Abstract
Tumor invasion underlies further metastasis, the leading cause for cancer-related deaths. Deregulation of microRNAs has been identified associated with the malignant behavior of various cancers, including lung adenocarcinoma (LUAD), the major subtype of lung cancer. Here, we showed the significantly positive correlation between miR-629-5p level and tumor invasion in LUAD specimens (n = 49). In a human LUAD metastasis mouse model, H1650 cells (high level of miR-629-5p) were more aggressive than A549 cells (low level of miR-629-5p) in vivo, including higher incidence of vascular invasion and pulmonary colonization. Ectopic expression of miR-629-5p in A549 cells also increased their invasive capability. Then we identified that miR-629-5p promotes LUAD invasion in a mode of dual regulation via tumor cells invasion and endothelial cells permeability, respectively. In tumor cells, miR-629-5p enhanced motility and invasiveness of tumor cells by directly targeting PPWD1 (a cyclophilin), which clinically related to tumor invasion in LUAD specimens. Restoring PPWD1 protein significantly attenuated the invasion-promoting effects of miR-629-5p. Besides, exosomal-miR-629-5p secreted from tumor cells could be transferred to endothelial cells and increased endothelial monolayers permeability by suppressing CELSR1 (a nonclassic-type cadherin), which had a low level in the endothelial cells of invasive LUAD specimens. Activating the expression of CELSR1 in endothelial cells markedly blocked the effect of miR-629-5p. Our study suggests the dual roles of miR-629-5p in tumor cells and endothelial cells for LUAD invasion, implying a therapeutic option to targeting miR-629-5p using the “one stone, two birds” strategy in LUAD.
Similar content being viewed by others
Introduction
Lung cancer occupies the most percent of frequency in cancer morbidity and mortality worldwide [1], about 90% of lung cancer-related deaths arise from metastasis. In recent decades, lung adenocarcinoma (LUAD) becomes the primary histological subtype of lung cancer with the high incidence of metastasis [2, 3]. There are no visible symptoms during the early stages of LUAD, and tumor invasion or even metastasis had happened when it was detected [4]. Tumor metastasis is a complicated and multistep process, where tumor invasion turns on the switch for further metastatic steps [5, 6]. However, the mechanism of tumor invasion remains elusive. Thus, finding the potential targets for inhibiting tumor invasion might offer new strategies to prevent the metastasis of LUAD.
There is a growing body of the evidence showing that microRNAs (miRNAs) participate extensively in the tumorigenesis and progression [7,8,9]. MiRNAs are endogenous, highly conserved, and single-stranded noncoding RNAs within 25 nucleotides, functioning as the gene regulators at transcriptional and posttranscriptional levels [10]. Deregulated miRNAs in tumor cells have been identified to regulate the signaling pathways related to tumor proliferation, tumor invasion, and metastasis such as NF-κB, PI3K/AKT, and TGF-β pathways [11,12,13]. In addition, miRNAs in exosomes secreted from tumor cells play a critical role in the interactions among tumor cells and other components within tumor microenvironment, especially for the immune escape, tumor vascular invasion, and metastasis [14,15,16]. Therefore, it is well worth exploring the multiple molecular mechanisms of ectopic miRNAs in tumor invasion.
In this study, we found miR-629-5p promotes the cell migration and invasion in LUADs via inhibiting the gene expression level of PPWD1 (Peptidyl Prolyl isomerase domain and WD repeat-containing protein 1), which has been reported related to tumor metastasis [17, 18] in tumor cells. Besides, miR-629-5p secreted by tumor cells could downregulate CELSR1 (Cadherin EGF LAG seven-pass G-type receptors 1) in endothelial cells, leading to the increase of endothelial monolayers permeability. The dual roles of miR-629-5p in tumor cells and endothelial cells for tumor cell invasion in LUAD have been demonstrated. A therapeutic option for LUAD invasion to target miR-629-5p using the “one stone, two birds” strategy is indicated.
Results
MiR-629-5p was upregulated in LUAD and associated with LUAD invasion
To identify the LUAD invasion-related miRNAs, we performed migration and cell viability assays on A549 cells with 27 miRNAs, which were selected according to the reads per million and the ratio of mean value from TCGA datasets (Fig. 1a), and the results showed that miR-629-5p increased the number of migrated A549 cells more than three times compared with negative control, without promoting cell proliferation (Fig. 1b–d and Supplementary Fig. 1A). We also noticed the promotion effect of miR-21-5p in A549 cells migration, which had been confirmed previously [19,20,21]. To validate the upregulation of miR-629-5p in LUAD, we examined its expression levels in another 49 human LUADs and their paired adjacent peritumoral tissues, including 17 preinvasive specimens and 32 invasive specimens. The results from qRT-PCR with TaqMan MiRNA assay kit showed that miR-629-5p expression significantly upregulated in invasive tumor tissues compared with adjacent peritumoral tissues (Fig. 1e). Pathological analysis of clinic samples suggested tumor with high miR-629-5p showed an invasive histologic pattern with vascular invasions and spreads through air spaces [22], compared with noninvasive histologic pattern with no vascular invasions and no spread through air spaces in tumor with low miR-629-5p (Fig. 1f). We also assessed the expression of miR-629-5p in six LUAD cell lines (A549, H1975, H1299, Calu-3, H1650, and H358) and bronchial epithelial cells (BEAS-2B), the results showed that miR-629-5p expression is significantly higher in cell lines derived from invasive tumor compared with cell lines derived from primary tumor and BEAS-2B. The miR-629-5p expression level in H1650 cells was nearly 3.5-fold higher than A549 cells (Fig. 1g). These results implied a potential positive correlation between miR-629-5p and LUAD invasion.
a Schematic of the workflow for the identification of invasive associated miRNAs. T and N refer to the mean value of miRNA expression in tumor (n = 519) and normal tissues (n = 46) unpaired, respectively. RPM refers to read per million miRNAs. b Transwell migration assays of A549 cells transfected with miRNA mimics. c Images of transwell migration assays. Scale bar represents 10 μm. d Analysis of miR-629-5p expression of LUAD in pairs from the TCGA database (n = 46). e Analysis of miR-629-5p expression in paired LUAD tumor and normal tissues (n = 49). f Hematoxylin and eosin (HE) staining of LUAD tissue. Arrow in area 1 shows tumor cells that infiltrated the blood vessel. Arrows in area 2 show the air spaces spread of tumor cells. Scale bar represents 250 μm. g The miR-629-5p expression in tumor cells of three independent qPCR with different lysates. Mean ± SD are provided. *p < 0.05; **p < 0.01; ***p < 0.001.
