Abstract
Multiple myeloma (MM) is a largely incurable haematological malignancy defined by the clonal proliferation of malignant plasma cells (PCs) within the bone marrow. Clonal heterogeneity has recently been established as a feature in MM, however, the subclonal evolution associated with disease progression has not been described. Here, we performed whole-exome sequencing of serial samples from 10 patients, providing new insights into the progression from monoclonal gammopathy of undetermined significance (MGUS) and smouldering MM (SMM), to symptomatic MM. We confirm that intraclonal genetic heterogeneity is a common feature at diagnosis and that the driving events involved in disease progression are more subtle than previously reported. We reveal that MM evolution is mainly characterised by the phenomenon of clonal stability, where the transformed subclonal PC populations identified at MM are already present in the asymptomatic MGUS/SMM stages. Our findings highlight the possibility that PC extrinsic factors may play a role in subclonal evolution and MGUS/SMM to MM progression.
Similar content being viewed by others
Introduction
Multiple myeloma (MM) is a haematological malignancy characterised by the uncontrolled proliferation of neoplastic plasma cells (PCs) within the bone marrow (BM). MM accounts for ~10% of all haematological malignancies [1], with a median survival rate of 5.2 years [2]. Although ten new therapeutic agents for MM have been approved in the last 20 years, and patient outcomes have improved significantly, individual responses to therapy and overall survival are varied [3]. MM remains largely incurable, with relapse being a common feature of disease.
The development of MM has been classically viewed as a multistage process involving the acquisition of multiple genetic mutations, with immunoglobulin heavy chain translocations and hyperdiploidy known to be common initiating events that deregulate normal PC behaviour leading to the development of monoclonal gammopathy of undetermined significance (MGUS) [4,5,6,7]. Further mutational load leads to an intermediate stage of smouldering multiple myeloma (SMM) [4, 8]. However, these common initiating events are insufficient to cause MM transformation, as MGUS/SMM patients commonly harbour these abnormalities and show no clinical symptoms of MM [9, 10]. Studies have shown that progression to MM is associated with additional genetic changes including aneuploidy, chromosomal translocations, single nucleotide variants, small insertions and deletions, and copy number variants affecting one or more genes, with mutations present at a frequency of 0.1 to 10 per megabase [11].
Recent studies show that MM patients display complex mutational landscapes involving intraclonal genetic heterogeneity at the bulk tumour level, where mutations are acquired in a non-linear branching pattern [12,13,14,15,16,17]. Intraclonal heterogeneity has been observed at all stages of MM, suggesting that disease progression may be mediated through inter-subclone competition and outgrowth of the fittest of these subclones. Genomic studies on large cohorts of unmatched MGUS-SMM-MM patients have led to the discovery of recurrently mutated genes, of which KRAS, NRAS, TP53, BRAF, FAM46C and DIS3 are believed to be drivers of MM transformation [10, 18,19,20,21]. While clonal heterogeneity is now an established feature in MM, the subclonal evolution associated with MGUS/SMM to MM progression remains poorly understood.
A comprehensive approach to identifying the key somatic mutations and infer the subclonal evolution associated with MM transformation, involves the longitudinal study of sequential MGUS-MM or SMM-MM samples from the same patient. However, because MGUS is often an incidental finding, it is extremely rare to have diagnostic BM samples from the same patient at both the MGUS and MM stages. In addition, because there are no cell line or mouse models of MGUS or SMM, there has been limited opportunity to study the specific genetic changes and molecular mechanisms that characterise the progression from MGUS/SMM to MM.
Here, we report a longitudinal genomics investigation of MM, based on paired MGUS-MM (n = 5) or SMM-MM (n = 5) patient samples obtained from the same patient when initially diagnosed at MGUS/SMM, and subsequently when they developed MM. Using whole-exome sequencing, we have obtained a detailed description of the genomic complexity and subclonal evolution underlying progression from MGUS/SMM to MM.
Materials and methods
Clinical samples
Bone marrow mononuclear cell aspirates were collected from patients at MGUS/SMM, and subsequently at later diagnosis of MM (MGUS-MM (n = 5) and SMM-MM (n = 5)). The median time to progression of MGUS to MM was 3.2 years (range 1–13 years) and SMM to MM was 1.2 years (range 0.48–4.1 years). Where available, the cytogenetic status of patients is reported in Supplementary Table 1. Samples were collected from patients prior to treatment. All patients provided informed consent in accordance with the Declaration of Helsinki. Samples were cryopreserved by the South Australian Cancer Research Biobank (SACRB) at SA Pathology. The studies were approved by the Royal Adelaide Hospital Human Research Ethics Committee (HREC/13/RAH/569 No: 131133). Samples were collected over a period of 22 years, and we initially began this study with paired-samples from 18 patients. However, due to our strict criteria for sample purity and mutation calling resolution, final analysis was only performed on samples from 10 patients.
