Abstract
Patients with major depressive disorder (MDD) exhibit concurrent deficits in both sensory and higher-order cognitive processing. Connectome studies have suggested a principal primary-to-transmodal gradient in functional brain networks, supporting the spectrum from sensation to cognition. However, whether this gradient structure is disrupted in patients with MDD and how this disruption associates with gene expression profiles and treatment outcome remain unknown. Using a large cohort of resting-state fMRI data from 2227 participants (1148 MDD patients and 1079 healthy controls) recruited at nine sites, we investigated MDD-related alterations in the principal connectome gradient. We further used Neurosynth, postmortem gene expression, and an 8-week antidepressant treatment (20 MDD patients) data to assess the meta-analytic cognitive functions, transcriptional profiles, and treatment outcomes related to MDD gradient alterations, respectively. Relative to the controls, MDD patients exhibited global topographic alterations in the principal primary-to-transmodal gradient, including reduced explanation ratio, gradient range, and gradient variation (Cohen’s d = 0.16–0.21), and focal alterations mainly in the primary and transmodal systems (d = 0.18–0.25). These gradient alterations were significantly correlated with meta-analytic terms involving sensory processing and higher-order cognition. The transcriptional profiles explained 53.9% variance of the altered gradient pattern, with the most correlated genes enriched in transsynaptic signaling and calcium ion binding. The baseline gradient maps of patients significantly predicted symptomatic improvement after treatment. These results highlight the connectome gradient dysfunction in MDD and its linkage with gene expression profiles and clinical management, providing insight into the neurobiological underpinnings and potential biomarkers for treatment evaluation in this disorder.
Similar content being viewed by others
Introduction
Major depressive disorder (MDD) is among the most common and burdensome psychiatric disorders globally [1]. In addition to the clinical symptoms, including low mood, loss of interest, and fatigue, neuropsychological studies suggest that patients with MDD exhibit deficits in low-level sensory processing, such as visual processing and pain perception [2,3,4], and higher-order cognitive functions, such as memory, social interaction, and reasoning [5,6,7,8]. Although many prior studies reported widespread abnormalities and treatment effects in the brain structure and function in MDD [9,10,11,12,13,14], the neurobiological mechanism underlying these sensory-cognitive deficits remains to be elucidated.
A hierarchical architecture is one of the fundamental organizational principles of the human brain, allowing information encoding and integration from sensation to cognition [15]. Using resting-state functional MRI (R-fMRI) [16] and the gradient decomposition framework [17], researchers can decompose the functional brain networks into different gradient components to capture the topography of the connectomes. Such a gradient-based method presents the relative differences in connectivity patterns among regions along a continuous spectrum [17, 18]. In healthy adults, the principal gradient spans the primary sensory networks to the transmodal default mode network (DMN), which corresponds to the hierarchical synaptic distributions in postmortem brains [15]. This spatial pattern also captures the functional spectrum of activation maps from perception to abstract cognition [17] and is spatially similar to the cortical myelin content map [19]. Although many R-fMRI studies revealed alterations in functional activity and connectivity involving the primary [9, 20,21,22] and transmodal [20, 21, 23,24,25] systems in patients with MDD, no studies reported whether and how the principal primary-to-transmodal gradient of the functional connectome is disrupted in this clinical population and whether this disruption provides valuable information for treatment outcomes. The characterization of the principal connectome gradient in MDD could provide insight into the hierarchical network mechanisms for understanding the concurrence of sensory-cognitive impairments in MDD patients and offer potential biomarkers for treatment evaluations in this disorder.
Many studies indicated that MDD is a moderately heritable disorder [26]. Genome-wide association studies identified several risk variants of genes linked to MDD, some of which play key roles in the biological functions of presynaptic differentiation and neuroinflammation [27]. Recently, the transcriptome-connectome association studies provide a critical opportunity to bridge the gap between the microlevel transcriptome profile and the macroscale brain network [28,29,30,31]. Specifically, the functional architectures of the connectomes are associated with gene expression profiles involving ion channel activity and oxidative metabolism [29, 32]. Therefore, if patients with MDD exhibit disturbances in the macroscale connectome gradient, these abnormalities might be associated with transcriptome profiles. The elucidation of such an association would enhance our understanding of the molecular genetic underpinnings of the dysfunctional connectome gradient in MDD.
To address these gaps, we employed a large multisite R-fMRI dataset of 2227 individuals and postmortem gene expression data to investigate the functional connectome gradients in MDD and establish their associations with the transcriptome profile and treatment outcomes. Specifically, we hypothesized that (i) the principal primary-to-transmodal gradient is disrupted in patients with MDD in both the global gradient topography and the focal gradient scores of primary and transmodal systems; (ii) the regions with MDD-related alterations of connectome gradient are associated with meta-analytic maps involving multiple functional domains, and with gene expression profiles enriched in particular biological processes (e.g., synapse-related functions); and (iii) the connectome gradients in MDD patients could predict clinical outcomes after antidepressant treatment.
Materials and methods
Imaging dataset and preprocessing
We used two imaging datasets with a strict quality control. Dataset 1 included 1148 patients with MDD and 1079 controls (Table 1) recruited from nine research centers through the Disease Imaging Data Archiving—Major Depressive Disorder Working Group (DIDA-MDD). Dataset 2 included 20 first-episode drug-naïve patients (Supplementary Table 1) recruited from Peking University Sixth Hospital [13]. Each patient in dataset 2 received an 8-week antidepressant treatment with escitalopram (an antidepressant of selective serotonin reuptake inhibitor, SSRI), and the treatment outcomes were recorded. All participants were diagnosed by experienced psychiatrists using structured clinical interviews. The patients met the Diagnostic and Statistical Manual of Mental Disorders-IV (DSM-IV) diagnostic criteria for MDD [33] and did not meet the criteria for any other Axis I disorder or current substance use. The severity of depression was rated using the Hamilton depression rating scale (HDRS) [34]. The controls did not have a current or lifetime history of Axis I disorder. The study was approved by the ethics committees of each research center, and written informed consent was obtained from each participant. For each participant, the R-fMRI data were obtained using 3.0-T MRI scanners (Supplementary Table 2) and were preprocessed using a standard pipeline described in our previous work [9].