Endogenous miR-629-5p promoted LUAD cells invasion in vitro and in vivo
To test whether miR-629-5p participates in the process of LUAD invasion, we first compared the migration and invasion ability of H1650 cells (high expression of miR-629-5p) and A549 cells (low expression of miR-629-5p) in vitro by transwell assay and wound healing assay. The results demonstrated that H1650 cells had the higher capability in cell migration and invasion than A549 cells (Fig. 2a and Supplementary Fig. 2A), and when endogenous miR-629-5p was blocked by transfecting the miR-629-5p specific antisense oligonucleotides (miR-629-5p inhibitor), the number of migrated H1650 cells was decreased about 50% (p < 0.001), and the invasion ability of H1650 cells were nearly 40% inhibited (p < 0.01) without affecting the cell cycle (Fig. 2b and Supplementary Fig. 2B–D). The same trend was also observed in A549 cells (Supplementary Fig. 2E, F). The results showed above indicate that miR-629-5p could elevate motility and invasiveness of LUAD cells in vitro. Next, we compared the two cell lines on the LUAD invasion in vivo. A549 cells and H1650 cells were injected into BALB/c nude mice via tail veins and sacrificed after 7 weeks. The lung metastasis incidence of H1650 cells was 100% versus 20% of A549 cells. Also, the number of metastatic nodules on the lung was markedly higher in mice injected with H1650 cells than mice injected with A549 cells (Fig. 2c). Notably, vascular invasions of tumor lesions were only observed in H1650 xenografts (Fig. 2d). These results suggest that endogenous miR-629-5p might be involved in the process of LUAD cells invasion and vascular invasion.
a Transwell migration and invasion assays of A549 cells and H1650 cells. Scale bar represents 10 μm. Result is the average of three independent experiments. b Transwell migration and invasion assays of H1650 cells transfected with miR-629-5p inhibitor for 48 h. Scale bar represents 10 μm. Result is the average of three independent experiments. c Representative lung images of mice injected with A549 cells and H1650 cells for 7 weeks. Arrows indicate the metastasis nodules on the lung. d HE staining of lung sections. Arrows indicate the tumor cells that invaded the blood vessel. Scale bar represents 200 μm. Mean ± SD are provided. *p < 0.05; **p < 0.01; ***p < 0.001.
Overexpression of miR-629-5p promoted A549 cells invasion in vitro and in vivo
To further explore the effect of miR-629-5p on LUAD invasion, we constructed a miR-629-5p-overexpressed A549 cell line (A549 Lv-629) using lentivirus (Supplementary Fig. 3A). Transwell assay and wound healing assay in vitro showed that overexpressing miR-629-5p increased over two times of migrated and invaded numbers of A549 cells (p < 0.001) (Fig. 3a and Supplementary Fig. 3B). Then A549 Lv-629 cells and A549 Lv-NC cells were injected intravenously into nude mice and kept for 7 weeks. The results showed that miR-629-5p overexpression significantly increased the incidence of lung metastasis and the number of metastasis foci (Fig. 3b). Vascular invasion also occurred in the sections of miR-629-5p overexpression group consistent with the results shown in Fig. 2d (Fig. 3c). Altogether, these results indicate that miR-629-5p is a positive regulator of LUAD invasion.
a Transwell migration and invasion assays of stably transduced A549 cells. Scale bar represents 10 μm. Result is the average of three independent experiments. b Representative lung images of mice injected with stably transduced A549 cells. Arrows indicated the metastasis nodules. c HE staining of lung sections. Arrow indicates the tumor cells that invaded the blood vessel. Scale bar represents 100 μm. Scale bar represents 250 μm. Mean ± SD are provided. *p < 0.05; **p < 0.01; ***p < 0.001.
PPWD1 was a downstream target of miR-629-5p in LUAD cells
To clarify the potential molecular mechanisms of miR-629-5p in LUAD invasion, we performed the proteomics analysis in A549 Lv-629 and A549 Lv-NC cells. Results showed that miR-629-5p inhibited the expression of PPWD1 manifestly (Fig. 4a). PPWD1 has been reported downregulation in gastroenteropancreatic neuroendocrine tumors metastases from the pancreas as the primary site [17, 18]. Therefore, first, we confirmed the downregulation effect of PPWD1 by miR-629-5p. We transfected miR-629-5p mimics into A549 cells for 48 h. There was a significant attenuation on the expression of PPWD1 (Fig. 4b). Besides, when we transfected miR-629-5p inhibitors into H1650 cells for 48 h, the mRNA and protein expression level of PPWD1 was markedly upregulated (Fig. 4c). To confirm whether endogenous miR-629-5p regulates PPWD1 expression, we tested the endogenous level of PPWD1 in LUAD cell lines. Results showed that H1650 and H358 cells exhibited less PPWD1 expression than A549 and H1299 cells (Supplementary Fig. 4A). To further examine whether miR-629-5p inhibits PPWD1 by targeting 3′UTR of PPWD1, we constructed the PPWD1 3′UTR reporter Luciferase assays in HEK293 cells showed that miR-629-5p suppressed the luciferase activities of PPWD1 3′UTR in a concentration-dependent way (Fig. 4d).
a The protein expression level of stably transduced A549 cells. b PPWD1 mRNA and protein levels in A549 cells transfected with miR-629-5p. Result is the average of three independent experiments. c PPWD1 mRNA and protein levels in H1650 cells transfected with miR-629-5p inhibitor. Result is the average of three independent experiments. d Luciferase assay of PPWD1 3′UTR in HEK293 cells transfected with miR-629-5p. Result is the average of three independent experiments. e The PPWD1 protein level in A549 cells transfected with PPWD1 siRNAs. f The promotion effects of PPWD1 siRNAs on cell migration and invasion were verified in A549 cells. Scale bar represents 10 μm. Result is the average of three independent experiments. g The expression of PPWD1 mRNA and protein in H1650 cells transfected with flag-PPWD1 vector or control vector. Result is the average of three independent experiments. h Overexpression of flag-PPWD1 inhibited the migration and invasion of H1650 cells. Scale bar represents 10 μm. Result is the average of three independent experiments. i The protein level in stably overexpression A549 cells transfected with flag-PPWD1 vector or control vector. j PPWD1 overexpression reversed the promotion effects of miR-629-5p on cell migration and invasion. Scale bar represents 10 μm. Result is the average of three independent experiments. Mean ± SD are provided. *p < 0.05; **p < 0.01; ***p < 0.001; ###p < 0.01.
