Abstract
Despite major progress in breast cancer research, the functional contribution of distinct cancer cell clones to malignant tumor progression and metastasis remains largely elusive. We have assessed clonal heterogeneity within individual primary tumors and metastases and also during the distinct stages of malignant tumor progression using clonal tracking of cancer cells in the MMTV–PyMT mouse model of metastatic breast cancer. Comparative gene expression analysis of clonal subpopulations reveals a substantial level of heterogeneity across and also within the various stages of breast carcinogenesis. The intra-stage heterogeneity is primarily manifested by differences in cell proliferation, also found within invasive carcinomas of luminal A-, luminal B-, and HER2-enriched human breast cancer. Surprisingly, in the mouse model of clonal tracing of cancer cells, chemotherapy mainly targets the slow-proliferative clonal populations and fails to efficiently repress the fast-proliferative populations. These insights may have considerable impact on therapy selection and response in breast cancer patients.
Similar content being viewed by others
Introduction
In the past years, we have gained increasing knowledge about the extent and the potential impact of tumor heterogeneity on clinical outcome. Tumor heterogeneity is known to be conferred by differences in various intrinsic parameters, e.g., the genetic profile and the transcriptomic landscape of a cell, the interplay between the genome, epigenome, and transcriptome, and variances in the extent of cancer cell plasticity. Besides these cell-autonomous traits, extrinsic factors in the tumor microenvironment affect tumor heterogeneity, including the tumor vasculature, oxygen supply, and tumor cell interactions with stromal and immune cells [1,2,3,4,5,6]. DNA sequencing and gene expression profiling on bulk and single cells of primary tumors and metastatic lesions with and without chemotherapeutic regimens have revealed substantial differences in mutational profiles and large transcriptomic variations between clones of primary tumors and metastases [1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17]. This heterogeneity ultimately results in the formation of subclones with different biological capabilities, which will follow Darwin’s principle of “survival of the fittest”. Nowell [18] has described the idea of a clonal evolution, where subclones with a growth advantage in a certain ecosystem will persist, survive, and eventually expand, whereas less fit subclones will become extinct [19]. Multicolor clonal tracing studies in mouse models of breast, pancreas, and skin squamous cell carcinoma and of sarcoma have identified the emergence of clonal dominance and clonal loss during carcinogenesis and the existence of mono- or polyclonal metastases [10, 11, 20,21,22,23]. However, the clonal diversification or, in other words, the topographical expansion of multiple cancer clones within a patient’s primary tumor and at secondary sites, remains largely elusive. In particular, the gene expression signatures of clonal populations within the distinct stages of oncogenesis and their functional contribution to malignant tumor progression and metastasis formation remain unknown.
Here, we have exploited the MMTV–PyMT mouse line as a model for stage-wise progression of metastatic breast cancer. In particular, we have combined the mammary tumor progression of these mice with Confetti cassette-driven stochastic expression of four different fluorescence proteins [24,25,26], laser capture microscopy, and next-generation RNA sequencing to interrogate clonal variation during breast cancer progression. Comparative analysis of gene expression profiles indicates a substantial level of heterogeneity across and also within the specific stages of neoplastic progression. The intra-stage differences are mainly due to varying capacities in cell proliferation and survival and in oxidative metabolism, differences which have also been observed in clonal subpopulations of luminal A-, luminal B-, and HER2-enriched human breast cancers. However, contrary to the expectation that chemotherapy specifically targets fast-proliferating tumor cells, in the various tumor stages of MMTV–PyMT mice chemotherapy mainly depleted the slow-proliferative and not the fast-proliferative cancer cell subpopulations.
Results
Confetti clonal tracking in murine mammary tumors
To assess clonal expansion and tumor heterogeneity during mammary gland carcinogenesis, the MMTV–PyMT mouse model of metastatic breast cancer was intercrossed with the R26-CBW Confetti reporter mouse line and the K8-Cre-ERT2 driver mouse line [24,25,26] to generate MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice (Fig. 1a). In MMTV–PyMT mice, the mouse mammary tumor virus (MMTV) promoter drives the mammary gland-specific expression of the polyoma virus middle T antigen (PyMT) oncogene, which results into the malignant transformation of the mammary gland epithelium, ultimately leading to a stepwise progression from hyperplasia to adenoma, luminal mammary adenocarcinoma and pulmonary metastasis within 14 weeks of age [24, 25]. The cytokeratin 8 promoter (K8) was used to drive the expression of a tamoxifen-inducible Cre recombinase (K8-Cre-ERT2) in luminal epithelial cells of mammary glands to allow the clonal tracking of tumor cells early on [27, 28]. To validate the K8 promoter as an adequate Cre-driver, MMTV–PyMT primary tumors were first analyzed for K8/K18 expression by immunofluorescence microscopy, demonstrating K8/18 expression in all the different stages of murine mammary gland tumorigenesis, including hyperplasia, adenoma, and carcinoma (Supplementary Fig. 1a).
a Clonal tracking strategy for a cytokeratin 8 (K8) promoter-driven color randomization during mammary gland tumorigenesis. In triple-transgenic mice, the MMTV promoter was used to drive the mammary gland-specific expression of the Polyoma virus middle T oncogene (MMTV–PyMT) and a stepwise cancer progression. The K8 promoter was used to drive the expression of tamoxifen-inducible Cre recombinase (K8-CreERT2) in a portion of mammary gland cells. The Confetti expression cassette integrated into the Rosa26 locus (R26-CBW) was used to clonally trace individual cells. Upon tamoxifen treatment of triple-transgenic female mice, activation of Cre recombinase led to the stochastic and stable expression of one of the four fluorescent proteins CFP, GFP, YFP, and RFP in K8-expressing luminal epithelial cells and their progeny in the mammary gland, allowing clonal tracking. b Schematic illustration of tamoxifen administration of 5-week-old female MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice to allow clonal tracking for 1, 4.5, or 8 weeks or until the maximum tumor volume had been reached. c Microscopic images of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mammary gland tumors following 1, 4.5, and 8 weeks of tracing or until the maximum tumor volume had been reached. In addition to visualization of CFP (Cyan), GFP (green), YFP (yellow), and RFP (red), sections have been immuno-stained for K8 (magenta) and DAPI (gray) (n = 3 mice per time point). Scale bars, 50 µm.
