Abstract
Cancer cells associated with radioresistance are likely to give rise to local recurrence and distant metastatic relapse. However, it remains unclear whether specific miRNAs have direct roles in radioresistance and/or prognosis. In this study, we find that miR-339-5p promotes radiosensitivity, and is downregulated in radioresistant subpopulations of esophageal cancer cells. Notably, miR-339-5p was selectively secreted into blood via exosomes, and that higher serum miR-339-5p levels were positively associated with radiotherapy sensitivity and good survival. Moreover, miR-339-5p expression was downregulated in the T3/T4 stage compared with T1/T2 stage in esophageal squamous cell carcinoma (ESCC) patients (P = 0.04), and low miR-339-5p expression in tissue was significantly associated with poor overall survival (P = 0.036) and disease-free survival (P = 0.037). Overexpression of miR-339-5p enhanced radiosensitivity in vitro and in vivo. Mechanistically, miR-339-5p enhances radiosensitivity by targeting Cdc25A, and is transcriptionally regulated by Runx3. Correlations were observed between miR-339-5p levels and Cdc25A/Runx3 levels in tissue samples. Intriguingly, combined analysis of miR-339-5p expression with Runx3 increased the separation of the survival curves obtained for either gene alone in the TCGA datasets (P = 0.009). Overall, exosome-derived miR-339-5p mediates radiosensitivity through downregulation of Cdc25A, and predicts pathological response to preoperative radiotherapy in locally advanced ESCC, suggesting it could be a promising non-invasive biomarker for facilitating personalized treatments.
Similar content being viewed by others

Introduction
Esophageal cancer remains one of the most virulent malignancies with a 5-year overall survival (OS) rate of under 20% [1, 2]. Surgery is a standard treatment for patients with resectable tumors, but surgery alone is inadequate therapy for patients with T3 & T4 or node-positive diseases [3]. Neoadjuvant radiotherapy or chemoradiation therapy (CRT) has emerged as a promising strategy for locally advanced esophageal squamous cell carcinoma (ESCC) [4, 5]. Approximately 25% of patients are highly sensitive to chemoradiation, resulting in pathologic complete response (pCR). pCR was achieved in 38.7% of patients with stage II and 20% of patients with stage III ESCC [6, 7]. However, some patients do not derive clinical benefit from it because their tumors are constitutively insensitive to radiotherapy. Without a prior knowledge of whether patients are sensitized to radiotherapy, there is inevitable overtreatment with agents associated with toxic side effects. Therefore, it would be advantageous to obtain more information on radiotherapy response before treatment initiation, which tends to stratify ESCC into different response categories, eventually optimizing a response-based therapeutic option.
MicroRNAs (miRNAs) bind to the 3′-untranslated region of particular (target) mRNAs, which leads to translational repression or reduced stability of the specific mRNAs [8]. Of note, miRNAs have previously been shown critical in genome integrity and stability via the regulation of DNA damage response (DDR) [9, 10]. Restoration of tumor-suppressive miRNAs reduces tumorigenic properties including cell growth and invasion by promoting apoptosis and sensitivity to therapy [11]. MiR-205 inhibits DNA damage repair and radiosensitizes cancer cells by targeting ZEB1 and Ubc13 in breast cancer [12]. Thus, the delivery of miR-205, in combination with radiotherapy may represent a new strategy for cancer treatment. It was recently reported that several miRNAs were associated with therapeutic response and clinic outcomes in ESCC. Both miR-192 and miR-194 expression were significantly correlated with pathologic response in locally advanced ESCC [13]. These two miRNAs were transcriptionally regulated by p53 and induced cell-cycle arrest by targeting a number of transcripts that regulate the G1/S and G2/M checkpoints [14, 15].
However, whether specific miRNAs can regulate radioresisitance and could be used as tumor radiosensitizers in ESCC remains unclear. In this study, we find that miR-339-5p promotes radiosensitivity through downregulation of Cdc25A, and is transcriptionally regulated by Runx3. Notably, miR-339-5p was selectively secreted into blood via exosomes, and that higher serum miR-339-5p levels were positively associated with radiotherapy sensitivity. MiR-339-5p expression in tumor tissue and serum significantly associated with clinical outcomes. Our findings suggest that exosome-derived miR-339-5p can serve as a non-invasive biomarker for predicting response to radiotherapy and prognosis in ESCC.
Results
Differential miRNA expression profiling in radioresistant cancer cells
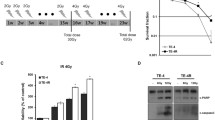
To establish a radioresistant cell model, we treated human ESCC cell lines (KYSE30, KYSE70 and KYSE180) by applying continuous 2 Gy fractionated irradiations to 60 Gy, and finally obtained a radioresistant subpopulation (designated as KYSE30R, KYSE70R and KYSE180R) (Fig. 1a). The clonogenic survival curves for the two pairs of cell lines at different doses are shown in Fig. 1b. A significant increase of radioresistance in KYSE30R, KYSE70R and KYSE180R cells was observed compared to the parental KYSE30, KYSE70 and KYSE180 cells, respectively.
Radioresistant cell model was established and the differentially expressed miRNAs were identified using miRNA microarray. a The schematic representation of the generation of radioresistance sublines (KYSE30R, KYSE70R and KYSE180R) from the parental cells (KYSE30, KYSE70 and KYSE180, respectively). b Clonogenic survival assays from the radioresistant and parental cells. n = 3 wells per group. c The radioresistant and parental cells were treated with or without 6 Gy of X-IR, and allowed to recover for 72 h. The PARP and Caspase-3 protein levels were examined by Western blot. d qRT-PCR assay validated miRNA microarray findings that miR-181a-2-3p, miR-193b-5p, miR-339-5p and miR-491-5p in the radioresistant cells (KYSE30R) and parental cells (KYSE30). U6 small nuclear RNA was used as an internal control. n = 3 samples per group. e KYSE30 cells were transiently transfected with 4 candidate miRNA mimics and negative control (NC) respectively. 24 h after transfection, cells were exposed to 6 Gy of X-IR, and allowed to recover for the indicated time. The PARP and Caspase-3 protein levels were detected by Western blot. The data (b, d) are the mean of biological replicates a representative experiment, and error bars indicate S.E.M. Statistical significance was determined by a two-tailed, unpaired Student’s t test. The experiments were repeated three times. *P < 0.05
Radiation is known to cause DNA double-strand breaks (DSBs), which lead to the formation of γH2AX foci, and γH2AX is required for DNA damage signaling and DNA repair [16, 17]. At 72 h after X-irradiation (X-IR), γH2AX foci remained in the parental cells, but reduced in the radioresistant cells (Supplementary Fig. S1a), suggesting that the radioresistant cells have an enhanced ability to clear DNA breaks. Radiation affects cells by generating DSBs resulting in a cellular response including DNA replication, DNA repair, and programmed cell death. We observed that the radioresistant cells decreased the activity of Caspase-3 and PARP cleavage, and inhibited cellular apoptosis via DNA damage response in vitro (Fig. 1c, Supplementary Fig. S1b, c).
