Abstract
A large body of evidence suggests that B-cell lymphomas with enhanced Myc expression are associated with an aggressive phenotype and poor prognosis, which makes Myc a compelling therapeutic target. Phosphodiesterase 4B (PDE4B), a main hydrolyzer of cyclic AMP (cAMP) in B cells, was shown to be involved in cell survival and drug resistance in diffuse large B cell lymphomas (DLBCL). However, the interrelationship between Myc and PDE4B remains unclear. Here, we first demonstrate the presence of the Myc-PDE4B feed-forward loop, in which Myc and PDE4B mutually reinforce the expression of each other. Next, the combined targeting of Myc and PDE4 synergistically prevented the proliferation and survival of B lymphoma cells in vitro and in a mouse xenograft model. We finally recapitulated this combinatorial effect in Eμ-myc transgenic mice; co-inhibition of Myc and PDE4 suppressed lymphomagenesis and restored B cell development to the wild type level that was associated with marked reduction in Myc levels, unveiling the critical role of the Myc-PDE4B amplification loop in the regulation of Myc expression and the pathogenesis of B cell lymphoma. These findings suggest that the disruption of the Myc-PDE4B circuitry can be exploited in the treatment of B cell malignancies.
Highlights
- 1.
Myc and PDE4B activate expression of each other, forming a positive feedback loop.
- 2.
Co-inhibition of Myc and PDE4 induces synergistic killing of B lymphoma cells.
Similar content being viewed by others
Introduction
Myc is a basic helix-loop-helix leucine zipper transcription factor that heterodimerizes with Max and binds the E-box sequence. The activation of Myc is thought to regulate an estimated 10–15% of the global transcriptome and has been associated with many aspects of biological functions in normal cells, encompassing cell cycle progression, cell growth, survival, apoptosis, angiogenesis and stem cell self-renewal [1,2,3,4]. Indeed, defects in the regulation of this gene are frequently found in a broad range of human cancers, including B-cell lymphomas [4], making Myc an attractive therapeutic target. The inhibitors of bromodomain and extra-terminal (BET) bromodomain-containing proteins have emerged from recent insights into the regulatory mechanism of Myc expression [5]. For example, JQ1, a prototype drug, and the I-BET series, resulted in a remarkable decrease in Myc protein abundance and associated tumor regression in murine lymphoma models, suggesting that Myc targeting with BET inhibitors may prove feasible in B-cell neoplasia [2, 6]. However, their efficacy in patients remains unknown.
Cyclic adenosine 3′,5′-monophosphate (cAMP) plays crucial roles in transmitting signals from many hormones and neurotransmitters. The regulation of intracellular cAMP concentrations is accomplished by the opposing actions of adenylyl cyclase (AC) and phosphodiesterase (PDE) that promote the generation and degradation of cAMP, respectively. The PDE4 family (PDE4A-PDE4D) is cAMP-specific, and observations that increment in cAMP levels induces cell death in hematologic neoplasia provided a strong rationale for exploring PDE4 inhibitors as antitumor agents in these types of cancer. Indeed, they have been successfully tested in CLL and acute lymphoblastic leukemia (ALL) cells [7, 8].
An association between PDE4B and B-cell lymphoma was first suggested by genome-wide profiling studies, wherein it was highly expressed in chemotherapy-resistant patients with DLBCL [9]. Subsequent studies have shown that cAMP downmodulates the PI3K/AKT pathway to promote cell death and that this effect was abolished by the hydrolysis of cAMP by PDE4B [10]. Patients with advanced B-cell malignancies showed favorable responses to a combination of roflumilast and prednisone in a recent phase Ib clinical trial, warranting further studies to improve combination strategies that include PDE4 inhibitors [11, 12].
Herein, we report that Myc and PDE4B mutually reinforce the expression of each other, forming a positive feedback circuitry, and that PDE4 inhibitors in combination with BET inhibitors can be effectively used as a therapy for Myc-induced B-cell lymphomas.
Materials and methods
See Supplementary Materials and Methods for detailed methodology.
Xenograft studies and Eμ-myc transgenic mice
Athymic nude mice (BALB/c-nu) were injected subcutaneously with 1.0 × 107 Ramos cells, followed by treatment with indicated drugs. Drugs were injected intraperitoneally daily and tumors were measured using a digital caliper [13]. Tumor tissues were immunostained with antibodies against Myc or Ki-67. Eμ-myc transgenic mice were obtained from the Jackson Laboratory (Bar Harbor, ME, USA). The animal protocol used in this study was reviewed and approved by the Pusan National University-Institutional Animal Care and Use Committee.
H&E and immunohistochemical (IHC) staining
H&E and IHC staining were performed as previously described [14]. For IHC staining of Myc in tissues, a polyclonal anti-Myc antibody and a polymer-horseradish peroxidase anti-rabbit antibody were used as a primary and a secondary antibody, respectively, followed by 3,3-diaminobenzidine treatment to visualize the proteins.
Isolation of murine B cells
After the lysis of red blood cells using ammonium chloride-based red cell lysis buffer, B cell isolation was performed using EasySep kit (STEMCELL Technologies) according to the manufacturer’s instruction. Myc and PDE4B protein levels were analyzed by western blotting.
Assessment of B cell development by flow cytometry
To assess B cell developmental stages, bone marrow and splenic cells were collected from Eμ-myc transgenic mice. Cells were stained with PE-Cy7 anti-mouse IgM, PE anti-mouse 45R/B220, and BB515 anti-mouse CD43 (BD biosciences) antibodies, followed by FACS analysis. The developmental stages were defined by the following key cell-surface markers (Fig. S1): pro-B cells (B220+CD43+IgM−) and pre-B cells (B220+CD43lowIgM−) [15].
Chromatin Immunoprecipitation (ChIP) and qRT-PCR
ChIP assays were performed as previously described [16]. DNA obtained from the ChIP assay was analyzed by PCR using the primers shown in Supplementary Table 1. The expression of Myc target genes and PDE4B splice variants in B lymphoma cells, following treatment with indicated drugs, was analyzed by qRT-PCR using the primers listed in Supplementary Tables 2, 3.
