaurralde, E. et al. The asymmetric distribution of the constituents of the Ran system is essential for transport into and out of the nucleus. EMBO J. 16, 6535–6547 (1997).
Oka, M. & Yoneda, Y. Importin alpha: functions as a nuclear transport factor and beyond. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 94, 259–274 (2018).
Damizia, M., Altieri, L. & Lavia, P. Non-transport roles of nuclear import receptors: In need of the right balance. Front Cell Dev Biol. 10, 1041938 (2022).
Zhang, Z. M. et al. SAPCD2 promotes neuroblastoma progression by altering the subcellular distribution of E2F7. Cell Death Dis. 13, 174 (2022).
Maik-Rachline, G. et al. The nuclear translocation of the kinases p38 and JNK promotes inflammation-induced cancer. Sci Signal. 11, eaao3428 (2018).
Watson, M. L. Further observations on the nuclear envelope of the animal cell. J. Biophys. Biochem. Cytol. 6, 147–156 (1959).
Stevens, B. J. & Swift, H. RNA transport from nucleus to cytoplasm in Chironomus salivary glands. J. Cell Biol. 31, 55–77 (1966).
Moreland, R. B., Nam, H. G., Hereford, L. M. & Fried, H. M. Identification of a nuclear localization signal of a yeast ribosomal protein. Proc. Natl. Acad. Sci. USA. 82, 6561–6565 (1985).
Chi, N. C., Adam, E. J. & Adam, S. A. Sequence and characterization of cytoplasmic nuclear protein import factor p97. J. Cell Biol. 130, 265–274 (1995).
Goldfarb, D. S., Gariepy, J., Schoolnik, G. & Kornberg, R. D. Synthetic peptides as nuclear localization signals. Nature. 322, 641–644 (1986).
Fornerod, M., Ohno, M., Yoshida, M. & Mattaj, I. W. CRM1 is an export receptor for leucine-rich nuclear export signals. Cell. 90, 1051–1060 (1997).
Stade, K., Ford, C. S., Guthrie, C. & Weis, K. Exportin 1 (Crm1p) is an essential nuclear export factor. Cell. 90, 1041–1050 (1997).
Wen, W., Meinkoth, J. L., Tsien, R. Y. & Taylor, S. S. Identification of a signal for rapid export of proteins from the nucleus. Cell. 82, 463–473 (1995).
Gorlich, D. & Kutay, U. Transport between the cell nucleus and the cytoplasm. Annu. Rev. Cell Dev. Biol. 15, 607–660 (1999).
Vetter, I. R. et al. Structural view of the Ran-Importin beta interaction at 2.3 A resolution. Cell. 97, 635–646 (1999).
Chook, Y. M. & Blobel, G. Structure of the nuclear transport complex karyopherin-beta2-Ran x GppNHp. Nature. 399, 230–237 (1999).
Alber, F. et al. The molecular architecture of the nuclear pore complex. Nature. 450, 695–701 (2007).
Cowburn, D. & Rout, M. Improving the hole picture: towards a consensus on the mechanism of nuclear transport. Biochem. Soc. Trans. 51, 871–886 (2023).
Tai, L., Yin, G., Sun, F. & Zhu, Y. Cryo-electron microscopy reveals the structure of the nuclear pore complex. J. Mol. Biol. 435, 168051 (2023).
Akhtar, A. & Gasser, S. M. The nuclear envelope and transcriptional control. Nat. Rev. Genet. 8, 507–517 (2007).
Jakel, S. et al. Importins fulfil a dual function as nuclear import receptors and cytoplasmic chaperones for exposed basic domains. EMBO J. 21, 377–386 (2002).
Pennisi, E. Nuclear transport protein does double duty in mitosis. Science. 284, 1260–1261 (1999).
Chahine, M. N. & Pierce, G. N. Therapeutic targeting of nuclear protein import in pathological cell conditions. Pharmacol Rev. 61, 358–372 (2009).
Syed, Y. Y. Selinexor: First global approval. Drugs. 79, 1485–1494 (2019).
Maul, G. G. & Deaven, L. Quantitative determination of nuclear pore complexes in cycling cells with differing DNA content. J. Cell Biol. 73, 748–760 (1977).
Lin, D. H. et al. Architecture of the symmetric core of the nuclear pore. Science. 352, aaf1015 (2016).
Reichelt, R. et al. Correlation between structure and mass distribution of the nuclear pore complex and of distinct pore complex components. J. Cell Biol. 110, 883–894 (1990).
Hampoelz, B., Andres-Pons, A., Kastritis, P. & Beck, M. Structure and assembly of the nuclear pore complex. Annu. Rev. Biophys. 48, 515–536 (2019).
Lin, D. H. & Hoelz, A. The structure of the nuclear pore complex (an update). Annu. Rev. Biochem. 88, 725–783 (2019).
Akey, C. W. et al. Comprehensive structure and functional adaptations of the yeast nuclear pore complex. Cell. 185, 361–378.e325 (2022).
Kosinski, J. et al. Molecular architecture of the inner ring scaffold of the human nuclear pore complex. Science. 352, 363–365 (2016).
Li, Z. et al. Near-atomic structure of the inner ring of the Saccharomyces cerevisiae nuclear pore complex. Cell Res. 32, 437–450 (2022).
Zimmerli, C. E. et al. Nuclear pores dilate and constrict in cellulo. Science. 374, eabd9776 (2021).
Petrovic, S. et al. Architecture of the linker-scaffold in the nuclear pore. Science. 376, eabm9798 (2022).
Eisenhardt, N., Redolfi, J. & Antonin, W. Interaction of Nup53 with Ndc1 and Nup155 is required for nuclear pore complex assembly. J. Cell Sci. 127, 908–921 (2014).
Mauro, M. S. et al. Ndc1 drives nuclear pore complex assembly independent of membrane biogenesis to promote nuclear formation and growth. Elife. 11, e75513 (2022).
Huang, G. et al. Cryo-EM structure of the inner ring from the Xenopus laevis nuclear pore complex. Cell Res. 32, 451–460 (2022).
Mosalaganti, S. et al. AI-based structure prediction empowers integrative structural analysis of human nuclear pores. Science. 376, eabm9506 (2022).
Wong, R. W. Nuclear pore complex: From structural view to chemical tools. Chem Biol. 22, 1285–1287 (2015).
Fischer, J. et al. Linker Nups connect the nuclear pore complex inner ring with the outer ring and transport channel. Nat. Struct. Mol. Biol. 22, 774–781 (2015).
Ng, S. C., Guttler, T. & Gorlich, D. Recapitulation of selective nuclear import and export with a perfectly repeated 12mer GLFG peptide. Nat. Commun. 12, 4047 (2021).
Celetti, G. et al. The liquid state of FG-nucleoporins mimics permeability barrier properties of nuclear pore complexes. J. Cell Biol. 219, e201907157 (2020).
Stuwe, T. et al. Evidence for an evolutionary relationship between the large adaptor nucleoporin Nup192 and karyopherins. Proc. Natl. Acad. Sci. USA. 111, 2530–2535 (2014).
Onischenko, E. et al. Natively unfolded FG repeats stabilize the structure of the nuclear pore complex. Cell. 171, 904–917.e919 (2017).
Huang, G. et al. Cryo-EM structure of the nuclear ring from Xenopus laevis nuclear pore complex. Cell Res. 32, 349–358 (2022).
Tai, L. et al. 8 A structure of the outer rings of the Xenopus laevis nuclear pore complex obtained by cryo-EM and AI. Protein Cell. 13, 760–777 (2022).
Bley, C. J. et al. Architecture of the cytoplasmic face of the nuclear pore. Science. 376, eabm9129 (2022).
Fontana, P. et al. Structure of cytoplasmic ring of nuclear pore complex by integrative cryo-EM and AlphaFold. Science. 376, eabm9326 (2022).
Nordeen, S. A., Turman, D. L. & Schwartz, T. U. Yeast Nup84-Nup133 complex structure details flexibility and reveals conservation of the membrane anchoring ALPS motif. Nat Commun. 11, 6060 (2020).
Richards, L. et al. Nucleoporins facilitate ORC loading onto chromatin. Cell Rep. 41, 111590 (2022).
Kuhn, T. M. et al. Chromatin targeting of nuclear pore proteins induces chromatin decondensation. J. Cell Biol. 218, 2945–2961 (2019).
Zhu, X. et al. Structure of the cytoplasmic ring of the Xenopus laevis nuclear pore complex. Science. 376, eabl8280 (2022).
Upla, P. et al. Molecular architecture of the major membrane ring component of the nuclear pore complex. Structure. 25, 434–445 (2017).
Zhang, Y. et al. Molecular architecture of the luminal ring of the Xenopus laevis nuclear pore complex. Cell Res. 30, 532–540 (2020).
Kim, S. J. et al. Integrative structure and functional anatomy of a nuclear pore complex. Nature. 555, 475–482 (2018).
Stavru, F., Nautrup-Pedersen, G., Cordes, V. C. & Gorlich, D. Nuclear pore complex assembly and maintenance in POM121- and gp210-deficient cells. J. Cell Biol. 173, 477–483 (2006).
Mitchell, J. M. et al. Pom121 links two essential subcomplexes of the nuclear pore complex core to the membrane. J. Cell Biol. 191, 505–521 (2010).
Chen, S. et al. Association of RanGAP to nuclear pore complex component, RanBP2/Nup358, is required for pupal development in Drosophila. Cell Rep. 37, 110151 (2021).
Fernandez-Martinez, J. & Rout, M. P. One ring to rule them all? structural and functional diversity in the nuclear pore complex. Trends Biochem Sci. 46, 595–607 (2021).
Gray, S., Cao, W., Montpetit, B. & De La Cruz, E. M. The nucleoporin Gle1 activates DEAD-box protein 5 (Dbp5) by promoting ATP binding and accelerating rate limiting phosphate release. Nucleic Acids Res. 50, 3998–4011 (2022).
von Appen, A. et al. In situ structural analysis of the human nuclear pore complex. Nature. 526, 140–143 (2015).
Huang, G. et al. Structure of the cytoplasmic ring of the Xenopus laevis nuclear pore complex by cryo-electron microscopy single particle analysis. Cell Res. 30, 520–531 (2020).
Ritterhoff, T. et al. The RanBP2/RanGAP1*SUMO1/Ubc9 SUMO E3 ligase is a disassembly machine for Crm1-dependent nuclear export complexes. Nat Commun. 7, 11482 (2016).
Port, S. A. et al. Structural and functional characterization of CRM1-Nup214 interactions reveals multiple FG-binding sites involved in nuclear export. Cell Rep. 13, 690–702 (2015).
Mahajan, R. et al. A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell. 88, 97–107 (1997).
Yokoyama, N. et al. A giant nucleopore protein that binds Ran/TC4. Nature. 376, 184–188 (1995).
Partridge, J. R. & Schwartz, T. U. Crystallographic and biochemical analysis of the Ran-binding zinc finger domain. J. Mol. Biol. 391, 375–389 (2009).
Bernad, R., van der Velde, H., Fornerod, M. & Pickersgill, H. Nup358/RanBP2 attaches to the nuclear pore complex via association with Nup88 and Nup214/CAN and plays a supporting role in CRM1-mediated nuclear protein export. Mol. Cell Biol. 24, 2373–2384 (2004).