Cre recombinase activity was then induced in K8-expressing luminal epithelial cells of 5-week-old female MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice by injection of one bolus of tamoxifen, similar to other clonal tracing studies in this mouse model [23]. Cre recombinase-mediated recombination of the Confetti cassette led to the random and stable expression of one color of the four Confetti reporter fluorescence proteins in individual mammary gland tumor cells and their progenies: membranous cyan fluorescent protein, nuclear green fluorescent protein, cytoplasmic yellow fluorescent protein, and cytoplasmic red fluorescent protein (Supplementary Fig. 2). The outgrowth of clonal cell populations within primary tumors was then analyzed after 1, 4.5, and 8 weeks of tamoxifen injection, or when the maximum primary tumor volumes had been reached. These different time points allowed the tracing of clonal subpopulations in the various breast cancer stages, such as hyperplasia, adenoma, carcinoma, and pulmonary metastases (Fig. 1b). Immunofluorescence staining of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 primary tumors for K8/18 expression verified the co-localization of recombined cells with K8/18 expression throughout tumor progression (Fig. 1c). The applied bolus of tamoxifen did not substantially affect primary tumor growth or malignant progression, tumor cell proliferation, apoptosis, immune cell infiltration, or tumor microvessel density, as has been reported previously [23] (Supplementary Fig. 1b).
Tumor progression is associated with clonal dominance
To decipher tumor heterogeneity during the various stages of breast carcinogenesis, female MMTV–PyMT;R26-CBW K8-CreERT2 mice were color-induced at 5 weeks of age, sacrificed at different time points (6, 9.5, and 13 weeks of age or once maximum tumor volume had been reached), and clonal populations defined by the continuous and homogenous expression of one-color fluorescence protein were analyzed. FACS analysis and whole-mount imaging of mammary tumors of MMTV–PyMT;R26-CBW;K8-CreERT2 mice 1 week after tamoxifen-induced color randomization revealed single recombined luminal progenitor cells with a rather low total labeling efficiency of around 3% in all four colors (but with a slight bias in frequency toward GFP and RFP), enabling clonal tracking over time (Supplementary Fig. 3).
Longitudinal imaging of MMTV–PyMT;R26-CBW;K8-CreERT2 mice harboring fluorescence-labeled primary tumors throughout progression to malignancy was associated with the emergence of clonal populations in the early stages of primary tumor development and culminated in a dominance of large individual clones in the later stages of primary tumor progression (Fig. 2a). This clonal restriction was manifested by a decrease in the number of clones and an increase in the area of clonal populations over time (Fig. 2b). Of note, it could not be excluded that clonal populations in proximity to each other and expressing the same color fluorescent protein have merged during clonal outgrowth.
a Representative fluorescence microscopy images (upper panels) and 3D reconstructions (lower panels) of color-induced mammary gland tumors of MMTV–PyMT;R26-CBW;K8-CreERT2 mice at 6, 9.5, and 13 weeks of age or when the maximum tumor volume had been reached. DAPI, gray; n = 7–17 mice per time point. Scale bars, 1000 µm. 3D reconstructions: DAPI, blue; n = 3–6 mice per time point. Scale bars, 100 µm. b Quantification of the numbers (left panel) and area sizes of clonal tumor cell populations (right panel) within tumors of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice at 6, 9.5, and 13 weeks of age or when the maximum tumor volume had been reached (log ten scale). n = 6–15 mice per time point. *p < 0.05, ***p < 0.001, using unpaired, two-tailed Student’s t test. c Z-stacks of primary tumors of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice when the maximum tumor volume had been reached were quantified for the percentage events in the following categories: disseminated single tumor cells, monochromatic tumor cell clusters (TCCs) (without and in close proximity to single cells), and polychromatic TCCs. n = 5 with one to three fields of view each. ***p < 0.001, using paired, two-tailed Student’s t test. d Representative fluorescence microscopy images (upper panels) and 3D reconstructions (lower panels) of an invasive and disseminating single cell (GFP) and mono- and polychromatic tumor cell clusters at the invasive front of late-stage tumors of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice. DAPI, blue; n = 5. Scale bars, upper panels, 250 µm; rectangle, 50 µm. Scale bars, middle and lower panels, 30 µm.
We next assessed the level of tumor cell heterogeneity during tumor invasion, the initial step in the metastatic dissemination of tumor cells. Confocal microscopy imaging of thick cryosections of tamoxifen-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mammary gland tumors and subsequent 3D reconstructions revealed three modes of dissemination at the invasive fronts of primary tumors: disseminating single cells, monochromatic tumor cell clusters, and polychromatic tumor cell clusters harboring at least two differently colored clonal populations. Quantification of the mode of dissemination at the invasive front revealed an equal contribution of disseminated single tumor cells and tumor cell clusters. However, polychromatic tumor cell clusters seemed to occur less frequently as compared to disseminated single tumor cells and monochromatic tumor cell clusters (Fig. 2c, d).
Polyclonal metastatic spread
To address the functional effects of the observed tumor cell heterogeneity at the invasive front in the subsequent steps of the metastatic cascade, we next analyzed circulating tumor cells (CTCs) in the bloodstream and pulmonary metastases in MMTV–PyMT;R26-CBW;K8-CreERT2 mice bearing fluorescence-labeled late-stage tumors. For CTC analysis, up to 1.3 ml of blood was isolated by cardiac puncture and CTCs were captured through size-based microfluidic enrichment. Microscopic analysis of microfluidic cassettes identified CTCs expressing one of the four fluorescence markers in only 3 out of 12 mice analyzed. In these three mice, we observed on average one single CTC and five CTC clusters harboring on average 23 cells per ml of blood (Fig. 3a and Supplementary Fig. 4a), in line with previous findings with the MMTV–PyMT mouse model [29]. The CTC clusters were either monochromatic or polychromatic (with at least two different colors) and most of them were found to cluster with non-recombined tumor cells or other non-recombined cell types, presumably leukocytes [29] (Fig. 3a and Supplementary Fig. 4a).
a Representative fluorescence microscopy images of single circulating tumor cells (CTCs) and monochromatic, polychromatic, and poly-cell type CTC clusters of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice at maximum tumor volume (left panels). Pie chart indicating that primary mammary gland tumors of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice shed few single CTCs and mainly CTC clusters (right panel). n = 3, out of 12 mice analyzed. Scale bars, 25 µm. b Representative fluorescence microscopy images of small or large and mono- or polychromatic metastatic lesions of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice. n = 31. Scale bars, 50 µm. c Quantification of the numbers and average metastatic areas of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice at 13 weeks of age and when the maximum tumor volume had been reached. n = 6–12 mice per time point. *p < 0.05, using unpaired, two-tailed Student’s t test. d Quantification of the numbers and average area of monochromatic vs. polychromatic metastatic lesions of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice when the maximum tumor volume had been reached (left and middle panel). Quantification of the average metastatic area of macroscopic monochromatic vs. polychromatic metastatic lesions is shown on the right. n = 12. *p < 0.05, **p < 0.01, using paired, two-tailed Student’s t test.