Cloning of the PDE4B2 and B3 promoter region, site-directed mutagenesis, and reporter assays
The promoter regions of the PDE4B splice variants PDE4B2 and B3, containing potential Myc-binding sites were PCR-amplified and cloned into the pGL3 basic vector (Promega). To mutate the Myc E-box in the PDE4B promoter region, site-directed mutagenesis was performed as described [17]. Primers for cloning and site-directed mutagenesis are listed in Supplementary Tables 4, 5. Reporter assays using the Dual Luciferase Reporter Assay System (Promega) was described previously [18].
Genetic modulation of PDE4B in DLBCL cell lines
For ectopic expression of wild type (WT) or phosphodiesterase-inactive (PI) mutant PDE4B2 in PDE4B-low DHL4 cells, PDE4B-WT and PDE4B-PI gene were cloned into the pCDH-CMV-MCS-EF1-copGFP lentiviral vector (Systems Biosciences) [10, 19]. For PDE4B knockdown, PDE4B-targeting shRNA constructs (Dharmacon) were delivered into PDE4B-high Ly1 cells, followed by sorting the GFP-positive cells using FACS.
Combination index values and statistical analysis
Combination index (CI) values were calculated using the CompuSyn software. To identify statistically significant differences (p < 0.05), a non-parametric Mann–Whitney U-test, Fisher’s exact test, or one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test was performed as indicated in the figure legends.
Analysis of the immunoglobulin heavy chain (IgH) rearrangements
Analysis of IgH rearrangements as a measure of clonality of tumor tissues was performed as described previously [20].
Results
cAMP/PDE4B modulates Myc expression
Previously, it was shown that Myc expression is regulated by cAMP in a cell-type specific manner [21,22,23]. To investigate whether cAMP/PDE4B regulates Myc levels in malignant B cells, we dichotomized six well-characterized B lymphoma cell lines according to PDE4B expression [24,25,26] (Fig. S2). Induction of intracellular cAMP by exposure to frs, an adenylyl cyclase activator, diminished Myc expression in PDE4B-low (DHL4 and DHL6), but not PDE4B-high (Ly1, Ly10, and Ramos), cells (Fig. 1a). In support of the critical role of PDE4B in Myc regulation in PDE4B-high cells, Myc levels were downregulated by frs when the PDE4B-high cells were co-treated with rolipram or roflumilast, pan-PDE4 inhibitors (Fig. 1b, S2, and S3). This implies that PDE4B upregulates Myc expression by degrading cAMP and thereby limiting its negative effect on Myc. Decreased Myc levels are tightly correlated with a reduction in Myc-dependent transcriptional activation, which is manifested by diminished expression of Myc target genes in B cells [27] (Fig. 1c). Treatment with 8-Br-cAMP the cell-permeable cAMP analog reproduced the data obtained with frs and/or a PDE4 inhibitor treatment, confirming that the modulation of Myc expression by these drugs is derived from elevation of cAMP levels (Fig. 1d). To genetically confirm the role of PDE4B in Myc regulation, we stably expressed either wild type (WT) or phosphodiesterase-inactive (PI) mutant PDE4B in PDE4B-low DHL4 cells [10] (Fig. 1e). Forskolin did not influence Myc expression in PDE4B-WT cells; however, it efficiently suppressed Myc in PDE4B-PI cells, the response being similar to that of the DHL4 parental cells. Conversely, introduction of shRNAs targeting two different sequences of PDE4B (sh-PDE4B#2 and #3) in PDE4B-high Ly1 cells decreased Myc levels upon frs treatment (Fig. 1f), corroborating the data regarding the pharmacological inhibition of PDE4B in PDE4B-high cells. Together, these results suggest that in B lymphoma cells cAMP inhibits Myc expression and PDE4B negates the negative impact of cAMP on Myc.
cAMP/PDE4B regulates Myc levels. a Six B lymphoma cell lines were dichotomized into PDE4B-low (DHL4 and DHL6) and PDE4B-high (Ly1, Ly10, and Ramos) cells and treated with frs (0, 20, or 40 μM) for 16 h. Cell lysates were prepared and Myc protein levels were analyzed by western blotting. β-actin was used as a loading control. b Ly1, Ly10, and Ramos cells were exposed to DMSO, frs, rolipram or frs plus rolipram for 16 h, as indicated. Myc protein levels were analyzed by western blotting. c DHL4 and Ly1 cells were treated with frs (40 μM, 16 h) and frs (40 μM, 16 h)/rolipram (40 μM, 16 h), respectively, and expression of indicated Myc target genes was analyzed by qRT-PCR. The data are presented as mean ± SD and representative of three independent experiments (*p < 0.05, Mann–Whitney U-test). d DHL4, DHL6, Ly1, and Ly10 cells were exposed to a cell-permeable cAMP analog 8-Br-cAMP (0–500 μM, 48 h) and protein levels of Myc were examined by western blotting. β-actin confirmed equal loading. As is the case with frs and/or PDE4 inhibitors, 8-Br-cAMP diminished Myc expression in multiple B lymphoma cells, suggesting that the effect of frs and/or PDE4 inhibitors on Myc expression stems from elevation of intracellular cAMP concentration. e PDE4B-low DHL4 cells stably expressing either PDE4B-WT or PDE4B-–PI were generated using lentiviral constructs and their ectopic expression was confirmed via western blotting (Upper panel). Forskolin-mediated elevation in cAMP levels reduced Myc protein levels in PDE4B-PI cells; however, Myc expression remained unchanged in PDE4B-WT cells (Lower panel). f PDE4B-targeting lentiviral shRNA constructs (sh-PDE4B#2 and #3) were stably transduced in PDE4B-high Ly1 cells, and PDE4B knockdown was confirmed via western blotting (Left panels). Forskolin treatment alone efficiently downregulated Myc in sh-PDE4B cells; however, it did not affect sh-Control cells (Right panels)