Lin, D. H. et al. Structural and functional analysis of mRNA export regulation by the nuclear pore complex. Nat Commun. 9, 2319 (2018).
Krull, S. et al. Nucleoporins as components of the nuclear pore complex core structure and Tpr as the architectural element of the nuclear basket. Mol. Biol. Cell. 15, 4261–4277 (2004).
Aksenova, V. et al. Nucleoporin TPR is an integral component of the TREX-2 mRNA export pathway. Nat Commun. 11, 4577 (2020).
Duheron, V. et al. Structural characterization of altered nucleoporin Nup153 expression in human cells by thin-section electron microscopy. Nucleus. 5, 601–612 (2014).
Mackay, D. R., Makise, M. & Ullman, K. S. Defects in nuclear pore assembly lead to activation of an Aurora B-mediated abscission checkpoint. J. Cell Biol. 191, 923–931 (2010).
Cibulka, J. et al. Assembly principle of a membrane-anchored nuclear pore basket scaffold. Sci. Adv. 8, eabl6863 (2022).
Souquet, B. et al. Nup133 is required for proper nuclear pore basket assembly and dynamics in embryonic stem cells. Cell Rep. 23, 2443–2454 (2018).
Meszaros, N. et al. Nuclear pore basket proteins are tethered to the nuclear envelope and can regulate membrane curvature. Dev Cell. 33, 285–298 (2015).
Petrovic, S. et al. Structure and function of the nuclear pore complex. Cold Spring Harb. Perspect. Biol. 14:a041264 (2022).
Gallardo, P., Salas-Pino, S. & Daga, R. R. A new role for the nuclear basket network. Microb Cell. 4, 423–425 (2017).
Pasha, T. et al. Karyopherin abnormalities in neurodegenerative proteinopathies. Brain. 144, 2915–2932 (2021).
Koepp, D. M. & Silver, P. A. A GTPase controlling nuclear trafficking: running the right way or walking RANdomly? Cell. 87, 1–4 (1996).
Stewart, M. Insights into the molecular mechanism of nuclear trafficking using nuclear transport factor 2 (NTF2). Cell Struct. Funct. 25, 217–225 (2000).
Goldfarb, D. S. et al. Importin alpha: a multipurpose nuclear-transport receptor. Trends Cell Biol. 14, 505–514 (2004).
Cagatay, T. & Chook, Y. M. Karyopherins in cancer. Curr Opin Cell Biol. 52, 30–42 (2018).
Kimura, M. et al. Extensive cargo identification reveals distinct biological roles of the 12 importin pathways. Elife. 6, e21184 (2017).
Baade, I. et al. The RNA-binding protein FUS is chaperoned and imported into the nucleus by a network of import receptors. J. Biol. Chem. 296, 100659 (2021).
Zhang, Z. C., Satterly, N., Fontoura, B. M. & Chook, Y. M. Evolutionary development of redundant nuclear localization signals in the mRNA export factor NXF1. Mol. Biol. Cell. 22, 4657–4668 (2011).
Gorlich, D., Henklein, P., Laskey, R. A. & Hartmann, E. A 41 amino acid motif in importin-alpha confers binding to importin-beta and hence transit into the nucleus. EMBO J. 15, 1810–1817 (1996).
Huber, J. et al. Snurportin1, an m3G-cap-specific nuclear import receptor with a novel domain structure. EMBO J. 17, 4114–4126 (1998).
Lott, K. & Cingolani, G. The importin beta binding domain as a master regulator of nucleocytoplasmic transport. Biochim. Biophys. Acta. 1813, 1578–1592 (2011).
Lee, S. J. et al. The structure of importin-beta bound to SREBP-2: nuclear import of a transcription factor. Science. 302, 1571–1575 (2003).
Ye, Z. et al. PCDH1 promotes progression of pancreatic ductal adenocarcinoma via activation of NF-kappaB signalling by interacting with KPNB1. Cell Death Dis. 13, 633 (2022).
Wang, S. M. et al. Nucleoporin POM121 signals TFEB-mediated autophagy via activation of SIGMAR1/sigma-1 receptor chaperone by pridopidine. Autophagy, 1-26, (2022).
Du, W. et al. KPNB1-mediated nuclear translocation of PD-L1 promotes non-small cell lung cancer cell proliferation via the Gas6/MerTK signaling pathway. Cell Death Differ. 28, 1284–1300 (2021).
Mahipal, A. & Malafa, M. Importins and exportins as therapeutic targets in cancer. Pharmacol Ther. 164, 135–143 (2016).
Twyffels, L., Gueydan, C. & Kruys, V. Transportin-1 and Transportin-2: protein nuclear import and beyond. FEBS Lett. 588, 1857–1868 (2014).
Nanaura, H. et al. C9orf72-derived arginine-rich poly-dipeptides impede phase modifiers. Nat Commun. 12, 5301 (2021).
Bourgeois, B. et al. Nonclassical nuclear localization signals mediate nuclear import of CIRBP. Proc. Natl. Acad. Sci. USA. 117, 8503–8514 (2020).
Niaki, A. G. et al. Loss of dynamic RNA interaction and aberrant phase separation induced by two distinct types of ALS/FTD-linked FUS mutations. Mol Cell. 77, 82–94.e84 (2020).
Guo, L., Fare, C. M. & Shorter, J. Therapeutic dissolution of aberrant phases by nuclear-import receptors. Trends Cell Biol. 29, 308–322 (2019).
Yang, T. J. et al. Tumor suppressor BAP1 nuclear import is governed by transportin-1. J. Cell Biol. 221, e202201094 (2022).
Hwang, W. Y. et al. Kap-beta2/Transportin mediates beta-catenin nuclear transport in Wnt signaling. Elife. 11, e70495 (2022).
Fernandez, J. et al. Transportin-1 binds to the HIV-1 capsid via a nuclear localization signal and triggers uncoating. Nat Microbiol. 4, 1840–1850 (2019).
Miyake, Y. et al. Influenza virus uses transportin 1 for vRNP debundling during cell entry. Nat Microbiol. 4, 578–586 (2019).
Shamsher, M. K., Ploski, J. & Radu, A. Karyopherin beta 2B participates in mRNA export from the nucleus. Proc. Natl. Acad. Sci. USA. 99, 14195–14199 (2002).
Kataoka, N., Bachorik, J. L. & Dreyfuss, G. Transportin-SR, a nuclear import receptor for SR proteins. J. Cell Biol. 145, 1145–1152 (1999).
Lai, M. C., Lin, R. I. & Tarn, W. Y. Transportin-SR2 mediates nuclear import of phosphorylated SR proteins. Proc. Natl. Acad. Sci. USA. 98, 10154–10159 (2001).
Maertens, G. N. et al. Structural basis for nuclear import of splicing factors by human Transportin 3. Proc. Natl. Acad. Sci. USA. 111, 2728–2733 (2014).
Janssens, J. et al. CRISPR/Cas9-induced mutagenesis corroborates the role of transportin-SR2 in HIV-1 nuclear import. Microbiol Spectr. 9, e0133621 (2021).
Rodriguez-Mora, S. et al. The mutation of Transportin 3 gene that causes limb girdle muscular dystrophy 1 F induces protection against HIV-1 infection. PLoS Pathog. 15, e1007958 (2019).
Kalab, P., Solc, P. & Motlik, J. The role of RanGTP gradient in vertebrate oocyte maturation. Results Probl. Cell Differ. 53, 235–267 (2011).
Azmi, A. S., Uddin, M. H. & Mohammad, R. M. The nuclear export protein XPO1 - from biology to targeted therapy. Nat. Rev. Clin. Oncol. 18, 152–169 (2021).
Sun, H. et al. A nuclear export signal is required for cGAS to sense cytosolic DNA. Cell Rep. 34, 108586 (2021).
Napolitano, G. et al. mTOR-dependent phosphorylation controls TFEB nuclear export. Nat Commun. 9, 3312 (2018).
Camus, V. et al. XPO1 in B cell hematological malignancies: from recurrent somatic mutations to targeted therapy. J. Hematol Oncol. 10, 47 (2017).
Kirli, K. et al. A deep proteomics perspective on CRM1-mediated nuclear export and nucleocytoplasmic partitioning. Elife. 4, e11466 (2015).
Thakar, K. et al. Identification of CRM1-dependent nuclear export cargos using quantitative mass spectrometry. Mol. Cell Proteomics. 12, 664–678 (2013).
Fung, H. Y., Fu, S. C. & Chook, Y. M. Nuclear export receptor CRM1 recognizes diverse conformations in nuclear export signals. Elife. 6, e23961 (2017).
Dong, X., Biswas, A. & Chook, Y. M. Structural basis for assembly and disassembly of the CRM1 nuclear export complex. Nat. Struct. Mol. Biol. 16, 558–560 (2009).
Koyama, M. & Matsuura, Y. Mechanistic insights from the recent structures of the CRM1 nuclear export complex and its disassembly intermediate. Biophys. (Nagoya-shi). 8, 145–150 (2012).
Okamura, M., Inose, H. & Masuda, S. RNA export through the NPC in eukaryotes. Genes (Basel). 6, 124–149 (2015).
ElMaghraby, M. F. et al. A heterochromatin-specific RNA export pathway facilitates piRNA production. Cell. 178, 964–979 e920 (2019).
Azizian, N. G. & Li, Y. XPO1-dependent nuclear export as a target for cancer therapy. J. Hematol Oncol. 13, 61 (2020).
Kutay, U. et al. Export of importin alpha from the nucleus is mediated by a specific nuclear transport factor. Cell. 90, 1061–1071 (1997).
Matsuura, Y. & Stewart, M. Structural basis for the assembly of a nuclear export complex. Nature. 432, 872–877 (2004).
Dong, Q. et al. Roles of the CSE1L-mediated nuclear import pathway in epigenetic silencing. Proc. Natl. Acad. Sci. USA. 115, E4013–E4022 (2018).
Zhang, X. et al. CSE1L, as a novel prognostic marker, promotes pancreatic cancer proliferation by regulating the AKT/mTOR signaling pathway. J. Cancer. 12, 2797–2806 (2021).
Nachmias, B. & Schimmer, A. D. Targeting nuclear import and export in hematological malignancies. Leukemia. 34, 2875–2886 (2020).
Wu, K., He, J., Pu, W. & Peng, Y. The role of exportin-5 in MicroRNA biogenesis and cancer. Genomics Proteomics Bioinformatics. 16, 120–126 (2018).
Lund, E. et al. Nuclear export of microRNA precursors. Science. 303, 95–98 (2004).
Okada, C. et al. A high-resolution structure of the pre-microRNA nuclear export machinery. Science. 326, 1275–1279 (2009).
Wild, T. et al. A protein inventory of human ribosome biogenesis reveals an essential function of exportin 5 in 60 S subunit export. PLoS Biol. 8, e1000522 (2010).
Yi, R., Qin, Y., Macara, I. G. & Cullen, B. R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016 (2003).
Calado, A. et al. Exportin-5-mediated nuclear export of eukaryotic elongation factor 1 A and tRNA. EMBO J. 21, 6216–6224 (2002).
