Abstract
Circular RNAs are a large group of noncoding RNAs that are widely expressed in mammalian cells. Genome-wide analyses have revealed abundant and evolutionarily conserved circular RNAs across species, which suggest specific physiological roles of these species. Using a microarray approach, we detected increased expression of a circular RNA circ-Dnmt1 in eight breast cancer cell lines and in patients with breast carcinoma. Silencing circ-Dnmt1 inhibited cell proliferation and survival. Ectopic circ-Dnmt1 increased the proliferative and survival capacities of breast cancer cells by stimulating cellular autophagy. We found that circ-Dnmt1-mediated autophagy was essential in inhibiting cellular senescence and increasing tumor xenograft growth. We further found that ectopically expressed circ-Dnmt1 could interact with both p53 and AUF1, promoting the nuclear translocation of both proteins. Nuclear translocation of p53 induced cellular autophagy while AUF1 nuclear translocation reduced Dnmt1 mRNA instability, resulting in increased Dnmt1 translation. From here, functional Dnmt1 could then translocate into the nucleus, inhibiting p53 transcription. Computational algorithms revealed that both p53 and AUF1 could bind to different regions of circ-Dnmt1 RNA. Our results showed that the highly expressed circular RNA circ-Dnmt1 could bind to and regulate oncogenic proteins in breast cancer cells. Thus circ-Dnmt1 appears to be an oncogenic circular RNA with potential for further preclinical research.
Similar content being viewed by others
Introduction
CircRNAs are a large class of single-stranded RNAs that have recently emerged as a new class of noncoding RNAs with a widespread expression pattern [1,2,3]. Genome-wide analyses of RNA sequencing data have revealed evolutionary conservation and abundance of circular RNAs [4, 5]. Circular RNAs can be generated from exons, called exonic circular RNAs [3, 4], or from introns, called intronic circular RNAs or a mix of both called ciRNAs [6]. Some circular RNAs are generated from intron and exon, producing exon–intron circRNAs or EIciRNAs [7]. Studies on human circRNAs have revealed that circular RNAs are usually composed of 1–5 exons, which can be three times longer than the average exon [4, 8]. Typically, the second exon acts as the acceptor exon [9]. The introns involved in the circularization are also longer than those which are not involved [9, 10]. It has been shown that exonic circular RNAs are predominantly detected in the cytoplasm, while intronic RNAs are mainly localized to the nuclei [8].
Studies from mouse and human cells have shown that many orthologous circular RNAs are evolutionarily conserved, suggesting specific roles of circular RNAs in cellular physiology [8]. One mode of action for circular RNAs appears to be as endogenous miRNA sponges since some circular RNAs possess miRNA binding sites, allowing them to arrest miRNA activity [11,12,13]. Circular RNAs can then regulate cancer development [14,15,16]. CircRNAs are proposed to be more effective sponges than linear transcripts due to their increased abundance and relative stability [2, 15]. Some circular RNAs may interact with RNA-binding proteins which form RNA–protein complex and regulate their activity [9, 17]. We have found that some circ-Foxo3 binds different proteins and plays roles in different pathways [18,19,20]. It has been proposed that circular RNAs may be relevant to the development of atherosclerosis, Parkinson’s disease and cancer, although this remains to be further explored [21, 22]. To explore the involvement of circular RNAs in breast cancer development, we analyzed levels of circular RNAs in human breast cancer cells and non-cancer breast cells by microarray, and found that a circular RNA circ-Dnmt1 was upregulated in all of the cancer cell lines. This study aimed to explore the biological role of circ-Dnmt1 in breast cancer oncogenesis and development.
Results
Circ-Dnmt1 enhances autophagy
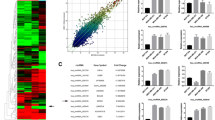
We analyzed levels of circular RNAs in human breast cancer cells and non-cancer breast cells by microarray and found that circ-Dnmt1 (hsa_circRNA_102439) was upregulated in all cancer cell lines relative to the non-cancer cells (Fig. 1a). Real-time PCR analysis showed that most of the cancer cell lines expressed higher levels of this isoform compared with the non-cancer cell lines (Fig. 1b, No 1–4). By real-time PCR, we detected significant upregulation of circ-Dnmt1 in the tumors of breast carcinoma patients compared to the adjacent tissues (Fig. 1c).
circ-Dnmt1 enhanced cell survival. a Comparison of circRNA levels in eight human breast cancer cell lines relative to human breast cell line MCF-10A by using the Agilent Human circular RNA microarray. b RT-PCR showed that expression of circ-Dnmt1 in human non-cancer cell lines (HaCaT, 293T, BEAS2B, Jurkat, HUVEC, and MCF10A), breast cancer cell lines (MCF-7, MDA-MB-468, MDA-MB-231, HTB126, and SK-BR-3), and other tumor cell lines (H460, LnCap, Du145, PC3, JHH1, SNU449, HepG2, and Hela). c Total RNAs were isolated from tumor tissues and adjacent tissues of breast carcinoma patients and subjected to qPCR. The levels of circ-Dnmt1 were significantly higher in the tumor tissues relative to the adjacent tissues. d Structure of circ-Dnmt1 and a designed circ-Dnmt1 expression construct. e Total RNA was extracted from mock- or circ-Dnmt1-transfected MCF-7 cells and incubated with or without RNAse R at 37 °C for 10 min, followed by gel electrophoresis and northern hybridization using circ-Dnmt1 sequence as a probe to confirm overexpression of circ-Dnmt1 and its resistance to RNAse R treatment. The lower portion was the photo of Ethidium Bromide staining. f Left, RT-PCR showing product from vector- and circ-Dnmt1-transfected cells. Right, The PCR product was cloned and sequenced. This confirmed the correct junction (arrow) sequence of circ-Dnmt1. g Circ-Dnmt1-, vector-transfected, and wild-type MCF-7 cells were seeded in 96-well plates at the density of 1 cell per well in DMEM containing 10% FBS. Cell numbers were counted for up to 13 days
Circ-Dnmt1 is the circularized product of Exons 6 and 7 of the mRNA NM_001130823.1 (Fig. 1d). To explore the function of circ-Dnmt1, we generated circ-Dnmt1 expression construct (Fig. 1d). Expression of circ-Dnmt1 was confirmed by real-time PCR (Fig S1a). To confirm circularization of the expressed product, we treated RNAs extracted from vector- and circ-Dnmt1-transfected MCF-7 cells without or with RNAse R, followed by Northern blot and qPCR. By northern blotting, we showed that transfection with circ-Dnmt1 produced higher levels of circ-Dnmt1 than the control (Fig. 1e). By qPCR, we confirmed that treatment with RNAse R decreased levels of Dnmt1 linear mRNA, but it did not affect circ-Dnmt1 levels (Fig S1b). By RT-PCR, we detected the expected size of PCR product (Fig. 1f, left). The PCR product was cloned and sequenced. It revealed the exact junction sequence (Fig. 1f, right). We tested whether ectopic transfection of circ-Dnmt1 had effect on host locus. While transfection with circ-Dnmt1 increased Dnmt1 mRNA levels, it had no effect on Dnmt1 pre-mRNA levels (Fig S1c).
