Abstract
SMARCE1 gene, encoding a core subunit of SWI/SNF chromatin remodeling complex, is situated on chromosome 17q21-ter region that is frequently gained in neuroblastoma. However, its role in the tumorigenesis remains unknown. Here, we showed that high expression of SMARCE1 was associated with poor prognosis of patients with neuroblastoma, especially those with MYCN amplification. Knockdown of SMARCE1 reduced proliferation, colony formation, and tumorigenicity of neuroblastoma cells. Mechanistically, SMARCE1 directly interacted with MYCN, which was necessary for MYCN-mediated transcriptional activation of downstream target genes including PLK1, ODC1, and E2F2. Overexpression of PLK1, ODC1 or E2F2 significantly reversed the inhibiting effect of SMARCE1 knockdown on the proliferation, colony formation, and tumorigenicity of MYCN-amplified neuroblastoma cells. Moreover, we revealed that MYCN directly regulated SMARCE1 transcription through binding to a non-canonical E-box of SMARCE1 promoter, thus enhancing SMARCE1-MYCN cooperativity. These findings establish SMARCE1 is a critical oncogenic factor in neuroblastoma and provide a new potential target for treatment of neuroblastoma with 17q21-ter gain and MYCN amplification.
Similar content being viewed by others
Introduction
Neuroblastoma is the most common extra cranial solid tumor in early childhood that arises from neuronal crest cells, which accounts for 15% of all pediatric cancer fatalities [1, 2]. Chromosome 17q21-ter gain is one of the most commonly gene copy number alteration in neuroblastoma, which shows statistically significant correlation with MYCN amplification and portends a grim prognosis [3, 4]. The insulin growth factor 2 binding protein 1, JMJD6, survivin and ALYREF genes located in this chromosomal region have been reported to facilitate the survival and proliferation of neuroblastoma cells [5,6,7,8].
SMARCE1 gene is located on chromosome 17q21.2, and encodes a core subunit of ATP-dependent chromatin remodeling complex SWI/SNF that functions in many biological processes [9, 10]. SMARCE1 contains a DNA-binding HMG domain and modulates gene expression in tumors [11, 12]. Recent researches reported that SMARCE1 functions as a transcriptional co-activator. In breast tumors, SMARCE1 formed a complex with HIF1A to cooperatively activate the PTK2 expression. Furthermore, SMARCE1 assists RUNX1 access the silencer and repress CD4 in early thymocytes [13, 14]. However, biological function of SMARCE1 in neuroblastoma and its mechanism have not been studied.
Amplification of the MYCN oncogene compose 25% of neuroblastoma patients, and correlate with adverse clinical outcomes and poor overall patient survival rates [15, 16]. MYCN is a member of the MYC oncogene family and primarily expressed in neural tissues [17, 18]. Like C-MYC, MYCN functions as a major transcriptional regulator and plays multiple regulatory roles in neuroblastoma including differentiation, proliferation, survival, self-renewal, metabolism, metastasis and angiogenesis [18]. MYCN regulates gene transcription by heterodimerizing with MAX to bind E-box DNA motifs [19, 20] and recruits co-factors to alter the structure of the chromatin [21].
In this study, we investigated the potential role of SMARCE1 in neuroblastoma. We found that SMARCE1 was highly expressed in MYCN-amplified tumors and cells. Suppression of SMARCE1 reduced the proliferation and clonogenic capacity and impaired tumorigenic potential of MYCN-amplified cells. Our data demonstrated that MYCN regulated SMARCE1 expression by binding to its promoter. Further investigation revealed that SMARCE1 and MYCN formed co-activators to regulate the transcription of three MYCN target genes (PLK1, ODC1 and E2F2). These results demonstrated that SMARCE1 is a key player in tumorigenesis of MYCN-amplified neuroblastoma.
Result
High SMARCE1 expression correlates with poor prognosis in neuroblastoma
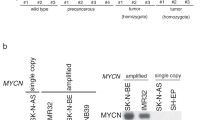
As SMARCE1 gene is localized at chromosome 17q21-ter region (Fig. 1A), we next examined whether chromosome 17q gain affects SMARCE1 expression in database. Analysis of the TARGET [22] and Jagannathan [23] datasets showed that SMARCE1 expression in tumor with 17q gain was significantly higher than that in 17q diploid tumor (Supplementary Fig. S1A). Interestingly, analysis using datasets of Kocak [24] and Delattre [25] showed that the mRNA level of SMARCE1 was higher in MYCN-amplified tumors than that in MYCN non-amplified tumors (Supplementary Fig. S1B). Consistently, immunoblots showed that the expression of SMARCE1 was higher in neuroblastoma cell lines with MYCN-amplification (IMR32, SK-N-BE(2)C, and SK-N-DZ) than that in neuroblastoma cell lines without the amplification (SHEP1, SH-SY5Y) (Fig. 1B). The gene expression of pan-cancer cell lines (RNAseq) were further examined in Cancer Cell Line Encyclopedia/Cancer Dependency Map [26]. We found that neuroblastoma cell lines have higher level of SMARCE1 expression compared to other tumor types (Supplementary Fig. S1C).
A SMARCE1 locus on chromosome 17q21-ter region. B SMARCE1 expression was analyzed by immunoblot in SK-N-BE(2)C, IMR32, SK-N-DZ, SH-SY5Y and SH-EP1 neuroblastoma cell lines. C Kaplan–Meier curves showed the probability of overall survival of patients according to SMARCE1 mRNA expression levels in the 66 MYCN-amplified neuroblastoma samples. Data are from the Kocak dataset. D Representative images of immunohistochemical staining of SMARCE1 in neuroblastoma patient tissues. E Quantitative analysis of SMARCE1 protein expression in neuroblastoma patient tissues at INSS stages 1and 2 versus stages 2B and 4. All data were shown as the mean ± SD, *p < 0.05, **p < 0.01, ***p < 0.001, ns not significant.
To validate the clinically relevance of SMARCE1 in patients with neuroblastoma. Kaplan–Meier analyses of overall survival probability of patients in Kocak, SEQC [27] and Versteeg [28] datasets were performed. The results showed that high SMARCE1 expression was associated with poor prognosis (Supplementary Fig. S1D). This is consistent with the literature reported previously [29, 30]. Furthermore, high SMARCE1 expression in the MYCN-amplified neuroblastoma patients was significantly correlated with worse outcome in Kocak and SEQC datasets (Fig. 1C and Supplementary Fig. S1E). We next examined SMARCE1 expression in 3 normal brain and 27 neuroblastoma samples by immunohistochemical staining. The quantified results showed SMARCE1 overexpression in neuroblastoma (Fig. 1D and Supplementary Fig. S1F). Moreover, high SMARCE1 protein expression was associated with the International Neuroblastoma Staging System (INSS) stage, an established risk factor (Fig. 1E). Similarly, SMARCE1 expression was closely associated with neuroblastoma INSS stage in Kocak datasets (Supplementary Fig. S1G).
