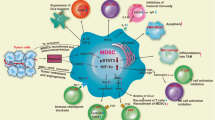
Abstract
Myeloid-derived suppressor cells (MDSCs) suppress antitumor immune activities and facilitate cancer progression. Although the concept of immunosuppressive MDSCs is well established, the mechanism that MDSCs regulate non-small cell lung cancer (NSCLC) progression through the paracrine signals is still lacking. Here, we reported that the infiltration of MDSCs within NSCLC tissues was associated with the progression of cancer status, and was positively correlated with the Patient-derived xenograft model establishment, and poor patient prognosis. Intratumoral MDSCs directly promoted NSCLC metastasis and highly expressed chemokines that promote NSCLC cells invasion, including CCL11. CCL11 was capable of activating the AKT and ERK signaling pathways to promote NSCLC metastasis through the epithelial-mesenchymal transition (EMT) process. Moreover, high expression of CCL11 was associated with a poor prognosis in lung cancer as well as other types of cancer. Our findings underscore that MDSCs produce CCL11 to promote NSCLC metastasis via activation of ERK and AKT signaling and induction of EMT, suggesting that the MDSCs-CCL11-ERK/AKT-EMT axis contains potential targets for NSCLC metastasis treatment.
Similar content being viewed by others
Introduction
Lung cancer remains a daunting health problem with high incidence and mortality rates worldwide [1], more than 80% of cases are non-small cell lung cancer (NSCLC) [2], most cancer-related deaths result from metastatic tumor progression. The origins of the invasive and metastatic cancer cells remain the subjects of intense investigation. Some studies attribute the metastatic phenotypes to cell-autonomous alterations by genomic mutations of cancer cells, whereas alternative views suggest that these traits are acquired through the paracrine signals that cancer cells receive from tumor-associated microenvironment, which has been found to play crucial roles in cancer development and metastasis [3]. The tumor programs cellular infiltrates in the microenvironment to maintain dysregulated inflammation that is favorable to tumor development and progression. Recent reports indicated that the myeloid-derived suppressor cells (MDSCs) are recruited to the environment of progressive tumors [4]. To better understand the role of MDSCs in tumorigenesis, we set out to determine whether MDSCs can provide contextual signals to promote NSCLC metastasis.
MDSCs are broadly defined as a population of immature myeloid cells [5]. In mice, MDSCs can be defined as CD11b+Gr1+ cells with immune suppressive functions. In human, granulocytic MDSCs (G-MDSCs) are predominantly CD15+ and monocytic MDSCs (M-MDSCs) are CD14+, both being HLA-DR-CD33+CD11b+ [6]. MDSCs are associated with a more advanced cancer stage and with reduced survival rate, supporting a role for MDSCs in tumor metastasis [7,8,9,10,11,12]. It has been well documented that MDSCs are able to suppress immune cell functions and contribute to tumor escape from immune surveillance [13,14,15]. Cancer xenograft models in immunodeficient mice indicate that MDSCs have immune-independent effects and may also play important roles in tumor angiogenesis and premetastatic niches [16]. Indeed, as we have demonstrated, mouse stroma cells prepared from developing human A549 or patient-derived lung cancer xenografts are enriched with immature myeloid cells which have the ability to support NSCLC progression [17]. Although the concept of immunosuppressive MDSCs is well established, an understanding of the crosstalk between MDSCs and NSCLC cells is lacking. In this study, our goal was to investigate the mechanism of MDSCs regulating NSCLC metastasis.
Results
Gene expression profiles associated with NSCLC metastasis and PDX model engraftment
To depict the gene expression profiles related to NSCLC metastasis, tumors from NSCLC patients (34 invasive samples and 41 noninvasive samples) were subjected to RNA-Seq analysis. Gene expression patterns were different between invasive and noninvasive samples (Supplementary Fig. 1A). Four distinct groups of genes were identified as myeloid leukocyte activation, myeloid leukocyte mediated immunity, ECM disassembly, and cell migration (Fig. 1A). Noticeably, the expression of S100 proteins (S100A8, S100A9, and S100A12), which were identified as possible markers of MDSCs [12, 18], were found to be enriched in the invasive tumors. The expression of matrix metalloproteinase (MMPs), the essential factors for MDSCs to induce cancer metastasis, were also enriched in the invasive tumors (Supplementary Fig. 1B). We further evaluated MDSCs in resected primary tumors and observed that invasive tumors contained more infiltrated MDSCs (Fig. 1B, C, Supplementary Table 1). These observations indicated that MDSCs are enriched in invasive NSCLC samples.
A Heatmap of differential gene expression analysis of snap-frozen banked patient tumors (34 invasive and 41 noninvasive samples); color pattern represents the row Z-score. B Immunohistochemical staining and analyses of CD33 myeloid cell in the invasive and noninvasive tumor samples. Scale bar: 100 μm. Staining intensity were showed as mean IOD (4 random sights/sample), data are presented as the mean ± SEM. Two-way ANOVA. C Flow cytometry gating strategy for the analysis of human MDSCs; summary of the percentages of human MDSCs (M-MDSCs: HLA-DR-/loCD33+CD11b+CD14+CD15-; G-MDSCs: HLA-DR-/loCD33+CD11b+CD14-CD15+) in the invasive (n = 34) and noninvasive samples (n = 38). D Hematoxylin and eosin (HE) staining and immunohistochemical analyses of cytokeratin 7 (CK7), transcription termination factor 1 (TTF1), and Napsin A expression in LC2 and matched PDX samples. Scale bar: 100 μm. Staining intensity were showed as mean IOD (4 random sights/sample), data are presented as the mean±SEM. Unpaired two-tailed t test. E Heatmap of differential gene expression analysis of snap-frozen banked patient tumors (33 engrafted and 42 nonengrafted samples); color pattern represents the column Z-score. F summary of the percentages of human MDSCs (M-MDSCs: HLA-DR-/loCD33 + CD11b + CD14 + CD15-; G-MDSCs: HLA-DR-/loCD33 + CD11b + CD14-CD15+) in the engrafted (n = 39) and nonengrafted samples (n = 33), and samples from survival patients (n = 37) or deaths (n = 35). Data are presented as the mean ± SEM. Unpaired two-tailed t test. ns: no significant difference; *p < 0.05, **p < 0.01, and ***p < 0.001.
