Abstract
The urothelium is a stratified epithelium composed of basal cells, one or more layers of intermediate cells, and an upper layer of differentiated umbrella cells. Most bladder cancers (BLCA) are urothelial carcinomas. Loss of urothelial lineage fidelity results in altered differentiation, highlighted by the taxonomic classification into basal and luminal tumors. There is a need to better understand the urothelial transcriptional networks. To systematically identify transcription factors (TFs) relevant for urothelial identity, we defined highly expressed TFs in normal human bladder using RNA-Seq data and inferred their genomic binding using ATAC-Seq data. To focus on epithelial TFs, we analyzed RNA-Seq data from patient-derived organoids recapitulating features of basal/luminal tumors. We classified TFs as “luminal-enriched”, “basal-enriched” or “common” according to expression in organoids. We validated our classification by differential gene expression analysis in Luminal Papillary vs. Basal/Squamous tumors. Genomic analyses revealed well-known TFs associated with luminal (e.g., PPARG, GATA3, FOXA1) and basal (e.g., TP63, TFAP2) phenotypes and novel candidates to play a role in urothelial differentiation or BLCA (e.g., MECOM, TBX3). We also identified TF families (e.g., KLFs, AP1, circadian clock, sex hormone receptors) for which there is suggestive evidence of their involvement in urothelial differentiation and/or BLCA. Genomic alterations in these TFs are associated with BLCA. We uncover a TF network involved in urothelial cell identity and BLCA. We identify novel candidate TFs involved in differentiation and cancer that provide opportunities for a better understanding of the underlying biology and therapeutic intervention.
Similar content being viewed by others
Introduction
The urothelium is a highly specialized epithelium lining the lower urinary tract, from the renal pelvis to the proximal urethra. It is composed of 3–8 cell layers [1]. Cells in the basal layer are small and lack expression of genes related to urothelial identity. Urothelial stem cells are thought to reside in this layer. Cells in the intermediate layers engage in differentiation and display increasing levels of uroplakins (UPKs), transmembrane proteins involved in the formation of urothelial plaques. The superficial layer is composed of umbrella cells, which are large [2], multinucleated, long-lived [3], differentiated cells generated by endoreplication [4]. Umbrella cells contain discoidal-shaped/fusiform vesicles that allow apical membrane expansion in response to bladder filling [5] and provide barrier impermeability [6]. Acquisition of urothelial lineage identity involves the extinction of the basal program-which is reminiscent of the epidermis and is characterized by expression of KRT5 - and the expression of urothelial markers, most notably UPKs and KRT20 [7, 8]. KRT14 is expressed by a small fraction of basal cells thought to have stem cell properties [9] (Fig. 1).
The urothelium is a slow-renewal epithelium: in homeostatic conditions, <1% of cells express Ki67 or are labeled with BrdU [10]. The lifespan of a rodent urothelial cell is estimated to be ~200 days [11]. However, upon injury, rapid regeneration ensues: basal cells secrete Sonic hedgehog (SHH) which stimulates stromal expression of Wnt agonists [12], promoting cell proliferation. Little is known about the spatial arrangements of the cellular neighborhood(s) in which these signaling loops take place. KRT14+ basal cells are involved in regeneration upon cyclophosphamide injury [9]. The Mendelsohn lab has proposed that cells other than basal are also important in urothelial regeneration [8]. SHH-expressing cells include both KRT5+ basal cells, as described by Shin et al., and two KRT5- UPK+ cell types: P-cells, present only in the embryo and I-cells, also present in the adult. Lineage tracing shows that transient intermediate urothelial cells act as progenitors both in the embryo and in the adult [8]. The emerging scenario is one where the urothelial lineage may not simply progress in a linear fashion, from basal to intermediate to umbrella cells; urothelial progenitors may instead display a wider range of phenotypes (or be in a wider range of states), depending on the extent/type of damage.
A deeper understanding of these aspects of urothelial biology is required to dissect the regulatory mechanisms involved in diseases of the bladder. Here, we combine a comprehensive review of the literature with new analyses of public data to provide a better understanding of the landscape of alterations in the regulation of urothelial differentiation associated with bladder cancer (BLCA) through the identification of well-established and novel candidate transcription factors (TFs) involved therein.
An overview of bladder cancer genetics
The majority of BLCA arise in the urothelium and >90% are urothelial carcinomas. Conventionally, BLCA are categorized into non-muscle invasive BLCA (NMIBC) (ca. 75%) and muscle-invasive BLCA (MIBC) (ca. 25%). NMIBC are often indolent, papillary tumors. Most of them are genomically stable, unlike MIBC tumors which are life-threatening and genomically unstable [13, 14]. It is thought that BLCA arises through two pathways: papillary tumors constitute the majority of NMIBC and develop from papillary hyperplasia displaying lineage fidelity, whereas the majority of MIBC arise from dysplasia/carcinoma in situ. Recent work suggests that MIBC develops through preneoplastic lesions (carcinoma in situ) that can either maintain or lose urothelial lineage fidelity [15, 16], the latter resulting from epigenetic changes [17] (Fig. 2).
Topological scheme depicting the involvement of well-established (highlighted in dark background) and novel candidate (highlighted in discontinuous outline) TFs participating in the luminal vs. basal differentiation programs. Color code: purple for “Luminal-enriched”, beige for “Common” and teal for “Basal-enriched” TFs.
Mutations in some BLCA-relevant genes—such as STAG2, RBM10, and KDM6A—can already be detected in normal urothelial cells from subjects without BLCA and undergo positive selection [18]. TERT hotspot promoter mutations and deletions in chr. 9 are shared by tumors evolving through both BLCA progression pathways, suggesting that they are early events in tumorigenesis. Papillary NMIBC commonly harbor activating mutations in FGFR3 and PIK3CA, and loss-of-function mutations in the tumor suppressors STAG2 and KDM6A [13, 19, 20]. Alterations in the RB (e.g., through inactivation of CDKN2A) and p53 pathways drive progression to MIBC [21]. Non-papillary tumors often invade muscle and are characterized by inactivation of tumor suppressors (e.g., TP53, RB1, ARID1A): TP53 mutations are highly frequent in urothelial dysplasia [22] and the RB pathway is altered in the majority of MIBC through a variety of genetic mechanisms [23]. Mutations in genes involved in chromatin regulation (e.g., MLL2, ARID1A, KDM6A, EP300) are enriched in MIBC more than in any other malignancy [23], 76% of the samples being mutated for at least one epigenetic regulator. Figure 2 summarizes current knowledge on the morpho-genetic pathways leading to BLCA.
Identification of candidate TFs relevant for urothelial lineage identity and carcinogenesis
The disruption of cell differentiation during BLCA development and progression is reflected by the loss of lineage fidelity, underlined by the transcriptional taxonomic classification of tumors. Consequently, an improved understanding of the gene regulatory networks (GRN) and epigenomic mechanisms involved therein is necessary.
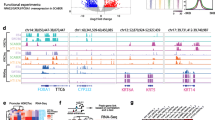
To systematically identify candidate TFs relevant for urothelial differentiation and carcinogenesis, we assessed the expression of all TFs from the consensus list of Lambert et al. [24] in patient-derived tumor organoids [25], recapitulating features of luminal and basal BLCA subtypes (Supplementary Fig. 1). We classified TFs as “luminal-enriched” or “basal-enriched” if they were: 1) present among the top 100 highest-expressed TFs only in luminal or basal BLCA organoids, respectively, 2) significantly differentially expressed in luminal vs. basal organoids (log2 fold change cutoff of ±1 and FDR < 0.05), and 3) significantly differentially expressed in Luminal Papillary vs. Basal/Squamous tumors from the TCGA series (FDR < 0.05). We also defined a category of “Common TFs” including those that were among the top 100 highest-expressed TFs in both basal and luminal organoids. Common TFs were analyzed further if a significant enrichment (P values < 0.05) of their associated motif was found in ATAC-Seq peaks in normal human bladder [26] (Table 1 and Fig. 3). This strategy unveiled known, as well as novel, TFs involved in urothelial lineage specification. See Box 1 for more details. To further validate the role of these TFs, we assessed their activity according to target expression and transcription factor enrichment analysis with differentially expressed genes from luminal vs. basal BLCA organoids, with the ChEA 2022 database through the Enrichr website and filtering by TFs from the consensus list of Lambert et al. [24]. The results show significant enrichment of selected well-established urothelial TFs, thus validating the overall strategy used (Supplementary Fig. 2). Tables 2 and 3 provide detailed information on the molecular features of the TFs selected and their relevance to cancer.
Normal urothelial cells acquire somatic mutations during aging, some of which endow them with clonal selection capacity. [18] A subset of the progeny acquires additional mutations that promote tumor development, associated with specific phenotypes. Non-invasive preneoplastic lesions arising through the papillary pathway (upper track) have a lower propensity to invade muscle, unlike those arising through the non-papillary pathway (lower tracks). Carcinoma in situ, a precursor participating in the non-papillary pathway, can display either luminal or basal features, [15, 16] possibly accounting for the existence of luminal as well as basal tumors along this track. However, this model does not rule out that—in some cases—luminal tumors arising through the papillary pathway can evolve to acquire basal features. It is also possible that some of the CIS-like lesions represent the intraluminal expansion of tumor cells along the urothelial lining (surface “cancerization”).
Additionally, we assessed the expression of all consensus TFs in normal human bladder samples from GTEx, investigated genomic TF binding motifs in open chromatin regions identified from ATAC-Seq data of normal human urinary bladder mucosa [26], and confirmed their expression in normal urothelial cells using single cell RNA-Seq [27]. Highly expressed TFs ( > 4000 DESeq2-normalized counts) with significant enrichment of their associated motif (P values < 0.05) were categorized as “normal-enriched TFs” and selected for further analysis (Supplementary Table 1).
TFs associated with luminal differentiation (“luminal-enriched”)
PPARG and RXRA
PPARG is a member of the peroxisome proliferator-activated receptor subfamily of nuclear receptors, together with PPARA and PPARD. Its N-terminus contains the ligand-independent activation function domain (AF-1), which is poorly conserved between the 3 family members. The highly conserved central region contains the DNA-binding domain (DBD) with two zinc fingers that bind to PPAR response elements (PPREs) in target genes and a flexible hinge region that separates the DBD from the highly conserved ligand-binding domain (LBD). The C-terminus contains a ligand-dependent activation domain, AF-2.
PPARG heterodimerizes with retinoid X receptor alpha (RXRA) and regulates transcription by binding to PPREs present in regulatory regions of target genes. RXRA homo- or hetero-dimerizes with 14 nuclear receptors, including PPARG, PPARA, and PPARD [28]. In the absence of ligand, PPARG/RXRA dimers bind to co-repressors such as NCoR1 and SMRT, which recruit histone deacetylases. Upon ligand binding, co-repressors are replaced by co-activators such as mediator complex subunit 1 (MED1) and PPARG coactivator 1 alpha. Natural PPARG ligands include fatty acids, arachidonic acid and its metabolites. PPARG is expressed in adipose tissue and several epithelia and it plays important roles in adipogenesis and glucose homeostasis. Synthetic agonists (e.g., thiazolidinediones) acting as insulin sensitizers are used to treat type 2 diabetes [29]. There is some evidence that they can increase BLCA risk [30] although the results are controversial.
In the urothelium, PPARG is expressed in all cell layers and is a key regulator of differentiation. PPARG activation, together with EGFR inhibition, promotes the differentiation of normal human urothelial cells [31], mouse urothelial organoids [32], and human pluripotent stem cells induced to acquire an urothelial fate [33]. In vivo, it is required for terminal differentiation of umbrella cells in the developing ureter [34]. Pparg deletion throughout the urothelium (using the Shh-Cre driver allele), or in basal (Krt5-CreERT2) or suprabasal (Upk2-CreERT2) cells, reveals its critical role in mitochondrial biogenesis, fatty acid transport, and urothelial identity. In the absence of PPARG, basal cells undergo squamous-like differentiation and umbrella cells fail to mature [35]. This is reminiscent of the squamous metaplasia reported to occur upon retinoid deficiency [36]. PPARG is also required for homeostatic regeneration: upon injury/uropathogenic E. coli infection, NF-kB signaling is transiently activated and PPARG is required to dampen NF-kB signaling, which otherwise results in chronic inflammation [35, 37]. Ectopic expression of an activated form of PPARG in basal urothelial cells in mice induces enhanced differentiation but not tumor formation. However, upon N-butyl-N-(4-hydroxybutyl)-nitrosamine (BBN) administration, luminal, immune-cold tumors develop, likely through reduced NF-kB signaling [38]. A subset of luminal tumors loses canonical identity and acquires basal features over time, in association with PPARG down-regulation [38].
PPARG focal amplifications occur in ~10% of MIBC [39] leading to overexpression of the protein and its target genes. In addition, seven recurrent PPARG point mutations have been identified, five of which confer gain-of-function. Those affecting the LBD (M280I, I290M and T475M) endow the protein with ligand-independent activity and favor its interaction with coregulators [40]. PPARG mutations and amplifications are associated with luminal tumors, which display an active PPARG regulon. In the consensus publication of BLCA subtypes, PPARG gains/amplifications or fusions were found in 76% of LumNS tumors and in 89% of LumU tumors [23]. RXRA hotspot mutations (S242F/Y) have also been identified in 5% of MIBCs, mostly in the luminal subgroup [39]. These gain-of-function mutations confer enhanced binding affinity for PPARG and promote ligand-independent activation [41, 42]. Despite these important data, and work supporting that retinoic signaling promotes luminal differentiation [8], there is no direct evidence showing a role for RAR-RXR heterodimers in the regulation of luminal cell identity.
The role of the PPARG/RXRA axis in BLCA is context-dependent: luminal tumors are addicted to PPARG/RXRA signaling whereas Basal-Squamous (Ba/Sq)-like tumors display down-regulation of this pathway. PPARG loss-of-function mutations, hemizygous deletions, and DNA hypermethylation - associated with decreased PPARG activity - have been described in Ba/Sq-type MIBC [43]. Overexpression of PPARG in low-expressing BLCA cells activated PPARG signaling and inhibited growth, partly by down-regulating EGFR expression, while no effect was seen in PPARG high-expressing cells. Transcriptomic analysis revealed increased activity of lipid metabolism and urothelial differentiation pathways, and decreased activity of immunity/inflammation pathways [43]. These effects may require the cooperation of GATA3 and/or FOXA1 [44]. Conversely, a recent genome-wide CRISPR knockout screening identified GATA3, SPT6, and cohesin complex components as upstream positive regulators of PPARG expression in luminal BLCA cells [45].