Next, we attempted to identify the signaling pathway responsible for mediating the cAMP/PDE4B regulation of Myc. We considered the SYK/AKT/mTOR axis as a plausible candidate [10, 24, 25, 28]. As previously demonstrated, the elevation of cAMP by frs and/or rolipram efficiently inhibited the phosphorylation of SYK, AKT, and 4EBP1, a downstream effector of mTOR; moreover, this was found to be associated with Myc suppression (Fig. 2a). To isolate the roles of SYK, AKT, and mTOR in the regulation of Myc, we examined Myc levels in the presence of inhibitors of these molecules, bay 61–3606, wortmannin and rapamycin, respectively. These inhibitors decreased the phosphorylation of SYK, AKT and 4EBP1 in a dose-dependent manner that was highly correlated with the repression of Myc (Fig. 2b–d, and S4). Although we cannot exclude the possibility that other signaling pathways might play a role, these results clearly show that SYK, AKT and mTOR connect cAMP/PDE4B to Myc. mTOR as a catalytic subunit of the mTOR complex 1 (mTROC1) serves as a central growth regulator by potentiating anabolic processes such as protein synthesis [29, 30]. Previous studies reported that rapamycin and mTORC1 regulate internal ribosome entry site (IRES)- and cap-dependent translation of Myc; however, Myc protein stability or mRNA levels was not affected [31,32,33]. These studies collectively suggest that SYK/AKT/mTOR signals downstream of cAMP/PDE4B may regulate Myc expression primarily at the translational level. Therefore, we decided to investigate how Myc transcription is controlled by cAMP/PDE4B. A host of transcription factors that directly bind to the Myc promoter have been identified and we hypothesized that Myc status would have a significant impact on the transcriptional regulation of the gene. Ly10 and Ramos cell lines were chosen for this experiment because the Myc status in these cells are different and well characterized (Table S6); Ly10 has unperturbed Myc loci with an intact Myc promoter region, whereas Ramos harbors a t(8;14) chromosomal translocation, disrupting the promoter region of the gene. Consistent with our hypothesis, frs/rolipram or frs/roflumilast repressed Myc mRNA levels in Ly10 cells but did not influence in Ramos cells, demonstrating that the transcriptional regulation of Myc by cAMP/PDE4B requires normal Myc promoter (Fig. 2e and S5). To examine the potential role of the SYK/AKT/mTOR pathway in Myc transcription, Ly10 and Ramos cells were treated with bay 61–3606, wortmannin or rapamycin, followed by the analysis of Myc mRNA levels via qRT-PCR. As reported previously [31], rapamycin did not influence Myc mRNA levels in either cell lines, whereas bay 61–3606 and wortmannin significantly reduced Myc transcription in Ly10, but not Ramos, suggesting that transcriptional control of Myc by SYK and AKT depends upon unperturbed Myc promoter region (Fig. 2f). Transcription factors that are activated by AKT and modulate Myc transcription include STAT3, β-catenin, E2F, and NF-κB among others. We suspect that regulation of SYK/AKT signals by cAMP/PDE4B eventually leads to the transcriptional control of Myc via activation/inactivation of some of these transcription factors. Together, these data collectively imply that cAMP/PDE4B controls Myc expression at the transcriptional and translational levels.
Myc is regulated at the transcriptional and translational levels. a DHL4, DHL6, and Ly1 cells were exposed to frs (40 μM) or frs (40 μM)/rolipram (40 μM) for 16 h and levels of pSYK, pAKT, p4EBP1, and Myc were detected by western blotting. Equal loading was confirmed by β-actin. b–d Myc, pSYK, pAKT and p4EBP1 levels were analyzed in DHL4, DHL6, and Ly1 cells following treatment with a SYK inhibitor bay 61–3606 (0–10 μM, 24 h), a PI3K inhibitor wortmannin (0–1000 nM, 24 h) or an mTOR inhibitor rapamycin (0–100 nM, 24 h). β-actin served as a loading control. Myc expression was downregulated upon suppression of SYK, AKT, and mTOR activities. Representative data from at least three independent experiments. e DMSO, forskolin (40 μM, 16 h), rolipram (40 μM, 16 h), or frs/rolipram was added in Ly10 and Ramos, followed by the measurement of Myc mRNA levels via qRT-PCR. The data are presented as mean ± SD and representative of three independent experiments (*p < 0.05, One-way ANOVA). f Ly10 and Ramos were exposed to bay 61–3606 (0–10 μM, 16 h), wortmannin (0–1000 nM, 16 h), or rapamycin (0–100 nM, 16 h), followed by the analysis of Myc transcription via qRT-PCR. The data are presented as mean ± SD and representative of three independent experiments (*p < 0.05, One-way ANOVA)
Considering the critical role of BCR/SYK signaling and Myc in proliferation of B cells [34], we suspected that Myc suppression might be due to indirect effects of cAMP on cell cycle progression, which was disproved by our results that addition of frs/rolipram or bay 61–3606 elicited no differences in the cell cycle profile (Fig. S6).
Myc directly regulates PDE4B expression
It was previously demonstrated that cAMP, LPS, and miR-124 are included among the regulators of PDE4B expression [19, 35,36,37]. However, the transcriptional regulation of this gene remains largely unknown. We found that major isoforms of PDE4B found in B lymphoma cells (PDE4B2 and B3) have noncanonical E-boxes in their promoter regions [38, 39] (Fig. 3a). ChIP was performed to explore whether the Myc protein could bind to E-box sequences. Antibodies against Myc efficiently immunoprecipitated the promoter regions of PDE4B isoforms, but not negative control DNA region, suggesting that Myc directly binds to the E-box in the PDE4B promoter (Fig. 3b, c). To validate ChIP results, reporter assays were carried out using luciferase constructs containing the PDE4B promoter region. It was found that the ectopic expression of Myc increased the luciferase activity of the PDE4B promoter constructs containing E-box in the wild type (pGL3-PDE4B-wt) configuration whereas an empty vector control (pGL3-basic) or the PDE4B promoter constructs with mutant E-box (pGL3-PDE4B-mut) did not respond to Myc transfection (Fig. 3d). This implies that the Myc E-box in the PDE4B promoter region was functional, and that Myc upregulates PDE4B levels by binding to the E-box. These results are in line with the observation that the pharmacological suppression of Myc by JQ1 or 10058-F4 (a Myc-Max dimerization inhibitor) diminished PDE4B expression in multiple B-lymphoma cell lines, whereas Myc overexpression augmented PDE4B levels in Ly1 cells and Eμ-myc mice, unveiling that PDE4B is a direct target of Myc (Fig. 3e–g, S7–9). Together with our data from the previous section, these results point to the presence of a Myc-PDE4B positive feedback loop, whereby Myc directly influences PDE4B expression, and eventually upregulates its own expression by attenuating inhibitory effect of cAMP on the SYK, AKT, and mTOR. This idea is supported by our finding that Myc depletion by JQ1, bay 61–3606, wortmannin and rapamycin resulted in suppression of PDE4B levels, which was accompanied by the elevation of intracellular cAMP concentrations and attenuation of the SYK, AKT, and mTOR (Fig. 3h and S10). Additionally, analysis of public datasets revealed that Myc expression was positively correlated with the expression of PDE4B in DLBCL specimen, further supporting the notion that Myc and PDE4B enhance the expression of each other (Fig. 3i).