Once maximum tumor volume had been reached in color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice, pulmonary metastases were analyzed by histological microscopy analysis of serial sections of whole lungs. Approximately 30 metastatic lesions were detected on average in the lungs of each mouse which ranged in size from small microscopic to large macroscopic nodules and exhibited mono- and polychromatic cellular composition (with at least two differently colored clonal populations) (Fig. 3b, c). Interestingly, polychromatic metastatic lesions were found larger in size but less frequent in numbers than the monochromatic ones in the triple-transgenic mice (Fig. 3d). To validate these findings in an additional mouse model, immunocompromised NSG mice were grafted in the mammary fat pad with a primary tumor fragment of triple-transgenic mice. Also in this setting, polychromatic metastatic lesions were found larger than monochromatic metastatic lesions (Supplementary Fig. 4b). The observation of larger polychromatic metastatic lesions might be the consequence of an outgrowth benefit of polyclonal cell populations at the late stages of tumor progression, when metastatic outgrowth is maximal, or reflecting an earlier arrival of polyclonal cell populations to the lungs (Fig. 3c, d and Supplementary Fig. 4c).
Substantial heterogeneity across and within tumor stages
To investigate molecular tumor heterogeneity during multi-stage oncogenesis in color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice at the gene expression level, zonal patches of clonal populations of various stages within primary tumors as well as metastases were isolated by laser capture microdissection and subjected to RNA sequencing (Fig. 4a and Supplementary Fig. 5a). Since CFP expression was underrepresented in late-stage tumors [30], CFP-expressing clonal populations were excluded from further analysis (Supplementary Fig. 3c, right panel). To avoid potential confounders in the RNA-sequencing analysis, identical size samples were isolated from one-color clonal tumor patches (Supplementary Fig. 5b). Unsupervised hierarchical clustering of the top 500 most variable genes specific for hyperplasia, adenoma, carcinoma, and metastatic lesions suggested a substantial level of heterogeneity across the different stages of carcinogenesis (Fig. 4b). Comparing gene expression in each tumor stage vs. gene expression in normal mammary glands revealed stage-specific gene sets and a consistent decrease in extracellular matrix organization and collagen degradation in every tumor stage (reactome) (Supplementary Fig. 5c). Interestingly, every stage with the exception of carcinoma harbored cell cycle-related terms in their top 10 reactome terms (Fig. 4c, left panels, highlighted in bold). Nevertheless, an assessment of functional contribution of tumor heterogeneity to metastatic breast cancer progression may not only require a stage-wise comparison, but also an interrogation of the level of heterogeneity between various subpopulations within a specific tumor stage.
a Outgrowth of clonal cell populations was analyzed when the maximum tumor volume comprising all tumor stages (normal mammary gland, hyperplasia, adenoma, carcinoma, pulmonary metastases) had been reached in color-induced female MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice. Zonal patches of clonal populations of the primary tumor as well as metastases were isolated by laser capture microdissection (encircled and boxed regions) and samples obtained were subjected to RNA sequencing. n = 3 mice. Scale bars, 150 µm. b Unsupervised hierarchical clustering of the top 500 most variable genes across the various stages of mammary gland carcinoma progression. Hyperplasia, n = 5; adenoma, n = 8; carcinoma, n = 9; metastases, n = 7. c List of the top 10 reactome terms (percentage of expressed genes per term) of the top 500 most variable genes within each tumor stage are depicted in the left panels. Cell cycle-related terms are highlighted in bold. Unsupervised hierarchical clustering of the top 500 most variable genes within each stage (right panels) revealed stage-specific subpopulations (highlighted in color-coded rectangles). d Bar graph representing the percentage of all upregulated cell cycle-related reactome terms resulting from the comparisons of stage-specific subpopulations vs. normal mammary gland. n = 2 for each tumor stage. Reactome terms related to cell cycle regulation are highlighted in bold. The following color code was used throughout the figure: hyperplasia, yellow; adenoma, orange; carcinoma, red; metastasis, blue.
Strikingly, unsupervised hierarchical clustering of the top 500 most variable genes of each tumor stage revealed a tremendous heterogeneity in gene expression profiles and the identification of distinct subpopulations within each tumor stage (Fig. 4c). Intriguingly, when each tumor stage-specific subpopulation was compared to gene expression profiles of normal murine mammary glands, each tumor stage was found to harbor one subpopulation which was enriched for cell cycle-related reactome terms (Fig. 4d). Based on this comparison, we classified each stage-specific subpopulation as fast-proliferating or slow-proliferating for further analysis. A comparison of the pathways (top 10 KEGG terms) of all fast-proliferating subpopulations of all tumor stages to all slow-proliferating subpopulations validated their prominent vs. absent proliferative phenotype, respectively (Supplementary Fig. 6a). Integrated Motif Activity Response Analysis on all subpopulations pointed toward a stage-specific transcriptional regulation of the fast-proliferating vs. slow-proliferating phenotype. This analysis revealed the transcription factor binding motifs Etv1/Etv5/Gabpa (highlighted in red) and Gtf2i/Gtf2f1 (highlighted in green) as the top regulatory hubs orchestrating the fast- and slow-proliferating phenotypes, respectively, of all tumor stages (Supplementary Fig. 6b).
Intra-tumor stage heterogeneity: differences in cell proliferation
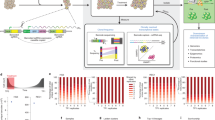
In line with the fast-proliferating vs. slow-proliferating clonal populations in all tumor stages of MMTV–PyMT;R26-CBW;K8-CreERT2 mice, gene-set enrichment analysis of each tumor stage confirmed for the fast-proliferating subpopulations of each tumor stage a significant enrichment in genes implicated in the positive regulation of cell cycle (Fig. 5a). Immunofluorescence staining on histological sections of primary tumors and lung metastases for phospho-histone H3 (pH3), a marker for proliferating cells, revealed the existence of fast- vs. slow-proliferating subpopulations in Confetti-labeled tumors of MMTV–PyMT;R26-CBW;K8-CreERT2 mice. Within one tumor section, several fast-proliferating clones in close proximity to slow-proliferating clones could be detected in hyperplasia, adenoma, carcinoma, and metastases (Fig. 5b and Supplementary Fig. 7a). Quantification of pH3+-nuclei per clonal area verified the existence of fast- vs. slow-proliferating clones within each tumor stage, which was accompanied by bigger vs. smaller clonal areas, respectively (Fig. 5b and Supplementary Fig. 7b). Immunofluorescence staining on histological sections of primary tumors for Ki67, another well-established marker for proliferating cells, revealed a similar expression pattern for pH3 and Ki67 (Supplementary Fig. 7c) and a comparable distinction in fast and slow-proliferating subclones in the different tumor stages of Confetti-labeled MMTV–PyMT;R26-CBW;K8-CreERT2 mice (Supplementary Fig. 7d). This separation of fast- and slow-proliferating subclones has also been observed with monoclonal cell lines established from primary tumors and from pulmonary metastatic lesions of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice (Fig. 5c). Notably, the clonal differences in the cell lines’ proliferation rates were found independent of their epithelial-to-mesenchymal transition state (data not shown).