In NMIBC, PPARG activity and immune cell infiltration are anticorrelated. Consistently, PPARG/RXRA activation in cancer cells inhibits secretion of pro-inflammatory cytokines, such as IL6 and IL8, that function as chemoattractants of effector T cells. Tumors from MBT2 murine cells overexpressing RXRA (S242F/Y) have substantially fewer CD3+ and CD8+ T cells and are partially resistant to immunotherapy. Down-modulating PPARG or RXRA significantly increases cytokine expression, suggesting therapeutic strategies that could sensitize patients to immunotherapies [42]. Pharmacological or genetic PPARG inhibition reduces proliferation, migration, and invasion of BLCA cells, especially those harboring PPARG amplification or RXRA activating mutations, highlighting the potential of PPARG as a therapeutic target in luminal BLCA [46, 47].
GATA proteins
GATA2 and GATA3 are Zn finger TFs that bind the GATA consensus sequence [48]. GATA proteins play fundamental roles in development and differentiation, contributing to cell lineage specification and morphogenesis in a wide variety of tissues. In the adult, GATA2 is expressed broadly while GATA3 displays a more tissue-restricted expression pattern including skin, breast, and urothelium. They can act as “pioneer factors”, being able to access silenced chromatin and facilitating the binding of other TFs. GATA3 germline mutations cause Congenital Anomalies of the Kidney and Urinary Tract (CAKUT) syndrome, characterized by urogenital morphogenesis defects [49].
Gata3 ablation in the adult murine mammary gland results in luminal cell death, disruption of epithelial architecture, and expansion of an undifferentiated luminal cell population [50]. In the bladder, GATA3 is expressed in all urothelial layers and it is a well-established urothelial lineage marker [51]. In mice, inactivation of Gata2 and Gata3 in the urogenital system leads to morphological alterations that recapitulate the human CAKUT phenotype [52, 53]. GATA3, together with FOXA1, plays a central role in differentiation through PPARG signaling. GATA3 silencing prevents the differentiation-associated down-regulation of TP63 and the expression of intermediate/late urothelial markers [54]. Ectopic GATA3 expression in T24 and mesenchymal-like UMUC3 BLCA cells reduces proliferation [55] and the combined overexpression of GATA3 and FOXA1 in cells with a basal phenotype leads to the expression of a luminal-specific program [44]. GATA3 binding motifs are enriched in chromatin regions that become accessible upon differentiation of normal human urothelial [54] and BLCA cells [56].
GATA3 has been proposed as one of the best markers of urothelial cell identity [57]. It is amplified in 9.8% of MIBC [14]. Expression is associated with luminal differentiation and down-regulation is common in high-grade tumors [58, 59], most notably in the Ba/Sq subtype. GATA2 is also one of the most down-regulated genes in the Ba/Sq subtype [60] and promoter hypermethylation is significantly associated with MIBC progression [61].
FOXA1
FOXA1 (also known as HNF3α) is the founding member of the Forkhead box (Fox) superfamily. It contains a winged-helix DNA binding domain, with a helix-turn-helix core flanked by two loops that stabilize chromatin binding, and is structurally similar to linker histones H1 and H5 [62, 63]. FOXA1 typically binds as a monomer and acts as a pioneer factor [64]. FOXA1 cooperates with nuclear hormone receptors, most notably with the estrogen receptor (ER) and androgen receptor (AR) in breast and prostate epithelium, respectively, and displays sexual dimorphism [65]. It also interacts with the repressor NR0B2 [66] and with the glucocorticoid receptor (GR or NR3C1) [67].
FOXA1 is required for the development/differentiation of endoderm-derived organs such as liver, lung, pancreas, or bladder and its expression is maintained throughout adulthood [68, 69]. In the bladder, FOXA1 is expressed in all urothelial layers [54] and it cooperates with PPARG to drive differentiation [70]. In normal human urothelial (NHU) cells induced to differentiate with troglitazone and EGFR inhibition, FOXA1 is up-regulated, binds to the promoters of UPKs, and activates their expression [70].
FOXA1 is expressed in luminal BLCA and its knockdown in RT4 cells enhances proliferation while its overexpression in T24 cells decreases cell proliferation and invasion, and increases E-cadherin expression [71]. GABPA has recently been proposed as an upstream activator of FOXA1 transcription in BLCA, dictating luminal differentiation and suppressing stem cell traits and invasion. GABPA expression correlates positively with luminal signatures and with better patient survival [72]. Foxa1 knockout mice develop preneoplastic urothelial lesions in adulthood: males develop urothelial hyperplasia while females show keratinizing squamous metaplasia [73]. The combined deletion of Foxa1 and Pten in superficial and intermediate cells leads to squamous tumors [74].
FOXA1 has been proposed to have mainly a tumor suppressor role but it can also act as an oncogene in acute myeloid leukemia, esophageal, or lung adenocarcinoma [75]. It is a significant cancer driver in prostate and breast tumors [76, 77] and it is mutated in 5% of BLCA-TCGA cases [14]. FOXA1 is detected in the majority of early-stage BLCA and in luminal-type MIBC and its expression is reduced/lost in a subset of MIBC with Ba/Sq or neuroendocrine features [14, 23, 57, 59, 71, 78].
ELF3
ELF3, a member of the ETS family of TFs, is highly expressed in the bladder as well as in other endodermal tissues. ELF3 is induced during epidermal differentiation, where it regulates the expression of SPRR2A, a marker of terminal maturation [79]. Constitutive Elf3 inactivation leads to embryonic or early postnatal lethality, associated with intestinal alterations [80]. In the urothelium, ELF3 expression increases with differentiation; in NHU cells, its up-regulation is driven by PPARG and is required for the induction of FOXA1 and GRHL3 and the expression of UPK3A [81].
ELF3 is a downstream component of WNT7B signaling, which is down-regulated in high-grade BLCA and associated with poor prognosis. Loss- and gain-of-function studies indicate that the WNT7B/FZD5-ELF3-NOTCH1 axis suppresses the epithelial-mesenchymal transition (EMT) and stem-like properties in BLCA cells [82]. ELF3 overexpression in UMUC3 mesenchymal-like cells induces an epithelial phenotype and reduces invasiveness [83].
ELF3 is a mutational driver in BLCA and cholangiocarcinoma: truncating, splice site and missense mutations, and deletions, occur in up to 14% of BLCA consistent with a tumor suppressor role. ELF3 alterations are enriched in the LumNS consensus subtype, as are PPARG amplifications [23]. ELF3 is expressed in low-grade human BLCA and, together with PPARG, FOXA1, GATA3, and GATA2, it is among the most down-regulated TFs in basal tumors [60].
GRHL3
GRHL3 (or GET1) is a Grainyhead family member that can act both as an activator or a repressor [84, 85]. GRHL3 is highly expressed in the esophagus, skin, vagina, and bladder. In the mouse bladder, GRHL3 is mainly expressed in luminal and umbrella cells; at E16.5 and E18.5, Grhl3−/− mice show defective urothelial barrier formation and down-regulation of UPKs, which are direct target genes [86]. The Southgate laboratory showed that, together with FOXA1, it is a downstream effector of PPARG and ELF3 in NHU cells [81]. Up-regulation of GRHL3 has also been described during the urothelial differentiation of human embryonic stem cells and induced pluripotent cells [87]. It is expressed at high levels in differentiated human BLCA cells. GRHL3 knockdown in luminal RT4 cells promotes a more invasive phenotype while overexpression in T24 cells reduces migration and invasiveness but it does not impact on proliferation/survival. These results suggest a tumor suppressor role [88].
Our analyses additionally identified BHLHE41, MECOM, MYCL, NCOA1, NR2F2, NR2F6, REPIN1, SREBF1, TBX2, TBX3, TRAFD1, and ZBTB7C as novel putative TFs that are enriched in the luminal transcriptional program and for which there is suggestive evidence for having a role in urothelial differentiation and/or BLCA (Box 2). Therefore, they merit additional study in BLCA.
TFs associated with basal/squamous differentiation (“basal-enriched”)
TP63
TP63 is a member of the P53 family that contains an extended C-terminal region thought to have repressor activity [89]. At least six isoforms (TAp63α, TAp63β, TAp63γ, ΔNp63α, ΔNp63β, and ΔNp63γ) are generated through alternative splicing and alternative promoter usage [90,91,92]. The latter leads to TAp63 or ΔNp63, containing—or lacking—a N-terminal transactivation domain, respectively. TAp63 can transactivate P53 target genes (e.g., CDKN1A, BAX and MDM2), whereas ΔNp63 has both activator and repressor functions [92,93,94]. Both isoforms regulate transcriptional programs related to cell cycle [93], differentiation [95], apoptosis [92], and senescence [96]. Cell type-specific cofactors may contribute to determine transcriptional outputs of ΔNp63 [97]. Using epigenomic profiling, most ΔNp63 binding sites were found in enhancers, where ΔNp63 has been proposed to bookmark genes in stratified epithelia and in the squamous lineage in pancreatic cancer [98, 99]. TP63 levels are also controlled post-translationally, DNA damage being a major regulator.
TP63 proteins are expressed in the basal layer of stratified epithelia [100] and are required for epidermal development and differentiation [95]. In mice, TAp63 isoforms are detected in the urothelium at E16.5, where ΔNp63 is first detected at postnatal day 1. In the adult murine and human urothelium, TP63 isoforms are expressed in the basal/intermediate layers [101]. The diversity of TP63 isoforms has hampered acquiring a precise understanding of their spatiotemporal expression and caution is suggested when making cross-species extrapolations. Trp63 knockout mice have severe ectodermal and heart defects and die shortly after birth [102,103,104]. They develop a single-layered, umbrella cell-like, urothelium [105,106,107]. Genetic defects in human TP63 have similar, but milder, phenotypic consequences [108,109,110,111].
There is little functional evidence on how TP63 proteins regulate differentiation in BLCA at the genomic level. Studies in pancreatic cancer show that, for the basal program to emerge, a concomitant down-regulation of lineage identity programs (i.e., GATA/FOXA) and an up-regulation of ΔNp63 is required [112].
TP63 is a mutational driver in BLCA, largely through missense mutations. In MIBC, ΔNp63α is the most abundant transcript, present at highest levels in Ba/Sq tumors and at low/undetectable levels in luminal and neuronal subtypes [113]. ΔNp63 is up-regulated in NMIBC and MIBC compared to normal urothelium while TAp63 is undetectable [114]. Similar findings were made when analyzing total TP63 in the UROMOL and TCGA-BLCA cohorts [14, 21, 114]. Expression of TP63 isoforms has been associated with tumor stage and molecular subtype: ΔNp63- tumors are KRT20+ while ΔNp63+ tumors are KRT5+ [101]. The association of TP63 expression with patient outcome is controversial [101, 113]. Sarcomatoid MIBC lack ΔNp63 expression in association with the worst outcome [115].
TFAP2s
TFAP2 family members (A-E) contain a N-terminal transactivation domain, a C-terminal helix-span-helix domain, and a central basic region. They are expressed broadly and play important roles in mouse development [116]. TFAP2s have been implicated in epidermal differentiation in mice [117] and humans [118]: TAp63α directly induces Tfap2c during differentiation and its knock-down blunts Krt14 expression [116]. In keratinocytes, functional TFAP2 binding sites have been identified in the promoter of Krt5 and other differentiation genes [117,118,119] and TFAP2A and TFAP2C co-regulate a subset of ΔNp63 targets [120]. In the murine urothelium, expression of Tfap2a, Krt6a, Krt14, and Krt15 is repressed by PPARG, further suggesting that TFAP2A is part of the basal program [35].
TFAP2A and TFAP2C are up-regulated in Ba/Sq BLCA cells and tumors and in areas of squamous metaplasia [121]. In BLCA patients treated with cisplatin, TFAP2A expression is an independent predictive marker of response and survival. Neither TPAP2A nor TFAP2C are identified as mutational driver genes in BLCA.
MYC
MYC is a basic helix-loop-helix (bHLH) and leucine zipper (LZ) protein that is ubiquitously expressed and plays a central role as an oncogene in almost all tumor types as it is crucially required for cell proliferation [122]. It regulates the expression of ~10% of the protein-coding genes.
Aberrant MYC activity in BLCA occurs through a wide variety of mechanisms such as genetic alterations [123], transcriptional and post-transcriptional regulation [124,125,126], or altered protein stability [127]. MYC amplifications occur in ~9% of MIBC and it is overexpressed—and its program is enriched—in a subset of T1 BLCA [128]. A recent study identified MYC as a key effector of mutant FGFR3 signaling and FGFR3 as a direct transcriptional MYC target. Consistently, MYC and FGFR3 levels were directly correlated in FGFR3-mutant tumors [125]. However, other work has shown an enrichment of MYC expression and activity in basal tumors [59, 129] and functional studies in human BLCA cells revealed that MYC expression is controlled by TP63 [130]. These findings highlight complex, context-dependent, roles of MYC in BLCA biology.
NOTCH
NOTCH is involved in a wide range of biological processes and mechanistically links cell-cell interactions to transcriptional responses. In our analyses, its critical effector RBPJ is among the top expressed TFs in normal urothelium and BLCA organoids and its target HES2 is significantly enriched in basal BLCA organoids. Notch signaling is required at multiple levels during development.
In normal urothelium, NOTCH1 is up-regulated during differentiation [131] and, in normal murine urothelial organoids, Notch pathway inhibition results in up-regulation of Tp63 and reduced expression of luminal markers [32]. Tissue-specific inducible inactivation of Notch in the mouse urothelium led to hyperplasia, inflammation, and mucosal sloughing [132].
Both oncogenic and tumor-suppressive roles have been described for the Notch pathway in a context-dependent manner in several tumor types. In BLCA, NOTCH1 inactivation is more common in Ba/Sq tumors and results in increased MAPK signaling [131, 133, 134]. In contrast, NOTCH2 appears to act as an oncogene [135]. Both tumor-suppressive and oncogenic roles have been attributed to NOTCH3 [135, 136]. Mutations in NOTCH1, NOTCH2, and NOTCH3 occur in 6%, 12% and 4% of samples in the TCGA-BLCA cohort [14], respectively. Missense mutations are the most frequent alteration in NOTCH1 and NOTCH3 though truncating mutations and amplifications are also reported. NOTCH2 copy number gains/amplifications have been reported in Ba/Sq tumors, associated with lower survival in the TCGA cohort [135]. Other NOTCH pathway genes such as MAML1, NCSTN, PSEN1 are also significantly mutated in BLCA, regardless of stage or grade [133].