The PDE4B gene is a direct target of Myc. a Schematic diagrams of luciferase reporter constructs showing the relative locations of potential Myc E-boxes (CATGTG, CACATG) in the promoters of PDE4B2 and B3 that were identified using the TRANSFAC database [18]. Arrow heads indicate the primers used in the ChIP assays. b ChIP assays to monitor occupancy of Myc on the PDE4B promoters in Ly1, Ly10, and Ramos cells. ChIP-enriched DNA was PCR-amplified using primers listed in supplementary Table 1 and run on an agarose gel. The IgG antibody was used as a negative control. c ChIP assays to monitor occupancy of Myc on PDE4B promoters or negative control DNA regions that are located 100–200 bp up- or down-stream of the Myc E-boxes. ChIP-enriched DNA was quantified by qRT-PCR. The data are presented as mean ± SD and representative of three independent experiments (*p < 0.05, One-way ANOVA). d Reporter assays were performed using luciferase constructs with Myc E-boxes in wild type (pGL3-PDE4B2 or B3-wt) or mutant (pGL3-PDE4B2 or B3-mut) configurations, or without the PDE4B promoter sequences (pGL3-basic). The luciferase activity of the reporter constructs containing PDE4B promoters increased by 2.5 to 3.5-fold when Myc was cotransfected, while that of the pGL3-PDE4B-mut or the pGL3-basic did not (Top panel). Ectopic Myc expression was confirmed by western blotting upon transfection of pcDNA-Myc as compared with empty vector (Bottom panel). The data are presented as mean ± SD and representative of three independent experiments (*p < 0.05, Mann–Whitney U-test). e, f PDE4B-high cell lines Ly1 and Ly10 were treated with increasing doses of the BET bromodomain inhibitor JQ1 (0–1000 nM) for 16 h. Myc and PDE4B mRNA and protein levels were measured by qRT-PCR and western blotting, respectively. β-actin served as an internal control. An arrow in the western blot for PDE4B in Ly10 cells denotes a PDE4B band. The data are presented as mean ± SD and representative of three independent experiments (*p < 0.05, One-way ANOVA). g Myc and PDE4B protein levels were measured by western blotting in Eμ-myc mice. High Myc expression was associated with elevated PDE4B expression. W and T represent B cells from wild type and transgenic mice, respectively. h Ly1 (PDE4B-high) cells were exposed to either vehicle or JQ1 (250 nM, 48 h), followed by frs (20 μM, 2 h). After the treatment, the same number of cells were used for the measurement of intracellular cAMP levels (Left panel). Expression of Myc and PDE4B and phosphorylation levels of SYK, AKT, and 4EBP1 in Ly1 cells were examined by western blotting, following treatment with JQ1 (0–250 nM) and frs (20 μM) for 48 h (Right panel). The data are presented as mean ± SD and representative of three independent experiments (*p < 0.05, Mann–Whitney U-test). i The relation between Myc and PDE4B at the mRNA level was investigated in 48 DLBCL cases (R2: Genomics Analysis and Visualization Platform, http://r2/amc.nl). The transcript levels of Myc and PDE4B were positively correlated (r = 0.319, p = 0.03)
Myc inhibition synergizes with PDE4 inhibition to suppress Myc expression and the survival of B lymphoma cells
In light of the important role of Myc and PDE4B for cell survival in B-cell malignancies, the disruption of the Myc-PDE4B loop may represent a promising therapeutic approach for the treatment of this type of tumor. We reasoned that the loop would collapse more effectively upon the simultaneous inhibition of these molecules. To test the proof of principle for the efficient disruption of the loop, we compared the levels of Myc following a single-drug treatment with those after combination therapy. Treatment with JQ1 or frs/rolipram alone exhibited a mild effect on Myc expression, whereas Myc levels were substantially reduced when these drugs were combined, highlighting the cooperative effects of these drugs in Myc suppression (Fig. S11). To directly ascertain whether Myc inhibition synergizes therapeutically with PDE4 inhibition, we measured cell viability (MTS assay) and apoptotic rate (PI staining) after suppression of Myc and/or PDE4 using pharmacological means. JQ1 or frs/rolipram alone had no or only a limited effect on cell survival or death, when compared with the combination treatment, which elicited a significant impact on cell viability or apoptosis (Fig. 4a, b). Moreover, we sought to quantitatively determine the interactions between JQ1 and frs/rolipram, or the drug-induced cytotoxic synergy. To that end, CI values were generated using CompuSyn software. Most data points indicated a strong or very strong synergism when Myc and PDE4 were concomitantly inhibited, reflecting that frs/rolipram substantially increased the antitumor effect of JQ1 in a highly synergistic manner and vice versa (Fig. 4c). JQ1 exhibited a similar combinatorial effect when co-administered with frs/roflumilast (Fig. S12). It has been shown that JQ1 can reduce the expression of a few hundred genes cell type-dependently [6], including genes implicated in survival and lymphomagenesis, such as Bcl2 and Bcl6, in addition to Myc. Thus, we employed 10058-F4 as a selective Myc inhibitor to dissect whether the proapoptotic effect of JQ1 was mainly attributed to the suppression of Myc in this setting. Overall, 10058-F4 reproduced the effect JQ1 on cell viability, albeit to a somewhat lesser extent (Fig. S13). These results have led us to postulate that the latter drug may deplete some other prosurvival factors or transcription factors than Myc to induce an apoptotic response in B lymphoma cells. Indeed, combinatorial treatment using JQ1 and frs/rolipram markedly downregulated Bcl2, Bcl6, BclXL, and Mcl1 (Fig. 4d); however, the effect of combination treatment with 10058-F4 and frs/rolipram was limited to Mcl1 (Fig. S14), thereby at least partly accounting for the better response to co-treatment with JQ1 and frs/rolipram. This data highlights that, although the Myc-PDE4B loop plays a critical role in regulating survival of B lymphoma cells, inclusion of JQ1 may have inhibitory effect on the expression of survival genes outside the loop, providing better therapeutic benefits. Altogether, these results suggest that the interplay between Myc and PDE4 plays an important role in the survival of B lymphoma cells.