a Gene-set enrichment analysis (GSEA) revealed an enrichment in gene sets involved in a positive regulation of cell cycle for the fast-proliferating subpopulations of each tumor stage. NES normalized enrichment score. b Representative immunofluorescence microscopy images of MMTV-PyMTR26-CBW; K8-CreERT2 mammary gland tumors, stained for phospho-histone H3 (pH3, arrowheads, and magenta) and DAPI (blue) (left panels). Lesions below or above the average of pH3+ nuclei/mm2 per tumor stage per mouse were classified as slow- or fast-proliferating clonal populations, respectively. Differences in pH3+ nuclei per area of slow- and fast-proliferating clonal populations within each stage are depicted on the right. n = 4 mice. ****p < 0.0001, using unpaired, two-tailed Mann–Whitney U test. Scale bars, 50 µm. c Microscopic images of monoclonal cell lines derived from primary tumors and metastatic lesions of color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice were labeled for pH3 (arrowheads; green) and DAPI (blue) (left panels). Slow- and fast-proliferating cell lines isolated from tumor (red, n = 9) and metastatic lesions (blue, n = 9) differed in their percentage of pH3+ cells (right panel). *p < 0.05, **p < 0.01, using unpaired, two-tailed Mann–Whitney U test. Scale bars, 50 µm. d Representative light microscopy image of a human breast cancer biopsy (invasive ductal carcinoma, luminal B) immunolabeled for Ki67 indicated intra-stage heterogeneity due to proliferation differences (left panels). Lesions below or above the average of Ki67+ nuclei/mm2 per tumor stage per sample were classified as slow- or fast-proliferating clonal populations, respectively. Quantification of Ki67+ nuclei per area of slow- and fast-proliferating clonal populations within invasive carcinoma lesions of luminal A- (Lum A), luminal B- (Lum B), and HER2-enriched (HER2) breast cancer samples is shown on the right. n = 9 patients, with one to four sections each; luminal A: n = 3; luminal B: n = 5; HER2 enriched: n = 1. *p < 0.05, ****p < 0.0001, using unpaired, two-tailed Mann–Whitney U test (fast vs. slow) and unpaired, two-tailed Student’s t test (fast vs. fast). Scale bar, 2000 µm. The following color code was used throughout the figure: hyperplasia, yellow; adenoma, orange; carcinoma, red; metastasis, blue.
To assess whether a comparable distinction in fast- and slow-proliferating cancer cell subpopulations is also found in tumors of breast cancer patients, we analyzed luminal A-, luminal B-, and HER2-enriched human breast cancer samples, which had been stained for the proliferation marker Ki67 in the course of clinical follow-up. Here, invasive lesions harboring Ki67+ cells clustering in close proximity to each other were defined as clonal outgrowth (Fig. 5d, left panel). Consistent with the intra-stage proliferative heterogeneity observed in murine mammary gland tumors, fast-proliferating and slow-proliferating lesions were also observed with high significance in invasive carcinomas of luminal A-, luminal B-, and HER2-enriched human breast cancer samples. The differences in the Ki67 index between luminal A-, luminal B-, and HER2-enriched breast cancer samples were mainly due to an increase in fast-proliferative clonal populations in luminal B- and HER2-enriched breast cancer samples when compared to the luminal A subtype (Fig. 5d, right panel). Yet also here, considerable inter- and intra-tumoral heterogeneity is observed in all samples of the breast cancer subtypes (Supplementary Fig. 7e).
Intra-tumor stage heterogeneity: differences in oxidative phosphorylation, apoptosis, hypoxia response, and reactive oxygen species (ROS) detoxification
Further gene-set enrichment analysis revealed additional specific characteristics of the fast-proliferating and slow-proliferating subpopulations. While the slow-proliferating populations of tumor cells did not apparently correlate with any specific gene expression profile or gene-set enrichments, the fast-proliferating subpopulations were further enriched for oxidative phosphorylation, apoptosis, cellular responses to hypoxia, and detoxification of ROS (Fig. 6a–d and Supplementary Fig. 8). These computational results were validated by immunofluorescence staining of primary tumor sections of single-transgenic MMTV–PyMT mice for Ndufa2 and with TdT-mediated dUTP-X nick end labeling (TUNEL), markers for complex I of the oxidative phosphorylation system and for DNA strand breaks generated during apoptosis, respectively (Fig. 6a, b). Indeed, the fast-proliferating clonal populations, particularly from adenoma lesions, displayed a specific increase in Ndufa2 staining intensity as compared to the slow-proliferating populations. Furthermore, fast-proliferating clonal populations, especially from adenoma lesions, harbored areas that were specifically high in TUNEL staining, indicating a high extent of apoptosis. Furthermore, gene-set enrichment analysis for all fast-proliferating clonal populations vs. all slow-proliferating clones of each stage indicated a significant enrichment in gene sets involved in TGFβ receptor signaling in EMT, cell adhesion molecule binding, tumor invasiveness, and metastasis in the fast-proliferative clonal populations of the carcinoma and metastasis lesions (Supplementary Fig. 8).
a Gene-set enrichment analysis (GSEA) revealed for all fast-proliferating clonal lesions (n = 8) vs. all slow-proliferating lesions ones (n = 9) an enrichment in gene sets involved in oxidative phosphorylation (left panel). NES normalized enrichment score. Lesions below or above the average of pH3+ nuclei/mm2 per tumor stage per MMTV–PyMT mouse were classified as slow- or fast-proliferating clonal populations, respectively. Quantification of Ndufa2 mean intensity confirmed an increased capacity for oxidative phosphorylation in fast-proliferating clones (right panel, black labels: total). n = 3. *p < 0.05, using unpaired, two-tailed Mann–Whitney U test. b GSEA for all fast-proliferating clonal populations (n = 8) vs. all slow-proliferating clones (n = 9) revealed an enrichment in gene sets involved in the regulation of apoptosis (left panel). NES normalized enrichment score. Lesions below or above the average of pH3+ nuclei/mm2 per tumor stage per MMTV–PyMT mouse were classified as slow- or fast-proliferating clonal populations, respectively. Quantification of TUNEL+ cells per area validates the high incidence of cell death in fast-proliferative populations in primary tumors of MMTV–PyMT mice (right panel, black labels: total). n = 3. *p < 0.05, using unpaired, two-tailed Mann–Whitney U test. c GSEA for all fast-proliferating clonal populations (n = 8) vs. all slow-proliferating clones (n = 9) indicated an enrichment in gene sets involved in the cellular response to hypoxia. NES normalized enrichment score. d GSEA for all fast-proliferating clonal populations (n = 8) vs. all slow-proliferating clones (n = 9) suggested an enrichment in gene sets involved in the detoxification of reactive oxygen species (ROS). NES normalized enrichment score. e Lesions below or above the average of pH3+ nuclei/mm2 per tumor stage per MMTV–PyMT mouse (13 weeks) were classified as slow- or fast-proliferating clonal populations, respectively. Quantification of CD45+ cells per area (mm2) indicated that the proliferation phenotype is not attributable to variations in immune cell infiltrates. n = 3. *p < 0.05, ns = not significant, using unpaired, two-tailed Mann–Whitney U test. f Lesions below or above the average of pH3+ nuclei/mm2 per tumor stage per MMTV–PyMT mouse (13 weeks) were classified as slow- or fast-proliferating clonal populations, respectively. Quantification of CD31+ vessels per area (mm2) revealed that the proliferation phenotype is not attributable to tumor microvessel density. n = 3. ns = not significant, using unpaired, two-tailed Mann–Whitney U test. The following color code was used throughout the figure: hyperplasia, yellow; adenoma, orange; carcinoma, red; metastasis, blue.