ETS family
ETS1 and ETS2 are characterized by the presence of a conserved ETS DNA-binding domain that recognizes the core consensus DNA sequence GGAA/T in target genes. They can act as transcriptional activators or repressors and play a key role in the regulation of cell proliferation, differentiation, survival, invasion, and angiogenesis [137,138,139].
ETS1 is expressed in the normal urothelium and its down-regulation is associated with high-grade/stage BLCA [140]. However, another study reported high levels of ETS1 mRNA in BLCA, compared to paired normal tissue. ETS1 silencing in BLCA cells inhibits cell migration and invasion, suggesting an oncogenic role [141]. Two microRNAs (miR-193b-3p and mir-106a) whose down-regulation is associated to tumor progression suppress ETS1 expression and, as a consequence, proliferation, migration and invasion of BLCA cells [142, 143]. Deep deletions of ETS1 and ETS2, as well as ETS2 amplification, have been reported in <1% tumors from the TCGA.
As shown in Box 2 and Table 1, our analyses additionally identified ARNTL2, BACH1, ELK3, HES2, HMGA2, KLF7, KLF13, NR3C1/GR, SNAI2 and ZBED2 as significantly associated with the basal transcriptional program and there is suggestive evidence of their role in urothelial differentiation and/or BLCA. Therefore, these TFs merit further investigation in BLCA.
TFs common to luminal and basal subtypes
KLF4 and KLF5
Krüppel-like factors (KLFs) act either as activators or repressors and participate in a wide range of cellular processes, including epithelial differentiation, in multiple tissues.
Klf5 deletion in the developing murine bladder impaired urothelial stratification and differentiation [144], accompanied by reduced expression of Pparg, Elf3, and Grhl3, supporting its participation in a urothelial GRN [144]. Similarly, KLF5 is required for basal-to-luminal differentiation in the prostate [145]. In BLCA cells, KLF5 overexpression increases proliferation, lamellipodia formation, and cell migration by direct promoter binding and up-regulation of the FYN kinase [146]. KLF5 may also promote angiogenesis by directly regulating VEGFA [147].
Genetic alterations in KLF4 (3%; amplifications and missense mutations), KLF5 (8%; amplifications, deletions, missense mutations), and KLF6 (9%; amplifications) are most frequent in the TCGA-BLCA cohort. KLF4 and KLF5 are abundant in normal urothelium [148]. KLF4 is frequently down-regulated in BLCA cells and tissues through promoter methylation [149]. Its overexpression triggers apoptosis in vitro [148, 150] and inhibits tumor growth in vivo [151], consistent with a tumor suppressor role.
SP1 and SP3
The specificity protein (Sp) TFs belong to the Sp/KLF family and consist of four members in humans: Sp1, Sp2, Sp3, and Sp4 [152]. Sp1 and Sp3, which emerge as “common” TFs in our analyses, are ubiquitous TFs implicated in the control of a wide variety of cellular processes [153]. In BLCA, they have been reported to play a role in cell invasion through regulating the expression of the metalloproteinase MMP2 [154, 155]. In tumors, SP1 expression is associated with poor prognosis and progression [156].
Biological processes/pathways involved in BLCA urothelial lineage fidelity
Our analyses have identified other TFs associated with biological processes for which there is evidence of their involvement in urothelial differentiation and/or BLCA. The majority of them were highly expressed in both luminal and basal organoids—hence classified as “common TFs” (Table 1).
Xenobiotic metabolism and oxidative stress. The P450 cytochromes, epoxide hydrolase, and glutathione S-transferases (GST) are involved in xenobiotic metabolism, protecting urothelial cells from damage impinged by the continued contact of urothelial cells with urine and exposure to carcinogens. Consequently, polymorphisms in genes involved in xenobiotic metabolism are associated with an increased BLCA risk (reviewed in [157]). Tobacco and urinary infections, which are known to cause oxidative stress, are also associated with increased BLCA risk [158].
AHR encodes a bHLH protein that acts as a “sensor” through binding to a wide variety of xenobiotics (e.g., halogenated aromatic hydrocarbons and polycyclic aromatic hydrocarbons) as well as endogenous molecules (e.g., flavonoids and metabolites of tryptophan) (reviewed in [159]). Upon ligand binding and nuclear translocation, AHR dimerizes with ARNT and regulates expression of multiple phase I (e.g., CYP1A1, CYP1A2, and CYP1B1) and phase II genes (e.g., NAD(P)H Quinone Dehydrogenase 1, NQO1 and GSTs) [160,161,162].
AHR is expressed at high levels in the urothelium (GTEx), in NMIBC [163], and in luminal MIBC [164] and the gene is mutated in 12% of MIBC in the TCGA cohort, 52% of which are amplifications. Recurrent in-frame deletions of AHR exons 8 and 9 occur in primary and metastatic BLCA [165]. Mutant AHR is constitutively active, down-regulates differentiation markers, up-regulates stem cell markers, and confers anchorage independent growth to bladder organoids [166]. Upon BBN administration, Ahr-null mice develop MIBC at a much higher frequency than Ahr wild-type mice suggesting a tumor-suppressive role in normal urothelium [163].
NFE2L2 (Nuclear factor, erythroid 2 like 2 or NRF2) is a master regulator of the response to oxidative stress. NRF2 regulates the expression of AHR as well as genes coding for proteins with antioxidant activity (e.g., NQO1, SQSTM1, GSTA1, GSTM1, and GSTP1) that may be important in bladder carcinogenesis [166]. NRF2 amplifications or hotspot missense mutations in 7% of MIBC in the TCGA cohort suggest an oncogenic role in BLCA, but little is known at the mechanistic level. Additional processes may lead to NRF2 hyperactivation including down-regulation of the KEAP1-binding protein GULP1 as a result of promoter methylation, especially in MIBC [167]. BACH1, a basic LZ TF that regulates the expression of genes involved in the oxidative stress pathway mediated by NFE2L2, is enriched in basal-like organoids and tumors (Table 1). Our analysis also found NFE2L1 (or NRF1) among the highest expressed TF in both BLCA organoids and normal bladder, although there is virtually no information on its role in the urothelium or in BLCA (Box 2).
The circadian clock machinery harmonizes physiological processes to provide homeostasis within the 24h cycle and is modulated by clock genes. The diurnal variation in bladder function and the time-dependent expression of these genes support a role in urothelial function [168], consistent with the robust expression of NPAS2, ARNTL2/BMAL2, and BHLHE41/DEC2 in BLCA organoids (Table 1). Altered expression of clock-related genes has been reported in BLCA [169]. Iyyanki et al. have recently described NPAS2 as a novel luminal TF that binds and regulates the FOXA1 promoter and is, in turn, up-regulated upon overexpression of FOXA1 or GATA3. This regulatory loop may contribute to BLCA cell proliferation and migration [56]. BHLHE41/DEC2 is also luminally-enriched, interacts with ARNTL and other nuclear receptors (e.g., RXRA, LXR, and VDR), and may contribute to repress the expression of genes controlled through the circadian rhythm pathway [170].
Inflammatory pathways are critically involved in cancer and a transcriptional link between cell differentiation and inflammation is increasingly recognized [171]. Several families of TFs involved in inflammation appeared in our analysis, including Activator protein 1 (AP-1), Interferon response factors (IRFs), Nuclear Factors of Activated T-cells (NFATs), and Signal Transducers and Activators of Transcription (STATs) (Fig. 2, Table 2).
AP-1 includes a ubiquitous type of homo- or heterodimeric complexes composed of members of four gene families: JUN, FOS, ATF, and MAF, all of which were represented in our analysis both in BLCA organoids and normal bladder (Table 1 and Supplementary Table 1). AP-1 signaling is activated in response to multiple stimuli (e.g., inflammation, stress, pathogens) and participates in proliferation, differentiation, and apoptosis [172]. The effects of AP-1 are highly context-dependent. In the urothelium, its activation by tobacco smoke results in abnormal differentiation [173]. AP-1 cooperates with several of the major BLCA TFs. Genomic regions bound by FOXA1, GATA3, and KDM6A are enriched in AP-1 motifs in BLCA cells [44, 56, 174, 175]. A recent epigenomic map of co-regulated enhancers and associated transcription factors identified AP-1, as well as SMAD2/3, NF-kB, and STAT3, as potential regulators of basal enhancers [176]. This is consistent with our findings of a significant up-regulation of AP-1 target genes in basal organoids (Supplementary Fig. 2).
Amplifications are the most common alterations in AP-1 genes in tumors. Highly frequent copy number gains of FOSL2/FRA2 significantly associate with advanced stage, high grade, and low disease-free survival in both MIBC and NMIBC patients [177].
IRFs activate or repress gene expression by binding to the interferon-stimulated response element. IRF6 and IRF3 are among the highest expressed TFs in basal-like BLCA organoids and normal bladder, respectively. IRF6 is involved in epithelial differentiation and cell cycle regulation, partly synergizing with SMAD4 in response to TGF-β [178, 179] and NOTCH signaling [180]. In NHU cells, IRF1 binds the promoter of differentiation-associated genes (e.g., UPK) and enables their expression upon PPARG activation [70].
STATs are expressed broadly in the bladder, in epithelial and non-epithelial cells, and mediate cellular immunity, proliferation, apoptosis, and differentiation. They activate gene expression upon phosphorylation by Janus kinases, emphasizing that post-translational modifications add an important layer of regulation that cannot be fully captured through the sole analysis of genomic data. In our analyses, STAT1, STAT3, and STAT6 appeared as relevant TFs for normal bladder and both basal and luminal tumors (Table 1). They are altered in 4%, 1.7%, and 2.9% of the TCGA-BLCA samples, respectively, mainly through missense mutations and amplifications. In this cohort, STAT1 was identified as a key regulator of an immune GRN [181, 182]. STAT1 expression positively correlates with levels of PD-L1 [183].
Several studies point to STAT3 as tumor-promoting. p-STAT3 levels are significantly higher in basal BLCA, particularly in SCC-like tumors [60, 184, 185]. Consistently, our differential gene expression analysis in luminal vs. basal organoids reveals a significant up-regulation of STAT targets in basal urothelial cells (data not shown). STAT3 is critical for the regulation of basal subtype-specific genes [56] and STAT3 regulon activity is significantly enriched in Ba/Sq tumors [23]. Increased expression of STAT3 targets predicts basal-type phenotype and associates with worse prognosis [186]. Furthermore, STAT3 signaling is activated in response to urological infections, which may increase the risk of BLCA, and is required for their resolution [187, 188]. Collectively, these data suggest that STAT signaling may play a pro-tumorigenic role in BLCA at early stages of tumor development.
Two transcriptional pathways that did not stand out in our analyses but that are relevant to differentiation and BLCA are sex hormones and the HOX developmental gene family.
Sex hormones. The gender bias associated with BLCA has long raised interest in the role of sex hormones. In MIBC, males are enriched in the LumP and neuroendocrine-like subtypes and females are enriched in the Ba/Sq-like subtype. Sex-specific differences in AR activity have been reported among the luminal-like subtypes, suggesting that luminal differentiation could be partly driven by AR signaling. The activity of the ER pathway has been associated with the luminal MIBC subtype [59].
Several studies have reported an inverse association between AR expression and tumor stage [189,190,191]. Moreover, AR co-regulators (e.g., NCOA1, NCOA2, NCOA3, and CREBBP) are enriched in BLCA [189]. Functional analysis suggests that high AR activity could increase the susceptibility of BLCA cells to carcinogens, as well as exert a direct pro-tumoral effect [192, 193]. Ar knockout mice of both sexes were completely protected from BBN-induced BLCA and androgen deprivation suppressed tumor growth in vivo [192].
The involvement of ER in BLCA is more complex. ERβ expression has been positively associated with stage and grade [194]. Regarding ERα, there is controversy on its association with tumor grade, stage, and outcome. Studies in knockout mice have reported opposing roles for ERα and ERβ, the former having a protective role and the latter promoting carcinogenesis [195, 196].
Developmental HOX genes. HOX genes are expressed upon urothelial differentiation and their levels are higher in the urethra than in the bladder urothelium. In our analyses, HOXA13 was found to be expressed at highest levels in normal bladder. In mice, Hoxa13 is expressed in the embryonic urinary tract and postnatally, and it is essential for the morphogenesis of the urogenital and digestive tracts [197]. Mutations in HOXA13 are associated with hand-foot-genital syndrome and urinary tract malformations [198].
In NMIBC, a transcriptional network of cell cycle dysregulation pointed to HOX gene activity [199]. Marzouka et al. reported an inverse association between HOXB and late cell-cycle gene expression in BLCA [200]. Approximately 20% of MIBC from the TCGA cohort harbor mutations and/or copy number changes in HOX genes, amplification being the most frequent alteration. Genome-wide methylation analyses revealed an association between gene copy number gains, high methylation, and low expression for HOX genes in MIBC [201]. This highlights the importance of DNA methylation for HOX gene regulation and points to a tumor suppressor-like activity of the HOX family. Consistently, Aine et al. reported a correlation between epigenetic inactivation of HOX genes and tumor differentiation [202]. In the TCGA cohort, expression of 24 HOX genes (e.g., HOXA13, HOXB1-8) was significantly higher in LumP than in Ba/Sq tumors (FDR < 0.05), whereas eight genes were significantly down-regulated (e.g., SHOX2, PHOX2A, HOXD10-13). This is consistent with the fact that luminal BLCA organoids express significantly higher levels of HOXB genes, the expression of which is activated upon retinoid-induced differentiation [203]. In contrast, expression of HOXD genes was reduced. Altogether, these findings provide support to the role of this gene family in urothelial differentiation and BLCA.
BLCA gene regulatory networks
Few studies have aimed at building BLCA GRNs, possibly due to the scarcity of functional genomics data. The CoRegNet (Co-regulatory network) tool integrates transcriptomic, gene copy number, and ChIP data [204, 205]. When applied to BLCA, CoRegNet confirmed the driver role of PPARG, FOXA1, and GATA3. This is consistent with another study combining chromatin accessibility and gene expression data to build a network of TFs modulating urothelial differentiation in vitro upon PPARG activation. This work also found that GATA3 and FOXA1 cooperate, driving expression of luminal genes, which were repressed by TP63 [54].
Champion et al. have built a reference GRN from NHU cells and analyzed perturbations thereof in BLCA [206]. They inferred GRNs for the TCGA molecular subtypes, computed a deregulation score for each target gene in each subtype, and identified deregulated TFs. This approach highlighted 108 and 137 deregulated TFs in Ba/Sq and LumP samples, respectively. PPARG and NOTCH4 were among the 10 most important TFs accounting for transcriptomic deregulation in LumP and Ba/Sq BLCA, consistent with previous knowledge. Liang et al. have used H3K27ac ChIP-Seq data to identify super-enhancer (SE)-regulated genes and reconstruct a network including SMAD3, ETS1, and HOXB2 as core TFs in BLCA cells [207].
