Abstract
The mineralocorticoid receptor (MR) plays a central role in sodium homoeostasis by transducing the response to aldosterone in the distal nephron and other sodium transporting epithelia. The MR is a member of the nuclear receptor family of ligand-dependent transcription factors; it is unusual in being the receptor for two steroid hormones aldosterone and cortisol (which also binds to the closely related glucocorticoid receptor). Less well recognised is that progesterone also binds to the MR with high affinity. The conformation of the ligand-bound receptor is determined by the ligand including whether the conformation is agonist or antagonist. An agonist MR conformation then enables interactions with DNA, other MR (homodimerization) and coregulatory molecules to regulate gene expression. Insights into the structural determinants of an agonist response to ligand come from studies of the evolution of the MR. Progesterone is an agonist in the fish MR, but antagonist in the MR of terrestrial vertebrates; this switch results from the loss of a critical leucine that mediates a leucine:leucine interaction between helix 1 and helix 8 which enables the agonist response to progesterone. The insights into the intramolecular dynamics of activation suggest novel ways in which MR antagonism may be achieved beyond the current, progesterone-based antagonists in clinical use.
Similar content being viewed by others
Introduction
Hypertension is a major contributor to the global burden of disease through diverse end-organ effects. Although many factors are known to contribute to the maintenance of a normal blood pressure, the aetiology of the hypertension in most patients remains to be determined. A range of studies including those of monogenic hypertension [1] demonstrate the fundamental role of renal sodium handling in the physiology and pathophysiology of blood pressure homoeostasis. After glomerular filtration, the kidney resorbs sodium along the nephron with the bulk of sodium resorption occurring in the proximal nephron. Although only 2–5% is absorbed in the cortical collecting tubule, it is the final and therefore most critical point of modulation. It is at this point that the primary defender of plasma volume, the renin–angiotensin–aldosterone system impacts sodium resorption through the actions of the mineralocorticoid hormone, aldosterone. Aldosterone acts in the epithelial cells of the distal nephron to increase the number of open apical epithelial sodium channels enabling a transepithelial sodium flux in which the ATP-dependent basolateral pump, sodium–potassium ATPase, drives the efflux of sodium from the cells of the distal nephron [2]. Parallel to this aldosterone-induced influx of sodium is an efflux of potassium ions with potassium status also impacting adrenal aldosterone synthesis [3]. At the centre of this elegant, homoeostatic mechanism is the receptor for aldosterone, the mineralocorticoid receptor (MR): its role is pivotal. Loss of MR function, usually a haploinsufficiency resulting from an inactivating mutation in one allele of the MR [4], is seen in patients with pseudohypoaldosteronism. In a transgenic mouse with loss (knock-out) of both MR alleles, there is profound salt-wasting [5]. The MR therefore plays a central obligatory role in sodium homoeostasis and the regulation of blood pressure.
Mineralocorticoid receptor structure
The structure of the MR was elucidated by Arriza et al. [6] over 30 years ago; it is a member of the nuclear hormone receptor superfamily of ligand-activated transcription factors [3, 7]. Nuclear hormone receptors have a modular structure (Fig. 1) [8]. Central to each receptor is the DNA-binding domain (DBD), which binds to conserved response elements in regulated genes. The DBD has two anti-parallel α-helices co-ordinated by a cluster of four cysteine residues interacting with two zinc atoms to form so-called, zinc fingers [9], a motif that is highly conserved across evolution; indeed, this structure defines the nuclear receptor superfamily [8, 9]. The N-terminal domain (NTD) contains the ligand-independent activation function-1 (AF-1), which lacks a distinct structural correlate as the N-terminus is “naturally disordered” with a flexibility that allows multiple protein–protein interactions with the cellular transcriptional machinery [3, 8]. AF-1 simply requires receptor dimerisation and DNA-binding to enable juxtaposition with the transcriptional apparatus, it is in that sense, ligand-independent. The NTD has very little sequence identity or similarity with those of the other steroid hormone receptors, but is conserved between species. The ligand-binding domain (LBD) consists of 11 α-helices (designated 1–12; the region between helices 1 and 3 is unstructured in the MR) organized in an antiparallel helical sandwich [10,11,12]. The LBD binds both agonist and antagonist ligands. The MR undergoes conformational change upon aldosterone binding such that helix 12 forms a stable interaction with helices 3, 4 and 5 to create AF-2, a hydrophobic cleft on the surface of the LBD, which serves as a docking platform for transcriptional coactivators. Many coactivators bind to the AF-2 region via a conserved “NR-box” that contains one or more LxxLL (where L is leucine and x is any amino acid) motifs [12]. Co activators may bind exclusively to AF-1 or AF-2 or often, both.