Gene expression dataset and preprocessing
The microarray-based gene expression data were downloaded from the Allen Human Brain Atlas (AHBA) [28, 35]. The tissue samples in this dataset were collected from the brains of six adult donors (mean age: 42.5 years, 1 female, Supplementary Table 3). Given the large transcriptional differences between cortical and subcortical regions [35], we conducted gene expression analysis within the cortical regions. We preprocessed the gene expression microarray data according to a recommended pipeline, including mapping the samples to a cortical parcellation with 360 regions [36], probe reannotation and selection, and normalization across donors [37].
Connectome gradient analysis
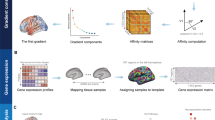
For each individual, we first constructed the voxelwise functional network (18,933 nodes) and then applied the diffusion map embedding approach [17, 38] to estimate the connectome gradient. Briefly, the top 10% connections of each node were retained, and the cosine similarity between each pair of nodes was calculated. The similarity matrix was further scaled into a normalized angle matrix to avoid negative values [39, 40]. Then, diffusion map embedding was applied to capture the gradient components explaining the variance in the connectivity pattern of the functional connectome. The resultant gradient maps were further aligned across individuals using iterative Procrustes rotation [38]. Given that the principal primary-to-transmodal gradient is closely associated with the neuronal microstructure and cognitive functions [17, 18], we primarily focused on MDD-related alterations in the principal gradients. Three global metrics, including gradient explanation ratio, range, and variation, were calculated. Finally, we utilized ComBat to correct the site effects on the gradient maps and metrics [9, 41]. The case-control differences in the connectome gradient were assessed by using a general linear model (GLM) with age and sex as covariates. To further explore the influence of sex on the group gradient differences, we used an additional GLM to examine possible sex-by-group interactions. For the global gradient metrics, the statistical significance threshold was set to P < 0.05. For the regional gradient score maps, the significance threshold was set to P < 0.001 at the voxel level, followed by Gaussian random field (GRF) correction at the cluster level of P < 0.05 [42].
Association between the functional network topology and connectome gradients in MDD
We further examined the association between MDD-related alterations in the connectome gradients and the network topology changes. Specifically, we computed the clustering coefficient (Cp) and the characteristic shortest path length (Lp) of each participant in the abovementioned whole-brain networks. Notably, Cp and Lp quantify brain network segregation and integration, respectively [43, 44]. The between-group differences in Cp and Lp were assessed using a GLM after controlling for age and sex. For the network metrics exhibiting significant between-group differences, we further examined the across-subject correlations to the global gradient metrics.
Association analysis between meta-analytic cognitive terms and MDD-related gradient alterations
We used Neurosynth (https://neurosynth.org/) [45] to examine the association between the meta-analytic cognitive terms and the MDD-related gradient alterations. The thresholded Z-maps derived from the between-group comparisons of regional gradient scores were first divided into MDD-positive (i.e., MDD > controls) and MDD-negative (i.e., MDD < controls) maps. We used the “decoder” function in Neurosynth to examine the spatial correlations between these MDD-positive or MDD-negative maps and the meta-analytic map of each term in the database. The significance of the correlation coefficients was determined by permutation tests in which spatial autocorrelations were corrected [46].
Association analysis between gene expression and MDD-related gradient alterations
We used a partial least squares (PLS) regression analysis [47] to explore the association between the transcriptional profiles and between-group differences in the principal gradient map. We first aligned the gene expression data (10,027 genes) and between-group difference Z-map of the principal gradient to a cortical parcellation atlas [36]. The gene expression data and the Z-map of the between-group differences of the principal gradient were set as the predictor variables and the response variable, respectively. We adopted a spatial autocorrelation corrected permutation test to examine whether the R2 of the PLS component was significantly greater than that expected by chance. For each significant component, we used a bootstrapping method to correct estimation error of the weight of each gene [48]. We then ranked the genes according to their corrected weights, which represent their contribution to the PLS regression component. Both the descending and ascending sequences were included in the gene enrichment analysis to identify enriched Gene Ontology terms by using GOrilla (http://cbl-gorilla.cs.technion.ac.il/) [49]. All three ontology categories, including biological process, molecular function, and cellular component, were considered. Significant enrichment was set to Benjamini-Hochberg false discovery rate (FDR)-corrected q < 0.05 [48, 50]. We also validated the sex and hemisphere effects on this connectome-transcriptome association analysis.
To further determine whether the MDD-related genes identified in previous gene studies contributed to the PLS model more than other disorder-related genes, we analyzed the disorder-related genes from the in situ hybridization (ISH) gene expression data provided by the AHBA (https://help.brain-map.org/display/humanbrain/Documentation). We selected previously reported MDD-related genes and genes related to other six brain disorders from the gene list. We compared the PLS weights between genes of MDD and those of other disorders using permutation tests. Moreover, we examined the correlation between the expression profile of each gene and the between-group Z-map of the principal gradient. The percentage of significantly correlated genes was compared between MDD and other disorders using permutation tests.