More recently, Neyret-Kahn et al. integrated RNA-Seq with ChIP-Seq data of active (H3K27ac) and repressive (H3K27me3, H3K9me3) histone marks in human primary tumors and cultures of BLCA and primary NHU cells [176]. They found chromatin states distinguishing basal and luminal samples that were associated with molecular BLCA subtypes. By analyzing H3K27ac, they inferred subsets of cancer-specific and subtype-specific SE and created SE-regulated networks. They confirmed FOXA1 as a major driver, provided evidence indicating that it also suppresses inflammatory programs, and discovered ZBED2 as a novel basal-specific TF that dampens IFN responses through STAT2. This is consistent with the enrichment we find for ZBED2 in basal-like organoids. These findings support that cell identity and inflammatory programs are part of the same TF networks, as previously described in the pancreas [171].
To expand this knowledge, we have used PPARG, RXRA, FOXA1, and GATA3 ChIP-Seq data to assess their binding to the promoter, gene body, and intergenic regions of the genes coding for the BLCA TFs identified in our analysis (Fig. 4). RXRA and PPARG show the highest and the lowest number of targets in our network, respectively. However, ChIP-Seq for them was performed in basal but not in luminal BLCA cells, which could affect these findings. In RT4 cells, FOXA1 and GATA3 binding was enriched in gene bodies and intergenic regions, compared to promoters, consistent with their reported role at distal enhancers [56]. The paucity of available data calls for expanding functional genomics analyses to establish the landscape of transcriptional regulation mechanisms in BLCA.
Regulatory network of TFs involved in luminal BLCA. ChIP-Seq data for the core nodes of the BLCA and RXRA (in bold) unveil genomic positions in promoters and intergenic regions of luminal-enriched (purple), basal-enriched (teal) and common (beige) BLCA TF nodes. The network displayed was constructed using Cytoscape.
Transcriptional regulation of BLCA plasticity and tumor heterogeneity
Histological and genomic analyses of BLCA have provided insight into cellular heterogeneity and plasticity, two prominent contributors to tumor progression and therapy resistance. The underlying evidences include: (1) histological heterogeneity and morphological changes occur during the disease course, (2) subclonal mutations undergo clonal selection over time, and (3) tumors with distinct gene expression signatures cluster into molecular subtypes. However, bulk tissue analyses fail to capture the diversity of individual cells and subclones, cannot fully explain the molecular underpinnings of heterogeneity and plasticity, and fail to provide information on the spatial configuration of cell-cell interactions. Single-cell technologies and multiplex analyses - at the RNA or protein level - provide resolution at the individual or quasi-individual cell level, allowing discovery of novel cell populations or states following gene ontologies. Single cell RNA-Seq (scRNA-Seq) has been used to infer cell lineage plasticity, hierarchies, and developmental trajectories. Understanding intermediate states in tumor evolution is of particular interest as this can lead to discovering key TFs acting as drivers of this process.
To delineate BLCA plasticity and tumor heterogeneity, Sfakianos et al. performed scRNA-Seq in human MIBC and in murine BBN-induced tumors [208]. Immune cell and non-immune cells were sorted from BBN-treated urothelium and scRNA-Seq revealed the existence of multiple cell populations within the tumor compartment. Differential expression analysis between the clusters showed their correspondence with well-known transcriptomic subtypes. For example, distinct populations expressed high levels of luminal-type TFs (Pparg, Foxa1, and Gata3) while others expressed basal-type (Trp63) or EMT-like (Zeb1, Zeb2, Snai2, and Twist1) TFs. These tumors are, thus, composed by a mosaic of heterogeneous cell populations. When analyzing human samples, they further identified individual epithelial cells with gene expression patterns characteristic of luminal, basal, and EMT-like transcriptomic subtypes. Using patient-derived xenografts, human BLCA cell populations with basal and EMT-like subtypes that can undergo lineage plasticity and subtype switch were identified [208].
In another study, scRNA-Seq combined with scATAC-Seq were used to resolve cell types, cell states, and tumor heterogeneity in BLCA [209]. Cancer stem cell markers were enriched in basal cell clusters. Regulon analysis revealed two groups of BLCA stem cells: one expressing CHD2, SIN3A, YY1, and KDM5B and another group expressing EZH2, SOX15, ATF3, and KLF10. EZH2 is a component of the Polycomb Repressive Complex 2 responsible for the H3K27me3 mark associated with transcriptional repression. Depletion of EZH2 in BLCA cells compromised cell proliferation, colony formation, and migration. EZH2-null BLCA xenografts in nude mice had decreased size and reduced proliferation. Loss of EZH2 was associated with enhanced chromatin accessibility near up-regulated genes, including those involved in stemness, differentiation, and cell-adhesion. Conversely, CD44, NCAM1, CDH2, and VIM genes showed reduced accessibility. DNA footprinting analysis of open chromatin regions specific for EZH2 wild-type cells further identified the TCFL5, TCF4, and CEBPB motifs, suggesting that EZH2 regulates BLCA stemness through these TFs. Our analyses revealed that CEBPB expression was highly enriched in normal and cancer urothelial cells according to bulk and scRNA-Seq data. Altogether, these data suggest that EZH2 is important for the stemness phenotype of certain BLCA populations and is relevant to patient outcome [210].
We have also analyzed public scRNA-Seq data on eight BLCA tumor samples [27]. We identified three clusters of urothelial cells defined as “Luminal_FABP4+”, “Luminal_UPK+” and “Basal_KRT5+” (Fig. 5A, B). The basal cluster shows high expression of genes enriched in the Ba/Sq subtype signature [23], whereas the luminal clusters are enriched in the LumP gene signature [23]. TFs identified by our analysis (see Box 1) as “Basal-enriched” and “Luminal-enriched” are highly expressed in the basal and the luminal clusters, respectively (Fig. 5C–F). In contrast, “Common” TFs are expressed widely (Fig. 5G). The candidate TFs identified here display distinct expression patterns. For example, ELF3 is highly expressed in the “Luminal_UPK+” cluster but it is undetectable in the “Luminal_FABP4+” cluster, which shows higher levels of GATA3, PPARG, and TBX3 (Fig. 5E, F; Supplementary Fig. 3). These findings highlight the existing heterogeneity even within molecular subtypes, which may be governed by different transcriptional programs. Moreover, we find an enrichment of novel candidate TFs, such as HMGA2 or MYCL, in basal and luminal cells, respectively (Fig. 5E, F), further supporting the relevance of the proposed novel candidate TFs in urothelial differentiation.
A, B UMAP plots of urothelial cells disclose a cluster of basal cells enriched in KRT5 expression, a cluster of luminal cells enriched in UPK expression, and a cluster of luminal cells enriched in FABP4 expression (B). C UMAP plots depicting the activity of the Ba/Sq signature from Kamoun et al. [23] (left) and the “basal-enriched TF” gene set (right) from our analysis. D. UMAP plots depicting the activity of the Luminal papillary signature from Kamoun et al. [23] (left) and the “luminal-enriched TF” gene set (right) from our analysis. E UMAP plots showing the expression of TP63 and selected basal-enriched novel candidate TFs (ARNTL2 and HMGA2). F UMAP plots showing the expression of ELF3, GATA3, FOXA1 and selected luminal-enriched novel candidate TFs (SREBF1 and MYCL): differential expression among the luminal clusters. G UMAP plots depicting expression activity of the “Common TFs” gene set and expression of KLF5 and NFE2L2 and novel candidate TF YY1.
TF-based opportunities for improved BLCA management
As discussed above, TFs can display both oncogenic or tumor suppressor function. Rescue of loss-of-function defects is more challenging than pharmacological inhibition and, with the exception of nuclear receptors, most oncogenic TFs have classically been considered undruggable. MYC is a prototypic TF the inhibition of which has long been sought because of its central role in oncogenesis. A variety of strategies to inhibit MYC have been applied [211] and a cell-penetrating miniprotein that acts as a dominant negative for MYC has antitumor activity in vitro and in vivo preclinical models [212]. A clinical trial in patients with advanced solid tumors is ongoing (https://clinicaltrials.gov/ct2/show/NCT04808362).
Several of the TFs discussed here are attractive pharmacological targets in BLCA, including PPARG and RXRA. The oncogenic activity of the PPARG/RXRA heterodimer in BLCA resembles that of ER/PR in breast cancer and AR in prostate cancer. In homeostatic conditions, these TF play an important role in the respective tissues, including the promotion of cell differentiation. As discussed above, PPARG can also promote tumorigenesis with both a direct role at the tumor cell level by controlling transcription and, indirectly, through effects on the immune TME [38]. We are not aware of any studies using PPARG antagonists in BLCA, possibly because of the broad metabolic functions of this target. On the other hand, the activating point mutations in RXRA and the recently described AHR deletion mutations, both present in a small proportion of BLCA, may provide therapeutic selectivity.
Regulon analyses of RNA-Seq data from NMIBC and MIBC suggest that sex hormone receptors may play a significant role in BLCA. In NMIBC, the ERβ regulon is up-regulated in Class 2a tumors whereas the ERα and PR regulons are up-regulated in a subset of Class 2b tumors, and the AR regulon is broadly up-regulated in class 3, FGFR3-mutant, tumors [213]. Consistent with these findings, increased activity of the AR regulon has been found in the three MIBC consensus papillary subtypes. The ERβ regulon is also activated in luminal tumors whereas the PR and ERα regulons are up-regulated in the stromal-rich and Ba/Sq-like subtypes [23]. However, these bioinformatics analyses need to be followed by rigorous experimental validation.
In the last few years, there has been renewed interest in TFs as drug targets, most notably thanks to the development of “degraders”, small molecules that induce target protein degradation by the proteasome [214]. Proteolysis targeting chimerics (PROTAC) are small heterobifunctional molecules that can bind a protein of interest and an E3 ubiquitin ligase, leading to target degradation. PROTACs may, but do not need to, involve the active or ligand-binding site of a protein but they must have target protein binding selectivity (target must be ligandable). PROTAC-type degraders have been developed to a variety of oncogenic molecules including fusion proteins (e.g., BCR-ABL), kinases (e.g., BTK, CDK6), and TFs (e.g., AR and ER) [215]. Molecular glues are degraders that act as protein-protein scaffolds, involve highly cooperative E3-neosubstrate interactions, and are discovered agnostically through molecular screening. Natural examples of such molecules are thalidomide and the plant signaling hormones auxin and jasmonate. Several degrader molecules targeting TF, namely the AR for prostate cancer and the ER for breast cancer, have entered clinical trials, most of them being of the molecular glue type. The degrader strategy may thus be more broadly applied to TFs acting through gain-of-function.
Concluding remarks
The evidence presented here underlies the notion that a profound dysregulation of transcriptional programs involved in cell differentiation contributes to BLCA. Furthermore, the GRNs involved play a key role in tumor cell heterogeneity and in the plasticity associated with tumor evolution and therapeutic resistance. Several of the key TFs participating in these processes display features that are amenable to therapeutic targeting. The high frequency of alterations in genes coding for proteins involved in chromatin function suggests that an in-depth analysis of epigenetic changes is mandatory. Acquiring a more profound mechanistic knowledge of both GRN and epigenetic regulation should provide improved opportunities to understand heterogeneity and leverage cell plasticity in the stratified management of BLCA.
References
Hicks RM. The mammalian urinary bladder: an accommodating organ. Biol Rev Camb Philos Soc. 1975;50:215–46.
Truschel ST, Clayton DR, Beckel JM, Yabes JG, Yao Y, Wolf-Johnston A, et al. Age-related endolysosome dysfunction in the rat urothelium. PLoS One. 2018;13:e0198817.
Jost SP, Potten CS. Urothelial proliferation in growing mice. Cell Tissue Kinet. 1986;19:155–60.
Wang J, Batourina E, Schneider K, Souza S, Swayne T, Liu C, et al. Polyploid superficial cells that maintain the urothelial barrier are produced via incomplete cytokinesis and endoreplication. Cell Rep. 2019;25:464–.e4.
Hudoklin S, Jezernik K, Neumüller J, Pavelka M, Romih R. Electron tomography of fusiform vesicles and their organization in urothelial cells. PLoS One. 2012;7:e32935.
Varley CL, Garthwaite MAE, Cross W, Hinley J, Trejdosiewicz LK, Southgate J. PPARgamma-regulated tight junction development during human urothelial cytodifferentiation. J Cell Physiol. 2006;208:407–17.
Harnden P, Eardley I, Joyce AD, Southgate J. Cytokeratin 20 as an objective marker of urothelial dysplasia. Br J Urol. 1996;78:870–5.
Gandhi D, Molotkov A, Batourina E, Schneider K, Dan H, Reiley M, et al. Retinoid signaling in progenitors controls specification and regeneration of the urothelium. Dev Cell. 2013;26:469–82.
Papafotiou G, Paraskevopoulou V, Vasilaki E, Kanaki Z, Paschalidis N, Klinakis A. KRT14 marks a subpopulation of bladder basal cells with pivotal role in regeneration and tumorigenesis. Nat Commun. 2016;7:11914.
Colopy SA, Bjorling DE, Mulligan WA, Bushman W. A population of progenitor cells in the basal and intermediate layers of the murine bladder urothelium contributes to urothelial development and regeneration. Dev Dyn. 2014;243:988–98.
Jost SP. Cell cycle of normal bladder urothelium in developing and adult mice. Virchows Arch B Cell Pathol. 1989;57:27–36.
Shin K, Lee J, Guo N, Kim J, Lim A, Qu L, et al. Hedgehog/Wnt feedback supports regenerative proliferation of epithelial stem cells in bladder. Nature. 2011;472:110–4.
Hurst CD, Alder O, Platt FM, Droop A, Stead LF, Burns JE, et al. Genomic subtypes of non-invasive bladder cancer with distinct metabolic profile and female gender bias in KDM6A mutation frequency. Cancer Cell. 2017;32:701–.e7.
Robertson AG, Kim J, Al-Ahmadie H, Bellmunt J, Guo G, Cherniack AD, et al. Comprehensive molecular characterization of muscle-invasive bladder. Cancer Cell. 2017;171:540–.e25.
Wullweber A, Strick R, Lange F, Sikic D, Taubert H, Wach S, et al. Bladder tumor subtype commitment occurs in carcinoma in situ driven by key signaling pathways including ECM remodeling. Cancer Res. 2021;81:1552–66.