To investigate miRNAs that regulate radiosensitivity, we performed a human miRNA expression array analysis to identify miRNAs upregulated or downregulated in the radioresistant cells, and found that a small subset of miRNAs were differentially expressed (Fold change > 2 or < 0.5) between the radioresistant cells and their parental cells (GSE124784). To further confirm the results, we selected 4 candidate miRNAs according to the fold change and function of miRNAs, and detected their expression levels in KYSE30 and KYSE30R cells using qRT-PCR (Fig. 1d). To further investigate whether the differentially expressed miRNAs contribute to cellular apoptosis, we transiently transfected these 4 candidate miRNAs mimics into KYSE30 cells. After 24 h, the cells were subsequently exposed to 6 Gy of X-IR. As shown in Fig. 1e, overexpression of these miRNAs increased the activity of PARP and caspase-3 in DNA damage response. Among all the interrogated miRNAs, miR-339-5p enhanced the activation of cleaved PARP and Caspase-3 to a much greater extent. Jansson et al. showed that miR-339-5p, by directly targeting MDM2, regulates p53 and impacts p53-governed cellular response such as proliferation arrest and senescence in several types of cancer cells [18]. Thereby, we selected miR-339-5p for further investigation.
MiR-339-5p exhibits tumor-suppressive effects and mediates radiosensitivity in ESCC
To determine the clinical relevance of miR-339-5p in ESCC, we examined the expression levels of miR-339-5p using in situ hybridization (ISH) in ESCC samples (Fig. 2a), but did not find any significant differences between tumor and adjacent normal mucosa tissues (Fig. 2b). However, we observed miR-339-5p downregulation in T3/T4 stage compared with T1/T2 stage (P = 0.04, Fig. 2c). MiR-339-5p expression was associated with esophageal cancer recurrence (P = 0.039, Fig. 2d). But, there was no significant association between miR-339-5p expression and the metastasis status of esophageal cancer patients (P = 0.890, Supplementary Fig. S2). We further assessed whether miR-339-5p mediated radiosensitivity in vivo. As shown in Fig. 2e–g, overexpression of miR-339-5p and PLVX-control grew similarly without X-IR treatment. It significantly decreased tumor growth compared with the control on X-IR treatment. Together, these findings indicate that miR-339-5p exhibits tumor-suppressive effects, and overcomes resistance of esophageal cancer to radiation therapy.
Expression status of miR-339-5p in human clinical specimens and ectopic expression of miR-339-5p enhanced radiosensitivity. a In situ hybridization (ISH) was used to detect mature miR-339-5p in cancer (C) and adjacent normal tissues (N) using LNA-miRNA probes. Positive control (U6 snRNA) and negative control (scrambled miRNA) were included in each hybridization reaction. b MiR-339-5p levels in 25 paired tumor and adjacent normal mucosa was analyzed using paired sample t tests. c MiR-339-5p levels in different TNM stages of ESCC (n = 180). d Association of miR-339-5p expression with recurrence in ESCC patients (n = 129). e KYSE30 cells (2 × 106/mouse) stably infected with a miR-339-5p overexpression vector or a control lentiviral vector, and were subcutaneously injected into the right leg of 5-week-old BALB/C nude mice. When tumors reached approximately 5 mm in diameter, the tumors were irradiated with a single 12 Gy dose of X-IR (dashed arrow indicates the time of irradiation). Each group was composed of 5 mice. Tumor sizes were measured every 3 days. Data are presented as tumor growth curves. f The tumor was shown after X-IR treatment. g Tumor was harvested and weighed, and analyzed by a two-tailed, unpaired Student’s t test
MiR-339-5p mediates radiosensitivity
To further investigate the function of miR-339-5p in radiosensitivity, we first examined miR-339-5p expression in ESCC cell lines (Fig. 3a). Indeed, cells with lower levels of miR-339-5p exhibited higher clonogenic survival after X-IR (Fig. 3b). Next, we performed gain-of-function and loss-of-function analysis with miR-339-5p, and found that miR-339-5p overexpression sensitized KYSE30 cells to radiation. Conversely, miR-339-5p inhibition conferred radioresistance (Fig. 3c). MiR-339-5p expression does not impair cell viability as detected by the Cell Counting Kit-8 (Supplementary Fig. S3a). We attempted to understand how miR-339-5p promotes radiosensitivity and showed that overexpression of miR-339-5p in KYSE30 cells drastically increased the activation of cleaved PARP and Caspase-3 in DNA damage response (Fig. 3d). Similar effects were also observed in KYSE180 cells (Supplementary Fig. S3b). Moreover, KYSE30 cells were also transiently transfected with miR-339-5p mimic or negative control and then treated with or without X-IR. MiR-339-5p also increased the activity of Caspase-3 and PARP cleavage (Fig. 3e). We observed more γH2AX foci in miR-339-5p-overexpressing cells than in control cells at 72 h after exposure to 6 Gy (Fig. 3f–h), suggesting that miR-339-5p might be involved in DNA damage response. Similar behaviours were observed in KYSE180 cells (Supplementary Fig. S3c–f). Our results indicate that miR-339-5p promotes radiosensitivity through DNA damage-induced cellular apoptosis.