Bohnsack, M. T. et al. Exp5 exports eEF1A via tRNA from nuclei and synergizes with other transport pathways to confine translation to the cytoplasm. EMBO J. 21, 6205–6215 (2002).
Li, J. et al. Pin1 impairs microRNA biogenesis by mediating conformation change of XPO5 in hepatocellular carcinoma. Cell Death Differ. 25, 1612–1624 (2018).
Shigeyasu, K. et al. Exportin-5 functions as an oncogene and a potential therapeutic target in colorectal cancer. Clin. Cancer Res. 23, 1312–1322 (2017).
Choi, H. K. et al. Programmed cell death 5 mediates HDAC3 decay to promote genotoxic stress response. Nat Commun. 6, 7390 (2015).
Grunwald, M., Lazzaretti, D. & Bono, F. Structural basis for the nuclear export activity of Importin13. EMBO J. 32, 899–913 (2013).
Tao, T. et al. Importin 13 regulates nuclear import of the glucocorticoid receptor in airway epithelial cells. Am. J. Respir Cell Mol. Biol. 35, 668–680 (2006).
Baade, I. et al. Extensive identification and in-depth validation of importin 13 cargoes. Mol. Cell Proteomics. 17, 1337–1353 (2018).
Grunwald, M. & Bono, F. Structure of Importin13-Ubc9 complex: nuclear import and release of a key regulator of sumoylation. EMBO J. 30, 427–438 (2011).
Zohud, B. A. et al. Importin 13 promotes NSCLC progression by mediating RFPL3 nuclear translocation and hTERT expression upregulation. Cell Death Dis. 11, 879 (2020).
Viswanathan, S. R. et al. Genome-scale analysis identifies paralog lethality as a vulnerability of chromosome 1p loss in cancer. Nat Genet. 50, 937–943 (2018).
Huang, X. F. et al. Mutation of IPO13 causes recessive ocular coloboma, microphthalmia, and cataract. Exp Mol Med. 50, 1–11 (2018).
Bhardwaj, A. & Das, S. SIRT6 deacetylates PKM2 to suppress its nuclear localization and oncogenic functions. Proc. Natl. Acad. Sci. USA. 113, E538–E547 (2016).
Chen, Y. S., Racca, J. D., Phillips, N. B. & Weiss, M. A. Inherited human sex reversal due to impaired nucleocytoplasmic trafficking of SRY defines a male transcriptional threshold. Proc. Natl. Acad. Sci. USA. 110, E3567–E3576 (2013).
Chen, L. et al. Exportin 4 depletion leads to nuclear accumulation of a subset of circular RNAs. Nat Commun. 13, 5769 (2022).
Aksu, M., Trakhanov, S. & Gorlich, D. Structure of the exportin Xpo4 in complex with RanGTP and the hypusine-containing translation factor eIF5A. Nat Commun. 7, 11952 (2016).
Metwally, M. et al. Copy number variation and expression of exportin-4 associates with severity of fibrosis in metabolic associated fatty liver disease. EBioMedicine. 70, 103521 (2021).
Mingot, J. M., Bohnsack, M. T., Jakle, U. & Gorlich, D. Exportin 7 defines a novel general nuclear export pathway. EMBO J. 23, 3227–3236 (2004).
Liang, P. et al. KPNB1, XPO7 and IPO8 mediate the translocation ofNF-kappaB/p65 into the nucleus. Traffic. 14, 1132–1143 (2013).
Aksu, M. et al. Xpo7 is a broad-spectrum exportin and a nuclear import receptor. J. Cell Biol. 217, 2329–2340 (2018).
Innes, A. J. et al. XPO7 is a tumor suppressor regulating p21(CIP1)-dependent senescence. Genes Dev. 35, 379–391 (2021).
Bayliss, R. et al. Structural basis for the interaction between NTF2 and nucleoporin FxFG repeats. EMBO J. 21, 2843–2853 (2002).
Clouse, K. N., Luo, M. J., Zhou, Z. & Reed, R. A Ran-independent pathway for export of spliced mRNA. Nat Cell Biol. 3, 97–99 (2001).
Ribbeck, K. et al. NTF2 mediates nuclear import of Ran. EMBO J. 17, 6587–6598 (1998).
Stewart, M., Kent, H. M. & McCoy, A. J. Structural basis for molecular recognition between nuclear transport factor 2 (NTF2) and the GDP-bound form of the Ras-family GTPase Ran. J. Mol. Biol. 277, 635–646 (1998).
Bullock, T. L., Clarkson, W. D., Kent, H. M. & Stewart, M. The 1.6 angstroms resolution crystal structure of nuclear transport factor 2 (NTF2). J. Mol. Biol. 260, 422–431 (1996).
Yamada, M., Tachibana, T., Imamoto, N. & Yoneda, Y. Nuclear transport factor p10/NTF2 functions as a Ran-GDP dissociation inhibitor (Ran-GDI). Curr. Biol. 8, 1339–1342 (1998).
Van Impe, K. et al. A new role for nuclear transport factor 2 and Ran: nuclear import of CapG. Traffic. 9, 695–707 (2008).
Lu, M. et al. A code for RanGDP binding in ankyrin repeats defines a nuclear import pathway. Cell. 157, 1130–1145 (2014).
Fribourg, S., Braun, I. C., Izaurralde, E. & Conti, E. Structural basis for the recognition of a nucleoporin FG repeat by the NTF2-like domain of the TAP/p15 mRNA nuclear export factor. Mol. Cell. 8, 645–656 (2001).
Aibara, S., Katahira, J., Valkov, E. & Stewart, M. The principal mRNA nuclear export factor NXF1:NXT1 forms a symmetric binding platform that facilitates export of retroviral CTE-RNA. Nucleic Acids Res. 43, 1883–1893 (2015).
Dimitrova, L. et al. Structural characterization of the chaetomium thermophilum TREX-2 complex and its interaction with the mRNA nuclear export factor Mex67:Mtr2. Structure. 23, 1246–1257 (2015).
Pasquinelli, A. E. et al. The constitutive transport element (CTE) of Mason-Pfizer monkey virus (MPMV) accesses a cellular mRNA export pathway. EMBO J. 16, 7500–7510 (1997).
Kerkow, D. E. et al. The structure of the NXF2/NXT1 heterodimeric complex reveals the combined specificity and versatility of the NTF2-like fold. J. Mol Biol. 415, 649–665 (2012).
Tan, W. et al. Identification and characterization of the mouse nuclear export factor (Nxf) family members. Nucleic Acids Res. 33, 3855–3865 (2005).
Sasaki, M. et al. Molecular cloning and functional characterization of mouse Nxf family gene products. Genomics. 85, 641–653 (2005).
Kose, S., Furuta, M. & Imamoto, N. Hikeshi, a nuclear import carrier for Hsp70s, protects cells from heat shock-induced nuclear damage. Cell. 149, 578–589 (2012).
Song, J. et al. Structural and functional analysis of Hikeshi, a new nuclear transport receptor of Hsp70s. Acta Crystallogr D Biol Crystallogr. 71, 473–483 (2015).
Imamoto, N. Heat stress-induced nuclear transport mediated by Hikeshi confers nuclear function of Hsp70s. Curr. Opin. Cell Biol. 52, 82–87 (2018).
Jullien, D., Gorlich, D., Laemmli, U. K. & Adachi, Y. Nuclear import of RPA in Xenopus egg extracts requires a novel protein XRIPalpha but not importin alpha. EMBO J. 18, 4348–4358 (1999).
Kosugi, S. et al. Design of peptide inhibitors for the importin alpha/beta nuclear import pathway by activity-based profiling. Chem. Biol. 15, 940–949 (2008).
Catimel, B. et al. Biophysical characterization of interactions involving importin-alpha during nuclear import. J. Biol. Chem. 276, 34189–34198 (2001).
Kobe, B. Autoinhibition by an internal nuclear localization signal revealed by the crystal structure of mammalian importin alpha. Nat Struct. Biol. 6, 388–397 (1999).
Pumroy, R. A. & Cingolani, G. Diversification of importin-alpha isoforms in cellular trafficking and disease states. Biochem. J. 466, 13–28 (2015).
Panayotis, N. et al. Importin alpha5 Regulates Anxiety through MeCP2 and Sphingosine Kinase 1. Cell Rep. 25, 3169–3179 e3167 (2018).
Sankhala, R. S. et al. Three-dimensional context rather than NLS amino acid sequence determines importin alpha subtype specificity for RCC1. Nat Commun. 8, 979 (2017).
Smith, K. M. et al. Structural basis for importin alpha 3 specificity of W proteins in Hendra and Nipah viruses. Nat Commun. 9, 3703 (2018).
Hazawa, M. et al. Disease-specific alteration of karyopherin-alpha subtype establishes feed-forward oncogenic signaling in head and neck squamous cell carcinoma. Oncogene. 39, 2212–2223 (2020).
Florio, T. J. et al. Differential recognition of canonical NF-kappaB dimers by Importin alpha3. Nat Commun. 13, 1207 (2022).
Delaforge, E. et al. Large-Scale Conformational Dynamics Control H5N1 Influenza Polymerase PB2 Binding to Importin alpha. J. Am. Chem. Soc. 137, 15122–15134 (2015).
Xu, W. et al. Ebola virus VP24 targets a unique NLS binding site on karyopherin alpha 5 to selectively compete with nuclear import of phosphorylated STAT1. Cell Host Microbe. 16, 187–200 (2014).
Kawashima, T. et al. Rac1 and a GTPase-activating protein, MgcRacGAP, are required for nuclear translocation of STAT transcription factors. J. Cell Biol. 175, 937–946 (2006).
Mitrousis, G., Olia, A. S., Walker-Kopp, N. & Cingolani, G. Molecular basis for the recognition of snurportin 1 by importin beta. J. Biol. Chem. 283, 7877–7884 (2008).
Paraskeva, E. et al. CRM1-mediated recycling of snurportin 1 to the cytoplasm. J. Cell Biol. 145, 255–264 (1999).
Dong, X. et al. Structural basis for leucine-rich nuclear export signal recognition by CRM1. Nature. 458, 1136–1141 (2009).
Dumont, J. & Verlhac, M. H. Using FRET to study RanGTP gradients in live mouse oocytes. Methods Mol. Biol. 957, 107–120 (2013).
Cole, C. N. & Hammell, C. M. Nucleocytoplasmic transport: driving and directing transport. Curr. Biol. 8, R368–R372 (1998).
Langer, K. et al. Insights into the function of the CRM1 cofactor RanBP3 from the structure of its Ran-binding domain. PLoS One. 6, e17011 (2011).
Bischoff, F. R. & Ponstingl, H. Catalysis of guanine nucleotide exchange on Ran by the mitotic regulator RCC1. Nature. 354, 80–82 (1991).
Cingolani, G., Petosa, C., Weis, K. & Muller, C. W. Structure of importin-beta bound to the IBB domain of importin-alpha. Nature. 399, 221–229 (1999).
Moroianu, J., Blobel, G. & Radu, A. Nuclear protein import: Ran-GTP dissociates the karyopherin alphabeta heterodimer by displacing alpha from an overlapping binding site on beta. Proc. Natl. Acad. Sci. USA. 93, 7059–7062 (1996).
Fu, S. C. et al. Correlation of CRM1-NES affinity with nuclear export activity. Mol. Biol. Cell. 29, 2037–2044 (2018).