Though miR-629-5p inhibited the PPWD1 expression, it was unclear whether PPWD1 participates in LUAD cells migration and invasion process. So we transfected siRNAs targeting PPWD1 into A549 cells, and the results showed that PPWD1 mRNA and protein expression were inhibited almost totally by PPWD1 siRNAs (Supplementary Fig. 4B and Fig. 4e). Next, we tested the effects of PPWD1 inhibition by transwell assay and wound healing assay. Notably, migration and invasion of A549 cells were enhanced when siRNA blocked PPWD1 expression (Fig. 4f and Supplementary Fig. 4C). Based on these results, we constructed a FLAG tagged-PPWD1 overexpression vector and transfected the vectors into H1650 cells, then we performed qRT-PCR and Western blot to test the expression of PPWD1, the level of PPWD1 mRNA and protein increased markedly after transfecting with flag-PPWD1 vectors compared with control vectors (Fig. 4g). Moreover, the migration and invasion of H1650 cells were reduced remarkably after PPWD1 overexpression with no influence on cell cycle (Fig. 4h and Supplementary Fig. 4D, E). Moreover, we also examined the function of PPWD1 on A549 Lv-629 cells, and results showed that PPWD1 overexpression reversed the promotion effect of miR-629-5p on cell migration and invasion abilities (Supplementary Fig. 4F–H and Fig. 4i, j). Hence, the loss of function and gain of function assays suggest PPWD1 is a downstream target of miR-629-5p in LUAD cells.
Next, we further validated the expression of PPWD1 in 49 paired LUAD clinic specimens. In keeping with Fig. 1e, the expression of PPWD1 significantly downregulated in invasive tumor tissues compared with paratumor tissues, no apparent change between preinvasive tumor tissues and paratumor tissues (Fig. 5a). Besides, a significant negative correlation exists expectedly between miR-629-5p expression and PPWD1 level within 49 paired LUAD clinic samples (Fig. 5b). Moreover, Immunohistochemistry (IHC) results showed that PPWD1 level was low in invasive LUAD tissues with higher miR-629-5p (Fig. 5c). Together, these data indicate that PPWD1 is a functional target of miR-629-5p in the process of LUAD cells migration and invasion.
LUAD-secreted exosomal-miR-629-5p induced HUVEC monolayers leakiness
Vascular invasion is an essential step for tumor metastasis [23,24,25,26], and exosomal miRNAs derived from tumor cells has been found playing a crucial part in the vascular invasion process such as miR-105 [16] and miR-25-3p [27]. As we have shown in Fig. 3c, vascular invasion occurred in lung sections of mice injected with miR-629-5p overexpression A549 cells. Therefore, we want to figure out whether miR-629-5p was involved in the process of LUAD intravasation. We first purified the exosomes from the conditioned media (CM) of LUAD cell lines and A549 Lv-629 cells (Supplementary Fig. 5A–C). Then we analyzed the expression of miR-629-5p by qRT-PCR, using cel-miR-39 as a loading control [28]. It was interesting that the expression of miR-629-5p in exosomes was consistent with the expression of cell lines and stably transduced A549 cells (Fig. 6a, b). Then we performed endothelial permeability assays to examine the effect of exosomal-miR-629-5p on the Human Umbilical Vein Endothelial Cells (HUVECs) (Fig. 6c). As the results showed, the miR-629-5p level significantly increased in HUVECs cocultured with A549 Lv-629 cells, and there was no significant change of miR-629-5p level in HUVECs when pretreating A549 cells with 20 μM GW4869, a neutral sphingomyelinase inhibitor [29] (Fig. 6d). The permeability of HUVEC monolayers was increased cocultured with A549 Lv-629 cells. However, this effect vanished when pretreating A549 with GW4869 and miR-629-5p inhibitor (Fig. 6e, f). Further, transendothelial invasion assays (Fig. 6g) indicated that miR-629-5p promoted the invasion of A549-GFP(Lv-629) through HUVEC monolayers significantly, and the number of invasive A549-GFP decreased when the cocultured A549 cells pretreated with GW4869 or miR-629-5p inhibitor (Fig. 6h, i). Also, we transfected miR-629-5p mimics into HUVECs directly, the effects of miR-629-5p mimics on HUVECs were consistent with the results showed in Fig. 6 (Supplementary Fig. 6A, B). These data indicate that HUVECs could catch the miR-629-5p secreted by LUAD cells and induce HUVEC monolayers leakiness.
a The expression of miR-629-5p in exosomes derived from tumor cells. Result is the average of three independent experiments. b The expression of miR-629-5p in stably overexpression A549-secreted exosomes. Result is the average of three independent experiments. c Experimental scheme of endothelial permeability assay. d Analysis of miR-629-5p expression in HUVECs cocultured with stably transduced A549 cells. Result is the average of three independent experiments. e, f Permeability of the HUVEC monolayers after cocultured with stably transduced A549 cells treated with GW4869 or transfected with miR-629-5p inhibitor, respectively. Result is the average of three independent experiments. g Experimental scheme of transendothelial invasion assay. h, i A549-GFP(Lv-629) transendothelial invasion in HUVEC treated with GW4869 or transfected with miR-629-5p inhibitor, respectively. Scale bar represents 10 μm. Result is the average of three independent experiments. Mean ± SD are provided. *p < 0.05; **p < 0.01; #p < 0.05; ##p < 0.01.
LUAD-derived miR-629-5p targeted CELSR1 gene in HUVECs
To explore the molecular mechanism of miR-629-5p in HUVECs, we detected 11 potential targets related to the integrity of endotheliocytes [30,31,32,33,34,35,36,37]. The results showed that mRNA of CELSR1 downregulated after transfected with miR-629-5p (Fig. 7a). We further confirmed the inhibition of CELSR1 by miR-629-5p using Western blot and luciferase assays (Fig. 7b, c). Then, siRNAs targeting CELSR1 were used to investigate the role of CELSR1 for endothelial cell integrity. The results showed that the permeability and the number of invasive tumor cells significantly increased when inhibiting CELSR1 by CELSR1 siRNA (Fig. 7d–f). Moreover, we activated the expression of CELSR1 by CRISPR/dcad9 activation plasmids (Fig. 7g). As the results showed, the effects of miR-629-5p were reversed almost totally when the expression of CELSR1 was activated (Fig. 7h, i). Then we tested the CELSR1 level of endothelial cells in LUAD specimens. The results showed that the expression of CELSR1 in endothelial cells was downregulated obviously in invasive LUAD samples with a higher level of miR-629-5p (Fig. 8a). Finally, our data show that the miR-629-5p regulation on CELSR1 expression in endothelial cells may contribute to the vascular invasion of LUAD cells.
a The mRNA level in HUVECs transfected with miR-629-5p. Result is the average of three independent experiments. b The protein level of CELSR1 in HUVECs transfected with miR-629-5p. c Luciferase assay of CELSR1 3′UTR in HEK293 cells transfected with miR-629-5p. Result is the average of three independent experiments. d The mRNA and protein expression of CELSR1 in HUVECs transfected with CELSR1 siRNAs. Result is the average of three independent experiments. Permeability (e) and transendothelial invasion assay (f) of the HUVEC monolayers after transfected with CELSR1 siRNAs. Scale bar represents 10 μm. Result is the average of three independent experiments. h, i Activating CELSR1 expression with CRISPR/dcas9 activation plasmid in HUVEC reversed the promotion effects of miR-629-5p on permeability and transendothelial invasion. Scale bar represents 10 μm. Result is the average of three independent experiments. Mean ± SD are provided. *p < 0.05; **p < 0.01; #p < 0.05; ##p < 0.01.