Immunofluorescence staining of primary tumor sections for CD45 and CD31 showed that the observed fast- and slow-proliferating phenotypes were not attributable to differences in immune cell infiltrates or tumor microvessel densities (Fig. 6e, f). Moreover, computational analysis for the number of total reads/genes expressed in all laser capture-microdissected clonal populations excluded a confounding factor by the number of total reads and genes expressed per sample (Supplementary Fig. 9).
Thus, throughout progression to malignancy, each tumor stage comprised not only inert but also highly dynamic clones, which were characterized by increased proliferation, oxidative phosphorylation, a high cellular turnover, hypoxia response, ROS detoxification, and larger clone sizes.
Chemotherapeutic reagents fail to repress fast-proliferative clonal populations
To elucidate whether the intra-stage proliferative tumor heterogeneity may have an impact on therapy response, color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice were treated with chemotherapeutic reagents, including paclitaxel, cyclophosphamide, and doxorubicin, for 3 consecutive weeks after first tumor palpation. Analysis of primary tumor weight and of serial histological sections of the lung revealed cyclophosphamide as the most efficient chemotherapeutic reagent in this mouse model; it decreased average tumor weight, metastatic seeding (number metastatic lesions), and metastatic outgrowth (metastatic area). Paclitaxel treatment reduced metastatic seeding and outgrowth to a lesser extent, while doxorubicin merely inhibited metastatic outgrowth (Fig. 7a, b). However, immunofluorescence staining of histological sections of primary tumors for pH3 and Ki67 identified a significant reduction of pH3+ cells and a nonsignificant reduction of Ki67+ cells only with doxorubicin, suggesting that the proliferative cell populations are not substantially affected by chemotherapy and thus are not explaining the reduction in tumor burden by chemotherapy (Fig. 7c and Supplementary Fig. 10a). Only the quantification of pH3+-nuclei per clonal area, but not of Ki67+-nuclei per clonal area, showed a decrease in proliferation especially in clonal populations of adenoma and carcinoma but not in hyperplasia (Fig. 7d and Supplementary Fig. 10b). Analysis of slow- and fast-proliferating clonal populations within the various tumor stages revealed different modes of action for the various chemotherapies. While cyclophosphamide mainly targeted slow-proliferating populations within adenoma and carcinoma, paclitaxel inhibited the proliferation of fast-proliferating clonal populations in the adenoma stage. On the other hand, doxorubicin efficiently decreased slow-proliferating populations in hyperplasia, adenoma, and carcinoma, and fast-proliferating populations only in the adenoma stage (Fig. 7e). Hence, although conventional chemotherapies were thought to specifically target fast-proliferating tumor cells, the results indicated that the observed therapeutic effects were mostly due to a depletion of slow-proliferative clonal populations, whereas the fast-proliferative clonal populations, especially in carcinoma lesions, remained unaffected (Fig. 7e and Supplementary Fig. 10c). Interestingly, the chemotherapeutic effects on tumor burden were not attributable to changes in programmed cell death between slow-proliferative and fast-proliferative clonal cell populations, as determined by immunofluorescence staining of tumor sections for cleaved caspase 3 (Supplementary Fig. 10d–f).
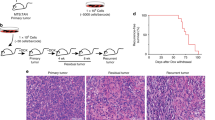
a Once tumor growth was initiated, color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice were treated with paclitaxel, cyclophosphamide, and doxorubicin for 3 consecutive weeks and primary tumors were collected. Cyclophosphamide was most efficient in reducing average tumor weight. n = 3 mice per treatment group. *p < 0.05, using unpaired, two-tailed Student’s t test. b Lungs were collected and serially sectioned to determine numbers and average metastatic areas under chemotherapy administration, with paclitaxel and cyclophosphamide being the most efficient in inhibiting these parameters. n = 3 mice per treatment group. ***p < 0.001, using unpaired, two-tailed Student’s t test. c Immunofluorescence staining and quantification of primary tumor sections of MMTV-PyMTR26-CBW; K8-CreERT2 for pH3+ nuclei/mm2 indicated doxorubicin as a potent proliferation inhibitor. n = 3 mice per treatment group. *p < 0.05, using unpaired, two-tailed Student’s t test. d Quantification of primary tumor sections of MMTV-PyMTR26-CBW; K8-CreERT2 for pH3+ nuclei/mm2 revealed differences in pH3+ nuclei within each stage due to chemotherapy administration. n = 3 mice per treatment group. *p < 0.05, **p < 0.01, using unpaired, two-tailed Student’s t test. e Lesions below or above the average of pH3+ nuclei/mm2 per tumor stage per mouse were classified as slow- or fast-proliferating clonal populations, respectively. The graph depicts differences in pH3+ nuclei per area of slow- and fast-proliferating clonal populations within each stage following chemotherapy treatment (Veh vehicle, Pax paclitaxel, Cyc cyclophosphamide, Dox doxorubicin). n = 3 mice per treatment group. *p < 0.05, ***p < 0.001, ****p < 0.0001, using unpaired, two-tailed Student’s t test. The following color code was used throughout the figure: hyperplasia, yellow; adenoma, orange; carcinoma, red.
Discussion
Using multicolor clonal tracking in combination with laser capture microdissection and RNA sequencing in MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice [24,25,26,27,28, 31,32,33,34,35], we have assessed clonal tumor heterogeneity not only across the various stages of neoplastic progression but also within individual tumor stages of metastatic breast cancer (Supplementary Fig. 11). Longitudinal imaging of Confetti-traced tumors identified clonal populations of various stages in primary tumors with the emergence of clonal dominance. Yet, metastatic spread appears associated with polyclonality at the primary tumor invasive front, in CTCs in the bloodstream, and in metastatic lesions in the lungs. Comparative gene expression analysis indicates a substantial level of heterogeneity between the different oncogenic stages and, even more intriguing, within the same tumor stages. Unexpectedly, this intra-stage tumor heterogeneity is due to differences in tumor cell proliferation: each tumor stage harbors slow proliferating, inert subpopulations as well as fast proliferating, highly dynamic clones. The fast-proliferating clonal populations are further characterized by an enrichment in the expression of genes active in oxidative phosphorylation, programmed cell death, a cellular response to hypoxia, detoxification of ROS and by increased clonal overgrowth.