Cell sorting
PCs and non-tumour cells were purified using multicolour flow cytometry as previously described [21]. Briefly, ~1 × 105 mononuclear cells were prepared for single-stain antibody control (CD138-PE (Beckman Coulter #A54190) and CD38-PE-Cy7 (Biolegend #303515)) and compensation/FMO tubes (1: unstained; 2: hydroxystilbamidine (FluoroGold; Life Technologies) only; 3: CD38-PE-Cy7 + FluroGold; 4: CD138-PE + FluroGold; and 5: CD38-PE-Cy7 + CD138-PE). The sort sample was stained with CD138-PE and CD38-PE-Cy7 antibody at 1 μL/100 μL cells. Cells were stained with FluoroGold immediately prior to sorting. Viable PCs (CD138+CD38+ and FluoroGold negative) and non-tumour cells were sorted on the FACSAria Fusion sorter (BD Biosciences). FACS purity check was carried out on sorted cells, using 100–500 cells from each sample.
DNA isolation, QC and sequencing
DNA was isolated from purified PC and non-tumour populations using the All Prep DNA/RNA Micro Kit (QIAGEN) as per manufacturers’ instructions. Yields and quality was assessed using the NanoDrop 8000 and Qubit 2.0 fluorometer (Thermo Fisher Scientific).
A total of 115 ng of gDNA was used as an input for fragmentation on the Covaris E220, followed by end-repair/A-tailing and ligation of SureSelect Adapter Oligos (Agilent). Pre-Capture PCR amplification of 10 cycles, or 12 cycles for low input samples, were performed. A total of 750 ng of each sample was hybridised to SureSelect XT Clinical Research Exome (Agilent) probes overnight. Captured DNA was amplified with 11 cycles of post-capture PCR incorporating index barcodes. Sequencing was performed on the Illumina HiSeq4000 (2 × 100 bp paired-end reads) and NextSeq 500 (2 × 150 bp paired-end reads). Samples were sequenced to a minimum depth of ~140 × mean coverage. Isolated non-tumour cells were also sequenced to a similar average depth (138x).
Analysis of data
Sequencing reads were mapped to the human decoy genome (hs37d5) using Novoalign (v3.02.08), followed by post-processing according to GATK best practices [22]. Somatic single nucleotide and small indel variants were called using MuTect2 [23] and multiSNV [24]. Variants were filtered based on: 10+ reads covering the variant site; 5+ reads covering the variant in the tumour sample. Variants were annotated with SnpEff [25].
R 3.3.2 was used throughout for analyses. Somatic copy number variants were called using CNVkit [26] v0.7.11 and custom in-house methods developed to support highly aneuploid genomes to perform segmentation and calculate log2 changes.
Clonal evolution was investigated using PhyloWGS [27] and visualised using fish plot in R [28]. PhyloWGS is noted to inflate the number of subclones, thus we recognise that subclone numbers may be overestimated. All phylogenetic trees constructed were based on the assumption that there is a single founder clone.
Additional information on sequencing and somatic mutation analysis is given in the supplementary methods.
Data deposition
All raw sequencing reads have been deposited in the EGA repository (Accession number: EGAS00001002850).
Code availability
Custom script generated for CNV analysis is available on request from corresponding authors.
Results
A changing spectrum of acquired mutations, not mutational load, is associated with MM progression
Whole-exome sequencing was performed on paired MGUS/SMM to MM patients (detailed in Supplementary Table 1) to a minimum average depth of 140x (Supplementary Table 2). A total of 1614 somatic non-synonymous single nucleotide variants (NS-SNVs) were identified across the MGUS/SMM samples (range 30–220) with a median 161 per patient. Interestingly, in the MM samples, we identified a total 1508 somatic NS-SNVs (range 59–226), with a median 152 per patient. There was an average of 27 NS-SNVs that were shared between the MGUS/SMM and MM stage (range 0–53) (Supplementary Table 3). We observed a moderately higher mutation load compared to previous larger cohort studies of MM, which identified median SNV numbers of 31 (range 15–46) [4], and 52 mutations per patient (range 2–488) [18].
Recent sequencing studies of unpaired MM samples have described an increasing median NS-SNV burden from MGUS to SMM to MM, with MGUS patients harbouring approximately half the number of NS-SNVs when compared to unmatched MM patients [4], with an average of 35 at the MM stage [10]. Here, we observed the opposite upon progression from MGUS/SMM to MM, where seven out of ten patients showed a decrease in total mutational load (Fig. 1a). While the total mutational burden is not considerably different between MGUS/SMM and MM, the presence of intraclonal heterogeneity and changes in the spectrum of mutated genes between disease stages, suggests that there is waxing and waning of subclones over time [29].