Meanwhile, we established lung cancer PDX model by resected human NSCLC samples, tumor samples directly resected from patients were designated P0 and implanted into NSI mice (P1). A tumor was considered engrafted if P1 tumor could generate P2 tumor. 75 samples were engrafted successfully (73/131, 55.7%), characteristics of tumors which were established PDX models were summarized in Supplementary Table 2. The established PDX tumors were routinely subjected to histological review. HE staining and immunohistochemical analysis of CK7, TTF1, and Napsin A revealed that the PDX tumors matched the primary tumors (Fig. 1D).
Gene expression patterns were different between engrafted and nonengrafted samples (Supplementary Fig. 1C). Notably, we observed that gene expression patterns in engrafted tumors were similar to invasive tumors, there were four distinct groups of upregulated genes in engrafted tumors (Fig. 1E). In addition, if a set of genes could stimulate the upregulation of keratin proteins, then they might enable tumor cells to accommodate a new epithelial environment (Supplementary Fig. 1D). We further analyzed the characteristics of these tumors (Table 1), and found that the success rates of PDX establishments were positively correlated with lymphatic invasion. The tumor samples from patients with lymph node metastases (42/56, 75%) were more likely to grow after implantation, and tumors dissected from the patients with progressive disease were more likely to successfully engraft (Stage I: 17/44, 38.6%; Stage II: 36/53, 67.9%; Stage III-IV: 20/34, 58.8%). In addition, cell differentiation state and tumor diameter were also significantly correlated with PDX establishments. In accord with observations, MDSCs were enriched in engrafted tumors and progressive tumors (Fig. 1F). Taken together, these observations indicated that the enrichment of MDSCs within tumors might favor NSCLC metastasis and PDX model engraftment, which may be associated with poor prognosis in NSCLC.
MDSCs enhance NSCLC metastasis
To investigate the role of MDSCs in promoting NSCLC metastasis, we established a xenograft model in which A549 human NSCLC cells were mixed with MDSCs derived from resected primary human NSCLC samples. More metastatic tumors were observed in the kidneys, livers, and lungs in A549-MDSCs bearing mice compared to A549-control mice (Fig. 2A, B). Also, counts of the metastatic nodules in these organs were folded enhancements in A549-MDSCs bearing mice compared to A549-control mice (Fig. 2C), and more metastatic dissemination of tumor cells were detected in these organs of A549-MDSCs bearing mice (Fig. 2D). We also established NSCLC PDX models in NSI mice as reported previously (LC18, LC85, LC88) [19]. Circulating tumor cells (CTCs) were detected in the peripheral blood (Supplementary Fig. 2A–C), and metastatic dissemination of tumor cells were detected in organs of the PDX models (Supplementary Fig. 2D). Furthermore, we found the percentages of CTCs in the mice model were positively correlated with MDSCs infiltration in primary tumors (Supplementary Fig. 2E). Since NSI mice exhibit impaired development of T cells, B cells, and natural killer cells [20], our data indicated that MDSCs or derivatives thereof might directly enhance NSCLC metastasis.
A Enhancement of cancer cells metastasis in mice that implanted A549 cells mixed with MDSCs (n = 5). Black arrowheads indicated the metastatic tumors. B Hematoxylin-eosin staining of the kidneys, livers and lungs of A549 tumor-bearing mice; scale bar, 1000 μm. Black arrowheads indicated the metastatic area. C The cancer metastasis index for each mouse was calculated as the ratio of the number of colonies observed in the organs divided by the colon numbers of their A549-control littermates (n = 5). Data are presented as the mean ± SD. Unpaired two-tailed t test. D Summary of the flow cytometry analysis of metastatic tumor cells (GFP+) in A549 xenograft mice (n = 5). Data are presented as the mean ± SD. Unpaired two-tailed t test; *p < 0.05, **p < 0.01, and ***p < 0.001.
Tumoral MDSCs having high expression of CCL11 could induce NSCLC cell invasion
To further investigate how tumor-associated MDSCs promote tumor metastasis, we purified MDSCs from PDX mice and their homologous cells from tumor-free mice and performed transcriptomic profiling of these populations using RNA-Seq (Supplementary Fig. 3A). A comparison of the fold changes in mRNA expression levels among cell populations revealed significant changes in gene expression levels that distinguished the tumoral MDSCs from the other populations (Supplementary Fig. 3B, Supplementary Table 3). Gene ontology (GO) analysis of the upregulated genes in tumoral MDSCs revealed that the enriched genes were associated with cell migration (Supplementary Fig. 3C, Supplementary Table 4). Previous studies in murine tumor models have reported that MDSCs induce cancer cell invasion and metastasis by IL-6 induced signals and S100A8/9 induced signals [21, 22]. We also found that IL-6 was upregulated in the tumoral MDSCs, however, the expression of other genes related to IL-6 signaling were not significantly upregulated in the invasive tumors (Supplementary Fig. 3D, E). Similarly, although we found high expression of S100A8/9 in the invasive tumors, S100A8/9 were not specifically expressed in the tumoral MDSCs; in addition, the expression of other genes related to S100A8/9 signaling were also not significantly upregulated in the invasive tumors (Supplementary Fig. 3D, E). Therefore, IL-6 signaling or S100A8/9 signaling might not responsible for MDSCs-induced metastasis in human NSCLC cells.