A schematic of the linear MR structure is shown with activation function 1 (AF-1) and activation function 2 (AF-2) indicated. The N-terminal domain is largely unstructured, whereas the DNA-binding domain (DBD) has a published crystal structure which is represented below the linear representation. It shows two DBD homodimers interacting with the DNA at a consensus motif (sequence shown below). The DBD consists of two α-helices (green ribbons), each co-ordinated by a zinc ion (grey ball). The ligand-binding domain (LBD) tertiary structure with its 11 α-helices is shown below the linear structure; aldosterone is in the binding-pocket in the lower third of the crystal structure. Helix 12 (orange) is shown in the agonist configuration common to all of the characterised nuclear receptors when forming the AF-2 region. The representations of the crystal structures are adapted from Hudson et al. [9] and Bledsoe et al. [11], respectively. (color figure online).
Mineralocorticoid receptor signalling: a series of molecular interactions
Early models of steroid hormone action conceptualized a rigid, linear process in which once ligand-bound, a transformation occurred in the receptor enabling the receptor to move into the nucleus, then interact with DNA and regulate gene expression; however, the mechanisms are much more complex and nuanced. The ligand–receptor interaction is not rigid but dynamic such that receptor conformation is determined in part, by the ligand. Signal transduction is then determined by conformation-specific interactions with an extensive, cell-specific repertoire of coregulatory molecules. Our work has focused on four functionally significant interactions of the MR: ligand-binding, DNA-binding, inter-domain interactions, and interactions with co-regulatory molecules. This review will focus on structural and evolutionary aspects of ligand-binding as the obligatory first step in this signalling cascade in which the ligands input is transduced into a unique, tissue-specific repertoire of changes in gene expression, the aldosterone-MR transcriptome [13].
Ligand-binding to the MR
The MR is unique amongst the steroid receptors in being the physiological receptor for more than one class of steroid hormone. In addition to its role as a receptor for “mineralocorticoid” hormones, transducing the response to aldosterone or the less potent mineralocorticoid, deoxycorticosterone (DOC), the MR also acts as a receptor for the physiological glucocorticoids, cortisol in most species or corticosterone in rodents. These glucocorticoids, but not aldosterone, also act through the closely related glucocorticoid receptor (GR) [3]. Aldosterone specificity in sodium transporting epithelial cells is maintained by 11β-hydroxysteroid dehydrogenase type 2 (HSD2), which converts cortisol/corticosterone to the inactive metabolites, cortisone/11-dehydrocorticosterone. HSD2 protection of the MR also appears operative in vascular beds [3] and discrete subpopulations of hypothalamic neurons [14]. The MR, unprotected by HSD2, plays a fundamental role in macrophage [15] and cardiomyocyte function [16]; the MR also acts as a receptor for cortisol in a range of other tissues [17] including adipocytes [3] and hippocampal neurones [18]. Conversely and less well recognised, progesterone is a physiological antagonist of the MR.
Agonism versus antagonism at the MR
Progesterone is both an early precursor in adrenal steroid hormone synthesis and the ligand for the progesterone receptor (PR) as well as a physiological antagonist of the MR. Indeed, the widely used MR antagonist (MRA), spironolactone, is based on the progesterone molecule as is the MR-selective, second generation MRA, eplerenone. The steroidal progestogen, drospirenone is also a potent MRA. These antagonists are “competitive antagonists” in that they compete with the agonist for MR binding. Both agonist and antagonist ligands therefore require high affinity binding to the MR. The unliganded MR is held in an a transcriptionally inactive, high binding affinity state by the cytoplasmic heat shock-binding protein (hsp90)-cochaperone complex [19]; ligand-binding results in a conformational change in which the hsp90-complex dissociates and the MR moves to the nucleus. The distinction between agonist versus antagonist is determined not by ligand-binding or the hsp90–complex interaction, which determines binding affinity and indeed specificity [19], but by a component of the ligand-induced conformational change. Attempts to model this distinction in the MR have been compromised by the intrinsic instability of the antagonist-bound complex as reflected in experiments many years ago showing the latter to be more sensitive to trypsin digestion than the agonist-bound complex.