SMARCE1 is required for proliferation and tumorigenicity of MYCN-amplified neuroblastoma cell
To determine whether SMARCE1 is required for neuroblastoma cell proliferation and survival, we designed two independent short hairpin RNAs (shRNAs) (shSMARCE1-1# and shSMARCE1-2#) targeting different regions of SMARCE1 mRNA. SK-N-BE(2)C and IMR32 cells were stably transfected with the shRNA vectors for effective knockdown of SMARCE1 expression (Fig. 2A). Consequently, cell proliferation was significantly suppressed after down-regulating SMARCE1 expression (Fig. 2B). Moreover, colony-formation assay showed that colony formation was diminished in the downregulated SMARCE1 group (Supplementary Fig. S2A). To determine whether downregulation of SMARCE1 expression has any effects on the cell cycle, we examined the cell cycle by flow cytometry. Silence of SMARCE1 resulted in G2/M-phase cell-cycle arrest of SK-N-BE(2)C and IMR32 cells (Supplementary Fig. S2B). In addition, the expression of CDK1 and Cyclin B1, G2/M related proteins, was markedly reduced in shSMARCE1 cells (Supplementary Fig. S2C).
A Real-time qPCR and immunoblot analyses of SMARCE1 in SK-N-BE(2)C and IMR32 cells with SMARCE1 shRNA knockdown. B Cell viability of SMARCE1 shRNA and the GFP control in SK-N-BE(2)C and IMR32 cells were assessed. C, D Xenograft assay of BE(2)-C and IMR32 cells with inducible expression of shGFP or shSMARCE1. Tumor weight was analyzed by scatterplot with horizontal lines indicating the mean. Tumor volume was measured and indicated by curves. E–G IHC staining of SMARCE1 and Ki67 for the tumors isolated from the indicated treatment. E Representative image of the staining; F, G percentage of SMARCE1 and Ki67 positive cells were quantified from three independent samples. All data were shown as the mean ± SD, *p < 0.05, **p < 0.01, ***p < 0.001, ns not significant.
Then, we implanted neuroblastoma cells subcutaneously in nonobese diabetic (NOD)/severe combined immunodeficient (SCID) mice. Both the volume and weight of the xenografts in the shSMARCE1 SK-N-BE(2)C and IMR32 cells were markedly smaller than those in the shGFP control cells (Fig. 2C, D). Immunohistochemical staining of the subcutaneous tumors indicated that Ki-67 expression, a proliferation marker, was significantly reduced in tumors expressing shSMARCE1 than that expressing shGFP (Fig. 2E–G). Taken together, these results underlined the strong oncogenic potential of SMARCE1 expression in neuroblastoma, and supported that SMARCE1 is critical to maintain clonogenic capacity, proliferation and survival of neuroblastoma cell.
MYCN regulates SMARCE1 transcription by binding to its promoter
Myc oncoproteins bind the canonical and non-canonical E-box sequences and stimulate transcription of target gene [31]. Bioinformatic analysis identified a non-canonical E-box upstream of the transcription start site of SMARCE1 gene (Supplementary Fig. S3A) and showed positive correlation between mRNA levels of MYCN and SMARCE1 (Supplementary Fig. S3B), which implied MYCN is involved in the regulation of SMARCE1 expression. To probe this possibility, we evaluated the effect of MYCN on SMARCE1 expression. Downregulation of MYCN led to reduced SMARCE1 mRNA and protein level in SK-N-BE(2)C and IMR32 cells (Fig. 3A, B). In addition, a dual-luciferase report assay was designed to examine the impact of knocking down MYCN on the SMARCE1 target promoter activity, and the results showed that the SMARCE1 promoter activity was markedly decreased upon silence of MYCN in SK-N-BE(2)C and IMR32 cells (Fig. 3C, D). Then, we performed chromatin immunoprecipitation (ChIP) assays using MYCN antibody in SK-N-BE(2)C and IMR32 cells. The experiments showed MYCN protein was enriched in the −782 to −408 bp region (Amplicon C) of the SMARCE1 promoter (Fig. 3E, F), consisting with previously published ChIP results [32, 33] that MYCN binds to the SMARCE1 promoter in other neuroblastoma cells (Supplementary Fig. S3C).
A Real-time qPCR analysis of MYCN and SMARCE1 expression upon MYCN shRNA knockdown in SK-N-BE(2)C and IMR32 cells. B Immunoblot detection of SMARCE1 protein levels after MYCN shRNA knockdown. C, D Relative luciferase activity of reporter constructs containing the SMARCE1 promoter in SK-N-BE(2) and IMR32 cells transfected with MYCN shRNA. E Upper panel, a total of five sets of primers were designed within the human SMARCE1 promoter; lower panel, ChIP-qPCR analysis using anti-IgG and anti-MYCN antibodies in SK-N-BE(2) cells. F ChIP-qPCR analysis using anti-IgG and anti-MYCN antibodies in IMR32 cells. G Predicted sequences of E-box in SMARCE1 promoter by JASPAR. H Luciferase assays performed using SMARCE1 wild-type and SMARCE1 mutant promoter. All data were shown as the mean ± SD, *p < 0.05, **p < 0.01, ***p < 0.001, ns not significant.
As the predicted E-box is located in the −782 to −408 bp region of the SMARCE1 promoter, we hypothesized this site is crucial for the MYCN binding and function. To test this hypothesis, we mutated the E-box sequence and tested its response to the knockdown of MYCN (Fig. 3G, H). The dual-luciferase report assay showed that the SMARCE1 promoter activity was significantly reduced upon MYCN knockdown, while such response was disappeared when the mutant SMARCE1 promoter sequence was used. These data indicated that MYCN regulated SMARCE1 transcription by binding to the non-canonical E-box of SMARCE1 promoter.