MiR-339-5p promotes radiosensitivity through DNA damage-induced apoptosis. a MiR-339-5p expression in ESCC cell lines and human normal esophageal epithelial cells (Het-1A) was examined by qRT-PCR. b Radiation clonogenic survival assays was performed in ESCC cell lines. c KYSE30 cells were infected with PLVX-miR-339-5p, PLVX (left panel) and miRzip-miR-339-5p or GFP-control (right panel) lentivirus, and screened by G418 at 72 h after infection. Clonogenic survival assays were performed on stable miR-339-5p overexpression or inhibition cells after increasing doses of X-IR. d MiR-339-5p overexpressing or inhibiting KYSE30 cells were treated with a single dose of 6 Gy of X-IR. After 72 h, PARP and Caspase-3 were detected by Western blot. e KYSE30 cells were transiently transfected with miR-339-5p mimics or negative control (NC), after 24 h, and treated with 6 Gy of X-IR. PARP and Caspase-3 were evaluated by Western blot. f KYSE30 cells were exposed to 6 Gy of X-IR. Then, 72 h after recovery, γH2AX was detected by western blot. g γH2AX and DAPI staining of KYSE30-miR-339-5p and KYSE30-PLVX control cells at 72 h after 6 Gy of X-IR. h γH2AX foci was count, and analyzed by a two-tailed, unpaired Student’s t test. The picture is representative of one experiment in (a, b, d–f). Statistical significance was performed on three independent experiments. *P < 0.05
MiR-339-5p directly targets Cdc25A
miRNAs are known to induce degradation of specific target mRNA molecules or inhibit their translation. To identify putative targets of miR-339-5p responsible for radiosensitivity, we analyzed the target of miR-339-5p using TargetScan and PicTar. Interestingly, among the targets, Cdc25A phosphatase inactivates cyclin-dependent kinase 2 (Cdk2) which is needed for DNA synthesis, but is degraded in response to DNA damage or stalled replication [19]. One conserved miR-339-5p recognition site was found in the Cdc25A 3′-UTR, and mutation sites were colored red (Fig. 4a). Moreover, Cdc25A mRNA and protein expression were increased in KYSE30R and KYSE180R cells (Supplementary Fig. S4a). On the contrary, miR-339-5p was downregulated in the radioresistant cells. Thus, we speculate miR-339-5p might be involved in the regulation of Cdc25A expression.
Cdc25A is a direct target of miR-339-5p, and Cdc25A protein expression and miR-339-5p expression was inversely correlated in ESCC. a The 3′ UTR sequence of Cdc25A contains miR-339-5p binding site. The mutant sites were colored red. b KYSE30 and KYSE180 cells were transfected with wild- or mutant-Cdc25A 3′-UTR, miR-339-5p (50 nM), negative control (NC, 50 nM) and pRL-TK as indicated. 24 h after transfection, dual luciferase activity was recorded. c KYSE30 and KYSE180 cells were stably infected with miR-339-5p (overexpression) or miRzip-339 (inhibition) lentivirus, and Cdc25A expression was determined by Western blot. d Cdc25A expression was detected by qRT-PCR in 30 pairs of tumor (T) and adjacent normal tissues (N). e Cdc25A expression was detected in 18 pairs of tumor and adjacent normal tissues by Western blot (ten representative pairs). f Cdc25A protein expression was quantified using ImageJ and normalized against β-actin. The correlation between miR-339-5p expression and Cdc25A protein expression in 18 pairs of ESCC tumor and adjacent normal tissues was analyzed using SPSS17.0. Statistical analysis was performed on -ΔCt of triplicates of one experiment. g KYSE30 cells were infected with miR-339-5p, negative control, Cdc25A (open reading frame lacking 3′-UTR) and pCMV3 as indicated. 24 h later, they were treated with 6 Gy of X-IR. PARP, Caspase-3 and Cdc25A were evaluated by Western blot. h Cellular apoptosis was detected by Annexin V-FITC staining. i Clonogenic survival assays for the indicated cells were performed. j Cdc25A expression were examined by Western blot in miR-339-5p overexpressed or inhibited KYSE30 cells after treatment with 6 Gy of X-IR. The data (b, h, i) are the mean of biological replicates a representative experiment, and error bars indicate S.E.M Statistical significance was determined by a two-tailed, unpaired Student’s t test. The experiments were repeated three times. *P < 0.05
To determine whether Cdc25A is a direct target of miR-339-5p, we used the dual-luciferase reporter assay system. The activity of luciferase reporter fused to a wild-type Cdc25A 3′-UTR, but not that of the mutant Cdc25A 3′-UTR, was significantly reduced by miR-339-5p in KYSE30 and KYSE180 cells (Fig. 4b), thereby confirming that miR-339-5p can directly bind to the 3′-UTR of Cdc25A, and repress its expression (Fig. 4c). Furthermore, we observed a significant increase in Cdc25A in ESCC tumor tissues compared with adjacent normal tissues at both RNA and protein levels (Fig. 4d, e, Supplementary Fig. 4b). Cdc25A expression was significantly upregulated in the ESCC tumors in the Gene Expression Omnibus (GEO) database GSE23400 (P = 5.9E-8) and GSE20347 (P = 4.88E-5; Supplementary Fig. S4c). Notably, miR-339-5p expression was inversely correlated with Cdc25A protein expression in tumor and adjacent normal mucosa tissues (R = -0.363, P = 0.023; Fig. 4f). These results suggested that miR-339-5p directly targets Cdc25A, and inversely correlated with Cdc25A in ESCC.
MiR-339-5p promotes radiosensitivity by targeting Cdc25A
To determine whether miR-339-5p promotes apoptosis by directly repressing Cdc25A, we found that Cdc25A overexpression inhibited DNA damage-induced apoptosis (Supplementary Fig. S4d–f). Additionally, we ectopically overexpressed Cdc25A using an expression vector containing the entire Cdc25A coding sequence but lacking its 3′-UTR in miR-339-5p-overexpressing KYSE30 cells. Cdc25A overexpression partially rescued miR-339-5p-induced cellular apoptosis and radiosensitivity in DNA damage response (Fig. 4g–i). Moreover, overexpression of miR-339-5p partly inhibited Cdc25A protein expression in DNA damage response. Conversely, inhibition of miR-339-5p could increase Cdc25A protein expression. MiR-339-5p overexpression could increase p-Chk2 expression in DNA damage response. In contrast, a decrease in p-Chk2 was observed in miRzip-339-5p cells, suggesting that miR-339-5p may be involved in the ATM-Chk2-Cdc25A pathways (Fig. 4j, Supplementary Fig. S4g). Moreover, miR-339-5p still could regulate Cdc25A expression after Chk2 inhibition in KYSE30 cells (Supplementary Fig. S4h). Collectively, these results suggest that Cdc25A is a functional target of miR-339-5p, and that miR-339-5p promotes radiosensitivity in part by targeting Cdc25A.
Runx3 activates miR-339-5p transcription
DNA hypermethylation in miRNA CpG islands contributes to transcriptional downregulation in human tumors [20, 21]. In this study, we found miR-339-5p was downregulated in the radioresistant cells. However, whether DNA methylation regulates miR-339-5p expression remains unclear. We found that human miR-339-5p on chromosome 7p22.3 is embedded in a CpG island using the UCSC database. Analysis of the TCGA esophageal cancer datasets revealed that 9 CpG sites were consistently methylated in miR-339-5p promoter region, which illustrates that expression of miR-339-5p may not depend on the extent of DNA methylation (Supplementary Fig. S5a). Moreover, we observed that miR-339-5p expression was slightly increased in ESCC cell lines treated with the DNA methyltransferases inhibitor 5-Aza-2-dC (Supplementary Fig. S5b). Thereby, we further explore the transcriptional regulation mechanism of miR-339-5p.