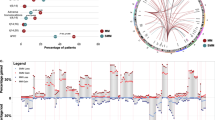
Pattern of genetic mutations in MGUS/SMM to MM progression. a Total NS-SNV mutational load associated with progression from MGUS/SMM to MM in individual patients. b Waterfall diagram indicating the 10 most frequently mutated genes associated with progression from MGUS/SMM to MM. c We identified mutations in 15 reported recurrently mutated MM genes in our MGUS/SMM to MM samples. However, individual patients harbour a heterogeneous genetic architecture, with a combination of mutations in known driver (KRAS, NRAS and DIS3) and other candidate genes. Mutations in RAS/MAPK pathway genes are most prevalent. d Gradient diagram across all patients indicating the variant allele frequencies (VAF) of identified known cancer drivers. RAS mutations are observed to mainly occur subclonally
We next examined the changing mutational landscape associated with MGUS/SMM to MM progression, to identify the genetic aberrations associated with this process, including both previously reported ‘drivers’ of MM and frequently acquired mutations present at MM transition. To this end, we identified 2566 unique genes with acquired variants at MM transition across all patients. The most common genes harbouring mutations at MM include KRAS, KMT2D, RYR2, DNAH5, PCDH8, RP4-669P10.16, DIS3, FAT3, PKHD1 and SP140 (Fig. 1b). Moreover, we identified 15 previously reported recurrently mutated genes: KRAS, FAT3, DIS3, TRAF3, SP140, RB1, PTEN, ROBO1, PRDM1, NRAS, MYC, MAGED1, IRF4, HLA-A and CDK4 [10, 18,19,20, 30] (Fig. 1c), including mutations in three known ‘drivers’ of MM: KRAS, NRAS and DIS3 (Supplementary Table 4). In our samples, mutations in KRAS and NRAS, were mutually exclusive, consistent with previous observations that report the rare co-occurrence of mutations in these genes (in 2% of patients) [20].
The RAS/MAPK pathway was highly mutated with 40% of patients at MGUS/SMM, and 70% at MM, harbouring mutations in KRAS and NRAS. DIS3 was mutated in 30% of patients at the MM stage only (Fig. 1d). This highlights that driver mutations can be acquired at both the asymptomatic stages and be maintained during progression to MM, or be acquired later at MM. However, we found that acquisitions of driver mutations are subclonal in nature. Low variant allele frequencies were identified for RAS pathway mutations (KRAS range 0.024–0.53, NRAS 0.03–0.28), suggesting that these mutations were present in subclonal PC populations during progression (P01, P02, P05 and P06) (Fig. 1d). Interestingly, acquisition of DIS3 mutations at the MM stage in patients P07 and P10 was observed to be clonal in nature [31] (Fig. 1d).
We also characterised the copy number variation (CNV) landscape associated with MGUS/SMM to MM progression, finding copy changes to be widespread (Fig. 2a). This contrasts a recent small longitudinal study of SMM to MM transformation that showed that copy changes are a feature of early stage of MM disease and not associated with progression [4]. We observed that MGUS/SMM patients harbour a similar frequency of chromosomal loci copy gains and losses than MM patients, with a median of 70 at MGUS/SMM (range 19 to 114), and 67.5 at MM (range 43 to 103) (Supplementary Table 5). Upon progression, we observed known frequent chromosomal copy number abnormalities in MM, including amplifications on chromosome arms 1q, 3p, 6p, 9p, 11q, 19p, 19q and 21q, coupled with losses on chromosome arms 1p, 6q, 8p, 13q, 16q and 22q across patients in the cohort (Fig. 2b). While we performed a gene level copy number analysis, we did not find any genes that were consistently gained or lost in our cohort upon progression. Interestingly, we also observed that many of the cytogenetic abnormalities associated with MM are present at MGUS/SMM stages and that standard cytogenetic methods did not accurately capture these abnormalities (Supplementary Figure 4).
The chromosomal copy number variation landscape associated with MGUS/SMM to MM progression. a Chromosomal copy number landscape plot illustrating the proportion of patients with copy amplifications (blue) and deletions (red) of chromosomes across all patients associated with MGUS/SMM to MM progression. Grey traces the average in the cohort as a whole, where below 0 indicates loss and above 0 represents gain. b Frequent previously reported copy number changes of MM were identified, including gains on 1q, 3p, 6p, 9p, 11q, 19p, 19q and 21q; and losses on 6q, 8p, 13q, 16q and 22q being present at the asymptomatic stages and maintained with progression to MM
The subclonal architecture required for MM progression exists at MGUS/SMM diagnosis
While clonal heterogeneity is an established feature in MM, the subclonal evolution associated with disease progression has not been well explored. Due to the nature of our paired longitudinal patient samples, we were able to directly examine the relationship between genetic variegation and clonal structure to construct evolutionary trajectories accompanying progression to MM in eight individual patients.
Comparisons of unpaired MGUS/SMM and MM samples have shown that MGUS/SMM exhibits mutational similarity with MM, but many mutations were present in a smaller proportion of aberrant PCs [32, 33]. Similarly, small paired-sample studies examining the evolution over time of asymptomatic monoclonal gammopathies (AMGs) to MM (n = 4) [34], and high-risk SMM to MM patients (n = 4) [4], have also found that most somatic changes required for MM were present at the asymptomatic stages, with the clinically dominant MM subclone present at the SMM stage. Therefore, the occurrence of clonal evolution in MM represents a change in clonal heterogeneity over time from the asymptomatic stages to MM [13].