Effects of clinical factors
To investigate the effects of clinical factors on the connectome gradient, we classified the patients into different pairs of subgroups according to their clinical information, including patients with an early-onset age less than or equal to 21 years vs. greater than 21 years (this cutoff is consistent with the DSM-IV text revision definition of the early-onset of unipolar depressive disorders and previous studies [11, 51]; a cutoff age of 18 years was applied for validation), patients suffering from their first episode vs. recurrent episodes, and patients receiving medication vs. not receiving medication. Then, we compared the gradient metrics between each corresponding pair of subgroups by using a GLM. Moreover, we explored the relationships between the gradient metrics and HDRS using partial correlations with age and sex controlled.
Prediction of treatment outcome
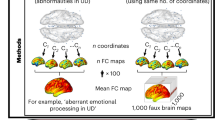
A support vector regression (SVR) was conducted to examine the prediction ability of connectome gradient for treatment response (i.e., changes of HDRS) in patients. The baseline principle gradient maps were served as predictive features, and the model was validated using an embedded 5-fold cross-validation procedure and permutation tests. For comparison purposes, we also estimated the prediction ability of the functional connectivity strength (FCS, i.e., summed connectivity for each voxel) map for treatment response.
Validation analysis
We validated our results by considering six potential influencing factors, including specific scanning site, age, Fisher’s r-to-z transformation to the correlation matrix, head motion, different gradient alignment methods, and non-parametric permutation tests. The Jaccard index was used to examine the similarity between each validation result and the main finding.
Results
The principal primary-to-transmodal gradient explained 11.9 ± 3.1% of the total connectivity variance (MDD, 11.7 ± 3.2%; HC, 12.1 ± 3.0%, Supplementary Fig. 1), which was organized along a gradual axis from the primary visual/sensorimotor networks (VIS/SMN) to the DMN (Fig. 1a), replicating the recent observation of connectome gradients from the primary to the transmodal cortices in healthy adults [17]. The spatial patterns of the group-averaged principal gradient maps were remarkably similar between the MDD and HC groups (Spearman’s ρ = 0.999, P < 0.0001, permutation tests with spatial autocorrelation corrected, Supplementary Fig. 2). A visual inspection of the histogram revealed that the extremes of the primary-to-transmodal gradient were contracted in MDD relative to the control range (Fig. 1b). Specifically, for each system, we compared the gradient score between the group-averaged maps of the MDD group and control groups by using paired t-tests across the voxel. Compared with the HC group, the MDD group had higher gradient scores in the VIS and SMN but lower in the other subnetworks (FDR-corrected q < 0.05, Fig. 1c and Supplementary Table 4).
a The principal gradient was organized along a gradual axis from the primary visual/sensorimotor networks to the default mode network. Surface rendering was generated using BrainNet Viewer (www.nitrc.org/projects/bnv/) [68] with the inflated cortical 32 K surface [36]. b Global and c system-based histograms showing that the extreme values were contracted in the patients with MDD relative to those in the healthy controls. Arrows in c indicate the direction of the significant differences between the MDD patients and controls. The whole-brain voxels were assigned to eight systems according to a 7-system cortical parcellation [69] and a subcortical system of the AAL atlas [70]. VIS visual network, SMN sensorimotor network, DAN dorsal attention network, VAN ventral attention network, SUB subcortical regions, LIB limbic network, FPN frontoparietal network, DMN default mode network.
Alterations in connectome gradients in MDD
The between-group statistical comparisons showed that the primary-to-transmodal gradient in the MDD group explained less variance in the functional connectome than that in the HC group (Cohen’s d = −0.16, P = 0.0002, Fig. 2a and Supplementary Table 5). Moreover, the patients with MDD showed a narrower range of scores (d = −0.21, P = 0.0000008) and less spatial variation (d = −0.20, P = 0.000002) than the HCs (Fig. 2a and Supplementary Table 5). Regionally, the MDD group showed lower gradient scores mostly in the DMN (76.6%) but higher scores mainly in the VIS (59.2%) and SMN (40.1%) than the HC group (absolute d = 0.18–0.25, voxel-level P < 0.001, GRF-corrected P < 0.05, Fig. 2b, Supplementary Table 6). No significant sex-group interaction effect was observed on either the global or local gradient measures. The results of the second and third gradients are shown in Supplementary Figs. 3, 4 and Supplementary Tables 5, 7, 8.
a Case-control differences in the global metrics of the principal gradient. b Voxelwise statistical comparisons between the healthy controls and patients with MDD and distribution of the regional case-control difference in different systems. Higher/lower values in MDD are presented as warm/cold colors. The statistical significance level was set as voxel-level P < 0.001 and Gaussian random field cluster level-corrected P < 0.05. c Case-control differences in the clustering coefficient (Cp) and correlations between the Cp and global metrics of the principal gradient across all patients. d Word clouds of cognitive functions associated with brain regions that exhibited higher (red) or lower (blue) gradient scores in MDD. The font size of a given cognitive term corresponds to the correlation coefficient (r) of the between-group Z-map of the principal gradient and meta-analytic map of that term generated by Neurosynth. The boxes below the word clouds indicate the correspondence between the font size and the correlation coefficient. All correlations are significant (FDR q < 0.05). ***P < 0.001. VIS visual network, SMN sensorimotor network, DAN dorsal attention network, VAN ventral attention network, SUB subcortical regions, LIB limbic network, FPN frontoparietal network, DMN default mode network.
Correlation between connectome gradients and network topology in MDD
The patients with MDD exhibited lower Cp than the HCs (d = −0.14, P = 0.0008), indicating lower functional segregation of brain networks in MDD. Moreover, Cp was correlated with the global metrics of the principal gradient across the individuals with MDD (explained ratio: r = 0.336, P < 0.001; gradient range: r = 0.257, P < 0.001; spatial variance: r = 0.356, P < 0.001, Fig. 2c).