Discussion
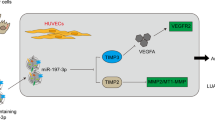
Epigenetic changes including miRNA dysregulation are critical events in the acquiring of invasive ability in cancer cells [38,39,40,41]. Here, we disclosed the dual roles of miR-629-5p in this process via enhancing the invasiveness of cancer cell itself and exosomally increasing the permeability of endothelial barrier (Fig. 8b). MiR-629-5p specifically upregulated in tumor tissues and cell lines of invasive LUAD. Endogenous and ectopic expression of miR-629-5p enhanced the invasion of LUAD cells in vitro and in vivo. Inhibition of PPWD1 by miR-629-5p subsequently promotes LUAD cells migration and invasion. Besides, LUAD cells derived exosomal-miR-629-5p is caught by endothelial cell and inhibits CELSR1 expression, inducing the leakiness of endothelial barrier.
Emerging evidence suggests that the mode of dual or multiple targets exists widely in the miRNAs regulation of tumor progression, including tumor proliferation, tumor metastasis, and multidrug resistance of tumor. Previous study showed that miR-1254 could attenuate nonsmall cell lung cancer (NSCLC) growth via suppressing HO-1 and TFAP2A expression [42]. MiR-140-5p was identified as a negative regulator of tumor growth and metastasis by targeting TGF-β and FGF9 in hepatocellular carcinoma [13]. In breast cancer, miR-200a promoted chemoresistance of breast cancer cells by antagonizing TP53INP1 and YAP1 [43]. As the direct targets of miR-21, PIK3R1, TIMP3, EGF, and TM1 were involved in the migration and invasion of breast cancer cells [44,45,46,47]. These modes of dual targets and multitargets imply the coordination and stages of miRNAs regulation, which develop new perspective for understanding the mechanisms of tumor progression.
Tumor invasion-metastasis cascade is a multistep process where tumor invasion takes the first step for metastasis [5, 6]. Vascular invasion of tumor cells is the dominating feature of tumor invasion, which provides precondition for the survival of tumor cells in distant sites [22]. Previous studies suggested that miR-629-5p involved in the process of metastatic dissemination in various types of cancers including NSCLC, but the roles of miR-629-5p in LUAD and the correlation between miR-629-5p level and the change of LUAD pathological characteristics has not been clarified clearly. Previously, miR-629-5p was found to promote tumor cell metastasis in clear cell renal cell carcinoma by targeting tripartite motif-containing 33, a suppressor of the TGFβ/Smad pathway [48]. Yang et al. found a functional polymorphism at the miR-629-binding site in the 3′UTR of the NBS1 gene and related to the risk of lung cancer [49]. MiR-629 was also found promoting tumor progression in colorectal cancer, ovarian cancer, and pancreatic cancer [50,51,52]. In our study, we revealed the positive correlation between miR-629-5p level and LUAD invasive characteristic in a certain regulation of dual-targets mode, which expanded the underlying mechanism of LUAD invasion regarding the new targets of miR-629-5p within tumor microenvironment. As the direct target of miR-629-5p in tumor cells, PPWD1 is a multidomain cyclophilin, which participates in the process of chromatin modification, transcription, and pre-mRNA splicing [53, 54]. Previous studies found that testing the expression of PPWD1 in metastases could predict the localization of the primary tumor in gastroenteropancreatic neuroendocrine neoplasm [18]. We examined the inhibition effects of PPWD1 on cell migration and invasion by overexpressing and knocking down PPWD1 in LUAD cells, showing the critical role of PPWD1 in LUAD cells migration and invasion. Meanwhile, RUNX3 was reported as the functional target of miR-629-5p in NSCLC cells invasion [55], and we also verified the inhibition effect of miR-629-5p on RUNX3 in our study (Supplementary Fig. 7), which offered another proof to extend the multitargets modes of miRNA regulation. Moreover, we identified that CELSR1 is the functional target of miR-629-5p in endothelial cells. CELSR1 belongs to atypical cadherins family [56], and plays a vital role in the epithelial planar cell polarity [57], the growth and differentiation of kidney and fate decision of neural progenitor [58], indicating CELSR1 is very important for normal physiological function. In our study, we found miR-629-5p inhibited CELSR1 expression markedly in HUVECs and examined the function of CELSR1 for endothelial integrity by suppressing and overexpressing CELSR1 in HUVECs, indicating the role of CELSR1 in LUAD cells vascular invasion. Still, there are some details during this process remain unclear, such as the downstream mechanisms of PPWD1, and the possible other effects of miR-629-5p within the tumor microenvironment. Exploration of these questions is meaningful for understanding the mechanisms of LUAD invasion in depth.
It remains mostly incurable for manifest metastasis. Thus, early intervention of tumor invasion may be an effective strategy for metastasis prevention. Our experimental analyses and clinical studies demonstrate the key roles of miR-629-5p in LUAD invasion, indicating the possibility of miR-629-5p as LUAD invasion-related marker and targeting miR-629-5p might offer a “one stone, two birds” strategy for the oncotherapy of LUAD invasion.
Materials and methods
Human LUAD samples
Collections of the human LUAD specimens for this study were approved by the ethics committees of Huadong Hospital, Fudan University. These samples were stored in liquid nitrogen for further analyses.
Cell lines
All cell lines in this study were obtained from the American Type Culture Collection (ATCC, USA) and incubated at 37 °C with 5% CO2. The human LUAD cell lines, including A549, H1975, H1299, Calu-3, H1650, and H358, were cultured in RPMI1640 (Gibco, USA) with 10% fetal bovine serum (Gibco). The bronchial epithelial cell line (BEAS-2B) was cultured with BEBM medium (Lonza, Switzerland). The HUVEC was cultured in ECM medium (ScienCell, USA). The human embryonic kidney cell line (HEK293) was cultured in DMEM (Hyclone, USA) with 10% fetal bovine serum. All cell lines were authenticated by STR and tested for mycoplasma contamination.