In 2005, three groups published the crystal structure of the MR LBD complexed with various ligands including progesterone and spironolactone [10,11,12]. There is, however, a critical caveat. To achieve stabilisation of the MR LBD for crystallization, two groups used the approach applied to obtain GR LBD crystals [20] with a cysteine in helix-5 mutated to serine [11, 12]; the crystals obtained by Fagart et al. [10] contained additional substitutions, Ser810Leu [21] and Cys910Ala. All X-ray crystal structures therefore constrain the MR LBD to a functionally active (agonist) form, which means that an antagonist conformation for the MR LBD is not represented by any of the published structures, irrespective of the bound ligand [21].
Progesterone as an MR agonist
In 2000, Geller et al. [22] described a kindred with low-renin early onset severe hypertension which was exacerbated during pregnancy. Despite features of mineralocorticoid-induced hypertension in the index case, a 15-year-old boy, his aldosterone levels were low and no alternate MR ligands were identified. He was demonstrated to be heterozygous for substitution of a leucine for a serine at position 810 in the MR LBD; they found progesterone to be an agonist in this mutant MR which explained exacerbations in pregnancy in the kindred. A subsequent study found cortisone to also be agonist providing a possible explanation for the MR activation in the boy [23], although other progesterone derivatives such as 17 hydroxyprogesterone may also have been contributing [22]. Spironolactone was also agonist, ruling it out as a therapeutic option. In an elegant series of studies Geller [22, 24] demonstrated that the leucine substitution at position 810 in helix-5 of the MR LBD enables contact with an alanine at position 773 in helix-3. They argued that the helix-3:helix-5 interaction enables ligand-dependent steroid receptor activation [24].
Insights from evolution
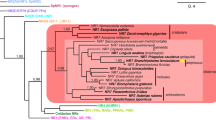
In 2006, Sturm et al. [25] cloned and characterized the MR from the rainbow trout (Oncorhynchus mykiss). When analysed in a transactivation assay, the responses to the agonist ligands, aldosterone and cortisol were remarkably similar to those for the human (h)MR, however, rather unexpectedly both progesterone and spironolactone were also agonists. We subsequently reported similar findings for the zebra fish (Danio rerio) MR [26]. In both the trout and the zebrafish MR, a serine is conserved at the equivalent position to amino acid 810 in the hMR. In a series of detailed analyses, Baker and Katsu [27] have reported progesterone to be an MR agonist in a range of teleost fish. By contrast these same authors have found the MR in amphibians and reptiles to exhibit a pattern of responses to agonist and antagonist ligands equivalent to that in the various mammalian MR that have been characterized [28]. An unexpected and currently unexplained anomaly is the avian MR in which progesterone is also an agonist [28].
The MR appeared with the GR through a gene duplication from an ancestral corticosteroid receptor >450 million years ago [29]. The MR has been remarkably conserved across evolution in that the DBD of the zebra fish (z)MR shares 97% identity at the DBD, 77% at the LBD and 33% at the N-terminus with the hMR [26]. Despite this conservation of sequence and indeed structure, the response to progesterone and related ligands differs across evolution.
In a recent study using a chimeric approach, we were able to demonstrate first that the agonist response of the zMR to progesterone and spironolactone was, perhaps not unexpectedly, mediated by the LBD [30]. Candidate regions and amino acids previously implicated in spironolactone or progesterone binding and activation were conserved between the zMR and the hMR. An exhaustive (and exhausting) series of chimeras coupled with site-directed mutagenesis enable us to identify a threonine at position 870 in the hMR which when changed to a leucine, as found at the equivalent position in the zMR (amino acid 856), enabled spironolactone and progesterone to act as agonists whilst reciprocal mutagenesis in the zMR converted the responses to antagonist. In all cases, the equivalent agonist responses for aldosterone and cortisol were maintained.
As in all previous studies, the transactivation assay used a mammalian cell line at 37 °C, which is of course not physiological for a zebrafish; we were able to confirm the responses in a zebrafish derived cell line at 28 °C [30]. The critical residues lie in helix-8, a region of the LBD not previously associated with ligand-binding or activation; the leucine is conserved across all of fish MR including the lungfish although the latter MR is yet to be functionally characterized. The evolutionary emergence or otherwise of aldosterone in the lungfish remains controversial.