Bondaruk J, Jaksik R, Wang Z, Cogdell D, Lee S, Chen Y, et al. The origin of bladder cancer from mucosal field effects. iScience. 2022;25:104551.
Majewski T, Yao H, Bondaruk J, Chung W, Lee S, Lee JG, et al. Whole-organ genomic characterization of mucosal field effects initiating bladder carcinogenesis. Cell Rep. 2019;26:2241–.e4.
Lawson ARJ, Abascal F, Coorens THH, Hooks Y, O’Neill L, Latimer C, et al. Extensive heterogeneity in somatic mutation and selection in the human bladder. Science. 2020;370:75–82.
López-Knowles E, Hernández S, Malats N, Kogevinas M, Lloreta J, Carrato A, et al. PIK3CA mutations are an early genetic alteration associated with FGFR3 mutations in superficial papillary bladder tumors. Cancer Res. 2006;66:7401–4.
Taylor CF, Platt FM, Hurst CD, Thygesen HH, Knowles MA. Frequent inactivating mutations of STAG2 in bladder cancer are associated with low tumour grade and stage and inversely related to chromosomal copy number changes. Hum Mol Genet. 2014;23:1964–74.
Hedegaard J, Lamy P, Nordentoft I, Algaba F, Høyer S, Ulhøi BP, et al. Comprehensive transcriptional analysis of early-stage urothelial carcinoma. Cancer Cell. 2016;30:27–42.
Hartmann A, Schlake G, Zaak D, Hungerhuber E, Hofstetter A, Hofstaedter F, et al. Occurrence of chromosome 9 and p53 alterations in multifocal dysplasia and carcinoma in situ of human urinary bladder. Cancer Res. 2002;62:809–18.
Kamoun A, de Reyniès A, Allory Y, Sjödahl G, Robertson AG, Seiler R, et al. A consensus molecular classification of muscle-invasive bladder cancer. Eur Urol. 2020;77:420–33.
Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, et al. The human transcription factors. Cell. 2018;172:650–65.
Lee SH, Hu W, Matulay JT, Silva MV, Owczarek TB, Kim K, et al. Tumor evolution and drug response in patient-derived organoid models of bladder. Cancer Cell. 2018;173:515–.e17.
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74.
Chen Z, Zhou L, Liu L, Hou Y, Xiong M, Yang Y, et al. Single-cell RNA sequencing highlights the role of inflammatory cancer-associated fibroblasts in bladder urothelial carcinoma. Nat Commun. 2020;11:5077.
Evans RM, Mangelsdorf DJ. Nuclear receptors, RXR, and the big bang. Cell. 2014;157:255–66.
Lehmann JM, Moore LB, Smith-Oliver TA, Wilkison WO, Willson TM, Kliewer SA. An antidiabetic thiazolidinedione is a high-affinity ligand for peroxisome proliferator-activated receptor gamma (PPAR gamma). J Biol Chem. 1995;270:12953–6.
Mamtani R, Haynes K, Bilker WB, Vaughn DJ, Strom BL, Glanz K, et al. Association between longer therapy with thiazolidinediones and risk of bladder cancer: a cohort study. J Natl Cancer Inst. 2012;104:1411–21.
Varley CL, Stahlschmidt J, Lee W-C, Holder J, Diggle C, Selby PJ, et al. Role of PPARgamma and EGFR signalling in the urothelial terminal differentiation programme. J Cell Sci. 2004;117:2029–36.
Santos CP, Lapi E, Martínez de Villarreal J, Álvaro-Espinosa L, Fernández-Barral A, Barbáchano A, et al. Urothelial organoids originating from Cd49fhigh mouse stem cells display Notch-dependent differentiation capacity. Nat Commun. 2019;10:4407.
Suzuki K, Koyanagi-Aoi M, Uehara K, Hinata N, Fujisawa M, Aoi T. Directed differentiation of human induced pluripotent stem cells into mature stratified bladder urothelium. Sci Rep. 2019;9:10506.
Weiss RM, Guo S, Shan A, Shi H, Romano R-A, Sinha S, et al. Brg1 determines urothelial cell fate during ureter development. J Am Soc Nephrol. 2013;24:618–26.
Liu C, Tate T, Batourina E, Truschel ST, Potter S, Adam M, et al. Pparg promotes differentiation and regulates mitochondrial gene expression in bladder epithelial cells. Nat Commun. 2019;10:4589.
Liang F-X, Bosland MC, Huang H, Romih R, Baptiste S, Deng F-M, et al. Cellular basis of urothelial squamous metaplasia: roles of lineage heterogeneity and cell replacement. J Cell Biol. 2005;171:835–44.
Pascual G, Fong AL, Ogawa S, Gamliel A, Li AC, Perissi V, et al. A SUMOylation-dependent pathway mediates transrepression of inflammatory response genes by PPAR-gamma. Nature. 2005;437:759–63.
Tate T, Xiang T, Wobker SE, Zhou M, Chen X, Kim H, et al. Pparg signaling controls bladder cancer subtype and immune exclusion. Nat Commun. 2021;12:6160.
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature. 2014;507:315–22.
Rochel N, Krucker C, Coutos-Thévenot L, Osz J, Zhang R, Guyon E, et al. Recurrent activating mutations of PPARγ associated with luminal bladder tumors. Nat Commun. 2019;10:253.
Halstead AM, Kapadia CD, Bolzenius J, Chu CE, Schriefer A, Wartman LD, et al. Bladder-cancer-associated mutations in RXRA activate peroxisome proliferator-activated receptors to drive urothelial proliferation. eLife. 2017;6:e30862.
Korpal M, Puyang X, Jeremy Wu Z, Seiler R, Furman C, Oo HZ, et al. Evasion of immunosurveillance by genomic alterations of PPARγ/RXRα in bladder cancer. Nat Commun. 2017;8:103.
Coutos-Thévenot L, Beji S, Neyret-Kahn H, Pippo Q, Fontugne J, Osz J, et al. PPARγ is a tumor suppressor in basal bladder tumors offering new potential therapeutic opportunities. BioRxiv. 2019; https://doi.org/10.1101/868190.
Warrick JI, Walter V, Yamashita H, Chung E, Shuman L, Amponsa VO, et al. FOXA1, GATA3 and ppar? cooperate to drive luminal subtype in bladder cancer: a molecular analysis of established human cell lines. Sci Rep. 2016;6:38531.
Tortora D, Roberts ME, Kumar G, Kotapalli SS, Ritch E, Scurll JM, et al. A genome-wide CRISPR screen maps endogenous regulators of PPARG gene expression in bladder cancer. iScience. 2023;26:106525.
Goldstein JT, Berger AC, Shih J, Duke FF, Furst L, Kwiatkowski DJ, et al. Genomic activation of PPARG reveals a candidate therapeutic axis in bladder cancer. Cancer Res. 2017;77:6987–98.
Sanchez DJ, Missiaen R, Skuli N, Steger DJ, Simon MC. Cell-intrinsic tumorigenic functions of PPARγ in bladder urothelial carcinoma. Mol Cancer Res. 2021;19:598–611.
Ko LJ, Engel JD. DNA-binding specificities of the GATA transcription factor family. Mol Cell Biol. 1993;13:4011–22.
van der Ven AT, Connaughton DM, Ityel H, Mann N, Nakayama M, Chen J, et al. Whole-exome sequencing identifies causative mutations in families with congenital anomalies of the kidney and urinary tract. J Am Soc Nephrol. 2018;29:2348–61.
Kouros-Mehr H, Slorach EM, Sternlicht MD, Werb Z. GATA-3 maintains the differentiation of the luminal cell fate in the mammary gland. Cell. 2006;127:1041–55.
Higgins JPT, Kaygusuz G, Wang L, Montgomery K, Mason V, Zhu SX, et al. Placental S100 (S100P) and GATA3: markers for transitional epithelium and urothelial carcinoma discovered by complementary DNA microarray. Am J Surg Pathol. 2007;31:673–80.
Ainoya K, Moriguchi T, Ohmori S, Souma T, Takai J, Morita M, et al. UG4 enhancer-driven GATA-2 and bone morphogenetic protein 4 complementation remedies the CAKUT phenotype in Gata2 hypomorphic mutant mice. Mol Cell Biol. 2012;32:2312–22.
Hoshino T, Shimizu R, Ohmori S, Nagano M, Pan X, Ohneda O, et al. Reduced BMP4 abundance in Gata2 hypomorphic mutant mice result in uropathies resembling human CAKUT. Genes Cells. 2008;13:159–70.
Fishwick C, Higgins J, Percival-Alwyn L, Hustler A, Pearson J, Bastkowski S, et al. Heterarchy of transcription factors driving basal and luminal cell phenotypes in human urothelium. Cell Death Differ. 2017;24:809–18.
Wang C, Yang S, Jin L, Dai G, Yao Q, Xiang H, et al. Biological and clinical significance of GATA3 detected from TCGA database and FFPE sample in bladder cancer patients. Onco Targets Ther. 2020;13:945–58.
Iyyanki T, Zhang B, Wang Q, Hou Y, Jin Q, Xu J, et al. Subtype-associated epigenomic landscape and 3D genome structure in bladder cancer. Genome Biol. 2021;22:105.
Lerner SP, McConkey DJ, Hoadley KA, Chan KS, Kim WY, Radvanyi F, et al. Bladder cancer molecular taxonomy: summary from a consensus meeting. Bladder Cancer. 2016;2:37–47.
Miyamoto H, Izumi K, Yao JL, Li Y, Yang Q, McMahon LA, et al. GATA binding protein 3 is down-regulated in bladder cancer yet strong expression is an independent predictor of poor prognosis in invasive tumor. Hum Pathol. 2012;43:2033–40.
Choi W, Porten S, Kim S, Willis D, Plimack ER, Hoffman-Censits J, et al. Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell. 2014;25:152–65.
Eriksson P, Aine M, Veerla S, Liedberg F, Sjödahl G, Höglund M. Molecular subtypes of urothelial carcinoma are defined by specific gene regulatory systems. BMC Med Genom. 2015;8:25.
van Kessel KEM, van der Keur KA, Dyrskjøt L, Algaba F, Welvaart NYC, Beukers W, et al. Molecular markers increase precision of the European Association of Urology non-muscle-invasive bladder cancer progression risk groups. Clin Cancer Res. 2018;24:1586–93.
Clark KL, Halay ED, Lai E, Burley SK. Co-crystal structure of the HNF-3/fork head DNA-recognition motif resembles histone H5. Nature. 1993;364:412–20.
Cirillo LA, McPherson CE, Bossard P, Stevens K, Cherian S, Shim EY, et al. Binding of the winged-helix transcription factor HNF3 to a linker histone site on the nucleosome. EMBO J. 1998;17:244–54.
Cirillo LA, Lin FR, Cuesta I, Friedman D, Jarnik M, Zaret KS. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol Cell. 2002;9:279–89.
Li Z, Tuteja G, Schug J, Kaestner KH. Foxa1 and Foxa2 are essential for sexual dimorphism in liver cancer. Cell. 2012;148:72–83.
Tsuchiya H, da Costa K-A, Lee S, Renga B, Jaeschke H, Yang Z, et al. Interactions between nuclear receptor SHP and FOXA1 maintain oscillatory homocysteine homeostasis in mice. Gastroenterology. 2015;148:1012–.e14.
Belikov S, Astrand C, Wrange O. FoxA1 binding directs chromatin structure and the functional response of a glucocorticoid receptor-regulated promoter. Mol Cell Biol. 2009;29:5413–25.
Besnard V, Wert SE, Hull WM, Whitsett JA. Immunohistochemical localization of Foxa1 and Foxa2 in mouse embryos and adult tissues. Gene Expr Patterns. 2004;5:193–208.
Oottamasathien S, Wang Y, Williams K, Franco OE, Wills ML, Thomas JC, et al. Directed differentiation of embryonic stem cells into bladder tissue. Dev Biol. 2007;304:556–66.
Varley CL, Bacon EJ, Holder JC, Southgate J. FOXA1 and IRF-1 intermediary transcriptional regulators of PPARgamma-induced urothelial cytodifferentiation. Cell Death Differ. 2009;16:103–14.
DeGraff DJ, Clark PE, Cates JM, Yamashita H, Robinson VL, Yu X, et al. Loss of the urothelial differentiation marker FOXA1 is associated with high grade, late stage bladder cancer and increased tumor proliferation. PLoS One. 2012;7:e36669.
Guo Y, Yuan X, Li K, Dai M, Zhang L, Wu Y, et al. GABPA is a master regulator of luminal identity and restrains aggressive diseases in bladder cancer. Cell Death Differ. 2020;27:1862–77.
Reddy OL, Cates JM, Gellert LL, Crist HS, Yang Z, Yamashita H, et al. Loss of FOXA1 drives sexually dimorphic changes in urothelial differentiation and is an independent predictor of poor prognosis in bladder cancer. Am J Pathol. 2015;185:1385–95.
Osei-Amponsa V, Buckwalter JM, Shuman L, Zheng Z, Yamashita H, Walter V, et al. Hypermethylation of FOXA1 and allelic loss of PTEN drive squamous differentiation and promote heterogeneity in bladder cancer. Oncogene. 2020;39:1302–17.
Bernardo GM, Keri RA. FOXA1: a transcription factor with parallel functions in development and cancer. Biosci Rep. 2012;32:113–30.
Robinson D, Van Allen EM, Wu Y-M, Schultz N, Lonigro RJ, Mosquera J-M, et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015;161:1215–28.
Ciriello G, Gatza ML, Beck AH, Wilkerson MD, Rhie SK, Pastore A, et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell. 2015;163:506–19.
Sikic D, Eckstein M, Wirtz RM, Jarczyk J, Worst TS, Porubsky S, et al. FOXA1 gene expression for defining molecular subtypes of muscle-invasive bladder cancer after radical cystectomy. J Clin Med. 2020;9:994.
Oettgen P, Alani RM, Barcinski MA, Brown L, Akbarali Y, Boltax J, et al. Isolation and characterization of a novel epithelium-specific transcription factor, ESE-1, a member of the ets family. Mol Cell Biol. 1997;17:4419–33.
Ng AY-N, Waring P, Ristevski S, Wang C, Wilson T, Pritchard M, et al. Inactivation of the transcription factor Elf3 in mice results in dysmorphogenesis and altered differentiation of intestinal epithelium. Gastroenterology. 2002;122:1455–66.
Böck M, Hinley J, Schmitt C, Wahlicht T, Kramer S, Southgate J. Identification of ELF3 as an early transcriptional regulator of human urothelium. Dev Biol. 2014;386:321–30.