Clarke, P. R. & Zhang, C. Spatial and temporal coordination of mitosis by Ran GTPase. Nat. Rev. Mol. Cell Biol. 9, 464–477 (2008).
Kalab, P., Weis, K. & Heald, R. Visualization of a Ran-GTP gradient in interphase and mitotic Xenopus egg extracts. Science. 295, 2452–2456 (2002).
Dasso, M. The Ran GTPase: theme and variations. Curr. Biol. 12, R502–R508 (2002).
Nemergut, M. E. et al. Chromatin docking and exchange activity enhancement of RCC1 by histones H2A and H2B. Science. 292, 1540–1543 (2001).
Makde, R. D., England, J. R., Yennawar, H. P. & Tan, S. Structure of RCC1 chromatin factor bound to the nucleosome core particle. Nature. 467, 562–566 (2010).
Chen, T. et al. N-terminal alpha-methylation of RCC1 is necessary for stable chromatin association and normal mitosis. Nat Cell Biol. 9, 596–603 (2007).
Klebe, C., Prinz, H., Wittinghofer, A. & Goody, R. S. The kinetic mechanism of Ran–nucleotide exchange catalyzed by RCC1. Biochemistry. 34, 12543–12552 (1995).
Renault, L., Kuhlmann, J., Henkel, A. & Wittinghofer, A. Structural basis for guanine nucleotide exchange on Ran by the regulator of chromosome condensation (RCC1). Cell. 105, 245–255 (2001).
Yamada, M., Mattaj, I. W. & Yoneda, Y. An ATP-dependent activity that releases RanGDP from NTF2. J. Biol. Chem. 279, 36228–36234 (2004).
Matunis, M. J., Coutavas, E. & Blobel, G. A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J. Cell Biol. 135, 1457–1470 (1996).
Bischoff, F. R. et al. RanGAP1 induces GTPase activity of nuclear Ras-related Ran. Proc. Natl. Acad. Sci. USA. 91, 2587–2591 (1994).
Seewald, M. J., Korner, C., Wittinghofer, A. & Vetter, I. R. RanGAP mediates GTP hydrolysis without an arginine finger. Nature. 415, 662–666 (2002).
Gong, Y. et al. Loss of RanGAP1 drives chromosome instability and rapid tumorigenesis of osteosarcoma. Dev Cell. 58, 192–210.e111 (2023).
Roscioli, E. et al. Importin-beta negatively regulates multiple aspects of mitosis including RANGAP1 recruitment to kinetochores. J. Cell Biol. 196, 435–450 (2012).
Li, Y. et al. Distinct RanBP1 nuclear export and cargo dissociation mechanisms between fungi and animals. Elife. 8, e41331 (2019).
Kehlenbach, R. H. et al. A role for RanBP1 in the release of CRM1 from the nuclear pore complex in a terminal step of nuclear export. J. Cell Biol. 145, 645–657 (1999).
Bischoff, F. R. et al. Co-activation of RanGTPase and inhibition of GTP dissociation by Ran-GTP binding protein RanBP1. EMBO J. 14, 705–715 (1995).
Kuhlmann, J., Macara, I. & Wittinghofer, A. Dynamic and equilibrium studies on the interaction of Ran with its effector, RanBP1. Biochemistry. 36, 12027–12035 (1997).
Richards, S. A., Lounsbury, K. M., Carey, K. L. & Macara, I. G. A nuclear export signal is essential for the cytosolic localization of the Ran binding protein, RanBP1. J Cell Biol. 134, 1157–1168 (1996).
Vetter, I. R. et al. Structure of a Ran-binding domain complexed with Ran bound to a GTP analogue: implications for nuclear transport. Nature. 398, 39–46 (1999).
Lindsay, M. E. et al. Ran-binding protein 3 is a cofactor for Crm1-mediated nuclear protein export. J. Cell Biol. 153, 1391–1402 (2001).
Koyama, M., Shirai, N. & Matsuura, Y. Structural insights into how Yrb2p accelerates the assembly of the Xpo1p nuclear export complex. Cell Rep. 9, 983–995 (2014).
Guttler, T. & Gorlich, D. Ran-dependent nuclear export mediators: a structural perspective. EMBO J. 30, 3457–3474 (2011).
Nemergut, M. E., Lindsay, M. E., Brownawell, A. M. & Macara, I. G. Ran-binding protein 3 links Crm1 to the Ran guanine nucleotide exchange factor. J. Biol. Chem. 277, 17385–17388 (2002).
Holzer, G. et al. The nucleoporin Nup50 activates the Ran guanine nucleotide exchange factor RCC1 to promote NPC assembly at the end of mitosis. EMBO J. 40, e108788 (2021).
Lindsay, M. E. et al. Npap60/Nup50 is a tri-stable switch that stimulates importin-alpha:beta-mediated nuclear protein import. Cell. 110, 349–360 (2002).
Werner, A., Flotho, A. & Melchior, F. The RanBP2/RanGAP1*SUMO1/Ubc9 complex is a multisubunit SUMO E3 ligase. Mol. Cell. 46, 287–298 (2012).
Pichler, A. & Melchior, F. Ubiquitin-related modifier SUMO1 and nucleocytoplasmic transport. Traffic. 3, 381–387 (2002).
Pichler, A. et al. The nucleoporin RanBP2 has SUMO1 E3 ligase activity. Cell. 108, 109–120 (2002).
Du, J. X., Bialkowska, A. B., McConnell, B. B. & Yang, V. W. SUMOylation regulates nuclear localization of Kruppel-like factor 5. J. Biol. Chem. 283, 31991–32002 (2008).
Terui, Y. et al. Dual role of sumoylation in the nuclear localization and transcriptional activation of NFAT1. J. Biol. Chem. 279, 28257–28265 (2004).
Ptak, C. & Wozniak, R. W. SUMO and nucleocytoplasmic transport. Adv. Exp. Med. Biol. 963, 111–126 (2017).
Rodriguez, M. S., Dargemont, C. & Hay, R. T. SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J. Biol. Chem. 276, 12654–12659 (2001).
Sternsdorf, T., Jensen, K., Reich, B. & Will, H. The nuclear dot protein sp100, characterization of domains necessary for dimerization, subcellular localization, and modification by small ubiquitin-like modifiers. J. Biol. Chem. 274, 12555–12566 (1999).
Panse, V. G. et al. A proteome-wide approach identifies sumoylated substrate proteins in yeast. J. Biol. Chem. 279, 41346–41351 (2004).
Melchior, F., Schergaut, M. & Pichler, A. SUMO: ligases, isopeptidases and nuclear pores. Trends Biochem Sci. 28, 612–618 (2003).
Di Cesare, E. et al. Aurora B SUMOylation is restricted to centromeres in early mitosis and requires RANBP2. Cells. 12, 372 (2023).
Panse, V. G. et al. Formation and nuclear export of preribosomes are functionally linked to the small-ubiquitin-related modifier pathway. Traffic. 7, 1311–1321 (2006).
Zhang, H., Saitoh, H. & Matunis, M. J. Enzymes of the SUMO modification pathway localize to filaments of the nuclear pore complex. Mol. Cell Biol. 22, 6498–6508 (2002).
Vertegaal, A. C. O. Signalling mechanisms and cellular functions of SUMO. Nat. Rev. Mol. Cell Biol. 23, 715–731 (2022).
Chang, H. M. & Yeh, E. T. H. SUMO: From bench to bedside. Physiol Rev. 100, 1599–1619 (2020).
Jamali, T., Jamali, Y., Mehrbod, M. & Mofrad, M. R. Nuclear pore complex: biochemistry and biophysics of nucleocytoplasmic transport in health and disease. Int Rev Cell Mol Biol. 287, 233–286 (2011).
Timney, B. L. et al. Simple rules for passive diffusion through the nuclear pore complex. J. Cell Biol. 215, 57–76 (2016).
Huang, K. & Szleifer, I. Modeling the nucleoporins that form the hairy pores. Biochem. Soc. Trans. 48, 1447–1461 (2020).
Donnaloja, F., Jacchetti, E., Soncini, M. & Raimondi, M. T. Mechanosensing at the nuclear envelope by nuclear pore complex stretch activation and its effect in physiology and pathology. Front Physiol. 10, 896 (2019).
Frey, S., Richter, R. P. & Gorlich, D. FG-rich repeats of nuclear pore proteins form a three-dimensional meshwork with hydrogel-like properties. Science. 314, 815–817 (2006).
Peters, R. Translocation through the nuclear pore complex: selectivity and speed by reduction-of-dimensionality. Traffic. 6, 421–427 (2005).
Lim, R. Y. et al. Flexible phenylalanine-glycine nucleoporins as entropic barriers to nucleocytoplasmic transport. Proc Natl. Acad. Sci. USA. 103, 9512–9517 (2006).
Milles, S. et al. Facilitated aggregation of FG nucleoporins under molecular crowding conditions. EMBO Rep. 14, 178–183 (2013).
Schmidt, H. B. & Gorlich, D. Nup98 FG domains from diverse species spontaneously phase-separate into particles with nuclear pore-like permselectivity. Elife. 4, e04251 (2015).
Lim, R. Y. et al. Nanomechanical basis of selective gating by the nuclear pore complex. Science. 318, 640–643 (2007).
Hoogenboom, B. W. et al. Physics of the nuclear pore complex: Theory, modeling and experiment. Phys Rep. 921, 1–53 (2021).
Frey, S. & Gorlich, D. FG/FxFG as well as GLFG repeats form a selective permeability barrier with self-healing properties. EMBO J. 28, 2554–2567 (2009).
Frey, S. & Gorlich, D. A saturated FG-repeat hydrogel can reproduce the permeability properties of nuclear pore complexes. Cell. 130, 512–523 (2007).
Ghavami, A., Veenhoff, L. M., van der Giessen, E. & Onck, P. R. Probing the disordered domain of the nuclear pore complex through coarse-grained molecular dynamics simulations. Biophys. J. 107, 1393–1402 (2014).
Ng, S. C. & Gorlich, D. A simple thermodynamic description of phase separation of Nup98 FG domains. Nat Commun. 13, 6172 (2022).
Sakiyama, Y., Mazur, A., Kapinos, L. E. & Lim, R. Y. Spatiotemporal dynamics of the nuclear pore complex transport barrier resolved by high-speed atomic force microscopy. Nat. Nanotechnol. 11, 719–723 (2016).
Rout, M. P., Aitchison, J. D., Magnasco, M. O. & Chait, B. T. Virtual gating and nuclear transport: the hole picture. Trends Cell Biol. 13, 622–628 (2003).
Zheng, T. & Zilman, A. Self-regulation of the nuclear pore complex enables clogging-free crowded transport. Proc. Natl. Acad. Sci. USA. 120, e2212874120 (2023).
Kalita, J. et al. Karyopherin enrichment and compensation fortifies the nuclear pore complex against nucleocytoplasmic leakage. J. Cell Biol. 221, e202108107 (2022).
Kapinos, L. E., Huang, B., Rencurel, C. & Lim, R. Y. H. Karyopherins regulate nuclear pore complex barrier and transport function. J. Cell Biol. 216, 3609–3624 (2017).