To examine the effects of circ-Dnmt1, we performed a gross growth assay and found that expression of circ-Dnmt1 enhanced cell proliferation (Fig S1d) and survival (Fig S1e). We then examine these effects in greater detail by using a single-cell culture. Individual circ-Dnmt1-, vector-transfected, or wild-type MCF-7 cells were cultured into 96-well plates (1 cell per well). Quantification was performed for 13 days. Ectopic expression of circ-Dnmt1 increased proliferation relative to vector-transfected or wild-type cells (Fig. 1g). Single-cell growth curves showed that only the circ-Dnmt1-transfected cells reached an exponential growth phase in our set time points. We also found that circ-Dnmt1-transfected cells survived longer than the control cells (Fig S1f).
Our results showed that serum played important roles in circ-Dnmt1-mediated cell proliferation. We observed that following a similar number of replicative cycles, the vector-transfected cells revealed a senescent phenotype and characteristic beta-galactosidase staining. We also found that circ-Dnmt1 overexpression increased the number of replicative cycles required for induction of a similar senescent phenotype (Fig. 2a, Fig S2a). Cell lysates prepared from circ-Dnmt1-, and vector-transfected, MCF-7 cells were analyzed by western blotting for LC3B, a marker of autophagy. We detected increased LC3B expression in the circ-Dnmt1-transfected MCF-7 cells (Fig. 2b).
circ-Dnmt1 enhanced autophagy. a Circ-Dnmt1-transfected MCF-7 cells were treated with 150 µM H2O2 for 2 h, and then cultured in basal medium for 48 h, followed by β-gal staining. Transfection with circ-Dnmt1 decreased β-gal staining (n = 6). b Cell lysates prepared from circ-Dnmt1- and vector-transfected cells was assayed on western blot that was probed with anti-LC3B antibody. LC3B levels increased in the circ-Dnmt1-transfected MCF-7 cells. c circ-Dnmt1- and vector-transfected MCF-7 cells were cultured with autophagy inhibitors SP600125 (50 µM), Chloroquine (10 µM), LY294002 (10 µM), and Wartmannin (10 µM) for 24 h, and circ-Dnmt1 levels were further enhanced by autophagy inhibitors. d Autophagy inhibitors could abolished circ-Dnmt1 repressed senescence and enhanced survival (n = 6). e Cell lysates were prepared from MDA-MB-231 cells transfected with or without (wild type, WT) control oligo 1 (olig1), oligo 2 (olig2), circ-Dnmt1 siRNA1 (si-Dn1), siRNA2 (si-Dn2), and siRNA1 + siRNA2, showed that silencing circ-Dnmt1 decreased circ-Dnmt1 expression (left), decreased Dnmt1 mRNA levels (middle), but did not affect pre-mRNA of Dnmt1 (right). f Silencing circ-Dnmt1 decreased expression of Dnmt1 and LC3B. g The cells were also stained with phalloidins, DAPI, and either Dnmt1 or LC3B. Image J analyses showed that circ-Dnmt1 siRNA decreased Dnmt1 and LC3B expression (n = 6). h Left, circ-Dnmt1- and vector-transfected MCF-7 cells were cultured with LY294002 (10 µM) for 24 h, LY294002 abolished circ-DNMT1 enhanced DNMT1 and LC3B. Right, circ-Dnmt1 siRNA- and control oligo-transfected MDA-MB-231 cells were cultured with LY294002 (10 µM) for 24 h, LY294002 repressed DNMT1 and LC3B expression (n = 6). i circ-Dnmt1 siRNA- and control oligo-transfected MB-231 cells were cultured in serum-free medium for 9 days. Autophagy inhibitors repressed control cell survival but not the siRNA-transfected cells (n = 6)
To test whether autophagy was essential in mediating circ-Dnmt1 functions, we treated the circ-Dnmt1- and vector-transfected MCF-7 cells with autophagy inhibitors SP600125, Chloroquine, LY294002, and Wartmannin, and found that circ-Dnmt1 levels were promoted by autophagy inhibitors (Fig. 2c), while LC3B levels decreased by treatment with these autophagy inhibitors (Fig S2b). The effects of circ-Dnmt1 on cellular senescence and survival were abolished in the presence of these autophagy inhibitors (Fig. 2d). Our results suggested that cellular autophagy was central in mediating the effects of circ-Dnmt1 on cellular senescence and survival.
We designed two siRNAs that targeted circ-Dnmt1. Transfection with these siRNAs greatly decreased circ-Dnmt1 and Dnmt1 levels, but had no effect on Dnmt1 transcription, measured by primers located in an intron (Fig. 2e). Silencing circ-Dnmt1 decreased expression of Dnmt1 and LC3B (Fig. 2f) but increased β-gal staining in MDA-MB-231 (Fig S2c) and MCF-7 (Fig S2d) cells treated with H2O2. These appeared to be the silencing effect of circ-Dnmt1 by the siRNAs.
Immunostaining and Image J analyses confirmed decreased expression of Dnmt1 and LC3B in the circ-Dnmt1 siRNA-transfected cells (Fig. 2g, Fig S2e). There was a positive correlation between circ-Dnmt1 and LC3B levels (Fig S2f). In the presence of the authophagy inhibitors, the effects of circ-Dnmt1 on expression of Dnmt1 and LC3B (Fig. 2h), cell survival (Fig. 2i), and senescence (Fig S2g) were abolished.
Circ-Dnmt1 promoted p53 and Auf1 nuclear translocation
Using immunofluorescence, we examined expression of LC3B in circ-Dnmt1- and vector-transfected MCF-7 cells. Alongside the overexpressed nuclear circ-Dnmt1, we found increased cytoplasmic LC3B (Fig. 3a, full panel of staining in Fig S3a). To confirm nuclear expression of circ-Dnmt1, we isolated total RNAs from the circ-Dnmt1- and vector-transfected MCF-7 cells and we found increased nuclear circ-Dnmt1 in the circ-Dnmt1-transfected cells (Fig. 3b).
