Abstract
Remarkable advances have been made in schizophrenia (SCZ) GWAS, but gleaning biological insight from these loci is challenging. Genetic influences on gene expression (e.g., eQTLs) are cell type-specific, but most studies that attempt to clarify GWAS loci’s influence on gene expression have employed tissues with mixed cell compositions that can obscure cell-specific effects. Furthermore, enriched SCZ heritability in the fetal brain underscores the need to study the impact of SCZ risk loci in specific developing neurons. MGE-derived cortical interneurons (cINs) are consistently affected in SCZ brains and show enriched SCZ heritability in human fetal brains. We identified SCZ GWAS risk genes that are dysregulated in iPSC-derived homogeneous populations of developing SCZ cINs. These SCZ GWAS loci differential expression (DE) genes converge on the PKC pathway. Their disruption results in PKC hyperactivity in developing cINs, leading to arborization deficits. We show that the fine-mapped GWAS locus in the ATP2A2 gene of the PKC pathway harbors enhancer marks by ATACseq and ChIPseq, and regulates ATP2A2 expression. We also generated developing glutamatergic neurons (GNs), another population with enriched SCZ heritability, and confirmed their functionality after transplantation into the mouse brain. Then, we identified SCZ GWAS risk genes that are dysregulated in developing SCZ GNs. GN-specific SCZ GWAS loci DE genes converge on the ion transporter pathway, distinct from those for cINs. Disruption of the pathway gene CACNA1D resulted in deficits of Ca2+ currents in developing GNs, suggesting compromised neuronal function by GWAS loci pathway deficits during development. This study allows us to identify cell type-specific and developmental stage-specific mechanisms of SCZ risk gene function, and may aid in identifying mechanism-based novel therapeutic targets.
Similar content being viewed by others
Introduction
SCZ is a highly heritable [1] neurodevelopmental disorder [2] that is characterized by positive symptoms (e.g., hallucinations and delusions), negative symptoms (e.g., apathy, anhedonia), and cognitive symptoms (e.g., memory, executive functions and attention) [3]. Recent large-scale SCZ GWAS have convincingly identified more than 100 risk loci with genome-wide significance [4], opening the way to a deeper understanding of SCZ pathogenic mechanisms by enabling the identification of potentially causal abnormalities encoded in the genome, rather than by-products of the disease pathogenetic cascade. However, gleaning biological insights on how these risk loci affect SCZ risk and pathogenesis is a nontrivial task, since they are often in non-coding regions. Many risk loci are enriched in enhancer regions [4], suggesting their role in gene regulation. Genetic influence on expression (e.g., eQTLs) is cell type-specific and sometimes has opposite effects in different cell types [5, 6], pointing to the importance of using disease-relevant and specific cell populations to clearly understand SCZ risk loci function. The postmortem brain tissues often used to derive eQTLs are heterogeneous cell populations which are likely to mask cell type-specific impacts of genetic loci, and can be confounded by effects from age, illness and treatment. Furthermore, genomic data sets, based on the enrichment of SCZ risk loci within active chromatin regions [7,8,9,10], suggest enriched SCZ heritability in fetal brains, stressing the need to study developing cells to understand how SCZ risk loci influence gene expression and alter neuronal function to increase SCZ risk. In addition, considering the modest effects of each SCZ GWAS locus, it is imperative to study the pathway that they converge onto to gain more meaningful biological insights into their function.
MGE-type cINs are one of the most substantially affected neural types in SCZ [11,12,13,14,15,16,17,18,19,20,21,22]. Accordingly, experimental evidence suggests a role for altered GABAergic neurotransmission in SCZ [23]. In particular, PV-expressing cINs regulate gamma oscillation [24, 25], which is important for cognitive function [26] but has been found to be defective in SCZ patients [27]. In addition, GNs are also significantly affected in SCZ, with evidence of altered glutamatergic neurotransmission [28]. Recent genomic analyses showed both cINs and GNs harbor enriched SCZ heritability in fetal brains [29,30,31,32], unlike other neuronal subtypes without such enrichments. Thus, to understand the functions of SCZ risk loci, it is imperative to study converging SCZ genetic influence on gene expression in these cell populations during the developmental stage where SCZ heritability is enriched.
The ability to study the critical pathogenic events in the developing human nervous system once seemed unimaginable due to insurmountable ethical and technical barriers. However, recent advances in iPSC technologies offer the possibility of generating SCZ-specific developmental brain cell populations with the same genetic makeup as the patient brains, providing the opportunity to analyze cellular and molecular abnormalities resulting from the SCZ genetic burden [33,34,35,36,37]. Differentiation of human PSCs well recapitulates human embryonic development [31, 38,39,40,41,42,43,44,45,46,47,48,49] and follows a relevant in vivo developmental timeline [50, 51], enabling us to study early developmental processes of neurons in the gestational stage. This is a critical time period, during which they actively integrate into cortical circuitry [52], the brain becomes more susceptible to environmental challenges [53,54,55], and many known SCZ risk genes are highly expressed [56,57,58]. Thus, to gain a deeper understanding of SCZ GWAS risk function during development, we have generated homogeneous populations of developing MGE-type cINs and GNs from human iPSCs. Using well-verified healthy control (HC) vs SCZ cINs [49,50,51, 59,60,61,62], we observed SCZ GWAS loci genes that are dysregulated in developing SCZ cINs converge onto the PKC pathway. Dysregulation of PKC pathway genes resulted in PKC hyperactivity, accompanied by deficits in arborization. Fine-mapped GWAS locus in the ATP2A2 gene of the PKC pathway harbors enhancer marks shown by ATAC-seq and ChIP-seq and regulates ATP2A2 expression. We also generated developing glutamatergic neurons (GNs), another population with enriched SCZ heritability, and confirmed their functionality after transplantation into the mouse brain. Then, we identified SCZ GWAS risk genes that are dysregulated in developing SCZ GNs. Distinct from cINs, GN-specific SCZ GWAS loci DE genes converge onto the ion channel pathway, stressing the need for cell type-specific studies to unravel SCZ GWAS loci function. Disruption of CACNA1D within GWAS loci resulted in reduced Ca2+ currents in these neurons, suggesting neuronal functional deficits by dysregulation of SCZ GWAS loci DE genes. Understanding cell type-specific and developmental stage-specific mechanisms of SCZ risk loci function will aid in identifying novel therapeutic targets based on the mechanistic insight of disease pathogenesis.
Results
SCZ DE genes within SCZ GWAS loci converge onto PKC pathway in developing cINs
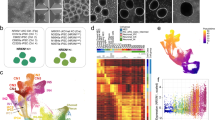
Recent large scale SCZ GWAS convincingly identified a large number of SCZ risk loci. However, which loci are operational in which cells, at what developmental stages and with what impact, still needs to be determined. With the purpose of identifying SCZ risk loci that are active in specific cell types with enriched SCZ heritability during development, first we identified SCZ GWAS loci genes that are dysregulated in homogeneous populations of SCZ developing cINs. Based on previously published RNA-seq analysis [49] on HC vs SCZ developing cINs (Fig. 1A, B and Supplementary Fig. 1), we identified 25 SCZ cIN DE genes within SCZ GWAS loci with genome-wide significance [63] (Fig. 1C and Supplementary Table 1). Analysis of convergence of these SCZ GWAS loci DE genes identified the PKC pathway as the most significantly enriched (Fig. 1C–E, Supplementary Fig. 2 and Supplementary Table 2), followed by RNA processing. All 6 PKC pathway DE genes (marked with red asterisks in Fig. 1C and in red rectangles in Fig. 1D) displayed a direction of change (Fig. 1E, F and [49] for clustered PCDH family members) that is expected to increase PKC activity in SCZ cINs by directly regulating levels of PKC activators, Ca2+ and DAG [64, 65], or via signaling cascades [49, 66, 67] as illustrated in Fig. 1E. Interestingly, PKC hyperactivity has been associated with numerous brain abnormalities [68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86]. We also confirmed the dysregulation of these genes in SCZ cINs by qPCR (Fig. 1G). Each of the 6 PKC genes have 3–7 independent studies that identify them as high confidence SCZ risk genes by various genomic analyses such as fine mapping, eQTL, colocalization, TWAS, ATAC-seq and Hi-C [6, 10, 47, 87,88,89,90,91,92,93,94] as summarized in Supplementary Table 3, giving us confidence to study the pathway on which they converge. Overall, these results suggest convergence of SCZ GWAS loci DE genes onto the PKC pathway in developing cINs.
A Scheme for analyzing SCZ GWAS loci DE genes in developing cINs. B Table of subjects analyzed in RNA-seq study and scheme for generating developing cINs from human iPSCs. HC refers to healthy control subjects and SCZ refers to people with SCZ. SRM: serum replacement media, LDN: 100 nM LDN193189, SB: 10 μM SB431542, SAG: 0.1 μM Smoothened agonist, and IWP2: 5 μM Inhibitor of Wnt production-2. C SCZ GWAS loci genes that are differentially expressed in SCZ developing cINs. Genes that can modulate PKC activity was marked using *. D Pathway analysis of SCZ GWAS loci DE genes in cINs. Left panel shows the % gene numbers among total SCZ risk loci DE genes and right panel shows enrichment test using Fisher’s exact test in -logP format. E Schematic diagram showing the function of SCZ risk loci DE genes in modulating PKC activity. F RNA-seq analysis of HC vs SCZ cINs, showing dysregulation of PKC SCZ risk loci genes. Gene expression is shown as TPM, obtained from Kallisto. Differentially expressed genes were analyzed by DESeq2 (Wald test for two-sided significance testing, n = 28 independent differentiations from 14 HC lines and 14 SCZ lines). Error bars are SEM. G qPCR analysis of PKC SCZ risk loci genes in cINs. Data were normalized by GAPDH expression and are presented as mean ± SEM. Two-tailed unpaired t-tests were used for analysis (N = 14 lines for HC and n = 14 lines for SCZ; Each data point is averaged from 2 independent differentiations, ATP2A2: t = 4.279, df = 26, DGKi: t = 2.316, df = 26, RGS6: t = 2.149, df = 26).
Functional impact of PKC pathway SCZ risk loci DE gene dysregulation on developing cINs
To understand the functional impact of PKC pathway DE gene dysregulation, we knocked out ATP2A2 or DGKI using the CRISPR/CAS9 system (Fig. 2A), since these 2 genes more directly regulate PKC activator level [64, 65]. Analysis of PKC activity in control vs knockout (KO) cINs derived from 6 different subject iPSCs showed that there was significant increase in PKC activity by KO of ATP2A2 or DGKI alone or together (Fig. 2B and Supplementary Fig. 3A). Though the directions of KO impact were same in different lines, the baseline PKC activity and fold increases were different in each line (Supplementary Fig. 3B), showing genetic background dependence of KO impacts. We did not see additive effects by KO of both genes, possibly due to crosstalk between Ca2+ pathway and DAG pathway, which have been shown to activate each other [95,96,97,98]. Since PKC hyperactivity was correlated with various neuronal deficits including arborization deficits [68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86], we examined whether downregulation of PKC pathway SCZ risk loci DE genes can result in compromised arborization of developing cINs. KO of either ATP2A2 or DGKI alone or together significantly reduced neurite length (Fig. 2C and Supplementary Fig. 4A, B). Again, though the directions of KO impact were the same across the different lines, baseline neurite lengths and fold changes were different in each line (Supplementary Fig. 4C). Next we tested whether the arborization deficit resulting from KO of PKC DE genes can be rescued by inhibition of PKC. Thus we treated developing cINs with ATP2A2 and DGKI KO with Go6893, a pan-PKC inhibitor. Treatment of KO cINs with PKC inhibitor restored arborization deficits, showing the deficits were mediated by hyperactive PKC (Fig. 2C). Interestingly, in our previous study, we observed a significant decrease in neurite length in SCZ cINs compared to HC cINs, which was reverted by PKC inhibitor treatment [49]. Overall, these results suggest that dysregulated SCZ risk loci DE genes could result in functional deficits in developing cINs.
