Abstract
Background
Still 30–40% of pediatric acute myeloid leukemia (pedAML) patients relapse. Delineation of the transcriptomic profile of leukemic subpopulations could aid in a better understanding of molecular biology and provide novel biomarkers.
Methods
Using microarray profiling and quantitative PCR validation, transcript expression was measured in leukemic stem cells (LSC, n = 24) and leukemic blasts (L-blast, n = 25) from pedAML patients in comparison to hematopoietic stem cells (HSCs, n = 19) and control myeloblasts (C-blast, n = 20) sorted from healthy subjects. Gene set enrichment analysis was performed to identify relevant gene set enrichment signatures, and functional protein associations were identified by STRING analysis.
Results
Highly significantly overexpressed genes in LSC and L-blast were identified with a vast majority not studied in AML. CDKN1A, CFP, and CFD (LSC) and HOMER3, CTSA, and GADD45B (L-blast) represent potentially interesting biomarkers and therapeutic targets. Eleven LSC downregulated targets were identified that potentially qualify as tumor suppressor genes, with MYCT1, PBX1, and PTPRD of highest interest. Inflammatory and immune dysregulation appeared to be perturbed biological networks in LSC, whereas dysregulated metabolic profiles were observed in L-blast.
Conclusion
Our study illustrates the power of taking into account cell population heterogeneity and reveals novel targets eligible for functional evaluation and therapy in pedAML.
Impact
-
Novel transcriptional targets were discovered showing a significant differential expression in LSCs and blasts from pedAML patients compared to their normal counterparts from healthy controls.
-
Deregulated pathways, including immune and metabolic dysregulation, were addressed for the first time in children, offering a deeper understanding of the molecular pathogenesis.
-
These novel targets have the potential of acting as biomarkers for risk stratification, follow-up, and targeted therapy.
-
Multiple LSC-downregulated targets endow tumor suppressor roles in other cancer entities, and further investigation whether hypomethylating therapy could result into LSC eradication in pedAML is warranted.
Similar content being viewed by others
Introduction
Pediatric acute myeloid leukemia (pedAML) is a rare hematological disease that accounts for 20% of all pediatric leukemias.1 Cytogenetic risk stratification combined with response-guided therapeutic decisions considerably improved prognostication.2,3 Unfortunately, still 30–40% of the good responders experience relapse.2 During the past decade, ample evidence showed that relapse is associated with a high leukemic stem cell (LSC) load at diagnosis as well as LSC persistence during an apparent state of remission.4,5,6,7,8 However, relapse still occurs in a considerable part of LSClow pedAML patients identified by flow cytometry,7 emphasizing a high need for a more profound molecular and phenotypic characterization of LSC.
Hitherto, most pedAML gene expression profiles (GEPs) were established in bulk leukemic samples,9,10,11,12,13,14,15 not taking into account cellular heterogeneity, and thus fail to identify critical LSC-specific genes and pathways. In adult AML, by contrast, several LSC gene signatures have been developed these past years.16,17,18,19,20,21,22,23 Interestingly, the LSC17 signature by Ng et al.16 also held a prognostic value in pedAML.24,25 Moreover, it was used to develop a pediatric-specific LSC6 score able to identify high-risk (HR) pedAML patients.26 Unfortunately, this LSC signature also contains genes that are expressed in hematopoietic stem cells (HSCs) and lack the inclusion of downregulated targets.27 It was however shown that adding PCD17, an LSC-specific downregulated tumor suppressor gene (TSG), to the LSC17 score improved risk stratification in adult AML.28 Hence, the identification of novel differentially expressed genes (DEGs) in pedAML leukemic subpopulations could aid in providing novel biomarkers for risk stratification, follow-up, and targeted therapy.
Here, we describe LSC and leukemic blast (L-blast) targets in pedAML discovered by microarray profiling followed by quantitative PCR (qPCR) validation following a cancer vs. normal (CvN) approach. We reveal deregulated pathways that have not yet been addressed in children, and aid in a further understanding of the pedAML molecular biology.
Methods
Patients and controls
Bone marrow (BM) and/or peripheral blood (PB) from a total of 28 pedAML patients were selected based on cell availability (>50 × 106 after routine work-up) and CD34 positivity (≥1%). For 21/28 patients, both LSC and L-blast fractions were available, whereas for 3/28 and 4/28 patients only LSC or L-blast, respectively, could be evaluated. Demographics of patients used for LSC (n = 24) and L-blast (n = 25) expression evaluation are shown in Table 1. Details on treatment protocols and outcome definitions are described in Supplementary information. In addition, HSC and control myeloblasts (C-blast) were sorted from 19 and 20 healthy controls, respectively. Pediatric normal BM (NBM, n = 9, 12–18 years) was collected from posterior iliac crest during scoliosis surgery. Cord blood (CB, n = 11) was obtained after full-term delivery. All subjects gave their informed consent for inclusion before participation. The study was conducted in accordance with the Declaration of Helsinki, and the protocol was approved by the Ethics Committee of the University Hospital of Ghent (EC2015-1443 and EC2019-0294).