SMARCE1 interacts with MYCN
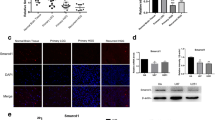
It has been reported that SWI/SNF complex interacts with c-MYC and is necessary for c-MYC-mediated transactivation [34], which inspired us to investigate the potential interaction between SMARCE1 and MYCN. We first performed Mass spectrometric assay for SMARCE1-interacting proteins. This assay identified MYCN as a bona fide interacting partner with SMARCE1 (Supplementary Fig. S4A). In addition, a number of previously reported SMARCE1-interacting proteins, including the SMARCC2, SMARCA4 and ILF3, were enriched in the interactome [35,36,37]. We then performed co-immunoprecipitation with MYCN or SMARCE1 antibody in SK-N-BE(2)C cells. The immunoblot experiments showed SMARCE1 was co-immunoprecipitated with MYCN, and vice versa (Fig. 4A). Similarly, co-immunoprecipitation between SMARCE1 and MYCN was observed in HEK-293FT cells transfected with Flag-SMARCE1 and HA-MYCN (Fig. 4B). To test whether the interaction between SMARCE1 and MYCN was direct, we performed GST pull-down assays. The result showed that SMARCE1 directly interacted with MYCN in vitro (Fig. 4C). Then we used a proximity ligation assay (PLA) to visualize the protein-protein interaction in vivo. The obvious PLA dot signal in the two neuroblastoma cell lines underpinned the interaction between SMARCE1 and MYCN (Fig. 4D and Supplementary Fig. S4B).
A Endogenous interaction between MYCN with SMARCE1 was confirmed by immunoprecipitation using anti-MYCN antibody and anti-SMARCE1 antibody in SK-N-BE(2)C cells. B HEK-293FT cells were ectopically overexpressed HA-MYCN and Flag-SMARCE1. The interaction was detected by co-immunoprecipitation using anti-HA antibody and anti-Flag antibody. C GST pull-down assay of MYCN using the indicated proteins expressed in bacteria. D Protein-protein interactions between SMARCE1 and MYCN were evaluated by using Duolink in situ PLA technology. Scale bars, 5 μm. E Schematic representation of the SMARCE1 deletion mutants. F Representative immunoblot analysis for ectopically expressed MYCN from HEK-293FT cells after IP of the indicated Flag-SMARCE1 mutants. G Schematic representation of the MYCN deletion mutants. H Representative immunoblot analysis for ectopically expressed SMARCE1 from HEK-293FT cells after IP of the indicated HA-MYCN mutants.
To better understand the interaction, we generated a series of truncated mutants of SMARCE1, including N-terminal truncated mutant, C-terminal truncated mutant, HMG domain-deletion mutant and coiled-coil domain-deletion mutant (Fig. 4D). Co-immunoprecipitation revealed C-terminal truncated and HMG domain-deletion mutants were unable to detect the co-precipitation of MYCN, demonstrating that HMG domain was crucial for the binding to MYCN (Fig. 4E). On the other hand, we constructed a series of MYCN truncating mutants and found only full length MYCN (1-464) and MYCN-M (141-345) were able to co-immunoprecipitated with SMARCE1 (Fig. 4F, G), suggesting the small region (amino acids 141-350) of MYCN is responsible for binding to SMARCE1.
SMARCE1 and MYCN cooperate to regulate the expression PLK1, ODC1 and E2F2
We next examined whether SMARCE1 is necessary for MYCN-mediated transcriptional regulation. Gene set enrichment analysis of the GSE16476 [38] cohort showed enriched MYCN target genes, PLK1 pathway and E2F pathway in samples with relatively high SMARCE1 expression (Fig. 5A). We focused on the effect of SMARCE1 on the three known target genes of Myc, PLK1, ODC1 and E2F2 [33, 22,23,24], whose high expression was associated with poor prognosis of neuroblastoma patients in Kocak and Versteeg datasets (Fig. 5B and Supplementary Fig. S5A). Quantitative PCR and immunoblotting assay showed that the mRNA and protein level of PLK1, ODC1 and E2F2 was significantly reduced after knockdown of SMARCE1 in SK-N-BE(2)C and IMR32 cells (Fig. 5C, D). Of course, the mRNA and protein level of MYCN did not change after knockdown of SMARCE1 in SK-N-BE(2)C and IMR32 cells. In addition, the mRNA and protein level of three target genes were restored upon re-expression of wild-type SMARCE1 in SK-N-BE(2)C cells (Supplementary Fig. S5B, C). Consistently, immunohistochemical staining of subcutaneous tumor tissues showed that the intensity of PLK1, ODC1 and E2F2 signal in the shSMARCE1-treated group was significantly reduced compared to that of shGFP-treated group (Supplementary Fig. S5D). We also analyzed the correlation between the expression of SMARCE1 and the three-target genes in Versteeg dataset and TARGET database. The results showed that SMARCE1 expression positively correlated with PLK1, ODC1 and E2F2 expression (Fig. 5E and Supplementary Fig. S5E).
A GSEA plots depicting enrichment of the MYCN target gene signature and PLK1 pathway in the transcriptional profiling of samples with high SMARCE1 expression in the GSE16476 cohort. FDR false discovery rate, NES normalized enrichment score. B Kaplan–Meier curves showed the probability of overall survival of patients according to PLK1, ODC1 and E2F2 mRNA expression levels in the 88 neuroblastoma samples in the Versteeg dataset. C Real-time qPCR analysis of PLK1, ODC1, E2F2 and MYCN expression upon SMARCE1 shRNA knockdown in SK-N-BE(2)C and IMR32 cells. D Immunoblot detection of PLK1, ODC1, E2F2 and MYCN protein levels after SMARCE1 shRNA knockdown. E The expression of SMARCE1 positively correlated with the expression of PLK1, ODC1 and E2F2 in the Target public database. All data were shown as the mean ± SD, *p < 0.05, **p < 0.01, ***p < 0.001, ns not significant.
Similarly, dual-luciferase assays using the PLK1, ODC1 and E2F2 promoters displayed that SMARCE1 deficiency significantly downregulated the activity of the three promoters (Fig. 6A–C). We further carried out ChIP experiments to determine the exact mechanism of transcriptional regulation. Using SMARCE1 antibody, we found SMARCE1 protein was mainly enriched within −132 to +156 bp region (Amplicon E) of PLK1, −335 to +35 bp region (Amplicon D) of ODC1, and −257 to +55 bp region (Amplicon D) of E2F2 (Fig. 6D–F). Similarly, MYCN was mainly recruited to Amplicon E of PLK1, Amplicon D of ODC1, and Amplicon D of E2F2 when used MYCN antibody in the ChIP experiments (Supplementary Fig. S6A). Supported of this, canonical and non-canonical E-boxes are found in these regions (Supplementary Fig. S6B). Analysis of publicly available ChIP sequencing data [39, 32, 33] revealed that MYCN and SMARCE1 could bind to the PLK1, ODC1 and E2F2 promoter and the binding peaks overlapped in other cancer cells (Supplementary Fig. S6C). Importantly, SMARCE1 knockdown attenuated the promoter occupancy of MYCN, suggesting that the recruitment of MYCN protein to promoters requires SMARCE1 protein (Fig. 6G and Supplementary Fig. S6D). Re-ChIP (sequential ChIP) experiments further demonstrated that SMARCE1 cooperated with MYCN to occupy the PLK1, ODC1 and E2F2 promoters (Fig. 6H).