Na L, Wang Z, Bai Y, Sun Y, Dong D, Wang W, et al. WNT7B represses epithelial-mesenchymal transition and stem-like properties in bladder urothelial carcinoma. Biochim Biophys Acta Mol Basis Dis. 2022;1868:166271.
Gondkar K, Patel K, Krishnappa S, Patil A, Nair B, Sundaram GM, et al. E74 like ETS transcription factor 3 (ELF3) is a negative regulator of epithelial- mesenchymal transition in bladder carcinoma. Cancer Biomark. 2019;25:223–32.
Ting SB, Wilanowski T, Cerruti L, Zhao L-L, Cunningham JM, Jane SM. The identification and characterization of human Sister-of-Mammalian Grainyhead (SOM) expands the grainyhead-like family of developmental transcription factors. Biochem J. 2003;370:953–62.
Traylor-Knowles N, Hansen U, Dubuc TQ, Martindale MQ, Kaufman L, Finnerty JR. The evolutionary diversification of LSF and Grainyhead transcription factors preceded the radiation of basal animal lineages. BMC Evol Biol. 2010;10:101.
Yu Z, Mannik J, Soto A, Lin KK, Andersen B. The epidermal differentiation-associated Grainyhead gene Get1/Grhl3 also regulates urothelial differentiation. EMBO J. 2009;28:1890–903.
Osborn SL, Thangappan R, Luria A, Lee JH, Nolta J, Kurzrock EA. Induction of human embryonic and induced pluripotent stem cells into urothelium. Stem Cells Transl Med. 2014;3:610–9.
Wezel F, Lustig J, Azoitei A, Liu J, Meessen S, Najjar G, et al. Grainyhead-like 3 influences migration and invasion of urothelial carcinoma cells. Int J Mol Sci. 2021;22:2959.
Ghioni P, Bolognese F, Duijf PHG, Van Bokhoven H, Mantovani R, Guerrini L. Complex transcriptional effects of p63 isoforms: identification of novel activation and repression domains. Mol Cell Biol. 2002;22:8659–68.
Augustin M, Bamberger C, Paul D, Schmale H. Cloning and chromosomal mapping of the human p53-related KET gene to chromosome 3q27 and its murine homolog Ket to mouse chromosome 16. Mamm Genome. 1998;9:899–902.
Osada M, Ohba M, Kawahara C, Ishioka C, Kanamaru R, Katoh I, et al. Cloning and functional analysis of human p51, which structurally and functionally resembles p53. Nat Med. 1998;4:839–43.
Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dötsch V, et al. p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell. 1998;2:305–16.
Dohn M, Zhang S, Chen X. p63alpha and DeltaNp63alpha can induce cell cycle arrest and apoptosis and differentially regulate p53 target genes. Oncogene. 2001;20:3193–205.
Lohrum MA, Vousden KH. Regulation and function of the p53-related proteins: same family, different rules. Trends Cell Biol. 2000;10:197–202.
Koster MI, Kim S, Mills AA, DeMayo FJ, Roop DR. p63 is the molecular switch for initiation of an epithelial stratification program. Genes Dev. 2004;18:126–31.
Guo X, Keyes WM, Papazoglu C, Zuber J, Li W, Lowe SW, et al. TAp63 induces senescence and suppresses tumorigenesis in vivo. Nat Cell Biol. 2009;11:1451–7.
Helton ES, Zhu J, Chen X. The unique NH2-terminally deleted (DeltaN) residues, the PXXP motif, and the PPXY motif are required for the transcriptional activity of the DeltaN variant of p63. J Biol Chem. 2006;281:2533–42.
Kouwenhoven EN, Oti M, Niehues H, van Heeringen SJ, Schalkwijk J, Stunnenberg HG, et al. Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation. EMBO Rep. 2015;16:863–78.
Somerville TDD, Xu Y, Miyabayashi K, Tiriac H, Cleary CR, Maia-Silva D, et al. TP63-mediated enhancer reprogramming drives the squamous subtype of pancreatic ductal adenocarcinoma. Cell Rep. 2018;25:1741–.e7.
Yang A, McKeon F. P63 and P73: P53 mimics, menaces and more. Nat Rev Mol Cell Biol. 2000;1:199–207.
Karni-Schmidt O, Castillo-Martin M, Shen TH, Gladoun N, Domingo-Domenech J, Sanchez-Carbayo M, et al. Distinct expression profiles of p63 variants during urothelial development and bladder cancer progression. Am J Pathol. 2011;178:1350–60.
Mills AA, Zheng B, Wang XJ, Vogel H, Roop DR, Bradley A. p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature. 1999;398:708–13.
Yang A, Schweitzer R, Sun D, Kaghad M, Walker N, Bronson RT, et al. p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature. 1999;398:714–8.
Paris M, Rouleau M, Pucéat M, Aberdam D. Regulation of skin aging and heart development by TAp63. Cell Death Differ. 2012;19:186–93.
Urist MJ, Di Como CJ, Lu M-L, Charytonowicz E, Verbel D, Crum CP, et al. Loss of p63 expression is associated with tumor progression in bladder cancer. Am J Pathol. 2002;161:1199–206.
Signoretti S, Pires MM, Lindauer M, Horner JW, Grisanzio C, Dhar S, et al. p63 regulates commitment to the prostate cell lineage. Proc Natl Acad Sci USA. 2005;102:11355–60.
Cheng W, Jacobs WB, Zhang JJR, Moro A, Park J-H, Kushida M, et al. DeltaNp63 plays an anti-apoptotic role in ventral bladder development. Development. 2006;133:4783–92.
Celli J, Duijf P, Hamel BC, Bamshad M, Kramer B, Smits AP, et al. Heterozygous germline mutations in the p53 homolog p63 are the cause of EEC syndrome. Cell. 1999;99:143–53.
van Bokhoven H, Hamel BC, Bamshad M, Sangiorgi E, Gurrieri F, Duijf PH, et al. p63 Gene mutations in eec syndrome, limb-mammary syndrome, and isolated split hand-split foot malformation suggest a genotype-phenotype correlation. Am J Hum Genet. 2001;69:481–92.
Duijf PHG, Vanmolkot KRJ, Propping P, Friedl W, Krieger E, McKeon F, et al. Gain-of-function mutation in ADULT syndrome reveals the presence of a second transactivation domain in p63. Hum Mol Genet. 2002;11:799–804.
McGrath JA, Duijf PH, Doetsch V, Irvine AD, de Waal R, Vanmolkot KR, et al. Hay-Wells syndrome is caused by heterozygous missense mutations in the SAM domain of p63. Hum Mol Genet. 2001;10:221–9.
Kloesch B, Ionasz V, Paliwal S, Hruschka N, Martinez de Villarreal J, Öllinger R, et al. A GATA6-centred gene regulatory network involving HNFs and ΔNp63 controls plasticity and immune escape in pancreatic cancer. Gut. 2022;71:766–77.
Bankhead A, McMaster T, Wang Y, Boonstra PS, Palmbos PL. TP63 isoform expression is linked with distinct clinical outcomes in cancer. EBioMedicine. 2020;51:102561.
Papadimitriou M-A, Avgeris M, Levis PK, Tokas T, Stravodimos K, Scorilas A. ΔNp63 transcript loss in bladder cancer constitutes an independent molecular predictor of TaT1 patients post-treatment relapse and progression. J Cancer Res Clin Oncol. 2019;145:3075–87.
Guo CC, Majewski T, Zhang L, Yao H, Bondaruk J, Wang Y, et al. Dysregulation of EMT drives the progression to clinically aggressive sarcomatoid bladder cancer. Cell Rep. 2019;27:1781–.e4.
Eckert D, Buhl S, Weber S, Jäger R, Schorle H. The AP-2 family of transcription factors. Genome Biol. 2005;6:246.
Sinha S, Fuchs E. Identification and dissection of an enhancer controlling epithelial gene expression in skin. Proc Natl Acad Sci USA. 2001;98:2455–60.
Kaufman CK, Sinha S, Bolotin D, Fan J, Fuchs E. Dissection of a complex enhancer element: maintenance of keratinocyte specificity but loss of differentiation specificity. Mol Cell Biol. 2002;22:4293–308.
Leask A, Byrne C, Fuchs E. Transcription factor AP2 and its role in epidermal-specific gene expression. Proc Natl Acad Sci USA. 1991;88:7948–52.
McDade SS, Henry AE, Pivato GP, Kozarewa I, Mitsopoulos C, Fenwick K, et al. Genome-wide analysis of p63 binding sites identifies AP-2 factors as co-regulators of epidermal differentiation. Nucleic Acids Res. 2012;40:7190–206.
Yamashita H, Kawasawa YI, Shuman L, Zheng Z, Tran T, Walter V, et al. Repression of transcription factor AP-2 alpha by PPARγ reveals a novel transcriptional circuit in basal-squamous bladder cancer. Oncogenesis. 2019;8:69.
Davis AC, Wims M, Spotts GD, Hann SR, Bradley A. A null c-myc mutation causes lethality before 10.5 days of gestation in homozygotes and reduced fertility in heterozygous female mice. Genes Dev. 1993;7:671–82.
Watters AD, Latif Z, Forsyth A, Dunn I, Underwood MA, Grigor KM, et al. Genetic aberrations of c-myc and CCND1 in the development of invasive bladder cancer. Br J Cancer. 2002;87:654–8.
Li Y, Liu H, Lai C, Du X, Su Z, Gao S. The Lin28/let-7a/c-Myc pathway plays a role in non-muscle invasive bladder cancer. Cell Tissue Res. 2013;354:533–41.
Mahe M, Dufour F, Neyret-Kahn H, Moreno-Vega A, Beraud C, Shi M, et al. An FGFR3/MYC positive feedback loop provides new opportunities for targeted therapies in bladder cancers. EMBO Mol Med. 2018;10:e8163.
Zhuang C, Ma Q, Zhuang C, Ye J, Zhang F, Gui Y. LncRNA GClnc1 promotes proliferation and invasion of bladder cancer through activation of MYC. FASEB J. 2019;33:11045–59.
Jiang G, Huang C, Liao X, Li J, Wu X-R, Zeng F, et al. The RING domain in the anti-apoptotic protein XIAP stabilizes c-Myc protein and preserves anchorage-independent growth of bladder cancer cells. J Biol Chem. 2019;294:5935–44.
Robertson AG, Groeneveld CS, Jordan B, Lin X, McLaughlin KA, Das A, et al. Identification of differential tumor subtypes of T1 bladder cancer. Eur Urol. 2020;78:533–7.
Fontugne J, Wong J, Cabel L, Neyret-Kahn H, Karboul N, Maillé P, et al. Progression-associated molecular changes in basal/squamous and sarcomatoid bladder carcinogenesis. J Pathol. 2023;259:455–67.
Marquis L, Tran M, Choi W, Lee I-L, Huszar D, Siefker-Radtke A, et al. p63 expression correlates with sensitivity to the Eg5 inhibitor ZD4877 in bladder cancer cells. Cancer Biol Ther. 2012;13:477–86.
Greife A, Jankowiak S, Steinbring J, Nikpour P, Niegisch G, Hoffmann MJ, et al. Canonical Notch signalling is inactive in urothelial carcinoma. BMC Cancer. 2014;14:628.
Paraskevopoulou V, Bonis V, Dionellis VS, Paschalidis N, Melissa P, Chavdoula E, et al. Notch controls urothelial integrity in the mouse bladder. JCI Insight. 2020;5:e133232.
Rampias T, Vgenopoulou P, Avgeris M, Polyzos A, Stravodimos K, Valavanis C, et al. A new tumor suppressor role for the Notch pathway in bladder cancer. Nat Med. 2014;20:1199–205.
Maraver A, Fernandez-Marcos PJ, Cash TP, Mendez-Pertuz M, Dueñas M, Maietta P, et al. NOTCH pathway inactivation promotes bladder cancer progression. J Clin Investig. 2015;125:824–30.
Hayashi T, Gust KM, Wyatt AW, Goriki A, Jäger W, Awrey S, et al. Not all NOTCH is created equal: the oncogenic role of NOTCH2 in bladder cancer and its implications for targeted therapy. Clin Cancer Res. 2016;22:2981–92.
Zhang L, Sha J, Yang G, Huang X, Bo J, Huang Y. Activation of Notch pathway is linked with epithelial-mesenchymal transition in prostate cancer cells. Cell Cycle. 2017;16:999–1007.
Watabe T, Yoshida K, Shindoh M, Kaya M, Fujikawa K, Sato H, et al. The Ets-1 and Ets-2 transcription factors activate the promoters for invasion-associated urokinase and collagenase genes in response to epidermal growth factor. Int J Cancer. 1998;77:128–37.
Westermarck J, Seth A, Kähäri VM. Differential regulation of interstitial collagenase (MMP-1) gene expression by ETS transcription factors. Oncogene. 1997;14:2651–60.
Hashiya N, Jo N, Aoki M, Matsumoto K, Nakamura T, Sato Y, et al. In vivo evidence of angiogenesis induced by transcription factor Ets-1: Ets-1 is located upstream of angiogenesis cascade. Circulation. 2004;109:3035–41.
Sari A, Calli A, Gorgel SN, Altinboga AA, Kara C, Dincel C, et al. Immunohistochemical determination of ETS-1 oncoprotein expression in urothelial carcinomas of the urinary bladder. Appl Immunohistochem Mol Morphol. 2012;20:153–8.
Liu L, Liu Y, Zhang X, Chen M, Wu H, Lin M, et al. Inhibiting cell migration and cell invasion by silencing the transcription factor ETS-1 in human bladder cancer. Oncotarget. 2016;7:25125–34.
Lin S-R, Yeh H-C, Wang W-J, Ke H-L, Lin H-H, Hsu W-C, et al. MiR-193b mediates CEBPD-induced cisplatin sensitization through targeting ETS1 and cyclin D1 in human urothelial carcinoma cells. J Cell Biochem. 2017;118:1563–73.
Shin S-S, Park S-S, Hwang B, Kim WT, Choi YH, Kim W-J, et al. MicroRNA-106a suppresses proliferation, migration, and invasion of bladder cancer cells by modulating MAPK signaling, cell cycle regulators, and Ets-1-mediated MMP-2 expression. Oncol Rep. 2016;36:2421–9.
Bell SM, Zhang L, Mendell A, Xu Y, Haitchi HM, Lessard JL, et al. Kruppel-like factor 5 is required for formation and differentiation of the bladder urothelium. Dev Biol. 2011;358:79–90.