Cell sorting
Mononuclear cells were isolated by Ficoll density gradient (Axis-Shield), complemented by CD34 isolation if expression was <50% (CD34 MicroBead Kit, Miltenyi). Cell sorting was performed to isolate CD34+/CD38− and CD34+/CD38+ cells from patients and controls, defined as LSC and HSC, and L-blast and C-blast, respectively. Availability of both PB and BM yielded a total of 35 cell fractions in the LSC cohort (24 patients, 11 PB–BM couples) and in the L-blast cohort (25 patients, 10 PB–BM couples). The LSC compartment of seven patients could additionally be phenotypical distinguished based on differential expression of CD45RA (n = 3), CLL-1 (n = 2), CD123 (n = 1), or GPR56 (n = 1), yielding a total of 42 LSC fractions. Staining and sorting strategy are described in Supplementary information. Monoclonal antibodies are described in Supplementary Table S1. Sorted cells were collected in RPMI supplemented with 50% fetal calf serum, with a minimum post-sort purity >90%, spun down (10 min, 3000 r.p.m., 4 °C), and resuspended in 700 µL TRIzol for RNA extraction and cDNA synthesis as described in Supplementary information.
Microarray downstream analyses
Microarray profiling was performed on LSC and L-blast sorted from 3/24 and 4/25 pedAML patients (Supplementary Table S2), respectively, next to two HSC and three C-blast fractions. Technical details are described in Supplementary information. DEGs were identified based on |log 2 FC| > 2 and adjusted P values (adj. P) < 0.05. Functional networks between protein–protein associations encoded by DEGs were identified by STRING at a high evidence level.29 Only KEGG (Kyoto Encyclopedia of Genes and Genomes) annotated pathways were derived from significant pathway analysis. Gene set enrichment analysis (GSEA) was performed by combining independent omics datasets through pathway enrichment meta-analysis in order to obtain gene set enrichment signatures.30 Unsupervised organization and visualization of enriched gene sets was performed in Cytoscape.31 At least two clustered gene sets based on P < 0.05, false discovery rate (FDR) < 0.25 and Jaccard overlap combined index 0.375 were required for node visualization.
In addition, we re-analyzed the publicly available GSE 17054 microarray dataset,17 containing GEPs of nine LSC from adult AML patients and four HSC from healthy adults.
Real-time quantitative PCR
Ninety-four targets were selected for qPCR validation based on the magnitude and significance of differential expression and feasibility of primer development. Available cell fractions were subdivided into three cohorts based on sample availability, while respecting a balanced distribution of the genetic variations, and subsequently measured by qPCR in three steps. Cell fractions with only limited material available were reserved for the most significant DEGs and evaluated in the third step, hereby avoiding sample paucity to present as an issue. Analytical details of qPCR experiments are described in Supplementary information, primers are shown in Supplementary Table S3. Data analysis were performed according to state-of-the-art methods30 as follows: Ct values were corrected for primer efficiency and expressed as relative quantities. Normalized relative quantities (NRQs) were calculated against the expression of housekeeping genes GAPD, HPRT1, and TBP. To allow inter-run comparison, calibrated NRQ values (CNRQ) were generated by taking into account the expression of an inter-run calibrator. Target-specific cut-offs for overexpression were calculated based on the average expression plus two standard deviations measured in the respective normal counterparts.
Results
In order to identify novel targets in LSC and L-blast, a CvN approach integrating different experimental and analysis techniques was used (Fig. 1). Each set of DEGs was explored in the context of intertwining pathways and databases to define the top deregulated pathways, and GSEA to gain insight into (anti-) correlated biological pathways. Full lists of significant DEGs and (anti-)correlated gene sets identified between LSC vs. HSC and L-blasts vs. C-blasts are shown in Supplementary Tables S4–S13. Below, we discuss the top-ranked significant DEGs and (anti-)correlated gene sets for each comparison separately.
Step-by-step workflow illustrating the experimental and data processing steps pursued to filter out highly significant DEGs in leukemic subpopulations starting from a microarray profiling dataset. PedAML pediatric acute myeloid leukemia, CB cord blood, FC fold change, P P value, DEGs differentially expressed genes, LSC leukemic stem cell, HSC hematopoietic stem cell, L-blast leukemic blast, C-blast control blast.
Immune dysregulation separates LSC in pedAML from HSC
The expression of 295 targets significantly differed between LSC and HSC, with 83 targets up- and 212 downregulated in LSC. The top 50 ranked DEGs is shown in Fig. 2a. Well-known oncogenes were present among the highest LSC-overexpressed targets (e.g., CFD, ANXA2, NLRP3) next to genes with yet undefined roles in AML (e.g., PLIN2, CRIP1). The top 10 most downregulated genes also contained targets for which no role was yet reported in pedAML (e.g., ATP9A, PLCB4, COL5A1).
a Visualization of DEGs identified between LSC (n = 3) and HSC (n = 2). Genes are plotted in a volcano plot as log 2 FC values against −log10 adj. P values. Thresholds |log 2 FC| > 2 and −log 10 adj. P < 0.05 are shown as dashed lines. Genes selected as significantly different are highlighted in red. The top 50-ranked downregulated genes (left) and upregulated genes (right), annotated with gene symbols, are sorted by log 2 FC values. b Top 10 most correlated (top) and anti-correlated (bottom) gene sets identified through GSEA. The number of concordantly expressed (CE) genes/total genes and normalized enrichment scores (NESs) is shown for each gene set individually. FC fold change, DEGs differentially expressed genes, LSC leukemic stem cell, HSC hematopoietic stem cell.