A KO of ATP2A2, DGKI or both analyzed by qPCR. Data were normalized by GAPDH expression and are presented as mean ± SEM. Repeated measures One-way ANOVA was used for analysis (N = 3 batches), followed by Dunnett’s multiple comparisons as a post hoc analysis. B Western blot analysis of control, ATP2A2 KO, DGKI KO or both KO cINs. Repeated measures one-way ANOVA was used for analysis (N = 6 lines), followed by Dunnett’s multiple comparison as a post hoc analysis. C Arborization analysis of control, ATP2A2 KO, DGKI KO, both KO or both KO + Go6983-treated cINs. Left panel: Representative images of arborization analysis of cINs with or without KO (Scale bar = 50 μm). Right panel: quantification of neurite lengths as analyzed using Image J with the Neuron J plugin. Center and error bars show mean ± SEM. Data were collected from 6 lines, each line with 10 neurons (N = 60 neurons). Two-Level Hierarchical Linear Mixed Effect Model after log-transformation was used for analysis of neurite length, followed by Dunnett’s test as a post hoc analysis.
SCZ GWAS loci regulate the expression of ATP2A2 in developing cINs
Next we analyzed the role of SCZ GWAS loci in regulating gene expression levels. To do this, we first examined fine mapping results from recent large SCZ GWAS. Among fine-mapped loci near PKC DE genes, a single SNP in the ATP2A2 gene intron rs4766428 has an almost perfect probability to be causal [probability of 1 [63] or 0.99 [99]], whereas other PKC GWAS loci have multiple fine mapped SNPs, each with moderate probability (Fig. 3A and Supplementary Table. 4). The SCZ GWAS p-value for this SNP was p = 2.61e−17 [99]. rs4766428 was identified as an eQTL for ATP2A2 in postmortem hippocampus (Fig. 3A and Supplementary Table. 4). ATAC-seq showed peaks around rs4766428 in developing cINs but not developing GNs (Fig. 3B and Supplementary Table 5), suggesting it is located in an active open chromatin region specifically in cINs but not in GNs. As a reference, when compared with GNs and cINs in human fetal brains [30], our ATAC-seq peaks from iPSC-derived cINs showed a significantly higher level of overlap with fetal cIN enhancers than fetal excitatory neuronal enhancers (Fisher’s exact test, p-value < 2.2e−16, OR = 1.5, 95% CI = 1.47–1.53), demonstrating their in vivo relevance. We further analyzed enhancer marks around this SNP by ChIP-seq and observed H3K4me1 peaks and H3K27Ac peaks at this locus (Fig. 3C), suggesting that this region may be involved in gene regulation. Consistent with this observation, Haploreg [100] search showed many enhancer epigenetic marks in rs4766428 in many different neural tissues as summarized in Supplementary Table 6. To test the impact of this locus in regulating ATP2A2 expression, we used CRISPRa and CRISPRi assays [101, 102] (Fig. 3D). When infected along with ATP2A2 risk loci gRNA, CRISPRa significantly increased expression level of ATP2A2, whereas CRISPRi along with ATP2A2 risk loci gRNA significantly reduced ATP2A2 expression (Fig. 3D). These results suggest this fine-mapped locus in the second intron of the ATP2A2 gene is involved in the regulation of ATP2A2 expression.
A ATP2A2 loci with fine mapped SCZ GWAS SNP marked by red dotted lines. GWAS p-value in -logP, fine mapping probability and eQTL p-value in -logP of this fine mapped SNP were marked by red dots. B ATAC-seq in loci surrounding fine mapped SCZ GWAS SNP in cINs shows open chromatin area compared to GNs. C ChIP-seq analysis in loci surrounding fine mapped SCZ GWAS SNP shows enhancer marks (overlap of H3K27ac peaks and H3K4me1 peaks). D CRISPRa and CRISPRi from SCZ risk loci modulate the expression of ATP2A2, as analyzed by qPCR. Data were normalized by GAPDH expression and are presented as mean±SEM. Two-tailed paired t-test was used for analysis (n = 8 lines; Each data point is averaged from 3 independent differentiations).
SCZ DE genes within SCZ GWAS loci converge onto Ion transporter pathway in developing GNs
Since SCZ heritability is enriched both in GNs [30, 32] and cINs [29,30,31] during fetal development, we also sought to find converging SCZ GWAS loci DE gene pathways in developing GNs. To do this, we did RNA-seq analysis on HC vs SCZ developing GNs (Fig. 4A, B, Supplementary Figs. 5–8 and Supplementary Table 7). Immunocytochemistry and cell counting analyses demonstrated the highly homogeneous phenotype of generated GNs, as shown by high expression of relevant genes and low expression of non-relevant markers (Supplementary Fig. 5). To further confirm functionalities of generated GNs, we transplanted ChR2-GFP-expressing GNs into the prefrontal cortex of Nod Scid Gamma (NSG) mice and analyzed them by electrophysiology. Whole-cell patch-clamp recordings were used to examine neuronal and synaptic properties of grafted human GNs. Nine months after transplantation, grafted cells were identified with green fluorescence in acute brain slices (GFP+ cells). Grafted cells displayed passive membrane properties of glutamatergic neurons [resting membrane potential (RMP) = −75.1 ± 5.1 mV; membrane resistance (Rm) = 0.272 ± 0.045 GΩ; membrane capacitance (Cm) = 109 ± 17 pF; average ± SEM; n = 6 cells; Supplementary Fig. 6A]. When depolarizing voltage pulses were applied in voltage-clamp mode, all 7 grafted cells showed rapidly desensitizing inward currents and sustained outward currents (Supplementary Fig. 6B, C). This suggests the expression of voltage-gated Na+ channels and K+ channels in grafted cells. In current-clamp mode, the injection of depolarizing currents induced action potential (AP) firings in all recorded grafted cells with the AP threshold of −49.9 ± 1.5 mV and the afterhyperpolarization (AHP) of 15.8 ± 2.6 mV (6 cells; Supplementary Fig. 6D, E). Moreover, all 6 grafted cells displayed spontaneous excitatory postsynaptic currents (sEPSC) at −85 mV in voltage-clamp mode (Fig. 4C top left panel), indicating that they received excitatory inputs from host glutamatergic neurons. Consistently, grafted human GNs contained dendritic spines, postsynaptic sites of excitatory synapses (Fig. 4C top right panel). We then applied blue light illumination to activate ChR2-GFP-expressing grafted cells for glutamate release, which induced excitatory synaptic responses in GFP– host neurons in the PFC (Fig. 4C bottom panels). Short pulses of blue light illumination induced excitatory postsynaptic currents (EPSC) in 3 out of 4 host pyramidal neurons at −85 mV in voltage-clamp mode. These EPSCs were inhibited by NBQX, an AMPA receptor antagonist, indicating that they were mediated by excitatory neurotransmitter glutamate. Together, these results suggest neuronal properties of human glutamatergic neurons transplanted into the mouse PFC as well as functional synaptic integration of grafted cells into host PFC circuits. After the confirmation of their functionalities, we proceeded to RNA-seq analysis using generated GNs. RNA-seq analysis also confirmed high expression of relevant markers and low expression of non-relevant markers (Supplementary Fig. 8A). Principal component analysis of RNA-seq data showed developing GNs are well separated from developing cINs or iPSCs or fibroblasts, but there was no clear separation between HC GNs and SCZ GNs, as expected from overall mild SCZ phenotypes (Supplementary Fig. 7A). Pair-wise comparison of RNA-seq samples showed high similarity among samples with r2 > 0.96, suggesting reliable generation of homogeneous populations of developing GNs (Supplementary Fig. 7B). HC GNs and SCZ GNs were highly similar in the pairwise comparison (Supplementary Fig. 7B), again confirming the overall mild phenotype of SCZ GNs. We identified 29 SCZ glutamatergic neuron DE genes within SCZ GWAS loci (Fig. 4D, Supplementary Fig. 7C and Supplementary Table 7). Pathway analysis of these SCZ GWAS loci DE genes using g:Profiler showed that they are enriched for ion channel pathway genes (Fig. 4D, E). Fisher exact analysis p-value for ion channel pathway in GNs was 0.000104 (Supplementary Fig. 8B) whereas that for PKC pathway in GNs was 0.999999, displaying cell type-specific pathway enrichment. On the other hand, Fisher’s exact analysis p-value for ion channel pathway in cINs were 0.338145. Among the 6 genes associated with the ion channel pathway, NDUFA4L2 is a mitochondrial electron transport protein, whereas all the others are expressed in the cellular membrane [103, 104], with the possibility of regulating cellular membrane properties of neurons, leading us to focus on these 5 cellular membrane proteins. All 5 ion channel pathway DE genes were downregulated in SCZ developing GNs (Fig. 4F), which was also confirmed by qPCR using 7 HC vs 7 SCZ lines (Fig. 4G). All of SCZ GWAS loci DE genes in developing GNs were not dysregulated in developing cINs nor were SCZ GWAS loci DE genes in developing cINs dysregulated in developing GNs (Supplementary Fig. 9), suggesting the importance to study cell type-specific genetic influence on gene expression. By and large, these results show SCZ risk loci DE genes converge onto ion channel pathway in developing GNs, unlike in the case of developing cINs.
A Scheme for analyzing GCZ GWAS loci DE genes in developing glutamatergic neurons. B Table of subjects analyzed in RNA-seq study and scheme for generating developing glutamatergic neurons from human iPSCs. C Electrophysiological analysis of developing GNs. Top left panel: ChR2-GFP+ grafted cells receive synaptic inputs from host glutamatergic neurons as recorded in acute brain slices. A representative trace of postsynaptic responses recorded in a GFP+ grafted cell at −85 mV in voltage-clamp mode. sEPSC trace marked with a dotted square box was magnified in the bottom trace. Top right panel: Confocal microscopic images showing dendrites of ChR2-GFP+ grafted cells. A dotted area is magnified in the right image, showing dendritic spines. Bottom left panel: blue light illumination induced glutamate release at axon terminals, generating excitatory postsynaptic responses recorded in GFP− host neurons (inset). Bottom right panel, representative traces of excitatory postsynaptic currents (EPSC) evoked by photostimulation and recorded in a GFP− host neuron. Postsynaptic responses were recorded at −85 mV in voltage-clamp mode (gray and black traces) and induced by photostimulations (36.1 mW/mm2, 10 ms duration, blue horizontal line). These EPSCs were blocked inhibited by NBQX (10 μM) in the same neuron (blue traces). D SCZ GWAS loci genes that are differentially expressed in SCZ developing GNs. Genes in ion channel pathways was marked using *. E Pathway analysis of SCZ GWAS loci DE genes in GNs analyzed by g:Profiler [137]. F RNA-seq analysis of HC vs SCZ glutamatergic neurons, showing dysregulation of ion channel pathway genes within SCZ risk loci. Gene expression is shown as TPM, obtained from Kallisto. Differentially expressed genes were analyzed by DESeq2 (Wald test for two-sided significance testing, N = 4 HC lines and 5 SCZ lines). Error bars are SEM. G qPCR analysis of SCZ risk loci DE genes in GNs. Data were normalized by GAPDH expression and are presented as mean ± SEM. Two-tailed unpaired t-test was used for analysis (N = 7 lines).