We further investigated genes related to cell migration and found that the upregulated genes in tumoral MDSCs included many chemokines (Supplementary Fig. 3F), that regulate the initiation or maintenance of tumor metastasis [23,24,25]. In line with these observations, we found that chemokines were enriched in the invasive tumors (Fig. 3A). These results indicated that tumoral MDSCs might promote NSCLC metastasis through chemokines. We found several chemokines (CXCL11, CCL11, CCL24, CCL7, and CCL4) that had not yet been reported or had not been well studied in the crosstalk between MDSCs and NSCLC cells during the process of metastasis. Therefore, we chose these chemokines for further functional studies in the context of tumor metastasis.
A Heatmap of chemokines expression analysis of the invasive and noninvasive tumors (34 invasive and 41 noninvasive samples); color pattern represents the column Z-score. B Experimental scheme for transwell assays. Tumor cells were placed in the upper chambers with an extracellular matrix (ECM), and 293 T cells were placed in the lower chambers. C Representative images of transwell assay results; scale bar, 100 μm. And quantitative analysis of invasive cells, the histogram indicates the invasive ability of tumor cells in the transwell cell invasion assays (n = 3). Data are presented as the mean ± SEM. One-way ANOVA. D Immunohistochemical staining for CCL4, CCL7, CCL11, CCL24, and CXCL11 in resected human NSCLC samples (Invasive samples: LC18, LC85, and LC88; and Noninvasive samples: LC15, LC43, and LC55); scale bar, 100 μm. Staining intensity were showed as mean IOD (4 random sights/sample), data are presented as the mean ± SEM. Two-way ANOVA; ns: no significant difference, *p < 0.05, **p < 0.01, and ***p < 0.001. E Kaplan–Meier survival curves of lung cancer patients (overall survival, 1925 patients).
First of all, we confirmed the expression of their receptors in NSCLC cells (Supplementary Fig. 3G, H). To determine whether these chemokines are responsible for tumor metastasis, we overexpressed the candidate chemokines in 293T cells (Supplementary Fig. 3I) and performed invasion assays with transwell chambers (Fig. 3B). Although all of the tested chemokines exhibited the capacity to enhance tumor invasion, CCL7 and CCL11 markedly increased the invasive capacity of lung cells (Fig. 3C). In addition, expression of these chemokines was examined by immunohistochemical staining in resected NSCLC samples, expression level of CCL11 and CXCL11 were significantly elevated in the invasive tumors (Fig. 3D). Furthermore, we found that increased CCL11 expression was associated with a significantly poor survival rate in lung cancer patients (Fig. 3E). Therefore, we focused our efforts on evaluating the importance of CCL11 in MDSCs-induced metastasis.
CCL11 promotes NSCLC metastasis
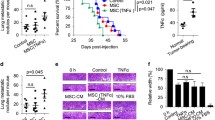
Several observations have linked CCL11 signaling in association with lung cancer [26, 27]. However, the contributions of CCL11 to NSCLC development, especially the crosstalk between MDSCs and NSCLC cells are poorly understood. To investigate further the possible causal role of CCL11 in NSCLC cell metastasis, we first performed an evaluated transwell assay, which showed that CCL11 promoted NSCLC invasion (Supplementary Fig. 4A). Then we established PDX mouse models and intraperitoneally treated them with CCL11 once per week (Fig. 4A). The percentage of CTCs significantly increased after 3 weeks of CCL11 treatment in the PDX models (Fig. 4B, Supplementary Fig. 4B). In addition, flow cytometric analysis showed that increased numbers of metastatic tumor cells were found in the spleen, liver, and lungs in two PDX models (Supplementary Fig. 4B). Histopathological analyses indicated that treatment with CCL11 could robustly increase tumor metastasis in both PDX models (Fig. 4C). Collectively, these observations point out the importance of CCL11 in NSCLC metastasis.
A Diagram of the assessment of CCL11 in PDX mouse models. Tumors were subcutaneously implanted, and CCL11 was intraperitoneally injected into the mice once per week. CTCs were detectable 3 weeks after tumor implantation, the mice were sacrificed after 4 weeks after CTCs were detectable. B FACS analysis of tumor cell frequencies in the peripheral blood, spleen, liver, and lungs of LC-PDX1 and LC-PDX2 mice (n = 3/group). Data are presented as the mean ± SD. Unpaired two-tailed t test. C Histopathological analysis of tumor metastasis in PDX models, immunohistochemical staining for hHLA in the spleen, liver, and lungs of two LC-PDX model mice; scale bar, 50 μm. Staining intensity were showed as mean IOD of hHLA in each group (4 random sights/sample). Data are presented as the mean ± SEM. Unpaired two-tailed t test. D Hematoxylin-eosin staining of the kidneys, livers, and lungs of A549 tumor-bearing mice; scale bar, 1000 μm. Black arrowheads indicated the metastatic area. And summary of the flow cytometry analysis of metastatic tumor cells in A549 xenograft mice (n = 5). Data are presented as the mean ± SD. One-way ANOVA; *p < 0.05, **p < 0.01, and ***p < 0.001.
To further explore the causal role of CCL11 in MDSCs-induced metastasis, we established a xenograft model in which A549 human NSCLC cells were mixed with MDSCs. We found that CCL11 treatment or the presence of MDSCs could robustly enhance A549 cancer cells dissemination. Neutralization of CCL11 protein using intraperitoneal injections of an anti-human CCL11 monoclonal antibody abrogated the MDSCs-induced metastasis in A549 cells (Fig. 4D). We noted that this enhanced metastatic ability was not due to CCL11’s effects on cellular proliferation (Supplementary Fig. 4C) or the contamination of tumor cells with MDSCs (Fig. 4D). These results indicated that the actions of CCL11 were responsible for much, if not all, of the observed MDSCs-induced metastasis in human NSCLC cells.
