Abstract
Acute myeloid leukemia (AML) is an aggressive disease for which only few targeted therapies are available. Using high-throughput RNA interference (RNAi) screening in AML cell lines, we identified LIM kinase 1 (LIMK1) as a potential novel target for AML treatment. High LIMK1 expression was significantly correlated with shorter survival of AML patients and coincided with FLT3 mutations, KMT2A rearrangements, and elevated HOX gene expression. RNAi- and CRISPR-Cas9-mediated suppression as well as pharmacologic inhibition of LIMK1 and its close homolog LIMK2 reduced colony formation and decreased proliferation due to slowed cell-cycle progression of KMT2A-rearranged AML cell lines and patient-derived xenograft (PDX) samples. This was accompanied by morphologic changes indicative of myeloid differentiation. Transcriptome analysis showed upregulation of several tumor suppressor genes as well as downregulation of HOXA9 targets and mitosis-associated genes in response to LIMK1 suppression, providing a potential mechanistic basis for the anti-leukemic phenotype. Finally, we observed a reciprocal regulation between LIM kinases (LIMK) and CDK6, a kinase known to be involved in the differentiation block of KMT2A-rearranged AML, and addition of the CDK6 inhibitor palbociclib further enhanced the anti-proliferative effect of LIMK inhibition. Together, these data suggest that LIMK are promising targets for AML therapy.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is an aggressive malignancy with an average five-year survival of ~28% [1], although prognosis varies considerably between genetic subtypes [2]. Despite an improved understanding of AML pathogenesis, standard treatment still relies on cytotoxic chemotherapy, which induces complete remission in up to 85% of patients [3]; however, 50% of younger and 80–90% of older patients relapse within one year [4]. Furthermore, older individuals, who constitute the majority of AML patients, are often not eligible for intensive treatment [5]. As a consequence, targeted therapies that selectively inhibit malignant cells, but spare healthy tissue, are urgently needed. Recent approvals of agents such as the FLT3 inhibitor midostaurin for FLT3-mutant AML [6] or the IDH inhibitors ivosidenib and enasidenib for IDH-mutant AML [7, 8] highlight the promise of targeted treatment approaches. However, a significant proportion of patients are refractory or develop resistance to these inhibitors. In addition, ~50% of patients harbor neither FLT3 nor IDH mutations [9], underscoring that additional molecular targets are required, especially in the context of currently “undruggable” driver alterations, such as NPM1, TP53, and RAS mutations or many oncogenic fusion proteins.
LIM kinase 1 and 2 (LIMK1/LIMK2) are ubiquitously expressed dual-specificity kinases that are primarily known for their role in regulating cell motility via phosphorylation of the actin depolymerization factor cofilin 1 (CFL1) at serine 3 (ref. 10, 11). LIM kinases (LIMK) have been shown to drive tumor cell migration, invasion, and metastasis formation in several epithelial cancers [12,13,14,15]. In addition, they can promote proliferation and tumor growth [15, 16], at least partially by regulating microtubule dynamics and mitotic spindle formation [17,18,19]. Given these pro-tumorigenic functions, several LIMK inhibitors have been developed; however, none of them have been sufficiently optimized to enter clinical trials [20]. A potential role of LIMK in leukemia, and AML in particular, is less clear. In 2014, Oku et al. showed that pharmacologic inhibition of LIMK and their upstream regulator Rho Associated Coiled-Coil Containing Protein Kinase (ROCK) reduced proliferation and induced apoptosis in T-cell leukemia cells [21]. Recently, the RHOB small GTPase was found overexpressed in children with relapsed t(8;21)-positive AML, which resulted in elevated cofilin phosphorylation – presumably in a LIMK-dependent manner – and was paralleled by enhanced migration and drug resistance [22].
In this study, we used RNA interference (RNAi) screening to identify LIMK as novel dependencies of AML cell lines and PDX samples, and found that high LIMK1 expression correlates with disease-defining genetic aberrations and poor patient outcome. Furthermore, we observed a reciprocal regulation between LIMK and CDK6, providing a rationale for a combination therapy approach.
Results
LIMK1 expression correlates with AML patient outcome and genetic subtypes
To discover novel essential genes in AML, we analyzed RNAi screens previously conducted in three AML cell lines that harbor no immediately targetable genetic alterations (U937, HL-60, and OCI-AML3) [23]. We identified 13 genes whose targeting by at least three independent short hairpin RNAs (shRNAs) reduced viability in all three cell lines (Fig. 1a). From these candidates, we excluded one core-essential gene (RIOK1) as defined previously [24, 25]. To eliminate additional false positives, we evaluated, using publicly available RNA sequencing (RNA-seq) data [26, 27], whether the remaining candidate genes were expressed in the screening cell lines. Except for EGFR, all candidate genes showed sufficient expression levels across all three cell lines (Fig. 1a), indicating that our analysis pipeline identified a high-confidence gene set. LIMK1 was the only gene that scored with at least four different shRNAs in each cell line, and we found a significant association between high expression and shorter overall survival for LIMK1 (Fig. 1b), but not the other candidate genes (Fig. S1A), in The Cancer Genome Atlas (TCGA) AML dataset [9]. Taking further into consideration that tumorigenic functions of LIMK had been reported in several cancer entities, whereas their role had not been investigated in AML, and that pre-clinical small-molecule inhibitors were already available, we prioritized LIMK1 for further validation.
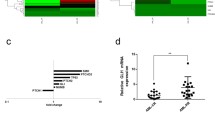
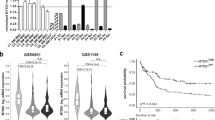
a Results of RNAi screens in HL-60, U937, and OCI-AML3 cell lines. The Venn diagram shows the number of candidate essential genes scoring with at least three independent shRNAs in each cell line. The heatmaps display the number of scoring shRNAs (blue) and mRNA expression using RNA-seq data from the Cancer Cell Line Encyclopedia (red; downloaded from www.depmap.org, version 19Q1) of twelve common candidate genes. TPM, transcripts per million. b Survival of AML patients from the TCGA study (n = 132) with LIMK1 mRNA expression above the median (LIMK1high) and below the median (LIMK1low). Statistical significance was assessed by log-rank test. c LIMK1 mRNA expression according to cytogenetic subtype from the Leukemia MILE study (MLL = KMT2A). Data were downloaded from www.bloodspot.eu. Black bars indicate the median. Statistical significance was assessed by one-way ANOVA with Dunnett’s correction for multiple comparisons (each condition vs. healthy bone marrow). The non-indicated comparisons were not significant (P > 0.05). ****P < 0.0001. d Multivariable regression analysis with backward variable selection to assess associations between recurrent AML driver mutations and LIMK1 expression. Coef, coefficient, S.E., standard error. e Enrichment of gene expression signatures associated with rearranged KMT2A (MLL = KMT2A, left panel) and mutant NPM1 (right panel) in TCGA AML patients with high LIMK1 expression. The analysis was performed with the 10% highest (n = 13) and 10% lowest (n = 13) LIMK1-expressing TCGA AML patients.