In both MGUS-MM (P01, P04 and P10) and SMM-MM (P02, P03, P05, P06 and P08) progression, we find a prevailing model of evolution defined by clonal stability. This is where the transformed subclonal PC populations identified at MM were already present in the asymptomatic MGUS/SMM stages. Progression to MM involved subtle changes in the existent subclonal structure from MGUS/SMM, coupled with a degree of emergence and/or extinction of child subclonal branches (Figs. 3 and 4). Of note, we observed that subclonal evolution has already begun prior to MGUS/SMM sampling. While multiple subclonal populations are present at MGUS/SMM diagnosis, each patient harbours unique set of oncogenic mutations driving MM progression.
The subclonal tumour evolution associated with MGUS to MM progression. Fishtail plots illustrate the subclonal architecture in MGUS-MM of two patients (a: P01 and b: P04), which was defined by the existence of between 5 and 8 PC subclones at MGUS diagnosis. These subclonal populations generally progress to MM in a stable manner, in combination with the coupled emergence and/or extinction of child subclones. Key mutations in the founder clone and subclones are highlighted, with mutations in driver genes identified at both the clonal and subclonal level. The full-annotated subclonal genetic architecture for all patients can be found in Supplementary Figure 1
The subclonal tumour evolution associated with SMM to MM progression. Fishtail plots illustrating the subclonality in SMM-MM of two patients (a: P02 and b: P03), with the existence of between 5 and 11 PC subclones at the SMM stage. Notably, in comparison to the subclonal architecture at MGUS diagnosis, we observe a similar number of subclones present at SMM. Similar to the MGUS subclones, these SMM subclonal populations generally progress to MM in a stable manner, in combination with the coupled emergence and/or extinction of child subclones. Key mutations in MM genes in the founder clone and subclones are highlighted, where mutations in driver genes were found to be both clonal and subclonal in nature. The full-annotated subclonal genetic architecture for all patients can be found in Supplementary Figure 2
Subclonal tumour evolution in MGUS-MM patients
Three patients (P01, P04 and P10) were initially diagnosed with MGUS, and subsequently with MM. Typically, an average of 7 subclones were identified in MGUS sample pairs. We describe two examples, with the full-annotated subclonal architecture for all MGUS-MM patients found in Supplementary Figure 1/Appendix 1.
Patient P01 exhibited a modest increase in NS-SNV mutations with progression and was composed of eight subclones at diagnosis. The founder clone had a copy number change on chromosome 1. Interestingly, while P01 mainly exhibited stable progression of subclones from MGUS to MM, we observed KRAS mutations to be newly acquired in multiple child subclones. Subclone 7 (brown) harboured a mutation causing an amino acid change at Q61L, with a resultant neutral growth observed. Furthermore, we identified mutations occurring in a nested fashion, with outgrowth of subclone 8 (grey from <1 to ~6%) harbouring mutations at G12D and G12S, with further emergence of child subclone 9 (green) harbouring additional change at Q61H with MM progression. This was coupled with the extinction of child subclonal branches of subclone 2 (purple) (Fig. 3a).
Patient P04 exhibited an interesting subclonal evolution pattern, where initially one subclone (subclone 2 purple) evolved from the founder clone, which was followed by substantial branching evolution resulting in six child subclones involved in MM progression. The founder clone harboured mutations in MYCBP2 (F22L) and TOP2A (K1199N) and copy number changes on chromosomes 9, 11, 13, 14 and 18. While most of the child subclones exhibit stability, subclone 3 (orange from <1 to ~18%) and subclone 9 (green from <1 to ~9%) appear to have a selective advantage and showed emergence towards MM (Fig. 3b). Similarly, P10 was composed of eight subclones at MGUS with the neutral growth of subclonal populations coupled with the emergence of multiple subclones (subclone 4 yellow from ~3 to ~25%, and subclone 5 blue from ~1 to ~10%) and extinction of child subclone 3 (orange from ~5 to <1%) with progression (Supplementary Figure 1c).
Subclonal tumour evolution in SMM-MM patients
Five patients (P02, P03, P05, P06 and P08) were diagnosed for SMM, and then subsequently MM at a later time point. Generally, an average of 8 subclones were identified in SMM-MM pairs. We report two examples, with the full-annotated subclonal architecture for all SMM-MM patients found in Supplementary Figure 2/Appendix 1.
Patient P02 was composed of eleven subclones at diagnosis and exhibited stable growth during progression with mainly the emergence of child subclone 5 and its branches (blue from ~5 to ~13%) and extinction of subclone 9 (dark green from ~6 to <1%) (Fig. 4a). The founder clone showed copy changes on chromosomes 6, 8 and 13, and mutations in HERPUD1 (STOP gain), FGFR3 (809S) and DAPK1 (K435R). Furthermore, we identified a KRAS mutation (A146P) in subclone 11, whose population proportion size, interestingly, did not change during MM progression.
Patient P03 displayed an interesting evolution pattern with massive extinction of subclone 2 (purple) from ~47 to ~6%, and almost all of its child subclones, by MM diagnosis. The founder clone harboured mutations in NOD2 (STOP gain) and CNVs on chromosomes 1, 6, 9, 13 and 16. Furthermore, two individual subclones that contained distinct DIS3 mutants M566K and R689P were identified at SMM diagnosis in subclone 8 (black) and child subclone 11 (dark purple), respectively (Fig. 4b). While recent single-cell analysis has demonstrated parallel evolution of the RAS/MAPK pathway in MM through the occurrence of RAS mutations in individual clones leading to distinct subclonal populations [35], here we uniquely identify parallel evolution of DIS3, with the resultant emergence of both subclonal lineages with MM progression. Additionally, subclone 13 and its child subclones exhibited outgrowth with a mutation in NEK2 (L39H) (light green from <1 to ~7%).