Essential role of the CCL11-CCR3 interaction in NSCLC metastasis
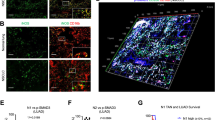
CCR3 has been determined to be the main receptor for CCL11 in cancer cells [28], therefore, we focused our efforts on evaluating the importance of the CCL11-CCR3 interaction in NSCLC metastasis. We inhibited CCR3 expression in lung cancer cells through shRNA knockdown (Supplementary Fig. 5A). Then we performed gene expression analysis of CCR3 knockdown in A549 cells as well as A549 cells treated with CCL11 using RNA-Seq. The complete list of expressed genes is shown in Supplementary Table 5. Differentially expressed genes distinguish differently treated cells (Supplementary Fig. 5B). KEGG pathway analysis of the differentially expressed genes identified 77 pathways for genes with upregulated expression after CCL11 treatment, and 71 pathways for genes with downregulated expression after CCR3 knockdown (Supplementary Table 6). PI3K-AKT signaling pathway and Ras-MAPK-ERK signaling pathway were identified as significantly important pathways in CCL11-treated cells (Fig. 5A, C) and CCR3-knockdown cells (Fig. 5B, C), respectively. In addition, we confirmed the increased phosphorylation of ERK1/2 and AKT in invasive tumors (Fig. 5D). ERK and AKT served as the key switch of upstream signals that promote cellular motility [29, 30]. Taken together, these results indicated the critical importance of ERK and AKT signaling in CCL11-induced metastasis of NSCLC cells.
Bubble chart showing the results of KEGG pathway enrichment analysis. A The top 20 enriched pathways for upregulated genes in CCL11-treated cells versus untreated cells. B The top 20 enriched pathways for downregulated genes in CCR3-knockdown cells versus untreated cells. Bubble size indicates the number of genes associated with each term, and bubble color indicates p value significance. C Heatmap analysis of differentially regulated genes in A549 tumor cells (untreated cells; CCL11-treated cells; CCR3-knockdown cells, n = 3/group). The color pattern represents the column Z-score. D Immunohistochemical staining for p-ERK and p-AKT in resected human NSCLC samples (Invasive samples: LC84, LC86, and LC87; and Noninvasive samples: LC15, LC69, and LC78); scale bar, 100 μm. Staining intensity were showed as mean IOD (4 random sights/sample, 6 invasive samples: LC84, LC86, LC87, LC18, LC85, and LC88; 6 noninvasive samples: LC15, LC69, LC78, LC43, LC55, and LC26), data are presented as the mean ± SEM. Two-way ANOVA; *p < 0.05, **p < 0.01, and ***p < 0.001.
Invasion and metastasis of cancer cells are facilitated through the process termed epithelial-to-mesenchymal transition (EMT), in which cancer cells convert stationary epithelial traits to acquire mesenchymal markers that endow their motility [31]. This transition has also been identified as the direct downstream target of the AKT and ERK signaling pathways [32, 33]. CCL11 treatment or CCR3 knockdown led to a change in the expression of canonical EMT markers, including loss of the epithelial marker keratins, mucins, and collagens, and induction of the mesenchymal marker N-cadherin, vimentin, fibronectin, and MMPs (Fig. 5C), which implied that CCL11 induced NSCLC cell EMT.
To further confirm the role of ERK and AKT in mediating the actions of CCL11 on cellular motility, we treated A549 cells with CCL11, ERK1/2 phosphorylation inhibitor (GDC0994) as well as AKT phosphorylation inhibitor (MK2206). We found that CCL11 induced the phosphorylation of ERK1/2 and AKT and the phosphorylation was decreased by knocking down of CCR3 (Fig. 6A). CCL11 increased Vimentin and N-cadherin expression levels, and their expression was decreased after treating with GDC0994, MK2206, or CCR3 knockdown. In contrast, the expression of epithelial markers (E-cadherin and Fibronectin 1) was suppressed in CCL11-treated tumor cells and increased after treating with GDC0994, MK2206, or CCR3 knockdown (Fig. 6A, B). The expression level of key transcription factors that mediate EMT and MMPs was upregulated in CCL11-treated tumor cells, but downregulated in GDC0994-treated, MK2206-treated and CCR3-knockdown cells (Fig. 6B). Furthermore, we found that CCR3-knockdown cells changed from an elongated shape to a rounded shape (Supplementary Fig. 5C), which indicated declined mobility. In addition, A549 cells exhibited enhanced invasiveness when treated with CCL11, whereas CCR3 knockdown or ERK/AKT signaling blockade impeded the CCL11-induced NSCLC metastasis (Fig. 6C, D). Similar phenomena were detected in H1299 and H460 cancer cells (Supplementary Figs. 6, 7). Collectively, these observations suggested that CCL11-CCR3 interaction activated AKT and ERK signaling pathways to promote NSCLC metastasis through the EMT process.
A Western blot analysis of the phosphorylation of ERK1/2 and AKT and expression of E-cadherin and vimentin in A549 cells, CCR3-knockdown cells treated with or without CCL11, GDC0994, or MK2206. Data are presented as the mean ± SEM (n = 3). One-way ANOVA. B qRT-PCR analysis of the mRNA expression of the indicated genes. The results were normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) mRNA levels and are presented as the mean ± SEM (n = 3). One-way ANOVA. C Representative images of transwell assay results; scale bar, 100 μm. And quantitative analysis of invasive cells. Data are presented as the mean ± SEM (n = 3). One-way ANOVA. D Hematoxylin-eosin staining of the kidneys, livers and lungs of A549 tumor-bearing mice; scale bar, 1000 μm. Black arrowheads indicated the metastatic area. And summary of the flow cytometry analysis of metastatic tumor cells in A549 xenograft mice (n = 5). Data are presented as the mean ± SD. One-way ANOVA; *p < 0.05, **p < 0.01 and ***p < 0.001.