A threonine is found at the equivalent position to hMR870 in all the terrestrial vertebrate MR that have been sequenced except for the rodent MR where there is a serine at this position [30]. Although both threonine and serine are potentially subject to phosphorylation, substitution of amino acids which mimic phosphorylation does not alter the response to ligand nor indeed does an alanine substitution arguing very clearly that it is the presence of a leucine at this position in helix-8 of the MR that is critical for the agonist response to progesterone.
Molecular dynamic simulation based on the published hMR LBD structures predicted a series of subtle shifts in the alignment of the helices when the threonine at position 870 in the hMR is substituted by a leucine [30]. The modelling also predicted a hydrophobic network which included an interaction of the leucine at position 856 in the zMR with a highly conserved leucine in helix-1. Mutation of this leucine in helix-1 to an alanine did not alter the response to aldosterone but rendered an antagonist response to progesterone/spironolactone. This argues that as with stabilization of the helix-3:helix-5 interaction, a stable helix-1:helix-8 interaction is critical if progesterone is to act as an agonist. An intriguing finding from the molecular dynamic simulation is an impact of the leucine:leucine interaction between helices 1 and 8, on the position of helix-5 and consequently rotation of helix-7. It would seem that the helix-1:helix-8 stabilizing interaction is upstream of the helix3:helix-5 interaction characterized by Geller et al. [22] with a subsequent impact on helix-7. This series of subtle shifts through the full LBD [30] likely determines the ultimate stability of the AF-2 region to which helices 3 and 5 contribute. The stability of coactivator binding to the AF-2 region is critical to an agonist response.
Implications from evolution
The evolutionary switch from agonism to antagonism for progesterone at the MR corresponds to both the appearance of terrestrial vertebrates and aldosterone synthesis. It would seem that the greatly increased need to conserve sodium in a terrestrial environment is met by the appropriation of the MR by aldosterone to the exclusion of progesterone. These findings raise a number of questions about the biology of the MR in fish where it would that seem that rather than salt and water balance, the MR has roles in ontogenesis [31]. The question of whether the MR retains these roles in higher vertebrates, and the role of progesterone as a physiologic antagonist, remains to be determined. An important caveat, that appears sometimes to have been overlooked, is that the use of spironolactone as an antagonist in studies of fish is clearly incorrect and potentially very misleading.
The insights into the intramolecular dynamics of the MR during activation provided in nature by both accidents [22] and evolution [30] suggest novel ways in which antagonism may be achieved beyond the current, progesterone-based antagonists in clinical use.
The marked shifts in both helix-5, helix-7 and the unstructured loops between would be predicted from their location to impact extrinsic interactions of the MR LBD. Such interactions potentially include the NTD, coactivators or perhaps a dimer interface. The existence of LBD dimerization for the AR, PR, GR and MR has remained controversial with crystal structures variously described as reflecting packing artifacts or dimer interfaces. Nadal et al. [32] have identified a dimer interface in the AR LBD that involves helices 5 and 7 suggesting that this question should be revisited in the MR.
Interactions of the MR LBD enabled by ligand-binding
The ligand-induced conformational changes in the LBD enable a series of interactions crucial to mediating the cellular responses which result from activation of the MR. These have been previously reviewed [33] and include DNA-binding, inter-domain interactions and the interaction with co-regulatory molecules.
DNA-binding by the ligand-activated MR is obligatory for sodium homoeostasis. When we created a transgenic mouse homozygous for an MR DBD mutation in which a serine has been substituted for a critical cysteine in the first zinc-finger to preclude DNA-binding, the mice exhibited a phenotype virtually identical to that of the MR knock-out mice, i.e. profound salt-wasting with neonatal lethality [34]. The DBD binds to hormone response elements (HRE) in the genome; the classic consensus steroid receptor HRE is an inverted palindrome (Fig. 1).
Although the genome wide distribution of the HRE for the other steroid receptors, the cistrome, has been characterized, studies with the MR have proven technically challenging with only two studies reported to date, one using a human renal cell line stably expressing a tagged MR [35] and the other examined rat hippocampus using both MR- and GR-specific antibodies [36]. The findings were somewhat conflicting between the two studies so the precise definition of the MR cistrome remains a “work in progress”.