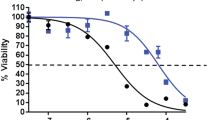
PDE4 and Myc inhibitors synergize in promoting cell death. a PDE4B-low (DHL4 and DHL6) and PDE4B-high (Ly1, Ly10, and Ramos) cells were exposed to frs (0–40 μM for 48 h)/JQ1 (0–600 nM for 48 h) either in the absence or presence of rolipram (40 μM for 48 h) as indicated, and MTS assays were performed to measure cell viability. The data are normalized to the respective values in vehicle-treated cells (no frs, rolipram, or JQ1). The results are presented as mean ± SD and representative of three independent experiments (*p < 0.05, One-way ANOVA). b Apoptotic rates were measured by PI-staining followed by FACS analysis in five B lymphoma cell lines treated with DMSO, frs (40 μM)/rolipram (40 μM), JQ1 (500 nM), or a combination of drugs for 48 h. The data represent the values after the subtraction of apoptosis rates in untreated cells. Representative data of three independent experiments showing similar trend. c Combination index (CI) values for quantifying drug synergy were obtained using the CompuSyn software using percent inhibition (fraction affected, Fa) from drug combinations as compared with single agents alone. Synergy levels are as follows: <0.1, very strong synergism; 0.1–0.3, strong synergism; 0.3–0.7, synergism; 0.7–0.85, moderate synergism; 0.85–0.90, slight synergism; 0.90–1.10, nearly additive; 1.10–1.20, slight antagonism; 1.20–1.45, moderate antagonism; and 1.45–3.30, antagonism. d Ly1 and Ly10 cells were treated with vehicle, frs (40 μM)/rolipram (40 μM), JQ1 (500 nM), or combination for 24 h and whole cell lysates were prepared. Expression of indicated pro-survival factors and a transcription factor Bcl6 was analyzed by western blotting. Representative data of at least three independent experiments are presented
Synergy between JQ1 and rolipram in vivo
To determine whether JQ1 interact with rolipram cooperatively in vivo, aggressive B lymphoma Ramos cells were subcutaneously implanted in athymic nude mice. Once the tumors became palpable, mice were randomly assigned to different treatment groups, followed by the daily administration of vehicle, JQ1, rolipram or a combination of JQ1 and rolipram. In agreement with our in vitro results, mice that received single agents showed a minimal response, whereas those receiving a combination of JQ1 and rolipram displayed a pronounced inhibition of tumor growth (Fig. 5a, b) and a decrease in the proportion of cells that stained positive for the nuclear antigen Ki-67, a cell proliferation marker (Fig. 5c). These observations were associated with a significant downregulation of Myc in mice co-treated with JQ1 and rolipram (Fig. 5d).
The combinatorial effect of rolipram and JQ1 in a B-cell lymphoma mouse xenograft model. a Five-week old athymic nude mice were subcutaneously inoculated with 1 × 107 Ramos cells. The size of the xenograft was calculated as πab/4 (a and b are the x, y diameters) [13]. The results represent mean ± SD and each point represents an average of 6 mice (*p < 0.05, One-way ANOVA; n = 6). b Representative xenograft tumors in each group are shown. c Xenograft tumors were extracted and processed for Ki-67 IHC staining (Left panel). Cells positive for Ki-67 were counted and the numbers were normalized to those obtained for the vehicle group (Right panel; *p < 0.05, One-way ANOVA). Cell proliferation was markedly inhibited by JQ1 and rolipram combination. d Myc IHC staining was performed in Ramos xenograft tumors (Left panel). Myc-positive cells were counted and the numbers were normalized to those for the vehicle group (Right panel; *p < 0.05, One-way ANOVA). Myc levels was substantially reduced upon exposure to the combination treatment
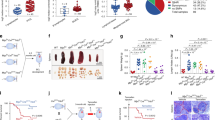
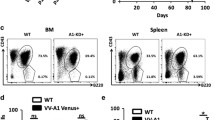
To verify these xenograft data in a more relevant setting of Myc-mediated lymphomagenesis, we utilized Eμ-myc transgenic mice that have been extensively used for studying Myc-driven aggressive B-cell lymphomas [40, 41]. Consistent with the widely accepted notion that the ectopic expression of Myc interferes with differentiation [3], Eμ-myc transgenic mice displayed strikingly elevated levels of pro-B and pre-B cells in the bone marrow and the spleen [41] (Fig. 6a, b, S1, and S15). To assess the role of PDE4 and its interaction with Myc in B lymphocyte development, we exposed the transgenic mice to vehicle, JQ1, rolipram or a combination treatment for 4 weeks and analyzed the B cell populations. The administration of rolipram plus JQ1 restored B cell differentiation to wild type levels, leading to decrease in pro-B and pre-B cells, while rolipram or JQ1 alone had a mild or negligible influence (Fig. 6a, b). Systemic toxicity of these agents was not found during the treatment; we did not observe any difference in body weights among four cohorts of Eμ-myc mice or wild type mice (Fig. S16). Additionally, the histopathologic analyses of spleens showed that the normal microscopic architecture of the spleen was completely collapsed by rapidly proliferating B cells (Ki-67+ and Myc+) in vehicle-treated Eμ-myc mice [14]; however, co-inhibition of Myc and PDE4 fully restored the normal architecture of the spleen (Fig. 6c; *p < 0.05, Fisher’s exact test), which was accompanied by substantial decline in Ki-67+ and Myc+ cells (Fig. 6d).