Meta-analytic cognitive functions related to gradient alterations in MDD
The regions with higher principal gradient scores in MDD were correlated with several meta-analytic cognitive terms mainly involved in sensorial and perceptional processes, such as visual, sensory, and audiovisual processes (Fig. 2d). The regions with lower principal gradient scores in MDD were correlated with several high-order cognitive terms, including memory, social interaction, and reasoning (Fig. 2d). For details, see Supplementary Tables 9–11.
Gene expression profiles related to gradient alterations in MDD
The first two components of the PLS regression explained 51.7% of the variance in the MDD-related alterations in the principal gradient (P < 0.0001 for component 1 and P = 0.0037 for component 2, permutation tests with spatial autocorrelation corrected, Supplementary Fig. 5). Component 1 represented a transcriptional profile characterized by high expression mainly in the posterior parietal-occipital areas but low expression in the prefrontal areas (Fig. 3a). Component 2 revealed a gene expression profile with high expression mainly in the sensorimotor, visual, and temporal cortices but low expression in the frontoparietal cortices (Fig. 3b). The regional mapping of these two components was positively correlated with the Z-map of the primary-to-transmodal gradient (component 1: r = 0.574, P < 0.0001; component 2: r = 0.456, P < 0.0001, permutation tests with spatial autocorrelation corrected, Fig. 3a, b). The Gene Ontology enrichment analysis revealed that the genes ranked in ascending order of the component 1 weight were enriched in biological processes related to transsynaptic signaling and molecular function of calcium ion binding (FDR-corrected q < 0.05, Fig. 3c and Supplementary Table 12). No significant enrichment of cellular components was observed. Most terms remained significant in separate analyses of the male participants or left hemisphere (Supplementary Tables 13, 14). The genes ranked according to the weight of component 2 did not show significant enrichment. Moreover, the MDD-related genes from the ISH survey exhibited stronger PLS weights than the genes related to other disorders (except for autism, all Ps < 0.05, Fig. 3d). The expression profiles of sixty-seven percent (8 of 12) of the MDD-related genes were correlated with the between-group Z-map of the primary-to-transmodal gradient (Supplementary Table 15). This percentage was higher than that in all other disorder categories (all Ps < 0.0004, Fig. 3e).
a The first PLS component (PLS 1) identified a gene expression profile with high expression mainly in the posterior parietal-occipital areas but low expression in the prefrontal areas. The transcriptional profiles were positively correlated with the between-group Z-map of the principal gradient (permutation tests with spatial autocorrelation corrected, 10,000 times). The shadow indicates the 95% confidence intervals. Each dot represents a region. b The second PLS component (PLS 2) revealed a gene expression profile with high expression mainly in the sensorimotor, visual, and temporal cortices but low expression in the frontoparietal cortices. The transcriptional profiles were positively correlated with the between-group Z-map of the principal gradient. c Genes ranked in ascending order of the PLS 1 weight were enriched in the biological process of transsynaptic signaling and molecular function of calcium ion binding (FDR q < 0.05). d The PLS weight of predefined genes related to different disorders based on the in situ hybridization (ISH) gene list provided by the AHBA. The asterisk indicates a significant difference between MDD and a specific disorder. **P < 0.01; *P < 0.05. e The percentage of disorder-related genes showing significant correlations between the expression profile and the between-group Z-map of the principal gradient. AD Alzheimer’s disease, ASD autism, EP epilepsy, ID intellectual disability, PD Parkinson’s disease, SCZ schizophrenia.
Clinical relations to connectome gradients in MDD
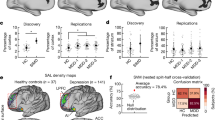
The patients with an onset in adolescence (age ≤ 21 years, N = 300) showed a narrower gradient range (d = 0.29, P = 0.0006) and smaller region variation (d = 0.19, P = 0.018) in the principal primary-to-transmodal gradient than the patients with an onset age older than 21 years (N = 292) (Fig. 4a). These results were also replicated using a cutoff age of 18 years (Supplementary Table 16). There were no significant differences in the topographic features of the principal gradient between the patients who were and those who were not taking medication or the patients in their first episode and recurrent patients (Supplementary Table 16). The voxelwise comparisons showed that there was no significant difference in the regional scores of the principal gradient between these clinical category pairs. The range of the primary gradient was correlated with HDRS across all patients (r = 0.080, P = 0.009, Fig. 4b).
a Group differences in the range and variance of the principal gradient between the patients with an onset age ≤ 21 years and those with an onset age > 21 years. ***P < 0.001; *P < 0.05. b The correlation between the range of the principal gradient and the HDRS across all patients. c Scatter plot presents the correlation between the observed HDRS change after 8-week antidepressant treatment and the predicted HDRS change derived from the SVR analysis (permutation-tests; P < 0.001). Each dot represents the data from one patient, and the dashes indicate the 95% prediction error bounds. d The absolute summed weights in 5-fold cross-validation were mapped onto the brain surface. Regions with higher/lower predictive power were colored in white/red. The radar map represents the distribution of predictive power in different systems. OA onset age, HDRS Hamilton depression rating scale, VIS visual network, SMN sensorimotor network, DAN dorsal attention network, VAN ventral attention network, SUB subcortical regions, LIB limbic network, FPN frontoparietal network, DMN default mode network.