Discussion
Certain models of metastatic progression indicate that cancer cell invasion and metastasis are strongly influenced by paracrine signals emanating from MDSCs. A syngeneic mouse model provided evidence that tumoral-mMDSCs facilitate tumor cell metastasis by inducing EMT phenotype, whereas pulmonary gMDSCs revert EMT phenotype to support the metastatic growth [34], which suggested that the traits of tumor cells to metastasize is a consequence of their education by MDSCs. Although murine models have been proposed to demonstrate that MDSCs induce cancer cell invasion and metastasis by some paracrine signals, such as IL-6 induced signals in breast cancer and S100A8/9 induced signals in colon cancer [21, 22], direct evidence of MDSCs regulating human NSCLC progression through the paracrine signals is still lacking. Our study extended the knowledge that MDSCs might serve to influence NSCLC metastasis. We noted that CCL11 which was associated with poor prognosis of NSCLC, was highly expressed in tumoral-MDSCs and enriched in invasive lung carcinomas. These observations argued strongly for a significant association between MDSCs, CCL11, levels and human invasive NSCLC.
CCL11-CCR3 interaction is involved in the recruitment of eosinophils, basophils, and T helper 2 lymphocytes which contribute to tumor angiogenesis [35] and Th2-dominant tumor environment [36]. We pointed out that CCL11 participated in MDSCs-induced NSCLC metastasis by interaction with CCR3, also CCL11 could activate ERK/AKT signaling to induce EMT phenotype. The CCL11-CCR3-ERK pathway has been shown to promote survival in lymphoma cells [37] and facilitate invasion in prostate cells [38], but no evidence has been shown that CCL11 is linked with EMT during NSCLC progression. These findings may extend our knowledge of the links between MDSCs and CCL11 and shed light on the mechanisms underlying how MDSCs facilitate human NSCLC metastasis by paracrine signals. Apart from human lung cancers, there are also a series of data which indicate the impact of CCL11 on cancer progression. Elevated CCL11 expression level in cancer patients is associated with progressive disease and could be used as a biomarker for poor prognosis (Supplementary Fig. 8) [39,40,41,42], which indicates the essential role of CCL11 signals in cancer metastasis.
PDX model has become a favored preclinical model to study tumor biology due to its high fidelity in reconstituting human tumors [43]. However, engraftment failure is still a bottleneck in PDX models. It has been reported that the increased success rate of PDX was associated with enhanced clinical aggressiveness in tumors [44, 45]. In our study, we also noticed that successful PDX establishment was associated with progressive tumor stage and poor prognosis, suggesting that progressive tumors are more likely to establish lung cancer PDX models. What’s more, stromal cells originally present in dissected tumors from patients were gradually replaced by murine stromal cells as the xenograft grows, which could obstruct the analysis of human tumor-stroma interactions. Our findings indicated that MDSCs could be a positive factor in progressive NSCLC for the establishment of a PDX model. Thus, transplanted pro-tumoral MDSCs or para-cancerous tissue might be beneficial for establishing lung cancer PDX models as well as the study of tumor biology. Furthermore, these findings suggest that targeting MDSCs as part of an approach to inhibit the metastatic process appears to be a very clinically promising strategy for NSCLC therapy.
In summary, our study provided evidence for the tumor-promoting function of MDSCs in NSCLC, and this function might represent a promotive factor for progressive NSCLC and favored the establishment of a PDX model. MDSCs produced CCL11 to promote NSCLC progression via the activation of ERK and AKT pathways as well as induction of EMT in NSCLC cells. These findings shed light on the mechanisms by which MDSCs could contribute to NSCLC metastasis in the primary tumor microenvironment. The MDSC-CCL11-ERK/AKT-EMT axis was essential for NSCLC metastasis and could be a potential target for metastatic NSCLC therapies.
Material and methods
Human tissues
This study was approved by the ethics committee of The Third Affiliated Hospital of Guangzhou Medical University. All human lung cancer tissues used to create PDX mouse models were obtained from Nanfang Hospital, Southern Medical University, and Guangdong General Hospital. The use of human tissue samples in this study was approved by the Committee for the Ethical Review of Research Involving Human Subjects at Nanfang Hospital, Southern Medical University, and Guangdong General Hospital. Written, informed consent was obtained for all involved patients.
Mice
Animal experiments were performed in the Laboratory Animal Center of the Guangzhou Institutes of Biomedicine and Health (GIBH), and all animal procedures were approved by the Animal Welfare Committee of GIBH. NOD-SCID-IL2Rg−/− (NSI) mice were derived at the Laboratory Animal Center of GIBH. Mice were maintained in specific pathogen-free cages and provided with autoclaved food and water. Adult male mice aged 6–8 weeks were used in this study. Protocols were approved by the relevant Institutional Animal Care and Use Committee.
Tumor models
For PDX models, the resected primary human NSCLC samples were implanted subcutaneously into the right and left flank of female NSI mice, the established PDX models (tumor volume >100 mm3) were selected for further analysis. The tumor-bearing mice were sacrificed when the tumors reached 1000 mm3 (as measured by calipers) to analyze MDSCs and CTCs. When evaluated CCL11 in PDX model, human cancer tissue blocks were subcutaneously transplanted into NSI mice treated with an intraperitoneal injection of PBS, or 40 μg/kg recombinant human CCL11 (ProSpec) once per week. CTCs were detectable 3 weeks after tumor implantation, the mice were sacrificed after 4 weeks after CTCs were detectable.
For A549 xenograft experiments, cancer cells were implanted alone (5 × 105 cells), or mixed with MDSCs derived from resected primary human NSCLC samples (1 × 105 cells) and injected intravenously into female NSI mice, the mice were treated with 40 μg/kg recombinant human CCL11 or 50 μg/kg anti-human CCL11 neutralizing Ab (R&D Systems), or equivalent amount of purified rat immunoglobulin G intraperitoneally, 50 mg/kg GDC0994 were given orally daily, 120 mg/kg MK2206 were oral given alternate days, 4 weeks after tumor implantation, the mice were sacrificed for analysis.