Next, we investigated if high LIMK1 expression is associated with specific AML subtypes. We therefore examined data from the Microarray Innovations in Leukemia (MILE) study [28,29,30] and observed significantly higher LIMK1 mRNA levels in normal karyotype and KMT2A-rearranged AML, but not in other genetic subtypes, compared to healthy bone marrow samples (Fig. 1c). RNA-seq data from the TCGA AML cohort confirmed this association (Fig. S1B). Since normal karyotype AML is frequently characterized by recurrent driver alterations, such as NPM1, FLT3, DNMT3A, IDH1/2, TET2, CEBPA, RAS, and RUNX1 mutations, we queried the TCGA AML dataset to evaluate whether the mutation status of these genes is linked to LIMK1 expression. Multivariable regression analysis confirmed the association between normal karyotype and high LIMK1 levels, and further revealed a significant correlation between FLT3 mutations and high LIMK1 expression, whereas IDH1 or IDH2 mutations showed a trend towards lower LIMK1 levels (Fig. 1d). Within the subset of FLT3-mutated samples (n = 35), additional DNMT3A and NPM1 mutations appeared to correlate with further increased LIMK1 levels (Fig. S1C), but these observations were not statistically significant, potentially due to low case numbers.
In a complementary approach to investigate if certain driver alterations are associated with LIMK1 expression, we performed Gene Set Enrichment Analysis (GSEA) [31] following differential gene expression analysis of TCGA AML patients with high and low LIMK1 expression. This revealed an enrichment of gene signatures driven by KMT2A rearrangements or mutant NPM1 in samples with high LIMK1 expression (Fig. 1e), suggesting that NPM1 mutations may in fact correlate with LIMK1 levels. Together, these results show that AML driver mutations affecting KMT2A, FLT3, and possibly NPM1 are associated with high LIMK1 expression.
LIMK depletion reduces proliferation and colony formation of KMT2A-rearranged and FLT3-mutant AML cells
Since we observed high LIMK1 expression in KMT2A-rearranged AML patient samples, we directed the subsequent functional characterization of LIMK1 to this aggressive subtype. To determine the dependency of KMT2A-rearranged cell lines on LIMK1, we performed competitive proliferation assays, in which cell lines were transduced with LIMK1- or non-targeting shRNA vectors co-expressing GFP, and the number of transduced, GFP-positive cells relative to non-transduced cells was monitored over time using flow cytometry. We found that LIMK1 depletion conferred a significant proliferation disadvantage compared to non-transduced cells in three KMT2A-rearranged cell lines, with the more efficient shRNA (shLIMK1_2) resulting in stronger cell reduction (Figs. 2a and S2A). Similarly, we observed a significant decrease in colony formation upon LIMK1 suppression (Figs. 2b and S2B). To verify these findings, we generated LIMK1 knockout cells by expanding single-cell clones from THP-1 cells transduced with three independent single guide RNAs (sgRNAs) targeting either LIMK1 or mCherry as a control. In accordance with the RNAi experiments, proliferation and colony formation were significantly reduced in LIMK1 knockout clones compared to the controls (Figs. 2c, d and S2C). In line with these observations, 5-ethynyl-2’-deoxyuridine (EdU) incorporation was decreased upon LIMK1 suppression, suggesting slowed cell-cycle progression (Fig. S2D). In addition, we observed distinct morphologic changes, such as vacuolization and increased cell size and cytoplasm-to-nucleus ratio, which are indicative of enhanced myeloid differentiation (Fig. 2e). Apoptotic cell death was not induced as neither PARP cleavage (Fig. S2E) nor an increase of annexin V-positive cells (data not shown) was detected.
a Competition assay of three KMT2A-rearranged AML cell lines transduced with two LIMK1-targeting shRNAs or NTC co-expressing GFP. GFP-positive cells were monitored by flow cytometry for 17 days starting three days post-transduction. Statistical analysis was performed on log-transformed data for the final day of the experiment by unpaired t-tests. *P < 0.05, ***P < 0.001, ****P < 0.0001. b Colony formation in methylcellulose after nine days of KMT2A-rearranged AML cell lines transduced with two LIMK1-targeting shRNAs or NTC. Statistical analysis was performed by unpaired t-tests. *P < 0.05, ***P < 0.001. ns, not significant (P > 0.05). c Proliferation determined by manual cell counting of THP-1-derived LIMK1 knockout and control (sgmCherry) clones. Each LIMK1 knockout clone was generated with a different sgRNA. Error bars indicate the standard deviation of three individual clones. Lines indicate the fitted model for exponential growth. Statistical significance for the corresponding proliferation constants was assessed by an unpaired t-test. ***P < 0.001. d Colony formation in methylcellulose of THP-1-derived LIMK1 knockout and control (sgmCherry) clones after ten days. Each LIMK1 knockout clone was generated with a different sgRNA. Statistical significance was assessed by an unpaired t-test. *P < 0.05. e May-Grünwald-Giemsa-stained cytospin preparations of KMT2A-rearranged AML cell lines eight days after transduction with two LIMK1-targeting shRNAs or NTC. Original magnification, x40. Insets show 1.5-fold magnified details of the corresponding photographs. f Competition assay of THP-1 cells transduced with two LIMK2-targeting shRNAs or NTC co-expressing GFP. GFP-positive cells were monitored for 17 days starting three days post-transduction. Statistical analysis was performed on log-transformed data for the final day of the experiment by unpaired t-tests. ****P < 0.0001. g Colony formation in methylcellulose of THP-1 cells transduced with LIMK2-targeting shRNAs or NTC after nine days. Statistical analysis was performed by unpaired t-tests. *P < 0.05. h Colony formation in methylcellulose after eleven days of two KMT2A-rearranged PDX samples transduced with LIMK1- or LIMK2-targeting shRNAs or NTC. For PDX AML-388, two technical replicates are shown. Statistical analysis was performed by unpaired t-tests. *P < 0.05, **P < 0.01.