Shen et al. indicated that a CpG island of the promoter region of miR-339-5p (–155 to + 187 bp, containing 27 CpG sites), and the epigenetic regulation might be a potential mechanism for miR-339-5p downregulation [22]. To further determine the miR-339-5p promoter region, we constructed a reporter plasmid by inserting a proximately 1000 bp promoter region fragment into pGL3-basic. This sequence significantly increased luciferase activity compared with the empty vector (Fig. 5a), suggesting that this region could be miR-339-5p promoter. By using bioinformatics analysis and the UCSC database, we verified the involvement of Runx3 in the transcriptional regulation of miR-339-5p. We also examined the genomic sequences flanking for the human miR-339-5p stem-loop (pre-miR-339-5p) and found five DNA binding sites for Runx3 located at upstream of miR-339-5p (Supplementary Fig S5c).
Runx3 activates miR-339-5p transcription. a Luciferase reporter assays were performed to identify the miR-339-5p promoter. b, c Runx3 expression was detected in KYSE30 and KYSE180 cells transiently transfected with pCMV3-Runx3 by Western blot (b) and qRT-PCR (c). d Pre-miR-339-5p (left panel) and mature miR-339-5p expression (right panel) was examined using qRT-PCR in Runx3 overexpressing cells. e ChIP assays were performed in KYSE30 and KYSE180 cells, and examined by PCR with primers specific for the Runx3 DNA binding sites. f KYSE30 cells were transfected with pGL3-Luc, miR-339-5p-wt, miR-339-5p-mt1-4, pCMV3 and Runx3 as indicated. The dual luciferase activity was detected after 24 h. g Runx3 expression in 30 paired tumor and adjacent normal tissues was detected by qRT-PCR. h Runx3 expression was determined in 20 paired tumor tissues and adjacent normal tissues by Western blot (ten representative pairs). i Pearson’s correlation analysis was used to determine correlation between miR-339-5p expression and Runx3 expression in ESCC specimens. j Correlation between Runx3 and Cdc25A protein expression was analyzed using SPSS17.0. The data (a, c–f are the mean of biological replicates a representative experiment, and error bars indicate S.E.M Statistical significance was determined by a two-tailed, unpaired Student’s t test. The experiments were repeated three times. *P < 0.05
To determine whether Runx3 is indeed a transcriptional activator of miR-339-5p, Runx3 was transfected in KYSE30 and KYSE180 cells, and examined by qRT-PCR and Western blot. Runx3 overexpression increased both pre-miR-339-5p and miR-339-5p expression levels (Fig. 5b–d). Conversely, Knockdown of Runx3 decreased pre-miR-339-5p and miR-339-5p expression levels in both cell lines Supplementary Fig. S5d, e). To verify the direct binding of Runx3 to the miR-339-5p promoter, we designed PCR amplicons to test for the presence of the two putative binding sites by chromatin immunoprecipitation (ChIP). The results revealed that the endogenous Runx3 protein bound to these sites upstream of the pre-miR-339-5p transcription start site (Fig. 5e). Furthermore, luciferase reporter assays demonstrated that overexpression of Runx3 significantly enhanced the activity of the putative miR-339-5p promoter. The mutations of DNA binding sites of Runx3 (-193 - -198bp, -201 - -207bp) dramatically suppressed the induction of transcriptional activity of miR-339-5p promoter, suggesting the two DNA binding sites might be play a vital role in the transcriptional regulation of miR-339-5p (Fig. 5f). Our results indicated that Runx3 was involved in the transcriptional regulation of miR-339-5p. Moreover, a previous study revealed that frequent silencing of Runx3 in ESCC is associated with radioresistance and poor prognosis [23]. Consistent with this previous study, Runx3 was downregulated in radioresistant cells (KYSE30R) compared to parental cells (Supplementary Fig. S6a). Runx3 was decreased in a time-dependent manner in KYSE30 and KYSE180 cells (Supplementary Fig. S6b, c).
To investigate the correlation between Runx3 and miR-339-5p in clinical samples, we examined the Runx3 expression by qRT-PCR and Western blot in 30 paired tumor and matched normal tissue samples, and observed a significant downregulation of Runx3 in the tumor samples at both mRNA and protein levels (Fig. 5g, h). Moreover, we found a positive correlation between miR-339-5p and Runx3 expression in ESCC (R = 0.343 P = 0.04; Fig. 5i). Notably, the inverse correlation between Runx3 and Cdc25A expression was also found in ESCC specimens (R = −0.359, P = 0.031; Fig. 5j). These results suggest that Runx3 is involved in the transcriptional regulation of miR-339-5p.
MiR-339-5p expression is associated with survival in ESCC
MiR-339-5p inhibited cell metastatic potential and mediated radiosensitivity, thus we further explore the correlation between miR-339-5p expression and survival of ESCC patients. We found that low miR-339-5p expression in tissue was significantly associated with poor overall survival (n = 138, P = 0.036; Fig. 6a) and disease-free survival (n = 122, P = 0.037; Fig. 6b). To validate our results, we checked the expression of miR-339-5p and Runx3 in The Cancer Genome Atlas (TCGA) data set containing 94 esophageal cancer patients with stages II/III, and found that patients with low miR-339-5p expression had shorter overall survival (OS) compared with those with high miR-339-5p levels (n = 94, P = 0.037; Fig. 6c); Moreover, the Runx3 low expression group had a significantly shorter OS compared with the Runx3 high expression group (n = 94, P = 0.019; Fig. 6d). Combined analysis of miR-339-5p with Runx3 expression increased the separation of the survival curves obtained by either gene alone, and patients with miR-339-5p (low)/Runx3 (low) had a significantly shorter survival than those with miR-339-5p (high)/Runx3 (high) (n = 49, P = 0.009; Fig. 6e). All these data suggest underlying Runx3 expression for offering a more accurate prognosis and even predicting recurrence.