Functional impact of Ion transporter pathway SCZ risk loci DE gene dysregulation in developing GNs
To understand the functional consequence of ion channel pathway dysregulation in developing GNs, we knocked out CACNA1D within the SCZ risk locus with the most significant SCZ associations among the 5 pathway genes (Fig. 5A and Supplementary Fig. 2C) and analyzed its impact on developing GNs by patch clamp analysis in vitro. KO of CACNA1D resulted in significant decrease of total Ca2+ currents in developing GNs (Fig. 5B–D and Supplementary Fig. 10D). Significant decrease of total Ca2+ currents was consistent when analyzed in HC lines and SCZ lines separately (Supplementary Fig. 10A) or analyzed using averages of each line (Supplementary Fig. 10C). We also observed significant decrease in high voltage-activated Ca2+ currents (Fig. 5D, E and Supplementary Fig. 10), consistent with those that are mediated L-type channels, which are located postsynaptically and involved in regulating neuronal excitability [105]. We also confirmed high voltage-activated Ca2+ current can be blocked with Nifedipine, a specific L-type Ca2+ channel blocker (Supplementary Fig. 10B). Considering the function of CACNA1D in regulating synaptic plasticity [106, 107], this result suggests suboptimal expression of ion channel genes may result in functional deficits in these neurons. In addition to CACNA1D, two additional GN DE genes identified within SCZ GWAS loci, P2RX4 and P2RX7, are both ATP-dependent cation channels and have preferences for Ca2+, suggesting the dysregulation of multiple genes that regulate Ca2+ transport in SCZ developing GNs. This may impact Ca2+influx in SCZ developing GNs compared to HC developing GNs. Patch clamp analysis showed that there is a significant decrease of total Ca2+ currents as well as high voltage-activated Ca2+ currents in SCZ developing GNs compared to HC GNs (Fig. 5E), which suggests functional deficits produced by converging ion channel pathway genes in SCZ GNs. Overall, this study provides novel mechanistic insights into how the influence of SCZ risk loci converges and alters early neurodevelopmental processes in human cINs and GNs, which were largely unknown previously.
A CACNA1D KO in developing GNs, analyzed by qPCR. Data were normalized by GAPDH expression and are presented as mean ± SEM. Repeated measures one-way ANOVA was used for analysis (N = 4 batches), followed by Dunnett’s multiple comparisons as a post hoc analysis. B Representative image of patched cINs. Scale bar = 10 μm. C Illustration of voltage scheme used to analyze total Ca2+ currents and high voltage-activated Ca2+ currents. D CACNA1D KO in developing GNs dysregulate total Ca2+ currents and high voltage-activated Ca2+ currents, as analyzed by patch clamp recording in vitro. Two-Level Hierarchical Linear Mixed Effect Model after log transformation was used for analysis (n = 47 cells from 6 Lines for Cas9 group and n = 42 cells from 6 lines). E SCZ GNs show compromised total Ca2+ currents and high voltage-activated Ca2+ currents, as analyzed by patch clamp recording in vitro. Two-tailed unpaired Kolmogorov–Smirnov test was used for analysis (n = 26 cells for HC and n = 21 cells for SCZ).
Discussion
In this study, we identified converging pathways of SCZ GWAS loci DE genes and observed distinct pathways are affected in cell type-specific manners. Most studies on genetic influences on gene expression have been done using postmortem brain tissues, that are not only confounded by effects from age, illness and treatment but are also composed of quite heterogeneous cell populations and thus unable to reveal the impacts on specific minority populations like cINs. Furthermore, considering genetic influence on gene expression is not only cell type-specific but also developmental stage-specific with enriched heritability in the fetal brain [7,8,9,10], the fetal tissue-specific influences may not be easily assessed. Recently, there have been studies using fetal brain transcriptomic data to understand genetic influences on expression [87, 91, 108], and this has allowed for identification of fetal stage-specific eQTLs. Still fetal brain tissue is not likely to provide insights into the mechanisms in minor, yet critical, populations like developing cINs. To address these gaps, we have utilized iPSC-derived homogeneous populations of developing cINs and GNs and identified SCZ GWAS loci genes that are dysregulated in these cells along with the pathways onto which they converge. This strategy gives the first insights on genetic influences in specific fetal cell populations with enriched SCZ heritability. Though eQTL analysis shows correlation between risk alleles and gene expression levels, it doesn’t allow us to determine which genes will be actually significantly dysregulated in disease genetic backgrounds. iPSC-derived fetal neural populations of HC compared to SCZ individuals provide such an opportunity, allowing for the analysis of the actual genes with sufficient effect size to show up as DE in disease samples during development. The observation that tissue-specific eQTLs have greater impact size than general eQTLs [91] also stresses the importance of using specific and homogeneous cell populations to identify these cell type-specific genetic influence with greater impact. Furthermore, these homogeneous developing cell populations also provide a disease-relevant model system where we can perturb the candidate gene expression to assess the functional consequences of disease-relevant gene perturbations.
In this study, we have used bulk analysis of homogeneous culture rather than single cell analysis approaches for cell type-specific studies. Single cell (sc) RNA-seq or scATAC-seq allow us to analyze specific cell types in the mixed cultures but it also has shortcomings that require further development down the road. scRNA-seq usually contains non-zero values for 10–50% expressed genes and low abundance genes can be under-represented [109, 110]. Also, recent studies showed confounder introduced by trypsinization during scRNA-seq sample preparation [111]. Similarly, scATAC-seq shows inherently sparse count matrices with each cell usually sampling only 2–3% peaks out of all peaks accessible in a given cell type [112, 113]. Although there are new bioinformatics methods to mitigate the issue of high noise and sparsity [114, 115], it still remains an area of active development. Thus, single cell analysis approaches and bulk analysis of homogeneous population, as employed in this study, can complement each other. Furthermore, monoculture system provides the opportunity to study cell-intrinsic regulation in the absence of any cell non-autonomous impact. Once cell-intrinsic regulations are identified, we can systematically add specific cell types as a co-culture to analyze specific non-cell autonomous regulations. In addition, in this study, we used the KO approach to address the question of whether the dysregulation of SCZ risk loci DE genes indeed affects functionality of developing cINs or developing GNs, with the purpose of obtaining clearer answers with a stronger impact. This KO approach is not exactly the same as in vivo situations where many genes with smaller impacts converge onto specific pathways; however, we were aiming to mimic functional consequences of converging pathway deficits from ‘the dysregulation of many genes with smaller impacts’ with pathway deficits from ‘the dysregulation of one gene with a larger impact’.
We observed that SCZ GWAS loci genes converge onto the PKC pathway in developing cINs. PKC is a family of serine/threonine kinases that are activated by DAG and Ca2+ and phosphorylate downstream substrate proteins [116]. In the brain, they regulate various neuronal functions such as arborization, neuronal excitability, neurotransmitter release, ion channels, growth and differentiation, apoptosis, and neuronal plasticity [117]. As expected from their critical roles in normal brain function, their dysregulation, especially their hyperactivity, has been associated with various neurodegenerative disorders [68,69,70,71], with some of the deficits reversed by PKC inhibitor treatment [83]. In addition to neurodegenerative disorders, PKC dysregulation has also been observed in psychiatric disorders. Increased PKC expression was reported in bipolar disorders (BD), which was normalized by medications that treat BD such as lithium and valproic acids [72]. Furthermore, PKC inhibitors were successfully used to treat acute mania in bipolar disorders in multiple small scale clinical trials [73,74,75]. PKC hyperactivity in the prefrontal cortex has been linked to aging- or stress-related working memory deficits, which were reversed by PKC inhibitor treatment [76,77,78]. These results point to the importance of maintaining proper PKC regulation for normal brain development and function. Various PKC isoforms were shown to regulate the actin cytoskeleton, [118] and thus, overactivation of PKCs results in spine loss and altered spine morphology in vitro [79]. In AD, increased phosphorylation was observed in PKC substrates [80], and abnormal spine formation in AD model mice was reversed by PKC inhibition [81]. PCDHG conditional knockout mice, which exhibit PKC hyperactivity, displayed increased PKC substrate phosphorylation accompanied by severely reduced arborization in cortical neurons, which is reversed by PKC inhibition [82]. PKC hyperactivity has also been shown to be associated with decreased length of basal dendrites of prefrontal cortical neurons and compromised working memory performance in aged rats. Both are reversed by PKC inhibitor treatment [77]. More recently, we have observed dysregulation of various PCDHA and PCDHG family members in SCZ cINs, accompanied by compromised arborization in these neurons that was reversed by PKC inhibitor GO6893 [49]. Recently, PKC activation was shown to reduce synaptic strength via cleavage of neuroligins [119], that hold the synaptic structure together. Taken together, these results all point to the role of a converging SCZ risk loci DE gene pathway in maintaining normal neuronal function.
On the other hand, SCZ GWAS loci DE genes converge onto ion channel pathways in developing GNs. CACNA1D is an L-type voltage-dependent Ca2+ channel that plays important roles in neuronal development, neuronal firing and synaptic plasticity [120] and thus its dysregulation was associated with various psychiatric disorders such as ASD, SCZ, BD and intellectual disability [120, 121]. CACNA1D KO results in decreased neuronal survival, decreased neurite outgrowth [122], impaired neurogenesis [123] and decreased firing frequency [124], depending on the cell types affected, resulting in cognitive deficits in these animals [122, 123]. Thus, it is conceivable that dysregulation of CACNA1D and thus defects in Ca2+ influx in GNs during development could result in dysfunctional neuronal development. In addition to CACNA1D, P2RX4 and P2RX7 are also involved in Ca2+ transport. P2RX7 is an ATP binding Ca2+ channel that is expressed at the presynaptic terminal, and is known to regulate synaptic transmission [125] and regulate axonal growth [126]. P2RX4 has also been shown to regulate Ca2+ influx [127, 128]. P2RX4 KO resulted in reduced long-term potentiation (LTP), indicating its role in synaptic plasticity [129]. Regulation of Ca2+ influx overall plays a critical role in neuronal development and function such as dendritic growth and dendritic spine formation [130], neuronal migration [131, 132], transcriptional regulation [133], synaptic plasticity [134] and neuronal survival/degeneration [135, 136]. Thus it is conceivable that suboptimal Ca2+ transport by converging SCZ GWAS loci DE genes in developing GNs would compromise their normal development and function.
In summary, we have shown converging influences of SCZ GWAS loci in specific developing cell types with enriched SCZ heritability. In the next phase of our study, this study can be expanded to other specific cell types with enriched SCZ heritability such as microglia [29]. Further understanding of tissue-specific, developmental stage-specific mechanism of SCZ GWAS loci function, either by bulk analysis of homogeneous cell types or single cell analysis approaches of mixed cells, or both, will aid the effort to identify novel therapeutic targets even preventive ones, based on neurobiological understanding of disease pathogenesis mechanism.
Material and methods
Differentiation of iPSCs
Generation of iPSCs were described in our previous publications [49, 59]. Male SCZ patient samples with age from 21–51 years old as well as age and gender-matched HC samples were used in this study as summarized in Figs. 1, 4, Supplementary Tables 7 and 8. Human fibroblasts were obtained from the laboratories of Dr. Bruce Cohen (McLean Hospital), Dr. Daniel Weinberger (Lieber Institute for Brain Development), and Dr. Judith Rapoport (National Institute of Mental Health). These study protocols were approved by the McLean Hospital/Partners Healthcare Institutional Review Board (IRB) and New York Medical College IRB. All procedures were performed in accordance with IRB’s guidelines and all human samples were obtained with informed consents. We have complied with all relevant ethical regulations. All non-McLean cohorts of iPSCs have been deposited to the NRGR biorepository. McLean cohorts of iPSCs will be available upon approval from the McLean Hospital/Partners Healthcare IRB. iPSC lines were validated by immunocytochemistry in our previous study [49]. All cell lines are routinely tested for mycoplasma contamination. Cell lines used in this study were verified to be mycoplasma negative before undertaking any experiment with them.