Kapinos, L. E. et al. Karyopherin-centric control of nuclear pores based on molecular occupancy and kinetic analysis of multivalent binding with FG nucleoporins. Biophys. J. 106, 1751–1762 (2014).
Lennon, K. M. et al. Characterizing binding interactions that are essential for selective transport through the nuclear pore complex. Int. J. Mol. Sci. 22, 10898 (2021).
Stanley, G. J., Fassati, A. & Hoogenboom, B. W. Biomechanics of the transport barrier in the nuclear pore complex. Semin Cell Dev Biol. 68, 42–51 (2017).
Yang, W. ‘Natively unfolded’ nucleoporins in nucleocytoplasmic transport: clustered or evenly distributed? Nucleus. 2, 10–16 (2011).
Moore, M. S. Npap60: a new player in nuclear protein import. Trends Cell Biol. 13, 61–64 (2003).
Milles, S. et al. Plasticity of an ultrafast interaction between nucleoporins and nuclear transport receptors. Cell. 163, 734–745 (2015).
Raveh, B. et al. Slide-and-exchange mechanism for rapid and selective transport through the nuclear pore complex. Proc. Natl. Acad. Sci. USA. 113, E2489–E2497 (2016).
Herrmann, M. et al. Near-field optical study of protein transport kinetics at a single nuclear pore. Nano Lett. 9, 3330–3336 (2009).
Yang, W., Gelles, J. & Musser, S. M. Imaging of single-molecule translocation through nuclear pore complexes. Proc. Natl. Acad. Sci. USA. 101, 12887–12892 (2004).
Nachury, M. V. & Weis, K. The direction of transport through the nuclear pore can be inverted. Proc. Natl. Acad. Sci. USA. 96, 9622–9627 (1999).
Huang, K. et al. Nanocompartmentalization of the nuclear pore lumen. Biophys. J. 118, 219–231 (2020).
Strawn, L. A. et al. Minimal nuclear pore complexes define FG repeat domains essential for transport. Nat Cell Biol. 6, 197–206 (2004).
Forler, D. et al. RanBP2/Nup358 provides a major binding site for NXF1-p15 dimers at the nuclear pore complex and functions in nuclear mRNA export. Mol Cell Biol. 24, 1155–1167 (2004).
Pyhtila, B. & Rexach, M. A gradient of affinity for the karyopherin Kap95p along the yeast nuclear pore complex. J. Biol. Chem. 278, 42699–42709 (2003).
Ben-Efraim, I. & Gerace, L. Gradient of increasing affinity of importin beta for nucleoporins along the pathway of nuclear import. J. Cell Biol. 152, 411–417 (2001).
Floer, M. & Blobel, G. Putative reaction intermediates in Crm1-mediated nuclear protein export. J. Biol. Chem. 274, 16279–16286 (1999).
Paci, G. et al. Molecular determinants of large cargo transport into the nucleus. Elife. 9, e55963 (2020).
Forbes, D. J., Travesa, A., Nord, M. S. & Bernis, C. Nuclear transport factors: global regulation of mitosis. Curr. Opin. Cell Biol. 35, 78–90 (2015).
Springhower, C. E., Rosen, M. K. & Chook, Y. M. Karyopherins and condensates. Curr. Opin. Cell Biol. 64, 112–123 (2020).
Sumner, M. C. & Brickner, J. The nuclear pore complex as a transcription regulator. Cold Spring Harb. Perspect. Biol. 14, a039438 (2022).
Endicott, S. J. & Brueckner, M. NUP98 sets the size-exclusion diffusion limit through the ciliary base. Curr. Biol. 28, 1643–1650 e1643 (2018).
Del Viso, F. et al. Congenital heart disease genetics uncovers context-dependent organization and function of nucleoporins at cilia. Dev Cell. 38, 478–492 (2016).
Jevtic, P. et al. The nucleoporin ELYS regulates nuclear size by controlling NPC number and nuclear import capacity. EMBO Rep. 20, e47283 (2019).
Brownlee, C. & Heald, R. Importin alpha partitioning to the plasma membrane regulates intracellular scaling. Cell. 176, 805–815.e808 (2019).
Bonner, A. M. & Hawley, R. S. A nucleoporin at the meiotic kinetochore. Dev Cell. 38, 447–448 (2016).
Hattersley, N. et al. MEL-28/ELYS and CENP-C coordinately control outer kinetochore assembly and meiotic chromosome-microtubule interactions. Curr. Biol. 32, 2563–2571.e2564 (2022).
Drutovic, D. et al. RanGTP and importin beta regulate meiosis I spindle assembly and function in mouse oocytes. EMBO J. 39, e101689 (2020).
Moore, W., Zhang, C. & Clarke, P. R. Targeting of RCC1 to chromosomes is required for proper mitotic spindle assembly in human cells. Curr. Biol. 12, 1442–1447 (2002).
Zhang, M. S., Arnaoutov, A. & Dasso, M. RanBP1 governs spindle assembly by defining mitotic Ran-GTP production. Dev Cell. 31, 393–404 (2014).
Cavazza, T. & Vernos, I. The RanGTP pathway: From nucleo-cytoplasmic transport to spindle assembly and beyond. Front Cell Dev Biol. 3, 82 (2015).
Chang, C. C. et al. Regulation of mitotic spindle assembly factor NuMA by importin-beta. J Cell Biol. 216, 3453–3462 (2017).
Ems-McClung, S. C. et al. RanGTP induces an effector gradient of XCTK2 and importin alpha/beta for spindle microtubule cross-linking. J. Cell Biol. 219, e201906045 (2020).
Chang, C. C. et al. Ran pathway-independent regulation of mitotic Golgi disassembly by Importin-alpha. Nat Commun. 10, 4307 (2019).
Eibes, S. et al. Nek9 phosphorylation defines a new role for TPX2 in Eg5-dependent centrosome separation before nuclear envelope breakdown. Curr. Biol. 28, 121–129.e124 (2018).
Lu, X. et al. Requirement for lamin B receptor and its regulation by importin beta and phosphorylation in nuclear envelope assembly during mitotic exit. J. Biol. Chem. 285, 33281–33293 (2010).
Wandke, C. & Kutay, U. Enclosing chromatin: reassembly of the nucleus after open mitosis. Cell. 152, 1222–1225 (2013).
Walther, T. C. et al. RanGTP mediates nuclear pore complex assembly. Nature. 424, 689–694 (2003).
Lau, C. K. et al. Transportin regulates major mitotic assembly events: from spindle to nuclear pore assembly. Mol. Biol. Cell. 20, 4043–4058 (2009).
Bernis, C. et al. Transportin acts to regulate mitotic assembly events by target binding rather than Ran sequestration. Mol. Biol. Cell. 25, 992–1009 (2014).
Arnaoutov, A. et al. Crm1 is a mitotic effector of Ran-GTP in somatic cells. Nat Cell Biol. 7, 626–632 (2005).
Bao, X. X. et al. Exportin Crm1 is repurposed as a docking protein to generate microtubule organizing centers at the nuclear pore. Elife. 7, e33465 (2018).
Budhu, A. S. & Wang, X. W. Loading and unloading: orchestrating centrosome duplication and spindle assembly by Ran/Crm1. Cell Cycle. 4, 1510–1514 (2005).
Wang, W., Budhu, A., Forgues, M. & Wang, X. W. Temporal and spatial control of nucleophosmin by the Ran-Crm1 complex in centrosome duplication. Nat Cell Biol. 7, 823–830 (2005).
Dawlaty, M. M. et al. Resolution of sister centromeres requires RanBP2-mediated SUMOylation of topoisomerase IIalpha. Cell. 133, 103–115 (2008).
Wozniak, R., Burke, B. & Doye, V. Nuclear transport and the mitotic apparatus: an evolving relationship. Cell Mol. Life Sci. 67, 2215–2230 (2010).
Yokoyama, H. et al. The nucleoporin MEL-28 promotes RanGTP-dependent gamma-tubulin recruitment and microtubule nucleation in mitotic spindle formation. Nat Commun. 5, 3270 (2014).
Mishra, R. K. et al. The Nup107-160 complex and gamma-TuRC regulate microtubule polymerization at kinetochores. Nat Cell Biol. 12, 164–169 (2010).
Zuccolo, M. et al. The human Nup107-160 nuclear pore subcomplex contributes to proper kinetochore functions. EMBO J. 26, 1853–1864 (2007).
Platani, M. et al. The Nup107-160 nucleoporin complex promotes mitotic events via control of the localization state of the chromosome passenger complex. Mol Biol Cell. 20, 5260–5275 (2009).
Hashizume, C., Nakano, H., Yoshida, K. & Wong, R. W. Characterization of the role of the tumor marker Nup88 in mitosis. Mol Cancer. 9, 119 (2010).
Jeganathan, K. B., Malureanu, L. & van Deursen, J. M. The Rae1-Nup98 complex prevents aneuploidy by inhibiting securin degradation. Nature. 438, 1036–1039 (2005).
Naylor, R. M., Jeganathan, K. B., Cao, X. & van Deursen, J. M. Nuclear pore protein NUP88 activates anaphase-promoting complex to promote aneuploidy. J. Clin. Invest. 126, 543–559 (2016).
Rodriguez-Bravo, V. et al. Nuclear pores protect genome integrity by assembling a premitotic and Mad1-dependent anaphase inhibitor. Cell. 156, 1017–1031 (2014).
Mossaid, I. & Fahrenkrog, B. Complex commingling: Nucleoporins and the spindle assembly checkpoint. Cells. 4, 706–725 (2015).
Jackman, M. et al. Cyclin B1-Cdk1 facilitates MAD1 release from the nuclear pore to ensure a robust spindle checkpoint. J. Cell Biol. 219, e201907082 (2020).
Schweizer, N. et al. Spindle assembly checkpoint robustness requires Tpr-mediated regulation of Mad1/Mad2 proteostasis. J. Cell Biol. 203, 883–893 (2013).
Dultz, E. et al. Systematic kinetic analysis of mitotic dis- and reassembly of the nuclear pore in living cells. J. Cell Biol. 180, 857–865 (2008).
Laurell, E. et al. Phosphorylation of Nup98 by multiple kinases is crucial for NPC disassembly during mitotic entry. Cell. 144, 539–550 (2011).
Linder, M. I. et al. Mitotic disassembly of nuclear pore complexes involves CDK1- and PLK1-mediated phosphorylation of key interconnecting nucleoporins. Dev Cell. 43, 141–156.e147 (2017).
Martino, L. et al. Channel nucleoporins recruit PLK-1 to nuclear pore complexes to direct nuclear envelope breakdown in C. elegans. Dev Cell. 43, 157–171 e157 (2017).
Chou, Y. Y. et al. Inherited nuclear pore substructures template post-mitotic pore assembly. Dev Cell. 56, 1786–1803.e1789 (2021).
Dultz, E., Huet, S. & Ellenberg, J. Formation of the nuclear envelope permeability barrier studied by sequential photoswitching and flux analysis. Biophys. J. 97, 1891–1897 (2009).
Bilokapic, S. & Schwartz, T. U. Structural and functional studies of the 252 kDa nucleoporin ELYS reveal distinct roles for its three tethered domains. Structure. 21, 572–580 (2013).
Hattersley, N. et al. A nucleoporin docks protein phosphatase 1 to direct meiotic chromosome segregation and nuclear assembly. Dev Cell. 38, 463–477 (2016).