Since LIMK2 is a close homolog of LIMK1, we evaluated if LIMK2 suppression results in a similar phenotype as LIMK1 depletion. In the initial RNAi screen, LIMK2 was only represented by three shRNAs and therefore did not score due to our stringent selection criteria. Similar to LIMK1 depletion, knockdown of LIMK2 significantly decreased proliferation and colony formation of THP-1 cells and induced morphologic changes (Figs. 2f, g and S2F, G), indicating that both LIMK family members are equally important for AML cells. To further substantiate our findings, we suppressed LIMK1 and LIMK2 in two patient-derived xenograft (PDX) AML samples harboring KMT2A translocations and measured colony formation in methylcellulose. In accordance with our observations in cell lines, loss of LIMK1 or LIMK2 resulted in significantly decreased colony formation in primary patient samples (Fig. 2h).
Finally, we investigated whether cell lines harboring genetic alterations other than KMT2A rearrangements would also respond to LIMK inhibition, with a focus on FLT3 mutations because of their association with high LIMK1 levels. We performed competition assays in three FLT3-mutant cell lines as well as K562 cells (driven by BCR-ABL1), and found that all FLT3-mutant lines were highly sensitive to depletion of LIMK1 or LIMK2 (Fig. S2H). K562 cells were not affected, indicating that LIMK suppression is not generally toxic, which is an important consideration with regard to possible clinical applications.
Together, these data indicate that LIMK1 and LIMK2 are involved in maintaining leukemic characteristics of AML cells of different genetic backgrounds, including KMT2A-rearrangements and FLT3 mutations, corroborating that LIMK represent promising therapeutic targets for a broad spectrum of AML patients.
LIMK1 regulates expression of tumor suppressor and HOXA9 target genes
To obtain insights into the molecular function of LIMK1 in AML cells, we determined transcriptional changes by RNA-seq in THP-1 cells transduced with a non-targeting control (NTC) or two independent shRNAs targeting LIMK1. We detected 362 deregulated genes, many of which are involved in oncogenic signaling and tumorigenesis [32,33,34,35,36,37,38,39,40,41,42,43,44,45,46] (Fig. 3a, Table S1). Among the top upregulated genes upon LIMK1 knockdown were several known tumor suppressor genes, including EGR1 (ref. 36,37,38), BTG2 (ref. 39,40,41), and BIN1 (ref. 42,43,44). In support of its tumor-suppressive function, high EGR1 expression correlated with better survival of AML patients, although this association was not statistically significant (Fig. 3b). On the other hand, LIMK1 suppression resulted in downregulation of LSP1, which has recently been implicated in leukemogenesis [45], and PAQR8, which has been described in the context of ovarian and endometrial cancer [46, 47]. The link between these two genes and LIMK1 was further substantiated by the observation that their expression was significantly higher in LIMK1high compared to LIMK1low patient samples (Fig. 3c). Furthermore, high expression of LSP1 or PAQR8 was significantly correlated with shorter survival of AML patients (Fig. 3d), indicating a pro-leukemogenic role of these genes.
a MA plots showing deregulated genes in THP-1 cells eight days after transduction with two independent LIMK1-targeting shRNAs in biological duplicates followed by RNA-seq. Colors indicate the significance level (red: P < 0.05, gray: P > 0.05). LIMK1, as well as selected genes known to play a role in tumorigenesis and oncogenic signaling, are highlighted. One gene (SMPDL3B; x = 1.37, y = −5.28, P = 0.028) is not included in the right panel to allow increased resolution of the plot. nrc, normalized read count. b Survival of TCGA AML patients with high (expression above median) and low (expression below median) EGR1 mRNA levels (n = 132). Statistical significance was assessed by log-rank test. c TCGA AML RNA-seq data for LSP1 and PAQR8 depending on LIMK1 expression status. The analysis was performed with the 10% highest (n = 13) and 10% lowest (n = 13) LIMK1-expressing patients. Statistical significance was assessed by an unpaired t-test. *P < 0.05, **P < 0.01. TPM, transcripts per million. d Survival of TCGA AML patients with high (above median) and low (below median) expression of LSP1 or PAQR8 (n = 132). Statistical significance was assessed by log-rank test. e Enrichment of HOXA9-related gene sets in the transcriptome of THP-1 cells after LIMK1 knockdown (shown in a). Genes were pre-ranked based on the mean log2(normalized read count shLIMK1/normalized read count NTC) of both shRNAs. KD, knockdown. f High expression of HOXA and HOXB genes in LIMK1high (n = 13) vs. LIMK1low (n = 13) TCGA AML patients. Significance levels (adjusted P value) are indicated by red or gray color, respectively. Missing HOX genes did not pass the expression cut-off (TPM > 0.41).
In consonance with the anti-proliferative effect of LIMK1 knockdown, GSEA revealed downregulation of genes involved in cell-cycle progression (Fig. S3A). In particular, genes associated with centrosome regulation and sister chromatid segregation and cohesion were significantly depleted following LIMK1 knockdown, suggesting induction of mitotic defects. GSEA further showed deregulation of HOXA9-associated gene sets upon LIMK1 depletion, as genes induced by conditional expression of Hoxa9 and Meis1 in hematopoietic precursor cells [48] were downregulated in LIMK1 knockdown samples (Fig. 3e, left panel). Similarly, genes showing reduced expression upon HOXA9 knockdown [49] were downregulated upon LIMK1 suppression (Fig. 3e, right panel), suggesting that LIMK1 promotes transcriptional activity of HOXA9. In support of a potential link between LIMK1 and HOX genes, differential gene expression analysis of AML patient samples from the TCGA study showed significantly higher expression of nearly all HOXA and HOXB genes in LIMK1high samples compared to LIMK1low samples (Figs. 3f and S3B, Table S2), and HOXA9 and LIMK1 mRNA levels were significantly correlated (Fig. S3C). However, mRNA levels of HOXA9 and its established cofactors were unchanged upon LIMK1 suppression according to our RNA-seq data (Fig. S3D), suggesting that LIMK1 might promote HOXA9 function by a post-transcriptional mechanism.