circ-Dnmt1 promoted p53 and Auf1 nuclear translocation. a The circ-Dnmt1- and vector-transfected MCF-7 cells were stained with fluorescence showing expression of Dnmt1 or LC3B. Image J analyses showed that circ-Dnmt1-transfected cells expressed increased levels of Dnmt1 and LC3B (n = 6). b Higher levels of circ-Dnmt1 were significantly detected in nuclei and cytoplasm of the circ-Dnmt1-transfected cells. c Western blotting of tumor lysates showing decreased expression of p53 and increased levels of Dnmt1 in the tumors formed by the circ-Dnmt1-transfected cells. d Protein lysates prepared from circ-Dnmt1- and vector-transfected MCF-7 cells were subject to Western blot. Levels of p53 decreased in the lysate and cytoplasm, but increased in the nuclei of the circ-Dnmt1-transfected cells. Level of Auf1 was not affected by circ-Dnmt1 expression, but decreased in the cytoplasm and increased in the nuclei of the circ-Dnmt1-transfected cells. e Silencing circ-Dnmt1 decreased nuclear translocation of p53 and Auf1
We also detected increased linear Dnmt1 mRNA in the cytoplasm. Since Auf1 and p53 are RNA-binding proteins and the former regulates stability of Dnmt1 mRNA, we examined levels of Auf1 and p53. We observed no change in Auf1 expression, decreased p53 but increased Dnmt1 in the tumors formed by the circ-Dnmt1-transfected cells (Fig. 3c). Protein lysates prepared from cytoplasm and nuclei of the circ-Dnmt1- and vector-transfected MCF-7 cells were subject to western blot. Total Auf1 levels were unaltered, but higher nuclear Auf1 was observed (Fig. 3d). Similarly, we found decreased p53 in the total lysate and cytoplasm, but increased p53 in the nuclei of the circ-Dnmt1-transfected cells (Fig S3b).
Silencing circ-Dnmt1 also increased cytoplasmic translocation but decreased nuclear Auf1 and p53 analyzed by Western blotting (Fig. 3e). The siRNA-transfected cells were used for confocal analysis of circ-Dnmt1, p53, and Auf1. Silencing circ-Dnmt1 decreased p53 and Auf1 in nuclei, but increased p53 levels in the cytosol (Fig S3c).
Autophagy enhances nuclear translocation of circ-Dnmt1, Auf1, and p53
We examined whether autophagy could induce nuclear translocation of circ-Dnmt1. MDA-MB-231 cells treated with autophagy inducers EBSS and Rapamycin displayed increased levels of circ-Dnmt1 in nuclei, but silencing circ-Dnmt1 abolished the effect of the inducers (Fig. 4a). These inducers increased protein levels of LC3B and Dnmt1 and nuclear translocation of Auf1 and p53 (Fig. 4b). In agreement with these results were the increased mRNA levels of Dnmt1 (Fig. 4c), decreased cell senescence (Fig. 4d) and increased viability (Fig. 4e). Silencing circ-Dnmt1 abolished the effects of the inducers. We tested the effect of Rapamycin on a number of circular RNAs in cytosol and observed specific effect of Rapamycin on circ-Dnmt1 (Fig S4a). Treatment with autophagy inhibitor ly294002 increased circ-Dnmt1 levels in the cytosol (Fig S4b).
Tumor cell autophagy activity depends on circ-Dnmt1 expression levels and subcellular distribution. a MB-231 cells were cultured in EBSS or 100 nM rapamycin for 24 h. EBSS and rapamycin promoted circ-Dnmt1 nuclear translocation, which could be abolished by circ-Dnmt1 silencing (n = 6). b EBSS and rapamycin enhanced LC3B and Dnmt1, and decreased p53 expression in cell lysis, while enhanced p53 and Auf1 expression in the nucleus, which could be abolished by circ-Dnmt1 silencing. c EBSS and rapamycin increased Dnmt1 mRNA levels in the cytoplasm, which could be abolished by circ-Dnmt1 silencing (n = 6). d The cells were treated with 100 µM H2O2 for 2 h, and then in basal medium for 48 h. EBSS and rapamycin treatment decreased β-gal staining, which could be abolished by circ-Dnmt1 silencing (n = 6). e The cells were treated with 0.9 mM H2O2 for 12 h. EBSS and rapamycin treatment enhanced cell survival, which could be abolished by circ-Dnmt1 silencing (n = 6). f Overexpression of circ-Dnmt1 increased the percentage of circ-Dnmt1 expression in the nucleus, which could be abolished by autophagy inhibitors (n = 6). g The correlation between circ-Dnmt1 levels and the percentage of circ-Dnmt1 in the cell nucleus (n = 12). h The correlation between the percentage of circ-Dnmt1 in the cell nucleus and expression of LC3B levels (n = 12). i circ-Dnmt1- and vector-transfected MCF-7 cells (left), or circ-Dnmt1 siRNA and control oligo-transfected MB-231 cells (right) were cultured with or without LY294002 (10 µM) for 24 h. Ly294002 abolished circ-Dnmt1 promoted p53 and Auf1 nuclear translocation
The increased circ-Dnmt1 in nuclei as a result of circ-Dnmt1 overexpression was abolished in the presence of the autophagy inhibitors (Fig. 4f). In these cells, while Dnmt1 mRNA levels increased in the cytosol of the circ-Dnmt1-transfected cells, treatment with the autophagy inhibitors suppressed Dnmt1 mRNA levels in the cytosol (Fig S4c, left). The total Dnmt1 mRNA levels were not increased as much (Fig S4c, right), suggesting accumulation of Dnmt1 mRNA in cytosol. The increased expression (Fig S4d), increased cytosol levels (Fig S4e), but decreased nuclear levels (Fig S4f) of circ-Dnmt1 as a result of treatment with autophagy inhibitors, were abolished by silencing endogenous circ-Dnmt1.
Increased expression of circ-Dnmt1 enhanced nuclear translocation of circ-Dnmt1 (Fig. 4g), and increased circ-Dnmt1 nuclear translocation promoted LC3B expression (Fig. 4h) leading to autophagy. Treatment with autophagy inhibitor Ly294002 inhibited nuclear translocation of p53 and Auf1 (Fig. 4i, left). Silencing circ-Dnmt1 further increased this effect (Fig. 4i, right). Cell lines expressing higher levels of circ-Dnmt1 (Fig S4g) also displayed increased levels of circ-Dnmt1 in nuclei (Fig S4h) and expressed higher levels of LC3B (Fig S4i).