Analysis of functional protein associations (STRING) showed upregulated pathways in LSC related to (breast) cancer, osteoclast differentiation, and apoptosis, whereas transcriptional misregulation, and Rap1/MAPK signaling were downregulated (Table 2). Myeloid cell activation networks involved in immune response were enriched in LSC (FDR 4.6e − 6), whereas networks related to stimuli responses, signaling, and cell communication were suppressed (FDR 5.9e − 3). From a total of 3650 signatures available through GSEA, 240 and 18 gene sets were significantly enriched or suppressed in LSC, respectively. The top 10 LSC-enriched gene sets involved LSC signatures, inflammatory response, apoptosis, immune suppression, and adipogenesis, whereas HSC signatures were anti-correlated (Fig. 2b). Unsupervised visualization (Cytoscape) identified LSC-upregulated pathways related to abnormal cell division, quiescence, autoimmune regulation, and environmental stress, while gene sets involved in normal quiescence and cell death signaling were downregulated (Supplementary Fig. S1A). Altogether, these data suggest that dysregulation of the immune system contributes to the leukemic transformation of stem cells in pedAML.
As pediatric and adult AML represent two genetically distinct diseases,32 we wondered if this heterogeneity is also reflected in the stem cell transcriptome. To this end, we re-analyzed the GSE 17054 microarray dataset from Majeti et al.17 and identified 486 significant DEGs between adult LSC and HSC (Supplementary Fig. S2A). Comparing the set of LSC-HSC DEGs identified in pediatric vs. adult AML revealed 71 common downregulated targets (Supplementary Fig. S2B), which was translated into mutual repressed pathways, for example, tight junction and MAPK signaling17 (Table 2). In sharp contrast, only three common LSC-upregulated transcripts were identified (TYROBP, CFP, and PTH2R).
Metabolic changes in pedAML L-blasts enhance proliferation compared to normal counterparts
We identified 157 and 332 significantly up- and downregulated targets in L-blast vs. C-blast. The top 50 upregulated transcripts is shown in Fig. 3a. Pathways enriched in L-blasts involved cancer transcriptional misregulation, FoxO signaling, and cytokine–cytokine receptor interaction (Table 2).
a Visualization of genes identified to be upregulated in L-blast (n = 4) compared to C-blast (n = 3). Genes are plotted in a volcano plot as log 2 FC values against −log10 adj. P values. Thresholds |log 2 FC| > 2 and −log 10 adj. P < 0.05 are shown as dashed lines, DEGs are highlighted in red. The top 50-ranked upregulated genes, sorted by log 2 FC values, are annotated with gene symbols on the right. b Top 10 most correlated (top) and anti-correlated (bottom) gene sets identified through GSEA, based on the DEGs between L-blast and C-blast. The number of concordantly expressed (CE) genes/total genes and normalized enrichment scores (NESs) is shown for each gene set individually. FC fold change, DEGs differentially expressed genes, L-blast leukemic blast, C-blast control blast.
Functional protein network analysis (STRING) illustrated a significant enrichment in L-blasts of stimuli responses (FDR 6.8e − 11) and metabolic processes (FDR 1.4e − 05). GSEA identified 163 enriched and 23 suppressed gene sets in L-blast vs. C-blast. The top-ranked adipogenesis gene set correlates with metabolic dysregulation, whereas increased EGF signaling and decreased stemness signatures relate to high proliferation (Fig. 3b). Among others, upregulated cancer and EGFR signaling, and downregulated death signaling, were confirmed by unsupervised clustering (Supplementary Fig. S1B).
Interestingly, the top-ranked (anti-)correlated gene sets identified in LSC (Fig. 2b) and L-blast (Fig. 3b) partially overlapped, and also enriched and suppressed pathways were recurrent (Table 2). Therefore, we sought similarities in the DEGs identified in LSC and L-blast. From the 83 and 157 significantly upregulated genes in LSC and L-blasts, respectively, 49 genes appeared to be common (Supplementary Fig. S3A). On the other hand, 134 targets were mutually downregulated from a total set of 212 and 332 transcripts, respectively (Supplementary Fig. S3B). Taken together, we conclude that LSC and L-blast share pan-leukemic molecular aberrancies compared to their normal counterparts.
Novel candidate targets in pedAML leukemic subpopulations validated by qPCR
A selection of the top-ranked up- and downregulated targets in LSC vs. HSC and upregulated targets in L-blast vs. C-blast, as identified by microarray analysis, were validated by qPCR (29/83, 23/212, and 42/157, respectively). A total of 94 targets were evaluated according to a three-step exclusion strategy, allowing the most significant DEGs to be evaluated in the highest number of cell fractions. Patients were dichotomized as high and low for the highest differentially expressed targets. Per target, overexpression was correlated to cytogenetic and molecular markers, and if patients were included in the NOPHO-DBH AML2012 protocol, correlated with event-free survival (EFS).