Thawed human iPSCs were maintained on Matrigel (BD, San Hose, CA) coated plates with Essential 8 (E8) media (Invitrogen, Carlsbad, CA). For differentiation, iPSCs were trypsinized and grown as floating spheres in low adherent flasks in KSR media (DMEM, 15% knockout serum replacement, 2 mM L-glutamine and 10 μM β-mercaptoethanol (all from Invitrogen) from day 0 to day 14. For neuroectoderm induction, cells were treated with LDN193189 (100 nM, Stemgent, Cambridge, MA) from day 0 to day 14 and SB431542 (10 μM, Tocris Cookson, Ellisville, MO) from day 0 to day 7. For MGE phenotype induction, media was supplemented with IWP2 (5 μM, EMD Millipore, Billerica, MA) from day 0 to day 7 and SAG (0.1 μM, EMD Millipore) from d0 to d21. From day 14, cells were grown in N2AA media (DMEM-F12 with N2-supplement (1:200, Invitrogen) and 200 μM ascorbic acid (AA, Sigma, St. Louis, MO). FGF8 (100 ng/ml, Peprotech, Rocky Hill, NJ) was added from day 14 to day 21 to induce the MGE phenotype at the expense of the CGE phenotype [50]. N2AA media was supplemented with 10 ng/ml glial cell derived neurotrophic factor (GDNF, Peprotech) and 10 ng/ml brain derived neurotrophic factor (BDNF, Peprotech) from day 21. After 8 weeks of differentiation, cIN spheres were trypsinized in the presence of 0.1M trehalose (Sigma) and then plated on polyornithine (PLO; 15 mg/ml; Sigma)- and fibronectin (FN; 1 mg/ml; Sigma)-coated plates in B27GB media (DMEM-F12 media with B27 supplement (1:100, Invitrogen), 10 ng/ml GDNF and 10 ng/ml BDNF).
For Glutamatergic neuron differentiation, iPSCs were trypsinized and grown as floating spheres in low adherent flasks in Neuronal induction media (DMEM-F12 with 15% knockout serum replacement, 1% MEM-NEAA, 100 µM β-mercaptoethanol (all from Invitrogen),100 nM LDN193189 (Stemgent), 10 µM SB431542 (Tocris Cookson) and 2 µM IWP2 (ApexBio)) from day 0 to day 10. From day 11, cells were grown in Neuronal differentiation media (DMEM-F12 with 0.25% N2, 0.5% MEM-NEAA, 50 µM β-mercaptoethanol (all from Invitrogen) and N21 MAX (R&D systems, Minneapolis, MN), until the spheres were trypsinized in the presence of 0.1M trehalose (Sigma) and plated on PLO/FN-coated plates in B27GB media after 8 weeks’ differentiation for analysis.
Subcloning to generate lentiviral vectors
Oligonucleotides to generate lentiviral (LV) vectors that express KO guide RNAs (gRNAs) as described in Supplementary Table 9 were synthesized by Thermo Fisher Scientific (Thermo Fisher, Waltham, MA, USA), and subcloned into lentiCRISPR v2 plasmid, a gift from Feng Zhang (Addgene plasmid # 52961), to obtain KO LV vectors (pLenti-U6 promoter-KO gRNA-EF1α-Cas9-P2A-Puro). For CRISPRi/CRSPRa assays, the gRNA in ATP2A2 SCZ GWAS loci was selected using BE-Designer tool (http://www.rgenome.net/be-designer/). The synthesized oligonucleotides for gRNA were subcloned into LentiGuide-Crimson plasmid (Addgene plasmid #217910) to obtain ATP2A2 SCZ GWAS loci gRNA plasmid (pLenti-U6 promoter-ATP2A2 gRNA-EF1α-Crimson).
Lentivirus package and infection
The 2nd generation lentiviral packaging plasmid (psPAX2, Addgene plasmid # 12260) and VSV-G envelope expressing plasmid (pMD2.G, Addgene plasmid # 12259), gifts from Didier Trono, were used to package the LV. These two packaging plasmids along with LV backbone plasmids were transfected to Lenti-HEK293T cells with polyethylenimine (PEI, Polyscience, Warrington, PA). Supernatants containing LVs were collected and concentrated using Lenti-X™ Concentrator (TaKaRa, San Jose, CA) according to the manufacturer’s instruction. Aliquots of the concentrated LVs were kept at −80 degree for later use.
For ATP2A2, DGKi or CACNA1D KOs, neurons were infected with LVs (MOI = 10) expressing Cas9 and KO gRNA (pLenti-U6 promoter-KO gRNA-EF1α-Cas9-P2A-Puro). After 24 hours’ infection, puromycin (1 μg/ml, InvivoGen, San Diego, CA) was added to the culture for one week to remove non-infected cells. Cells infected with LVs that express Cas9 only (pLenti-U6 promoter-EF1α-Cas9-P2A-Puro) were used as a negative control.
For the ATP2A2 CRISPRi/CRSPRa assay [138], cINs were infected with LVs expressing dCas9-VPR or dCas9-KRAB fusion proteins (lenti-EF1a-dCas9-VPR-Puro or lenti-EF1a-dCas9-KRAB-Puro) along with LVs expressing ATP2A2 SCZ GWAS loci gRNA (pLenti-U6-ATP2A2 gRNA-EF1α-Crimson). Puromycin was added to the culture for a week to select infected cells. Survived cells were also confirmed for Crimson expression. Lenti-EF1a-dCas9-VPR-Puro (Addgene plasmid # 99373) and lenti-EF1a-dCas9-KRAB-Puro (Addgene plasmid # 99372) were gifts from Kristen Brennand.
RNA purification and qPCR analysis
RNA samples were isolated using a TRIzol-reagent (Invitrogen) and 300 ng of the total RNA was used for cDNA synthesis using the Oligo(dT)12–18 primer (Gene Link, Hawthorne, NY), Recombinant RNAse Inhibitor (Takara, Japan), 0.5 mM dNTP mix (Thermo Scientific), 5 mM DTT (Invitrogen), RevertAid H minus RT (Thermo Scientific), and 5X Reaction Buffer (Thermo Scientific) in SimpliAmpTM Thermal Cycler (Applied Biosystems, Waltham, MA). The real time PCRs were performed using the QuantStudio 3 Real-Time PCR System (ThermoFisher), with 40 cycles of denaturation (95 °C for 15 s), annealing (55 °C for 30 s), and extension (72 °C for 30 s). Primer sequences are listed in Supplementary Table 9.
RNA-seq analysis
For RNA-seq, RNA quality was examined by the 2100 Bioanalyzer or 4200 TapeStation (Agilent Technologies, Santa Clara, CA) and RNA concentration was determined using the Qubit Fluorometric Quantitation (Thermo Fisher). Stranded cDNA libraries were prepared using the TruSeq Stranded LT mRNA kit (Illumina, San Diego, CA) in accordance with the manufacturer’s protocol using the poly-adenylated RNA isolation. Sequencing of paired-end reads (75 bp × 2) was performed on the NextSeq 550 system. Raw sequence reads were de-multiplexed and trimmed for adapters by using the Illumina bcl2fastq conversion software (v2.19). Sequence reads of each sample were pseudo-aligned to the human hg38 reference transcriptome and the gene transcript abundances were quantified by using Kallisto (v.0.46.1). Differential expression of genes and transcripts were achieved by using DESeq2 (v.1.26.0) in R Studio packages (v.1.2.1335 with R v.3.6.1) with a cut off criteria of adj p < 0.01. The Kallisto-Sleuth and Kallisto-DESeq2 workflows are based upon a previous publication [139]. For quantification, we used TPM (Transcripts Per Kilobase Million) as a measure of transcript abundance. For pair-wise comparisons, Pearson correlation was conducted in R Studio with DESeq2 normalized results. Pathways used for enrichment analysis by Fisher’s exact test is summarized in Supplementary Tables 1 and 11. Pathway analysis of SCZ GWAS DE genes in GNs were done using g:Profiler portal (https://biit.cs.ut.ee/gprofiler/gost). The RNA-seq data were deposited at the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) and the accession numbers are GSE121376 for cINs and GSE184102 for GNs.
Western blot
cINs infected with ATP2A2 or DGKI KO LVs were washed twice with ice-cold PBS and lysed with RIPA buffer (Thermo Fisher) containing protease inhibitor cocktail (bimake, Houston, TX) and phosphatase inhibitor cocktail (Santa Cruz Biotechnology, Dallas, TX). Protein concentration of cell lysate was determined by BCA protein assay kit (Thermo Fisher). The samples were run on 10% Precast Polyacrylamide Gels (Bio-rad, Hercules, CA) and then transferred to PVDF membrane (EMD Millipore). The PVDF membrane was blocked with 5% BSA (Cat#9048-46-8, Research Products International, Mount Prospect, IL) and incubated with antibody recognizing Phospho-PKC substrate (1:3000, Cell Signal Technology) and GAPDH (1:5000, Santa Cruz), respectively, at 4 °C overnight, followed by incubation with HRP-conjugated secondary antibody (1:10000, Vector) for 1 hour at room temperature. Detection was done using Super Signal West Pico Chemiluminescent Substrate (Pierce) and IBright FL1500 Imaging System (Thermo Fisher). Band intensities were quantified using Image J software (Version 1.51p, NIH, Bethesda, MD).
Immunocytochemistry/Cell counting/Arborization analysis
cINs and GNs on coverslips were fixed using 4% paraformaldehyde (PFA, Electron Microscopy Sciences, Hatfield, PA) for 10 min, washed with PBS and used for staining. The brain sections perfused with 4% formaldehyde and cryoprotected in the 30% sucrose (Thermo Fisher) overnight at 4 °C were cryosectioned at 40 µm using a Leica CM1850 cryostat (Leica Biosystem, Buffalo Grove, IL, USA). Fixed cells or brain sections were incubated with blocking/permeabilization buffer (PBS with 10% normal serum and 0.1% Triton X-100) for 10 min. The samples were then incubated in primary antibodies in antibody dilution buffer (PBS containing 2% normal serum) overnight at 4 °C. The detailed information of the antibodies used are as follows; GAD1 (sc7512, 1:1,000, Santa Cruz), SOX6 (AB5805, 1:1,000, Millipore, validated in ref. [49]), VGLUT1 (AB5905, 1:1,000, Millipore), GABA (A0310, 1:1,000, Sigma), b-Tubulin III (560381, 1:1,000, BD Bioscience), OLIG2 (AB9610, 1:1,000, Millipore, validated in ref. [49]), GFAP (73240, 1:1,000, UC Davis/NIH NeuroMab), CHAT (AB153, 1:1,000, Chemicon, validated in ref. [49]), TH (P40101-150, 1:1,000, Pel-Freez, validated in ref. [49]) and VIP (20077, 1:1,000, IMMUNOSTAR, validated in ref. [49]) as listed in Supplementary Table 10. After washing with PBS, cells were incubated with fluorescently labeled secondary antibodies and DAPI (Invitrogen) in antibody dilution buffer for 1 h at room temperature. Following the PBS wash, the samples were mounted with Fluoromount-G (SouthernBiotech, Birmingham, AL, USA). Fluorescent images were taken by the EVOS FL Auto microscope (Life Technologies, Carlsbad, CA) and Leica SPE Confocal Laser Scanning Microscopes (Leica microsystems, Wetzlar, Germany).
For cell counting analysis, Image J software (Version 1.51p, NIH, Bethesda, MD) was used to count the cell number using the multi point function. The experimenters were blind to the sample ID during data collection. Percentages of cells positive for each marker were quantified in relation to DAPI-stained nuclei from three independent differentiations, with a total of at least 500 cells counted for each line.
For arborization analysis, cINs plated onto the PLO/FN-coated coverslips were infected with a limiting titer (MOI = 0.001) of LV-UbiC-GFP virus [140] to label cells only scarcely, in addition to the KO LVs described above. After one week’s selection in Puromycin to remove non-infected cells, the images of GFP+ cells were collected by the EVOS microscope. The arborization of each GFP+ cell was analyzed using Image J software with the Neuron J plugin to get parameters of total neurite lengths.