Synergistic effect of rolipram and JQ1 in a Eμ-myc transgenic mouse model. a, b Eight-week old Eμ-myc male mice were randomly divided into four treatment groups, vehicle, rolipram (5 mg/kg), JQ1 (10 mg/kg), and rolipram/JQ1 combination. Indicated drugs were administered for 4 weeks and developmental stages of splenic and bone marrow B-cell populations were analyzed using flow cytometry with the indicated key B-cell markers (Fig. S1). FACS data from a representative group is shown (Left panel). Developmental stages of splenic and bone marrow B-cell populations from different treatment groups are plotted, n = 7 in each group (Right panel; *p < 0.05, One-way ANOVA). c Representative images of the H&E stained spleen sections of wild type or Eμ-myc mice treated with vehicle or rolipram/JQ1 combination. The microscopic architecture of spleen in vehicle-treated Eμ-myc was collapsed, as compared with that of wild type mice, which was reestablished upon the combination treatment. W and R represent white pulp and red pulp, respectively (Left panel). Pathological analysis of spleens of Eμ-myc mice treated with indicated drugs was conducted. Findings of blastic B-cell lymphomas that were indicated by the disruption of the microscopic architecture of spleen were counted. Treatment with rolipram/JQ1 combination prevented development of blastic B-cell lymphomas (Right panel; *p < 0.05, Fisher’s exact test), while administration of rolipram or JQ1 alone did not elicit statistically significant change. Scale bar, 200 μm. d Spleen sections of Eμ-myc mice were processed for Ki-67 and Myc IHC staining, following treatment with indicated drugs. Ki67-positive and Myc-positive cells were counted and the numbers were normalized to the vehicle group (Bottom panel; *p < 0.05, One-way ANOVA). Scale bar, 100 μm
Most tumors in Eμ-myc mice are derived from clonal expansions of B lymphoblasts, which can be evidenced by reduced IgH gene diversity [20, 40]. To investigate the clonality of the tumors, we analyzed H-chain VDJ rearrangements in tumor cells (Fig. S17). Tumors from vehicle-treated or single agent-treated mice showed single or few VDJ rearrangements, indicative of limited IgH gene diversity. In contrast, co-treatment with rolipram and JQ1 clearly increased IgH gene diversity and reduced the clonality. These data implicate a critical role for the Myc-PDE4B loop in the regulation of Myc expression, B cell development, and clonal expansions of B lymphocytes.
Discussion
Deregulation of Myc has been recognized as a hallmark in most human cancers, including lymphoid malignancies [2, 3]. The recent development of BET bromodomain inhibitors shows great promise for the treatment of Myc-dependent tumors. Several BET protein inhibitors are currently in early phase clinical trials for a number of cancers, but their efficacy remain to be determined [5]. In this study, we sought to explore new strategies to therapeutically target Myc that could be rapidly translated to the clinic. To that end, we have demonstrated that a combination of PDE4 and the BET bromodomain inhibitors displayed potent antitumor effects in multiple B lymphoma cell lines, murine xenograft models, and Eμ-myc transgenic mice. The unique synergism of these inhibitors was driven by the cooperative disruption of the Myc-PDE4B feedforward loop.
Irrespective of the mechanism, our results have clinical implications. First, PDE4 is a therapeutically relevant target in B-cell malignancies. An FDA-approved PDE4 inhibitor, roflumilast, was successfully tested in patients with advanced B-cell malignancies in a recent clinical trial. The patients exhibited a partial response, or stable disease with the associated repression of PI3K/AKT signaling [11, 12]. Given that Myc is a downstream target of the PI3K/AKT/mTOR signaling cascade and plays an important role in malignant B cells, it is tempting to speculate that the favorable response to the drug may be attributed, at least in part, to the suppression of Myc (Figs. 1, 2). PI3K, a downstream kinase of the B-cell receptor, is critical for the survival of mature B cells, and this pathway is deregulated in a subset of B-cell lymphomas. The PI3Kδ isoform inhibitor idelalisib was approved for relapsed CLL and follicular lymphoma (FL) [42]. Although the data is limited, the loss of Myc downmodulation and the activation of the PI3K pathway were found to contribute to idelalisib resistance [43]. Intriguingly, I-BET762 BET inhibitor induced cell death in idelalisib-resistant U2932 ABC-DLBCL cells [44], suggesting that resistance to idelalisib could be overcome by treatment with BET inhibitors. Based on our results that combinatorial treatment of JQ1 and frs/rolipram markedly influenced AKT phosphorylation/activation as well as Myc expression (Figs. S11, S18), we propose that roflumilast and a BET inhibitor may be included in therapeutics involving idelalisib to enhance its efficacy and prevent potential resistance to the drug. One valid question here is whether B-lymphoma patients with low PDE4B expression would benefit from the inclusion of roflumilast. The present in vitro data using PDE4B-low DHL4 and DHL6 cell lines is against the speculation; addition of a PDE4 inhibitor rolipram did not enhance JQ1-mediated cell death (Fig. S19). This result suggests that patients with B-cell lymphoma may need to be dichotomized into PDE4B-low and PDE4B-high groups and combinatorial therapies including roflumilast would be more effective for the latter group; this warrants further study.
Second, we found that the Myc-PDE4B loop had dramatic influence on the levels of Mcl1 and phosphorylation of AKT and mTOR, and less so on the levels of other pro-survival Bcl2 family proteins and Bcl6 transcription factor (Figs. 4, S14, and S18). This is intriguing because high expression of Mcl1 is frequently observed in cancer and has been shown to be implicated in the resistance to ABT-737 the Bcl2 small molecule inhibitor [45]. Moreover, PI3K/AKT/mTOR activation has been reported to confer resistance to treatment with ABT-199, a Bcl2-specific inhibitor [46]. These imply that the inhibition of the Myc-PDE4B loop could be an effective strategy for overcoming the chemoresistance imparted by overexpression of Mcl1 or activation of the PI3K/AKT/mTOR signaling.
In conclusion, our work uncovered the critical role of interplay between Myc and PDE4B in the biology of B-cell lymphomas. The direct regulation of PDE4B by Myc sustains SYK/AKT/mTOR signaling, which in turn leads to an increase in Myc levels and promotes the viability of tumor cells. A recent study has delineated the role of PDE4B in regulating the secretion of vascular endothelial growth factor (VEGF) and vessel development [26]. This and our work offer a unique opportunity for targeting both the oncogenic properties of B lymphoma cells and angiogenesis in the tumor microenvironment, via the disruption of the Myc-PDE4B interaction, which may elicit synergistic killing of malignant B cells [26, 47]. Further mechanistic insights into the Myc-PDE4B feedforward loop may help improve the efficacy of PDE4 inhibitors and the BET bromodomain inhibitors in the treatment of B-cell malignancies.