First, differential expression was confirmed at a significant level (P < 0.05) for 24/29 LSC upregulated targets (Supplementary Fig. S4A). Moreover, differential expression was significant at P < 0.01 with concomitant low HSC expression for 12/24 targets. Expression of these 12 targets was further evaluated using additional LSC and HSC fractions (Supplementary Fig. S4B). Too low LSC expression, or too high HSC expression, led to the exclusion of 4/12 targets. Finally, expression of PLIN2, CFD, EMP1, DUSP5, ANXA2, CRIP1, CDKN1A, and CFP was evaluated in all fractions and shown to be highly significantly overexpressed in LSC (n = 42) compared to HSC (n = 20) (Fig. 4a). Overexpression of these eight targets was observed in 38–67% of the patients, with CDKN1A, CRIP1, CFP, and CFD overexpressed in more than half of the patients (Table 3). Expression was averagely 4- to 12-fold higher in LSC compared to HSC, with CFD and CRIP1 the most upregulated targets. CFP overexpression significantly correlated to FLT3-ITD mutations, for example, 46% in CFP-high (n = 14) vs. 10% in CFP-low (n = 10) patients (P = 0.043). High ANXA2 levels beneficially impacted EFS at a borderline significant level (P = 0.061, 4 ANXA2-high vs. 10 ANXA2-low patients, Supplementary Fig. S5A).
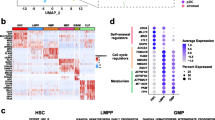
Targets with significant differential expression in LSC compared to HSC, shown by microarray analysis, were evaluated by qPCR analysis. Eight upregulated targets (a) and eleven downregulated targets (b) were evaluated in all available LSC (n = 42) and HSC (n = 20) fractions. Mean values are shown by horizontal lines, error bars indicate the 95% confidence interval of the mean, and the dotted line indicates the cut-off used to define overexpression, with the respective numbers of patients classified as high or low indicated above or below the line, respectively. P values (one-tailed) were calculated by the Mann–Whitney U test, and *, **, ***, or **** indicate the level of significance (P < 0.05, P < 0.01, P < 0.001 and P < 0.0001, respectively). LSC leukemic stem cell, HSC hematopoietic stem cell, DEG differentially expressed gene, CNRQ calibrated normalized relative quantities.
Second, significant LSC downregulation was confirmed for 21/23 targets (Supplementary Fig. S4C, P < 0.05). Among these, 15/21 targets were even downregulated at P < 0.01, with virtual absent expression in LSC. Including more sample fractions led to the exclusion of 4/15 targets due to too low differential expression levels (Supplementary Fig. S4D). Evaluation of all fractions illustrated that MECOM, HLF, PLCB4, PLAG1, ATP9A, PTPRD, COL5A1, BEX2, DSG2, MYCT1, and PBX1 transcripts are highly significantly repressed in LSC compared to HSC (P < 0.0001, Fig. 4b). These targets appeared to be suppressed in 75–100% of all patients. BEX2 downregulation was significantly anticorrelated to KMT2A-rearrangements (P < 0.01), as previously demonstrated using cell lines.33 On the other hand, the previously reported association between PLAG1 and inv16 (p13q22), or between MYCT1 and FAB classifications M1/M5/M6, was not confirmed (P > 0.05).34,35 Interestingly, 9/11 pedAML LSC-downregulated targets were also significantly suppressed in adult LSC (Supplementary Fig. S1B). Furthermore, PBX1, MYCT1, HLF, ATP9A, and PLCB4 appeared to be also suppressed in other pediatric malignancies, suggesting a possible tumor suppressor role (Supplementary Fig. S6).
Third, qPCR confirmed a significant upregulation in L-blast vs. C-blast for 16/42 targets (Supplementary Fig. S4E, P < 0.05). Further analysis using more samples showed that expression was too low in L-blasts, or too high in C-blasts, for 7/16 targets (Supplementary Fig. S4F). Evaluation of the remaining nine targets in all samples illustrated that DUSP6, HOMER3, ANXA2P1, CTSA, RHBDF2, EMP1, GADD45B, TYROBP, and PNP transcripts were highly significantly overexpressed in L-blasts (n = 36) vs. C-blasts (n = 19) (Fig. 5). Five out of nine targets (RHBDF2, HOMER3, ANXA2P1, GADD45B, and TYROBP) were overexpressed in more than two-thirds of the patients, with HOMER3 showing the highest differential expression (Table 3). Interestingly, HOMER3-high cases (n = 21) showed significantly less inv16 (p13q22) (P = 0.014) and FAB M4 (P = 0.031), compared to HOMER3-low pedAML (n = 4). DUSP6, overexpressed in one-third of the patients, was previously shown to be significantly associated with FLT3-ITD in adult AML,36 which we could not confirm in a pediatric setting (P = 0.49). PNP-high patients showed a significant lower EFS (Supplementary Fig. S5B), which was confirmed by Cox log-rank univariate analysis (hazard ratio 9.24, P = 0.04), but did not remain significant in multivariate analysis.