Clonal stability in tumour evolution is also exemplified in other SMM-MM patients (P05, P06 and P08), which were characterised by 7, 5 and 8 subclones at diagnosis, respectively, and exhibited coupled emergence and extinction of child subclones in the progression to MM. In P05, there were initially two subclones that progressed to MM with the emergence and extinction of child clones from subclonal branch 5 (blue), combined with the neutral growth from subclonal branch 2 (purple) (Supplementary Figure 2c). Similarly, in P06, with progression we observed the emergence of subclone 8 (black from <1 to ~23%) and its child subclone 9 (dark green from <1 to ~4%), and subclone 2 (purple from ~8 to ~28%) and its child subclones 6 (pink from <1 to ~10%) and 7 (brown from <1 to ~6%). The proportions of child subclonal population 3 (orange) remained unchanged between SMM and MM (Supplementary Figure 2d). Patient P08 exhibited neutral growth, which was coupled with the emergence of child subclone 9 (green from <1 to ~5%) and extinction of child subclone 8 (black from ~6 to <1%) with MM progression (Supplementary Figure 2e).
Our analysis reveals conclusive evidence of intraclonal heterogeneity and subclonality from the earliest MGUS/SMM stages, where most of the transformed subclonal populations involved in progression to MM are already present at diagnosis. Notably, we do not observe a remarkable difference in the subclonality characteristic at the initial asymptomatic MGUS stage (average 7 subclones) and the intermediate SMM stage (average 8 subclones). This suggests that major subclonal remodelling is also not a phenomenon associated with advancement between the asymptomatic stages.
Discussion
The longitudinal investigation of MGUS/SMM to MM samples using NGS has revealed a new understanding of the underlying genetic architecture and subclonal evolution associated with MM progression. Analysis of MGUS-MM, and SMM-MM transition has shown that intraclonal heterogeneity is present at the asymptomatic stages. We find that progression is associated with an altered landscape of acquired mutations, rather than an increased total mutational burden.
Cancer progression models propose either the sequential accumulation of key genetic mutations throughout progressive disease and clonal expansions (‘Darwinian’ evolution), or punctuated bursts of large-scale chromosomal alterations (‘Saltationist’ evolution) [36]. The current understanding of MM transformation involves a sequential nature of evolution from the well-defined asymptomatic stages of MGUS and SMM, characterised by clonal expansion of PCs, and branching ‘Darwinian’ evolution with the presence of 2 to 6 subclones, highlighting clonal heterogeneity at MM presentation [10, 18,19,20, 29, 30, 35, 37,38,39,40,41]. In this model it is recognised that progression from the asymptomatic stages is dependent on the rise and fall in dominance of PC subclones based on their clonal fitness. The acquisition of driver mutations confers a selective advantage and facilitates better survival properties allowing the subclones to survive the selective pressures of the microenvironment/immune system and progress to symptomatic MM.
Notably, our study establishes MM disease progression to be characterised by the phenomenon of clonal stability, where substantial remoulding of the subclonal populations from the asymptomatic stages is not a necessary prerequisite for progression to MM. We found the existence of multiple PC subclones (range 5–11) at both MGUS and SMM that were intrinsic in the development and progression of MM. Furthermore, by comparing patients at MGUS and SMM stages we identified no significant difference in the number of PC subclones present at diagnosis (with an average of 7 versus 8, respectively). This is striking, as progression between the asymptomatic stages of MGUS and SMM is currently distinguished by an increased BM PC% and monoclonal protein level. We also found no correlation between the extent of subclonality and BM PC% at the MM stage in patients (Supplementary Figure 3). Similarly, a recent study of four high-risk SMM to MM transformation patients revealed that clonal progression was the key feature of MM onset, where the invasive clinically predominant clone typical of MM, was already present at SMM [4]. Similarly, in their investigation, Walker et al. reported a shifting clonal structure with the outgrowth and reduction of subclonal populations from SMM to MM [4]. Taken together, we hypothesise that patients who progress within a short time frame, MGUS/SMM to MM transformation does not always required the acquisition of many additional mutations and clonal selection. These MGUS/SMM patients appear to be sufficiently genetically complex to be on the threshold of transformation to MM, which may possibly be driven by extrinsic factors.
Being able to define the crucial oncogenic events in the founder clonal population could facilitate treatment strategies for early intervention to arrest MM progression. However, as the subclones responsible for MM are evident at the asymptomatic stages this poses the question as to why these patients are not symptomatic. A strong possibility is that further to intrinsic genetic factors, extrinsic factors such as the tumour microenvironment may also play an important role in defining both the subclonal architecture and the overall tumour cell burden for progression to clinical malignancy. The complex interactions of the tumour microenvironment with subclones provide signals that may support tumour growth or dormancy, which may influence their transformation [42,43,44,45,46,47,48,49]. Of note, a recent study from our group which used the C57BL/KaLwRij mouse model of MM, demonstrated habitual clonal dominance, where only a few establishing MM cells subsequently contributed to tumour burden while most remained dormant. This illustrates the strong selection pressures present within the BM microenvironment which plays a role in defining the clonal architecture [46].