Kaplan–Meier survival analysis for miR-339-5p expression in tissue specimens from ESCC patients. a Overall survival of patients with ESCC based on tissue miR-339-5p expression using Kaplan–Meier curves (n = 138). b Disease-free survival of patients with ESCC based on miR-339-5p expression (n = 122). c Association of miR-339-5p and the overall survival in the TCGA datasets (n = 94). The group was defined according to the mean of miR-339-5p expression. d Association of Runx3 expression and the overall survival in the TCGA datasets (n = 94). The group was defined according to the mean of Runx3 expression. e Patient overall survival was analyzed according to the expression of miR-339-5p and Runx3 in the TCGA datasets (n = 49). Cum survival: cumulative survival
MiR-339-5p is found in exosomes
Our findings suggested that miR-339-5p is associated with patient survival, we speculated whether miR-339-5p can be secreted in the circulation. Villarroya-Beltri et al. demonstrated that sumoylated hnRNPA2B1 controls miRNAs sorting into exosomes through binding to specific motifs [24]. We found that miR-339-5p contains Exomotif sequence: GGAG (Fig. 7a). To determine whether miR-339-5p is a secretory miRNA, we examined the presence of miR-339-5p in culture medium. Exosomes were isolated from conditioned medium from three different ESCC cell lines using ultracentrifugation and examined via the Western blot detection of several exosome markers, including heat shock protein70 (HSP70), CD81, and TSG101. Both Western blot and transmission electron microscopy (TEM) confirmed the isolation of exosomes (Supplementary Fig. S7a, b). Using qRT–PCR, we found that miR-339-5p expression was decreased in exosome-depleted supernatants (Fig. 7b). Similarly, miR-339-5p was contained within exosomes derived from cells, and that its expression was directly proportional to culture duration and cell number (Fig. 7c, d). Furthermore, we blocked exosome formation by treating KYSE30 and KYSE180 cells with GW4869, a drug that hinders exosome biogenesis by blocking neutral sphingomyelinase 2 (nSMase2) and miRNA content in exosomes [25]. GW4869 blocked exosome secretion (Fig. 7e), and statistically significantly reduced miR-339-5p levels (Fig. 7f). Collectively, our data suggest that miR-339-5p is contained within exosomes.
miR-339-5p is contained in exosomes. a The predicted Exomotif sequences for miR-339-5p. b MiR-339-5p was detected in culture medium or exosome-depleted medium of KYSE30 cells using qRT-PCR. c, d MiR-339-5p expression was detected in exosomes from culture medium of three different ESCC cell lines (KYSE30, KYSE180 and KYSE510) by ultracentrifugation for the indicated time periods (c) and cell number (d). e KYSE180 cells were treated with 10 μM GW4869 or DMSO (as a control). After 72 h, culture medium was collected. Exosomes were isolated by ultracentrifugation (100,000 g), and detected by Western blot using exosome markers. f MiR-339-5p expression was detected in using qRT-PCR exosomes from KYSE30 cells culture medium with or without GW4869 treatment. The data (b–d, f) are the mean of biological replicates, and error bars indicate S.E. Statistical significance was determined by a two-tailed, unpaired Student’s t test. The experiments were repeated three times. *P < 0.05
Serum miR-339-5p as a non-invasive biomarker for pathological response to preoperative radiotherapy
To further determine whether miR-339-5p can be secreted into blood by exosomes, we obtained serum from 5 ESCC patients, purified the exosomes, and performed Western blot to confirm the presence of the exosomes (Supplementary Fig. S7c). As shown in Fig. 8a, miR-339-5p was detected in exosomes isolated from serum of ESCC patients. We further investigated the correlation between serum miR-339-5p expression and clinical pathological parameters, but did not observe any statistically significant differences in the serum miR-339-5p expression between ESCC patients and healthy control subjects (P = 0.874; Fig. 8b). Of note, we observed that ESCC patients that were non-responsive to radiotherapy had significantly lower mean serum miR-339-5p levels (P = 6.82E-5; Fig. 8c). Additionally, patients with low miR-339-5p expression had significantly shorter survival than those with higher miR-339-5p expression (P = 0.012; Fig. 8d). These results suggest that serum miR-339-5p is a promising non-invasive biomarker for predicating the response to radiotherapy and prognosis in ESCC.
Serum miR-339-5p serves as a non-invasive biomarker for pathological response to preoperative radiotherapy in ESCC. a Exosomes were isolated from 5 serums of ESCC patients by ultracentrifugation. MiR-339-5p expression was examined in serum, exosomes and exosome-depleted serum by qRT-PCR. Cel-miR-39 was used as an internal control. Each reaction included three technical replicates, and error bars indicate s.d. b Serum miR-339-5p level in 42 healthy control subjects and 41 ESCC patients was detected by qRT-PCR. c Comparison of miR-339-5p expression levels in 48 serum samples between non-response group and response group. d Kaplan–Meier analysis of overall survival of 59 patients with ESCC stratified according to admission serum miR-339-5p level
Discussion
In cancer cells, in response to DNA damage, cell cycle checkpoints and DNA repair pathways become activated. These mechanisms may ultimately promote radioresistance and cell survival [26, 27]. Many groups are dedicated to identify the agents that target DNA damage response pathways and mediate radioresistance; however, it still remains unclear whether these agents have direct roles in sensitivity or offer prognostic value to clinicians. This study demonstrates the potential role of exosome-derived miR-339-5p as a non-invasive biomarker for determining patient prognosis and predicting response to radiotherapy.
By establishing a radioresistant cell model, we found a set of significantly differentially expressed miRNAs associated with radioresistance (or radiosensitivity), and some of these miRNAs have been previously studied. Among them, miR-339-5p, which is known to be a putative tumor suppressor that modulates the expression of multiple cancer-related target genes such as MDM2, BACE1 and ICAM1, has been shown to inhibit cellular proliferation, migration and invasion in various human tumors. [18, 28,29,30] MiR-339-5p-5p expression was inversely associated with metastasis to lymph nodes, clinical stages and survival in breast cancer [31]. Our findings showed that miR-339-5p expression levels in ESCC tissue samples from patients with T1/T2 stage were significantly higher than those from patients with T3/T4 stage, and its expression was inversely correlated with recurrence. Likewise, analysis of the TCGA datasets revealed that miR-339-5p expression was positively correlated with survival. Our findings demonstrate that miR-339-5p may be a predicator for prognosis and recurrence in ESCC.