Targets with significant differential expression in L-blast compared to C-blast, shown by microarray analysis, were evaluated by qPCR analysis. Nine upregulated targets were evaluated in all available L-blast (n = 35) and C-blast (n = 19) fractions. Mean values are shown by horizontal lines, error bars indicate the 95% confidence interval of the mean, and the dotted line indicates the cut-off used to define overexpression, with the respective numbers of patients classified as high or low indicated above or below the line, respectively. P values (one-tailed) were calculated by the Mann–Whitney U test, and *, **, ***, or **** indicate the level of significance (P < 0.05, P < 0.01, P < 0.001 and P < 0.0001, respectively). L-blast leukemic blast, C-blast control blast, DEG differentially expressed gene, CNRQ calibrated normalized relative quantities.
Discussion
We here describe a novel set of differentially expressed targets in LSC and L-blast of pedAML patients identified based on a CvN approach. Moreover, we reveal previously unexplored deregulated pathways in these leukemic subpopulations of children with AML.
Eight targets were found highly significantly overexpressed in LSC compared to HSC. CDKN1A, ANXA2, EMP1, and CFD were previously linked to leukemogenesis, whereas the role of PLIN2, DUSP5, CRIP1, and CFP in AML remains elusive. CDKN1A, CRIP1, CFP, and CFD were found to be most frequently overexpressed (>50% of the patients), with CRIP1 and CFD showing the highest differential expression. CDKN1A might represent an interesting target for LSC eradication, since elevated expression was reported to maintain LSC activity,37,38 and CDKN1A knockdown indirectly reversed stem cell quiescence.39 Overexpression of CFP and CFD in LSC of pedAML patients suggest a disturbed complement pathway regulation. CFP overexpression was significantly associated with FLT3-ITD mutations in pedAML (P = 0.043) and concomitantly overexpressed in adult AML, suggesting a role as age-independent LSC marker in (high-risk) AML. CFD expression was previously linked to poor outcome in adult AML,40 and its prognostic value in children awaits validation. Last, we found that patients with high ANXA2 LSC expression (38%, 4.8-fold higher expression than HSC) show a trend towards prolonged EFS (P = 0.061). This finding is in agreement with a previously reported favorable prognostic effect of ANXA2 in bulk pedAML cells.41 Hence, ANXA2 could hold promise as a prognostic biomarker in pedAML. Although only the top-ranked DEGs were validated by qPCR in our study, microarray analysis additionally revealed interesting targets for flow cytometric validation. Our data confirm upregulated CD96 and CLECL12 expression in pedAML LSC compared to HSC.42,43,44 A potential role for targeting CD180 and CD68 in LSC, or their qualification as follow-up marker, deserves further attention.
Suppression of LSC-specific downregulated targets was highly consistent across the different genetic subgroups (75–100% of all patients). Since several of these targets endow tumor suppressor roles in other cancer entities and are inactivated upon promoter hypermethylation, further investigation whether hypomethylating therapy could result into LSC eradication in pedAML is warranted. MYCT1 was already identified as a TSG in AML,35 and deserves the highest attention since expression levels are 100-fold lower in LSC compared to HSC. PBX1 was previously reported to act as both an oncogene and TSG.45 Suppressed PTPRD levels in LSC is in agreement with a previous report on PTPRD suppression in pedAML bulk leukemic cells.46
Transcriptional misregulation in cancer, osteoclastogenesis, and tight junction pathways were dysregulated in LSC. The “cancer transcriptional misregulation” pathway is associated with myeloid leukemogenesis and held responsible for tumorigenic epigenetic abnormalities.47,48 Distortion of osteoclastogenesis and tight junction pathways might provide LSC an advantage over HSC during homing towards the endosteal–vascular niche. The observed immune dysregulation, separating LSC from HSC, strokes with a previous statement that multiple inflammatory signaling pathways are involved in the generation of pre-LSCs.49
L-blast upregulated targets were overexpressed in a larger portion of patients compared to LSC upregulated targets (36–88%, median 72% vs. 38–67%, median 50%, respectively). Among these, only DUSP6 and HOMER3 were previously addressed in AML. DUSP6 is an important cellular signaling regulator overexpressed in AML.36 HOMER3 relates to the occurrence and development of AML,50 and increased levels were significantly associated with favorable cytogenetics in adult AML.51 Since HOMER3 also showed the highest differential expression compared to C-blasts, targeting could be of therapeutic value. CTSA also represent an interesting target, since several other cathepsins were shown to have a diagnostic, prognostic, or therapeutic significance in AML.52,53,54,55 GADD45B is known to be involved in negative growth control during myeloid differentiation,56 and suggested to play a role in the tumorigenesis of colorectal carcinoma.57 Finally, high PNP expression was significantly associated with a worse EFS. Studies with PNP inhibitors in relapsed and refractory leukemias are ongoing,58 and investigation of their applicability in pedAML might be worthwhile.
Deregulated pathways identified in L-blast compared to their normal counterparts suggest that disturbed regulation of cell cycling, apoptosis, glycolysis/gluconeogenesis, and oxidative stress resistance promote the maintenance and proliferation of blasts in a leukemic setting. Indeed, adapting to hypoxic conditions and switching from oxidative phosphorylation towards glycolysis was shown to correlate with an aggressive disease course in solid cancers.59 Gluconeogenesis blocking, previously proposed as general antitumor therapeutic strategy, should be further explored in pedAML.60
EMP1 was the only target highly significantly upregulated in LSC and L-blast (38% and 68% of patients, respectively, P < 0.0001). Although Ng et al.16 did not elaborate on its role, EMP1 was included in the LSC17 score,25 but not retained in the pedAML-specific LSC6-score.26 However, based on the here observed CD38-independent overexpression, and the previously reported in vitro targetability of EMP1 in B-ALL,61 its role as a therapeutic target in pedAML should be further explored.