Cell culture
The human non-small cell lung carcinoma A549, H460, and H1299 cells were cultured in RPMI-1640 medium (Gibco) supplemented with 10% fetal bovine serum (FBS; Biochrom) and passaged at 80% confluence. Cells were maintained at 37 °C in a humidified-air atmosphere containing 5% CO2. The A549, H460, and H1299 cells used in this study were obtained from ATCC with identification by STR profiling and kept at a low passage number for experimental use. We routinely screened for mycoplasma infection with the Mycoplasma Detection Kit (Lonza) and discarded mycoplasma-positive cells. For in vitro experiment, tumor cells were cultured with RMPI-1640 supplemented with 10% FBS with or without 10 ng/mL human CCL11, 1 μM GDC0994, 1 μM MK2206.
For CCL11 culture experiments, 1000 A549 cells were cultured in a 96-well plate with RMPI-1640 supplemented with 10% FBS with or without 10 ng/mL human CCL11. Tumor cell numbers were measured by Cell Counting Kit-8 (CCK-8, DOJINDO, Japan) at the indicated times according to the manufacturer’s instructions.
Gene expression analysis
Tumor tissue or tumor cells were collected for RNA isolation. RNA preparation, library construction, and sequencing on a BGISEQ-500 platform were performed at the Beijing Genomics Institute (BGI). RNA was isolated from cell pellets using TRIzol™ Reagent (Invitrogen), and RNA quality was determined using the Bioanalyzer 2100 (Agilent). 23 S and 16 S rRNAs were depleted using a MicrobExpress kit (Ambion). Genomic DNA was removed with two digestions using amplification-grade DNase 1 (Invitrogen). The processed RNA was sheared and reverse transcribed using random primers to obtain cDNA, which was used for library construction. Library quality was determined by using the Bioanalyzer 2100 (Agilent). Then, the library was used for sequencing with the sequencing platform BGISEQ-500 (BGI).
The generated raw sequencing reads were filtered to remove reads with adapters, reads in which unknown bases accounted for more than 5% of bases, and low-quality reads. Clean reads were then obtained and stored in FASTQ format. HISAT [46] was used to map the clean reads to a reference genome. Gene expression levels were quantified by the software package RSEM [47]. The DEGseq method was used to screen differentially expressed genes between groups (cutoff criteria: fold change absolute value of log2-Ratio ≥1 and a false discovery rate ≤0.001). GO and pathway annotation and enrichment analyses were based on the Database for Annotation, Visualization, and Integrated Discovery (https://david.ncifcrf.gov/).
Data mining
The associations between chemokine expression in tissue and clinical features and outcomes of cancer patients were obtained by Kaplan–Meier plotter (http://www.kmplot.com) as previously described [48, 49]. Additional details of the study are available at Kaplan–Meier plotter.
Statistics
Sample sizes were estimated based on prior related studies [17, 19], samples and animals were random allocated and were unbiasedly included into the analysis unless specific mention. All cell culture experiments, transwell assay, western blot, real-time PCR and ELISA were performed in triplicate at least three independent times. Statistical analysis to determine group differences is determined by unpaired two-tailed t test comparing two groups, one-way or two ways ANOVA multiple tests comparing three or more groups using GraphPad Prism (GraphPad Software), overall survival of cancer patients was assessed using Cox proportional hazards and log-rank analyses from the database, all statistical analyses are described in the figure legends. The p values are considered as follows: *p < 0.05, **p < 0.01, and ***p < 0.001.
Data availability
RNA-Seq of human NSCLC samples, MDSCs and A549 cells have been uploaded to public databases (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE136949). Other data generated or analyzed during this study are included either in this article or in the supplementary information files.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Miller KD, Nogueira L, Mariotto AB, Rowland JH, Yabroff KR, Alfano CM, et al. Cancer treatment and survivorship statistics, 2019. CA Cancer J Clin. 2019;69:363–85.
Lambert AW, Pattabiraman DR, Weinberg RA. Emerging biological principles of metastasis. Cell. 2017;168:670–91.
Yang Z, Guo J, Weng L, Tang W, Jin S, Ma W. Myeloid-derived suppressor cells-new and exciting players in lung cancer. J Hematol Oncol. 2020;13:10.
Peranzoni E, Zilio S, Marigo I, Dolcetti L, Zanovello P, Mandruzzato S, et al. Myeloid-derived suppressor cell heterogeneity and subset definition. Curr Opin Immunol. 2010;22:238–44.
Marvel D, Gabrilovich DI. Myeloid-derived suppressor cells in the tumor microenvironment: expect the unexpected. J Clin Investig. 2015;125:3356–64.
Zhang G, Huang H, Zhu Y, Yu G, Gao X, Xu Y, et al. A novel subset of B7-H3(+)CD14(+)HLA-DR(-/low) myeloid-derived suppressor cells are associated with progression of human NSCLC. Oncoimmunology. 2015;4:e977164.
Meyer C, Cagnon L, Costa-Nunes CM, Baumgaertner P, Montandon N, Leyvraz L, et al. Frequencies of circulating MDSC correlate with clinical outcome of melanoma patients treated with ipilimumab. Cancer Immunol Immunother. 2014;63:247–57.
Huang A, Zhang B, Wang B, Zhang F, Fan KX, Guo YJ. Increased CD14(+)HLA-DR (-/low) myeloid-derived suppressor cells correlate with extrathoracic metastasis and poor response to chemotherapy in non-small cell lung cancer patients. Cancer Immunol Immunother. 2013;62:1439–51.
Diaz-Montero CM, Salem ML, Nishimura MI, Garrett-Mayer E, Cole DJ, Montero AJ. Increased circulating myeloid-derived suppressor cells correlate with clinical cancer stage, metastatic tumor burden, and doxorubicin-cyclophosphamide chemotherapy. Cancer Immunol Immunother. 2009;58:49–59.