In line with our findings, the use of Confetti clonal tracing systems in mouse models of breast and pancreatic carcinomas, skin squamous cell carcinomas and sarcomas has revealed a similar replacement of a high polyclonality in early tumor lesions by the overgrowth of few clonal populations in later tumor stages [10, 11, 21,22,23]. Furthermore, ours and other studies reveal the existence of polyclonal metastases in autochthonous and in tumor transplantation models [10, 20, 21]. However, whether clonal expansion and polyclonal spread are due to acquired mutations and/or to beneficial clonal cooperativity between various clonal subpopulations remain open [36]. The presence of polychromatic and poly-cell type CTC clusters identified by us and others [20, 22, 29, 37], as well as our observation of larger polychromatic metastatic lesions, suggests an outgrowth benefit for polyclonal lesions.
Consistent with previous findings, we observe a tremendous transcriptomic diversity between the different stages of carcinogenesis [10, 11, 38]. As expected, subclonal diversification and heterogeneity are high between the longitudinal stages of tumor progression. However, to adequately assess the functional contribution of tumor heterogeneity to breast cancer progression, it is important to additionally interrogate the extent of heterogeneity between subpopulations within the same tumor stages. Indeed, we find an intriguing heterogeneity between clonal subpopulations within individual stages of malignant tumor progression. This intra-stage heterogeneity is mainly based on differences in tumor cell proliferation, where the fast-proliferating subpopulation is further enriched in oxidative phosphorylation, cellular turnover, hypoxia response, and ROS detoxification. Consistent with our findings, bulk RNA sequencing of MMTV–PyMT primary tumors has previously reported a role of cell cycle-related genes in tumor progression [38]. Other studies have found slow-cycling subpopulations in other cancer types to be essential for continuous tumor growth, invasion, metastatic progression, and drug resistance [39,40,41,42]. Furthermore, the proliferative phenotype has been found predominantly accompanied by oxidative metabolism [43, 44]. Our study also reveals a tight association of the proliferative phenotype with a high oxidative capacity, ROS detoxification, and larger clonal outgrowth, yet also with an enrichment in cell turnover, altogether pointing to an increased metabolic activity in fast-proliferating tumor subpopulations. Since our study uses a K8-driven Cre recombinase in the MMTV–PyMT mouse model, representing luminal B breast cancer, other models need to be developed in future studies in order to explore tumor heterogeneity and clonal evolution also in other breast cancer subtypes.
The finding of slow and fast-proliferating subpopulations in the various stages of mammary gland tumorigenesis is also found to be translatable to human breast cancer patients. The analysis of cellular Ki67 levels confirms considerable inter- and intra-tumor stage heterogeneity in human breast cancer punch biopsies. Intriguingly, low or high zonal expression of the proliferation marker Ki67 is found within lesions of invasive carcinomas of luminal A-, luminal B-, and HER2-enriched breast cancer patients. Interestingly, the Ki67 index of the fast-proliferative clonal populations determines the respective breast cancer subtype, with luminal B and HER2-positive breast cancers displaying an increase in fast-proliferative clonal populations. Unexpectedly and clinically relevant, various chemotherapeutic reagents (paclitaxel, cyclophosphamide, and doxorubicin) have mainly depleted slow-proliferative clonal subpopulations in MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice, while the fast-proliferative clonal populations have remained unaffected. Yet, it is difficult to determine whether the fast-proliferative subpopulations have become slow proliferative during the course of the treatment or whether they have developed resistance against the chemotherapies, warranting further investigations. These data may be of great importance for a patient’s therapy response and tumor recurrence. To date, Ki67 evaluation is not implemented in the guidelines of the American Society of Clinical Oncology [45]. Currently, histological parameters, such as the morphological subtype in combination with the expression of hormone receptors, HER2 status, and further genomic and transcriptomic molecular profiling, diagnostically determine distinct breast cancer subtypes and their respective therapeutic options. Yet, to differentiate between luminal A and B breast cancers and to predict therapeutic options in a neoadjuvant setting, Ki67 expression levels are being considered [46, 47]. However, the temporal and spatial clonal variation and the intra-stage proliferation differences specified here further add to the complexity and highlight the need to define a Ki67-cutoff value for clinical decision-making [48]. Furthermore, the observed intra-stage heterogeneity in tumor cell proliferation may enhance a tumor’s robustness to anti-cancer therapies. As exemplified here, conventional chemotherapies seem to exert their therapeutic effect by specifically targeting slow-proliferating cells, yet leaving the fast-proliferating cells unaffected. Presumably, these fast-proliferating subpopulations may be responsible for the occurrence of therapy resistance and clinical progression. These insights need to be considered for clinical decision-making.
Methods
Mice and Confetti labeling
MMTV–PyMT, R26-Confetti (here: R26-CBW) and K8-CreERT2 mice have been previously described [24,25,26,27,28]. To generate MMTV–PyMT;R26-CBW;K8-CreERT2 triple-transgenic mice, male MMTV–PyMT;R26-CBW mice were crossed to female K8-CreERT2 mice. To stochastically express one of the four Confetti reporter fluorescent proteins, female MMTV–PyMT;R26-CBW;K8-CreERT2 mice of 5 weeks of age were induced intra-peritoneally with 5 mg tamoxifen (Sigma-Aldrich) dissolved in 200 µl sunflower oil (Sigma-Aldrich) containing 10% EtOH (Scharlau), similar to a previous clonal tracing study with the MMTV–PyMT, R26-Confetti model [23]. These animal experiments have been performed under approval by the Swiss Federal Veterinary Office and the Cantonal Veterinary Office, Basel-Stadt, Switzerland (permit numbers 1878, 1907, and 1908).
Treatment of MMTV–PyMT;R26-CBW;K8-CreERT2 mice with chemotherapeutic reagents
Once tumor growth was initiated, color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice were treated once a week intra-peritoneally with paclitaxel (20 mg/kg, T7191, Sigma-Aldrich), cyclophosphamide monohydrate (100 mg/kg, C7397, Sigma-Aldrich), doxorubicin hydrochloride (5 mg/kg, D1515, Sigma-Aldrich), and vehicle for 3 consecutive weeks. One-week post treatment, mice were sacrificed and primary tumors and lungs were isolated for further processing.