We detected a high molecular heterogeneity between pediatric and adult AML at the stem cell level. LSC populations from both entities shared 71 suppressed transcripts, but only three mutual upregulated targets (TYROBP, CFP, and PTH2R) were identified. TYROBP and CFP have not been functionally associated with AML, and their role in LSC transformation should be further explored. PTH2R, on the other hand, is known to be upregulated in AML and MDS,62 including adult AML LSC.63 Further research is warranted to evaluate whether these three targets could serve as pan-LSC targets, irrespective of the age of onset. Although these findings further underline the distinct biology between pediatric and adult AML,32 it should be taken into account that, due to the small number of patients evaluated, the genetic subtypes might also play a role besides the age of the patients.
Our investigation has some limitations, including its design as a single-center study recruiting patients from different clinical trials, and the lack of protein expression data. We acknowledge that extensive in vitro and in vivo validation of several of the identified targets is needed to fully unlock the potential of the presented dataset. Nevertheless, we here provide a repository to the pediatric acute leukemia community allowing to fulfill the high need for alternative therapeutic strategies in pediatric AML. The limited number of patient samples asks for data to be interpreted with caution within the framework of generalizability. Although promising, these data need confirmation in larger, preferentially multicenter trials, as survival analyses were performed in a limited number patients. It is important to acknowledge that the pedAML cohort included one secondary AML evolved from juvenile myelomonocytic leukemia, one acute promyelocytic leukemia, and two relapsed patients, all excluded from outcome analysis.
In conclusion, we here report a unique set of LSC and L-blast-specific overexpressed genes in pedAML. Most targets have not been studied in AML, and are involved in immune regulation, apoptosis, adhesion, or intracellular signaling, making them attractive candidates for functional studies, refining signatures, and targeted therapy. Inflammatory pathways and immune regulation are critical biological networks perturbed in pedAML LSC, and L-blast presents a high proliferative cell cycle activity combined with metabolic dysregulation. In addition, we identified novel LSC-specific downregulated targets, often described as TSGs in solid tumors, of which some are relevant in adult AML LSC or other pediatric hematological diseases.
References
Rasche, M. et al. Successes and challenges in the treatment of pediatric acute myeloid leukemia: a retrospective analysis of the AML-BFM trials from 1987 to 2012. Leukemia 32, 2167–2177 (2018).
De Moerloose, B. et al. Response-guided chemotherapy for pediatric acute myeloid leukemia without hematopoietic stem cell transplantation in first complete remission: results from protocol DB AML-01. Pediatr. Blood Cancer 66, e27605 (2019).
Abrahamsson, J. et al. Response-guided induction therapy in pediatric acute myeloid leukemia with excellent remission rate. J. Clin. Oncol. 29, 310–315 (2011).
De Kouchkovsky, I. & Abdul-Hay, M. Acute myeloid leukemia: a comprehensive review and 2016 update. Blood Cancer J. 6, e441 (2016).
Hope, K. J., Jin, L. & Dick, J. E. Acute myeloid leukemia originates from a hierarchy of leukemic stem cell classes that differ in self-renewal capacity. Nat. Immunol. 5, 738–743 (2004).
Shlush, L. I. et al. Tracing the origins of relapse in acute myeloid leukaemia to stem cells. Nature 547, 104–108 (2017).
Hanekamp, D. et al. Leukaemic stem cell load at diagnosis predicts the development of relapse in young acute myeloid leukaemia patients. Br. J. Hematol. 183, 512–516 (2018).
Witte, K. E. et al. High proportion of leukemic stem cells at diagnosis is correlated with unfavorable prognosis in childhood acute myeloid leukemia. Pediatr. Hematol. Oncol. 28, 91–99 (2011).
Bachas, C. et al. Gene expression profiles associated with pediatric relapsed AML. PLoS ONE 10, e0121730 (2015).
Pigazzi, M. et al. MLL partner genes drive distinct gene expression profiles and genomic alterations in pediatric acute myeloid leukemia: an AIEOP study. Leukemia 25, 560–563 (2011).
Jo, A. et al. High expression of EVI1 and MEL1 is a compelling poor prognostic marker of pediatric AML. Leukemia 29, 1076–1083 (2015).
Andersson, A. et al. Microarray-based classification of a consecutive series of 121 childhood acute leukemias: prediction of leukemic and genetic subtype as well as of minimal residual disease status. Leukemia 21, 1198–1203 (2007).
Balgobind, B. V. et al. Evaluation of gene expression signatures predictive of cytogenetic and molecular subtypes of pediatric acute myeloid leukemia. Haematologica 96, 221–230 (2011).