A–C Relative luciferase activity of reporter constructs containing the PLK1, ODC1 or E2F2 promoter in SK-N-BE(2) cells transfected with SMARCE1 shRNA. D Upper panel, a total of six sets of primers were designed within the human PLK1 promoter; lower panel, ChIP-qPCR analysis using anti-IgG and anti-SMARCE1 antibodies in SK-N-BE(2) cells. E, F Upper panel, a total of five sets of primers were designed within the human ODC1 or E2F2 promoter; lower panel, ChIP-qPCR analysis using anti-IgG and anti-SMARCE1 antibodies in SK-N-BE(2) cells. G ChIP analysis with MYCN and SMARCE1 antibodies upon SMARCE1 shRNA knockdown in SK-N-BE(2)C cells. H SK-N-BE(2) cells analyzed by re-ChIP to detect mutual binding of SMARCE1 and MYCN to promoters. The first round (initial ChIP) antibody (IgG or MYCN) is indicated below the plots and the second round (re-ChIP) antibody (IgG or SMARCE1) is shown in the color key. All data were shown as the mean ± SD, *p < 0.05, **p < 0.01, ***p < 0.001, ns not significant.
Finally, we overexpressed PLK1, ODC1 or E2F2 in SMARCE1-depleted SK-N-BE(2)C cells (Fig. 7A). As a result, the cell proliferation and colony formation were significantly restored upon the re-introduction of PLK1, ODC1 or E2F2 in the SMARCE1-depleted SK-N-BE(2)C cells (Fig. 7B–D). Next, we performed subcutaneous tumorigenesis assays to test tumorigenic potential in vivo, and the result indicated that overexpression of PLK1, ODC1 or E2F2 restored tumor growth after SMARCE1 knockdown (Fig. 7E–G).
A Immunoblot detection of SMARCE1, PLK1, ODC1 or E2F2 protein levels after overexpressing with PLK1, ODC1 or E2F2 in SMARCE1 shRNA knockdown SK-N-BE(2)C cells. B SK-N-BE(2)C cells stably expressing the negative control or SMARCE1 shRNA were further infected with a lentivirus overexpressing PLK1, ODC1 or E2F2. Cell viability was assessed. C, D The colony-formation ability was tested after overexpressing PLK1, ODC1 or E2F2 in SMARCE1 shRNA knockdown SK-N-BE(2)C cells. E–G Xenograft assay was performed after overexpressing PLK1, ODC1 or E2F2 in SMARCE1 shRNA knockdown SK-N-BE(2)C cells. Tumor weight was analyzed by scatterplot with horizontal lines indicating the mean. Tumor volume was measured and indicated by curves. H Model for MYCN-SMARCE1 cooperation in neuroblastoma. All data were shown as the mean ± SD, *p < 0.05, **p < 0.01, ***p < 0.001, ns not significant.
Discussion
Neuroblastoma is the most common malignancy of early childhood and arises in the developing sympathetic nervous system (from any neural crest element) [1]. In this study, we identified SMARCE1 as a potential target for prognosis and therapy of neuroblastoma, especially those with MYCN amplification. Mechanistically, we revealed MYCN-SMARCE1-PLK1/ODC1/E2F2 regulatory axis in the MYCN-amplified neuroblastoma cells, which is involved in the proliferation, self-renewal, and tumorigenicity of the tumor cells (Fig. 7H).
As the most common changes, MYCN amplification is closely correlated with poor outcomes and driving carcinogenic events in neuroblastoma [4]. MYC protein recruited transcriptional co-factors to regulate genes expression, such as TIP60 histone acetyltransferases or the p300/CBP acetyltransferase [40, 41]. Our data identified SMARCE1 as a novel binding protein of MYCN and demonstrated that the molecular interaction between SMARCE1 and MYCN might have important implications for efficient gene transcription. Consistent with this observation, SMARCE1 plays an important role in DNA replication and DNA repair, especially the transcriptional co-regulation [10, 37]. We confirmed that SMARCE1 protein interacted with MYCN protein at the truncated region of amino acids 141–350. It has been shown that MYC protein contains two highly conserved MYC boxes (MYC box III and IV) [42,43,44]. The boxes play a role in regulating DNA-binding activity and localize mostly in the binding regions of MYCN [45, 46]. Moreover, we identified that MYCN protein binds to the HMG DNA-binding domain of SMARCE1. Studies have implicated that HMG domain is required for SMARCE1 gene regulation [11, 12, 47]. SMARCE1 cooperated with MYCN to regulate the Myc target genes (PLK1, ODC1 and E2F2) transcription by binding to their gene promoters. Further research suggest that the MYCN-SMARCE1 interaction is an important factor that regulates PLK1 expression and cell proliferation (Supplementary Fig. S7A–C). Myc recruited co-activators to locally remodel the chromatin structure to achieve the function of transcriptional regulation [21, 48]. For example, MYCN interacted with p300/CBP to lead to an open chromatin structure that provides docking sites for additional proteins that promote transcription [40]. As a core subunit of ATP-dependent chromatin remodeling complex SWI/SNF, SMARCE1 might altered the nucleosome structure and chromatin accessibility for assisting the transcriptional regulation of MYCN. We suggested that the identity of the co-activator of SMARCE1 is biologically significant in MYCN-amplified neuroblastoma cells.
Previous studies have demonstrated that these chosen targets were under the regulation of MYCN and played an important role in neuroblastoma development [7, 49,50,51]. As essential regulators of cell cycle, PLK1 and E2F2 were enriched in correlated certain KEGG pathways [52, 53]. DOT1L was identified as a novel cofactor in MYCN–mediated transcriptional activation of E2F2 and ODC1 and neuroblastoma oncogenesis [51]. Moreover, C-MYC promoted cell proliferation and transformation by activating growth-promoting genes, including the ornithine decarboxylase (ODC1) and E2F2 [54].
By performing a dual-luciferase assay and ChIP analyses, we also found that MYCN regulated SMARCE1 expression by binding to its promoter. It is well known that MYCN activated the transcription of target genes through binding to Myc responsive element E-Boxes in promoters [31]. For example, recent evidence showed that MYCN regulated DOT1L expression through binding to Myc responsive element E-Boxes [51]. Interestingly, we identified an MYCN binding motif upstream of the transcription start site of the SMARCE1 promoter and confirmed MYCN protein enriching to the SMARCE1 promoter. Furthermore, the SMARCE1 promoter activity was not reduced after MYCN knockdown when the mutant SMARCE1 promoter sequence was used. Our data confirmed MYCN protein enriching to the SMARCE1 promoter and showed that SMARCE1 plays a key role in neuroblastoma due to the identity of a critical downstream factor of MYCN. Given the regulatory relationship between SMARCE1 and MYCN in this work, our finding also may extend implications to other tumor types with MYCN amplification or overexpression (e.g., retinoblastoma, glioblastoma, rhabdomyosarcoma and Wilms tumor) [55,56,57,58] (Supplementary Fig. S8A–D).