Zhang B, Li Y, Wu Q, Xie L, Barwick B, Fu C, et al. Acetylation of KLF5 maintains EMT and tumorigenicity to cause chemoresistant bone metastasis in prostate cancer. Nat Commun. 2021;12:1714.
Du C, Gao Y, Xu S, Jia J, Huang Z, Fan J, et al. KLF5 promotes cell migration by up-regulating FYN in bladder cancer cells. FEBS Lett. 2016;590:408–18.
Gao Y, Wu K, Chen Y, Zhou J, Du C, Shi Q, et al. Beyond proliferation: KLF5 promotes angiogenesis of bladder cancer through directly regulating VEGFA transcription. Oncotarget. 2015;6:43791–805.
Ohnishi S, Ohnami S, Laub F, Aoki K, Suzuki K, Kanai Y, et al. Downregulation and growth inhibitory effect of epithelial-type Krüppel-like transcription factor KLF4, but not KLF5, in bladder cancer. Biochem Biophys Res Commun. 2003;308:251–6.
Li H, Wang J, Xiao W, Xia D, Lang B, Wang T, et al. Epigenetic inactivation of KLF4 is associated with urothelial cancer progression and early recurrence. J Urol. 2014;191:493–501.
Ai X, Jia Z, Liu S, Wang J, Zhang X. Notch-1 regulates proliferation and differentiation of human bladder cancer cell lines by inhibiting expression of Krüppel-like factor 4. Oncol Rep. 2014;32:1459–64.
Xu X, Li J, Zhu Y, Xie B, Wang X, Wang S, et al. CRISPR-ON-Mediated KLF4 overexpression inhibits the proliferation, migration and invasion of urothelial bladder cancer in vitro and in vivo. Oncotarget. 2017;8:102078–87.
Suske G. The Sp-family of transcription factors. Gene. 1999;238:291–300.
Hagen G, Müller S, Beato M, Suske G. Sp1-mediated transcriptional activation is repressed by Sp3. EMBO J. 1994;13:3843–51.
Xu J, Hua X, Yang R, Jin H, Li J, Zhu J, et al. XIAP Interaction with E2F1 and Sp1 via its BIR2 and BIR3 domains specific activated MMP2 to promote bladder cancer invasion. Oncogenesis. 2019;8:71.
Huang H, Jin H, Zhao H, Wang J, Li X, Yan H, et al. RhoGDIβ promotes Sp1/MMP-2 expression and bladder cancer invasion through perturbing miR-200c-targeted JNK2 protein translation. Mol Oncol. 2017;11:1579–94.
Zhu J, Lu Z, Ke M, Cai X. Sp1 is overexpressed and associated with progression and poor prognosis in bladder urothelial carcinoma patients. Int Urol Nephrol. 2022;54:1505–12.
Malats N, Real FX. Epidemiology of bladder cancer. Hematol. Oncol. Clin. North Am. 2015;29:177–89.
Caliri AW, Tommasi S, Besaratinia A. Relationships among smoking, oxidative stress, inflammation, macromolecular damage, and cancer. Mutat Res Rev Mutat Res. 2021;787:108365.
Denison MS, Nagy SR. Activation of the aryl hydrocarbon receptor by structurally diverse exogenous and endogenous chemicals. Annu Rev Pharm Toxicol. 2003;43:309–34.
Pollenz RS, Sattler CA, Poland A. The aryl hydrocarbon receptor and aryl hydrocarbon receptor nuclear translocator protein show distinct subcellular localizations in Hepa 1c1c7 cells by immunofluorescence microscopy. Mol Pharm. 1994;45:428–38.
Hord NG, Perdew GH. Physicochemical and immunocytochemical analysis of the aryl hydrocarbon receptor nuclear translocator: characterization of two monoclonal antibodies to the aryl hydrocarbon receptor nuclear translocator. Mol Pharm. 1994;46:618–26.
Soshilov AA, Motta S, Bonati L, Denison MS. Transitional states in ligand-dependent transformation of the aryl hydrocarbon receptor into its DNA-binding form. Int J Mol Sci. 2020;21:2474.
Yu J, Lu Y, Muto S, Ide H, Horie S. The dual function of aryl hydrocarbon receptor in bladder carcinogenesis. Anticancer Res. 2020;40:1345–57.
Baker SC, Arlt VM, Indra R, Joel M, Stiborová M, Eardley I, et al. Differentiation-associated urothelial cytochrome P450 oxidoreductase predicates the xenobiotic-metabolizing activity of “luminal” muscle-invasive bladder cancers. Mol Carcinog. 2018;57:606–18.
Vlaar JM, Borgman A, Kalkhoven E, Westland D, Besselink N, Shale C, et al. Recurrent exon-deleting activating mutations in AHR act as drivers of urinary tract cancer. Sci Rep. 2022;12:10081.
Huang HC, Nguyen T, Pickett CB. Regulation of the antioxidant response element by protein kinase C-mediated phosphorylation of NF-E2-related factor 2. Proc Natl Acad Sci USA. 2000;97:12475–80.
Hayashi M, Guida E, Inokawa Y, Goldberg R, Reis LO, Ooki A, et al. GULP1 regulates the NRF2-KEAP1 signaling axis in urothelial carcinoma. Sci Signal. 2020;13:eaba0443.
Ihara T, Mitsui T, Nakamura Y, Kanda M, Tsuchiya S, Kira S, et al. The oscillation of intracellular Ca2+ influx associated with the circadian expression of Piezo1 and TRPV4 in the bladder urothelium. Sci Rep. 2018;8:5699.
Litlekalsoy J, Rostad K, Kalland K-H, Hostmark JG, Laerum OD. Expression of circadian clock genes and proteins in urothelial cancer is related to cancer-associated genes. BMC Cancer. 2016;16:549.
Honma S, Kawamoto T, Takagi Y, Fujimoto K, Sato F, Noshiro M, et al. Dec1 and Dec2 are regulators of the mammalian molecular clock. Nature. 2002;419:841–4.
Cobo I, Martinelli P, Flández M, Bakiri L, Zhang M, Carrillo-de-Santa-Pau E, et al. Transcriptional regulation by NR5A2 links differentiation and inflammation in the pancreas. Nature. 2018;554:533–7.
Bejjani F, Evanno E, Zibara K, Piechaczyk M, Jariel-Encontre I. The AP-1 transcriptional complex: Local switch or remote command? Biochim Biophys Acta Rev Cancer. 2019;1872:11–23.
Wang Y, Geng H, Zhao L, Zhang Z, Xie D, Zhang T, et al. Role of AP-1 in the tobacco smoke-induced urocystic abnormal cell differentiation and epithelial-mesenchymal transition in vivo. Int J Clin Exp Pathol. 2017;10:8243–52.
Barrows D, Feng L, Carroll TS, Allis CD. Loss of UTX/KDM6A and the activation of FGFR3 converge to regulate differentiation gene-expression programs in bladder cancer. Proc Natl Acad Sci USA. 2020;117:25732–41.
Qiu H, Makarov V, Bolzenius JK, Halstead A, Parker Y, Wang A, et al. KDM6A loss triggers an epigenetic switch that disrupts urothelial differentiation and drives cell proliferation in bladder cancer. Cancer Res. 2023;83:814–29.
Neyret-Kahn H, Fontugne J, Meng XY, Groeneveld CS, Cabel L, Ye T, et al. Epigenomic mapping identifies an enhancer repertoire that regulates cell identity in bladder cancer through distinct transcription factor networks. Oncogene. 2023;42:1524–42.
Cai Z, Chen H, Bai J, Zheng Y, Ma J, Cai X, et al. Copy number variations of CEP63, FOSL2 and PAQR6 serve as novel signatures for the prognosis of bladder cancer. Front Oncol. 2021;11:674933.
Iwata J, Suzuki A, Pelikan RC, Ho T-V, Sanchez-Lara PA, Urata M, et al. Smad4-Irf6 genetic interaction and TGFβ-mediated IRF6 signaling cascade are crucial for palatal fusion in mice. Development. 2013;140:1220–30.
Richardson RJ, Dixon J, Malhotra S, Hardman MJ, Knowles L, Boot-Handford RP, et al. Irf6 is a key determinant of the keratinocyte proliferation-differentiation switch. Nat Genet. 2006;38:1329–34.
Restivo G, Nguyen B-C, Dziunycz P, Ristorcelli E, Ryan RJH, Özuysal ÖY, et al. IRF6 is a mediator of Notch pro-differentiation and tumour suppressive function in keratinocytes. EMBO J. 2011;30:4571–85.
Weng H, Yuan S, Huang Q, Zeng X-T, Wang X-H. STAT1 is a key gene in a gene regulatory network related to immune phenotypes in bladder cancer: An integrative analysis of multi-omics data. J Cell Mol Med. 2021;25:3258–71.
Su Q, Sun Y, Zhang Z, Yang Z, Qiu Y, Li X, et al. Identification of prognostic immune genes in bladder urothelial carcinoma. Biomed Res Int. 2020;2020:7510120.
Kawahara T, Ishiguro Y, Ohtake S, Kato I, Ito Y, Ito H, et al. PD-1 and PD-L1 are more highly expressed in high-grade bladder cancer than in low-grade cases: PD-L1 might function as a mediator of stage progression in bladder cancer. BMC Urol. 2018;18:97.
Huang W-T, Yang S-F, Wu C-C, Chen W-T, Huang Y-C, Su Y-C, et al. Expression of signal transducer and activator of transcription 3 and suppressor of cytokine signaling 3 in urothelial carcinoma. Kaohsiung J Med Sci. 2009;25:640–6.
Hindupur SV, Schmid SC, Koch JA, Youssef A, Baur E-M, Wang D, et al. STAT3/5 inhibitors suppress proliferation in bladder cancer and enhance oncolytic adenovirus therapy. Int J Mol Sci. 2020;21:1106.
Gatta LB, Melocchi L, Bugatti M, Missale F, Lonardi S, Zanetti B, et al. Hyper-activation of STAT3 sustains progression of non-papillary basal-type bladder cancer via FOSL1 regulome. Cancers. 2019;11:1219.
Ching CB, Gupta S, Li B, Cortado H, Mayne N, Jackson AR, et al. Interleukin-6/Stat3 signaling has an essential role in the host antimicrobial response to urinary tract infection. Kidney Int. 2018;93:1320–9.
Schneidewind L, Neumann T, Plis A, Brückmann S, Keiser M, Krüger W, et al. Novel 3D organotypic urothelial cell culture model for identification of new therapeutic approaches in urological infections. J Clin Virol. 2020;124:104283.
Boorjian S, Ugras S, Mongan NP, Gudas LJ, You X, Tickoo SK, et al. Androgen receptor expression is inversely correlated with pathologic tumor stage in bladder cancer. Urology. 2004;64:383–8.
Tuygun C, Kankaya D, Imamoglu A, Sertcelik A, Zengin K, Oktay M, et al. Sex-specific hormone receptors in urothelial carcinomas of the human urinary bladder: a comparative analysis of clinicopathological features and survival outcomes according to receptor expression. Urol Oncol. 2011;29:43–51.
Miyamoto H, Yao JL, Chaux A, Zheng Y, Hsu I, Izumi K, et al. Expression of androgen and oestrogen receptors and its prognostic significance in urothelial neoplasm of the urinary bladder. BJU Int. 2012;109:1716–26.
Miyamoto H, Yang Z, Chen Y-T, Ishiguro H, Uemura H, Kubota Y, et al. Promotion of bladder cancer development and progression by androgen receptor signals. J Natl Cancer Inst. 2007;99:558–68.
Johnson AM, O’Connell MJ, Messing EM, Reeder JE. Decreased bladder cancer growth in parous mice. Urology. 2008;72:470–3.
Shen SS, Smith CL, Hsieh J-T, Yu J, Kim IY, Jian W, et al. Expression of estrogen receptors-alpha and -beta in bladder cancer cell lines and human bladder tumor tissue. Cancer. 2006;106:2610–6.
Hsu L-H, Liu K-J, Tsai M-F, Wu C-R, Feng A-C, Chu N-M, et al. Estrogen adversely affects the prognosis of patients with lung adenocarcinoma. Cancer Sci. 2015;106:51–9.
Ide H, Inoue S, Miyamoto H. Histopathological and prognostic significance of the expression of sex hormone receptors in bladder cancer: a meta-analysis of immunohistochemical studies. PLoS One. 2017;12:e0174746.
Warot X, Fromental-Ramain C, Fraulob V, Chambon P, Dollé P. Gene dosage-dependent effects of the Hoxa-13 and Hoxd-13 mutations on morphogenesis of the terminal parts of the digestive and urogenital tracts. Development. 1997;124:4781–91.
Mortlock DP, Innis JW. Mutation of HOXA13 in hand-foot-genital syndrome. Nat Genet. 1997;15:179–80.
Warrick JI, Knowles MA, Hurst CD, Shuman L, Raman JD, Walter V, et al. A transcriptional network of cell cycle dysregulation in noninvasive papillary urothelial carcinoma. Sci Rep. 2022;12:16538.
Marzouka N-A-D, Eriksson P, Bernardo C, Hurst CD, Knowles MA, Sjödahl G, et al. The lund molecular taxonomy applied to non-muscle-invasive urothelial carcinoma. J Mol Diagn. 2022;24:992–1008.
Lauss M, Aine M, Sjödahl G, Veerla S, Patschan O, Gudjonsson S, et al. DNA methylation analyses of urothelial carcinoma reveal distinct epigenetic subtypes and an association between gene copy number and methylation status. Epigenetics. 2012;7:858–67.
Aine M, Sjödahl G, Eriksson P, Veerla S, Lindgren D, Ringnér M, et al. Integrative epigenomic analysis of differential DNA methylation in urothelial carcinoma. Genome Med. 2015;7:23.
Bracken AP, Dietrich N, Pasini D, Hansen KH, Helin K. Genome-wide mapping of polycomb target genes unravels their roles in cell fate transitions. Genes Dev. 2006;20:1123–36.
Picchetti T, Chiquet J, Elati M, Neuvial P, Nicolle R, Birmelé E. A model for gene deregulation detection using expression data. BMC Syst Biol. 2015;9:S6.
Nicolle R, Radvanyi F, Elati M. CoRegNet: reconstruction and integrated analysis of co-regulatory networks. Bioinformatics. 2015;31:3066–8.
Champion M, Chiquet J, Neuvial P, Elati M, Radvanyi F, Birmelé E. Identification of deregulation mechanisms specific to cancer subtypes. J Bioinf Comput Biol. 2021;19:2140003.
Liang Y, Li L, Xin T, Li B, Zhang D. Superenhancer-transcription factor regulatory network in malignant tumors. Open Med (Wars). 2021;16:1564–82.