Sun HL, Zhou X, Xue YF, Wang K, Shen YF, Mao JJ, et al. Increased frequency and clinical significance of myeloid-derived suppressor cells in human colorectal carcinoma. World J Gastroenterol. 2012;18:3303–9.
Wang L, Chang EW, Wong SC, Ong SM, Chong DQ, Ling KL. Increased myeloid-derived suppressor cells in gastric cancer correlate with cancer stage and plasma S100A8/A9 proinflammatory proteins. J Immunol. 2013;190:794–804.
Lindau D, Gielen P, Kroesen M, Wesseling P, Adema GJ. The immunosuppressive tumour network: myeloid-derived suppressor cells, regulatory T cells and natural killer T cells. Immunology. 2013;138:105–15.
Mao Y, Poschke I, Wennerberg E, Pico de Coana Y, Egyhazi Brage S, Schultz I, et al. Melanoma-educated CD14+ cells acquire a myeloid-derived suppressor cell phenotype through COX-2-dependent mechanisms. Cancer Res. 2013;73:3877–87.
Rodriguez PC, Quiceno DG, Zabaleta J, Ortiz B, Zea AH, Piazuelo MB, et al. Arginase I production in the tumor microenvironment by mature myeloid cells inhibits T-cell receptor expression and antigen-specific T-cell responses. Cancer Res. 2004;64:5839–49.
Joyce JA, Pollard JW. Microenvironmental regulation of metastasis. Nat Rev Cancer. 2009;9:239–52.
Lin S, Huang G, Xiao Y, Sun W, Jiang Y, Deng Q, et al. CD215+ Myeloid Cells Respond to Interleukin 15 Stimulation and Promote Tumor Progression. Front Immunol. 2017;8:1713.
Zhao F, Hoechst B, Duffy A, Gamrekelashvili J, Fioravanti S, Manns MP, et al. S100A9 a new marker for monocytic human myeloid-derived suppressor cells. Immunology. 2012;136:176–83.
Lin S, Huang G, Cheng L, Li Z, Xiao Y, Deng Q, et al. Establishment of peripheral blood mononuclear cell-derived humanized lung cancer mouse models for studying efficacy of PD-L1/PD-1 targeted immunotherapy. MAbs. 2018;10:1301–11.
Ye W, Jiang Z, Li GX, Xiao Y, Lin S, Lai Y, et al. Quantitative evaluation of the immunodeficiency of a mouse strain by tumor engraftments. J Hematol Oncol. 2015;8:59.
Oh K, Lee OY, Shon SY, Nam O, Ryu PM, Seo MW. et al. A mutual activation loop between breast cancer cells and myeloid-derived suppressor cells facilitates spontaneous metastasis through IL-6 trans-signaling in a murine model. Breast Cancer Res. 2013;15:R79. https://doi.org/10.1186/bcr3473.
Ichikawa M, Williams R, Wang L, Vogl T, Srikrishna G. S100A8/A9 activate key genes and pathways in colon tumor progression. Mol Cancer Res. 2011;9:133–48.
Levina V, Nolen BM, Marrangoni AM, Cheng P, Marks JR, Szczepanski MJ, et al. Role of eotaxin-1 signaling in ovarian cancer. Clin Cancer Res. 2009;15:2647–56.
Balkwill F. Cancer and the chemokine network. Nat Rev Cancer. 2004;4:540–50.
Wang D, Sun H, Wei J, Cen B, DuBois RN. CXCL1 Is Critical for Premetastatic Niche Formation and Metastasis in Colorectal Cancer. Cancer Res. 2017;77:3655–65.
Tian W, Liu J, Pei B, Wang X, Guo Y, Yuan L. Identification of miRNAs and differentially expressed genes in early phase non-small cell lung cancer. Oncol Rep. 2016;35:2171–6.
Liu H, Zhao H. Prognosis related miRNAs, DNA methylation, and epigenetic interactions in lung adenocarcinoma. Neoplasma. 2019;66:487–93.
Zajkowska M, Mroczko B. Eotaxins and their receptor in colorectal cancer-a literature review. Cancers. 2020;12:1383. https://doi.org/10.3390/cancers12061383.
Alzahrani AS. PI3K/Akt/mTOR inhibitors in cancer: at the bench and bedside. Semin Cancer Biol. 2019;59:125–32.
Mendoza MC, Er EE, Blenis J. The Ras-ERK and PI3K-mTOR pathways: cross-talk and compensation. Trends Biochem Sci. 2011;36:320–8.
Tsai JH, Donaher JL, Murphy DA, Chau S, Yang J. Spatiotemporal regulation of epithelial-mesenchymal transition is essential for squamous cell carcinoma metastasis. Cancer cell. 2012;22:725–36.
Xu W, Yang Z, Lu N. A new role for the PI3K/Akt signaling pathway in the epithelial-mesenchymal transition. Cell Adh Migr. 2015;9:317–24.
Watanabe-Takano H, Takano K, Hatano M, Tokuhisa T, Endo T. DA-Raf-mediated suppression of the Ras-ERK pathway is essential for TGF-beta 1-induced epithelial-mesenchymal transition in alveolar epithelial type 2 cells. PLoS ONE. 2015;10:e0127888. https://doi.org/10.1371/journal.pone.0127888.
Ouzounova M, Lee E, Piranlioglu R, El Andaloussi A, Kolhe R, Demirci MF, et al. Monocytic and granulocytic myeloid derived suppressor cells differentially regulate spatiotemporal tumour plasticity during metastatic cascade. Nat Commun. 2017;8:14979.
Xu W, Qian J, Zeng F, Li S, Guo W, Chen L, et al. Protein kinase Ds promote tumor angiogenesis through mast cell recruitment and expression of angiogenic factors in prostate cancer microenvironment. J Exp Clin Cancer Res. 2019;38:114.