Treatment outcome prediction of connectome gradients in MDD
The principal gradient map of the patients at baseline could significantly predict their HDRS changes after 8-week antidepressant treatment (r = 0.652, P < 0.001, one-tailed permutation test, Fig. 4c). The most contributive features were located in the DMN (percentage of total feature weights in SVR: 26.4%), SMN (15.3%), and FPN (15.2%) (Fig. 4d). In contrast, the baseline FCS map of the patients could not predict their HDRS changes (r = −0.385, P = 0.902, one-tailed permutation test).
Validation results
Overall, the MDD-related alterations in the principal connectome gradient observed using different validation strategies remained highly similar to our main findings, with a Jaccard index = 0.90 ± 0.12 for the global metrics and 0.70 ± 0.17 for the regional gradient scores (Supplementary Figs. 7, 8 and Supplementary Tables 18–20). The differences in both gradient range and variation between MDD subgroups remained significant while considering different validation strategies (Supplementary Table 21).
Discussion
In this study, we for the first time report connectome gradient dysfunction in patients with MDD and its association with transcriptional profiles and treatment outcomes. These findings advance our understanding of the neurobiological mechanisms underlying sensory-cognitive deficits and provide potential biomarkers for treatment evaluations in MDD.
In patients with MDD, the lower explained ratio of the principal gradient represents less connectivity variance in the functional connectome, while the narrower range and smaller variance indicate a less differentiated connectivity pattern between the primary and transmodal areas. The DMN regions are geometrically located in the cortex at maximal spatial distances from the primary system, corresponding to their distribution in the embedded gradient axis [17]. The maximal differences in terms of the physical distance and connectivity dissimilarity ensure the complete processing route from mapping concrete stimuli to integrating abstract conceptions [52] while increasing divergence between the primary and transmodal areas to facilitate the formation of abstract cognition by avoiding the interference of input noise [53]. Our meta-analysis confirms that MDD-related alterations in the focal gradient are located in brain areas involving multidomain cognitive functions, including sensory and perceptual processes and higher-order functions. These functions have been reported to be impaired in patients with MDD [2,3,4,5,6,7,8]. Thus, our results provide a potential linkage between connectome gradient disruption and concurrence of sensory-cognitive impairments in MDD. We also show that the topography alteration of the principal gradient was correlated with reduced functional segregation (in terms of clustering coefficient) of the brain network in MDD. This reduction in functional segregation represents a shift toward a randomized network configuration, which has been observed in previous studies of MDD [22, 54, 55]. Here, our results established the connections between alterations of the network gradient and topology in MDD. Importantly, the principal gradient scores of the DMN contributed greatly to our prediction model for antidepressant outcomes. This finding is comparable with several previous reports, demonstrating that the DMN distinguishes remitters from non-remitters in antidepressant medication [56] and predicts patients’ response to sertraline [57]. Our results extended this knowledge that not only the DMN connectivity itself but also its differentiated patterns to other systems are potential markers for clinical outcomes in depression. Notably, the principal gradient explained on average 12% of the connectome variance at the individual level in the current study. This ratio is relatively low compared to gradients decomposed from a group-level matrix [17], however, it is comparable to some individual-level analyses [58].
Our connectome-transcriptome association analysis established a link between MDD-related changes in connectome gradients and gene expression enriched in transsynaptic signaling and calcium ion binding. Transsynaptic signaling is among the most fundamental biological processes that contributes to a series of critical molecular functions, including instructing the formation of synapses and regulating synaptic plasticity [59, 60]. Disruptions in transsynaptic signaling in many key pathways can influence the formation and stability of synapses and play critical roles in the pathology of depression [61]. Studies involving postmortem tissues and rodent models suggest that exposure to chronic stress disrupts the pathway of brain-derived neurotrophic factor (BDNF)-tropomyosin-related kinase B (TrkB) receptor signaling by reducing the downstream extracellular signal-regulated kinase (ERK) and Akt pathways in the hippocampus and prefrontal cortex [62, 63]. Disturbances in these signaling pathways can decrease the expression and function of BDNF and further cause neuronal atrophy in regions implicated in depression [64]. In addition to transsynaptic signaling, calcium ion binding is a molecular function crucial for intracellular signaling. Calcium ion binding can occur in signal transduction resulting from the activation of ion channels or as a second messenger in wide-ranging physiological pathways involving synaptic plasticity. Evidence from postmortem studies suggests that the density of calbindin-immunoreactive GABAergic neurons is reduced in the dorsolateral prefrontal cortex of MDD patients [65]. Currently, measuring regional gene expression in the brain in vivo is extremely difficult. Thus, our results offer a preliminary clue in explaining the relationship between these microscale biological events and macroscopic network alterations in MDD.
Limitations and future research
First, the current work is a retrospective study in which the cognitive performance of the patients was not measured. Alternatively, we examined the associations between the gradient alteration maps and meta-analytic cognitive terms from the Neurosynth database [45]. These results should be considered as evidence of an indirect brain-cognition association in MDD. Second, all patients involved in the prediction analysis were responders to escitalopram, given that patients who had a poor response discontinued the medication or changed their treatment plans. Future studies need to include more non-responders to establish prediction models for treatment‐resistant depression. Third, several critical variables related to the pathology of MDD, such as poor sleep, social stigma, and chronic stress, were not collected in the current dataset. These variables should be recorded in future studies to assess their effects on the functional connectome in MDD. Fourth, the gene expression data from the AIBS were sampled from donors without a diagnosis of MDD, and were susceptible to intersubject variability. A larger sample of whole-brain gene expression data from patients with MDD is required to further characterize and validate the connectome-transcriptome relationship. Fifth, this study applied the ComBat method to harmonize gradient measures across sites. However, potential nonlinear and site-by-group interactive effects remain to be elucidated, for example using traveling-subject datasets and nonlinear models. Finally, the observed effect size of the between-group gradient differences is relatively small (d = 0.16–0.21) but is comparable with previous large-sample imaging studies of MDD from the ENIGMA project [11, 66] and UK Biobank [67]. The small effect size might be due to the high heterogeneity among the MDD patients. Using multivariable statistical methods and considering the subtypes of MDD could provide more effective strategies for improving the statistical power.