The current standard of care at the asymptomatic stages involves monitoring patients, with no treatment options until they progress to symptomatic MM. Here our study has demonstrated that there is no significant shift in subclonal structure associated with MM progression. As such, subclonal populations present at MGUS/SMM diagnosis would be just as amenable to treatment, and eradication of these subclonal populations prior to disease transformation could delay progression and may provide the prospect of a durable cure [50]. However, we recognise that intraclonal heterogeneity has been shown to be characteristic of MGUS/SMM/MM, with multiple subclones having differing survival properties, therefore the risk of further mutation and tumour evolution due to drug selection pressures would eventually lead to relapse. Furthermore, intraclonal heterogeneity with clonal selection may not be the only defining evolution associated with progression of MM, with a recent study illustrating the involvement of spatial heterogeneity with regional site seeding and outgrowth resulting in progression [51]. Therefore, a combined longitudinal and spatial study of progression in patients would further elucidate genomic biomarkers of MM tumour evolution [52], although the ability to sample from multiple sites in asymptomatic patients has significant ethical and practical challenges.
Our findings reveal new insights into the genomic complexity and subclonal tumour evolution that is present from MGUS/SMM through to MM transformation. The existence of subclonality and clonal stability as a model of tumour evolution not only provides a more comprehensive understanding of the underlying biology of MM disease progression, but also new considerations required for patients at diagnosis and future therapeutic approaches to control this disease.
References
Rajkumar SV. Multiple myeloma: 2016 update on diagnosis, risk-stratification, and management. Am J Hematol. 2016;91:719–34.
Kumar SK, Dispenzieri A, Lacy MQ, Gertz MA, Buadi FK, Pandey S, et al. Continued improvement in survival in multiple myeloma: changes in early mortality and outcomes in older patients. Leukemia. 2014;28:1122–8.
Smith D, Yong K. Advances in understanding prognosis in myeloma. Br J Haematol. 2016;175:367–80.
Walker BA, Wardell CP, Melchor L, Brioli A, Johnson DC, Kaiser MF, et al. Intraclonal heterogeneity is a critical early event in the development of myeloma and precedes the development of clinical symptoms. Leukemia. 2014;28:384–90.
Palumbo A, Anderson K. Multiple myeloma. N Engl J Med. 2011;364:1046–60.
Landgren O, Kyle RA, Pfeiffer RM, Katzmann JA, Caporaso NE, Hayes RB, et al. Monoclonal gammopathy of undetermined significance (MGUS) consistently precedes multiple myeloma: a prospective study. Blood. 2009;113:5412–7.
Kyle RA, Therneau TM, Rajkumar SV, Offord JR, Larson DR, Plevak MF, et al. A long-term study of prognosis in monoclonal gammopathy of undetermined significance. N Engl J Med. 2002;346:564–9.
Kyle RA, Remstein ED, Therneau TM, Dispenzieri A, Kurtin PJ, Hodnefield JM, et al. Clinical course and prognosis of smoldering (asymptomatic) multiple myeloma. N Engl J Med. 2007;356:2582–90.
Fonseca R, Bailey RJ, Ahmann GJ, Rajkumar SV, Hoyer JD, Lust JA, et al. Genomic abnormalities in monoclonal gammopathy of undetermined significance. Blood. 2002;100:1417–24.
Chapman MA, Lawrence MS, Keats JJ, Cibulskis K, Sougnez C, Schinzel AC, et al. Initial genome sequencing and analysis of multiple myeloma. Nature. 2011;471:467–72.
Pawlyn C, Morgan GJ. Evolutionary biology of high-risk multiple myeloma. Nat Rev Cancer. 2017;17:543–56.
Dutta AK, Hewett DR, Fink JL, Grady JP, Zannettino ACW. Cutting edge genomics reveal new insights into tumour development, disease progression and therapeutic impacts in multiple myeloma. Br J Haematol. 2017;178:196–208.
Manier S, Salem KZ, Park J, Landau DA, Getz G, Ghobrial IM. Genomic complexity of multiple myeloma and its clinical implications. Nat Rev Clin Oncol. 2017;14:100–13.
Szalat R, Munshi NC. Genomic heterogeneity in multiple myeloma. Curr Opin Genet Dev. 2015;30:56–65.
Brioli A, Melchor L, Cavo M, Morgan GJ. The impact of intra-clonal heterogeneity on the treatment of multiple myeloma. Br J Haematol. 2014;165:441–54.
Greaves M, Maley CC. Clonal evolution in cancer. Nature. 2012;481:306–13.
Morgan GJ, Walker BA, Davies FE. The genetic architecture of multiple myeloma. Nat Rev Cancer. 2012;12:335–48.