Cell migration and invasion assays
The cell invasion and migration assays were performed using the Transwell Permeable Supports with 8 μm pore (Corning, USA). In total, 1.5 × 105 cells/cm2 were seeded into apical chambers with the basement membrane pretreated with (for cell invasion assay) or without (for cell migration assay) 20 μg matrigel matrix (Corning, USA). After 24 h, the transwell was fixed with 4% PFA for 15 min and stained with hematoxylin for 15 min and the cells on the upper side of basement membrane were gently removed by cotton swabs. The cells on the lower side of basement membrane were imaged for four random microscopic fields (magnification, 20×), and ImageJ was used to count the cell number.
Wound healing assay
For wound healing assay, 2 × 105 cells were seeded to confluence in a 12-well plate, scratch wounds were made using a 200 µl pipette tip. Then, cells were cultured in medium without fetal bovine serum. Images were captured at 24 h, 72 h, and wound healing percentage was calculated by ImageJ.
Flow cytometric analyses of cell cycle
For cell cycle analysis, treated cells were collected by trypsin, then, cells were incubated with propidium iodide (5 mg/ml) (Sigma, USA), and were analyzed using cytoFlEX (Beckman, USA).
Mass spectrometry analysis
Numerous proteins were markedly changed after overexpressing miR-629-5p from the quantitative protein profiling of A549 cells. Forty-three proteins were downregulated and 58 proteins were upregulated by more than 1.2-fold in miR-629-5p-overexpressed A549 cells (Fig. 4a).
Plasmids, viruses, and RNA oligonucleotides
Human PPWD1 cDNA without 3′UTR were cloned into the p3xFLAG-myc-CMVTM-24 vector (Sigma, USA) for the overexpression PPWD1, using the primers: PPWD1 forward, 5′-TTGCGGCCGCATGAGGAGAACGAAGAGCGCT-3′; PPWD1 reverse, 5′-GGGGTACCTCACTTGACAGTAATATTTATGATGCTG3′. CELSR1 CRISPR Activation Plasmids were purchased for the overexpression of CELSR1 (Santa Cruz, USA). The 3′UTR of PPWD1 and CELSR1 were cloned into the psi-CHECK-2 vector (Promega, Madison, WI, USA), respectively, to generate the reporter constructs for PPWD1 and CELSR1 gene, using the primers as follows:
PPWD1 3′UTR forward, 5′-CCTCGAGAACTCCATACATCACCTCTTACTCAGA-3′; PPWD1 3′UTR reverse, 5′-TTGCGGCCGCACATGATCTGTATATTCAGCAAGTCCT-3′; CELSR1 3′UTR forward, 5′-CCCTCGAGCTGTCAGACCTGGTGACAAGT-3′; CELSR1 3′UTR reverse, 5′-TTGCGGCCGCAACAGAGCCTCAGGGACCA-3′. The lentivirus-hsa-miR-629 (Genechem) was purchased to generate the miR-629-overexpressed cell lines. MiRNA mimics, anti-miR-629-5p, siRNA targeting PPWD1 and CELSR1, or their negative control RNAs were obtained from GenePharma (Suzhou, China). These RNAs (10 nM) were transfected with Lipofectamine RNAiMAX Transfection Reagent (Invitrogen, USA) The sequences were listed as follows: anti-miR-629-5p: 5′-AGUUCUCCCAACGUAAACCCA-3′; siPPWD1-1: 5′-GCUUAUCCAACCAGCGUAUTT-3′; siPPWD1-2: 5′-GCUGCAACUACUAUAGAAATT-3′; siCELSR1-1: 5′-CCACCUACAUCACUGUCUUTT-3′; siCELSR1-2: 5′-CCAACAAGUCCAACAGUUUTT-3′.
Reverse transcriptase (RT)-PCR and qRT-PCR
Total RNA was extracted from cells, tissues, or exosomes using Trizol reagent (Takara, Japan) according to the protocol. For mRNA expression, RT reactions were performed using PrimeScript RT Master Mix (Takara), qRT-PCR was performed with TB Green Premix Ex Taq (Takara) and analyzed on 7500 Fast Real-Time PCR System (Thermo Fisher Scientific, USA). GAPDH gene was used as a reference for mRNA expression. The primer sequences were listed as follows: PPWD1 forward, 5′-TGGCCGAGGAGATAACCAG-3′; PPWD1 reverse, 5′-CCCAGTCCAGTATTCAATCATCC-3′; CELSR1 forward, 5′-GGCGTTGTTTGAGAACGAACC-3′; CELSR1 reverse, 5′-AGAGTCGATTCGGAAGTAGCC-3′; CLDN1 forward, 5′-CCTCCTGGGAGGATAGCAAT-3′; CLDN1 reverse, 5′-GGCAACTAAAATAGCCAGACCT-3′; CLDN2 forward, 5′-GCCTCTGGATGGAATGTGCC-3′; CLDN2 reverse, 5′-GCTACCGCCACTCTGTCTTTG-3′; CLDN9 forward, 5′-CGGCTGCACTGCTTATGCT-3′; CLDN9 reverse, 5′-GAGGGGATGGAGTAGCCCA-3′; JAM2 forward, 5′-GCAGTAGAGTACCAAGAGGCT-3′; JAM2 reverse, 5′-AGACACTCCGACCCAGTTTCT-3′; JAM3 forward, 5′-CGGCTGCCTGACTTCTTCC-3′; JAM3 reverse, 5′-TGGGGTTCGATTGCTGGATTT-3′; OCLN forward, 5′-ACAAGCGGTTTTATCCAGAGTC-3′; OCLN reverse, 5′-GTCATCCACAGGCGAAGTTAAT-3′; ZO-1 forward, 5′-CAACATACAGTGACGCTTCACA-3′; ZO-1 reverse, 5′-CACTATTGACGTTTCCCCACTC-3′; ZO-2 forward, 5′-ATGGAAGAGCTGATATGGGAACA-3′; ZO-2 reverse, 5′-TGCTGAACTGCAAACGAATGAA-3′. For miRNA expression, RT reactions were performed using the TaqMan MiRNA assay kit (Thermo Fisher Scientific). U6 was used as a reference for cellular miRNA expression, and cel-miR-39 was used as a reference for exosomal miRNA.