Vagnarelli, P. et al. Repo-Man coordinates chromosomal reorganization with nuclear envelope reassembly during mitotic exit. Dev Cell. 21, 328–342 (2011).
Hawryluk-Gara, L. A. et al. Nup53 is required for nuclear envelope and nuclear pore complex assembly. Mol. Biol. Cell. 19, 1753–1762 (2008).
Walther, T. C. et al. The conserved Nup107-160 complex is critical for nuclear pore complex assembly. Cell. 113, 195–206 (2003).
Bodoor, K. et al. Sequential recruitment of NPC proteins to the nuclear periphery at the end of mitosis. J. Cell Sci. 112, 2253–2264 (1999).
Mansfeld, J. et al. The conserved transmembrane nucleoporin NDC1 is required for nuclear pore complex assembly in vertebrate cells. Mol. Cell. 22, 93–103 (2006).
Otsuka, S. et al. Postmitotic nuclear pore assembly proceeds by radial dilation of small membrane openings. Nat. Struct. Mol. Biol. 25, 21–28 (2018).
Kutay, U., Juhlen, R. & Antonin, W. Mitotic disassembly and reassembly of nuclear pore complexes. Trends Cell Biol. 31, 1019–1033 (2021).
Ren, H. et al. Postmitotic annulate lamellae assembly contributes to nuclear envelope reconstitution in daughter cells. J. Biol. Chem. 294, 10383–10391 (2019).
Hampoelz, B. et al. Pre-assembled nuclear pores insert into the nuclear envelope during early development. Cell. 166, 664–678 (2016).
Harel, A. & Forbes, D. J. Importin beta: conducting a much larger cellular symphony. Mol Cell. 16, 319–330 (2004).
Ganguly, A. et al. Importin-beta directly regulates the motor activity and turnover of a kinesin-4. Dev Cell. 44, 642–651.e645 (2018).
Chen, M. et al. The nuclear transport receptor Importin-11 is a tumor suppressor that maintains PTEN protein. J. Cell Biol. 216, 641–656 (2017).
Bernardes, N. E. et al. Structure of IMPORTIN-4 bound to the H3-H4-ASF1 histone-histone chaperone complex. Proc. Natl. Acad. Sci. USA. 119, e2207177119 (2022).
Pardal, A. J. & Bowman, A. J. A specific role for importin-5 and NASP in the import and nuclear hand-off of monomeric H3. Elife. 11, e81755 (2022).
Padavannil, A. et al. Importin-9 wraps around the H2A-H2B core to act as nuclear importer and histone chaperone. Elife. 8, e43630 (2019).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell. 149, 753–767 (2012).
Qamar, S. et al. FUS phase separation is modulated by a molecular chaperone and methylation of arginine cation-pi interactions. Cell. 173, 720–734 e715 (2018).
Hutten, S. et al. Nuclear import receptors directly bind to arginine-rich dipeptide repeat proteins and suppress their pathological interactions. Cell Rep. 33, 108538 (2020).
Guo, L. et al. Nuclear-import receptors reverse aberrant phase transitions of RNA-binding proteins with prion-like domains. Cell. 173, 677–692.e620 (2018).
Hofweber, M. et al. Phase separation of FUS is suppressed by its nuclear import receptor and arginine methylation. Cell. 173, 706–719.e713 (2018).
Yoshizawa, T. et al. Nuclear import receptor inhibits phase separation of FUS through binding to multiple sites. Cell. 173, 693–705.e622 (2018).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell. 174, 688–699.e616 (2018).
Ibanez de Opakua, A. et al. Molecular interactions of FG nucleoporin repeats at high resolution. Nat Chem. 14, 1278–1285 (2022).
Danilov, L. G. et al. The human NUP58 nucleoporin can form amyloids in vitro and in vivo. Biomedicines. 9, 1451 (2021).
Najbauer, E. E. et al. Atomic resolution dynamics of cohesive interactions in phase-separated Nup98 FG domains. Nat Commun. 13, 1494 (2022).
Davis, L. K., Ford, I. J., Saric, A. & Hoogenboom, B. W. Intrinsically disordered nuclear pore proteins show ideal-polymer morphologies and dynamics. Phys. Rev. E. 101, 022420 (2020).
Coyne, A. N. & Rothstein, J. D. Nuclear pore complexes—a doorway to neural injury in neurodegeneration. Nat Rev. Neurol. 18, 348–362 (2022).
Chou, C. C. et al. TDP-43 pathology disrupts nuclear pore complexes and nucleocytoplasmic transport in ALS/FTD. Nat Neurosci. 21, 228–239 (2018).
Diez, L. et al. Phosphorylation but not oligomerization drives the accumulation of tau with nucleoporin Nup98. Int. J. Mol. Sci. 23, 3495 (2022).
Eftekharzadeh, B. et al. Tau protein disrupts nucleocytoplasmic transport in Alzheimer’s disease. Neuron. 99, 925–940.e927 (2018).
Ahn, J. H. et al. Phase separation drives aberrant chromatin looping and cancer development. Nature. 595, 591–595 (2021).
Mendes, A. & Fahrenkrog, B. NUP214 in leukemia: It’s more than transport. Cells. 8, 76 (2019).
Rodriguez-Bravo, V. et al. Nuclear pores promote lethal prostate cancer by increasing POM121-driven E2F1, MYC, and AR nuclear import. Cell. 174, 1200–1215.e1220 (2018).
Raices, M. & D’Angelo, M. A. Nuclear pore complexes and regulation of gene expression. Curr. Opin. Cell Biol. 46, 26–32 (2017).
Pascual-Garcia, P. & Capelson, M. Nuclear pores in genome architecture and enhancer function. Curr. Opin. Cell Biol. 58, 126–133 (2019).
Ibarra, A. & Hetzer, M. W. Nuclear pore proteins and the control of genome functions. Genes Dev. 29, 337–349 (2015).
Kohler, A. & Hurt, E. Gene regulation by nucleoporins and links to cancer. Mol Cell. 38, 6–15 (2010).
Sun, J., Shi, Y. & Yildirim, E. The nuclear pore complex in cell type-specific chromatin structure and gene regulation. Trends Genet. 35, 579–588 (2019).
Pascual-Garcia, P. et al. Metazoan nuclear pores provide a scaffold for poised genes and mediate induced enhancer-promoter contacts. Mol. Cell. 66, 63–76 e66 (2017).
Blobel, G. Gene gating: a hypothesis. Proc. Natl. Acad. Sci. USA. 82, 8527–8529 (1985).
Jani, D. et al. Functional and structural characterization of the mammalian TREX-2 complex that links transcription with nuclear messenger RNA export. Nucleic Acids Res. 40, 4562–4573 (2012).
Kadota, S. et al. Nucleoporin 153 links nuclear pore complex to chromatin architecture by mediating CTCF and cohesin binding. Nat Commun. 11, 2606 (2020).
Aleman, J. R. et al. Correct dosage of X chromosome transcription is controlled by a nuclear pore component. Cell Rep. 35, 109236 (2021).
Ibarra, A. et al. Nucleoporin-mediated regulation of cell identity genes. Genes Dev. 30, 2253–2258 (2016).
Liu, Z. et al. Nucleoporin Seh1 interacts with Olig2/Brd7 to promote oligodendrocyte differentiation and myelination. Neuron. 102, 587–601 e587 (2019).
Kalverda, B., Pickersgill, H., Shloma, V. V. & Fornerod, M. Nucleoporins directly stimulate expression of developmental and cell-cycle genes inside the nucleoplasm. Cell. 140, 360–371 (2010).
Capelson, M. et al. Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. Cell. 140, 372–383 (2010).
Capitanio, J. S., Montpetit, B. & Wozniak, R. W. Human Nup98 regulates the localization and activity of DExH/D-box helicase DHX9. Elife. 6, e18825 (2017).
Chandra, B. et al. Phase separation mediates NUP98 fusion oncoprotein leukemic transformation. Cancer Discov. 12, 1152–1169 (2022).
Oka, M. et al. Chromatin-prebound Crm1 recruits Nup98-HoxA9 fusion to induce aberrant expression of Hox cluster genes. Elife. 5, e09540 (2016).
Jost, K. L., Bertulat, B. & Cardoso, M. C. Heterochromatin and gene positioning: inside, outside, any side? Chromosoma. 121, 555–563 (2012).
Gozalo, A. et al. Core components of the nuclear pore bind distinct states of chromatin and contribute to polycomb repression. Mol. Cell. 77, 67–81 e67 (2020).
Jacinto, F. V., Benner, C. & Hetzer, M. W. The nucleoporin Nup153 regulates embryonic stem cell pluripotency through gene silencing. Genes Dev. 29, 1224–1238 (2015).
Weiler, K. S. & Wakimoto, B. T. Heterochromatin and gene expression in Drosophila. Annu. Rev. Genet. 29, 577–605 (1995).
Gotta, M. et al. The clustering of telomeres and colocalization with Rap1, Sir3, and Sir4 proteins in wild-type Saccharomyces cerevisiae. J. Cell Biol. 134, 1349–1363 (1996).
Van de Vosse, D. W. et al. A role for the nucleoporin Nup170p in chromatin structure and gene silencing. Cell. 152, 969–983 (2013).
Galy, V. et al. Nuclear pore complexes in the organization of silent telomeric chromatin. Nature. 403, 108–112 (2000).
Lapetina, D. L., Ptak, C., Roesner, U. K. & Wozniak, R. W. Yeast silencing factor Sir4 and a subset of nucleoporins form a complex distinct from nuclear pore complexes. J. Cell Biol. 216, 3145–3159 (2017).
Feuerbach, F. et al. Nuclear architecture and spatial positioning help establish transcriptional states of telomeres in yeast. Nat Cell Biol. 4, 214–221 (2002).
Pombo, A. & Dillon, N. Three-dimensional genome architecture: players and mechanisms. Nat. Rev. Mol. Cell Biol. 16, 245–257 (2015).
Toda, T. et al. Nup153 interacts with Sox2 to enable bimodal gene regulation and maintenance of neural progenitor cells. Cell Stem Cell. 21, 618–634 e617 (2017).
Michmerhuizen, N. L., Klco, J. M. & Mullighan, C. G. Mechanistic insights and potential therapeutic approaches for NUP98-rearranged hematologic malignancies. Blood. 136, 2275–2289 (2020).
Xu, S. & Powers, M. A. Nuclear pore proteins and cancer. Semin Cell Dev Biol. 20, 620–630 (2009).
Martin, A. J. & Jans, D. A. Antivirals that target the host IMPalpha/beta1-virus interface. Biochem. Soc. Trans. 49, 281–295 (2021).
Uddin, M. H., Zonder, J. A. & Azmi, A. S. Exportin 1 inhibition as antiviral therapy. Drug Discov. Today. 25, 1775–1781 (2020).
Walker, J. S. et al. Recurrent XPO1 mutations alter pathogenesis of chronic lymphocytic leukemia. J. Hematol. Oncol. 14, 17 (2021).
Ishizawa, J. et al. Expression, function, and targeting of the nuclear exporter chromosome region maintenance 1 (CRM1) protein. Pharmacol Ther. 153, 25–35 (2015).