References
Ott G. Impact of MYC on malignant behavior. Hematol Am Soc Hematol Educ Program. 2014;2014:100–6.
Dang CV. MYC on the path to cancer. Cell. 2012;149:22–35.
Eilers M, Eisenman RN. Myc’s broad reach. Genes Dev. 2008;22:2755–66.
Sewastianik T, Prochorec-Sobieszek M, Chapuy B, Juszczynski P. MYC deregulation in lymphoid tumors: molecular mechanisms, clinical consequences and therapeutic implications. Biochim Biophys Acta. 2014;1846:457–67.
Delmore JE, Issa GC, Lemieux ME, Rahl PB, Shi J, Jacobs HM, et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell. 2011;146:904–17.
Chaidos A, Caputo V, Karadimitris A. Inhibition of bromodomain and extra-terminal proteins (BET) as a potential therapeutic approach in haematological malignancies: emerging preclinical and clinical evidence. Ther Adv Hematol. 2015;6:128–41.
Moon EY, Lerner A. PDE4 inhibitors activate a mitochondrial apoptotic pathway in chronic lymphocytic leukemia cells that is regulated by protein phosphatase 2A. Blood. 2003;101:4122–30.
Ogawa R, Streiff MB, Bugayenko A, Kato GJ. Inhibition of PDE4 phosphodiesterase activity induces growth suppression, apoptosis, glucocorticoid sensitivity, p53, and p21(WAF1/CIP1) proteins in human acute lymphoblastic leukemia cells. Blood. 2002;99:3390–7.
Shipp MA, Ross KN, Tamayo P, Weng AP, Kutok JL, Aguiar RC, et al. Diffuse large B-cell lymphoma outcome prediction by gene-expression profiling and supervised machine learning. Nat Med. 2002;8:68–74.
Smith PG, Wang F, Wilkinson KN, Savage KJ, Klein U, Neuberg DS, et al. The phosphodiesterase PDE4B limits cAMP-associated PI3K/AKT-dependent apoptosis in diffuse large B-cell lymphoma. Blood. 2005;105:308–16.
Cooney JD, Aguiar RC. Phosphodiesterase 4 inhibitors have wide-ranging activity in B-cell malignancies. Blood. 2016;128:2886–90.
Kelly K, Mejia A, Suhasini AN, Lin AP, Kuhn J, Karnad AB, et al. Safety and pharmacodynamics of the PDE4 inhibitor roflumilast in advanced B-cell malignancies. Clin Cancer Res. 2017;23:1186–92.
Kim EA, Kim SW, Nam J, Sung EG, Song IH, Kim JY, et al. Inhibition of c-FLIPL expression by miRNA-708 increases the sensitivity of renal cancer cells to anti-cancer drugs. Oncotarget. 2016;7:31832–46.
Kim E, Nam J, Chang W, Zulfugarov IS, Okhlopkova ZM, Olennikov D, et al. Angelica gigas Nakai and Decursin downregulate Myc expression to promote cell death in B-cell lymphoma. Sci Rep. 2018;8:10590.
Balkhi MY, Willette-Brown J, Zhu F, Chen Z, Liu S, Guttridge DC, et al. IKKalpha-mediated signaling circuitry regulates early B lymphopoiesis during hematopoiesis. Blood. 2012;119:5467–77.
Cho Y, Song SH, Lee JJ, Choi N, Kim CG, Dean A, et al. The role of transcriptional activator GATA-1 at human beta-globin HS2. Nucleic Acids Res. 2008;36:4521–8.
Kwak B, Kim DU, Kim TO, Kim HS, Kim SW. MicroRNA-552 links Wnt signaling to p53 tumor suppressor in colorectal cancer. Int J Oncol. 2018;53:1800–8.
Jeong D, Kim J, Nam J, Sun H, Lee YH, Lee TJ, et al. MicroRNA-124 links p53 to the NF-kappaB pathway in B-cell lymphomas. Leukemia. 2015;29:1868–74.
Kim J, Jeong D, Nam J, Aung TN, Gim JA, Park KU, et al. MicroRNA-124 regulates glucocorticoid sensitivity by targeting phosphodiesterase 4B in diffuse large B cell lymphoma. Gene. 2015;558:173–80.
Turner SD, Tooze R, Maclennan K, Alexander DR. Vav-promoter regulated oncogenic fusion protein NPM-ALK in transgenic mice causes B-cell lymphomas with hyperactive Jun kinase. Oncogene. 2003;22:7750–61.
Andersson KB, Tasken K, Blomhoff HK. Cyclic AMP downregulates c-myc expression by inhibition of transcript initiation in human B-precursor Reh cells. FEBS Lett. 1994;337:71–6.
Pirson I, Coulonval K, Lamy F, Dumont JE. c-Myc expression is controlled by the mitogenic cAMP-cascade in thyrocytes. J Cell Physiol. 1996;168:59–70.
Williamson EA, Burgess GS, Eder P, Litz-Jackson S, Boswell HS. Cyclic AMP negatively controls c-myc transcription and G1 cell cycle progression in p210 BCR-ABL transformed cells: inhibitory activity exerted through cyclin D1 and cdk4. Leukemia. 1997;11:73–85.
Kim SW, Rai D, Aguiar RC. Gene set enrichment analysis unveils the mechanism for the phosphodiesterase 4B control of glucocorticoid response in B-cell lymphoma. Clin Cancer Res. 2011;17:6723–32.
Kim SW, Rai D, McKeller MR, Aguiar RC. Rational combined targeting of phosphodiesterase 4B and SYK in DLBCL. Blood. 2009;113:6153–60.
Suhasini AN, Wang L, Holder KN, Lin AP, Bhatnagar H, Kim SW, et al. A phosphodiesterase 4B-dependent interplay between tumor cells and the microenvironment regulates angiogenesis in B-cell lymphoma. Leukemia. 2016;30:617–26.
Zeller KI, Zhao X, Lee CW, Chiu KP, Yao F, Yustein JT, et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc Natl Acad Sci USA. 2006;103:17834–9.