ATAC-seq analysis
Library generation and sequencing
ATAC-seq reactions were performed using an established protocol [141] with minor modifications. Media was removed and replaced with 200 µl of 1X Trypsin and 100 mM trehalose. Tubes were incubated at 37 °C for 10 min, after which dissociation was performed by gentle pipetting. Trypsin was neutralized by the addition of 200 µl DMEM +10% FBS containing 0.5 µl turbo DNase and samples incubated for an additional 20 min at 37 °C. Cells were then centrifuged at 300 × g for 10 min at 4 °C and pellets resuspended in 250 µl PBS + 5% BSA. DAPI (Thermo Fisher) was added to the cells and viable (DAPI-) cells sorted on a FACSAria flow cytometer (BD Biosciences). Following FACS, cell concentrations and viability were confirmed using a Countess automated cell counter (Life technologies) and, where available, 75,000 cells were carried forward for the ATAC-seq reactions. Cells were centrifuged at 300 × g for 10 min at 4 °C and, following removal of the supernatant, pellets were resuspended in lysis buffer (10 mM Tris-Cl pH7.4, 10 mM NaCl, 3 mM MgCl, 0.1% IGEPAL CA630) by pipetting 20 times. Following resuspension, samples were centrifuged at 500 × g for 10 min at 4 °C. Pellets were resuspended in transposase reaction mix (25 μL 2× TD Buffer (Illumina Cat #FC-121-1030) 2.5 μL Tn5 Transposase (Illumina Cat #FC-121-1030) and 22.5 μL Nuclease Free H2O) on ice. Reactions were incubated at 37 °C for 30 min before being purified using the MinElute Reaction Cleanup kit (Qiagen Cat #28204). Following purification, library fragments were amplified using the Nextera index kit (Illumina Cat #FC-121-1011) as previously described [141]. Library quality was assessed on a Tapestation instrument using D5000 ScreenTapes (Agilent technologies Cat# 5067-5588). Following amplification, libraries were resolved on 2% agarose gels and fragments ranging in size from 100 to 1000 bp were excised and purified (Qiagen Minelute Gel Extraction Kit – Qiagen Cat#28604). Before sequencing, library fragment sizes estimated using Tapestation D5000 ScreenTapes (Agilent technologies Cat# 5067–5588) and libraries quantified by quantitative PCR (KAPA Biosystems Cat#KK4873). Finally, libraries were sequenced on the NovaSeq6000 platform (Illumina) obtaining 2 × 50 paired-end reads.
Read mapping
Raw sequencing paired-end files were aligned by STAR (v2.7.0.e) (Dobin et al. 2013) to the hg38 reference genome with the pseudoautosomal region masked on chromosome Y using the following parameters: -alignIntronMax 1 -out FilterMismatchNmax 100 -alignEndsType EndToEnd -out FilterScoreMinOverLread 0.3 -out FilterMatchNminOverLread 0.3. This yielded for each sample a BAM file of mapped paired-end reads sorted by genomic coordinates. From these BAM-files, we removed reads that were mapped to multiple loci or to the mitochondrial genome using samtools (v0.1.19) [142] as well as the duplicated reads using PICARD (v2.24; http://broadinstitute.github.io/picard). The read coverage files (bedGraph) were generated by genomeCoverageBed from BEDTools (v2.25.0) (Quinlan and Hall 2010) and converted to the indexed binary format files (bigWig) using bedGraphToBigWig.
Quality control
To perform quality control, we collected the following metrics for all samples: the total number of initial reads; the number of uniquely mapped reads; the fraction of reads that were uniquely mapped; further mappability-related metrics from the STAR aligner; GC content, insert and duplication metrics from Picard; the rate of reads mapping to the mitochondrial genome; the PCR bottleneck coefficient (PBC), which approximates library complexity as uniquely mapped non-redundant reads divided by the number uniquely mapped reads; the relative strand cross-correlation coefficient (RSC) and the normalized strand cross-correlation coefficient (NSC), which are metrics that use cross-correlation of stranded read density profiles to evaluate the sample quality independently of peak calling; and, finally, the fraction of reads in peaks (FRiP), which is the fraction of reads that fall in peaks. To calculate FRiP, we called peaks for each BAM file with MACS2 [143] using the following parameters: -keep-dup all -shift −100 -extsize 200 -nomodel. Then, per-sample peaks were merged across all samples, retaining only peaks found in at least 2 samples. Subsequently, we used featureCounts from Rsubread package to calculate the fraction of reads that overlap consensus peaks. The main quality metrics are provided in Supplementary Table 5. All samples exceeded our quality controls thresholds, i.e., having more than 5 million paired-end reads (mean = 38.4 million, min = 16.8 million), mappability over 50% (mean = 83.2%, min = 50.5%), and FRiP over 5% (mean = 26.2%, min = 9.0%).
To compare regulatory landscape of iPSC-derived cINs within in vivo cell types, we compared ATAC-seq peaks of iPSC-derived cINs with ATAC-seq peaks from GNs and cINs sorted from the fetal cortex [30]. We used the findOverlap function in the GenomicRanges package to overlap iPSC-derived cIN ATAC-seq peaks with fetal excitatory and inhibitory neuronal ATAC-seq peaks. We then compared the proportion of fetal inhibitory neuronal peaks that overlap with cIN peaks with the proportion of fetal excitatory neuronal peaks that overlap with cIN peaks using a Fisher’s exact test.
Those samples were visually inspected in IGV. The data from this study have been submitted to the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) under accession number GSE184165.
ChIP-seq analysis
5 × 106 cINs were used for ChIP experiment using the Pierce Magnetic ChIP Kit (Cat #26157, Thermo Fisher) according to the manufacturer’s instruction. In brief, cells were fixed in 1% formaldehyde at room temperature for 15 min, followed by addition of Glycine. After 5 minutes’ incubation at room temperature, cells were washed twice with ice-cold PBS. Cells were scraped in ice-cold PBS supplemented with protease inhibitor cocktail, followed by centrifugation and resuspension in extraction buffer to lyse the cell membrane and collect nuclei. After centrifugation, obtained nuclei were resuspended with MNase digestion buffer and incubated in a 37 °C water bath for 15 min. After centrifugation, nuclei were resuspended with immunoprecipitation buffer containing protease inhibitors and phosphatase inhibitors. After sonication on ice, samples were incubated with either anti-H3K4me1 (Cat #710795, Thermo Fisher) or Anti- H3K27Ac antibody (ab4729, Abcam) overnight at 4 °C with rotation. The next day, magnetic beads were added to each sample and incubated for 2 h at 4 °C. After washing three times, DNAs were recovered in elution solution.
The purified DNAs were converted to small fragments using the NEBNext Ultra II FS Enzyme Mix (NEBNext Ultra II FS DNA library Prep Kit, New England Biolabs) at 37 °C for 10 min and 65 °C for 30 min. The target DNA with biotin labeling was pulled down using Streptavidin Dynabeads M270 (Invitrogen), washed once with Tween buffer (5 mM Tris-HCl, pH 8, 0.5 mM EDTA, 1M NaCl, 0.05% Tween-20) and three times with no-Tween buffer. The biotin-labeled DNA was then ligated to an adaptor with index (TruSeq Stranded Total RNA Kit, Illumina) at 20 °C for 15 min, washed twice with the no-Tween buffer, and PCR amplified (98 °C 30 s; 15 cycles of 98 °C 10 s, 60 °C 30 s and 72 °C 30 s; and at final 72 °C 5 min). The PCR product was purified with 0.8 volume of AMPure XP beads (Beckman Coulter), quantified with Qubit 2.0 fluorometer (Invitrogen), and loaded onto NextSeq 550 (Illumina) with its High Output reagents for 75 bp × 2 paired-end sequencing. The fastq files obtained from sequencing were aligned/mapped to the human reference genome GRCh38 using Bowtie2 [144], and further subjected to peak calling using MACS2 [145]. ChIP-seq data was submitted to the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) with the accession number GSE186738.
Whole-cell patch-clamp recording in brain slices
HC iPSC L9-derived developing GNs infected with ChR2-GFP-expressing lentivirus (Addgene #20945) were transplanted into prefrontal cortex of NSG mice at the following coordinate (AP: +1.9 mm, L: +0.4 mm, V:1.2). Nine months post transplantation, the neuronal and synaptic properties of human GNs were examined with whole-cell patch-clamp recording in brain slices. Acute brain slices containing the PFC were prepared using a vibrating microtome. After recovery, brain slices were placed in the recording chamber and continuously perfused at the rate of 1 mL per minute with the artificial cerebrospinal fluid (130 mM NaCl, 2.5 mM KCl, 2.5 mM CaCl2, 1 mM MgSO4, 1.25 mM NaH2PO4, 26 mM NaHCO3, and 10 mM glucose with 95% O2 and 5% CO2). Whole-cell patch-clamp recordings were performed at 30–32 °C using a Multiclamp 700B amplifier, a Digidata 1550A digitizer, and Clampex 10 software (Molecular Devices). The patch electrodes (2.5–3.5 MΩ resistance) were filled with solution containing 150 mM K-gluconate, 5 mM NaCl, 1 mM MgCl2, 10 mM HEPES, 0.2 EGTA, 2 mM MgATP, and 0.5 mM NaGTP (290 mOsm, adjusted to pH 7.3 with KOH). Liquid junction potential of 15.5 mV was corrected. The resting membrane potential was estimated as the average membrane potential recorded without current injection or withdrawal in current-clamp mode. To calculate membrane resistance (Rm) and membrane capacitance (Cm) of grafted human interneurons, we applied hyperpolarizing voltage pulses (−5 mV, 50 ms-long) from the holding voltage of −85 mV and recorded currents in voltage-clamp mode with a sampling rate of 100 kHz in grafted cells. The peak amplitude of the transient capacitive current (Ipeak) was used to estimate apparent access resistance (Ra) using the equation, Ra = −5 mV/Ipeak. Steady-state current (Iss) is the average current from the baseline for the last 5 ms of voltage pulse and was used to calculate membrane resistance (Rm) using the equation, Rm = (−5 mV/Iss) – Ra. Transient capacitive current was fitted with a double exponential function, I(t) = If exp(–t/τf) + Is exp(–t/τs), where If/τf and Is/τs are fast and slow components of peak amplitudes and decay time constants of transient capacitive current. The weighted mean decay time constant (τw) was calculated from the equation, τw = τf [If/(If + Is)] + τs [Is/(If + Is)] and then was used to calculate membrane capacitance (Cm) from the equation, Cm = τ w × (Ra + Rm)/(Ra × Rm). Whole-cell membrane capacitance (Cm) was compensated with the amplifier. Series (access) resistance was not compensated. The membrane potential was held at approximately −85 mV in current-clamp mode, and square pulses of depolarizing currents (0–300 pA with the increment of 50 pA, 0.5 s duration) were applied. The first APs induced by depolarizing current injection near threshold was used to calculate afterhyperpolarization. Spontaneous EPSC (sEPSC) were recorded in grafted cells at −85 mV in voltage-clamp mode. Blue collimated light-emitting diode (LED) with 470 nm peak wavelength (M470L2, Thorlabs) was used for photostimulation of grafted interneurons expressing ChR2-GFP. Brain slices in the recording chamber were illuminated through a 40× water-immersion objective lens (Olympus LUMPLFLN 40XW) and a 450–490 nm filter (Chroma). Illumination area was 0.17 mm2 and was centered at the cell patched for recording. NBQX disodium salt was purchased from Tocris Bioscience (Cat. No. 1044). Offline analysis of electrophysiological data was performed using Clampfit 10 program (Molecular Devices).