Together, these data indicate that LIMK1 promotes leukemogenesis by inhibiting the expression of tumor suppressor genes as well as by driving LSP1 and PAQR8 expression and promoting the activity of HOX transcription factors.
AML cells show reciprocal regulation of LIMK and CDK6
We previously identified an anti-differentiation function of CDK6 in KMT2A-rearranged AML cells, and genetic and pharmacologic CDK6 inhibition results in myeloid differentiation [23]. Since high LIMK1 expression is also associated with KMT2A-rearranged AML and LIMK suppression induces the same phenotypic changes as CDK6 inhibition in this AML subtype, we wondered whether there is a link between CDK6 and LIMK. In line with this hypothesis, we observed increased CDK6 expression upon LIMK1 or LIMK2 suppression in THP-1 (Fig. 4a) and NOMO-1 (Fig. S4A) cells. Interestingly, LIMK2 suppression was also accompanied by upregulation of LIMK1, indicating that LIMK1 can partially compensate the loss of LIMK2. On the other hand, CDK6 knockdown resulted in upregulation of LIMK1, but not LIMK2, and the more efficient shRNA also induced phosphorylation of LIMK’s major substrate CFL1 (Fig. 4b). The latter observation could be confirmed in a series of THP-1-derived CDK6 knockout clones, which showed strongly increased levels of phospho-CFL1 compared to control clones (Fig. 4c), suggesting that CDK6 depletion leads to induction of LIMK activity.
a Western blot of THP-1 cells transduced with LIMK1- or LIMK2-targeting shRNAs or NTC. Lysates were prepared eight days after transduction. b Western blot of THP-1 cells transduced with CDK6-targeting shRNAs or NTC. Lysates were prepared eight days after transduction. The result is representative of two independent experiments. c Western blot of THP-1-derived CDK6 knockout clones generated with two different sgRNAs and control clones (sgmCherry). d Western blot of a PDX sample five days after transduction with NTC or shRNAs targeting LIMK1 or CDK6. e Western blots of AML cells treated with the CDK6 inhibitor palbociclib or DMSO for 72 h (NOMO-1) or 120 h (MOLM-14 and THP-1). f Proliferation determined by manual cell counting (left panel) and colony formation in methylcellulose after ten days (right panel) of THP-1-derived LIMK1 knockout or sgmCherry control clones treated with palbociclib or DMSO. Proliferation data are presented as mean ± SD of three individual clones, and the connecting lines indicate the fitted model for exponential growth. Statistical significance was assessed for the corresponding proliferation constants by two-way ANOVA with repeated measurements (P value for palbociclib effect: <0.0001, P value for LIMK1 knockout effect: <0.001, P value for interaction effect: >0.05). For colony formation, statistical significance was also analyzed by two-way ANOVA with repeated measurements (P value for palbociclib effect: <0.05, P value for LIMK1 knockout effect: <0.001, P value for interaction effect: <0.05). Each LIMK1 knockout clone was generated with a different sgRNA. DMSO data are the same as in Fig. 2c and 2d, respectively. g May-Grünwald-Giemsa-stained cytospin preparations of THP-1 cells eight days after transduction with two shRNAs targeting LIMK1 or NTC and treatment with palbociclib or DMSO. Original magnification, x40. Insets show 1.5-fold magnified details of the corresponding photographs. DMSO data are the same as in Fig. 2e. h Proliferation indices as assessed by CFSE tracking for THP-1 cells treated for four days with DMSO, the CDK6 inhibitor palbociclib, or the LIMK inhibitor LIMKi3 alone, or in combination. Data were normalized to the DMSO sample for each replicate. Statistical significance for each treatment condition was assessed by one-sample t-test on log-transformed data (asterisks below data points). Statistical significance for single vs. combination treatment was assessed by unpaired t-tests on log-transformed data (asterisks above data points). *P < 0.05, **P < 0.01, ***P < 0.001.
A reciprocal regulation between LIMK1 and CDK6 was also evident in a KMT2A-rearranged PDX sample, as shRNA-mediated CDK6 knockdown resulted in increased LIMK1 expression and CFL1 phosphorylation, whereas LIMK1 depletion led to upregulation of CDK6 (Fig. 4d). Similarly, suppression of either Cdk6 or Limk1 in KMT2A-MLLT3 and NRASG12D-transformed murine bone marrow cells resulted in upregulation of the other gene as well as of Limk2, although these observations were dependent on the time point investigated (Fig. S4B). Finally, treatment of several KMT2A-rearranged AML cell lines with the CDK4/6 inhibitor palbociclib induced CFL1 phosphorylation (Fig. 4e), indicating that inhibition of CDK6’s kinase activity is sufficient for this effect. Together, these observations suggest that the reciprocal regulation between CDK6 and LIMK might serve as a compensatory mechanism in response to inhibition of either protein. Indeed, palbociclib treatment further reduced proliferation and colony formation of LIMK1 knockout clones (Fig. 4f). Two-way ANOVA revealed a significant interaction between palbociclib treatment and LIMK1 knockout for colony formation, indicating a synergistic effect; however, no interaction was observed for proliferation, suggesting an additive effect. Furthermore, the differentiated cell morphology induced by LIMK1 depletion was markedly enhanced by addition of palbociclib (Fig. 4g).
Finally, we tested whether the combinatorial effect could be recapitulated by using pharmacologic instead of genetic LIMK inhibition. Proliferation of THP-1 cells was significantly reduced upon treatment with palbociclib or a highly potent LIMK1/LIMK2 inhibitor (LIMKi3) [50] alone and was further decreased by the combination of both inhibitors (Figs. 4h and S4C). Of note, both the reduction in proliferation and the morphologic changes induced by genetic LIMK inhibition were more pronounced compared to LIMKi3 treatment (Figs. 4g and S4D), indicating that the effects of LIMK depletion might be at least partially attributable to a kinase-independent function. In summary, these data suggest that a compensatory feedback loop increases expression and/or activity of LIMK upon CDK6 inhibition and vice versa to preserve survival of AML cells, providing a rationale for a combination therapy approach.