Data availability
The core analysis code and resulting data are publicly available at github.com/mingruixia/MDD_ConnectomeGradient. For details of Materials and Methods, see Supplementary Material.
References
DALYs GBD, Collaborators H. Global, regional, and national disability-adjusted life-years (DALYs) for 359 diseases and injuries and healthy life expectancy (HALE) for 195 countries and territories, 1990-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet. 2018;392:1859–922.
Bubl E, Kern E, Ebert D, Bach M, Tebartz van Elst L. Seeing gray when feeling blue? Depression can be measured in the eye of the diseased. Biol Psychiatry. 2010;68:205–8.
Adler G, Gattaz WF. Pain perception threshold in major depression. Biol Psychiatry. 1993;34:687–9.
Fitzgerald PJ. Gray colored glasses: is major depression partially a sensory perceptual disorder? J Affect Disord. 2013;151:418–22.
Millan MJ, Agid Y, Brune M, Bullmore ET, Carter CS, Clayton NS, et al. Cognitive dysfunction in psychiatric disorders: characteristics, causes and the quest for improved therapy. Nat Rev Drug Discov. 2012;11:141–68.
Dillon DG, Pizzagalli DA. Mechanisms of Memory Disruption in Depression. Trends Neurosci. 2018;41:137–49.
Kupferberg A, Bicks L, Hasler G. Social functioning in major depressive disorder. Neurosci Biobehav Rev. 2016;69:313–32.
Huys QJ, Daw ND, Dayan P. Depression: a decision-theoretic analysis. Annu Rev Neurosci. 2015;38:1–23.
Xia M, Si T, Sun X, Ma Q, Liu B, Wang L, et al. Reproducibility of functional brain alterations in major depressive disorder: Evidence from a multisite resting-state functional MRI study with 1,434 individuals. Neuroimage. 2019;189:700–14.
Cheng W, Rolls ET, Qiu J, Liu W, Tang Y, Huang CC, et al. Medial reward and lateral non-reward orbitofrontal cortex circuits change in opposite directions in depression. Brain. 2016;139:3296–309.
Schmaal L, Hibar DP, Samann PG, Hall GB, Baune BT, Jahanshad N, et al. Cortical abnormalities in adults and adolescents with major depression based on brain scans from 20 cohorts worldwide in the ENIGMA Major Depressive Disorder Working Group. Mol Psychiatry. 2017;22:900–9.
Singh MK, Kesler SR, Hadi Hosseini SM, Kelley RG, Amatya D, Hamilton JP, et al. Anomalous gray matter structural networks in major depressive disorder. Biol Psychiatry. 2013;74:777–85.
Wang L, Xia M, Li K, Zeng Y, Su Y, Dai W, et al. The effects of antidepressant treatment on resting-state functional brain networks in patients with major depressive disorder. Hum Brain Mapp. 2015;36:768–78.
Perrin JS, Merz S, Bennett DM, Currie J, Steele DJ, Reid IC, et al. Electroconvulsive therapy reduces frontal cortical connectivity in severe depressive disorder. Proc Natl Acad Sci USA. 2012;109:5464–8.
Mesulam MM. From sensation to cognition. Brain. 1998;121:1013–52.
Biswal B, Yetkin FZ, Haughton VM, Hyde JS. Functional connectivity in the motor cortex of resting human brain using echo-planar MRI. Magn Reson Med. 1995;34:537–41.
Margulies DS, Ghosh SS, Goulas A, Falkiewicz M, Huntenburg JM, Langs G, et al. Situating the default-mode network along a principal gradient of macroscale cortical organization. Proc Natl Acad Sci USA. 2016;113:12574–9.
Huntenburg JM, Bazin PL, Margulies DS. Large-Scale Gradients in Human Cortical Organization. Trends Cogn Sci. 2018;22:21–31.
Huntenburg JM, Bazin PL, Goulas A, Tardif CL, Villringer A, Margulies DS. A Systematic Relationship Between Functional Connectivity and Intracortical Myelin in the Human Cerebral Cortex. Cereb Cortex. 2017;27:981–97.
Gong Q, He Y. Depression, neuroimaging and connectomics: a selective overview. Biol Psychiatry. 2015;77:223–35.
Ma Q, Tang Y, Wang F, Liao X, Jiang X, Wei S, et al. Transdiagnostic Dysfunctions in Brain Modules Across Patients with Schizophrenia, Bipolar Disorder, and Major Depressive Disorder: A Connectome-Based Study. Schizophr Bull. 2020;46:699–712.
Xia M, Womer FY, Chang M, Zhu Y, Zhou Q, Edmiston EK, et al. Shared and Distinct Functional Architectures of Brain Networks Across Psychiatric Disorders. Schizophr Bull. 2019;45:450–63.
Yan CG, Chen X, Li L, Castellanos FX, Bai TJ, Bo QJ, et al. Reduced default mode network functional connectivity in patients with recurrent major depressive disorder. Proc Natl Acad Sci USA. 2019;116:9078–83.
Yu M, Linn KA, Shinohara RT, Oathes DJ, Cook PA, Duprat R, et al. Childhood trauma history is linked to abnormal brain connectivity in major depression. Proc Natl Acad Sci USA. 2019;116:8582–90.