Western blot analysis
Total protein lysates were quantified with Pierce BCA Protein Assay Kit (Thermo Fisher Scientific), separated by 10% SDS-PAGE and transferred to PVDF membranes (Millipore, USA), then incubated with antibodies against PPWD1 (HPA019360, Sigma, USA), CELSR1 (ab225889, Abcam, UK), α-Tublin (OM239217, Cell Signaling Technology, USA), and RUNX3 (ab224641, Abcam, UK), respectively. Following incubated with Peroxidase-conjugated Affinipure Goat Anti-Mouse lgG (Jackson ImmunoResearch, USA) and Peroxidase-conjugated Affinipure Goat Anti-rabbit lgG (Jackson ImmunoResearch) at room temperature for 1 h. α-tublin was used as an inner loading control. The epitope was visualized by lumi Q ECL regent solution kit (Share-Bio, China) according to the manufacturer’s instructions.
Luciferase reporter assay
HEK293 cells seeded in 96-well plates were cotransfected with miR-629-5p and reporter constructs by lipofectamine 3000. After 24 h, the luciferase assay was performed using the Dual-Luciferase Reporter Assay System (Promega, USA). Luminescence was detected using the Biotek Synergy 4 (Biotex, USA).
Exosome purification, electron microscopy (EM), and nanoparticle tracking analysis (NTA)
Exosomes in CM were purified by ultracentrifugation as described [59]. For EM [60], PBS containing exosomes was fixed by 4% paraformaldehyde with equal volume, applied to the formvar-coated grids and adhered for 20 min, blotted up PBS by filter paper (Whatman, UK) gently, infiltrated the grid by 1% uranyl formate solution for 1 min and repeated two times. Images were acquired by a Transmission Electron Microscope (Tecnai G2 spirit 120 kV, FEI, USA). NTA was performed to estimate the particle size of exosomes by NanoSight NS300 (Malvern, UK).
Endothelial permeability and transendothelial invasion assays
In total, 1 × 106/cm2 HUVEC cells and 1.5 × 105/cm2 stably transduced A549 cells (A549 Lv-629/A549 Lv-NC) were seeded in the top well and bottom well of the Transwell Permeable Supports with 0.4 μm pore separately, and cocultured for 72 h. Tetramethylrhodamine isothiocyanate-Dextran (MW20000, Sigma) was added to the upper well at 10 mg/ml for 1 h, and the fluorescence intensity of the bottom well was monitored using Biotek Synergy 4 (Biotek, USA) (Ex = 550 nm, Em = 590 nm). For transendothelial invasion assay, HUVEC monolayers cocultured with stably transduced A549 cells (A549 Lv-629/A549 Lv-NC) for 72 h, 1.5 × 105/cm2 GFP-labeled A549 cells were seed into the top well of the Transwell Permeable Supports with 8 μm pore, after 24 h, cells on the lower side of the chamber were imaged and counted.
Immunohistochemistry (IHC) and immunofluorescence (IF) staining
Five-micrometer-thick paraffin sections of human LUAD patients were obtained from Huadong Hospital, Fudan University. Antigen repair was performed for 10 min in the citric acid buffer (PH 6.0), Then, the treated sections were incubated with primary antibodies at 4 °C overnight. For IF staining, CELSR1 antibody (ABT-119, Sigma, USA) and CD31 antibody (11265-1-AP, Proteintech, USA) were used as primary antibodies, and Alexa Fluor 488 (green) (Invitrogen, USA) was used for secondary antibody to CELSR1, Alexa Flour 555 (red) (Invitrogen) was used for secondary antibody to CD31. For IHC staining, MaxVision HRP-Polymer anti-Rabbit IHC Kit (MXB, China) was used for secondary antibody incubation for 30 min, and DAB Kit (Zhongshanjinqiao, China) was used for PPWD1 visualization. An IHC score was assessed by one pathologist (HL) according to percentage of immunostained cells (0–3) and staining intensity (0–3).
Animal studies
Animal studies were approved by the Institutional Animal Care and Use Committee of Shanghai Institute of Materia Medica, Chinese Academy of Sciences. In total, 4–6-week-old female BALB/c nude mice (Jihui, China) were used for metastasis assays. To compare the invasive ability of A549 cells and H1650 cells, 18 mice were randomly allocated to A549 group (n = 10) and H1650 group (n = 8). To explore the effect of miR-629-5p on LUAD invasion, 25 mice were randomly allocated to A549 Lv-NC group (n = 13) and A549 Lv-629 group (n = 14). After 1 week of acclimation, 2 × 106 cells resuspended in 200 μl saline were injected into the lateral tail veins of nude mice. After 7 weeks, mice sacrificed the lungs were collected to count the metastatic nodules on surface, and then these tissues were fixed in 4% paraformaldehyde for further hematoxylin and eosin staining.
Statistics
All data were analyzed by GraphPad Prism 6 software (GraphPad Software, USA), and were shown as mean ± standard deviation (SD). Student’s t test was used to measure the differences between two groups, all analyses were two tailed, and p < 0.05 was considered significant.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Cancer J Clin. 2018;68:394–424.
Riihimaki M, Hemminki A, Fallah M, Thomsen H, Sundquist K, Sundquist J, et al. Metastatic sites and survival in lung cancer. Lung Cancer. 2014;86:78–84.
Myers DJ, Wallen JM. Cancer, lung adenocarcinoma. Treasure Island, FL: StatPearls; 2019.
Zhao Y, Adjei AA. New strategies to develop new medications for lung cancer and metastasis. Cancer Metastasis Rev. 2015;34:265–75.
van Zijl F, Krupitza G, Mikulits W. Initial steps of metastasis: cell invasion and endothelial transmigration. Mutat Res. 2011;728:23–34.
Jin K, Li T, van Dam H, Zhou F, Zhang L. Molecular insights into tumour metastasis: tracing the dominant events. J Pathol. 2017;241:567–77.
Paliouras AR, Monteverde T, Garofalo M. Oncogene-induced regulation of microRNA expression: implications for cancer initiation, progression and therapy. Cancer Lett. 2018;421:152–60.
Chorti A, Bangeas P, Papavramidis TS, Tsoulfas G. Role of MicroRNA in the diagnosis and therapy of hepatic metastases from colorectal cancer. Microrna. 2018;7:167–77.
Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer. 2015;15:321–33.
Miao L, Yao H, Li C, Pu M, Yao X, Yang H, et al. A dual inhibition: microRNA-552 suppresses both transcription and translation of cytochrome P450 2E1. Biochim Biophys Acta. 2016;1859:650–62.