Although the three main functional domains in the steroid receptors have been viewed as “modular” and independent; it is now clear that an interaction between the NTD and the C-terminus/LBD, the N/C-interaction, can play a key role in defining the nature of the response. This N/C-interaction is best characterised in the androgen receptor where it has been used in the development of selective androgen receptor modulators. It is also observed for the PR and oestrogen receptor α, although curiously, not for the closely related GR [37]. We have characterized a ligand-specific N/C-interaction in the MR [37]. This interaction occurs in the presence of aldosterone. However, this aldosterone-mediated interaction is antagonised by the MR agonists cortisol and DOC together with the MRA [37]. The zebrafish (z)MR also exhibits an N/C-interaction [26], which argues for its functional significance although in the zMR, cortisol and DOC are also activate the N/C-interaction.
Coregulators interact with the activation functions, AF-1 and AF-2 (Fig. 1) and the coregulatory complex that links the receptor to the transcriptional apparatus. The importance of AF-1 and AF-2 coregulator interactions varies with the cell type, target-gene and ligand in the case of AF-2. In the MR, replacement of a highly conserved glutamic acid with the neutral amino acid alanine in helix 12 eliminates AF-2 function in vitro, whilst preserving ligand-binding and AF-1-mediated transactivation. We have confirmed that this mutation, hMR E962A, completely eliminates interactions with LxxLL motif-containing co-activators (SRC-1 and PGC1α), but has a limited impact on ligand-binding and the N/C-interaction [38].
Aside from the “generic” steroid receptor coactivators, the p160 family (SRC-1, 2 and 3) and PGC-1α, only a small number of coactivators had been described for the MR [39]; none showed tissue- or ligand-specificity. We took three complementary approaches to identify unique ligand-specific (aldosterone versus cortisol) MR interacting proteins: yeast 2-hybrid (Y-2-H) screens, M13 phage display and T7-phage display [39]. Y-2-H screens identified proteins that interact with the MR LBD to coactivate MR-mediated transactivation in the presence of aldosterone but not cortisol, including tesmin which contains LxxLL motifs that directly interact with the AF-2 region of the MR LBD. In parallel studies using recombinant full-length MR and M13 phage display, we identified novel, LxxLL motif-constrained, 19-mer peptides that interact with the MR in a ligand-dependent manner, including a conserved motif which predicted an interacting protein Gemin4 [40]. With the T7-phage libraries we identified interacting proteins that differ between cell type, ligand and promoter in their coregulation of the MR response [41].
Conclusions
The MR uniquely binds aldosterone and as such plays a central, fundamental role in sodium homoeostasis in terrestrial vertebrates including of course man. The appropriation by aldosterone through the evolutionary transition to terrestrial life is associated with a unique single amino acid switch that precludes progesterone acting as an agonist at the MR. These findings together with a series of other studies exploring interactions of the MR demonstrate the importance of the ligand–MR interaction in defining the confirmation of the receptor and the resulting interactions that define the nature of the signal induced. These nuances which define both ligand and cell specific differences in MR action provide targets for novel MR ligands with modulated rather than binary outputs. Modulating ligands are in clinical use (ER) or development (AR, GR, PR) for the other steroid receptors; the interacts identified for the MR provide opportunities for the development of similar therapeutics targeting the MR.
References
Lifton RP, Gharavi AG, Geller DS. Molecular mechanisms of human hypertension. Cell. 2001;104:545–66.
Hanukoglu I, Hanukoglu A. Epithelial sodium channel (ENaC) family: phylogeny, structure-function, tissue distribution, and associated inherited diseases. Gene. 2016;579:95–132.
Fuller PJ, Young MJ. Mechanisms of mineralocorticoid action. Hypertension. 2005;46:1227–35.
Zennaro MC, Fernandes-Rosa F. 30 years of the mineralocorticoid receptor: mineralocorticoid receptor mutations. J Endocrinol. 2017;234:T93–106.
Berger S, Bleich M, Schmid W, Cole TJ, Peters J, Watanabe H, et al. Mineralocorticoid receptor knockout mice: pathophysiology of Na+ metabolism. Proc Natl Acad Sci USA. 1998;95:9424–9.
Arriza JL, Weinberger C, Cerelli G, Glaser TM, Handelin BL, Housman DE, et al. Cloning of human mineralocorticoid receptor complementary DNA: structural and functional kinship with the glucocorticoid receptor. Science. 1987;237:268–75.