Discussion
Despite an improved understanding of AML pathogenesis, AML treatment still relies on cytotoxic chemotherapy in most cases. Particularly older individuals, who constitute the majority of AML patients, are often not eligible for intensive treatment, so that less toxic, molecularly targeted therapies are highly desirable. Here, we identified LIMK1 and LIMK2 as potential targets for AML therapy.
High LIMK1 expression was associated with several AML driver alterations such as KMT2A rearrangements as well as FLT3 and NPM1 mutations. Of note, NPM1 mutations and KMT2A rearrangements as well as DNMT3A mutations, which showed a trend toward further increased LIMK1 levels in FLT3mut/NPM1mut cases, have been associated with increased expression of the leukemogenic HOX family of transcription factors [51,52,53], and differential gene expression analysis showed significantly higher HOX mRNA levels in LIMK1high compared to LIMK1low AML patient samples. Although this correlation does not necessarily imply a causal link, it is intriguing to speculate that homeobox transcription factors may drive LIMK1 expression. Putative target genes of HOXA9 have been identified by transcriptome profiling following HOXA9 overexpression [48] or knockdown [49] in hematopoietic cells. Although LIMK were not found to be deregulated in those studies, specific genetic backgrounds may be required to observe such interactions. Furthermore, other HOX genes, which have not been studied in as much detail as HOXA9 but were also significantly higher expressed in LIMK1high compared to LIMK1low AML patient samples, may be responsible for driving LIMK1 expression. Another possibility could be that LIMK1 directly or indirectly drives HOX expression. Supporting this hypothesis, we found putative HOXA9 target genes to be downregulated upon LIMK1 depletion. Neither HOXA9 nor any of its established cofactors were downregulated upon LIMK1 suppression in our RNA-seq data, suggesting that LIMK1 might promote HOXA9 function by a post-transcriptional mechanism. Considering that LIMK1 knockdown led to increased expression of proteasomal genes (Fig. S5), we hypothesize that LIMK1 stabilizes HOXA9 and/or its cofactors by preventing their degradation, which might contribute to its leukemogenic function.
Two additional downstream effectors of LIMK1 relevant for its role in AML may be LSP1 and PAQR8, as we observed strong downregulation of both genes upon LIMK1 suppression. Expression of LSP1 and PAQR8 was significantly higher in LIMK1high compared to LIMK1low patient samples, and high expression was correlated with shorter survival of AML patients. Whereas PAQR8, a progesterone receptor, has not been studied in the context of hematologic malignancies, LSP1 has recently been identified as part of a three-gene signature that is predictive of AML outcome [45]. In this study, the authors found a robust correlation of high LSP1 levels and shorter survival not only in the TCGA dataset, but also in two additional AML cohorts, further strengthening the hypothesis that LSP1 may be involved in AML development.
Another potential mechanism by which LIMK1 might promote leukemogenesis is its apparent ability to suppress certain tumor suppressor genes, which were significantly upregulated upon LIMK1 knockdown. For example, EGR1 has been shown to induce differentiation and apoptosis in AML [36, 37], and loss of EGR1 has been associated with accelerated progression of chronic myeloid leukemia and AML with deletion of chromosome 5 or 5q [38]. Similarly, loss of BTG2 enhances transformation and tumor progression [40, 41], and its overexpression prevents proliferation in cells of several tissue backgrounds [39, 54]. Finally, BIN1 expression is frequently lost in carcinomas, particularly in aggressive and metastatic cases, and its overexpression induces apoptosis and inhibits MYC-induced transformation as well as proliferation in various tumor models [42,43,44]. In line with the upregulation of the aforementioned tumor suppressors, we observed slowed cell-cycle dynamics, as assessed by decreased EdU incorporation, upon LIMK1 suppression. LIMK have previously been shown to directly regulate centrosome positioning and mitotic spindle formation by physically interacting with microtubules and regulating Aurora kinase [17,18,19]. Our transcriptomic data suggest that in addition to these protein-protein interactions, LIMK may also regulate expression levels of crucial mitosis-associated genes.
Importantly, depending on the cellular system, we observed upregulation of LIMK1 upon LIMK2 depletion or vice versa, indicating that loss of one family member can be partially compensated by the other. Therefore, we believe that future efforts to develop small-molecule inhibitors should ideally aim to target both LIMK1 and LIMK2 to ensure efficient inhibition of downstream signaling pathways.
Finally, we observed a mutual co-regulation of CDK6 and LIMK, as inhibition of either protein induced the kinase activity and/or expression of the other. Since both CDK6 and LIMK drive cell-cycle progression, this phenomenon may present an escape route for AML cells to prevent more detrimental effects upon inhibition of either protein, although the underlying mechanism remains to be determined. In line with this hypothesis, simultaneous inhibition of CDK6 and LIMK1 resulted in additive or synergistic effects, depending on the cellular characteristics tested. Such a combination approach may not only show superior treatment efficacy in AML, but also in other malignancies for which palbociclib is currently being evaluated in clinical trials or has already been approved, i.e. hormone receptor-positive, HER2-negative breast cancer. Given that several studies have implicated LIMK in progression and metastasis formation of breast cancer [14, 16, 55], combining LIMK and CDK6 inhibition may be particularly interesting in this tumor entity.
In conclusion, this study provides the first comprehensive characterization of LIMK in AML. We found that high LIMK1 expression is associated with specific AML subtypes and poor clinical outcome, and demonstrate that genetic and/or pharmacologic inhibition of LIMK results in decreased proliferation and colony formation of AML cell lines and PDX samples. Furthermore, our in-depth analysis of transcriptomic changes induced by LIMK1 suppression revealed potential novel downstream effectors mediating LIMK1’s leukemogenic function. Finally, LIMK and CDK6 show a reciprocal regulation indicative of an escape mechanism, which could be exploited by a combination therapy approach not only in AML, but potentially also in other tumor entities. Future experiments should focus on the verification of these findings in vivo as well as on the refinement of LIMK inhibitors for evaluation in clinical trials.