Kaiser RH, Andrews-Hanna JR, Wager TD, Pizzagalli DA. Large-Scale Network Dysfunction in Major Depressive Disorder: A Meta-analysis of Resting-State Functional Connectivity. JAMA Psychiatry. 2015;72:603–11.
Flint J, Kendler KS. The Genetics of Major Depression. Neuron. 2014;81:1214.
Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne EM, Abdellaoui A, et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet. 2018;50:668–81.
Hawrylycz MJ, Lein ES, Guillozet-Bongaarts AL, Shen EH, Ng L, Miller JA, et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature. 2012;489:391–9.
Richiardi J, Altmann A, Milazzo AC, Chang C, Chakravarty MM, Banaschewski T, et al. BRAIN NETWORKS. Correlated gene expression supports synchronous activity in brain networks. Science. 2015;348:1241–4.
Fornito A, Arnatkeviciute A, Fulcher BD. Bridging the Gap between Connectome and Transcriptome. Trends Cogn Sci. 2019;23:34–50.
Liu J, Xia M, Wang X, Liao X, He Y. The spatial organization of the chronnectome associates with cortical hierarchy and transcriptional profiles in the human brain. Neuroimage. 2020;222:117296.
Vertes PE, Rittman T, Whitaker KJ, Romero-Garcia R, Vasa F, Kitzbichler MG, et al. Gene transcription profiles associated with inter-modular hubs and connection distance in human functional magnetic resonance imaging networks. Philos Trans R Soc Lond B Biol Sci. 2016;371:20150362.
First M, Spitzer R, Gibbon M, Williams J. Structured Clinical Interview for DSM-IV Axis I Disorders. Washington, DC: American Psychiatric Publishing; 1997.
Williams JB. A structured interview guide for the Hamilton Depression Rating Scale. Arch Gen Psychiatry. 1988;45:742–7.
Hawrylycz M, Miller JA, Menon V, Feng D, Dolbeare T, Guillozet-Bongaarts AL, et al. Canonical genetic signatures of the adult human brain. Nat Neurosci. 2015;18:1832–44.
Glasser MF, Coalson TS, Robinson EC, Hacker CD, Harwell J, Yacoub E, et al. A multi-modal parcellation of human cerebral cortex. Nature. 2016;536:171–8.
Arnatkeviciute A, Fulcher BD, Fornito A. A practical guide to linking brain-wide gene expression and neuroimaging data. Neuroimage. 2019;189:353–67.
Hong SJ, Vos de Wael R, Bethlehem RAI, Lariviere S, Paquola C, Valk SL, et al. Atypical functional connectome hierarchy in autism. Nat Commun. 2019;10:1022.
Lariviere S, Vos de Wael R, Hong SJ, Paquola C, Tavakol S, Lowe AJ, et al. Multiscale Structure-Function Gradients in the Neonatal Connectome. Cereb Cortex. 2020;30:47–58.
Paquola C, Vos De Wael R, Wagstyl K, Bethlehem RAI, Hong SJ, Seidlitz J, et al. Microstructural and functional gradients are increasingly dissociated in transmodal cortices. PLoS Biol. 2019;17:e3000284.
Yu M, Linn KA, Cook PA, Phillips ML, McInnis M, Fava M, et al. Statistical harmonization corrects site effects in functional connectivity measurements from multi-site fMRI data. Hum Brain Mapp. 2018;39:4213–27.
Eklund A, Nichols TE, Knutsson H. Cluster failure: Why fMRI inferences for spatial extent have inflated false-positive rates. Proc Natl Acad Sci USA. 2016;113:7900–5.
Watts DJ, Strogatz SH. Collective dynamics of ‘small-world’ networks. Nature. 1998;393:440–2.
Liao X, Vasilakos AV, He Y. Small-world human brain networks: Perspectives and challenges. Neurosci Biobehav Rev. 2017;77:286–300.
Yarkoni T, Poldrack RA, Nichols TE, Van Essen DC, Wager TD. Large-scale automated synthesis of human functional neuroimaging data. Nat Methods. 2011;8:665–70.
Burt JB, Helmer M, Shinn M, Anticevic A, Murray JD. Generative modeling of brain maps with spatial autocorrelation. Neuroimage. 2020;220:117038.
Abdi H. Partial least squares regression and projection on latent structure regression (PLS Regression). Wiley Interdiscip Rev: Computational Stat. 2010;2:97–106.
Whitaker KJ, Vertes PE, Romero-Garcia R, Vasa F, Moutoussis M, Prabhu G, et al. Adolescence is associated with genomically patterned consolidation of the hubs of the human brain connectome. Proc Natl Acad Sci USA. 2016;113:9105–10.
Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinforma. 2009;10:48.
Romero-Garcia R, Warrier V, Bullmore ET, Baron-Cohen S, Bethlehem RAI. Synaptic and transcriptionally downregulated genes are associated with cortical thickness differences in autism. Mol psychiatry. 2019;24:1053–64.
Benazzi F. Classifying mood disorders by age-at-onset instead of polarity. Prog Neuro-Psychopharmacol Biol Psychiatry. 2009;33:86–93.
Smallwood J, Bernhardt BC, Leech R, Bzdok D, Jefferies E, Margulies DS. The default mode network in cognition: a topographical perspective. Nat Rev Neurosci. 2021;22:503–13.
Buckner RL, Krienen FM. The evolution of distributed association networks in the human brain. Trends Cogn Sci. 2013;17:648–65.
Zhang J, Wang J, Wu Q, Kuang W, Huang X, He Y, et al. Disrupted brain connectivity networks in drug-naive, first-episode major depressive disorder. Biol Psychiatry. 2011;70:334–42.