Ca2+ currents analysis of iPSC-derived GNs
After 3 months’ differentiation, human GNs on PLO/FN-coated glass coverslips were used for electrophygiological experiments. Microelectrodes were pulled from standard-wall borosilicate glass with a flaming/brown micropipette puller (P-97, Sutter Instruments). Patch pipettes with a tip resistance of 4–5 M when they were filled with internal solution. GNs on coverslips were transferred to the recording chamber which was perfused with B27GB media added with 3 mM CsCl2 and 0.0005 mM TTX. The internal pipette solution contained 20 U/ml phosphocreatine kinase, 100 mM CsMeSO3, 10 mM EGTA, 1 mM CaC12, 10 mM HEPES, 2 mM MgATP, 0.1 mM leupeptin, and 20mM phosphocreatine. Electrophysiological signals were recorded from the cultured GNs which typically showed neuronal dendritic processes as illustrated in the Fig. 5B. Total Ca2+ currents and high voltage-activated Ca2+ currents were recorded using whole-cell patch clamp in voltage-clamp mode. Electric signals were filtered at 10 kH and digitized at 3 kHz with Axon DigiData 1550 (Molecular Devices, California) that was controlled with a MultiClamp 700B (Molecular Devices) and run by Clampex (v10). After establishing a whole-cell configuration, the membrane potential was clamped at −70 mV. Electrophysiological data were analyzed initially with Clampfit (v10) (Molecular Devices). Analyzed data were further processed and presented with Igor (v6) and presented with CorelDraw (2021) (Corel, Ottawa, Canada) programs.
when they were filled with internal solution. GNs on coverslips were transferred to the recording chamber which was perfused with B27GB media added with 3 mM CsCl2 and 0.0005 mM TTX. The internal pipette solution contained 20 U/ml phosphocreatine kinase, 100 mM CsMeSO3, 10 mM EGTA, 1 mM CaC12, 10 mM HEPES, 2 mM MgATP, 0.1 mM leupeptin, and 20mM phosphocreatine. Electrophysiological signals were recorded from the cultured GNs which typically showed neuronal dendritic processes as illustrated in the Fig. 5B. Total Ca2+ currents and high voltage-activated Ca2+ currents were recorded using whole-cell patch clamp in voltage-clamp mode. Electric signals were filtered at 10 kH and digitized at 3 kHz with Axon DigiData 1550 (Molecular Devices, California) that was controlled with a MultiClamp 700B (Molecular Devices) and run by Clampex (v10). After establishing a whole-cell configuration, the membrane potential was clamped at −70 mV. Electrophysiological data were analyzed initially with Clampfit (v10) (Molecular Devices). Analyzed data were further processed and presented with Igor (v6) and presented with CorelDraw (2021) (Corel, Ottawa, Canada) programs.
Statistics
All statistical analyses were performed using GraphPad Prism8 (GraphPad Software, La Jolla, CA), and SAS statistical software (9.4, SAS Institute, Cary, NC). For each figure, the statistical test was justified as appropriate, meeting the assumption of the tests as summarized in Supplementary Table 14. The normal distribution of data was tested using the Shapiro–Wilks test. Data with significant Shapiro–Wilks tests were analyzed after log-transformation or using non-parametric tests as indicated in Supplementary Table 14. We tested equal variance with the F-test, and if there was any significant difference, we used the t-test with Welch’s correction. To compare the difference between two means, a two-tailed paired or unpaired t-test was used as shown in Supplementary Table 14. To handle clustering and covariance among correlated samples [146, 147], a mixed effect model was used as summarized in Supplementary Table 14. For the comparison of group means for more than 2 groups, one-way ANOVA or Two-Level Hierarchical Linear Mixed Effect Model were used depending on data clustering, and if there was significant difference, posthoc analysis was performed using Dunnett’s multiple comparisons. P-values < 0.05 were considered to be statistically significant. Experimental cohorts were chosen based on our selection criteria (Caucasian male patients treated with Clozapine vs. age- and gender-matched Caucasian male healthy controls) without randomization to reduce variation caused by age, ethnicity and gender. Blinding was used during cell counting and arborization analysis. No data was excluded and no statistical method was used to pre-determine the sample sizes.
References
Sullivan PF, Kendler KS, Neale MC. Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Arch Gen Psychiatry. 2003;60:1187–92.
Weinberger DR. Implications of normal brain development for the pathogenesis of schizophrenia. Arch Gen Psychiatry. 1987;44:660–9.
Carpenter WT Jr., Buchanan RW. Schizophrenia. N Engl J Med. 1994;330:681–90.
Ripke S, Neale BM, Corvin A, Walters JTR, Farh KH, Holmans PA, et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Wainberg M, Sinnott-Armstrong N, Mancuso N, Barbeira AN, Knowles DA, Golan D, et al. Opportunities and challenges for transcriptome-wide association studies. Nat Genet. 2019;51:592–9.
Collado-Torres L, Burke EE, Peterson A, Shin J, Straub RE, Rajpurohit A et al. Regional heterogeneity in gene expression, regulation, and coherence in the frontal cortex and hippocampus across development and schizophrenia. Neuron. 2019;103:203–216.e8.
Zhang K, Hocker JD, Miller M, Hou X, Chiou J, Poirion OB, et al. A single-cell atlas of chromatin accessibility in the human genome. Cell. 2021;184:5985–6001.e5919.
Finucane HK, Bulik-Sullivan B, Gusev A, Trynka G, Reshef Y, Loh PR, et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet. 2015;47:1228–35.
Schork AJ, Won H, Appadurai V, Nudel R, Gandal M, Delaneau O, et al. A genome-wide association study of shared risk across psychiatric disorders implicates gene regulation during fetal neurodevelopment. Nat Neurosci. 2019;22:353–61.
Sey NYA, Hu B, Mah W, Fauni H, McAfee JC, Rajarajan P, et al. A computational tool (H-MAGMA) for improved prediction of brain-disorder risk genes by incorporating brain chromatin interaction profiles. Nat Neurosci. 2020;23:583–93.
Marin O. Interneuron dysfunction in psychiatric disorders. Nat Rev Neurosci. 2012;13:107–20.
Kalus P, Bondzio J, Federspiel A, Muller TJ, Zuschratter W. Cell-type specific alterations of cortical interneurons in schizophrenic patients. Neuroreport. 2002;13:713–7.
Guidotti A, Auta J, Davis JM, Di-Giorgi-Gerevini V, Dwivedi Y, Grayson DR, et al. Decrease in reelin and glutamic acid decarboxylase67 (GAD67) expression in schizophrenia and bipolar disorder: a postmortem brain study. Arch Gen Psychiatry. 2000;57:1061–9.
Volk D, Austin M, Pierri J, Sampson A, Lewis D. GABA transporter-1 mRNA in the prefrontal cortex in schizophrenia: decreased expression in a subset of neurons. Am J Psychiatry. 2001;158:256–65.
Reynolds GP, Abdul-Monim Z, Neill JC, Zhang ZJ. Calcium binding protein markers of GABA deficits in schizophrenia-postmortem studies and animal models. Neurotox Res. 2004;6:57–61.
Fung SJ, Webster MJ, Sivagnanasundaram S, Duncan C, Elashoff M, Weickert CS. Expression of interneuron markers in the dorsolateral prefrontal cortex of the developing human and in schizophrenia. Am J Psychiatry. 2010;167:1479–88.
Hashimoto T, Arion D, Unger T, Maldonado-Aviles JG, Morris HM, Volk DW, et al. Alterations in GABA-related transcriptome in the dorsolateral prefrontal cortex of subjects with schizophrenia. Mol Psychiatry. 2008;13:147–61.
Hoftman GD, Volk DW, Bazmi HH, Li S, Sampson AR, Lewis DA. Altered cortical expression of GABA-related genes in schizophrenia: illness progression vs developmental disturbance. Schizophrenia Bull. 2013;41:180–91.
Wang AY, Lohmann KM, Yang CK, Zimmerman EI, Pantazopoulos H, Herring N, et al. Bipolar disorder type 1 and schizophrenia are accompanied by decreased density of parvalbumin- and somatostatin-positive interneurons in the parahippocampal region. Acta Neuropathol. 2011;122:615–26.
Morris HM, Hashimoto T, Lewis DA. Alterations in somatostatin mRNA expression in the dorsolateral prefrontal cortex of subjects with schizophrenia or schizoaffective disorder. Cereb Cortex. 2008;18:1575–87.
Meyer-Lindenberg A. From maps to mechanisms through neuroimaging of schizophrenia. Nature. 2010;468:194–202.
Jaaro-Peled H, Ayhan Y, Pletnikov MV, Sawa A. Review of pathological hallmarks of schizophrenia: comparison of genetic models with patients and nongenetic models. Schizophrenia Bull. 2010;36:301–13.
Lewis DA, Hashimoto T, Volk DW. Cortical inhibitory neurons and schizophrenia. Nat Rev Neurosci. 2005;6:312–24.
Fuchs EC, Zivkovic AR, Cunningham MO, Middleton S, Lebeau FE, Bannerman DM, et al. Recruitment of parvalbumin-positive interneurons determines hippocampal function and associated behavior. Neuron. 2007;53:591–604.
Sohal VS, Zhang F, Yizhar O, Deisseroth K. Parvalbumin neurons and gamma rhythms enhance cortical circuit performance. Nature. 2009;459:698–702.
Williams S, Boksa P. Gamma oscillations and schizophrenia. J Psychiatry Neurosci. 2010;35:75–77.
Dienel SJ, Lewis DA. Alterations in cortical interneurons and cognitive function in schizophrenia. Neurobiol Dis. 2019;131:104208.
Lewis DA, Glantz LA, Pierri JN, Sweet RA. Altered cortical glutamate neurotransmission in schizophrenia: evidence from morphological studies of pyramidal neurons. Ann N Y Acad Sci. 2003;1003:102–12.
Polioudakis D, de la Torre-Ubieta L, Langerman J, Elkins AG, Shi X, Stein JL, et al. A single-cell transcriptomic atlas of human neocortical development during mid-gestation. Neuron. 2019;103:785–801.e788.
Song M, Pebworth M-P, Yang X, Abnousi A, Fan C, Wen J et al. Cell-type-specific 3D epigenomes in the developing human cortex. Nature 2020;587:644–9.
Trevino AE, Sinnott-Armstrong N, Andersen J, Yoon S-J, Huber N, Pritchard JK, et al. Chromatin accessibility dynamics in a model of human forebrain development. Science. 2020;367:eaay1645.
Ziffra RS, Kim CN, Ross JM, Wilfert A, Turner TN, Haeussler M et al. Single-cell epigenomics reveals mechanisms of human cortical development. Nature 2021;598:205–13.
Benson CA, Powell HR, Liput M, Dinham S, Freedman DA, Ignatowski TA, et al. Immune factor, TNFα, disrupts human brain organoid development similar to schizophrenia-schizophrenia increases developmental vulnerability to TNFα. Front Cell Neurosci. 2020;14:233.
Brennand KJ, Simone A, Jou J, Gelboin-Burkhart C, Tran N, Sangar S, et al. Modelling schizophrenia using human induced pluripotent stem cells. Nature. 2011;473:221–5.
Forrest MP, Zhang H, Moy W, McGowan H, Leites C, Dionisio LE, et al. Open chromatin profiling in hiPSC-derived neurons prioritizes functional noncoding psychiatric risk variants and highlights neurodevelopmental loci. Cell Stem Cell. 2017;21:305–31.e308.
Robicsek O, Karry R, Petit I, Salman-Kesner N, Müller FJ, Klein E, et al. Abnormal neuronal differentiation and mitochondrial dysfunction in hair follicle-derived induced pluripotent stem cells of schizophrenia patients. Mol Psychiatry. 2013;18:1067–76.
Li J, Tran OT, Crowley TB, Moore TM, Zackai EH, Emanuel BS, et al. Association of mitochondrial biogenesis with variable penetrance of schizophrenia. JAMA Psychiatry. 2021;78:911–21.
Linaro D, Vermaercke B, Iwata R, Ramaswamy A, Libé-Philippot B, Boubakar L et al. Xenotransplanted human cortical neurons reveal species-specific development and functional integration into mouse visual circuits. Neuron. 2019;104:972–86.e6.
Camp JG, Badsha F, Florio M, Kanton S, Gerber T, Wilsch-Brauninger M, et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc Natl Acad Sci USA 2015;112:15672–7.
Velasco S, Kedaigle AJ, Simmons SK, Nash A, Rocha M, Quadrato G, et al. Individual brain organoids reproducibly form cell diversity of the human cerebral cortex. Nature. 2019;570:523–7.
Luo C, Lancaster MA, Castanon R, Nery JR, Knoblich JA, Ecker JR. Cerebral organoids recapitulate epigenomic signatures of the human fetal brain. Cell Rep. 2016;17:3369–84.