Zangrando, A., Dell’orto, M. C., Te Kronnie, G. & Basso, G. MLL rearrangements in pediatric acute lymphoblastic and myeloblastic leukemias: MLL specific and lineage specific signatures. BMC Med. Genomics 2, 36 (2009).
Ross, M. E. et al. Gene expression profiling of pediatric acute myelogenous leukemia. Blood 104, 3679–3687 (2004).
Ng, S. W. et al. A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature 540, 433–437 (2016).
Majeti, R. et al. Dysregulated gene expression networks in human acute myelogenous leukemia stem cells. Proc. Natl Acad. Sci. USA 106, 3396–3401 (2009).
Eppert, K. et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat. Med. 17, 1086–1093 (2011).
Forsberg, E. C. et al. Molecular signatures of quiescent, mobilized and leukemia-initiating hematopoietic stem cells. PLoS ONE 5, e8785 (2010).
Gal, H. et al. Gene expression profiles of AML derived stem cells; similarity to hematopoietic stem cells. Leukemia 20, 2147–2154 (2006).
Gentles, A. J., Plevritis, S. K., Majeti, R. & Alizadeh, A. A. Association of a leukemic stem cell gene expression signature with clinical outcomes in acute myeloid leukemia. JAMA 304, 2706–2715 (2010).
Saito, Y. et al. Identification of therapeutic targets for quiescent, chemotherapy-resistant human leukemia stem cells. Sci. Transl. Med. 2, 17ra9 (2010).
de Leeuw, D. C. et al. Attenuation of microRNA-126 expression that drives CD34+38- stem/progenitor cells in acute myeloid leukemia leads to tumor eradication. Cancer Res. 74, 2094–2105 (2014).
Duployez, N., Preudhomme, C. & Cheok, M. A 17-gene-expression profile to improve prognosis prediction in childhood acute myeloid leukemia. Oncotarget 9, 33869–33870 (2018).
Duployez, N. et al. The stem cell-associated gene expression signature allows risk stratification in pediatric acute myeloid leukemia. Leukemia 33, 348–357 (2018).
Elsayed, A. H. et al. A six-gene leukemic stem cell score identifies high risk pediatric acute myeloid leukemia. Leukemia https://doi.org/10.1038/s41375-019-0604-8 (2019).
Richard-Carpentier, G. & Sauvageau, G. Bringing a leukemic stem cell gene signature into clinics: are we there yet? Cell Stem Cell 20, 300–301 (2017).
Xu, Z. J. et al. Reduced protocadherin17 expression in leukemia stem cells: the clinical and biological effect in acute myeloid leukemia. J. Transl. Med. 17, 102 (2019).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, 607–613 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protoc. 14, 482–517 (2019).
Bolouri, H. et al. The molecular landscape of pediatric acute myeloid leukemia reveals recurrent structural alterations and age-specific mutational interactions. Nat. Med. 24, 103–112 (2018).
Rohrs, S. et al. Hypomethylation and expression of BEX2, IGSF4 and TIMP3 indicative of MLL translocations in acute myeloid leukemia. Mol. Cancer 8, 86 (2009).
Landrette, S. F. et al. Plag1 and Plagl2 are oncogenes that induce acute myeloid leukemia in cooperation with Cbfb-MYH11. Blood 105, 2900–2907 (2005).
Fu, S. et al. Overexpression of MYCT1 inhibits proliferation and induces apoptosis in human acute myeloid leukemia HL-60 and KG-1a cells in vitro and in vivo. Front. Pharm. 9, 1045 (2018).
Arora, D. et al. Expression of protein-tyrosine phosphatases in acute myeloid leukemia cells: FLT3 ITD sustains high levels of DUSP6 expression. Cell Commun. Signal. 10, 19 (2012).
Peng, L. et al. Structural maintenance of chromosomes 4 is required for leukemia stem cell maintenance in MLL-AF9 induced acute myeloid leukemia. Leuk. Lymph. 59, 2423–2430 (2018).
Andries, V. et al. NBPF1, a tumor suppressor candidate in neuroblastoma, exerts growth inhibitory effects by inducing a G1 cell cycle arrest. BMC Cancer 15, 391 (2015).
Korthuis, P. M. et al. CITED2-mediated human hematopoietic stem cell maintenance is critical for acute myeloid leukemia. Leukemia 29, 625–635 (2015).
Laverdiere, I. et al. Complement cascade gene expression defines novel prognostic subgroups of acute myeloid leukemia. Exp. Hematol. 44, 1039–1043 (2016).
Niu, Y. et al. Distinct prognostic values of Annexin family members expression in acute myeloid leukemia. Clin. Transl. Oncol. 21, 1186–1196 (2019).
Chavez-Gonzalez, A. et al. Expression of CD90, CD96, CD117, and CD123 on different hematopoietic cell populations from pediatric patients with acute myeloid leukemia. Arch. Med. Res. 45, 343–350 (2014).
Darwish, N. H. et al. Acute myeloid leukemia stem cell markers in prognosis and targeted therapy: potential impact of BMI-1, TIM-3 and CLL-1. Oncotarget 7, 57811–57820 (2016).
Hanekamp, D., Cloos, J. & Schuurhuis, G. J. Leukemic stem cells: identification and clinical application. Int. J. Hematol. 105, 549–557 (2017).