Borchardt V, Lord AR, Li M, van der Meer J, Heinze HJ, Bogerts B, et al. Preprocessing strategy influences graph-based exploration of altered functional networks in major depression. Hum Brain Mapp. 2016;37:1422–42.
Korgaonkar MS, Goldstein-Piekarski AN, Fornito A, Williams LM. Intrinsic connectomes are a predictive biomarker of remission in major depressive disorder. Mol Psychiatry. 2020;25:1537–49.
Chin Fatt CR, Jha MK, Cooper CM, Fonzo G, South C, Grannemann B, et al. Effect of Intrinsic Patterns of Functional Brain Connectivity in Moderating Antidepressant Treatment Response in Major Depression. Am J Psychiatry. 2020;177:143–54.
Bethlehem RAI, Paquola C, Seidlitz J, Ronan L, Bernhardt B, Consortium CC, et al. Dispersion of functional gradients across the adult lifespan. Neuroimage. 2020;222:117299.
Fossati M, Assendorp N, Gemin O, Colasse S, Dingli F, Arras G, et al. Trans-Synaptic Signaling through the Glutamate Receptor Delta-1 Mediates Inhibitory Synapse Formation in Cortical Pyramidal. Neurons Neuron. 2019;104:1081–94 e1087.
de Wit J, Ghosh A. Specification of synaptic connectivity by cell surface interactions. Nat Rev Neurosci. 2016;17:22–35.
Duman RS, Aghajanian GK. Synaptic dysfunction in depression: potential therapeutic targets. Science. 2012;338:68–72.
Duman RS, Voleti B. Signaling pathways underlying the pathophysiology and treatment of depression: novel mechanisms for rapid-acting agents. Trends Neurosci. 2012;35:47–56.
Duric V, Banasr M, Licznerski P, Schmidt HD, Stockmeier CA, Simen AA, et al. A negative regulator of MAP kinase causes depressive behavior. Nat Med. 2010;16:1328–32.
Krishnan V, Nestler EJ. The molecular neurobiology of depression. Nature. 2008;455:894–902.
Rajkowska G, O’Dwyer G, Teleki Z, Stockmeier CA, Miguel-Hidalgo JJ. GABAergic neurons immunoreactive for calcium binding proteins are reduced in the prefrontal cortex in major depression. Neuropsychopharmacol: Off Publ Am Coll Neuropsychopharmacol. 2007;32:471–82.
Schmaal L, Veltman DJ, van Erp TG, Samann PG, Frodl T, Jahanshad N, et al. Subcortical brain alterations in major depressive disorder: findings from the ENIGMA Major Depressive Disorder working group. Mol Psychiatry. 2016;21:806–12.
Anderson KM, Collins MA, Kong R, Fang K, Li J, He T, et al. Convergent molecular, cellular, and cortical neuroimaging signatures of major depressive disorder. Proc Natl Acad Sci USA. 2020;117:25138–49.
Xia M, Wang J, He Y. BrainNet Viewer: a network visualization tool for human brain connectomics. PLoS One. 2013;8:e68910.
Yeo BT, Krienen FM, Sepulcre J, Sabuncu MR, Lashkari D, Hollinshead M, et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J Neurophysiol. 2011;106:1125–65.
Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N, et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage. 2002;15:273–89.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82071998 and 81671767 to MX; 82021004, 81620108016, and 91432115 to YH; 81171286 and 91232714 to LL; 31771231 and 31271087 to JQ; 81571331 to FW; 81271499 and 81571311 to YT; 81920108019 and 81771344 to SQ; 81630031 to TS; 81621003 to QG; 81660237 to XX), the Beijing Nova Program (Z191100001119023 to MX), Fundamental Research Funds for the Central Universities (2020NTST29 to MX), the National Key R&D Program of China (2018YFA0701400 to YH), the Changjiang Scholar Professorship Award (T2015027 to YH), the National Science and Technologic Program of China (2015BAI13B02 to LL), the National Basic Research Program of China (2013CB835100 LL), the Natural Science Foundation of Chongqing (cstc2019jcyj-msxmX0520 to JQ), the National High Tech Development Plan (863) (2015AA020513 to FW), the Medical Science and Technology Research Project of Henan Province (201701011 to JC), and Shanghai Science and Technology Innovation Plan (17JC1404105 and 17JC1404101 to C-PL). The authors thank the Allen Institute for Brain Science for providing the gene expression data.
Author information
Authors and Affiliations
Consortia
Contributions
MX and YH designed research; XW, DW, YChen, BL, C-CH, YZ, YW, TC, YCheng, XX, QG, TS, SQ, C-PL, JC, YT, FW, JQ, PX, and LL collected data; MX, XS, and QM performed data quality control; MX and JL analyzed data; and MX, JL, AM, and YH, wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Xia, M., Liu, J., Mechelli, A. et al. Connectome gradient dysfunction in major depression and its association with gene expression profiles and treatment outcomes. Mol Psychiatry 27, 1384–1393 (2022). https://doi.org/10.1038/s41380-022-01519-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01519-5
- Springer Nature Limited
This article is cited by
-
Is depression a global brain disorder with topographic dynamic reorganization?
Translational Psychiatry (2024)
-
Transcriptional patterns of amygdala functional connectivity in first-episode, drug-naïve major depressive disorder
Translational Psychiatry (2024)
-
Reshaping the Cortical Connectivity Gradient by Long-Term Cognitive Training During Development
Neuroscience Bulletin (2024)
-
Tau follows principal axes of functional and structural brain organization in Alzheimer’s disease
Nature Communications (2024)
-
Brain-wide changes in excitation-inhibition balance of major depressive disorder: a systematic review of topographic patterns of GABA- and glutamatergic alterations
Molecular Psychiatry (2023)