Circ-Dnmt1 interacted with p53 and Auf1
To test whether circ-Dnmt1 interacted with p53 and Auf1, we prepared lysates from MB-231 cells, followed by co-precipitation using antibodies against p53, Auf1 and PABP that binds mRNAs but not circular RNAs. The precipitated products were subject to western blotting and qPCR with primers amplifying circ-Dnmt1. After confirming that the antibodies pulled down the relative protein (Fig S5a), real-time qPCR indicated that anti-p53 and anti-Auf1 antibodies pulled down increased levels of circ-Dnmt1 from circ-Dnmt1-transfected cells relative to control (Fig. 5a). In the untransfected cells, anti-p53 and anti-Auf1 antibodies precipitated high levels of circ-Dnmt1 compared to the linear Dnmt1 (Fig S5b). However, the anti-p53 and anti-Auf1 antibodies did not precipitate other circular RNAs (Fig S5c).
circ-Dnmt1 modulated expression of p53 and Dnmt1 via autophagy. a Antibodies against p53 and Auf1 pulled down higher levels of circ-Dnmt1 than Dnmt1 mRNA. PABP antibody (mRNA positive control) only pulled down Dnmt1 mRNA, but not circ-Dnmt1. b Upper, lysates were prepared from MCF-7 cells that were transfected with vector and circ-Dnmt1. The lysates were subjected to western blot analysis. Expression of circ-Dnmt1 decreased p53. Lower, In pull-down assay, circ-Dnmt1 probe co-precipitated p53 and AUF1 in cells transfected with circ-Dnmt1. c Anti-p53 and anti-Auf1 antibodies pulled down decreased levels of circ-Dnmt1 in the cells transfected with circ-Dnmt1 siRNAs. d The circ-Dnmt1 probe pulled down decreased amounts of p53 and Auf1 when circ-Dnmt1 was knocked down by the siRNAs. e The diagram shows that circ-Dnmt1 promotes p53 and Auf1 nuclear translocation. Auf1 nuclear translocation increases Dnmt1 translation, leading to inhibiting p53 transcription. As a result, the ectopic circ-Dnmt1 expression enhanced cellular autophagy and repressed senescence, leading to increased tumor growth. f Left, ectopic expression of circ-Dnmt1 in MCF-7 cells decreased p53 mRNA levels. Right, transfection with circ-Dnmt1 siRNA increased p53 mRNA levels in MDA-MB-231 cells. g Upper, cell nuclear lysis prepared from MCF-7 cells were subject to immuno-precipitation. RIP assays indicated that anti-p53 and anti-Auf1 antibodies pulled down much more circ-Dnmt1 than linear RNA, which was significant when the samples were pre-treated with cross-linking. Lower, RIP assays indicated anti-p53 and anti-Auf1 antibodies pulled down more circ-Dnmt1 from cells transfected with circ-Dnmt1 than from control, which was significant when the samples were subjected to cross-linking (n = 6)
The lysates were also incubated with a probe of circ-Dnmt1, followed by RNA pull-down assay. We confirmed that the circ-Dnmt1 probe pulled down higher levels of circ-Dnmt1 in the cells that were transfected with circ-Dnmt1 (Fig S5d). This probe that pulled down circ-Dnmt1, also precipitated p53 and Auf1 (Fig. 5b). In the lysates prepared from circ-Dnmt1 siRNA-transfected MB-231 cells, after confirming the silencing effect (Fig S5e), antibodies against p53 and Auf1 pulled down less circ-Dnmt1 (Fig. 5c). The probe also pulled down less circ-Dnmt1 in the cells that were transfected with circ-Dnmt1 siRNAs (Fig S5f). Although silencing circ-Dnmt1 increased p53 levels, the circ-Dnmt1 probe co-precipitated decreased levels of p53 and Auf1 (Fig. 5d).
Our results showed that ectopic circ-Dnmt1 could bind to both p53 and Auf1, and promote p53 and Auf1 nuclear translocation. Nuclear translocation of p53 induced cellular autophagy. Auf1 nuclear translocation relieved its effect on instability of Dnmt1 mRNA resulting in increased Dnmt1 translation. Dnmt1 protein could then enter into the nucleus and inhibit p53 transcription by promoter methylation (Fig. 5e). We also detected significant reduction in p53 transcription relative to control (Fig. 5f, left). Silencing endogenous circ-Dnmt1 abrogated these effects (Fig. 5f, right). In the cross-linking assays, we found that anti- Auf1 and anti-p53 antibodies precipitated circ-Dnmt1 relative to the linear Dnmt1 mRNA (Fig. 5g, upper). Ectopic circ-Dnmt1 increased the pulled down levels (Fig. 5g, lower). As a consequence, the ectopically expressed circ-Dnmt1 enhanced cellular autophagy and repressed senescence. This was well correlated with increased Dnmt1 mRNA and protein (Fig S5g).
Computational algorithm of circ-Dnmt1 interacting with Auf1 and p53
The predicted secondary structure of circ-Dnmt1 was obtained by using the formula ΔG = ΔH–TΔS, ΔG = −14.70 kcal/mol at 37 °C, ΔS = −1177.8 cal/(K·mol), and ΔH = −380.00 kcal/mol, where T (K) is the absolute temperature while ΔH, ΔS, and ΔG denote the changes in enthalpy, entropy, and free energy, respectively. The secondary structure delineated in dot bracket notation was assayed by the software RNA composer for tertiary structure prediction. Finally, NPDock was used to perform the in-silico docking of circ-Dnmt1 with p53 (C-terminal regulatory domain) and Auf1 (N-terminal and C-terminal RNA-binding domains) (Fig S6a). The p53 C-terminus containing a negative regulatory domain has been shown to interact with RNA [23] that we used for docking analysis. The input was derived from the Protein Data Bank (PDB) entry 1DT7. The molecular simulation result supports that circ-Dnmt1 could dock p53 and predicted a minimal binding region of circ-Dnmt1 for p53: ‘aaccttcac’‘aac’‘aggaagaa’. The contact map (Fig S6b), the residue-level resolution contact maps (Fig S6c), the MC score (Fig S6d), and the contact distance (Table S1), the Accessible Surface Area (Table S2) all supported the conclusion that circ-Dnmt1 could dock the p53 C-terminal regulatory motif. The difference in the three-dimensional structures between circ-Dnmt1 (Fig S6e) and Dnmt1 linear mRNA (Fig S6f) also confirmed our results that p53 only bound to circ-Dnmr1 but not to Dnmt1 linear mRNA.
Auf1 contains two ribonucleoprotein (RNP)-type RNA-binding domains (RBDs), N-terminus, and glutamine-rich domain. The NMR structure of RBD1 (N-terminal) [24] and RBD2 (C-terminal) [25] have previously been resolved. Both N and C-terminal RBDs of Auf1 were used in the docking analysis. Auf1 (N-terminal RBD1) used in the docking procedure was derived from PDB entry 1HD0. The molecular simulation result supports that circ-Dnmt1 could perfectly dock Auf1 (N-terminal RBD1) and predicted a minimal binding site of circ-Dnmt1 for Auf1 at the junction of circularization: ‘aaccttcac’ ‘ttaca’ ‘c’ ‘c’ ‘gagt’ ‘ag’ ‘agaa’ (Fig S7a). The conclusion that circ-Dnmt1 sufficiently docked RBD1 was supported by the contact map (Fig S7b), residue-level maps (Fig S7c), MC score (Fig S7d), contact distance (Table S3), and Accessible Surface Area (Table S4).
Auf1 (C-terminal RBD2) inputs used in the docking were obtained from PDB entry 1X0F. The simulation result confirmed that circ-Dnmt1 perfectly docked RBD2 and predicts a minimal circ-Dnmt1 binding site for RBD2 at the junction of circularization: ‘tgaaccttc’ ‘tta’ ‘tca’ ‘ga’ ‘aggaagaaga’ (Fig S7e). The conclusion that circ-Dnmt1 sufficiently docked RBD2 was supported by the contact map (Fig S7f), residue-level contact maps (Fig S7g), MC score (Fig S7h), contact distance (Table S5), and Accessible Surface Area (Table S6).