Viengchareun S, Le Menuet D, Martinerie L, Munier M, Pascual-Le Tallec L, et al. The mineralocorticoid receptor: insights into its molecular and (patho)physiological biology. Nucl Recept Signal. 2007;5:e012.
Huang P, Chandra V, Rastinejad F. Structural overview of the nuclear receptor superfamily: insights into physiology and therapeutics. Annu Rev Physiol. 2010;72:247–72.
Hudson WH, Youn C, Ortlund EA. Crystal structure of the mineralocorticoid receptor DNA binding domain in complex with DNA. PLoS ONE. 2014;9:e107000.
Fagart J, Huyet J, Pinon GM, Rochel M, Mayer C, Rafestin-Oblin ME. Crystal structure of a mutant mineralocorticoid receptor responsible for hypertension. Nat Struct Mol Biol. 2005;12:554–5.
Bledsoe RK, Madauss KP, Holt JA, Apolito CJ, Lambert MH, Pearce KH, et al. A ligand-mediated hydrogen bond network required for the activation of the mineralocorticoid receptor. J Biol Chem. 2005;280:31283–93.
Li Y, Suino K, Daugherty J, Xu HE. Structural and biochemical mechanisms for the specificity of hormone binding and coactivator assembly by mineralocorticoid receptor. Mol Cell. 2005;19:367–80.
Poulsen SB, Limbutara K, Fenton RA, Pisitkun T, Christensen BM. RNA sequencing of kidney distal tubule cells reveals multiple mediators of chronic aldosterone action. Physiol Genomics. 2018;50:343–54.
Geerling JC, Kawata M. Aldosterone-sensitive neurons in the rat central nervous system. J Com Neurol. 2005;494:515–27.
Rickard AJ, Morgan J, Tesch G, Funder JW, Fuller PJ, Young MJ. Deletion of mineralocorticoid receptors from macrophages protects against DOC/salt-induced cardiac fibrosis and increased blood pressure. Hypertension. 2009;54:537–43.
Rickard AJ, Morgan J, Bienvenu LA, Fletcher EK, Cranston GA, Shen JZ, et al. Cardiomyocyte mineralocorticoid receptors are essential for deoxycorticosterone/salt-mediated inflammation and cardiac fibrosis. Hypertension. 2012;60:1443–50.
Jaisser F, Farman N. Emerging roles of the mineralocorticoid receptor in pathology: toward new paradigms in clinical pharmacology. Pharmacol Rev. 2016;68:49–75.
Joëls M, de Kloet ER. 30 years of the mineralocorticoid receptor: the brain mineralocorticoid receptor: a saga in three episodes. J Endocrinol. 2017;234:T49–66.
Cluning C, Ward BK, Rea SL, Arulpragasam A, Fuller PJ, Ratajczak T. The helix 1-3 loop in the glucocorticoid receptor LBD is a regulatory element for FKBP cochaperones. Mol Endocrinol. 2013;27:1020–35.
Bledsoe RK, Montana VG, Stanley TB, Delves CJ, Apolito CJ, McKee DD, et al. Crystal structure of the glucocorticoid receptor ligand binding domain reveals a novel mode of receptor dimerization and coactivator recognition. Cell. 2002;110:93–105.
Huyet J, Pinon GM, Fay MR, Rafestin-Oblin M-E, Fagart J. Structural determinants of ligand binding to the mineralocorticoid receptor. Mol Cell Endocrinol. 2012;350:187–95.
Geller DS, Farhi A, Pinkerton N, Fradley M, Moritz M, Spitzer A, et al. Activating mineralocorticoid receptor mutation in hypertension exacerbated by pregnancy. Science. 2000;289:119–23.
Rafestin-Oblin ME, Souque A, Bocchi B, Pinon G, Fagart J, Vandewalle A. The severe form of hypertension caused by the activating S810L mutation in the mineralocorticoid receptor is cortisone related. Endocrinology. 2003;144:528–33.
Zhang J, Simisky J, Tsai FT, Geller DS. A critical role of helix 3-helix 5 interaction in steroid hormone receptor function. Proc Natl Acad Sci USA. 2005;102:2707–12.
Sturm A, Bury N, Dengreville L, Fagart J, Flouriot G, Rafestin-Oblin M-E, et al. 11-Deoxycorticosterone is a potent agonist of the rainbow trout (Oncorhynchus mykiss) mineralocorticoid receptor. Endocrinology. 2005;146:47–55.