Accumulating data in recent years have convincingly demonstrated that circulating miRNAs reflect physiological and pathological alterations in patients, and may be used as promising biomarkers for the non-invasive detection of cancer at early stages [32, 33]. Circulating exosomic miRNAs, which are similar to those, originating from cancer cells, were found to be associated with therapy resistance or progression. Challagundla et al. indicated that a unique role for exosomic miR-21 and miR-155 in the cross-talk between neuroblastoma and human monocytes in chemoresistance through a novel miR-21/TLR8-NF-кB/miR-155/TERF1 signaling pathway [34]. In ESCC, exosomic miRNAs also showed potential as biomarkers for diagnoses. MiR-21 and miR-1246 could be secreted by cancer cells to exosomes. These two miRNAs were upregulated in serum and positively correlated with tumor progression and patient survival [35, 36]. However, with regards to ESCC, only a few circulating miRNAs have been reported to potentially play a role in radioresistance. A previous study illustrated that miRNAs are not randomly loaded into exosomes. Sumoylated hnRNPA2B1 specifically binds to miRNAs containing the ‘shuttling’ motif GGAG, leading to their upload into exosomes [24]. In this study, we systematically investigated miR-339-5p as a secretory miRNA. Firstly, miR-339-5p contains the Exomotif sequence GGAG, and secreted into the culture medium by exosome isolation, thus establishing its secretory potential. Subsequently, miR-339-5p expression level in exosomes was increased in a cell number- and time-dependent manner. To the best of knowledge, our results, for the first time, demonstrate that serum miR-339-5p was associated with overall survival and predicted the response to radiotherapy in locally advanced ESCC. Recent studies indicated that exosomes, carrying cancer cell-derived miRNAs, act as natural nano-sized membranous vesicles, and provide benefit of mediating gene delivery without inducing adverse immune reactions and pro-inflammatory response [37,38,39]. Thereby, future studies aim to investigate whether these miRNAs examined in our study act as potential therapeutic targets and to also understand if exosome-derived miRNAs could be a potential biomarker panel for predicting therapeutic response in ESCC.
Radiation affects cells by generating DSBs resulting in a cellular response including DNA replication, DNA repair, and programmed cell death [26]. Radiation-induced programmed cell death is a major form of death in tumors derived from lymphoid, hematopoietic and germ cells [40]. However, epithelial tumors show wide resistance to apoptosis induced by ionizing radiation. In our study, we found that miR-339-5p overexpression or inhibition does not impact esophageal cancer cell proliferation. However, miR-339-5p overexpression markedly enhances DNA damage-induced apoptosis in esophageal cancer cells. Likewise, miR-339-5p suppression inhibits its effect. Thus, our results demonstrate that miR-339-5p promotes radiosensitivity mainly through DNA damage-induced cellular apoptosis in ESCC cells.
Runx3 acts as a tumor suppressor and was associated with prognosis and radio- or chemosensitivity in several cancers. Frequent silencing of Runx3 was associated with radioresistance and poor prognosis in ESCC [23]. Runx3 facilitates p53 phosphorylation through the ATM/ATR pathway and p53 acetylation by p300 in DNA damage response [41, 42]. Moreover, the expression of miR-339-5p was positively correlated with Runx3 in ESCC specimens. Notably, our analysis of the TCGA datasets revealed a combination of miR-339-5p with Runx3 expression increased the separation of the survival curves. Our results demonstrate that Runx3 expression, in combination with miR-339-5p, may serve as potential prognostic markers in ESCC.
In conclusion, our findings identify that miR-339-5p as a radiosensitizing miRNA, can be released into circulation, and serves as a promising non-invasive biomarker for facilitating personalized treatments.
Materials and methods
Establishment of radioresistant cell model
The ESCC cell lines (KYSE series) were kindly provided by Dr. Yutaka Shimada [43]. Cells were cultured in RPMI1640 medium supplemented with 10% fetal bovine serum and 1% penicillin and streptomycin. The radioresistant cells were established according the following scheme. Cells (KYSE30, KYSE70 and KYSE180) were grown to 50% confluence, treated with 2 Gy of X-IR, and then incubated. Upon reaching 90% confluence, the cells were trypsinized and then incubated. When the cells reached 50% confluence, they were irradiated again (second fraction), and then this process repeated a total of 30 times to reach 60 Gy. The parental cells were subjected to identical trypsinization and culture conditions but were not irradiated. For all assays, there was at least a 4 or 5 months interval between the last 2 Gy fractionated irradiation. The irradiated cells were required to recovery for 2–3 weeks after the last irradiation, and used to functional assay.
Clinical specimens
Serum from patients with locally advanced ESCC treated with preoperative radiotherapy followed by surgery from 2013 to 2016 (n = 48) or radiotherapy alone (n = 100) and 42 healthy individuals from 2006 to 2010 at the Cancer Hospital Chinese Academy of Medical Sciences. 30 pairs of primary ESCC and adjacent normal tissue specimens were obtained from patients who were treated with surgical resection alone in 2014 at the Cancer Hospital Chinese Academy of Medical Sciences. All of patients never received any chemotherapy before radiotherapy. The clinical characteristics of the patients are summarized in Supplementary Table S1. This study was approved by the Institutional Review Board of the Cancer Hospital Chinese Academy of Medical Sciences.
Tissue microarray and in situ hybridization
Tissue microarray containing 180 tumor samples (duplicate 1.5 mm tissue cores for each ESCC) were constructed by Zhejiang Cancer Hospital (Supplementary Table S2). For in situ hybridization analysis, the tissue microarrays were hybridized with a miR-339-5p probe (locked nucleic acid (LNA)-modified and 5′-DIG-labelled and 3′-DIG-labelled oligonucleotide; Exiqon, Woburn, Massachusetts, USA) as described previously [44]. Positive (U6 snRNA) and negative controls (scrambled miRNA) were included in the hybridization reaction. After hybridization, the sections were scanned and imaged by a single investigator who was not informed of the clinical characteristics. The integral intensity value was measured with Aperio’s ImageScope software (Aperio, Vista, CA, USA).
Transfection and western blot
Transfections were performed using HiperFect (Qiagen, Valencia, CA) for miRNA mimics or Runx3 siRNAs (#E-012666-00-0005, Dharmacon, USA), and Lipofectamine 2000 (Thermo Fisher Scientific, Australia) for plasmids according to the manufacturers’ recommendations. Western blot was performed according to the standard protocol. β-actin was used as loading control. The intensity of protein bands was quantified by ImageJ. The antibodies were listed in Supplementary Table S3.
Radiation clonogenic survival assay
Cells were seeded in triplicate in six-well plates. After 24 h, the cells were exposed to different doses of X-IR (2, 4 or 6 Gy), followed by incubation at 37 °C for 9–12 days until the development of visible colonies. Colonies were stained with crystal violet staining solution and counted. The survival fraction was calculated as: (number of colonies / number of cells plated) irradiated / (number of colonies / number of cells plated) non-irradiated.
Exosomes isolation and transmission electron microscopy
A total of 6 × 106 Cells were grown in 10% exosome-depleted FBS medium. After 72 h, 30 ml supernatant was collected, and subsequently subjected to centrifugation at 3,500 g for 30 min. This resulting supernatant was then filtered using 0.22 μm filters, and ultracentrifuged at 100,000 × g for 2 h. Exosomes were wicked off to create a thin layer before addition of a thin layer of 2% urany acetate in water. Grids were allowed to dry overnight, and transmission electron microscopy (JEM-1400, 80KV) performed the next day.