Wang T, Xu X, Xu Q, Ren J, Shen S, Fan C, et al. miR-19a promotes colitis-associated colorectal cancer by regulating tumor necrosis factor alpha-induced protein 3-NF-kappaB feedback loops. Oncogene. 2017;36:3240–51.
Yang N, Chen J, Zhang H, Wang X, Yao H, Peng Y, et al. LncRNA OIP5-AS1 loss-induced microRNA-410 accumulation regulates cell proliferation and apoptosis by targeting KLF10 via activating PTEN/PI3K/AKT pathway in multiple myeloma. Cell Death Dis. 2017;8:e2975.
Yang H, Fang F, Chang R, Yang L. MicroRNA-140-5p suppresses tumor growth and metastasis by targeting transforming growth factor beta receptor 1 and fibroblast growth factor 9 in hepatocellular carcinoma. Hepatology. 2013;58:205–17.
Graner MW, Schnell S, Olin MR. Tumor-derived exosomes, microRNAs, and cancer immune suppression. Semin Immunopathol. 2018;40:505–15.
Chen L, Gibbons DL, Goswami S, Cortez MA, Ahn YH, Byers LA, et al. Metastasis is regulated via microRNA-200/ZEB1 axis control of tumour cell PD-L1 expression and intratumoral immunosuppression. Nat Commun. 2014;5:5241.
Zhou W, Fong MY, Min Y, Somlo G, Liu L, Palomares MR, et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell. 2014;25:501–15.
Kaemmerer D, Posorski N, von Eggeling F, Ernst G, Horsch D, Baum RP, et al. The search for the primary tumor in metastasized gastroenteropancreatic neuroendocrine neoplasm. Clin Exp Metastasis. 2014;31:817–27.
Posorski N, Kaemmerer D, Ernst G, Grabowski P, Hoersch D, Hommann M, et al. Localization of sporadic neuroendocrine tumors by gene expression analysis of their metastases. Clin Exp Metastasis. 2011;28:637–47.
Zhu S, Wu H, Wu F, Nie D, Sheng S, Mo YY. MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res. 2008;18:350–9.
Asangani IA, Rasheed SA, Nikolova DA, Leupold JH, Colburn NH, Post S, et al. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene. 2008;27:2128–36.
Bica-Pop C, Cojocneanu-Petric R, Magdo L, Raduly L, Gulei D, Berindan-Neagoe I. Overview upon miR-21 in lung cancer: focus on NSCLC. Cell Mol Life Sci. 2018;75:3539–51.
Travis WD, Brambilla E, Nicholson AG, Yatabe Y, Austin JHM, Beasley MB, et al. The 2015 world health organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol. 2015;10:1243–60.
Nieto MA. Epithelial plasticity: a common theme in embryonic and cancer cells. Science. 2013;342:1234850.
Singleton PA. Hyaluronan regulation of endothelial barrier function in cancer. Adv Cancer Res. 2014;123:191–209.
Bielenberg DR, Zetter BR. The contribution of angiogenesis to the process of metastasis. Cancer J. 2015;21:267–73.
Maishi N, Hida K. Tumor endothelial cells accelerate tumor metastasis. Cancer Sci. 2017;108:1921–6.
Zeng Z, Li Y, Pan Y, Lan X, Song F, Sun J, et al. Cancer-derived exosomal miR-25-3p promotes pre-metastatic niche formation by inducing vascular permeability and angiogenesis. Nat Commun. 2018;9:5395.
Poel D, Buffart TE, Oosterling-Jansen J, Verheul HM, Voortman J. Evaluation of several methodological challenges in circulating miRNA qPCR studies in patients with head and neck cancer. Exp Mol Med. 2018;50:e454.
Faict S, Muller J, De Veirman K, De Bruyne E, Maes K, Vrancken L, et al. Exosomes play a role in multiple myeloma bone disease and tumor development by targeting osteoclasts and osteoblasts. Blood Cancer J. 2018;8:105.
Zhan YH, Luo QC, Zhang XR, Xiao NA, Lu CX, Yue C, et al. CELSR1 is a positive regulator of endothelial cell migration and angiogenesis. Biochemistry. 2016;81:591–9.
Liu K, Wang XJ, Li YN, Li B, Qi JS, Zhang J, et al. Tongxinluo reverses the hypoxia-suppressed Claudin-9 in cardiac microvascular endothelial cells. Chin Med J. 2016;129:442–7.
Tornavaca O, Chia M, Dufton N, Almagro LO, Conway DE, Randi AM, et al. ZO-1 controls endothelial adherens junctions, cell-cell tension, angiogenesis, and barrier formation. J Cell Biol. 2015;208:821–38.
Lee SW, Kim WJ, Jun HO, Choi YK, Kim KW. Angiopoietin-1 reduces vascular endothelial growth factor-induced brain endothelial permeability via upregulation of ZO-2. Int J Mol Med. 2009;23:279–84.
Jenkinson EM, Livingston JH, O’Driscoll MC, Desguerre I, Nabbout R, Boddaert N, et al. Comprehensive molecular screening strategy of OCLN in band-like calcification with simplified gyration and polymicrogyria. Clin Genet. 2018;93:228–34.
Aurrand-Lions M, Duncan L, Ballestrem C, Imhof BA. JAM-2, a novel immunoglobulin superfamily molecule, expressed by endothelial and lymphatic cells. J Biol Chem. 2001;276:2733–41.
Naik MU, Naik UP. Junctional adhesion molecule-A-induced endothelial cell migration on vitronectin is integrin alpha v beta 3 specific. J Cell Sci. 2006;119:490–9.
Xiaohua T, Dongmei L, Xiaobo W, Yan Z, Yutao Y, Lei Y. Claudin-2 downregulation by KSHV infection is involved in the regulation of endothelial barrier function. J Cutan Pathol. 2014;41:630–9.
Ito M, Teshima K, Ikeda S, Kitadate A, Watanabe A, Nara M, et al. MicroRNA-150 inhibits tumor invasion and metastasis by targeting the chemokine receptor CCR6, in advanced cutaneous T-cell lymphoma. Blood. 2014;123:1499–511.
Zhang BG, Li JF, Yu BQ, Zhu ZG, Liu BY, Yan M. microRNA-21 promotes tumor proliferation and invasion in gastric cancer by targeting PTEN. Oncol Rep. 2012;27:1019–26.
Wen J, Zhao YK, Liu Y, Zhao JF. MicroRNA-34a inhibits tumor invasion and metastasis in osteosarcoma partly by effecting C-IAP2 and Bcl-2. Tumour Biol. 2017;39:1010428317705761.