Bolli N, Avet-Loiseau H, Wedge DC, Van Loo P, Alexandrov LB, Martincorena I, et al. Heterogeneity of genomic evolution and mutational profiles in multiple myeloma. Nat Commun. 2014;5:2997.
Lohr JG, Stojanov P, Carter SL, Cruz-Gordillo P, Lawrence MS, Auclair D, et al. Widespread genetic heterogeneity in multiple myeloma: implications for targeted therapy. Cancer Cell. 2014;25:91–101.
Walker BA, Boyle EM, Wardell CP, Murison A, Begum DB, Dahir NM, et al. Mutational spectrum, copy number changes, and outcome: results of a sequencing study of patients with newly diagnosed myeloma. J Clin Oncol. 2015;33:3911–20.
Vandyke K, Zeissig MN, Hewett DR, Martin SK, Mrozik KM, Cheong CM, et al. HIF-2alpha promotes dissemination of plasma cells in multiple myeloma by regulating CXCL12/CXCR4 and CCR1. Cancer Res. 2017;77:5452–63.
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, Del Angel G, Levy-Moonshine A, et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics 2013;43:11–33. 11 10
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31:213–9.
Josephidou M, Lynch AG, Tavare S. multiSNV: a probabilistic approach for improving detection of somatic point mutations from multiple related tumour samples. Nucleic Acids Res. 2015;43:e61.
Cingolani P, Platts A, Wang le L, Coon M, Nguyen T, Wang L, et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strainw1118; iso-2; iso-3. Fly. 2012;6:80–92.
Talevich E, Shain AH, Botton T, Bastian BC. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput Biol. 2016;12:e1004873.
Deshwar AG, Vembu S, Yung CK, Jang GH, Stein L, Morris Q. PhyloWGS: reconstructing subclonal composition and evolution from whole-genome sequencing of tumors. Genome Biol. 2015;16:35.
Miller CA, McMichael J, Dang HX, Maher CA, Ding L, Ley TJ, et al. Visualizing tumor evolution with the fishplot package for R. BMC Genomics 2016;17:880.
Egan JB, Shi CX, Tembe W, Christoforides A, Kurdoglu A, Sinari S, et al. Whole-genome sequencing of multiple myeloma from diagnosis to plasma cell leukemia reveals genomic initiating events, evolution, and clonal tides. Blood. 2012;120:1060–6.
Walker BA, Wardell CP, Melchor L, Hulkki S, Potter NE, Johnson DC, et al. Intraclonal heterogeneity and distinct molecular mechanisms characterize the development of t(4;14) and t(11;14) myeloma. Blood. 2012;120:1077–86.
Weissbach S, Langer C, Puppe B, Nedeva T, Bach E, Kull M, et al. The molecular spectrum and clinical impact of DIS3 mutations in multiple myeloma. Br J Haematol. 2015;169:57–70.
Lopez-Corral L, Sarasquete ME, Bea S, Garcia-Sanz R, Mateos MV, Corchete LA, et al. SNP-based mapping arrays reveal high genomic complexity in monoclonal gammopathies, from MGUS to myeloma status. Leukemia. 2012;26:2521–9.
Mikulasova A, Wardell CP, Murison A, Boyle EM, Jackson GH, Smetana J, et al. Somatic mutation spectrum in monoclonal gammopathy of undetermined significance indicates a less complex genomic landscape compared to multiple myeloma. Haematologica. 2017;102:1617–25.
Zhao S, Choi M, Heuck C, Mane S, Barlogie B, Lifton RP, et al. Serial exome analysis of disease progression in premalignant gammopathies. Leukemia. 2014;28:1548–52.
Melchor L, Brioli A, Wardell CP, Murison A, Potter NE, Kaiser MF, et al. Single-cell genetic analysis reveals the composition of initiating clones and phylogenetic patterns of branching and parallel evolution in myeloma. Leukemia. 2014;28:1705–15.
McGranahan N, Swanton C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell. 2017;168:613–28.
Lopez-Corral L, Gutierrez NC, Vidriales MB, Mateos MV, Rasillo A, Garcia-Sanz R, et al. The progression from MGUS to smoldering myeloma and eventually to multiple myeloma involves a clonal expansion of genetically abnormal plasma cells. Clin Cancer Res. 2011;17:1692–1700.
Keats JJ, Chesi M, Egan JB, Garbitt VM, Palmer SE, Braggio E, et al. Clonal competition with alternating dominance in multiple myeloma. Blood. 2012;120:1067–76.
Magrangeas F, Avet-Loiseau H, Gouraud W, Lode L, Decaux O, Godmer P, et al. Minor clone provides a reservoir for relapse in multiple myeloma. Leukemia. 2013;27:473–81.
Kortum KM, Langer C, Monge J, Bruins L, Zhu YX, Shi CX, et al. Longitudinal analysis of 25 sequential sample-pairs using a custom multiple myeloma mutation sequencing panel (M(3)P). Ann Hematol. 2015;94:1205–11.
Weston-Bell N, Gibson J, John M, Ennis S, Pfeifer S, Cezard T, et al. Exome sequencing in tracking clonal evolution in multiple myeloma following therapy. Leukemia. 2013;27:1188–91.