Luciferase reporter assay and chromatin immunoprecipitation
A Muta site Directed Mutagenesis kit was used to generate mutant constructs for Cdc25A 3′-UTR and for the promoter of miR-339-5p. Cells were seeded, and transiently cotransfected with Cdc25A 3′-UTR (wild or mutant type), pGL3-miR-339-5p (wild or mutant type), pGL3-Basic, pCMV3-Runx3, pCMV3, miR-339-5p mimcs and negative control (NC) as indicated. Details of luciferase reporter assays were performed according to the manufacturer’s protocol, and normalized for transfection efficiency by cotransfecting with pRL-TK Renilla. KYSE30 and KYSE180 cells were cross-linked in 1% formaldehyde for 10 min at 37 °C. DNA from the fixed-chromatin cells were then subjected to chromatin immunoprecipitation according to the manufacturer’s instructions (Thermo Fisher Scientific, Australia).
Animal studies
Male BALB/c Nude mice 5 weeks of age were subcutaneously injected into the right leg with miR-339-5p-overexpressing cells or PLVX-control cells (2 × 106). When the diameter of tumor reached approximately 5 mm, tumors were irradiated with a single 12 Gy dose of X-IR. Each group was composed of 5 mice, randomly chosen. Tumor sizes were measured with a digital caliper interval of two days after X-IR, and tumor volume was determined by the formula [length x width2] x 0.5. Data are presented as tumor growth curves. The tumor was harvested and weighed, and analyzed by a two-tailed, unpaired Student’s t test. The experimental procedures were approved by Institutional Animal Care and Use Committee of Chinese Academy of Medical Sciences Cancer Hospital.
Statistical analysis
Statistical analysis was performed using two-tailed, two independent or paired Student’s t tests with SPSS17.0. Correlations were analyzed using Pearson’s correlation coefficient. Survival curves were calculated by Kaplan–Meier method and compared using the log-rank test. An F-test was used to compare the variances between groups, and we found no significant difference. All the data meet normal distribution, and no samples were excluded. The high and low sets were defined according to the mean of miR-339-5p expression in ESCC and TCGA-ESCC data set. High (n = 42) and low group (n = 17) were defined according to the mean of miR-339-5p expression. P < 0.05 was considered statistically significant (*).
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66:7–30.
Pennathur A, Gibson MK, Jobe BA, Luketich JD. Oesophageal carcinoma. Lancet. 2013;381:400–12.
Davies L, Lewis WG, Arnold DT, Escofet X, Blackshaw G, Gwynne S, et al. Prognostic significance of age in the radical treatment of oesophageal cancer with surgery or chemoradiotherapy: a prospective observational cohort study. Clin Oncol. 2010;22:578–85.
Shapiro J, van Lanschot JJB, Hulshof M, van Hagen P, van Berge Henegouwen MI, Wijnhoven BPL, et al. Neoadjuvant chemoradiotherapy plus surgery versus surgery alone for oesophageal or junctional cancer (CROSS): long-term results of a randomised controlled trial. Lancet Oncol. 2015;16:1090–8.
Sjoquist KM, Burmeister BH, Smithers BM, Zalcberg JR, Simes RJ, Barbour A, et al. Survival after neoadjuvant chemotherapy or chemoradiotherapy for resectable oesophageal carcinoma: an updated meta-analysis. Lancet Oncol. 2011;12:681–92.
Kawai T, Kochi M, Fujii M, Song K, Hagiwara K, Watanabe M, et al. Neoadjuvant chemoradiotherapy for stage II or III esophageal squamous cell carcinoma. Anticancer Res. 2017;37:3301–6.
Hamai Y, Hihara J, Emi M, Furukawa T, Ibuki Y, Yamakita I, et al. Effects of neoadjuvant chemoradiotherapy on pathological tnm stage and their prognostic significance for surgically-treated esophageal squamous cell carcinoma. Anticancer Res. 2017;37:5639–46.
Pasquinelli AE. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet. 2012;13:271–82.
Chowdhury D, Choi YE, Brault ME. Charity begins at home: non-coding RNA functions in DNA repair. Nat Rev Mol Cell Biol. 2013;14:181–9.
Majidinia M, Yousefi B. DNA damage response regulation by microRNAs as a therapeutic target in cancer. DNA Repair. 2016;47:1–11.
Cheng CJ, Bahal R, Babar IA, Pincus Z, Barrera F, Liu C, et al. MicroRNA silencing for cancer therapy targeted to the tumour microenvironment. Nature. 2015;518:107–10.
Zhang P, Wang L, Rodriguez-Aguayo C, Yuan Y, Debeb BG, Chen D, et al. miR-205 acts as a tumour radiosensitizer by targeting ZEB1 and Ubc13. Nat Commun. 2014;5:5671.
Odenthal M, Bollschweiler E, Grimminger PP, Schroder W, Brabender J, Drebber U, et al. MicroRNA profiling in locally advanced esophageal cancer indicates a high potential of miR-192 in prediction of multimodality therapy response. Int J Cancer. 2013;133:2454–63.
Georges SA, Biery MC, Kim SY, Schelter JM, Guo J, Chang AN, et al. Coordinated regulation of cell cycle transcripts by p53-Inducible microRNAs, miR-192 and miR-215. Cancer Res. 2008;68:10105–12.
Pichiorri F, Suh SS, Rocci A, De Luca L, Taccioli C, Santhanam R, et al. Downregulation of p53-inducible microRNAs 192, 194, and 215 impairs the p53/MDM2 autoregulatory loop in multiple myeloma development. Cancer Cell. 2010;18:367–81.
Taneja N, Davis M, Choy JS, Beckett MA, Singh R, Kron SJ, et al. Histone H2AX phosphorylation as a predictor of radiosensitivity and target for radiotherapy. J Biol Chem. 2004;279:2273–80.
Banath JP, Macphail SH, Olive PL. Radiation sensitivity, H2AX phosphorylation, and kinetics of repair of DNA strand breaks in irradiated cervical cancer cell lines. Cancer Res. 2004;64:7144–9.
Jansson MD, Damas ND, Lees M, Jacobsen A, Lund AH. miR-339-5p regulates the p53 tumor-suppressor pathway by targeting MDM2. Oncogene. 2015;34:1908–18.
Falck J, Mailand N, Syljuasen RG, Bartek J, Lukas J. The ATM-Chk2-Cdc25A checkpoint pathway guards against radioresistant DNA synthesis. Nature. 2001;410:842–7.
Kim S, Lee UJ, Kim MN, Lee EJ, Kim JY, Lee MY, et al. MicroRNA miR-199a* regulates the MET proto-oncogene and the downstream extracellular signal-regulated kinase 2 (ERK2). J Biol Chem. 2008;283:18158–66.