Gravina, G. L. et al. Nucleo-cytoplasmic transport as a therapeutic target of cancer. J. Hematol Oncol. 7, 85 (2014).
Sun, Q. et al. Inhibiting cancer cell hallmark features through nuclear export inhibition. Signal Transduct Target Ther. 1, 16010 (2016).
Turner, J. G. et al. Melphalan and exportin 1 inhibitors exert synergistic antitumor effects in preclinical models of human multiple myeloma. Cancer Res. 80, 5344–5354 (2020).
Newlands, E. S., Rustin, G. J. & Brampton, M. H. Phase I trial of elactocin. Br J Cancer. 74, 648–649 (1996).
Sun, Q. et al. Nuclear export inhibition through covalent conjugation and hydrolysis of Leptomycin B by CRM1. Proc Natl. Acad. Sci USA. 110, 1303–1308 (2013).
Sakakibara, K. et al. CBS9106 is a novel reversible oral CRM1 inhibitor with CRM1 degrading activity. Blood. 118, 3922–3931 (2011).
Tian, X. et al. Small-molecule antagonist targeting exportin-1 via rational structure-based discovery. J Med. Chem. 63, 3881–3895 (2020).
Etchin, J. et al. KPT-330 inhibitor of CRM1 (XPO1)-mediated nuclear export has selective anti-leukaemic activity in preclinical models of T-cell acute lymphoblastic leukaemia and acute myeloid leukaemia. Br. J. Haematol. 161, 117–127 (2013).
Chari, A. et al. Oral selinexor-dexamethasone for triple-class refractory multiple myeloma. N. Engl. J. Med. 381, 727–738 (2019).
Lassman, A. B. et al. A phase II study of the efficacy and safety of oral selinexor in recurrent glioblastoma. Clin. Cancer Res. 28, 452–460 (2022).
Garzon, R. et al. A phase 1 clinical trial of single-agent selinexor in acute myeloid leukemia. Blood. 129, 3165–3174 (2017).
Etchin, J. et al. KPT-8602, a second-generation inhibitor of XPO1-mediated nuclear export, is well tolerated and highly active against AML blasts and leukemia-initiating cells. Leukemia. 31, 143–150 (2017).
Janssen, J. et al. Addition of the nuclear export inhibitor selinexor to standard intensive treatment for elderly patients with acute myeloid leukemia and high risk myelodysplastic syndrome. Leukemia. 36, 2189–2195 (2022).
Huang, J. L. et al. Discovery of highly potent daphnane diterpenoids uncovers importin-beta1 as a druggable vulnerability in castration-resistant prostate cancer. J. Am. Chem. Soc. 144, 17522–17532 (2022).
Liu, T. et al. LncRNA DLEU1 contributes to colorectal cancer progression via activation of KPNA3. Mol. Cancer. 17, 118 (2018).
Zhu, C. et al. Cancer-associated exportin-6 upregulation inhibits the transcriptionally repressive and anticancer effects of nuclear profilin-1. Cell Rep. 34, 108749 (2021).
Volpon, L. et al. Importin 8 mediates m7G cap-sensitive nuclear import of the eukaryotic translation initiation factor eIF4E. Proc. Natl. Acad. Sci. USA. 113, 5263–5268 (2016).
Nataraj, N. B. et al. Nucleoporin-93 reveals a common feature of aggressive breast cancers: robust nucleocytoplasmic transport of transcription factors. Cell Rep. 38, 110418 (2022).
Boudhraa, Z., Carmona, E., Provencher, D. & Mes-Masson, A. M. Ran GTPase: A key player in tumor progression and metastasis. Front Cell Dev. Biol. 8, 345 (2020).
Camus, V. et al. Detection and prognostic value of recurrent exportin 1 mutations in tumor and cell-free circulating DNA of patients with classical Hodgkin lymphoma. Haematologica. 101, 1094–1101 (2016).
Miloudi, H. et al. XPO1(E571K) Mutation Modifies Exportin 1 Localisation and Interactome in B-cell Lymphoma. Cancers (Basel). 12, 2829 (2020).
Garcia-Santisteban, I. et al. A cellular reporter to evaluate CRM1 nuclear export activity: functional analysis of the cancer-related mutant E571K. Cell Mol. Life Sci. 73, 4685–4699 (2016).
Benzeno, S. et al. Identification of mutations that disrupt phosphorylation-dependent nuclear export of cyclin D1. Oncogene. 25, 6291–6303 (2006).
Mariano, A. R. et al. Cytoplasmic localization of NPM in myeloid leukemias is dictated by gain-of-function mutations that create a functional nuclear export signal. Oncogene. 25, 4376–4380 (2006).
Siddiqui, N. & Borden, K. L. mRNA export and cancer. Wiley Interdiscip. Rev RNA. 3, 13–25 (2012).
Viphakone, N. et al. Luzp4 defines a new mRNA export pathway in cancer cells. Nucleic Acids Res. 43, 2353–2366 (2015).
Dominguez-Sanchez, M. S. et al. Differential expression of THOC1 and ALY mRNP biogenesis/export factors in human cancers. BMC Cancer. 11, 77 (2011).
Carroll, M. & Borden, K. L. The oncogene eIF4E: using biochemical insights to target cancer. J. Interferon Cytokine Res. 33, 227–238 (2013).
Culjkovic, B. et al. The eIF4E RNA regulon promotes the Akt signaling pathway. J. Cell Biol. 181, 51–63 (2008).
Borden, K. L. B. The nuclear pore complex and mRNA export in cancer. Cancers (Basel). 13, 42 (2020).
Malone, C. F. et al. Selective modulation of a pan-essential protein as a therapeutic strategy in cancer. Cancer Discov. 11, 2282–2299 (2021).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature. 435, 834–838 (2005).
Melo, S. A. et al. A genetic defect in exportin-5 traps precursor microRNAs in the nucleus of cancer cells. Cancer Cell. 18, 303–315 (2010).
Sun, H. L. et al. ERK activation globally downregulates mirnas through phosphorylating exportin-5. Cancer Cell. 30, 723–736 (2016).
Gough, S. M., Slape, C. I. & Aplan, P. D. NUP98 gene fusions and hematopoietic malignancies: common themes and new biologic insights. Blood. 118, 6247–6257 (2011).
Struski, S. et al. NUP98 is rearranged in 3.8% of pediatric AML forming a clinical and molecular homogenous group with a poor prognosis. Leukemia. 31, 565–572 (2017).
Lam, D. H. & Aplan, P. D. NUP98 gene fusions in hematologic malignancies. Leukemia. 15, 1689–1695 (2001).
Xu, H. et al. NUP98 fusion proteins interact with the NSL and MLL1 complexes to drive leukemogenesis. Cancer Cell. 30, 863–878 (2016).
Heikamp, E. B. et al. The menin-MLL1 interaction is a molecular dependency in NUP98-rearranged AML. Blood. 139, 894–906 (2022).
Schmoellerl, J. et al. CDK6 is an essential direct target of NUP98 fusion proteins in acute myeloid leukemia. Blood. 136, 387–400 (2020).
Oka, M. et al. Chromatin-bound CRM1 recruits SET-Nup214 and NPM1c onto HOX clusters causing aberrant HOX expression in leukemia cells. Elife. 8, e46667 (2019).
Ben Abdelali, R. et al. SET-NUP214 is a recurrent gammadelta lineage-specific fusion transcript associated with corticosteroid/chemotherapy resistance in adult T-ALL. Blood. 123, 1860–1863 (2014).
Vanden Bempt, M. et al. Cooperative enhancer activation by TLX1 and STAT5 drives development of NUP214-ABL1/TLX1-positive T cell acute lymphoblastic leukemia. Cancer Cell. 34, 271–285.e277 (2018).
Gough, S. M. et al. NUP98-PHF23 is a chromatin-modifying oncoprotein that causes a wide array of leukemias sensitive to inhibition of PHD histone reader function. Cancer Discov. 4, 564–577 (2014).
Tziortzouda, P., Van Den Bosch, L. & Hirth, F. Triad of TDP43 control in neurodegeneration: autoregulation, localization and aggregation. Nat Rev. Neurosci. 22, 197–208 (2021).
Hutten, S. & Dormann, D. Nucleocytoplasmic transport defects in neurodegeneration—Cause or consequence? Semin Cell Dev Biol. 99, 151–162 (2020).
Lester, E. et al. Tau aggregates are RNA-protein assemblies that mislocalize multiple nuclear speckle components. Neuron. 109, 1675–1691.e1679 (2021).
Kavanagh, T., Halder, A. & Drummond, E. Tau interactome and RNA binding proteins in neurodegenerative diseases. Mol. Neurodegener. 17, 66 (2022).
Boeynaems, S., Bogaert, E., Van Damme, P. & Van Den Bosch, L. Inside out: the role of nucleocytoplasmic transport in ALS and FTLD. Acta Neuropathol. 132, 159–173 (2016).
Tang, X. et al. Divergence, convergence, and therapeutic implications: A cell biology perspective of C9ORF72-ALS/FTD. Mol. Neurodegener. 15, 34 (2020).
Doll, S. G. et al. Recognition of the TDP-43 nuclear localization signal by importin alpha1/beta. Cell Rep. 39, 111007 (2022).
Ferreira, P. A. The coming-of-age of nucleocytoplasmic transport in motor neuron disease and neurodegeneration. Cell Mol. Life Sci. 76, 2247–2273 (2019).
Gasset-Rosa, F. et al. Cytoplasmic TDP-43 de-mixing independent of stress granules drives inhibition of nuclear import, loss of nuclear TDP-43, and cell death. Neuron. 102, 339–357.e337 (2019).
Eftekharzadeh, B. et al. Tau protein disrupts nucleocytoplasmic transport in Alzheimer’s disease. Neuron. 101, 349 (2019).
Park, J. H. et al. Cytosolic calcium regulates cytoplasmic accumulation of TDP-43 through Calpain-A and Importin alpha3. Elife. 9, e60132 (2020).
Zhang, K. et al. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature. 525, 56–61 (2015).
Yamashita, T. et al. Calpain-dependent disruption of nucleo-cytoplasmic transport in ALS motor neurons. Sci. Rep. 7, 39994 (2017).
Nishimura, A. L. et al. Nuclear import impairment causes cytoplasmic trans-activation response DNA-binding protein accumulation and is associated with frontotemporal lobar degeneration. Brain. 133, 1763–1771 (2010).
Hock, E. M. et al. Hypertonic stress causes cytoplasmic translocation of neuronal, but not astrocytic, FUS due to impaired transportin function. Cell Rep. 24, 987–1000.e1007 (2018).
Goodman, L. D. et al. TNPO2 variants associate with human developmental delays, neurologic deficits, and dysmorphic features and alter TNPO2 activity in Drosophila. Am. J. Hum. Genet. 108, 1669–1691 (2021).
Fallini, C., Khalil, B., Smith, C. L. & Rossoll, W. Traffic jam at the nuclear pore: All roads lead to nucleocytoplasmic transport defects in ALS/FTD. Neurobiol Dis. 140, 104835 (2020).
Freibaum, B. D. et al. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature. 525, 129–133 (2015).
Lin, Y. C. et al. Interactions between ALS-linked FUS and nucleoporins are associated with defects in the nucleocytoplasmic transport pathway. Nat Neurosci. 24, 1077–1088 (2021).