Dardaei, L., Longobardi, E. & Blasi, F. Prep1 and Meis1 competition for Pbx1 binding regulates protein stability and tumorigenesis. Proc. Natl Acad. Sci. USA 11, 896–905 (2014).
Song, L. et al. Protein tyrosine phosphatases receptor type D is a potential tumour suppressor gene inactivated by deoxyribonucleic acid methylation in paediatric acute myeloid leukaemia. Acta Paediatr. 105, 132–141 (2016).
Jasielec, J., Saloura, V. & Godley, L. A. The mechanistic role of DNA methylation in myeloid leukemogenesis. Leukemia 28, 1765–1773 (2014).
Yin, X. et al. Identification of long non-coding RNA competing interactions and biological pathways associated with prognosis in pediatric and adolescent cytogenetically normal acute myeloid leukemia. Cancer Cell Int. 18, 122 (2018).
Hemmati, S., Haque, T. & Gritsman, K. Inflammatory signaling pathways in preleukemic and leukemic stem cells. Front. Oncol. 7, 265 (2017).
Li, Z. et al. Growth and differentiation effects of Homer3 on a leukemia cell line. Asian Pac. J. Cancer Prev. 14, 2525–2528 (2013).
Stirewalt, D. L. et al. Identification of genes with abnormal expression changes in acute myeloid leukemia. Genes Chromosomes Cancer 47, 8–20 (2008).
Jain, M., Bakhshi, S., Shukla, A. A. & Chauhan, S. S. Cathepsins B and L in peripheral blood mononuclear cells of pediatric acute myeloid leukemia: potential poor prognostic markers. Ann. Hematol. 89, 1223–1232 (2010).
Pandey, G., Bakhshi, S., Thakur, B., Jain, P. & Chauhan, S. S. Prognostic significance of cathepsin L expression in pediatric acute myeloid leukemia. Leuk. Lymph. 59, 2175–2187 (2018).
Alatrash, G. Targeting cathepsin G in myeloid leukemia. Oncoimmunology 2, e23442 (2013).
Pandey, G. et al. Prognostic and therapeutic relevance of cathepsin B in pediatric acute myeloid leukemia. Am. J. Cancer Res. 9, 2634–2649 (2019).
Liebermann, D. A. & Hoffman, B. Myeloid differentiation (MyD) primary response genes in hematopoiesis. Oncogene 21, 3391–3402 (2002).
Wang, L. et al. Abnormal expression of GADD45B in human colorectal carcinoma. J. Transl. Med. 10, 215 (2012).
Korycka, A., Lech-Maranda, E. & Robak, T. Novel purine nucleoside analogues for hematological malignancies. Recent Pat. Anticancer Drug Discov. 3, 123–136 (2008).
Calle, E. E. & Kaaks, R. Overweight, obesity and cancer: epidemiological evidence and proposed mechanisms. Nat. Rev. Cancer 4, 579–591 (2004).
Zi, F. et al. Metformin and cancer: an existing drug for cancer prevention and therapy. Oncol. Lett. 15, 683–690 (2018).
Aries, I. M. et al. EMP1, a novel poor prognostic factor in pediatric leukemia regulates prednisolone resistance, cell proliferation, migration and adhesion. Leukemia 28, 1828–1837 (2014).
Zhang, Z. et al. Integrated bioinformatic analysis of microarray data reveals shared gene signature between MDS and AML. Oncol. Lett. 16, 5147–5159 (2018).
Bonardi, F., Fusetti, F., Deelen, P., van Gosliga, D. & Vellenga, E. A proteomics and transcriptomics approach to identify leukemic stem cell (LSC) markers. Mol. Cell Proteom. 12, 626–637 (2013).
Acknowledgements
Our gratitude goes to Dr. F. Plasschaert, C. Matthys of the Cord Blood Bank, and the Department of Pediatric Hematology and Oncology of the Ghent University Hospital (Ghent, Belgium) for providing samples. We thank all patients and their parents for their participation in the study, as well as the data managers involved in the clinical trials. This research was funded by the Belgian Foundation against Cancer (grant 2014-265), FOD-KankerPlan (Actie29, grant to J.P.), vzw Kinderkankerfonds (grant to T.L.), and the Research Foundation—Flanders (Fonds voor Wetenschappelijk Onderzoek Vlaanderen, FWO, grant 1113117 to B. Depreter).
Author information
Authors and Affiliations
Contributions
Conception and design, acquisition of data, or analysis and interpretation of data: B.Depreter, T.L., J.P., and M.H.; drafting the article or revising it critically for important intellectual content: B.D.M., E.T., K.V., L.D.; final approval of the version to be published: B.D.P., T.L., J.P., B.D.M., M.H., E.T., K.V., L.D., B.Denys, A.U., A.V.D., M.-F.D., J.V.d.W.T.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Patient consent
All patients gave consent for inclusion in this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Depreter, B., De Moerloose, B., Vandepoele, K. et al. Deciphering molecular heterogeneity in pediatric AML using a cancer vs. normal transcriptomic approach. Pediatr Res 89, 1695–1705 (2021). https://doi.org/10.1038/s41390-020-01199-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-020-01199-3
- Springer Nature America, Inc.