Dhodapkar MV. MGUS to myeloma: a mysterious gammopathy of underexplored significance. Blood. 2016;128:2599–606.
Ghobrial IM, Detappe A, Anderson KC, Steensma DP. The bone-marrow niche in MDS and MGUS: implications for AML and MM. Nat Rev Clin Oncol. 2018;15:219–33.
Noll JE, Williams SA, Purton LE, Zannettino AC. Tug of war in the haematopoietic stem cell niche: do myeloma plasma cells compete for the HSC niche? Blood Cancer J. 2012;2:e91.
Manier S, Sacco A, Leleu X, Ghobrial IM, Roccaro AM. Bone marrow microenvironment in multiple myeloma progression. J Biomed Biotechnol. 2012;2012:157496.
Hewett DR, Vandyke K, Lawrence DM, Friend N, Noll JE, Geoghegan JM, et al. DNA barcoding reveals habitual clonal dominance of myeloma plasma cells in the bone marrow microenvironment. Neoplasia. 2017;19:972–81.
Lawson MA, McDonald MM, Kovacic N, Hua Khoo W, Terry RL, Down J, et al. Osteoclasts control reactivation of dormant myeloma cells by remodelling the endosteal niche. Nat Commun. 2015;6:8983.
Das R, Strowig T, Verma R, Koduru S, Hafemann A, Hopf S, et al. Microenvironment-dependent growth of preneoplastic and malignant plasma cells in humanized mice. Nat Med. 2016;22:1351–7.
Mitsiades CS, Mitsiades NS, Munshi NC, Richardson PG, Anderson KC. The role of the bone microenvironment in the pathophysiology and therapeutic management of multiple myeloma: interplay of growth factors, their receptors and stromal interactions. Eur J Cancer. 2006;42:1564–73.
Barlogie B, Mitchell A, van Rhee F, Epstein J, Morgan GJ, Crowley J. Curing myeloma at last: defining criteria and providing the evidence. Blood. 2014;124:3043–51.
Rasche L, Chavan SS, Stephens OW, Patel PH, Tytarenko R, Ashby C, et al. Spatial genomic heterogeneity in multiple myeloma revealed by multi-region sequencing. Nat Commun. 2017;8:268.
Bustoros M, Mouhieddine TH, Detappe A, Ghobrial IM. Established and novel prognostic biomarkers in multiple myeloma. Am Soc Clin Oncol Educ Book. 2017;37:548–60.
Acknowledgements
This study was supported by funding from the Australian Government National Health and Medical Research Council (APP1065314). A.K.D was supported by a Leukaemia Foundation of Australia National Research Program PhD Scholarship and Florey Medical Research Foundation Project Grant in Cancer Research for PhD students. We would like to thank the South Australian Cancer Research Biobank (SACRB) for assistance with sourcing patient samples; Dr. Annie Chow for curation of patient samples used in the study; Dr. Stanley Cheung for curation of clinical data of patient samples used in the study; Dr. Randall Grose, SAHMRI Australian Cancer Research Foundation (ACRF) Innovative Imaging Centre for assistance with flow cytometry cell sorting; Lisa Anderson and team from UQDI/QUT for the exome library preparation and sequencing; Jeremy Dumsday, Agilent Technologies and Jenny Peters, Integrated Sciences for the CRE kit; Prof. Maher Gandhi, UQDI, for facilitating access to the MMRF CoMMpass raw sequencing data (dbGaP phs000348, phs000748); and members of the Myeloma Research Laboratory for their support and advice.
Authors contributions
Conception and design: A.C.W.Z, J.L.F, D.R.H, G.J.M, C.G.M, L.B.T, A.K.D. Acquisition of data: A.K.D; analysis and interpretation of data: J.L.F, J.P.G, A.K.D, D.R.H, A.C.W.Z. Writing, review, and/or revision and approval of the manuscript: A.K.D, J.L.F, D.R.H, A.C.W.Z, G.J.M, C.G.M, J.P.G, L.B.T. Study supervision: J.L.F, D.R.H, A.C.W.Z.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dutta, A.K., Fink, J.L., Grady, J.P. et al. Subclonal evolution in disease progression from MGUS/SMM to multiple myeloma is characterised by clonal stability. Leukemia 33, 457–468 (2019). https://doi.org/10.1038/s41375-018-0206-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-018-0206-x
- Springer Nature Limited
This article is cited by
-
How to Identify and Manage High-Risk Smoldering Multiple Myeloma
Current Oncology Reports (2024)
-
TERRA G-quadruplex stabilization as a new therapeutic strategy for multiple myeloma
Journal of Experimental & Clinical Cancer Research (2023)
-
Telomere length and hTERT genetic variants as potential prognostic markers in multiple myeloma
Scientific Reports (2023)
-
Mesenchymal stromal cell senescence in haematological malignancies
Cancer and Metastasis Reviews (2023)
-
Importance of circulating adipocytokines in multiple myeloma: a systematic review and meta-analysis based on case-control studies
BMC Endocrine Disorders (2022)