Many groups have explored the effect of targeting MYCN expression in neuroblastoma cells and indicated that direct targeting of MYCN could be promising for neuroblastoma therapy but remaining difficult [59,60,61]. To repress neuroblastoma growth effectively, the MYC pathway could potentially be targeted at different levels [62,63,64]. One strategy is to target critical downstream transcriptional targets of MYCN [65]. In this study, as a direct downstream factor of MYCN, SMARCE1 could be used as a potential molecular target for inhibiting MYCN signaling pathway in neuroblastoma. In addition, another strategy for neuroblastoma is to target the interaction of MYCN with other proteins, either with the obligate partner Max, or with co-factors recruited to the MYCN [66]. Thus, the interaction between MYCN and SMARCE1 could serve as potential targets for therapeutic interventions.
In summary, our study extends the knowledge about the key role of SMARCE1 in the tumor progress in MYCN-amplified and 17q21-ter gained neuroblastoma. The MYCN-SMARCE1-PLK1/ODC1/E2F2 regulatory axis identified in this study is valuable and could be a potential target for the treatment of neuroblastoma by exploiting the enforced SMARCE1 addiction present in MYCN-amplified tumors.
Materials and methods
Cell culture
Neuroblastoma cell lines SK-N-BE(2)C, IMR32, SHEP1, SH-SY5Y, SK-N-DZ and HEK-293FT were obtained from ATCC (Manassas, VA, USA). Cell culture methods were consistent with those described previously [67]. IMR32 were grown in alpha-minimum essential media (Gibco, New York, NY, USA). SK-N-BE(2)C were cultured in DME/F-12 1:1 media (Gibco, New York, NY, USA). SHEP1, SH-SY5Y, SK-N-DZ and HEK-293FT were cultured in Dulbecco’s modified Eagle’s medium (VivaCell, Shanghai, China). All media were supplemented with 10% fetal bovine serum.
Antibodies
The MYCN (51705S), Cyclin B1 (12231S) and PLK1 (4513S) antibodies were purchased from Cell Signaling Technology (Shanghai, China). The SMARCE1 antibody (#A300-810A) was obtained from Bethyl (Texas, USA). The ODC1 (28728-1-AP), HA (51064-2-AP) and CDK1 (19532-1-AP) antibodies were purchased from Proteintech Group (Wuhan, China). The Ki67 (ab92742), α-Tubulin (ab7291), E2F2 (ab138515) and Lamin B1 (ab16048) antibodies were purchased from Abcam (Shanghai, China). Anti-Flag (F3165) was purchased from Sigma (St Louis, MO, USA).
Plasmids, transfection, and infection
Human MYCN and SMARCE1 shRNA were purchased from Beijing Genomics Institute (BGI, Beijing, China) and then cloned into the PLKO.1 vector. The shRNA sequences were listed in Supplementary Table 1. Plasmids encoding human SMARCE1, MYCN, PLK1, ODC1, E2F2, truncated mutants of SMARCE1 and MYCN was obtained from Youbao Corporation (Changsha, China). Transfection and infection were performed essentially as previously described [68].
Immunoblot analysis
Cells were harvested and lysed with cell lysis buffer (Beyotime). Immunoblotting was performed as previously described [69].
Quantitative reverse-transcription PCR (qRT-PCR)
Total RNA was extracted using the standard TRIZOL method. Subsequently, cDNA was generated with the Reverse Transcription Kit (Promega). Quantitative real-time-PCR was performed as previously described [70]. The sequence of the primers used in this study were listed in Supplementary Table 2.
Flow cytometry analysis
Flow cytometry analysis was performed as described previously [70, 71].
Chromatin immunoprecipitation (ChIP) assays
ChIP assays were performed as previously described [72]. ChIP assay was performed with anti-MYCN antibody, anti-SMARCE1 antibody or control mouse IgG antibody (Beyotime). Quantitative real-time PCR was performed to analyze the precipitates. Primers used for qPCR were listed in Supplementary Table 3. For re-ChIP assays, we performed sequential pull-downs, first with anti-MYCN antibody followed by anti-SMARCE1 antibody using appropriate IgG controls.
GST pull-down and co-immunoprecipitation assays
GST pull-down assay was performed as described previously [73]. Briefly, SHEP1 or HEK-293FT cells were stably transfected with relevant proteins and lysed. The supernatant lysates were incubated overnight with GST or GST-bound protein immobilized on glutathione agarose beads. The beads were washed 4–5 times with buffer. The products were analyzed by immunoblotting analysis.
For endogenous MYCN or SMARCE1 immunoprecipitation assays, protein was extracted from SK-N-BE(2)C cells and co-immunoprecipitation was carried out using anti-MYCN antibody, anti-SMARCE1 antibody or control IgG as a negative control, followed by immunoblotting analysis. The appropriate bands were excised from protein gels were identified using MS/MS analysis.
For immunoprecipitation assays with ectopically expressed Flag and HA-tagged proteins, protein was extracted from HEK-293FT cells and co-immunoprecipitation was carried out using anti-Flag antibody, anti-HA antibody or control IgG as a negative control, followed by immunoblotting analysis.
Luciferase reporter assay
The SMARCE1, PLK1, ODC1, E2F2 and SMARCE1 mutant promoter sequences were synthesized by the BGI and subsequently ligated into the pGL3-basic vector (Promega). Luciferase reporter assay was performed as previously described [68].
Immunohistochemistry
Paraffin sections were dewaxed, hydrated and repaired with antigen. For antigen retrieval, sections were performed by microwave heating for 20 min in 10 mM citrate buffer (pH 6.0), followed incubating with SMARCE1, PLK1, ODC1, E2F2 and Ki-67 antibodies at 4 °C overnight. The target antigen was visualized using a horseradish peroxidase detection system (GBI Labs).
Cell growth and plate clone formation assays
The 2-(4,5-dimethyltriazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT; Sigma) colorimetric, plate clone formation assays were conducted as previously described [69, 74].
Animal studies
Five-week-old female nude (NOD/SCID) mice were purchased and housed in an SPF room. Total 1.5 × 106 SK-N-BE(2)C or IMR32 cells were collected and inoculated subcutaneously into the flanks of mice. Randomization and single-blinding were used for measurement. Tumor volumes were measured every 3 days with a caliper. Six weeks after injection of neuroblastoma cells, mice were euthanized and the tumors were resected. Animal studies were performed in accordance with the Guidelines of the Institute for Laboratory Animal Research, Southwest University (Chongqing, China).