Laser capture microdissection and transcriptome sequencing
The outgrowth of clonal cell populations has been analyzed when the maximum allowed tumor volume comprising all stages (normal, hyperplasia, adenoma, carcinoma, and pulmonary metastases) had been reached in color-induced MMTV–PyMT;R26-CBW;K8-CreERT2 mice. By combining the recently published Geo-seq protocol using SmartSeq2 for laser capture-microdissected samples with a recent report for parallel DNA and RNA sequencing from single cells, representative cDNA libraries of laser captured regions from fixed tumor sections were generated [31,32,33,34,35]. In brief, 7-µm-thick cryosections were mounted onto Menzel Superfrost Plus slides and placed in decreasing and increasing ethanol solutions. Zonal patches of clonal populations of various stages in the primary tumor as well as monochromatic metastases were microdissected using a Zeiss Laser Capture Microscope (energy: 49, focus: 67, delta-energy: 15, delta-focus: 5; CFP (474/23), GFP (510/20), YFP (535/22), and RFP (620/60)) in Auto LPC mode. Clonal populations of normal mammary glands were pooled to obtain a sufficient sample size. Collected samples were first lysed at 37 °C for 30 min by using a digestion buffer containing proteinase K (150 mM NaCl, 100 mM Tris pH7.5, 0.5% Igepal, 0.5 µg/µl proteinase K) and for further 30 min at 42 °C in a 4 M GuSCN solution (Thermo Fisher). Biotinylated oligo-dT30VN beads were used to prime the reverse-transcriptase reaction and cDNA preamplification on polyadenylated mRNA (exception to GeoSeq protocol: dNTP mix contained 10 mM each of dATP, dCTP, dGTP, dTTP and SmartSeq2 ISPCR oligos were used, 5′-AAGCAGTGGTATCAACGCAGAGT-3′). cDNA quality was assessed by a Qubit Fluorometer with a dsDNA HS assay kit (Thermo Fisher), an Agilent high-sensitivity DNA chip for cDNA size distribution, and quantitative real-time PCR to check for the expression of housekeeping genes and the absence of gDNA.
Statistical evaluation
Graphs and statistical analyses were generated using GraphPad Prism software Version 8.4.1. To determine statistical significance, the (un)paired, two-tailed Student’s t test, the Mann–Whitney U test, the Kruskal–Wallis test, and Spearman correlations were applied (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001). Quantitative data were depicted as means ± SEM.
Data availability
The data sets generated and analyzed within the current study are deposited at Gene Expression Omnibus (GEO; RNA sequencing data: accession number GSE139390).
References
Banerji S, Cibulskis K, Rangel-Escareno C, Brown KK, Carter SL, Frederick AM, et al. Sequence analysis of mutations and translocations across breast cancer subtypes. Nature. 2012;486:405–9.
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Casasent AK, Schalck A, Gao R, Sei E, Long A, Pangburn W, et al. Multiclonal invasion in breast tumors identified by topographic single cell sequencing. Cell. 2018;172:205–17.e12.
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486:346–52.
Eirew P, Steif A, Khattra J, Ha G, Yap D, Farahani H, et al. Dynamics of genomic clones in breast cancer patient xenografts at single-cell resolution. Nature. 2015;518:422–6.
Kim C, Gao R, Sei E, Brandt R, Hartman J, Hatschek T, et al. Chemoresistance evolution in triple-negative breast cancer delineated by single-cell sequencing. Cell. 2018;173:879–93.e13.
Nguyen LV, Pellacani D, Lefort S, Kannan N, Osako T, Makarem M, et al. Barcoding reveals complex clonal dynamics of de novo transformed human mammary cells. Nature. 2015;528:267–71.
Nik-Zainal S, Van Loo P, Wedge DC, Alexandrov LB, Greenman CD, Lau KW, et al. The life history of 21 breast cancers. Cell. 2012;149:994–1007.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, et al. Molecular portraits of human breast tumours. Nature. 2000;406:747–52.
Reeves MQ, Kandyba E, Harris S, Del Rosario R, Balmain A. Multicolour lineage tracing reveals clonal dynamics of squamous carcinoma evolution from initiation to metastasis. Nat Cell Biol. 2018;20:699–709.
Rios AC, Capaldo BD, Vaillant F, Pal B, van Ineveld R, Dawson CA, et al. Intraclonal plasticity in mammary tumors revealed through large-scale single-cell resolution 3D imaging. Cancer Cell. 2019;35:618–32.e6.
Shah SP, Morin RD, Khattra J, Prentice L, Pugh T, Burleigh A, et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature. 2009;461:809–13.
Shah SP, Roth A, Goya R, Oloumi A, Ha G, Zhao Y, et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature. 2012;486:395–9.
van ‘t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415:530–6.
Wang Y, Waters J, Leung ML, Unruh A, Roh W, Shi X, et al. Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature. 2014;512:155–60.
Yates LR, Gerstung M, Knappskog S, Desmedt C, Gundem G, Van Loo P, et al. Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat Med. 2015;21:751–9.
Yates LR, Knappskog S, Wedge D, Farmery JHR, Gonzalez S, Martincorena I, et al. Genomic evolution of breast cancer metastasis and relapse. Cancer Cell. 2017;32:169–84.e7.
Nowell PC. The clonal evolution of tumor cell populations. Science. 1976;194:23–8.
Greaves M, Maley CC. Clonal evolution in cancer. Nature. 2012;481:306–13.
Cheung KJ, Padmanaban V, Silvestri V, Schipper K, Cohen JD, Fairchild AN, et al. Polyclonal breast cancer metastases arise from collective dissemination of keratin 14-expressing tumor cell clusters. Proc Natl Acad Sci USA. 2016;113:E854–63.
Maddipati R, Stanger BZ. Pancreatic cancer metastases harbor evidence of polyclonality. Cancer Discov. 2015;5:1086–97.
Tang YJ, Huang J, Tsushima H, Ban GI, Zhang H, Oristian KM, et al. Tracing tumor evolution in sarcoma reveals clonal origin of advanced metastasis. Cell Rep. 2019;28:2837–50.e35.
Zomer A, Ellenbroek SI, Ritsma L, Beerling E, Vrisekoop N, Van Rheenen J. Intravital imaging of cancer stem cell plasticity in mammary tumors. Stem Cells. 2013;31:602–6.
Guy CT, Cardiff RD, Muller WJ. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol Cell Biol. 1992;12:954–61.
Lin EY, Jones JG, Li P, Zhu L, Whitney KD, Muller WJ, et al. Progression to malignancy in the polyoma middle T oncoprotein mouse breast cancer model provides a reliable model for human diseases. Am J Pathol. 2003;163:2113–26.
Livet J, Weissman TA, Kang H, Draft RW, Lu J, Bennis RA, et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature. 2007;450:56–62.
Koren S, Reavie L, Couto JP, De Silva D, Stadler MB, Roloff T, et al. PIK3CA(H1047R) induces multipotency and multi-lineage mammary tumours. Nature. 2015;525:114–8.