Miyagaki T, Sugaya M, Fujita H, Ohmatsu H, Kakinuma T, Kadono T, et al. Eotaxins and CCR3 interaction regulates the Th2 environment of cutaneous T-cell lymphoma. J Invest Dermatol. 2010;130:2304–11.
Miyagaki T, Sugaya M, Murakami T, Asano Y, Tada Y, Kadono T, et al. CCL11-CCR3 interactions promote survival of anaplastic large cell lymphoma cells via ERK1/2 activation. Cancer Res. 2011;71:2056–65.
Zhu F, Liu P, Li J, Zhang Y. Eotaxin-1 promotes prostate cancer cell invasion via activation of the CCR3-ERK pathway and upregulation of MMP-3 expression. Oncol Rep. 2014;31:2049–54.
Tian M, Chen LN, Ma L, Wang DD, Shao B, Wu JY, et al. Expression and prognostic significance of CCL11/CCR3 in glioblastoma. Oncotarget. 2016;7:32617–27.
Blank S, Nienhuser H, Dreikhausen L, Sisic L, Heger U, Ott K, et al. Inflammatory cytokines are associated with response and prognosis in patients with esophageal cancer. Oncotarget. 2017;8:47518–32.
Zohny SF, Fayed ST. Clinical utility of circulating matrix metalloproteinase-7 (MMP-7), CC chemokine ligand 18 (CCL18) and CC chemokine ligand 11 (CCL11) as markers for diagnosis of epithelial ovarian cancer. Med Oncol. 2010;27:1246–53.
Agarwal M, He C, Siddiqui J, Wei JT, Macoska JA. CCL11 (eotaxin-1): a new diagnostic serum marker for prostate cancer. Prostate. 2013;73:573–81.
Lai Y, Wei X, Lin S, Qin L, Cheng L, Li P. Current status and perspectives of patient-derived xenograft models in cancer research. J Hematol Oncol. 2017;10:106.
DeRose YS, Wang G, Lin YC, Bernard PS, Buys SS, Ebbert MT, et al. Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat Med. 2011;17:1514–20.
John T, Kohler D, Pintilie M, Yanagawa N, Pham NA, Li M, et al. The ability to form primary tumor xenografts is predictive of increased risk of disease recurrence in early-stage non-small cell lung cancer. Clin Cancer Res. 2011;17:134–41.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 2011;12:323.
Gyorffy B, Surowiak P, Budczies J, Lanczky A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PloS One. 2013;8:e82241.
Nagy A, Lanczky A, Menyhart O, Gyorffy B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep. 2018;8:9227.
Acknowledgements
We thank the cancer patients who donated their tissues. This work was supported by Strategic Priority Research Program of the Chinese Academy of Sciences (XDB19030205); and National Key Research and Development Plan (2017YFE0131600, 2019YFA0111500); and National Natural Science Foundation of China (81961128003; 81972672; 31872800; 81773301; 82003054); and China Postdoctoral Science Foundation (2018M640771); and Guangdong Provincial Significant New Drugs Development (2019B020202003); and Guangdong Basic and Applied Basic Research Foundation (2019A1515110084, 2019A1515010062, 2020A1515011516); and Guangdong Special Support Program (2017TX04R102); and Science and Technology Planning Project of Guangdong Province (2017B030314056); and Natural Science Foundation of Guangdong Province (2020A0505100062); and Guangdong Provincial Key Lab of Translational Medicine in Lung Cancer (2017B030314120); and Guangzhou City Science and Technology Key Topics Project (201904020025); and Guangzhou Science and Technology Plan Project (201907010042, 201904010473) and Foundation of Guangzhou Science and Information Technology of Guangzhou Key Project (201803040009); and Guangzhou Regenerative Medicine and Health Guangdong Laboratory Frontier Research Program (2018GZR110105003); and Clinical Innovation Research Program of Guangzhou Regenerative Medicine and Health Guangdong Laboratory (2018GZR0201002); and Research Program of the Hefei Institute of Stem Cell and Regenerative Medicine (2019YF001); and Science and Technology Program of Guangzhou (202002020083).
Author information
Authors and Affiliations
Contributions
SL and LC, contributed to the conception and design; the collection and/or assembly of data; data analysis and interpretation; and manuscript writing. XZ, GH, JL, and DZ contributed to the provision of study material or patient samples and the collection and/or assembly of data. BL, SL, SW, YL, and QW provided animal care and administrative support. WW, PL, DP, YY, SZC, ZW, and SC contributed to the conception and design of the study. XS, YY, YL, and PL contributed to the conception and design of the study; data analysis and interpretation; manuscript writing; and the final approval of the manuscript and provided financial support. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lin, S., Zhang, X., Huang, G. et al. Myeloid-derived suppressor cells promote lung cancer metastasis by CCL11 to activate ERK and AKT signaling and induce epithelial-mesenchymal transition in tumor cells. Oncogene 40, 1476–1489 (2021). https://doi.org/10.1038/s41388-020-01605-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01605-4
- Springer Nature Limited
This article is cited by
-
The role of stromal cells in epithelial–mesenchymal plasticity and its therapeutic potential
Discover Oncology (2024)
-
Exploring CCL11 in breast cancer: unraveling its anticancer potential and immune modulatory effects involving the Akt-S6 signaling
Journal of Cancer Research and Clinical Oncology (2024)
-
SFRP1 decreases WNT-Mediated M2 macrophage marker expression in breast tissue
Cancer Immunology, Immunotherapy (2024)
-
Cancer-associated fibroblast-derived PAI-1 promotes lymphatic metastasis via the induction of EndoMT in lymphatic endothelial cells
Journal of Experimental & Clinical Cancer Research (2023)
-
MacroH2A restricts inflammatory gene expression in melanoma cancer-associated fibroblasts by coordinating chromatin looping
Nature Cell Biology (2023)