Materials and methods
Cell lines and inhibitors
AML cell lines and HEK293T cells were maintained under standard conditions with up to 0.01% DMSO. Cell line identity was verified using the Multiplex Cell Authentication Test (Multiplexion, Friedrichshafen, Germany) or the Human Cell Line Authentication Service (Eurofins Genomics GmbH, Ebertsberg, Germany). All cell lines were routinely tested for mycoplasma contamination. AML PDX cells were isolated from the spleen of leukemic mice as described previously [56] and cultured in StemPro34 SFM (Thermo Fisher Scientific, Waltham, Massachusetts, USA) supplemented with provided nutrient supplement, 2% FBS, 1% penicillin/streptomycin and human FLT3 ligand, human thrombopoietin, human interleukin 3, and human stem cell factor (PeproTech, Princeton, New Jersey, USA; 10 ng/mL each). Murine cell lines were a gift from Stephen M. Sykes and were generated and cultured as described previously [57] (liquid culture) or in methylcellulose medium (MethoCult GF M3434, Stem Cell Technologies, Vancouver, Canada). Palbociclib (PD-0332991) and LIMKi3 (CAS number 1338247-35-0) were obtained from Selleck (Munich, Germany) or Merck Millipore (Burlington, Massachusetts, USA), respectively.
RNAi screening
RNAi screening of HL-60, U937, and OCI-AML3 cells has been described previously [23]. A gene was defined as essential if at least three shRNAs were associated with B-scores of <1 standard deviation below the median (HL-60, < −1.61; U937, < −1.71, OCI-AML3, < −1.58).
Differential gene expression analysis of LIMK1 knockdown samples
Differential gene expression analysis was performed with DESeq2 (ref. 58) (version 1.24.0). Only genes with an average TPM > 0.5 were considered. DESeq function was run with default parameters [58]. Batch correction for replicates, Independent Hypothesis Weighting [59] (IHW 1.12.0), and log-fold change shrinkage with apeglm [60] shrinkage estimator were applied. P value cut-off was set to 0.05. For subsequent GSEAPreranked (GSEA version 4.0.3), all genes that passed the TPM cut-off were ranked by mean log2-fold change (shLIMK1/NTC), and GSEAPreranked was performed using default parameters.
Differential gene expression analysis of TCGA data
AML patients with available RNA-seq and survival data (n = 132) were sorted according to LIMK1 expression. Log-transformed read count data were converted to raw read count values before analysis. The top 10% of highest and lowest LIMK1-expressing patients (based on FPKM values; n = 26) were used for differential gene expression analysis with DESeq2 (ref. 58) (version 1.24.0). Only genes with an average TPM > 0.41 were considered. DESeq function was run with default parameters [58]. Independent Hypothesis Weighting [59] (IHW 1.12.0) and log-fold change shrinkage with apeglm [60] shrinkage estimator were applied. P value cut-off was set to 0.01, and log-fold change cut-off was set to 1. For heatmap generation, FPKM values were converted to TPM values. Only protein-coding genes were considered for subsequent GSEA (version 4.0.3), which was performed using default parameters.
Survival analysis of TCGA data
AML patients with available RNA-seq and survival data (n = 132) were sorted according to candidate gene expression (based on FPKM values). The cohort was split into high and low expressers based on the median. Statistical analysis was performed in GraphPad Prism 6 (GraphPad Software) by log-rank test. Survival and expression data were downloaded from https://xenabrowser.net.
Code availability
R code used for differential gene expression analysis of LIMK1 knockdown experiments and TCGA data can be found in the supplemental information.
Statistics
Statistical analysis was performed using one-sample t-test, paired or unpaired two‐tailed t‐test with F test to ensure equal variances, one‐way ANOVA with Dunnett’s or Tukey’s post-hoc tests and Brown–Forsythe-test to ensure equal variances, Kruskal-Wallis test with Dunn’s post-hoc test, or two-way ANOVA with repeated measurements as appropriate. Details about multivariable regression analysis of TCGA data can be found in the supplemental information. P values < 0.05 were considered significant. Computations were performed using GraphPad Prism versions 6 or 8 (GraphPad Software) or R version 3.5. All experiments were performed at least three times unless indicated otherwise, and error bars indicate the standard deviation of the mean.
References
SEER*Explorer: An interactive website for SEER cancer statistics. Surveillance Research Program, National Cancer Institute. https://seer.cancer.gov/explorer/.
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND, et al. Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med. 2016;374:2209–21.
Döhner H, Weisdorf DJ, Bloomfield CD. Acute myeloid leukemia. N Engl J Med. 2015;373:1136–52.
Schlenk RF, Frech P, Weber D, Brossart P, Horst HA, Kraemer D, et al. Impact of pretreatment characteristics and salvage strategy on outcome in patients with relapsed acute myeloid leukemia. Leukemia. 2017;31:1217–20.
Webster JA, Pratz KW. Acute myeloid leukemia in the elderly: therapeutic options and choice. Leuk Lymphoma. 2018;59:274–87.
Stone RM, Mandrekar SJ, Sanford BL, Laumann K, Geyer S, Bloomfield CD, et al. Midostaurin plus chemotherapy for acute myeloid leukemia with a FLT3 mutation. N Engl J Med. 2017;377:454–64.
DiNardo CD, Stein EM, de Botton S, Roboz GJ, Altman JK, Mims AS, et al. Durable remissions with ivosidenib in IDH1-mutated relapsed or refractory AML. N Engl J Med. 2018;378:2386–98.
Stein EM, DiNardo CD, Pollyea DA, Fathi AT, Roboz GJ, Altman JK, et al. Enasidenib in mutant IDH2 relapsed or refractory acute myeloid leukemia. Blood. 2017;130:722–31.
The Cancer Genome Atlas Research Network, Ley TJ, Miller C, Ding L, Raphael BJ, Mungall AJ, et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–74.
Arber S, Barbayannis FA, Hanser H, Schneider C, Stanyon CA, Bernard O, et al. Regulation of actin dynamics through phosphorylation of cofilin by LIM-kinase. Nature. 1998;393:805–9.
Yang N, Higuchi O, Ohashi K, Nagata K, Wada A, Kangawa K, et al. Cofilin phosphorylation by LIM-kinase 1 and its role in Rac-mediated actin reorganization. Nature. 1998;393:809–12.
Yoshioka K, Foletta V, Bernard O, Itoh K. A role for LIM kinase in cancer invasion. Proc Natl Acad Sci USA. 2003;100:7247–52.
Su J, Zhou Y, Pan Z, Shi L, Yang J, Liao A, et al. Downregulation of LIMK1-ADF/cofilin by DADS inhibits the migration and invasion of colon cancer. Sci Rep. 2017;7:45624.