Duolink PLA
This technique was performed using Duolink In Situ Red Starter Kit (#DUO92101, Sigma-Aldrich) according to the manufacturer’s instructions. The detailed technique can be found online (www.olink.com/products-services/duolink/how-useduolink). Briefly, neuroblastoma cells were fixed in 4% paraformaldehyde for 20 min at room temperature and blocked for 1 h with Duolink blocking solution at 37 °C. Then, cells were incubated with primary antibodies overnight at 4 °C and conjugated with oligonucleotides (PLA probe MINUS and PLA probe PLUS) for 120 min at 37 °C. Finally, cells were incubated sequentially with ligation solution and amplification solution followed by analysis with fluorescence microscopy.
Patient data analysis and patient tumor tissues
Patient data and gene expression datasets were obtained from R2: microarray analysis and visualization platform (https://hgserver1.amc.nl/cgi-bin/r2/main.cgi). Kaplan–Meier analysis and survival curves were carried out by using GraphPad Prism (version 6.0, GraphPad Software, San Diego, CA, USA). All cutoff values for separating high and low expression groups were determined by the online R2 database algorithm.
Tumor samples were obtained from Chaoying Biotechnology Co., Ltd. (Henan, China). Tissue analysis was approved by the Ethics Committee of Southwest University of China. All the patients provided written informed consent to participate.
ChIP-seq analysis
We obtained several other publicly available ChIP-seq datasets (GSM2113542, GSM2214113, GSM2113529, GSM2113526 and ENCSR157TCS) [32, 33, 39] to complement our experimental results. These files were aligned to the human genome (GRCh38) by using Integrative Genomics Viewer (v2.11.1).
Statistical analysis
All observations were confirmed by at least three independent experiments, and both sample size and significance analysis are indicated in the legends. All the results acquired in this study were presented as the means ± standard deviation. Two-tailed Student’s t test was performed for paired samples; p < 0.05 was considered statistically significant.
Data availability
All of the data and material in this paper are available when requested. We obtained several other publicly available ChIP-seq datasets (GSM2113542, GSM2214113, GSM2113529, GSM2113526 and ENCSR157TCS) for analysis. We also obtained 88 human Neuroblastoma samples dataset (GSE16476) for analysis (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi; https://www.encodeproject.org/).
References
Matthay KK, Maris JM, Schleiermacher G, Nakagawara A, Mackall CL, Diller L, et al. Neuroblastoma. Nat Rev Dis Prim. 2016;2:16078.
Fetahu IS, Taschner-Mandl S. Neuroblastoma and the epigenome. Cancer Metastasis Rev. 2021;40:173–89.
Bown N, Cotterill S, Lastowska M, O’Neill S, Pearson AD, Plantaz D, et al. Gain of chromosome arm 17q and adverse outcome in patients with neuroblastoma. N Engl J Med. 1999;340:1954–61.
O’Neill S, Ekstrom L, Lastowska M, Roberts P, Brodeur GM, Kees UR, et al. MYCN amplification and 17q in neuroblastoma: evidence for structural association. Genes Chromosomes Cancer. 2001;30:87–90.
Bell JL, Turlapati R, Liu T, Schulte JH, Huttelmaier S. IGF2BP1 harbors prognostic significance by gene gain and diverse expression in neuroblastoma. J Clin Oncol. 2015;33:1285–93.
Islam A, Kageyama H, Takada N, Kawamoto T, Takayasu H, Isogai E, et al. High expression of Survivin, mapped to 17q25, is significantly associated with poor prognostic factors and promotes cell survival in human neuroblastoma. Oncogene. 2000;19:617–23.
Wong M, Sun Y, Xi Z, Milazzo G, Poulos RC, Bartenhagen C, et al. JMJD6 is a tumorigenic factor and therapeutic target in neuroblastoma. Nat Commun. 2019;10:3319.
Nagy Z, Seneviratne JA, Kanikevich M, Chang W, Mayoh C, Venkat P, et al. An ALYREF-MYCN coactivator complex drives neuroblastoma tumorigenesis through effects on USP3 and MYCN stability. Nat Commun. 2021;12:1881.
Clapier CR, Iwasa J, Cairns BR, Peterson CL. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat Rev Mol Cell Bio. 2017;18:407–22.
Tsukiyama T. The in vivo functions of ATP-dependent chromatin-remodelling factors. Nat Rev Mol Cell Bio. 2002;3:422–9.
Wattler F, Wattler S, Kelly M, Skinner HB, Nehls M. Cloning, chromosomal location, and expression analysis of murine Smarce1-related, a new member of the high-mobility 365 group gene family. Genomics. 1999;60:172–8.
Heo Y, Park JH, Kim J, Han J, Yun JH, Lee W. Crystal structure of the HMG domain of human BAF57 and its interaction with four-way junction DNA. Biochem Biophys Res Commun. 2020;533:919–24.
Sethuraman A, Brown M, Seagroves TN, Wu ZH, Pfeffer LM, Fan M. SMARCE1 regulates metastatic potential of breast cancer cells through the HIF1A/PTK2 pathway. Breast Cancer Res. 2016;18:81.
Wan M, Zhang J, Lai D, Jani A, Prestone-Hurlburt P, Zhao L, et al. Molecular basis of CD4 repression by the Swi/Snf-like BAF chromatin remodeling complex. Eur J Immunol. 2009;39:580–8.
Brodeur GM. Neuroblastoma: biological insights into a clinical enigma. Nat Rev Cancer. 2003;3:203–16.
Monclair T, Brodeur GM, Ambros PF, Brisse HJ, Cecchetto G, Holmes K, et al. The International Neuroblastoma Risk Group (INRG) staging system: an INRG Task Force report. J Clin Oncol. 2009;27:298–303.
Meyer N, Penn LZ. Reflecting on 25 years with MYC. Nat Rev Cancer. 2008;8:976–90.
Huang M, Weiss WA. Neuroblastoma and MYCN. Cold Spring Harb Perspect Med. 2013;3:a014415.
Baluapuri A, Wolf E, Eilers M. Target gene-independent functions of MYC oncoproteins. Nat Rev Mol Cell Biol. 2020;21:255–67.
Wenzel A, Schwab M. The mycN/max protein complex in neuroblastoma. Short Rev Eur J Cancer. 1995;31A:516–9.
Guccione E, Martinato F, Finocchiaro G, Luzi L, Tizzoni L, Dall’ Olio V, et al. Myc-binding-site recognition in the human genome is determined by chromatin context. Nat Cell Biol. 2006;8:764–70.
Pugh TJ, Morozova O, Attiyeh EF, Asgharzadeh S, Wei JS, Auclair D, et al. The genetic landscape of high-risk neuroblastoma. Nat Genet. 2013;45:279–84.