The buried surface area (BSA) is the primary and critical descriptor to be directly related to binding affinity and to the interaction based on the Chothia–Janin model [26, 27]. To further determine the binding affinity of circ-Dnmt1 for RBD1 and RBD2, we took into account various functional and structural features namely the network of inter-residue contacts (ICs) along with BSA. The circ-Dnmt1-RBD1 showed an increased binding affinity with a BSA (3692.3 A2) and ICs (83) as compared to circ-Dnmt1-RBD2 complex having a BSA of (3472.1 A2) and ICs (78).
To test the effects of the binding sites on mediating circ-Dnmt1 functions, we generated mutations abolishing the interaction with Auf1 and p53 by site-directed mutagenesis (Fig S8a) and designed blocking oligo of the binding sites (Fig S8b). Lysates prepared from MCF-7 cells were subjected to pull-down assay. Anti-Auf1 and anti-p53 antibodies could no longer precipitate circ-Dnmt1 when circ-Dnmt1 was mutated or blocked with the blocking oligo (Fig S8c). While the probe could pull-down circ-Dnmtl1 (Fig S8d), it did not precipitate Auf1 and p53 (Fig. 6a). As a consequence, MCF-7 cells transfected with the blocking oligo or the mutant showed decreased proliferation rates (Fig S8e), survival, but increased levels of senescence (Fig. 6b). This appeared to be the consequence of decreased nuclear translocation of Auf1 and p53 (Fig. 6c).
Computational analysis of the interaction between circ-Dnmt1 and p53 or Auf1. a Upper, expression of circ-Dnmt1 decreased p53, which could be abolished by the mutation or the blocking oligo. Lower, circ-Dnmt1 probe co-precipitated p53 and Auf1 in cells transfected with circ-Dnmt1, which could be abolished by the mutation or the blocking oligo. b Left, circ-Dnmt1 expression enhanced MCF-7 cell survival that was abolished by the mutation or the blocking oligo. Right, expression of circ-Dnmt1 decreased β-gal staining, which could be abolished by the mutation or the blocking oligo (n = 4). c Expression of circ-Dnmt1 decreased p53 levels, but increased Dnmt1 and LC3B levels, and promoted p53 and Auf1 nuclei translocation, which could be abolished by the mutation or the blocking oligo. d Left, the blocking oligo repressed cell survival. Right, the blocking oligo showed increased β-gal staining (n = 6). e Transfection with the blocking oligo increased p53 levels in cell lysates and cytoplasm but decreased p53 levels in the nucleus. Auf1 expression was not affected by the blocking oligo, but Auf1 translocation to cytoplasm was induced. The blocking oligo also repressed Dnmt1 and LC3B expression
To corroborate these results, we transfected the blocking oligo or the control into MB-231 cells and confirmed that delivery of the blocking oligo inhibited precipitation of circ-Dnmt1 by anti-Auf1 and anti-p53 antibodies (Fig S8f). Transfection of the blocking oligo decreased cell survival but increased senescence (Fig. 6d). It also decreased nuclear translocation of Auf1 and p53 (Fig. 6e), suggesting a role of endogenous circ-Dnmt1 in regulating cell activities by modulating Auf1 and p53 translocation.
Circ-Dnmt1 enhances tumor xenograft growth and decreased mouse survival
In vivo, we found that expression of circ-Dnmt1 significantly increased tumor xenograft volume relative to a vector-control (Fig. 7a, Fig S9a). After confirming upregulation of circ-Dnmt1, we detected increased levels of LC3B in tumors formed by the circ-Dnmt1-transfected cells (Fig. 7b). Tumor sections were also subject to in situ hybridization (Fig S9b-c), followed by Image J analysis (Fig S9d) and immunostaining (Fig S9e). Most of Auf1, p53, and circ-Dnmt1 were found translocated to nuclei.
In vivo analysis of circ-Dnmt1 functions. a Expression of circ-Dnmt1 enhanced tumor growth. b Left, real-time PCR confirmed increased expression of circ-Dnmt1 in the circ-Dnmt1 tumors. Right, western blotting showed that tumors from the circ-Dnmt1 group expressed higher levels of LC3B relative to the control group. c Mice were intraperitoneally injected with MB-231 cells and Kaplan–Meier of survival curves was obtained. Comparing to control oligo group, injection with circ-Dnmt1 siRNA, blocking oligo or autophagy inhibitor LY294002 prolonged the lifespan significantly (log-rank test, **, p < 0.0001). The EBSS group mice lived shorter lifespans than control group, which could be attenuated by delivery of circ-Dnmt1 siRNA (EBSS vs. oligo, **, p = 0.035). d Analysis with Image J revealed that delivery of circ-Dnmt1 siRNA, the blocking oligo or autophagy inhibitor LY294002 repressed AUF1 (upper) and p53 (lower) levels in the nuclei (n = 6). e Western blotting showed decreased nuclear translocation of Auf1 and p53 and decreased expression of LC3B and Dnmt1, when circ-Dnmt1 was silenced or blocked, which could be reversed by autophagy promotion. f Diagram showing that circ-Dnmt1 switches on/off autophagy in breast cancer progression
Since the delivering efficiency of plasmid and blocking oligo into the tumors was low, nude mice were injected with MB-231 cells or EBSS-treated MB-231 cells intraperitoneally to form ascites tumors, followed by delivering the control oligo, siRNA targeting circ-Dnmt1, the blocking oligo, and Ly294002 mixed with oligo. Mouse survival was monitored and the assay was terminated on day 43. We found that the lifespan of mice was significantly extended in mice delivered with the siRNA, the blocking oligo, and Ly294002 relative to the mice delivered with the control oligo and EBSS treatment (Fig. 7c, Fig S10).
Tumor sections were subject to immunohistochemical staining followed by confocal microscopic examination. Treatment with the siRNA decreased circ-Dnmt1 levels (Fig S11a), but Auf1 expression was not affected (Fig S11b). Nuclear translocation of circ-Dnmt1 decreased in the siRNA and autophagy inhibitor treated groups (Fig S11c). However, nuclear translocation of Auf1 and p53 was inhibited by the blocking oligo (Fig. 7d), while total levels of p53 were higher in these groups relative to the groups treated by the control oligo or EBSS (Fig S11d). Consistently, the levels of LC3B and Dnmt1 were lower as a result of lacking nuclear translocation of Auf1 and p53 (Fig S11e). Isolation of nuclei from the tumor tissues confirmed that nuclear translocation of circ-Dnmt1 was inhibited by treatment with siRNA and autophagy inhibitor Ly294002 (Fig S11f). Western blotting confirmed these results that nuclear translocation of Auf1 and p53 was inhibited by siRNA treatment, by the blocking oligo, and the autophagy inhibitor Ly294002 (Fig. 7e). In summary, increased circ-Dnmt1 expression would facilitate nuclear translocation of Auf1 and p53, decrease cytosolic levels of these two proteins, enhance expression of Dnmt1 and LC3B, leading to increased autophagy and tumor growth (Fig. 7f).