Amiri A, Coppola G, Scuderi S, Wu F, Roychowdhury T, Liu F, et al. Transcriptome and epigenome landscape of human cortical development modeled in organoids. Science. 2018;362:eaat6720.
Keller G. Embryonic stem cell differentiation: emergence of a new era in biology and medicine. Genes Dev. 2005;19:1129–55.
Zhu Z, Huangfu D. Human pluripotent stem cells: an emerging model in developmental biology. Development. 2013;140:705–17.
Qian X, Nguyen HaN, Song Mingxi M, Hadiono C, Ogden Sarah C, Hammack C, et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell. 2016;165:1238–54.
Paşca AM, Sloan SA, Clarke LE, Tian Y, Makinson CD, Huber N, et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat Methods. 2015;12:671–8.
Rajarajan P, Borrman T, Liao W, Schrode N, Flaherty E, Casino C et al. Neuron-specific signatures in the chromosomal connectome associated with schizophrenia risk. Science. 2018;362:eaat4311.
Cunningham M, Cho J-H, Leung A, Savvidis G, Ahn S, Moon M, et al. hPSC-derived maturing GABAergic interneurons ameliorate seizures and abnormal behavior in epileptic mice. Cell Stem Cell. 2014;15:559–73.
Shao Z, Noh H, Bin Kim W, Ni P, Nguyen C, Cote SE, et al. Dysregulated protocadherin-pathway activity as an intrinsic defect in induced pluripotent stem cell-derived cortical interneurons from subjects with schizophrenia. Nat Neurosci. 2019;22:229–42.
Kim T-G, Yao R, Monnell T, Cho J-H, Vasudevan A, Koh A, et al. Efficient specification of interneurons from human pluripotent stem cells by dorsoventral and rostrocaudal modulation. Stem Cells. 2014;32:1789–804.
Nicholas CR, Chen J, Tang Y, Southwell DG, Chalmers N, Vogt D, et al. Functional maturation of hPSC-derived forebrain interneurons requires an extended timeline and mimics human neural development. Cell Stem Cell. 2013;12:573–86.
Zecevic N, Hu F, Jakovcevski I. Interneurons in the developing human neocortex. Dev Neurobiol. 2011;71:18–33.
Aguilar-Valles A, Luheshi GN. Alterations in cognitive function and behavioral response to amphetamine induced by prenatal inflammation are dependent on the stage of pregnancy. Psychoneuroendocrinology. 2011;36:634–48.
Li Q, Cheung C, Wei R, Hui ES, Feldon J, Meyer U, et al. Prenatal immune challenge is an environmental risk factor for brain and behavior change relevant to schizophrenia: evidence from MRI in a mouse model. PLoS One. 2009;4:e6354.
Piper M, Beneyto M, Burne TH, Eyles DW, Lewis DA, McGrath JJ. The neurodevelopmental hypothesis of schizophrenia: convergent clues from epidemiology and neuropathology. Psychiatr Clin North Am. 2012;35:571–84.
Xu B, Ionita-Laza I, Roos JL, Boone B, Woodrick S, Sun Y, et al. De novo gene mutations highlight patterns of genetic and neural complexity in schizophrenia. Nat Genet. 2012;44:1365–9.
Lin M, Pedrosa E, Shah A, Hrabovsky A, Maqbool S, Zheng D, et al. RNA-Seq of human neurons derived from iPS cells reveals candidate long non-coding RNAs involved in neurogenesis and neuropsychiatric disorders. PLoS One. 2011;6:e23356.
Gilman SR, Chang J, Xu B, Bawa TS, Gogos JA, Karayiorgou M, et al. Diverse types of genetic variation converge on functional gene networks involved in schizophrenia. Nat Neurosci. 2012;15:1723–8.
Ni P, Noh H, Park G-H, Shao Z, Guan Y, Park JM et al. iPSC-derived homogeneous populations of developing schizophrenia cortical interneurons have compromised mitochondrial function. Mol Psychiatry. 2020;25:2873–88.
Ni P, Noh H, Shao Z, Zhu Q, Guan Y, Park JJ, et al. Large-scale generation and characterization of homogeneous populations of migratory cortical interneurons from human pluripotent stem cells. Mol Ther Methods Clin Dev. 2019;13:414–30.
Maroof AM, Keros S, Tyson JA, Ying SW, Ganat YM, Merkle FT, et al. Directed differentiation and functional maturation of cortical interneurons from human embryonic stem cells. Cell Stem Cell. 2013;12:559–72.
Liu Y, Weick JP, Liu H, Krencik R, Zhang X, Ma L, et al. Medial ganglionic eminence-like cells derived from human embryonic stem cells correct learning and memory deficits. Nat Biotechnol. 2013;31:440–7.
Pardinas AF, Holmans P, Pocklington AJ, Escott-Price V, Ripke S, Carrera N, et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat Genet. 2018;50:381–9.
Goto K, Kondo H. Diacylglycerol kinase in the central nervous system-molecular heterogeneity and gene expression. Chem Phys Lipids. 1999;98:109–17.
Nakajima K, Ishiwata M, Weitemier AZ, Shoji H, Monai H, Miyamoto H et al. Brain-specific heterozygous loss-of-function of ATP2A2, endoplasmic reticulum Ca2+ pump responsible for Darier’s disease, causes behavioral abnormalities and a hyper-dopaminergic state. Hum Mol Genet. 2021;30:1762–72.
Anderson A, Masuho I, Marron Fernandez de Velasco E, Nakano A, Birnbaumer L, Martemyanov KA, et al. GPCR-dependent biasing of GIRK channel signaling dynamics by RGS6 in mouse sinoatrial nodal cells. Proc Natl Acad Sci USA 2020;117:14522–31.
Ahlers KE, Chakravarti B, Fisher RA. RGS6 as a novel therapeutic target in CNS diseases and cancer. AAPS J. 2016;18:560–72.
Verbeek DS, Goedhart J, Bruinsma L, Sinke RJ, Reits EA. PKC gamma mutations in spinocerebellar ataxia type 14 affect C1 domain accessibility and kinase activity leading to aberrant MAPK signaling. J Cell Sci. 2008;121:2339–49.
Yamamoto K, Seki T, Adachi N, Takahashi T, Tanaka S, Hide I, et al. Mutant protein kinase C gamma that causes spinocerebellar ataxia type 14 (SCA14) is selectively degraded by autophagy. Genes Cells. 2010;15:425–38.
Alfonso SI, Callender JA, Hooli B, Antal CE, Mullin K, Sherman MA, et al. Gain-of-function mutations in protein kinase Cα (PKCα) may promote synaptic defects in Alzheimer’s disease. Sci Signal. 2016;9:ra47.
Battaini F. Protein kinase C isoforms as therapeutic targets in nervous system disease states. Pharmacol Res. 2001;44:353–61.
Manji HK, Moore GJ, Chen G. Bipolar disorder: leads from the molecular and cellular mechanisms of action of mood stabilizers. Br J Psychiatry Suppl. 2001;41:s107–119.
Amrollahi Z, Rezaei F, Salehi B, Modabbernia AH, Maroufi A, Esfandiari GR, et al. Double-blind, randomized, placebo-controlled 6-week study on the efficacy and safety of the tamoxifen adjunctive to lithium in acute bipolar mania. J Affect Disord. 2011;129:327–31.
Bebchuk JM, Arfken CL, Dolan-Manji S, Murphy J, Hasanat K, Manji HK. A preliminary investigation of a protein kinase C inhibitor in the treatment of acute mania. Arch Gen Psychiatry. 2000;57:95–97.
Yildiz A, Guleryuz S, Ankerst DP, Ongur D, Renshaw PF. Protein kinase C inhibition in the treatment of mania: a double-blind, placebo-controlled trial of tamoxifen. Arch Gen Psychiatry. 2008;65:255–63.
Birnbaum SG, Yuan PX, Wang M, Vijayraghavan S, Bloom AK, Davis DJ, et al. Protein kinase C overactivity impairs prefrontal cortical regulation of working memory. Science. 2004;306:882–4.
Brennan AR, Yuan P, Dickstein DL, Rocher AB, Hof PR, Manji H, et al. Protein kinase C activity is associated with prefrontal cortical decline in aging. Neurobiol Aging. 2009;30:782–92.
Runyan JD, Moore AN, Dash PK. A role for prefrontal calcium-sensitive protein phosphatase and kinase activities in working memory. Learn Mem. 2005;12:103–10.
Calabrese B, Halpain S. Essential role for the PKC target MARCKS in maintaining dendritic spine morphology. Neuron. 2005;48:77–90.
Callender JA, Newton AC. Conventional protein kinase C in the brain: 40 years later. Neuronal Signal. 2017;1:Ns20160005.
Tagawa K, Homma H, Saito A, Fujita K, Chen X, Imoto S, et al. Comprehensive phosphoproteome analysis unravels the core signaling network that initiates the earliest synapse pathology in preclinical Alzheimer’s disease brain. Hum Mol Genet. 2015;24:540–58.
Garrett AM, Schreiner D, Lobas MA, Weiner JA. γ-protocadherins control cortical dendrite arborization by regulating the activity of a FAK/PKC/MARCKS signaling pathway. Neuron. 2012;74:269–76.
Huang L, Wang C, Zhao S, Ge R, Guan S, Wang JH. PKC and CaMK-II inhibitions coordinately rescue ischemia-induced GABAergic neuron dysfunction. Oncotarget. 2017;8:39309–22.
Chen Y, Cantrell AR, Messing RO, Scheuer T, Catterall WA. Specific modulation of Na+ channels in hippocampal neurons by protein kinase C epsilon. J Neurosci. 2005;25:507–13.
Carr DB, Cooper DC, Ulrich SL, Spruston N, Surmeier DJ. Serotonin receptor activation inhibits sodium current and dendritic excitability in prefrontal cortex via a protein kinase C-dependent mechanism. J Neurosci. 2002;22:6846–55.
Castro-Alamancos MA, Arevalo MA, Torres-Aleman I. Involvement of protein kinase C and nitric oxide in the modulation by insulin-like growth factor-I of glutamate-induced GABA release in the cerebellum. Neuroscience. 1996;70:843–7.
Jaffe AE, Straub RE, Shin JH, Tao R, Gao Y, Collado-Torres L, et al. Developmental and genetic regulation of the human cortex transcriptome illuminate schizophrenia pathogenesis. Nat Neurosci. 2018;21:1117–25.
Gandal MJ, Zhang P, Hadjimichael E, Walker RL, Chen C, Liu S et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science. 2018;362:eaat8127.
Wang D, Liu S, Warrell J, Won H, Shi X, Navarro FCP, et al. Comprehensive functional genomic resource and integrative model for the human brain. Science. 2018;362:eaat8464.
Li M, Santpere G, Imamura Kawasawa Y, Evgrafov OV, Gulden FO, Pochareddy S, et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science. 2018;362:eaat7615.
Walker RL, Ramaswami G, Hartl C, Mancuso N, Gandal MJ, de la Torre-Ubieta L, et al. Genetic control of expression and splicing in developing human brain informs disease mechanisms. Cell. 2019;179:750–71.e722.
Wang Q, Chen R, Cheng F, Wei Q, Ji Y, Yang H, et al. A Bayesian framework that integrates multi-omics data and gene networks predicts risk genes from schizophrenia GWAS data. Nat Neurosci. 2019;22:691–9.
Gusev A, Mancuso N, Won H, Kousi M, Finucane HK, Reshef Y, et al. Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat Genet. 2018;50:538–48.
Won H, de la Torre-Ubieta L, Stein JL, Parikshak NN, Huang J, Opland CK, et al. Chromosome conformation elucidates regulatory relationships in developing human brain. Nature. 2016;538:523–7.
Hisatsune C, Nakamura K, Kuroda Y, Nakamura T, Mikoshiba K. Amplification of Ca2+ signaling by diacylglycerol-mediated inositol 1,4,5-trisphosphate production. J Biol Chem. 2005;280:11723–30.