Pippal JP, Cheung CMI, Yao YZ, Brennan FE, Fuller PJ. Characterization of the zebrafish (Danio rerio) mineralocorticoid receptor. Mol Cell Endocrinol. 2011;332:58–66.
Baker ME, Katsu K. 30 years of the mineralocorticoid receptor: evolution of the mineralocorticoid receptor: sequence, structure and function. J Endocrinol. 2017;234:T1–16.
Katsu Y, Oka K, Baker ME. Evolution of human, chicken, alligator, frog, and zebrafish mineralocorticoid receptors: allosteric influence on steroid specificity. Sci Signal. 2018;11:eaao1520.
Bridgham J, Carroll S, Thornton J. Evolution of hormone-receptor complexity by molecular exploitation. Science. 2006;312:97–101.
Fuller PJ, Yao YZ, Jin R, He S, Martín-Fernández B, Young MJ, et al. Molecular evolution of the switch for progesterone and spironolactone from mineralocorticoid receptor agonist to antagonist. Proc Natl Acad Sci USA. 2019;116:18578–83.
Kiilerich P, Geffroy B, Valotaire C, Prunet P. Endogenous regulation of 11-deoxycorticosterone (DOC) and corticosteroid receptors (CRs) during rainbow trout early development and the effects of corticosteroids on hatching. Gen Comp Endocrinol. 2018;265:22–30.
Nadal M, Prekovic S, Gallastegui N, Helsen C, Abella M, Zielinska K, et al. Structure of the homodimeric androgen receptor ligand-binding domain. Nat Commun. 2017;8:14388.
Yang J, Fuller PJ. Interactions of the mineralocorticoid receptor-within and without. Mol Cell Endocrinol. 2012;350:196–205.
Cole TJ, Terella L, Morgan J, Alexiadis M, Yao YZ, Enriori P, et al. (2015) Aldosterone-mediated renal sodium transport requires intact mineralocorticoid receptor DNA-binding in the mouse. Endocrinology. 2015;156:2958–68.
Le Billan F, Khan JA, Lamribet K, Viengchareun S, Bouligand J, Fagart J, et al. Cistrome of the aldosterone-activated mineralocorticoid receptor in human renal cells. FASEB J. 2015;29:3977–89.
van Weert LTCM, Buurstede JC, Mahfouz A, Braakhuis PSM, Polman JAE, Sips HCM, et al. NeuroD factors discriminate mineralocorticoid from glucocorticoid receptor DNA binding in the male rat brain. Endocrinology. 2017;158:1511–22.
Pippal JB, Yao Y, Rogerson FM, Fuller PJ. Structural and functional characterisation of the interdomain interaction in the mineralocorticoid receptor. Mol Endocrinol. 2009;23:1360–70.
Rogerson FM, Yao YZ, Young MJ, Fuller PJ. Identification and characterization of a ligand-selective mineralocorticoid receptor coactivator. FASEB J. 2014;28:4200–10.
Fuller PJ, Yang J, Young MJ. 30 years of the mineralocorticoid receptor: coregulators as mediators of mineralocorticoid receptor signalling diversity. J Endocrinol. 2017;234:T23–34.
Yang J, Fuller PJ, Morgan J, Shibata H, Clyne CD, Young MJ. GEMIN4 functions as a coregulator of the mineralocorticoid receptor. J Mol Endocrinol. 2015;54:149–60.
Yang J, Fuller PJ, Morgan J, Shibata H, McDonnell DP, Clyne CD, et al. Use of phage display to identify novel mineralocorticoid receptor-interacting proteins. Mol Endocrinol. 2014;28:1571–84.
Acknowledgements
The authors wish to thank Sue Panckridge for preparation of the figure. This work was supported by grants-in-aid from the National Health & Medical Research Council of Australia through Project Grants (#1058336 and #1143840) and a Senior Principal Research Fellowship to PJF (#1002559). The Hudson Institute is supported by the Victorian Government’s Operational Infrastructure Scheme.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fuller, P.J., Yao, YZ., Yang, J. et al. Structural determinants of activation of the mineralocorticoid receptor: an evolutionary perspective. J Hum Hypertens 35, 110–116 (2021). https://doi.org/10.1038/s41371-020-0360-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41371-020-0360-2
- Springer Nature Limited