Discussion
In this study, we found that circ-Dnmt1 is a highly expressed circular RNA in human breast cancer cells. Expression of circ-Dnmt1 not only increased cell proliferation and survival, but also stimulated cellular autophagy, inhibited cellular senescence and increased tumor growth. These effects were modulated by directly binding to p53 and Auf1, and promoting their nuclear translocation. Nuclear translocation of p53 and Auf1 induced cellular autophagy and increased Dnmt1 expression, inhibiting p53 transcription.
The effects of circ-Dnmt1 on cellular autophagy appeared to be central to its effects on oncogenesis. Without active autophagy, single cells expressing circ-Dnmt1 were unable to proliferate exponentially within our defined time points. This represented an approximate 10-fold increase in growth rate relative to the vector-transfected cells. Based on single-cell proliferation, we showed that circ-Dnmt1-transfected cells could induce auto-phagocytosis for intracellular homeostatic maintenance. This property was not seen in wild-type MCF-7 breast cancer cells. Understanding the heterogeneity of solid tumors, we found that a panel of breast cancer cell lines all expressed increased circ-Dnmt1. We further used a single-cell culture method to evaluate the proliferative capacity of these cells, accounting for phenotypic heterogeneity within these cell lines. With this model, the difference in proliferation and survival between the vector-transfected cells and the circ-Dnmt1-transfected cells remained significant. Relative to vector-transfected MCF-7 cells, circ-Dnmt1-transfected cells could better maintain their internal milieu via auto-phagocytosis. This in turn delayed the attainment of a senescent phenotype and increased the tumor cells’ ability to proliferate, survive, and form subcutaneous xenografts.
The importance of autophagy in mediating circ-Dnmt1’s effects was confirmed by autophagy inhibitors. In the presence of the autophagy inhibitors, the functions of circ-Dnmt1 on cell senescence and survival were completely abolished. However, it is not clear whether autophagy can regulate circular RNA biogenesis, translocation, and degradation. This awaits further investigation.
Dnmt1 is an enzyme encoded by the DNMT1 gene that plays a role in the regulation of tissue-specific methylation of cytosine residues [28]. Aberrant methylation of Dnmt1 is associated with development of abnormalities and human tumors [29]. It has been reported that Dnmt1 bound to the p53 regulatory motif and loss of Dnmt1 causes derepression of p53 expression [30]. Our results showed that expression of circ-Dnmt1 increased Dnmt1 expression resulting in the inhibition of p53 transcription. This appears to be the central role of circ-Dnmt1 in enhancing cell autophagy, decreasing cellular senescence, and promoting cell proliferation, survival, and tumor growth.
The tumor suppressor p53 has many tumor suppressive mechanisms contributing to apoptosis, genomic stability, and angiogenesis. Our results showed that downregulation of p53 via ectopic circ-Dnmt1 expression increased cellular autophagy, proliferation, survival, and tumor growth, and upregulation of p53 by silencing circ-Dnmt1 produced opposite effects. Circ-Dnmt1 could bind to p53 and induced nuclear translocation. It has been reported that cytoplasmic p53 represses autophagy, while nuclear p53 enhances autophagy [31, 32]. Our results also showed that nuclear translocation of p53 appeared to be key in mediating circ-Dnmt1’s role in autophagy. p53 possesses RNA-binding activity and has been shown to control mRNA translation that occurs in cytoplasm [33]. The precise nature of the interaction between p53 and circ-Dnmt1 awaits further investigation, however we hypothesize that it may be facilitated by Auf1. Whether the p53 downstream molecule PUMA is involved in mediating the effects of circ-Dnmt1 may be worth of further investigation.
Auf1 is an AU-rich RNA-binding protein encoded by the HNRNPD gene [34]. Alternative splicing produces four transcript variants. The hnRNPs proteins interact with pre-mRNAs in the nucleus where they affect pre-mRNA processing, metabolism, and translocation. While most of the hnRNPs are present in nucleus, a fraction will shuttle between cytoplasm and nucleus. Auf1 variations modulate DNA methylation by regulating stability of Dnmt1 mRNA [35]. The Auf1 protein has been reported to bind Dnmt1 mRNA 3′-UTR and promote Dnmt1 mRNA degradation [36]. It appears that nuclear translocation of Auf1, induced by circ-Dnmt1 binding, increases Dnmt1 mRNA stability and translation. This ultimately inhibits p53 transcription and promotes autophagy.
To validate the interaction of circ-Dnmt1 with p53 and Auf1, we employed computational algorithms that included scoring, clustering, docking, and further optimized the most promising models. The computational algorithms allowed us to elucidate the structure and functions of different RNA molecules, and also confirmed binding interaction between circular RNA and associated proteins [37]. In order to analyze the possible docking of circ-Dnmt1 with p53, 20,000 models were generated using the NPDock server, a protein–RNA docking analysis tool [38]. NPDock server combines GRAMM for global macromolecular docking, scoring, clustering and refinement of the highest scored docked complexes from the biggest clusters.
It should be noted that circular RNAs differ from linear ones in such a way that their 3′- and 5′-ends are joined together forming a covalently closed continuous loop [39]. It has been proposed that circular RNAs could adopt three-dimensional structures that are different from related linear molecules of the same sequence [2]. Consequently new protein binding sites could be generated by the sequences that are brought into closer proximity during circularization. Indeed, we identified a p53 binding site covering the junction sequence of circ-Dnmt1. In addition, the three-dimensional structure of circ-Dnmt1 was different from the Dnmt1 linear mRNA, and produced a binding site for Auf1. In addition to their canonical signaling, it is conceivable that RNA molecules also utilize three-dimensional structure and conformation as a regulatory strategy. It is thus anticipated that circular RNAs could impact gene expression using a separate mechanism from mRNAs of the same origin. The emergence of circular RNAs expressed by mammalian cells may shed further light on understanding gene expression, with relevance toward various biological paradigms.
Materials and methods
Constructs and primers
A construct expressing human circular RNA Dnmt1 (circ-Dnmt1, circ-Dnmt1.10 isoform) was generated using the plasmid described by us previously [40]. The plasmids contained a CMV promoter driving circ-Dnmt1 or a non-related sequence serving as a control. The green fluorescent protein (GFP) expression unit was driven by a separate CMV promoter. All primer sequences used are listed in Fig S11g. The anti-circ-Dnmt1 siRNA and oligo probes labeled with biotin or Cy5 were purchased from Integrated DNA Technologies.