Cole KA, Huggins J, Laquaglia M, Hulderman CE, Russell MR, Bosse K, et al. RNAi screen of the protein kinome identifies checkpoint kinase 1 (CHK1) as a therapeutic target in neuroblastoma. Proc Natl Acad Sci USA 2011;108:3336–41.
Kocak H, Ackermann S, Hero B, Kahlert Y, Oberthuer A, Juraeva D, et al. Hox-C9 activates the intrinsic pathway of apoptosis and is associated with spontaneous regression in neuroblastoma. Cell Death Dis. 2013;4:e586.
Bourdeaut F, Janoueix-Lerosey I, Lucchesi C, Paris R, Ribeiro A, de Pontual L, et al. Cholinergic switch associated with morphological differentiation in neuroblastoma. J Pathol. 2009;219:463–72.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Zhang W, Yu Y, Hertwig F, Thierry-Mieg J, Zhang W, Thierry-Mieg D, et al. Comparison of RNA-seq and microarray-based models for clinical endpoint prediction. Genome Biol. 2015;16:133.
Molenaar JJ, Koster J, Zwijnenburg DA, van Sluis P, Valentijn LJ, van der Ploeg I, et al. Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes. Nature. 2012;483:589–93.
Krstic A, Konietzny A, Halasz M, Cain P, Oppermann U, Kolch W, et al. A chemo-genomic approach identifies diverse epigenetic therapeutic vulnerabilities in MYCN-amplified neuroblastoma. Front Cell Dev Biol. 2021;9:612518.
Duffy DJ, Konietzny A, Krstic A, Mehta JP, Halasz M, Kolch W. Identification of a MYCN and Wnt-related VANGL2-ITLN1 fusion gene in neuroblastoma. Gene Rep. 2018;12:187–200.
Eilers M, Eisenman RN. Myc’s broad reach. Genes Dev. 2008;22:2755–66.
Gryder BE, Yohe ME, Chou HC, Zhang X, Marques J, Wachtel M, et al. PAX3-FOXO1 establishes myogenic super enhancers and confers BET bromodomain vulnerability. Cancer Disco. 2017;7:884–99.
Zeid R, Lawlor MA, Poon E, Reyes JM, Fulciniti M, Lopez MA, et al. Enhancer invasion shapes MYCN-dependent transcriptional amplification in neuroblastoma. Nat Genet. 2018;50:515–23.
Cheng SW, Davies KP, Yung E, Beltran RJ, Yu J, Kalpana GV. c-MYC interacts with INI1/hSNF5 and requires the SWI/SNF complex for transactivation function. Nat Genet. 1999;22:102–5.
Kadoch C, Crabtree GR. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci Adv. 2015;1:e1500447.
Nakayama RT, Pulice JL, Valencia AM, McBride MJ, McKenzie ZM, Gillespie MA, et al. SMARCB1 is required for widespread BAF complex-mediated activation of enhancers and bivalent promoters. Nat Genet. 2017;49:1613–23.
Sokol ES, Feng YX, Jin DX, Tizabi MD, Miller DH, Cohen MA, et al. SMARCE1 is required for the invasive progression of in situ cancers. Proc Natl Acad Sci USA 2017;114:4153–8.
Molenaar JJ, Domingo-Fernandez R, Ebus ME, Lindner S, Koster J, Drabek K, et al. LIN28B induces neuroblastoma and enhances MYCN levels via let-7 suppression. Nat Genet. 2012;44:1199–206.
Davis CA, Hitz BC, Sloan CA, Chan ET, Davidson JM, Gabdank I, et al. The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 2018;46:D794–801.
Vervoorts J, Luscher-Firzlaff JM, Rottmann S, Lilischkis R, Walsemann G, Dohmann K, et al. Stimulation of c-MYC transcriptional activity and acetylation by recruitment of the cofactor CBP. EMBO Rep. 2003;4:484–90.
Rust K, Tiwari MD, Mishra VK, Grawe F, Wodarz A. Myc and the Tip60 chromatin remodeling complex control neuroblast maintenance and polarity in Drosophila. Embo J. 2018;37:e98659.
Blackwell TK, Kretzner L, Blackwood EM, Eisenman RN, Weintraub H. Sequence-specific DNA binding by the c-Myc protein. Science. 1990;250:1149–51.
Blackwood EM, Eisenman RN. Max: a helix-loop-helix zipper protein that forms a sequence-specific DNA-binding complex with Myc. Science. 1991;251:1211–17.
Nair SK, Burley SK. X-ray structures of Myc-Max and Mad-Max recognizing DNA. Molecular bases of regulation by proto-oncogenic transcription factors. Cell. 2003;112:193–205.
Cowling VH, Chandriani S, Whitfield ML, Cole MD. A conserved Myc protein domain, MBIV, regulates DNA binding, apoptosis, transformation, and G2 arrest. Mol Cell Biol. 2006;26:4226–39.
Amente S, Milazzo G, Sorrentino MC, Ambrosio S, Di Palo G, Lania L, et al. Lysine-specific demethylase (LSD1/KDM1A) and MYCN cooperatively repress tumor suppressor genes in neuroblastoma. Oncotarget. 2015;6:14572–83.
Kumar P, Mistri TK. Transcription factors in SOX family: potent regulators for cancer initiation and development in the human body. Semin Cancer Biol. 2020;67:105–13.
Amente S, Zhang J, Lavadera ML, Lania L, Avvedimento EV, Majello B. Myc and PI3K/AKT signaling cooperatively repress FOXO3a-dependent PUMA and GADD45a gene expression. Nucleic Acids Res. 2011;39:9498–507.
Xiao D, Yue M, Su H, Ren P, Jiang J, Li F, et al. Polo-like Kinase-1 regulates Myc stabilization and activates a feedforward circuit promoting tumor cell survival. Mol Cell. 2016;64:493–506.
Hogarty MD, Norris MD, Davis K, Liu X, Evageliou NF, Hayes CS, et al. ODC1 is a critical determinant of MYCN oncogenesis and a therapeutic target in neuroblastoma. Cancer Res. 2008;68:9735–45.
Wong M, Tee AEL, Milazzo G, Bell JL, Poulos RC, Atmadibrata B, et al. The histone methyltransferase DOT1L promotes neuroblastoma by regulating gene transcription. Cancer Res. 2017;77:2522–33.
Sun J, Yi S, Qiu L, Fu W, Wang A, Liu F, et al. SATB1 defines a subtype of cutaneous CD30(+) lymphoproliferative disorders associated with a T-Helper 17 cytokine profile. J Invest Dermatol. 2018;138:1795–804.