Chang TH, Tsai MF, Gow CH, Wu SG, Liu YN, Chang YL, et al. Upregulation of microRNA-137 expression by Slug promotes tumor invasion and metastasis of non-small cell lung cancer cells through suppression of TFAP2C. Cancer Lett. 2017;402:190–202.
Pu M, Li C, Qi X, Chen J, Wang Y, Gao L, et al. MiR-1254 suppresses HO-1 expression through seed region-dependent silencing and non-seed interaction with TFAP2A transcript to attenuate NSCLC growth. PLoS Genet. 2017;13:e1006896.
Yu SJ, Yang L, Hong Q, Kuang XY, Di GH, Shao ZM. MicroRNA-200a confers chemoresistance by antagonizing TP53INP1 and YAP1 in human breast cancer. BMC Cancer. 2018;18:74.
Qi L, Bart J, Tan LP, Platteel I, Sluis T, Huitema S, et al. Expression of miR-21 and its targets (PTEN, PDCD4, TM1) in flat epithelial atypia of the breast in relation to ductal carcinoma in situ and invasive carcinoma. BMC Cancer. 2009;9:163.
Han M, Wang F, Gu Y, Pei X, Guo G, Yu C, et al. MicroRNA-21 induces breast cancer cell invasion and migration by suppressing smad7 via EGF and TGF-beta pathways. Oncol Rep. 2016;35:73–80.
Nagao Y, Hisaoka M, Matsuyama A, Kanemitsu S, Hamada T, Fukuyama T, et al. Association of microRNA-21 expression with its targets, PDCD4 and TIMP3, in pancreatic ductal adenocarcinoma. Mod Pathol. 2012;25:112–21.
Yan LX, Liu YH, Xiang JW, Wu QN, Xu LB, Luo XL, et al. PIK3R1 targeting by miR-21 suppresses tumor cell migration and invasion by reducing PI3K/AKT signaling and reversing EMT, and predicts clinical outcome of breast cancer. Int J Oncol. 2016;48:471–84.
Jingushi K, Ueda Y, Kitae K, Hase H, Egawa H, Ohshio I, et al. miR-629 targets TRIM33 to promote TGFbeta/Smad signaling and metastatic phenotypes in ccRCC. Mol Cancer Res. 2015;13:565–74.
Yang L, Li Y, Cheng M, Huang D, Zheng J, Liu B, et al. A functional polymorphism at microRNA-629-binding site in the 3’-untranslated region of NBS1 gene confers an increased risk of lung cancer in Southern and Eastern Chinese population. Carcinogenesis. 2012;33:338–47.
Shao L, Shen Z, Qian H, Zhou S, Chen Y. Knockdown of miR-629 inhibits ovarian cancer malignant behaviors by targeting testis-specific Y-Like protein 5. DNA Cell Biol. 2017;36:1108–16.
Yan H, Li Q, Wu J, Hu W, Jiang J, Shi L, et al. MiR-629 promotes human pancreatic cancer progression by targeting FOXO3. Cell Death Dis. 2017;8:e3154.
Lu J, Lu S, Li J, Yu Q, Liu L, Li Q. MiR-629-5p promotes colorectal cancer progression through targetting CXXC finger protein 4. Biosci Rep. 2018;38:BSR20180613.
Rajiv C, Davis TL. Structural and functional insights into human nuclear cyclophilins. Biomolecules. 2018;8:E161.
Davis TL, Walker JR, Ouyang H, MacKenzie F, Butler-Cole C, Newman EM, et al. The crystal structure of human WD40 repeat-containing peptidylprolyl isomerase (PPWD1). FEBS J. 2008;275:2283–95.
Zhuang B, Cheng Y. MicroRNA629 inhibition suppresses the viability and invasion of nonsmall cell lung cancer cells by directly targeting RUNX3. Mol Med Rep. 2019;19:3933–40.
Goffinet AM, Tissir F. Seven pass cadherins CELSR1-3. Semin Cell Dev Biol. 2017;69:102–10.
Duncan JS, Stoller ML, Francl AF, Tissir F, Devenport D, Deans MR. Celsr1 coordinates the planar polarity of vestibular hair cells during inner ear development. Dev Biol. 2017;423:126–37.
Brzoska HL, d’Esposito AM, Kolatsi-Joannou M, Patel V, Igarashi P, Lei Y, et al. Planar cell polarity genes Celsr1 and Vangl2 are necessary for kidney growth, differentiation, and rostrocaudal patterning. Kidney Int. 2016;90:1274–84.
Thery C, Amigorena S, Raposo G, Clayton A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol. 2006;3:3.22.1–29.
Kang Y, Kuybeda O, de Waal PW, Mukherjee S, Van Eps N, Dutka P, et al. Cryo-EM structure of human rhodopsin bound to an inhibitory G protein. Nature. 2018;558:553–8.
Acknowledgements
This work was supported by the National Science & Technology Major Project “Key New Drug Creation and Manufacturing Program,” China (2018ZX09101001-003-007 and 2018ZX09711002-010-001), “the National Natural Science Foundation of china (91859106),” One Hundred Talent Program of Chinese Academy of Sciences, the Fundamental and Open Research Funds from the State Key Laboratory of Drug Research, and the grant from Shanghai Committee of Science and Technology, China (Grant No.: 18DZ2290200). We thank the Electron Microscopy System and Animal Facility at the National Facility for Protein Science in Shanghai (NFPS), Zhangjiang Lab, China, and the Institutional Technology Service Center of Shanghai Institute of Materia Medica, Chinese Academy of Sciences for technical supports.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, Y., Zhang, H., Fan, L. et al. MiR-629-5p promotes the invasion of lung adenocarcinoma via increasing both tumor cell invasion and endothelial cell permeability. Oncogene 39, 3473–3488 (2020). https://doi.org/10.1038/s41388-020-1228-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-1228-1
- Springer Nature Limited
This article is cited by
-
The role of tRF-Val-CAC-010 in lung adenocarcinoma: implications for tumorigenesis and metastasis
BMC Cancer (2024)
-
The RNA-binding protein KSRP aggravates malignant progression of clear cell renal cell carcinoma through transcriptional inhibition and post-transcriptional destabilization of the NEDD4L ubiquitin ligase
Journal of Biomedical Science (2023)
-
A circulating microRNA panel as a novel dynamic monitor for oral squamous cell carcinoma
Scientific Reports (2023)
-
Exosomal HMGA2 protein from EBV-positive NPC cells destroys vascular endothelial barriers and induces endothelial-to-mesenchymal transition to promote metastasis
Cancer Gene Therapy (2022)
-
Noncoding RNAs in tumor metastasis: molecular and clinical perspectives
Cellular and Molecular Life Sciences (2021)