Sfakianos JP, Daza J, Hu Y, Anastos H, Bryant G, Bareja R, et al. Epithelial plasticity can generate multi-lineage phenotypes in human and murine bladder cancers. Nat Commun. 2020;11:2540.
Wang H, Mei Y, Luo C, Huang Q, Wang Z, Lu G-M, et al. Single-cell analyses reveal mechanisms of cancer stem cell maintenance and epithelial-mesenchymal transition in recurrent bladder cancer. Clin Cancer Res. 2021;27:6265–78.
Wang K-J, Wang C, Dai L-H, Yang J, Huang H, Ma X-J, et al. Targeting an autocrine regulatory loop in cancer stem-like cells impairs the progression and chemotherapy resistance of bladder cancer. Clin Cancer Res. 2019;25:1070–86.
Whitfield JR, Beaulieu M-E, Soucek L. Strategies to inhibit myc and their clinical applicability. Front Cell Dev Biol. 2017;5:10.
Beaulieu M-E, Jauset T, Massó-Vallés D, Martínez-Martín S, Rahl P, Maltais L, et al. Intrinsic cell-penetrating activity propels Omomyc from proof of concept to viable anti-MYC therapy. Sci Transl Med. 2019;11:eaar5012.
Lindskrog SV, Prip F, Lamy P, Taber A, Groeneveld CS, Birkenkamp-Demtröder K, et al. An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer. Nat Commun. 2021;12:2301.
Chamberlain PP, Hamann LG. Development of targeted protein degradation therapeutics. Nat Chem Biol. 2019;15:937–44.
Qi S-M, Dong J, Xu Z-Y, Cheng X-D, Zhang W-D, Qin J-J. PROTAC: an effective targeted protein degradation strategy for cancer therapy. Front Pharm. 2021;12:692574.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38:576–89.
Brooks DJ, Woodward S, Thompson FH, Dos Santos B, Russell M, Yang JM, et al. Expression of the zinc finger gene EVI-1 in ovarian and other cancers. Br J Cancer. 1996;74:1518–25.
Inoue Y, Kishida T, Kotani S-I, Akiyoshi M, Taga H, Seki M, et al. Direct conversion of fibroblasts into urothelial cells that may be recruited to regenerating mucosa of injured urinary bladder. Sci Rep. 2019;9:13850.
Primdahl H, von der Maase H, Christensen M, Wolf H, Orntoft TF. Allelic deletions of cell growth regulators during progression of bladder cancer. Cancer Res. 2000;60:6623–9.
Boorjian SA, Heemers HV, Frank I, Farmer SA, Schmidt LJ, Sebo TJ, et al. Expression and significance of androgen receptor coactivators in urothelial carcinoma of the bladder. Endocr Relat Cancer. 2009;16:123–37.
Dozmorov M, Stone R, Clifford JL, Sabichi AL, Engles CD, Hauser PJ, et al. System level changes in gene expression in maturing bladder mucosa. J Urol. 2011;185:1952–8.
Zhang Y, Dufau ML. Nuclear orphan receptors regulate transcription of the gene for the human luteinizing hormone receptor. J Biol Chem. 2000;275:2763–70.
Hermann-Kleiter N, Gruber T, Lutz-Nicoladoni C, Thuille N, Fresser F, Labi V, et al. The nuclear orphan receptor NR2F6 suppresses lymphocyte activation and T helper 17-dependent autoimmunity. Immunity. 2008;29:205–16.
Du X, Wang Q-R, Chan E, Merchant M, Liu J, French D, et al. FGFR3 stimulates stearoyl CoA desaturase 1 activity to promote bladder tumor growth. Cancer Res. 2012;72:5843–55.
Fu Y, Sun S, Bi J, Kong C, Yin L. Construction and analysis of a ceRNA network and patterns of immune infiltration in bladder cancer. Transl Androl Urol. 2021;10:1939–55.
Jiang A, Liu N, Bai S, Wang J, Gao H, Zheng X, et al. The construction and analysis of tumor-infiltrating immune cells and ceRNA networks in bladder cancer. Front Genet. 2020;11:605767.
Papaioannou VE. The T-box gene family: emerging roles in development, stem cells and cancer. Development. 2014;141:3819–33.
Lingbeek ME, Jacobs JJL, van Lohuizen M. The T-box repressors TBX2 and TBX3 specifically regulate the tumor suppressor gene p14ARF via a variant T-site in the initiator. J Biol Chem. 2002;277:26120–7.
Peres J, Davis E, Mowla S, Bennett DC, Li JA, Wansleben S, et al. The highly homologous T-box transcription Factors, TBX2 and TBX3, have distinct roles in the oncogenic process. Genes Cancer. 2010;1:272–82.
Ichijo R, Kobayashi H, Yoneda S, Iizuka Y, Kubo H, Matsumura S, et al. Tbx3-dependent amplifying stem cell progeny drives interfollicular epidermal expansion during pregnancy and regeneration. Nat Commun. 2017;8:508.
Ito A, Asamoto M, Hokaiwado N, Takahashi S, Shirai T. Tbx3 expression is related to apoptosis and cell proliferation in rat bladder both hyperplastic epithelial cells and carcinoma cells. Cancer Lett. 2005;219:105–12.
Aydogdu N, Rudat C, Trowe M-O, Kaiser M, Lüdtke TH, Taketo MM, et al. TBX2 and TBX3 act downstream of canonical WNT signaling in patterning and differentiation of the mouse ureteric mesenchyme. Development. 2018;145:dev171827.
Shi Z, Li X, Wu D, Tang R, Chen R, Xue S, et al. Silencing of HMGA2 suppresses cellular proliferation, migration, invasion, and epithelial-mesenchymal transition in bladder cancer. Tumour Biol. 2016;37:7515–23.
Chen Z, Li Q, Wang S, Zhang J. miR-485-5p inhibits bladder cancer metastasis by targeting HMGA2. Int J Mol Med. 2015;36:1136–42.
Ding X, Wang Y, Ma X, Guo H, Yan X, Chi Q, et al. Expression of HMGA2 in bladder cancer and its association with epithelial-to-mesenchymal transition. Cell Prolif. 2014;47:146–51.
Zhang Y, Luo G, You S, Zhang L, Liang C, Chen X. Exosomal LINC00355 derived from cancer-associated fibroblasts promotes bladder cancer cell proliferation and invasion by regulating miR-15a-5p/HMGA2 axis. Acta Biochim Biophys Sin. 2021;53:673–82.
Zhuang J, Shen L, Yang L, Huang X, Lu Q, Cui Y, et al. TGFβ1 promotes gemcitabine resistance through regulating the LncRNA-LET/NF90/miR-145 signaling axis in bladder cancer. Theranostics. 2017;7:3053–67.
Zheng Y, Izumi K, Yao JL, Miyamoto H. Dihydrotestosterone upregulates the expression of epidermal growth factor receptor and ERBB2 in androgen receptor-positive bladder cancer cells. Endocr Relat Cancer. 2011;18:451–64.
Xu C, Sun M, Zhang X, Xu Z, Miyamoto H, Zheng Y. Activation of glucocorticoid receptor inhibits the stem-like properties of bladder cancer via inactivating the β-catenin pathway. Front Oncol. 2020;10:1332.
Sun J, Hoshino H, Takaku K, Nakajima O, Muto A, Suzuki H, et al. Hemoprotein Bach1 regulates enhancer availability of heme oxygenase-1 gene. EMBO J. 2002;21:5216–24.
Kanno H, Ozawa H, Dohi Y, Sekiguchi A, Igarashi K, Itoi E. Genetic ablation of transcription repressor Bach1 reduces neural tissue damage and improves locomotor function after spinal cord injury in mice. J Neurotrauma. 2009;26:31–9.
Somerville TDD, Xu Y, Wu XS, Maia-Silva D, Hur SK, de Almeida LMN, et al. ZBED2 is an antagonist of interferon regulatory factor 1 and modifies cell identity in pancreatic cancer. Proc Natl Acad Sci USA. 2020;117:11471–82.
Finnegan A, Cho RJ, Luu A, Harirchian P, Lee J, Cheng JB, et al. Single-cell transcriptomics reveals spatial and temporal turnover of keratinocyte differentiation regulators. Front Genet. 2019;10:775.
Fossum SL, Mutolo MJ, Tugores A, Ghosh S, Randell SH, Jones LC, et al. Ets homologous factor (EHF) has critical roles in epithelial dysfunction in airway disease. J Biol Chem. 2017;292:10938–49.
Asai T, Morrison SL. The SRC family tyrosine kinase HCK and the ETS family transcription factors SPIB and EHF regulate transcytosis across a human follicle-associated epithelium model. J Biol Chem. 2013;288:10395–405.
Rubin AJ, Barajas BC, Furlan-Magaril M, Lopez-Pajares V, Mumbach MR, Howard I, et al. Lineage-specific dynamic and pre-established enhancer-promoter contacts cooperate in terminal differentiation. Nat Genet. 2017;49:1522–8.
Shi J, Qu Y, Li X, Sui F, Yao D, Yang Q, et al. Increased expression of EHF via gene amplification contributes to the activation of HER family signaling and associates with poor survival in gastric cancer. Cell Death Dis. 2016;7:e2442.
Taniue K, Oda T, Hayashi T, Okuno M, Akiyama T. A member of the ETS family, EHF, and the ATPase RUVBL1 inhibit p53-mediated apoptosis. EMBO Rep. 2011;12:682–9.
Lv Y, Sui F, Ma J, Ren X, Yang Q, Zhang Y, et al. Increased expression of EHF contributes to thyroid tumorigenesis through transcriptionally regulating HER2 and HER3. Oncotarget. 2016;7:57978–90.
Cheng Z, Guo J, Chen L, Luo N, Yang W, Qu X. Knockdown of EHF inhibited the proliferation, invasion and tumorigenesis of ovarian cancer cells. Mol Carcinog. 2016;55:1048–59.
Cangemi R, Mensah A, Albertini V, Jain A, Mello-Grand M, Chiorino G, et al. Reduced expression and tumor suppressor function of the ETS transcription factor ESE-3 in prostate cancer. Oncogene. 2008;27:2877–85.
Zhao T, Jiang W, Wang X, Wang H, Zheng C, Li Y, et al. ESE3 inhibits pancreatic cancer metastasis by upregulating E-cadherin. Cancer Res. 2017;77:874–85.
Wang L, Xing J, Cheng R, Shao Y, Li P, Zhu S, et al. Abnormal localization and tumor suppressor function of epithelial tissue-specific transcription factor ESE3 in esophageal squamous cell carcinoma. PLoS One. 2015;10:e0126319.
Fisher WG, Yang P-C, Medikonduri RK, Jafri MS. NFAT and NFkappaB activation in T lymphocytes: a model of differential activation of gene expression. Ann Biomed Eng. 2006;34:1712–28.
Lee JH, Kim M, Im YS, Choi W, Byeon SH, Lee HK. NFAT5 induction and its role in hyperosmolar stressed human limbal epithelial cells. Investig Ophthalmol Vis Sci. 2008;49:1827–35.
Johnsen O, Skammelsrud N, Luna L, Nishizawa M, Prydz H, Kolstø AB. Small Maf proteins interact with the human transcription factor TCF11/Nrf1/LCR-F1. Nucleic Acids Res. 1996;24:4289–97.
Johnsen O, Murphy P, Prydz H, Kolsto AB. Interaction of the CNC-bZIP factor TCF11/LCR-F1/Nrf1 with MafG: binding-site selection and regulation of transcription. Nucleic Acids Res. 1998;26:512–20.
Steffen J, Seeger M, Koch A, Krüger E. Proteasomal degradation is transcriptionally controlled by TCF11 via an ERAD-dependent feedback loop. Mol Cell. 2010;40:147–58.
Cui M, Atmanli A, Morales MG, Tan W, Chen K, Xiao X, et al. Nrf1 promotes heart regeneration and repair by regulating proteostasis and redox balance. Nat Commun. 2021;12:5270.
Xu Z, Chen L, Leung L, Yen TSB, Lee C, Chan JY. Liver-specific inactivation of the Nrf1 gene in adult mouse leads to nonalcoholic steatohepatitis and hepatic neoplasia. Proc Natl Acad Sci USA. 2005;102:4120–5.
Lu Q, Qiufang Y, Peng L, Xiaowen Z, Yonghui Y.Xiuman Z,et al. Function expansion of antitumor transcriptional activator NFE2L1 by the original discovery of its non-transcription factor activity. BioRxiv. 2020; https://doi.org/10.1101/2020.10.08.330597.
Xia W, Li Y, Wu Z, Wang Y, Xing N, Yang W, et al. Transcription factor YY1 mediates epithelial-mesenchymal transition through the TGFβ signaling pathway in bladder cancer. Med Oncol. 2020;37:93.
Chen W, Jiang T, Mao H, Gao R, Gao X, He Y, et al. Nodal promotes the migration and invasion of bladder cancer cells via regulation of snail. J Cancer. 2019;10:1511–9.
Li J, Xu X, Meng S, Liang Z, Wang X, Xu M, et al. MET/SMAD3/SNAIL circuit mediated by miR-323a-3p is involved in regulating epithelial-mesenchymal transition progression in bladder cancer. Cell Death Dis. 2017;8:e3010.
Acknowledgements
We thank David McConkey and Andrew Mason for critical review of a previous version of the manuscript, Jaime Martínez de Villarreal and other members of the Epithelial Carcinogenesis Group for valuable contributions.
Funding
This work was supported, in part, by a grant from Fundación Científica de la Asociación Española Contra el Cáncer to FXR and EL (PRYGN223005REAL). The project that gave rise to these results received the support of a fellowship from “la Caixa” Foundation (ID 100010434). The fellowship code is LCF/BQ/DR20/11790014. SC was supported by Fellowship PRE2018-085808 from Agencia Estatal de Investigación, co-financed by Fondo Social Europeo. CNIO is supported by Ministerio de Ciencia, Innovación y Universidades as a Centro de Excelencia Severo Ochoa SEV-2015-0510.
Author information
Authors and Affiliations
Contributions
All authors reviewed and summarized literature on the topic. MR also performed new bioinformatics analyses included in the manuscript. SC took the main responsibility for the illustrations.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ramal, M., Corral, S., Kalisz, M. et al. The urothelial gene regulatory network: understanding biology to improve bladder cancer management. Oncogene 43, 1–21 (2024). https://doi.org/10.1038/s41388-023-02876-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-023-02876-3
- Springer Nature Limited
This article is cited by
-
Molecular profiling of a bladder cancer with very high tumour mutational burden
Cell Death Discovery (2024)