Carnevale J, Ross L, Puissant A, Banerji V, Stone RM, DeAngelo DJ, et al. SYK regulates mTOR signaling in AML. Leukemia. 2013;27:2118–28.
Guertin DA, Sabatini DM. The pharmacology of mTOR inhibition. Sci Signal. 2009;2:pe24.
Saxton RA, Sabatini DM. mTOR signaling in growth, metabolism, and disease. Cell. 2017;169:361–71.
Wall M, Poortinga G, Hannan KM, Pearson RB, Hannan RD, McArthur GA. Translational control of c-MYC by rapamycin promotes terminal myeloid differentiation. Blood. 2008;112:2305–17.
Shi Y, Sharma A, Wu H, Lichtenstein A, Gera J. Cyclin D1 and c-myc internal ribosome entry site (IRES)-dependent translation is regulated by AKT activity and enhanced by rapamycin through a p38 MAPK- and ERK-dependent pathway. J Biol Chem. 2005;280:10964–73.
West MJ, Stoneley M, Willis AE. Translational induction of the c-myc oncogene via activation of the FRAP/TOR signalling pathway. Oncogene. 1998;17:769–80.
Wossning T, Herzog S, Kohler F, Meixlsperger S, Kulathu Y, Mittler G, et al. Deregulated Syk inhibits differentiation and induces growth factor-independent proliferation of pre-B cells. J Exp Med. 2006;203:2829–40.
D’Sa C, Tolbert LM, Conti M, Duman RS. Regulation of cAMP-specific phosphodiesterases type 4B and 4D (PDE4) splice variants by cAMP signaling in primary cortical neurons. J Neurochem. 2002;81:745–57.
Verghese MW, McConnell RT, Lenhard JM, Hamacher L, Jin SL. Regulation of distinct cyclic AMP-specific phosphodiesterase (phosphodiesterase type 4) isozymes in human monocytic cells. Mol Pharmacol. 1995;47:1164–71.
Jin SL, Conti M. Induction of the cyclic nucleotide phosphodiesterase PDE4B is essential for LPS-activated TNF-alpha responses. Proc Natl Acad Sci USA. 2002;99:7628–33.
Blackwell TK, Huang J, Ma A, Kretzner L, Alt FW, Eisenman RN, et al. Binding of myc proteins to canonical and noncanonical DNA sequences. Mol Cell Biol. 1993;13:5216–24.
Grandori C, Mac J, Siebelt F, Ayer DE, Eisenman RN. Myc-Max heterodimers activate a DEAD box gene and interact with multiple E box-related sites in vivo. EMBO J. 1996;15:4344–57.
Adams JM, Harris AW, Pinkert CA, Corcoran LM, Alexander WS, Cory S, et al. The c-myc oncogene driven by immunoglobulin enhancers induces lymphoid malignancy in transgenic mice. Nature. 1985;318:533–8.
Langdon WY, Harris AW, Cory S, Adams JM. The c-myc oncogene perturbs B lymphocyte development in E-mu-myc transgenic mice. Cell. 1986;47:11–8.
Cheah CY, Fowler NH. Idelalisib in the management of lymphoma. Blood. 2016;128:331–6.
Meadows S, Rick S, Anella Y, Liu J, Li L, Yue P, et al. Up-regulation of the PI3K signaling pathway mediates resistance to Idelalisib. Blood. 2015;126;3707.
Lasorsa E, Smonksey M, Kirk JS, Rosario S, Hernandez-Ilizaliturri FJ, Ellis L. Mitochondrial protection impairs BET bromodomain inhibitor-mediated cell death and provides rationale for combination therapeutic strategies. Cell Death Dis. 2015;6:e2014.
van Delft MF, Wei AH, Mason KD, Vandenberg CJ, Chen L, Czabotar PE, et al. The BH3 mimetic ABT-737 targets selective Bcl-2 proteins and efficiently induces apoptosis via Bak/Bax if Mcl-1 is neutralized. Cancer Cell. 2006;10:389–99.
Choudhary GS, Al-Harbi S, Mazumder S, Hill BT, Smith MR, Bodo J, et al. MCL-1 and BCL-xL-dependent resistance to the BCL-2 inhibitor ABT-199 can be overcome by preventing PI3K/AKT/mTOR activation in lymphoid malignancies. Cell Death Dis. 2015;6:e1593.
Lenz G, Wright G, Dave SS, Xiao W, Powell J, Zhao H, et al. Stromal gene signatures in large-B-cell lymphomas. N Engl J Med. 2008;359:2313–23.
Acknowledgements
pcDNA-Myc construct is a gift from Dr. Dongchul Kang (Hallym University, Korea) and Ly10 DLBCL cell line was kindly provided by Dr. Yoon Kyung Jeon (Seoul National University Hospital, Korea). This work was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (NRF-2013R1A1A2008838 and NRF-2016R1A2B4011758) to S-WK and (NRF-2014R1A5A2010008 and 2018R1A2B2007410) to Y-HL. IP was a recipient of the research fund 2018 through Youngsan University.
Author information
Authors and Affiliations
Contributions
JN performed experiments, analyzed the data, and wrote the paper; DK, EK, BK, MK, H-JS, IP and J-YJ performed experiments; A-YO and B-JP helped with mouse breeding; YWK and AK helped with ChIP experiments; HS performed statistical analysis; YJ and J-HL helped with tissue staining and qRT-PCR, respectively; D-KS and Y-HL supervised the study and wrote the manuscript; S- WK designed and supervised the study, planned experiments, analyzed the data and wrote the paper. All the authors edited the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Nam, J., Kim, D., Kim, E. et al. Disruption of the Myc-PDE4B regulatory circuitry impairs B-cell lymphoma survival. Leukemia 33, 2912–2923 (2019). https://doi.org/10.1038/s41375-019-0492-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-019-0492-y
- Springer Nature Limited
This article is cited by
-
C-MYC-activated lncRNA SNHG20 accelerates the proliferation of diffuse large B cell lymphoma via USP14-mediated deubiquitination of β-catenin
Biology Direct (2024)
-
Low NDRG2, regulated by the MYC/MIZ-1 complex and methylation, predicts poor outcomes in DLBCL patients
Annals of Hematology (2024)
-
Reimagining high-throughput profiling of reactive cysteines for cell-based screening of large electrophile libraries
Nature Biotechnology (2021)