Nuclear extract preparation
Cells were lysed in nuclear extraction buffer (1 ml containing 20 mM Hepes, pH 7.2, 2 mM MgCl2, 10 mM KCl, and PI). It was homogenized with a prechilled Dounce homogenizer with 20 strokes. The lysis was subject to centrifugation at 4200 rpm for 5 min. The supernatant was removed and the pellet was washed three times with PBS. The pellet was then resuspended in 100 μl lysis buffer containing 0.5 M NaCl. After centrifugation at 13,200 rpm for 10 min, the supernatants were used for analysis.
Tumor formation assay
Tumorigenesis was performed as described [41, 42]. All mouse assays were performed following the guideline approved by the Animal Care Committee at Sunnybrook Research Institute. The Protocol Number is AUP 15–076. Mice were kept in the Animal Core Facility at Sunnybrook Research Institute for 1 week before use. Four-week-old CD-1 female nude mice were injected subcutaneously with MCF-7 cells (4 × 107 cells in 200 µl 50% Matrigel) and injected subcutaneously with 50 µg β-estradiol 17-cypionate (in 50 µl sesame oil) every 3 days. Twenty days after tumor implantation, the mice were sacrificed and the tumors were harvested, followed by immunohistochemical staining as described [43].
In mouse survival assay, 4-week-old CD-1 nude mice were divided into six groups randomly including the control, siRNA, blocking, LY294002, EBSS, and EBSS + siRNA (10 mice in each). The mice were intraperitoneally injected with MB-231 cells (2 × 106 cells/mouse), which allowed up-take of ectopic plasmids by the tumor cells. EBSS and EBSS + siRNA groups were injected with cells cultured in medium containing EBSS for 24 h before injection. Next day after cell injection, circ-Dnmt1 siRNA, blocking oligo, or control oligo were conjugated with PEG-AuNP complexes. The complexes were injected at a dose of 5 µg/mouse intraperitoneally. The assay was repeated every 3 days when the assay was completed. In the control group, the mice were injected with 0.9% NaCl. LY294002 (100 mg/kg) was dissolved in 0.9% NaCl. Formulation of the complexes (oligo-PEG-AuNP) was conducted as previously described [15]. Briefly, 40 μg oligo was dissolved in a volume of 800 μl RNase-free H2O2. The mPEGSH (PG1-TH-2k, Nanocs) was mixed with the oligo at a ratio of 1:20. 10 nm gold nanoparticles (AuNP, Cytodiagnostics) were used to mix with the oligo (1:20). The mixture was gently shaken at 60 °C for 30 min before it was transferred into a syringe, followed by the injection.
Identification of binding site, docking simulations, and contact maps
We used the protein–RNA docking tool NPDOCK Server [38] to analyze 20,000 models of the Protein–RNA interaction. This resulted in determination of the interaction between circ-Dnmt1 and p53 or Auf1. NPDock Server combines GRAMM for molecular docking, scoring, and clustering. Residue contact maps of circ-Dnmt1-p53 and circ-Dnmt1-Auf1 complexes were determined using RNAmap2D [44]. The contact distances (between Cα atoms of protein residues and O5′ atoms of RNA strands) were tested. Two residues were placed in contact when their O5′–Cα distance is smaller than 10 Å. The distance-based approach was employed to identify the binding site for RNA–protein complex. Two atoms (one in RNA and the other in protein) were considered to be docking with each other if their distance was <3.5 Å.
General molecular and cell biology methods
Western blotting and immunohistochemistry were conducted as described [45, 46]. RNA–protein interaction assays were performed as described [18, 47]. Real-time PCR was performed as described [41, 48].
Statistical analysis
All assays were conducted in triplicate and the results were subjected to independent sample test (t-test). The values of significance were set at *p < 0.05 and **p < 0.01. Error bars means SD (n = 4 if no indicated).
References
Enuka Y, Lauriola M, Feldman ME, Sas-Chen A, Ulitsky I, Yarden Y. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 2016;44:1370–83.
Jeck WR, Sharpless NE. Detecting and characterizing circular RNAs. Nat Biotechnol. 2014;32:453–61.
Liu YC, Li JR, Sun CH, Andrews E, Chao RF, Lin FM, et al. CircNet: a database of circular RNAs derived from transcriptome sequencing data. Nucleic Acids Res. 2016;44:D209–215.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–8.
Zheng LL, Li JH, Wu J, Sun WJ, Liu S, Wang ZL, et al. deepBasev2.0: identification, expression, evolution and function of small RNAs, LncRNAs and circular RNAs from deep-sequencing data. Nucleic Acids Res. 2016;44:D196–202.
Zhang Y, Zhang XO, Chen T, Xiang JF, Yin QF, Xing YH, et al. Circular intronic long noncoding RNAs. Mol Cell. 2013;51:792–806.
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256–64.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–57.
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE. 2012;7:e30733.
Panda AC, De S, Grammatikakis I, Munk R, Yang X, Piao Y, et al. High-purity circular RNA isolation method (RPAD) reveals vast collection of intronic circRNAs. Nucleic Acids Res. 2017;45:e116.
Bai Y, Zhang Y, Han B, Yang L, Chen X, Huang R, et al. Circular RNA DLGAP4 ameliorates ischemic stroke outcomes by targeting miR-143 to regulate endothelial-mesenchymal transition associated with blood-brain barrier integrity. J Neurosci. 2017;38:32–50.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–8.
Huang R, Zhang Y, Han B, Bai Y, Zhou R, Gan G, et al. Circular RNA HIPK2 regulates astrocyte activation via cooperation of autophagy and ER stress by targeting MIR124-2HG. Autophagy. 2017;13:1722–41.
Hsiao KY, Lin YC, Gupta SK, Chang N, Yen L, Sun HS, et al. Noncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res. 2017;77:2339–50.
Yang W, Du WW, Li X, Yee AJ, Yang BB. Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene. 2016;35:3919–31.
Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F et al. Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. J Natl Cancer Inst. 2018;110, https://doi.org/10.1093/jnci/djx166.
Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell. 1993;73:1019–30.
Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016;44:2846–58.
Du WW, Fang L, Yang W, Wu N, Awan FM, Yang Z, et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 2017;24:357–70.
Du WW, Yang W, Chen Y, Wu ZK, Foster FS, Yang Z, et al. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J. 2017;38:1402–12.
Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z, Sharpless NE. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet. 2010;6:e1001233.
Hansen TB, Kjems J, Damgaard CK. Circular RNA and miR-7 in cancer. Cancer Res. 2013;73:5609–12.
Yoshida Y, Izumi H, Torigoe T, Ishiguchi H, Yoshida T, Itoh H, et al. Binding of RNA to p53 regulates its oligomerization and DNA-binding activity. Oncogene. 2004;23:4371–9.
Nagata T, Kurihara Y, Matsuda G, Saeki J, Kohno T, Yanagida Y, et al. Structure and interactions with RNA of the N-terminal UUAG-specific RNA-binding domain of hnRNP D0. J Mol Biol. 1999;287:221–37.