Bagheri-Yarmand R, Mazumdar A, Sahin AA, Kumar R. LIM kinase 1 increases tumor metastasis of human breast cancer cells via regulation of the urokinase-type plasminogen activator system. Int J Cancer. 2006;118:2703–10.
You T, Gao W, Wei J, Jin X, Zhao Z, Wang C, et al. Overexpression of LIMK1 promotes tumor growth and metastasis in gastric cancer. Biomed Pharmacother. 2015;69:96–101.
McConnell BV, Koto K, Gutierrez-Hartmann A. Nuclear and cytoplasmic LIMK1 enhances human breast cancer progression. Mol Cancer. 2011;10:75.
Kaji N, Muramoto A, Mizuno K. LIM kinase-mediated cofilin phosphorylation during mitosis is required for precise spindle positioning. J Biol Chem. 2008;283:4983–92.
Ou S, Tan MH, Weng T, Li H, Koh CG. LIM kinase1 regulates mitotic centrosome integrity via its activity on dynein light intermediate chains. Open Biol. 2018;8:170202.
Petrilli A, Copik A, Posadas M, Chang LS, Welling DB, Giovannini M, et al. LIM domain kinases as potential therapeutic targets for neurofibromatosis type 2. Oncogene. 2014;33:3571–82.
Manetti F. Recent advances in the rational design and development of LIM kinase inhibitors are not enough to enter clinical trials. Eur J Med Chem. 2018;155:445–58.
Oku Y, Tareyanagi C, Takaya S, Osaka S, Ujiie H, Yoshida K, et al. Multimodal effects of small molecule ROCK and LIMK inhibitors on mitosis, and their implication as anti-leukemia agents. PLoS One. 2014;9:e92402.
Zampini M, Tregnago C, Bisio V, Simula L, Borella G, Manara E, et al. Epigenetic heterogeneity affects the risk of relapse in children with t(8;21)RUNX1-RUNX1T1-rearranged AML. Leukemia. 2018;32:1124–34.
Placke T, Faber K, Nonami A, Putwain SL, Salih HR, Heidel FH, et al. Requirement for CDK6 in MLL-rearranged acute myeloid leukemia. Blood. 2014;124:13–23.
Hart T, Chandrashekhar M, Aregger M, Steinhart Z, Brown KR, MacLeod G, et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell. 2015;163:1515–26.
Wang T, Birsoy K, Hughes NW, Krupczak KM, Post Y, Wei JJ, et al. Identification and characterization of essential genes in the human genome. Science. 2015;350:1096–1101.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Cancer Cell Line Encyclopedia C, Genomics of Drug Sensitivity in Cancer C. Pharmacogenomic agreement between two cancer cell line data sets. Nature. 2015;528:84–7.
Haferlach T, Kohlmann A, Wieczorek L, Basso G, Kronnie GT, Bene MC, et al. Clinical utility of microarray-based gene expression profiling in the diagnosis and subclassification of leukemia: report from the International Microarray Innovations in Leukemia Study Group. J Clin Oncol. 2010;28:2529–37.
Kohlmann A, Kipps TJ, Rassenti LZ, Downing JR, Shurtleff SA, Mills KI, et al. An international standardization programme towards the application of gene expression profiling in routine leukaemia diagnostics: the Microarray Innovations in LEukemia study prephase. Br J Haematol. 2008;142:802–7.
Bagger FO, Sasivarevic D, Sohi SH, Laursen LG, Pundhir S, Sonderby CK, et al. BloodSpot: a database of gene expression profiles and transcriptional programs for healthy and malignant haematopoiesis. Nucleic Acids Res. 2016;44:D917–24.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Ahmad MK, Abdollah NA, Shafie NH, Yusof NM, Razak SRA. Dual-specificity phosphatase 6 (DUSP6): a review of its molecular characteristics and clinical relevance in cancer. Cancer Biol Med. 2018;15:14–28.
Schmid CA, Robinson MD, Scheifinger NA, Muller S, Cogliatti S, Tzankov A, et al. DUSP4 deficiency caused by promoter hypermethylation drives JNK signaling and tumor cell survival in diffuse large B cell lymphoma. J Exp Med. 2015;212:775–92.
Ben-Batalla I, Schultze A, Wroblewski M, Erdmann R, Heuser M, Waizenegger JS, et al. Axl, a prognostic and therapeutic target in acute myeloid leukemia mediates paracrine crosstalk of leukemia cells with bone marrow stroma. Blood. 2013;122:2443–52.
Raffel S, Falcone M, Kneisel N, Hansson J, Wang W, Lutz C, et al. BCAT1 restricts alphaKG levels in AML stem cells leading to IDHmut-like DNA hypermethylation. Nature. 2017;551:384–88.
Fu L, Huang W, Jing Y, Jiang M, Zhao Y, Shi J, et al. AML1-ETO triggers epigenetic activation of early growth response gene l, inducing apoptosis in t(8;21) acute myeloid leukemia. FEBS J. 2014;281:1123–31.
Gibbs JD, Liebermann DA, Hoffman B. Egr-1 abrogates the E2F-1 block in terminal myeloid differentiation and suppresses leukemia. Oncogene. 2008;27:98–106.
Joslin JM, Fernald AA, Tennant TR, Davis EM, Kogan SC, Anastasi J, et al. Haploinsufficiency of EGR1, a candidate gene in the del(5q), leads to the development of myeloid disorders. Blood. 2007;110:719–26.
Jiang H, Zhu Y, Zhou Z, Xu J, Jin S, Xu K, et al. PRMT5 promotes cell proliferation by inhibiting BTG2 expression via the ERK signaling pathway in hepatocellular carcinoma. Cancer Med. 2018;7:869–82.
Coppola V, Musumeci M, Patrizii M, Cannistraci A, Addario A, Maugeri-Sacca M, et al. BTG2 loss and miR-21 upregulation contribute to prostate cell transformation by inducing luminal markers expression and epithelial-mesenchymal transition. Oncogene. 2013;32:1843–53.
Takahashi F, Chiba N, Tajima K, Hayashida T, Shimada T, Takahashi M, et al. Breast tumor progression induced by loss of BTG2 expression is inhibited by targeted therapy with the ErbB/HER inhibitor lapatinib. Oncogene. 2011;30:3084–95.
Tajiri T, Liu X, Thompson PM, Tanaka S, Suita S, Zhao H, et al. Expression of a MYCN-interacting isoform of the tumor suppressor BIN1 is reduced in neuroblastomas with unfavorable biological features. Clin Cancer Res. 2003;9:3345–55.