Anderson, E. N. et al. Traumatic injury compromises nucleocytoplasmic transport and leads to TDP-43 pathology. Elife. 10, e67587 (2021).
Khalil, B. et al. Nuclear import receptors are recruited by FG-nucleoporins to rescue hallmarks of TDP-43 proteinopathy. Mol. Neurodegener. 17, 80 (2022).
Woerner, A. C. et al. Cytoplasmic protein aggregates interfere with nucleocytoplasmic transport of protein and RNA. Science. 351, 173–176 (2016).
Fujita, A. et al. Homozygous splicing mutation in NUP133 causes Galloway-Mowat syndrome. Ann Neurol. 84, 814–828 (2018).
Coyne, A. N. et al. Nuclear accumulation of CHMP7 initiates nuclear pore complex injury and subsequent TDP-43 dysfunction in sporadic and familial ALS. Sci. Transl. Med. 13, eabe1923 (2021).
Martins, F. et al. Nuclear envelope dysfunction and its contribution to the aging process. Aging Cell. 19, e13143 (2020).
Hayes, L. R. et al. C9orf72 arginine-rich dipeptide repeat proteins disrupt karyopherin-mediated nuclear import. Elife. 9, e51685 (2020).
Dubey, S. K., Maulding, K., Sung, H. & Lloyd, T. E. Nucleoporins are degraded via upregulation of ESCRT-III/Vps4 complex in Drosophila models of C9-ALS/FTD. Cell Rep. 40, 111379 (2022).
Zhang, S. et al. The ataxin-1 interactome reveals direct connection with multiple disrupted nuclear transport pathways. Nat Commun. 11, 3343 (2020).
Grima, J. C. et al. Mutant huntingtin disrupts the nuclear pore complex. Neuron. 94, 93–107.e106 (2017).
Coyne, A. N. et al. G(4)C(2) repeat RNA initiates a POM121-mediated reduction in specific nucleoporins in C9orf72 ALS/FTD. Neuron. 107, 1124–1140.e1111 (2020).
Gasset-Rosa, F. et al. Polyglutamine-expanded huntingtin exacerbates age-related disruption of nuclear integrity and nucleocytoplasmic transport. Neuron. 94, 48–57.e44 (2017).
Jovicic, A. et al. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nat Neurosci. 18, 1226–1229 (2015).
Zhang, K. et al. Stress granule assembly disrupts nucleocytoplasmic transport. Cell. 173, 958–971.e917 (2018).
Liu, S. et al. The XPO1 Inhibitor KPT-8602 Ameliorates Parkinson’s Disease by Inhibiting the NF-kappaB/NLRP3 Pathway. Front Pharmacol. 13, 847605 (2022).
Haines, J. D. et al. Nuclear export inhibitors avert progression in preclinical models of inflammatory demyelination. Nat Neurosci. 18, 511–520 (2015).
Shen, Q., Wang, Y. E. & Palazzo, A. F. Crosstalk between nucleocytoplasmic trafficking and the innate immune response to viral infection. J. Biol. Chem. 297, 100856 (2021).
Francis, A. C. & Melikyan, G. B. Single HIV-1 imaging reveals progression of infection through CA-dependent steps of docking at the nuclear pore, uncoating, and nuclear transport. Cell Host Microbe. 23, 536–548.e536 (2018).
Strunze, S. et al. Kinesin-1-mediated capsid disassembly and disruption of the nuclear pore complex promote virus infection. Cell Host Microbe. 10, 210–223 (2011).
Yamauchi, Y. Influenza A virus uncoating. Adv. Virus Res. 106, 1–38 (2020).
Whittaker, G. R. Virus nuclear import. Adv. Drug Deliv. Rev. 55, 733–747 (2003).
Wang, X. et al. Influenza A virus use of BinCARD1 to facilitate the binding of viral NP to importin alpha7 is counteracted by TBK1-p62 axis-mediated autophagy. Cell Mol. Immunol. 19, 1168–1184 (2022).
Flatt, J. W. & Greber, U. F. Viral mechanisms for docking and delivering at nuclear pore complexes. Semin Cell Dev Biol. 68, 59–71 (2017).
Shida, H. Role of nucleocytoplasmic RNA transport during the life cycle of retroviruses. Front Microbiol. 3, 179 (2012).
Mostafa-Hedeab, G. et al. Selinexor and COVID-19: The Neglected Warden. Front Pharmacol. 13, 884228 (2022).
Abid, H., Wu, J. F. & Abid, M. B. Risk for infections with selinexor in patients with relapsed/refractory multiple myeloma: a systematic review of clinical trials. Eur J Cancer. 154, 7–10 (2021).
Mogensen, T. H. IRF and STAT transcription factors - from basic biology to roles in infection, protective immunity, and primary immunodeficiencies. Front Immunol. 9, 3047 (2018).
Xia, H. et al. Evasion of type I interferon by SARS-CoV-2. Cell Rep. 33, 108234 (2020).
He, J. et al. Zika virus NS2A protein induces the degradation of KPNA2 (karyopherin subunit alpha 2) via chaperone-mediated autophagy. Autophagy. 16, 2238–2251 (2020).
Zhang, K. et al. Structural basis for influenza virus NS1 protein block of mRNA nuclear export. Nat Microbiol. 4, 1671–1679 (2019).
Park, N., Katikaneni, P., Skern, T. & Gustin, K. E. Differential targeting of nuclear pore complex proteins in poliovirus-infected cells. J. Virol. 82, 1647–1655 (2008).
Gustin, K. E. & Sarnow, P. Effects of poliovirus infection on nucleo-cytoplasmic trafficking and nuclear pore complex composition. EMBO J. 20, 240–249 (2001).
Bardina, M. V. et al. Mengovirus-induced rearrangement of the nuclear pore complex: hijacking cellular phosphorylation machinery. J. Virol. 83, 3150–3161 (2009).
Porter, F. W. & Palmenberg, A. C. Leader-induced phosphorylation of nucleoporins correlates with nuclear trafficking inhibition by cardioviruses. J. Virol. 83, 1941–1951 (2009).
Monette, A., Ajamian, L., Lopez-Lastra, M. & Mouland, A. J. Human immunodeficiency virus type 1 (HIV-1) induces the cytoplasmic retention of heterogeneous nuclear ribonucleoprotein A1 by disrupting nuclear import: implications for HIV-1 gene expression. J. Biol. Chem. 284, 31350–31362 (2009).
Simon, D. N. & Rout, M. P. Cancer and the nuclear pore complex. Adv. Exp. Med. Biol. 773, 285–307 (2014).
Feng, H. et al. Molecular mechanism underlying selective inhibition of mRNA nuclear export by herpesvirus protein ORF10. Proc. Natl. Acad. Sci. USA. 117, 26719–26727 (2020).
Gao, X. et al. Structural basis for Sarbecovirus ORF6 mediated blockage of nucleocytoplasmic transport. Nat Commun. 13, 4782 (2022).
Le Sage, V. & Mouland, A. J. Viral subversion of the nuclear pore complex. Viruses. 5, 2019–2042 (2013).
Liu, W. et al. Identification of a covalent importin-5 inhibitor, goyazensolide, from a collective synthesis of furanoheliangolides. ACS Cent Sci. 7, 954–962 (2021).
Lewin, J. et al. A phase 1b trial of selinexor, a first-in-class selective inhibitor of nuclear export (SINE), in combination with doxorubicin in patients with advanced soft tissue sarcomas (STS). Eur. J. Cancer. 144, 360–367 (2021).
Ho, J. et al. A phase 1 study of the safety, pharmacokinetics and pharmacodynamics of escalating doses followed by dose expansion of the selective inhibitor of nuclear export (SINE) selinexor in Asian patients with advanced or metastatic malignancies. Ther. Adv. Med. Oncol. 14, 17588359221087555 (2022).
Cornell, R. F. et al. A phase 1 clinical trial of oral eltanexor in patients with relapsed or refractory multiple myeloma. Am. J. Hematol. 97, E54–E58 (2022).
Martinez Sanchez, M. P. et al. A phase I trial of selinexor plus FLAG-Ida for the treatment of refractory/relapsed adult acute myeloid leukemia patients. Ann. Hematol. 100, 1497–1508 (2021).
Stephens, D. M. et al. Selinexor combined with ibrutinib demonstrates tolerability and safety in advanced B-cell malignancies: A phase I study. Clin. Cancer Res. 28, 3242–3247 (2022).
Westin, S. N. et al. Selinexor in combination with weekly paclitaxel in patients with metastatic solid tumors: Results of an open label, single-center, multi-arm phase 1b study with expansion phase in ovarian cancer. Gynecol Oncol. 168, 76–82 (2023).
Thein, K. Z. et al. Selinexor in combination with standard chemotherapy in patients with advanced or metastatic solid tumors. Exp. Hematol Oncol. 10, 59 (2021).
Grosicki, S. et al. Once-per-week selinexor, bortezomib, and dexamethasone versus twice-per-week bortezomib and dexamethasone in patients with multiple myeloma (BOSTON): a randomised, open-label, phase 3 trial. Lancet. 396, 1563–1573 (2020).
Lee, S. et al. Oral eltanexor treatment of patients with higher-risk myelodysplastic syndrome refractory to hypomethylating agents. J. Hematol Oncol. 15, 103 (2022).
Qiu, L. et al. Selinexor plus low-dose dexamethasone in Chinese patients with relapsed/refractory multiple myeloma previously treated with an immunomodulatory agent and a proteasome inhibitor (MARCH): a phase II, single-arm study. BMC Med. 20, 108 (2022).
Zijlstra, J. M. et al. The association between patient characteristics and the efficacy and safety of selinexor in diffuse large B-cell lymphoma in the SADAL study. Cancers (Basel). 14, 791 (2022).
Gounder, M. M. et al. Selinexor in advanced, metastatic dedifferentiated liposarcoma: a multinational, randomized, double-blind, placebo-controlled trial. J. Clin. Oncol. 40, 2479–2490 (2022).
Mateos, M. V. et al. Effect of prior treatments on selinexor, bortezomib, and dexamethasone in previously treated multiple myeloma. J. Hematol Oncol. 14, 59 (2021).
Acknowledgements
This work was funded by the National Natural Science Foundation of China (NSFC # 82273850).
Author information
Authors and Affiliations
Contributions
Q.S. wrote the draft. Y.Y., L.G., C.L., B.G., and D.J. provided relevant resources and revised the manuscript. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, Y., Guo, L., Chen, L. et al. Nuclear transport proteins: structure, function and disease relevance. Sig Transduct Target Ther 8, 425 (2023). https://doi.org/10.1038/s41392-023-01649-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-023-01649-4
- Springer Nature Limited
This article is cited by
-
Therapy-induced senescent tumor cell-derived extracellular vesicles promote colorectal cancer progression through SERPINE1-mediated NF-κB p65 nuclear translocation
Molecular Cancer (2024)
-
Selinexor targeting XPO1 promotes PEG3 nuclear accumulation and suppresses cholangiocarcinoma progression
Cancer Chemotherapy and Pharmacology (2024)
-
Space radiation damage rescued by inhibition of key spaceflight associated miRNAs
Nature Communications (2024)