Tellez CS, Juri DE, Do K, Picchi MA, Wang T, Liu G, et al. miR-196b Is epigenetically silenced during the premalignant stage of lung carcinogenesis. Cancer Res. 2016;76:4741–51.
Shen B, Zhang Y, Yu S, Yuan Y, Zhong Y, Lu J, et al. MicroRNA-339, an epigenetic modulating target is involved in human gastric carcinogenesis through targeting NOVA1. FEBS Lett. 2015;589:3205–11.
Sakakura C, Miyagawa K, Fukuda KI, Nakashima S, Yoshikawa T, Kin S, et al. Frequent silencing of RUNX3 in esophageal squamous cell carcinomas is associated with radioresistance and poor prognosis. Oncogene. 2007;26:5927–38.
Villarroya-Beltri C, Gutierrez-Vazquez C, Sanchez-Cabo F, Perez-Hernandez D, Vazquez J, Martin-Cofreces N, et al. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat Commun. 2013;4:2980.
Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem. 2010;285:17442–52.
Morgan MA, Lawrence TS. Molecular pathways: overcoming radiation resistance by targeting DNA damage response pathways. Clin Cancer Res. 2015;21:2898–904.
Maier P, Hartmann L, Wenz F, Herskind C. Cellular pathways in response to ionizing radiation and their targetability for tumor Radiosensitization. Int J Mol Sci. 2016;17:E102.
Long JM, Ray B, Lahiri DK. MicroRNA-339-5p down-regulates protein expression of beta-site amyloid precursor protein-cleaving enzyme 1 (BACE1) in human primary brain cultures and is reduced in brain tissue specimens of Alzheimer disease subjects. J Biol Chem. 2014;289:5184–98.
Ueda R, Kohanbash G, Sasaki K, Fujita M, Zhu X, Kastenhuber ER, et al. Dicer-regulated microRNAs 222 and 339 promote resistance of cancer cells to cytotoxic T-lymphocytes by down-regulation of ICAM-1. Proc Natl Acad Sci USA. 2009;106:10746–51.
Weber CE, Luo C, Hotz-Wagenblatt A, Gardyan A, Kordass T, Holland-Letz T, et al. miR-339-3p is a tumor suppressor in melanoma. Cancer Res. 2016;76:3562–71.
Wu ZS, Wu Q, Wang CQ, Wang XN, Wang Y, Zhao JJ, et al. MiR-339-5p inhibits breast cancer cell migration and invasion in vitro and may be a potential biomarker for breast cancer prognosis. BMC Cancer. 2010;10:542.
Schwarzenbach H, Nishida N, Calin GA, Pantel K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat Rev Clin Oncol. 2014;11:145–56.
Nedaeinia R, Manian M, Jazayeri MH, Ranjbar M, Salehi R, Sharifi M, et al. Circulating exosomes and exosomal microRNAs as biomarkers in gastrointestinal cancer. Cancer gene Ther. 2017;24:48–56.
Challagundla KB, Wise PM, Neviani P, Chava H, Murtadha M, Xu T, et al. Exosome-mediated transfer of microRNAs within the tumor microenvironment and neuroblastoma resistance to chemotherapy. J Natl Cancer Inst. 2015;107:djv135.
Tanaka Y, Kamohara H, Kinoshita K, Kurashige J, Ishimoto T, Iwatsuki M, et al. Clinical impact of serum exosomal microRNA-21 as a clinical biomarker in human esophageal squamous cell carcinoma. Cancer. 2013;119:1159–67.
Takeshita N, Hoshino I, Mori M, Akutsu Y, Hanari N, Yoneyama Y, et al. Serum microRNA expression profile: miR-1246 as a novel diagnostic and prognostic biomarker for oesophageal squamous cell carcinoma. Br J Cancer. 2013;108:644–52.
Melo SA, Sugimoto H, O’Connell JT, Kato N, Villanueva A, Vidal A, et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell. 2014;26:707–21.
Prathipati P, Nandi SS, Mishra PKStemCell-DerivedExosomes. Autophagy, extracellular matrix turnover, and miRNAs in cardiac regeneration during stem cell therapy. Stem Cell Rev. 2017;13:79–91.
Phinney DG, Pittenger MF. Concise review: MSC-derived exosomes for cell-free therapy. Stem Cells. 2017;35:851–8.
Eriksson D, Stigbrand T. Radiation-induced cell death mechanisms. Tumour Biol: J Int Soc Oncodev Biol Med. 2010;31:363–72.
Yamada C, Ozaki T, Ando K, Suenaga Y, Inoue K, Ito Y, et al. RUNX3 modulates DNA damage-mediated phosphorylation of tumor suppressor p53 at Ser-15 and acts as a co-activator for p53. J Biol Chem. 2010;285:16693–703.
Krishnan V, Ito Y. A regulatory role for RUNX1, RUNX3 in the maintenance of genomic integrity. Adv Exp Med Biol. 2017;962:491–510.
Shimada Y, Imamura M, Wagata T, Yamaguchi N, Tobe T. Characterization of 21 newly established esophageal cancer cell lines. Cancer. 1992;69:277–84.
Jorgensen S, Baker A, Moller S, Nielsen BS. Robust one-day in situ hybridization protocol for detection of microRNAs in paraffin samples using LNA probes. Methods. 2010;52:375–81.
Acknowledgements
We thank Dr. Yutaka Shimada for the generous gifts of ESCC cell lines (KYSE series). This research was supported by National Key R&D program of China (2016YFC1302100), National Natural Science Foundation of China (81472791, 81372418), and CAMS Initiative for Innovative Medicine (2016-I2M-1-001).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Luo, A., Zhou, X., Shi, X. et al. Exosome-derived miR-339-5p mediates radiosensitivity by targeting Cdc25A in locally advanced esophageal squamous cell carcinoma. Oncogene 38, 4990–5006 (2019). https://doi.org/10.1038/s41388-019-0771-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0771-0
- Springer Nature Limited
This article is cited by
-
Circular RNA circ-SLC7A5 Functions as a Competing Endogenous RNA to Impact Cell Biological Behaviors in Esophageal Squamous Cell Carcinoma (ESCC)
Cell Biochemistry and Biophysics (2024)
-
The roles of exosomes in esophageal cancer
Discover Oncology (2024)
-
Novel insights into the heterogeneity of FOXP3 + Treg cells in drug-induced allergic reactions through single-cell transcriptomics
Immunologic Research (2024)
-
Molecular mechanisms of AMPK/YAP/NLRP3 signaling pathway affecting the occurrence and development of ankylosing spondylitis
Journal of Orthopaedic Surgery and Research (2023)
-
Mechanisms of radiotherapy resistance and radiosensitization strategies for esophageal squamous cell carcinoma
Molecular Cancer (2023)