Elliott K, Ge K, Du W, Prendergast GC. The c-Myc-interacting adaptor protein Bin1 activates a caspase-independent cell death program. Oncogene. 2000;19:4669–84.
Sakamuro D, Elliott KJ, Wechsler-Reya R, Prendergast GC. BIN1 is a novel MYC-interacting protein with features of a tumour suppressor. Nat Genet. 1996;14:69–77.
Wagner S, Vadakekolathu J, Tasian SK, Altmann H, Bornhauser M, Pockley AG, et al. A parsimonious 3-gene signature predicts clinical outcomes in an acute myeloid leukemia multicohort study. Blood Adv. 2019;3:1330–46.
Charles NJ, Thomas P, Lange CA. Expression of membrane progesterone receptors (mPR/PAQR) in ovarian cancer cells: implications for progesterone-induced signaling events. Horm Cancer. 2010;1:167–76.
Sinreih M, Knific T, Thomas P, Frkovic Grazio S, Rizner TL. Membrane progesterone receptors beta and gamma have potential as prognostic biomarkers of endometrial cancer. J Steroid Biochem Mol Biol. 2018;178:303–11.
Hess JL, Bittner CB, Zeisig DT, Bach C, Fuchs U, Borkhardt A, et al. c-Myb is an essential downstream target for homeobox-mediated transformation of hematopoietic cells. Blood. 2006;108:297–304.
Faber J, Krivtsov AV, Stubbs MC, Wright R, Davis TN, van den Heuvel-Eibrink M, et al. HOXA9 is required for survival in human MLL-rearranged acute leukemias. Blood. 2009;113:2375–85.
Ross-Macdonald P, de Silva H, Guo Q, Xiao H, Hung CY, Penhallow B, et al. Identification of a nonkinase target mediating cytotoxicity of novel kinase inhibitors. Mol Cancer Ther. 2008;7:3490–98.
Armstrong SA, Staunton JE, Silverman LB, Pieters R, de Boer ML, Minden MD, et al. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat Genet. 2002;30:41–7.
Yan XJ, Xu J, Gu ZH, Pan CM, Lu G, Shen Y, et al. Exome sequencing identifies somatic mutations of DNA methyltransferase gene DNMT3A in acute monocytic leukemia. Nat Genet. 2011;43:309–15.
Mullighan CG, Kennedy A, Zhou X, Radtke I, Phillips LA, Shurtleff SA, et al. Pediatric acute myeloid leukemia with NPM1 mutations is characterized by a gene expression profile with dysregulated HOX gene expression distinct from MLL-rearranged leukemias. Leukemia. 2007;21:2000–9.
Dolezal E, Infantino S, Drepper F, Borsig T, Singh A, Wossning T, et al. The BTG2-PRMT1 module limits pre-B cell expansion by regulating the CDK4-Cyclin-D3 complex. Nat Immunol. 2017;18:911–20.
Prunier C, Josserand V, Vollaire J, Beerling E, Petropoulos C, Destaing O, et al. LIM Kinase Inhibitor Pyr1 Reduces the Growth and Metastatic Load of Breast Cancers. Cancer Res. 2016;76:3541–52.
Vick B, Rothenberg M, Sandhofer N, Carlet M, Finkenzeller C, Krupka C, et al. An advanced preclinical mouse model for acute myeloid leukemia using patients’ cells of various genetic subgroups and in vivo bioluminescence imaging. PLoS One. 2015;10:e0120925.
Rudat S, Pfaus A, Cheng YY, Holtmann J, Ellegast JM, Buhler C, et al. RET-mediated autophagy suppression as targetable co-dependence in acute myeloid leukemia. Leukemia. 2018;32:2189–202.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Ignatiadis N, Klaus B, Zaugg JB, Huber W. Data-driven hypothesis weighting increases detection power in genome-scale multiple testing. Nat Methods. 2016;13:577–80.
Zhu A, Ibrahim JG, Love MI. Heavy-tailed prior distributions for sequence count data: removing the noise and preserving large differences. Bioinformatics. 2019;35:2084–92.
Acknowledgements
The authors thank the DKFZ Genome and Proteome, Flow Cytometry, Light Microscopy, Omics IT and Data Management, and Tumor Models Core Facilities for excellent technical assistance; Cihan Erkut and Annette Kopp-Schneider for support with bioinformatic and statistical analysis; Stephen M. Sykes for providing transformed murine bone marrow cells; and Gina Walter-Bausch, Saskia Rudat, and Marie Groth for helpful discussions. Patrizia Jensen was supported by a grant from the German José Carreras Leukemia Foundation (DJCLS F 15/04). Ya-Yun Cheng was supported by a stipend from the Helmholtz International Graduate School for Cancer Research. Simon Weisemann was supported by a stipend from the German Academic Scholarship Foundation.
Author information
Authors and Affiliations
Contributions
PJ, SF, and CS designed the study and wrote the manuscript; PJ, MC, RFS, JK, AW, IB, GG, and SW performed experiments and/or analyzed data; MC and IJ provided human AML PDX samples, and MS and YYC gave conceptual input.
Corresponding authors
Ethics declarations
Conflict of interest
SF has had a consulting or advisory role, received honoraria, research funding, and/or travel/accommodation expenses funding from the following for-profit companies: Amgen, AstraZeneca, Bayer, Eli Lilly, Pfizer, PharmaMar, and Roche. RFS has had a consulting or advisory role, received honoraria, research funding, and/or travel/accommodation expenses funding from the following for-profit companies: Roche, Daiichi Sankyo, Pfizer, Novartis, PharmaMar, and AstraZeneca. The other authors declare no competing financial interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Jensen, P., Carlet, M., Schlenk, R.F. et al. Requirement for LIM kinases in acute myeloid leukemia. Leukemia 34, 3173–3185 (2020). https://doi.org/10.1038/s41375-020-0943-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-020-0943-5
- Springer Nature Limited
This article is cited by
-
Heterozygous variants in GATA2 contribute to DCML deficiency in mice by disrupting tandem protein binding
Communications Biology (2022)
-
Involvement of LIMK2 in actin cytoskeleton remodeling during the definitive endoderm differentiation
In Vitro Cellular & Developmental Biology - Animal (2021)