Van Keymeulen A, Rocha AS, Ousset M, Beck B, Bouvencourt G, Rock J, et al. Distinct stem cells contribute to mammary gland development and maintenance. Nature. 2011;479:189–93.
Szczerba BM, Castro-Giner F, Vetter M, Krol I, Gkountela S, Landin J, et al. Neutrophils escort circulating tumour cells to enable cell cycle progression. Nature. 2019;566:553–7.
Martinez RJ, Neeld DK, Evavold BD. Identification of T cell clones without the need for sequencing. J Immunol Methods. 2015;424:28–31.
Chen J, Suo S, Tam PP, Han JJ, Peng G, Jing N. Spatial transcriptomic analysis of cryosectioned tissue samples with Geo-seq. Nat Protoc. 2017;12:566–80.
Macaulay IC, Haerty W, Kumar P, Li YI, Hu TX, Teng MJ, et al. G&T-seq: parallel sequencing of single-cell genomes and transcriptomes. Nat Methods. 2015;12:519–22.
Macaulay IC, Teng MJ, Haerty W, Kumar P, Ponting CP, Voet T. Separation and parallel sequencing of the genomes and transcriptomes of single cells using G&T-seq. Nat Protoc. 2016;11:2081–103.
Picelli S, Bjorklund AK, Faridani OR, Sagasser S, Winberg G, Sandberg R. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat Methods. 2013;10:1096–8.
Picelli S, Faridani OR, Bjorklund AK, Winberg G, Sagasser S, Sandberg R. Full-length RNA-seq from single cells using Smart-seq2. Nat Protoc. 2014;9:171–81.
Tabassum DP, Polyak K. Tumorigenesis: it takes a village. Nat Rev Cancer. 2015;15:473–83.
Aceto N, Bardia A, Miyamoto DT, Donaldson MC, Wittner BS, Spencer JA, et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell. 2014;158:1110–22.
Cai Y, Nogales-Cadenas R, Zhang Q, Lin JR, Zhang W, O’Brien K, et al. Transcriptomic dynamics of breast cancer progression in the MMTV-PyMT mouse model. BMC Genomics. 2017;18:185.
Chen K, Cao W, Li J, Sprengers D, P YH, Kong X, et al. Differential sensitivities of fast- and slow-cycling cancer cells to inosine monophosphate dehydrogenase 2 inhibition by mycophenolic acid. Mol Med. 2016;21:792–802.
Perego M, Maurer M, Wang JX, Shaffer S, Muller AC, Parapatics K, et al. A slow-cycling subpopulation of melanoma cells with highly invasive properties. Oncogene. 2018;37:302–12.
Roesch A, Fukunaga-Kalabis M, Schmidt EC, Zabierowski SE, Brafford PA, Vultur A, et al. A temporarily distinct subpopulation of slow-cycling melanoma cells is required for continuous tumor growth. Cell. 2010;141:583–94.
Wu FH, Mu L, Li XL, Hu YB, Liu H, Han LT, et al. Characterization and functional analysis of a slow-cycling subpopulation in colorectal cancer enriched by cell cycle inducer combined chemotherapy. Oncotarget. 2017;8:78466–79.
Rodriguez-Enriquez S, Vital-Gonzalez PA, Flores-Rodriguez FL, Marin-Hernandez A, Ruiz-Azuara L, Moreno-Sanchez R. Control of cellular proliferation by modulation of oxidative phosphorylation in human and rodent fast-growing tumor cells. Toxicol Appl Pharm. 2006;215:208–17.
Yao CH, Wang R, Wang Y, Kung CP, Weber JD, Patti GJ. Mitochondrial fusion supports increased oxidative phosphorylation during cell proliferation. Elife. 2019;8:e41351.
Dowsett M, Nielsen TO, A’Hern R, Bartlett J, Coombes RC, Cuzick J, et al. Assessment of Ki67 in breast cancer: recommendations from the International Ki67 in Breast Cancer Working Group. J Natl Cancer Inst. 2011;103:1656–64.
Denkert C, Budczies J, von Minckwitz G, Wienert S, Loibl S, Klauschen F. Strategies for developing Ki67 as a useful biomarker in breast cancer. Breast. 2015;24:S67–72.
Yerushalmi R, Woods R, Ravdin PM, Hayes MM, Gelmon KA. Ki67 in breast cancer: prognostic and predictive potential. Lancet Oncol. 2010;11:174–83.
Cserni G, Voros A, Liepniece-Karele I, Bianchi S, Vezzosi V, Grabau D, et al. Distribution pattern of the Ki67 labelling index in breast cancer and its implications for choosing cut-off values. Breast. 2014;23:259–63.
Acknowledgements
We are grateful to V. Taylor and M. Bentires-Alj (DBM, University of Basel) for providing R26-CBW and K8-CreERT2 mice, respectively. We thank E. Panoussis, I. Galm, U. Schmieder, and T. Bürglin (DBM, University of Basel) for their technical assistance, P. Lorentz, the DBM microscopy core facility and N. Sugiyama (DBM, University of Basel) for expert support with microscopy (University of Basel), C. Beisel, E. Burcklen, K. Eschbach, and the Genomics Facility Basel (University of Basel) for RNA sequencing, and R. Ivanek for bioinformatic support. RNA-sequencing calculations were conducted at sciCORE scientific computing core facility, University of Basel.
Funding
This work was supported by the SystemsX.ch MTD project MetastasiX, the Swiss National Science Foundation, the Swiss Cancer League, and the Krebsliga Beider Basel.
Author information
Authors and Affiliations
Contributions
ST wrote the paper, designed the experiments, and conducted most of the in vivo experiments, analyses, laser capture microdissections, and validation studies, with support from FL, LvA, MH, BM, and JCM. RKRK performed all bioinformatics tools, including analysis of RNA-sequencing data, functional enrichment analysis, genome-wide predictions of regulatory sites, and gene-set enrichment analysis. BMS and NA performed microfluidic CTC isolation. TV provided human breast cancer samples and pathology expertise. GC conceived and designed the experiments, analyzed the data, and wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Tiede, S., Kalathur, R.K.R., Lüönd, F. et al. Multi-color clonal tracking reveals intra-stage proliferative heterogeneity during mammary tumor progression. Oncogene 40, 12–27 (2021). https://doi.org/10.1038/s41388-020-01508-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01508-4
- Springer Nature Limited
This article is cited by
-
Lineage plasticity enables low-ER luminal tumors to evolve and gain basal-like traits
Breast Cancer Research (2023)
-
Breast cancers as ecosystems: a metabolic perspective
Cellular and Molecular Life Sciences (2023)
-
Mastering the use of cellular barcoding to explore cancer heterogeneity
Nature Reviews Cancer (2022)
-
Spatial omics and multiplexed imaging to explore cancer biology
Nature Methods (2021)