Liu L, Yudin Y, Rohacs T. Diacylglycerol kinases regulate TRPV1 channel activity. J Biol Chem. 2020;295:8174–85.
Kunkel MT, Newton AC. Calcium transduces plasma membrane receptor signals to produce diacylglycerol at Golgi membranes. J Biol Chem. 2010;285:22748–52.
Young KW, Nash MS, Challiss RA, Nahorski SR. Role of Ca2+ feedback on single cell inositol 1,4,5-trisphosphate oscillations mediated by G-protein-coupled receptors. J Biol Chem. 2003;278:20753–60.
Trubetskoy V, Pardinas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 2022;604:502–8.
Ward LD, Kellis M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 2016;44:D877–881.
Schrode N, Ho SM, Yamamuro K, Dobbyn A, Huckins L, Matos MR, et al. Synergistic effects of common schizophrenia risk variants. Nat Genet. 2019;51:1475–85.
Tian R, Abarientos A, Hong J, Hashemi SH, Yan R, Drager N, et al. Genome-wide CRISPRi/a screens in human neurons link lysosomal failure to ferroptosis. Nat Neurosci. 2021;24:1020–34.
Lai RK, Xu IM, Chiu DK, Tse AP, Wei LL, Law CT, et al. NDUFA4L2 fine-tunes oxidative stress in hepatocellular carcinoma. Clin Cancer Res. 2016;22:3105–17.
Liu Z, Chaillou T, Santos Alves E, Mader T, Jude B, Ferreira DMS, et al. Mitochondrial NDUFA4L2 is a novel regulator of skeletal muscle mass and force. FASEB J. 2021;35:e22010.
Moore SJ, Murphy GG. The role of L-type calcium channels in neuronal excitability and aging. Neurobiol Learn Mem. 2020;173:107230.
Schierberl K, Hao J, Tropea TF, Ra S, Giordano TP, Xu Q, et al. Cav1.2 L-type Ca2+ channels mediate cocaine-induced GluA1 trafficking in the nucleus accumbens, a long-term adaptation dependent on ventral tegmental area Ca(v)1.3 channels. J Neurosci Off J Soc Neurosci. 2011;31:13562–75.
Shibasaki M, Kurokawa K, Ohkuma S. Upregulation of L-type Ca(v)1 channels in the development of psychological dependence. Synapse. 2010;64:440–4.
O’Brien HE, Hannon E, Hill MJ, Toste CC, Robertson MJ, Morgan JE, et al. Expression quantitative trait loci in the developing human brain and their enrichment in neuropsychiatric disorders. Genome Biol. 2018;19:194–194.
Ding J, Adiconis X, Simmons SK, Kowalczyk MS, Hession CC, Marjanovic ND, et al. Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat Biotechnol. 2020;38:737–46.
Mereu E, Lafzi A, Moutinho C, Ziegenhain C, McCarthy DJ, Alvarez-Varela A, et al. Benchmarking single-cell RNA-sequencing protocols for cell atlas projects. Nat Biotechnol. 2020;38:747–55.
Marsh SE, Walker AJ, Kamath T, Dissing-Olesen L, Hammond TR, de Soysa TY, et al. Dissection of artifactual and confounding glial signatures by single-cell sequencing of mouse and human brain. Nat Neurosci. 2022;25:306–16.
Li YE, Preissl S, Hou X, Zhang Z, Zhang K, Qiu Y, et al. An atlas of gene regulatory elements in adult mouse cerebrum. Nature. 2021;598:129–36.
Preissl S, Fang R, Huang H, Zhao Y, Raviram R, Gorkin DU, et al. Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation. Nat Neurosci. 2018;21:432–9.
Li Z, Kuppe C, Ziegler S, Cheng M, Kabgani N, Menzel S, et al. Chromatin-accessibility estimation from single-cell ATAC-seq data with scOpen. Nat Commun. 2021;12:6386.
Lal A, Chiang ZD, Yakovenko N, Duarte FM, Israeli J, Buenrostro JD. Deep learning-based enhancement of epigenomics data with AtacWorks. Nat Commun. 2021;12:1507.
Battaini F, Pascale A. Protein kinase C signal transduction regulation in physiological and pathological aging. Ann N Y Acad Sci. 2005;1057:177–92.
Huang KP. The mechanism of protein kinase C activation. Trends Neurosci. 1989;12:425–32.
Larsson C. Protein kinase C and the regulation of the actin cytoskeleton. Cell Signal. 2006;18:276–84.
Bemben MA, Nguyen TA, Li Y, Wang T, Nicoll RA, Roche KW. Isoform-specific cleavage of neuroligin-3 reduces synapse strength. Mol Psychiatry. 2019;24:145–60.
Kabir ZD, Lee AS, Rajadhyaksha AM. L-type Ca(2+) channels in mood, cognition and addiction: integrating human and rodent studies with a focus on behavioural endophenotypes. J Physiol. 2016;594:5823–37.
Pinggera A, Lieb A, Benedetti B, Lampert M, Monteleone S, Liedl KR, et al. CACNA1D de novo mutations in autism spectrum disorders activate Cav1.3 L-type calcium channels. Biol Psychiatry. 2015;77:816–22.
Kim SH, Park YR, Lee B, Choi B, Kim H, Kim CH. Reduction of Cav1.3 channels in dorsal hippocampus impairs the development of dentate gyrus newborn neurons and hippocampal-dependent memory tasks. PLoS One. 2017;12:e0181138.
Marschallinger J, Sah A, Schmuckermair C, Unger M, Rotheneichner P, Kharitonova M, et al. The L-type calcium channel Cav1.3 is required for proper hippocampal neurogenesis and cognitive functions. Cell Calcium. 2015;58:606–16.
Liu Y, Harding M, Pittman A, Dore J, Striessnig J, Rajadhyaksha A, et al. Cav1.2 and Cav1.3 L-type calcium channels regulate dopaminergic firing activity in the mouse ventral tegmental area. J Neurophysiol. 2014;112:1119–30.
Deuchars SA, Atkinson L, Brooke RE, Musa H, Milligan CJ, Batten TF, et al. Neuronal P2X7 receptors are targeted to presynaptic terminals in the central and peripheral nervous systems. J Neurosci Off J Soc Neurosci. 2001;21:7143–52.
Díaz-Hernandez M, del Puerto A, Díaz-Hernandez JI, Diez-Zaera M, Lucas JJ, Garrido JJ, et al. Inhibition of the ATP-gated P2X7 receptor promotes axonal growth and branching in cultured hippocampal neurons. J Cell Sci. 2008;121:3717–28.
Lalisse S, Hua J, Lenoir M, Linck N, Rassendren F, Ulmann L. Sensory neuronal P2RX4 receptors controls BDNF signaling in inflammatory pain. Sci Rep. 2018;8:964.
Khaira SK, Pouton CW, Haynes JM. P2X2, P2X4 and P2Y1 receptors elevate intracellular Ca2+ in mouse embryonic stem cell-derived GABAergic neurons. Br J Pharm. 2009;158:1922–31.
Sim JA, Chaumont S, Jo J, Ulmann L, Young MT, Cho K, et al. Altered hippocampal synaptic potentiation in P2X4 knock-out mice. J Neurosci Off J Soc Neurosci. 2006;26:9006–9.
Konur S, Ghosh A. Calcium signaling and the control of dendritic development. Neuron. 2005;46:401–5.
Lovisolo D, Ariano P, Distasi C. Calcium signaling in neuronal motility: pharmacological tools for investigating specific pathways. Curr Med Chem. 2012;19:5793–801.
Horigane SI, Ozawa Y, Yamada H, Takemoto-Kimura S. Calcium signalling: a key regulator of neuronal migration. J Biochem. 2019;165:401–9.
West AE, Chen WG, Dalva MB, Dolmetsch RE, Kornhauser JM, Shaywitz AJ, et al. Calcium regulation of neuronal gene expression. Proc Natl Acad Sci USA 2001;98:11024–31.
Mateos-Aparicio P, Rodríguez-Moreno A. Calcium dynamics and synaptic plasticity. Adv Exp Med Biol. 2020;1131:965–84.
Fairless R, Williams SK, Diem R. Dysfunction of neuronal calcium signalling in neuroinflammation and neurodegeneration. Cell Tissue Res. 2014;357:455–62.
Jaskova K, Pavlovicova M, Jurkovicova D. Calcium transporters and their role in the development of neuronal disease and neuronal damage. Gen Physiol Biophys. 2012;31:375–82.
Raudvere U, Kolberg L, Kuzmin I, Arak T, Adler P, Peterson H, et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 2019;47:W191–w198.
Ho SM, Hartley BJ, Flaherty E, Rajarajan P, Abdelaal R, Obiorah I, et al. Evaluating synthetic activation and repression of neuropsychiatric-related genes in hiPSC-derived NPCs, neurons, and astrocytes. Stem Cell Rep. 2017;9:615–28.
Bray NL, Pimentel H, Melsted P, Pachter L. Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol. 2016;34:525–7.
Hong S, Hwang DY, Yoon S, Isacson O, Ramezani A, Hawley RG, et al. Functional analysis of various promoters in lentiviral vectors at different stages of in vitro differentiation of mouse embryonic stem cells. Mol Ther. 2007;15:1630–9.
Buenrostro JD, Wu B, Chang HY, Greenleaf WJ. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr Protoc Mol Biol. 2015;109:21.29.21–21.29.29.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9:R137.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Feng J, Liu T, Qin B, Zhang Y, Liu XS. Identifying ChIP-seq enrichment using MACS. Nat Protoc. 2012;7:1728–40.
Herring AH. Applied Longitudinal Analysis, 2nd Edition, by Garrett M. Fitzmaurice, Nan M. Laird, and James H. Ware, John Wiley & Sons, 2011. J Biopharmaceutical Stat. 2013;23:940–1.
Laird NM, Ware JH. Random-effects models for longitudinal data. Biometrics. 1982;38:963–74.
Acknowledgements
We thank Karen F. Berman, MD and Jose Apud, MD Ph.D. at the National Institute of Mental Health for their contribution in providing patient fibroblast samples. We thank Daniel R. Weinberger, M.D. at the Lieber Institute for Brain Development for supplying patient fibroblast samples and for critical review of the manuscript.
Funding
This study was supported by MH125246 (PR), MH116442 (PR), MH118339 (J-HC). DP2MH122403 (HW), MH107884 (SC), NS121541 (SC) and NYSTEM C32607GG (SC).
Author information
Authors and Affiliations
Contributions
DL, AZ, AM, HN, G-HP, YQ, JB, JFF, PR, XZ, PS, J-HC and SC designed the experiments. DL, AZ, AM, HN, G-HP, YQ and JMP differentiated cortical interneurons and analyzed them by qPCR and immunocytochemistry. DL, JSP and FS performed arborization analysis. DL, AZ and JF performed synapse analysis. DL, AZ, OO and CA did Western blot analysis. JTC reviewed data interpretation and manuscript contents. ES provided SCZ GWAS fine mapping data. JB, JFF, PR and HW did ATAC-seq analysis. J-HC did patch clamp analysis in brain slices. XZ and PS did patch clamp analysis in culture. CY and WH did RNA-seq and ChiP-seq analysis. H-YK performed statistical analysis. DL, JB, JFF, PR, XZ, PKS, HW, J-HC and SC wrote the manuscript. PR, J-HC, HW and SC supported this study financially.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Liu, D., Zinski, A., Mishra, A. et al. Impact of schizophrenia GWAS loci converge onto distinct pathways in cortical interneurons vs glutamatergic neurons during development. Mol Psychiatry 27, 4218–4233 (2022). https://doi.org/10.1038/s41380-022-01654-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01654-z
- Springer Nature Limited
This article is cited by
-
Co-transplantation of autologous Treg cells in a cell therapy for Parkinson’s disease
Nature (2023)
-
Neurodevelopmental disturbances in schizophrenia: evidence from genetic and environmental factors
Journal of Neural Transmission (2023)