Wei J, Yin Y, Deng Q, Zhou J, Wang Y, Yin G, et al. Integrative analysis of MicroRNA and gene interactions for revealing candidate signatures in prostate cancer. Front Genet. 2020;11:176.
Mastronikolis N, Ragos V, Kyrodimos E, Chrysovergis A, Papanikolaou V, Mastronikolis S, et al. Mechanisms of C-myc oncogenic activity in head and neck squamous cell carcinoma. J BUON. 2019;24:2242–4.
Lee WH, Murphree AL, Benedict WF. Expression and amplification of the N-myc gene in primary retinoblastoma. Nature. 1984;309:458–60.
Hui AB, Lo KW, Yin XL, Poon WS, Ng HK. Detection of multiple gene amplifications in glioblastoma multiforme using array-based comparative genomic hybridization. Lab Invest. 2001;81:717–23.
Dias P, Kumar P, Marsden HB, Gattamaneni HR, Heighway J, Kumar S. N-myc gene is amplified in alveolar rhabdomyosarcomas (RMS) but not in embryonal RMS. Int J Cancer. 1990;45:593–6.
Nisen PD, Zimmerman KA, Cotter SV, Gilbert F, Alt FW. Enhanced expression of the N-myc gene in Wilms’ tumors. Cancer Res. 1986;46:6217–22.
Tonelli R, Purgato S, Camerin C, Fronza R, Bologna F, Alboresi S, et al. Anti-gene peptide nucleic acid specifically inhibits MYCN expression in human neuroblastoma cells leading to cell growth inhibition and apoptosis. Mol Cancer Ther. 2005;4:779–86.
Burkhart CA, Cheng AJ, Madafiglio J, Kavallaris M, Mili M, Marshall GM, et al. Effects of MYCN antisense oligonucleotide administration on tumorigenesis in a murine model of neuroblastoma. J Natl Cancer Inst. 2003;95:1394–403.
Otto T, Horn S, Brockmann M, Eilers U, Schuttrumpf L, Popov N, et al. Stabilization of N-Myc is a critical function of Aurora A in human neuroblastoma. Cancer Cell. 2009;15:67–78.
Albihn A, Johnsen JI, Henriksson MA. MYC in oncogenesis and as a target for cancer therapies. Adv Cancer Res. 2010;107:163–224.
Vita M, Henriksson M. The Myc oncoprotein as a therapeutic target for human cancer. Semin Cancer Biol. 2006;16:318–30.
Zafar A, Wang W, Liu G, Wang X, Xian W, McKeon F, et al. Molecular targeting therapies for neuroblastoma: progress and challenges. Med Res Rev. 2021;41:961–1021.
Bell E, Chen L, Liu T, Marshall GM, Lunec J, Tweddle DA. MYCN oncoprotein targets and their therapeutic potential. Cancer Lett. 2010;293:144–57.
Chen H, Liu H, Qing G. Targeting oncogenic Myc as a strategy for cancer treatment. Signal Transduct Target Ther. 2018;3:5.
Mao L, Ding J, Zha Y, Yang L, McCarthy BA, King W, et al. HOXC9 links cell-cycle exit and neuronal differentiation and is a prognostic marker in neuroblastoma. Cancer Res. 2011;71:4314–24.
Peng W, Shi S, Zhong J, Liang H, Hou J, Hu X, et al. CBX3 accelerates the malignant progression of glioblastoma multiforme by stabilizing EGFR expression. Oncogene. 2022;41:3051–63.
Zhang K, Fu G, Pan G, Li C, Shen L, Hu R, et al. Demethylzeylasteral inhibits glioma growth by regulating the miR-30e-5p/MYBL2 axis. Cell Death Dis. 2018;9:1035.
Hou J, Deng Q, Zhou J, Zou J, Zhang Y, Tan P, et al. CSN6 controls the proliferation and metastasis of glioblastoma by CHIP-mediated degradation of EGFR. Oncogene. 2017;36:1134–44.
Hu X, Pei D, Ci M, Zhang G, Li B, Wang J, et al. ACTL6A deficiency induces apoptosis through impairing DNA replication and inhibiting the ATR-Chk1 signaling in glioblastoma cells. Biochem Biophys Res Commun. 2022;599:148–55.
Yang R, Li X, Wu Y, Zhang G, Liu X, Li Y, et al. EGFR activates GDH1 transcription to promote glutamine metabolism through MEK/ERK/ELK1 pathway in glioblastoma. Oncogene. 2020;39:2975–86.
Lee TH, Tun-Kyi A, Shi R, Lim J, Soohoo C, Finn G, et al. Essential role of Pin1 in the regulation of TRF1 stability and telomere maintenance. Nat Cell Biol. 2009;11:97–105.
Li Y, Su Y, Zhao Y, Hu X, Zhao G, He J, et al. Demethylzeylasteral inhibits proliferation, migration, and invasion through FBXW7/c-Myc axis in gastric cancer. MedComm. 2021;2:467–80.
Acknowledgements
This research was supported by the Natural Science Foundation of Chongqing (cstc2019jcyj-zdxmX0033, cstc2022ycjh-bgzxm0145), the pilot program of Southwest University (SWU-XDZD22006).
Author information
Authors and Affiliations
Contributions
HC, PL and XH conceived and designed the experiments. XH, JH, WP, SW, YL, GZ and XZ performed the experiments, collected the data. XH, RL and MX analyzed the data and prepared the figures. XH and HC wrote the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All experiments involving cancer patients’ samples were obtained from Chaoying Biotechnology Co., Ltd. (Henan, China), and the studies were approved by the Medical Ethics Committee of Tongxu County People’s Hospital of Henan Province. All of the patients were informed consent.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41388_2022_2428_MOESM4_ESM.jpg
Supplementary Figure 2. SMARCE1 is required for the colony-formation ability and proper cell cycle progression of MYCN-amplified neuroblastoma cells
41388_2022_2428_MOESM8_ESM.jpg
Supplementary Figure 6. SMARCE1 cooperate with MYCN to transcriptionally regulates the expression of PLK1, ODC1 and E2F2
41388_2022_2428_MOESM9_ESM.jpg
Supplementary Figure 7. The MYCN-SMARCE1 interaction is an important factor that regulates PLK1 expression and cell proliferation.
41388_2022_2428_MOESM10_ESM.jpg
Supplementary Figure 8. Analysis of the correlation between MYCN and SMARCE1 expression levels in several MYCN-amplified or overexpressed cancers
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, X., Liu, R., Hou, J. et al. SMARCE1 promotes neuroblastoma tumorigenesis through assisting MYCN-mediated transcriptional activation. Oncogene 41, 4295–4306 (2022). https://doi.org/10.1038/s41388-022-02428-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02428-1
- Springer Nature Limited