Katahira M, Miyanoiri Y, Enokizono Y, Matsuda G, Nagata T, Ishikawa F, et al. Structure of the C-terminal RNA-binding domain of hnRNP D0 (AUF1), its interactions with RNA and DNA, and change in backbone dynamics upon complex formation with DNA. J Mol Biol. 2001;311:973–88.
Chen J, Sawyer N, Regan L. Protein-protein interactions: general trends in the relationship between binding affinity and interfacial buried surface area. Protein Sci. 2013;22:510–5.
Kastritis PL, Bonvin AM. On the binding affinity of macromolecular interactions: daring to ask why proteins interact. J R Soc Interface. 2013;10:20120835.
Yen RW, Vertino PM, Nelkin BD, Yu JJ, el-Deiry W, Cumaraswamy A, et al. Isolation and characterization of the cDNA encoding human DNA methyltransferase. Nucleic Acids Res. 1992;20:2287–91.
Klein CJ, Botuyan MV, Wu Y, Ward CJ, Nicholson GA, Hammans S, et al. Mutations in DNMT1 cause hereditary sensory neuropathy with dementia and hearing loss. Nat Genet. 2011;43:595–600.
Georgia S, Kanji M, Bhushan A. DNMT1 represses p53 to maintain progenitor cell survival during pancreatic organogenesis. Genes Dev. 2013;27:372–7.
Tasdemir E, Chiara Maiuri M, Morselli E, Criollo A, D’Amelio M, Djavaheri-Mergny M, et al. A dual role of p53 in the control of autophagy. Autophagy. 2008;4:810–4.
Tasdemir E, Maiuri MC, Galluzzi L, Vitale I, Djavaheri-Mergny M, D’Amelio M, et al. Regulation of autophagy by cytoplasmic p53. Nat Cell Biol. 2008;10:676–87.
Miller SJ, Suthiphongchai T, Zambetti GP, Ewen ME. p53 binds selectively to the 5′ untranslated region of cdk4, an RNA element necessary and sufficient for transforming growth factor beta- and p53-mediated translational inhibition of cdk4. Mol Cell Biol. 2000;20:8420–31.
Dempsey LA, Li MJ, DePace A, Bray-Ward P, Maizels N. The human HNRPD locus maps to 4q21 and encodes a highly conserved protein. Genomics. 1998;49:378–84.
Torrisani J, Unterberger A, Tendulkar SR, Shikimi K, Szyf M. AUF1 cell cycle variations define genomic DNA methylation by regulation of DNMT1 mRNA stability. Mol Cell Biol. 2007;27:395–410.
Gratacos FM, Brewer G. The role of AUF1 in regulated mRNA decay. Wiley Interdiscip Rev RNA. 2010;1:457–73.
Washietl S, Will S, Hendrix DA, Goff LA, Rinn JL, Berger B, et al. Computational analysis of noncoding RNAs. Wiley Interdiscip Rev RNA. 2012;3:759–78.
Tuszynska I, Magnus M, Jonak K, Dawson W, Bujnicki JM. NPDock: a web server for protein-nucleic acid docking. Nucleic Acids Res. 2015;43:W425–430.
Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T, et al. Correlation of circular RNA abundance with proliferation--exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep. 2015;5:8057.
Yang Q, Du WW, Wu N, Yang W, Awan FM, Fang L, et al. A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ. 2017;24:1609–20.
Rutnam ZJ, Du WW, Yang W, Yang X, Yang BB. The pseudogene TUSC2P promotes TUSC2 function by binding multiple microRNAs. Nat Commun. 2014;5:2914.
Yang X, Du WW, Li H, Liu F, Khorshidi A, Rutnam ZJ, et al. Both mature miR-17-5p and passenger strand miR-17-3p target TIMP3 and induce prostate tumor growth and invasion. Nucleic Acids Res. 2013;41:9688–704.
Shan SW, Lee DY, Deng Z, Shatseva T, Jeyapalan Z, Du WW, et al. MicroRNA MiR-17 retards tissue growth and represses fibronectin expression. Nat Cell Biol. 2009;11:1031–8.
Pietal MJ, Szostak N, Rother KM, Bujnicki JM. RNAmap2D—calculation, visualization and analysis of contact and distance maps for RNA and protein-RNA complex structures. BMC Bioinforma. 2012;13:333.
Du WW, Liu F, Shan SW, Ma XC, Gupta S, Jin T, et al. Inhibition of dexamethasone-induced fatty liver development by reducing miR-17-5p levels. Mol Ther. 2015;23:1222–33.
Yang BL, Cao L, Kiani C, Lee V, Zhang Y, Adams ME, et al. Tandem repeats are involved in G1 domain inhibition of versican expression and secretion and the G3 domain enhances glycosaminoglycan modification and product secretion via the complement-binding protein-like motif. J Biol Chem. 2000;275:21255–61.
Zeng Y, Du WW, Wu Y, Yang Z, Awan FM, Li X, et al. A circular RNA binds to and activates AKT phosphorylation and nuclear localization reducing apoptosis and enhancing cardiac repair. Theranostics. 2017;7:3842–55.
Shan SW, Fang L, Shatseva T, Rutnam ZJ, Yang X, Du W, et al. Mature miR-17-5p and passenger miR-17-3p induce hepatocellular carcinoma by targeting PTEN, GalNT7 and vimentin in different signal pathways. J Cell Sci. 2013;126:1517–30.
Acknowledgements
This work was supported by grants from Canadian Institutes of Health Research (PJT-153105), National Natural Science Foundation of Guangdong (2017A030310088), and Innovation and Entrepreneurship Leading Talent of Guangzhou Development Zone (2017-L181).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
These authors contributed equally: William W. Du, Weining Yang, Xiangmin Li, Faryal Mehwish Awan, Zhenguo Yang.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Du, W.W., Yang, W., Li, X. et al. A circular RNA circ-DNMT1 enhances breast cancer progression by activating autophagy. Oncogene 37, 5829–5842 (2018). https://doi.org/10.1038/s41388-018-0369-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0369-y
- Springer Nature Limited
This article is cited by
-
RUNX1, FUS, and ELAVL1-induced circPTPN22 promote gastric cancer cell proliferation, migration, and invasion through miR-6788-5p/PAK1 axis-mediated autophagy
Cellular & Molecular Biology Letters (2024)
-
Engineering circular RNA for molecular and metabolic reprogramming
Functional & Integrative Genomics (2024)
-
Role of circRNAs in regulating cell death in cancer: a comprehensive review
Cell Biochemistry and Biophysics (2024)
-
Circular RNA circHMCU promotes breast tumorigenesis through miR-4458/PGK1 regulatory cascade
Hereditas (2023)
-
Circular RNAs in renal cell carcinoma: from mechanistic to clinical perspective
Cancer Cell International (2023)