Abstract
Obesity and substantially increased risk of metabolic diseases have become a global epidemic. microRNAs have attracted a great deal of attention as a potential therapeutic target for obesity. MiR-143 has been known to specifically promote adipocyte differentiation by downregulating extracellular signal-regulated kinase 5. Our latest study found that miR-143 knockout is against diet-induced obesity by promoting brown adipose tissue thermogenesis and inhibiting white adipose tissue adipogenesis. Moreover, LPS- or IL-6-induced inhibition of miR-143 expression in brown adipocytes promotes thermogenesis by targeting adenylate cyclase 9. In this review, we will summarize the expression and functions of miR-143 in different tissues, the influence of obesity on miR-143 in various tissues, the important role of adipose-derived miR-143 in the development of obesity, the role of miR-143 in immune cells and thermoregulation and discuss the potential significance and application prospects of miR-143 in obesity management.
Similar content being viewed by others
Introduction
Tight and flexible control of energy homeostasis is fundamental for organism health and its dysregulation contributes to obesity, a worldwide epidemic in our current society [1]. Obesity is an energy imbalance between calories consumed and calories expended, and long-term excessive energy intake leads to metabolic disorders (especially glucose and lipid metabolism). In severe situations, obesity leads to a substantially increased risk of metabolic diseases (for example type 2 diabetes mellitus [2] and hyperlipidemia [3]), cardiovascular diseases [4], and some types of cancer (for example breast [5] and lung [6]). Since the amount or activity of brown/beige fat inversely correlates with body mass index in adult humans, brown/beige fat is believed to help reduce adiposity [7,8,9]. The epidemic of obesity and diabetes has greatly increased the interest in this metabolically active type of fat. In the past two decades, microRNAs (miRNAs) have attracted extensive attention due to being highly conserved across species and also being involved in many metabolic regulation activities by silencing target genes. Currently, miRNAs, particularly in brown/beige fat due to their regulatory functions in the initiation and progression of obesity, have gained extensive attention in exploring effective biomarkers for early and fast diagnosis and novel treatment targets for obesity [10, 11]. As a member of the miR-143/145 cluster, the gene of miR-143 is highly conserved across mammalian species. miR-143 displays a specific function in regulating metabolic activity, such as adipocyte differentiation [12,13,14,15,16,17], insulin resistance [18], glycolysis [19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34], smooth muscle cell proliferation [35], and intestinal epithelial regeneration [36]. The purpose of this review is to describe our current understanding of how miR-143 acts in the adipose tissue, liver, and body circulation to affect obesity. Firstly, we will summarize recent advances in understanding the expression pattern of miR-143 in various organs. Secondly, we will discuss expression changes of miR-143 that occur in the adipose tissue, liver, and blood circulation in the setting of obesity and highlight how these changes contribute to metabolic dysfunction. Thirdly, we will discuss the relationship between miR-143 and immune disorders and thermoregulation. Finally, we will evaluate the possibility of miR-143 as a biomarker of obesity and the potential therapeutic implications of targeting the miR-143 to treat obesity and associated diseases.
The expression pattern of miR-143 in various organs
In humans, miR-143 is located at sites 149428918–149429023 (in intron) on chromosome 5, and miR-143 traverses a canonical biogenesis pathway involving compartmentalized processing by two RNase III enzymes [37]. In the nucleus, primary miR-143 transcripts bearing inverted repeats are cleaved by Drosha to release pre-miR-143 hairpins. In the cytoplasm, they are cleaved by Dicer to yield miR-143-3p/miR-143-5p duplexes, which are then loaded onto Argonaute effector proteins [38]. Following the removal of miR-143-5p species, the single-stranded miR-143-3p-argonaute complex, in association with GW182/TNRC6 cofactors, seeks regulatory targets. We sorted out the sequences of miR-143 in different species from miRBase and found that the first 20 bases of miR-143 are conserved not only in mammals but also in non-mammals (for example Gallus gallus, Danio rerio, and Xenopus laevis) (Table 1).
As early as 2002, it was first reported that miR-143 was expressed in the mouse spleen, heart, cortex, and midbrain [39]. Following reports found that miR-143 is wildly expressed in different tissues [18, 36, 40,41,42], including the liver [18, 40], small intestine [42], and colon [36] where miR-143 expression has not been detected before [39]. In humans, miR-143 shows a similar expression pattern to that in murine [12]. But in pigs, miR-143 exhibits different expression patterns [43]. Expression patterns in different species and organs are summarized in Table 2. The expression of miR-143 still needs to be refined since inconsistent results are given in different reports. Although miR-143 has been proven to be wildly expressed and regulate a large number of metabolic activities, conditional overexpression or tissue-specific knockout of miR-143 or miR-143/145 did not overt developmental defects of tissue and organ under normal diet (ND) or normal conditions [18, 36, 40, 44].
MiR-143 regulates the adipogenesis
In obesity, the adipose tissue undergoes dynamic remodeling including an increase in size (hypertrophy) and the number of adipocytes (hyperplasia) [45, 46]. miR-143 may play an important regulatory role in this process. The miR-143 in mature adipocytes is significantly higher than that in stromal vascular fraction (SVF) preadipocytes [14], and the expression of miR-143 is increased during the differentiation of human white preadipocytes [12], 3T3-L1 cells [13,14,15,16], and adipose tissue-derived stromal cells (ADSCs) [17]. In addition to regulating adipocyte differentiation, miR-143 has also been reported to regulate milk fat synthesis in bovine mammary epithelial cells [47].
It is widely acknowledged that the MAPK signaling pathway plays a pivotal role in many essential cellular processes, including cell proliferation and differentiation [48]. Mitogen-activated protein 2 kinase kinase 5 (MAP2K5 or MEK5) and extracellular signal-regulated kinase 5 (ERK5) are important target genes for miR-143 to regulate adipocyte differentiation [12, 17]. The adipogenic differentiation process involves several successive steps, and the MEK5-ERK5 cascade regulated by miR-143 plays an important role in each step. During the clonal expansion stage, the overexpression of miR-143 inhibits the adipogenic differentiation by targeting MAPK signaling pathway [17]. In the growth arrest stage or terminal differentiation stage, the overexpression of miR-143 inactivates MEK5-ERK5 and promotes cell differentiation via the phosphorylation of peroxisome proliferator-activated receptor-gamma (PPARγ) [49, 50]. PPARγ, a member of the nuclear receptor superfamily, is a master regulator in lipid metabolism by controlling networks of gene expression for lipid accumulation [51], lipolysis [52, 53], and browning of white adipocyte [54]. It is phosphorylated within the AF1 region by MAPKs, inhibiting ligand binding and altering cofactor recruitment which represses its transcriptional activity [51, 55]. A recent study pointed out that by binding to nerve growth factor-induced B alpha (Nur77), PPARγ promotes the ubiquitination and degradation of Nur77 mediated by the ubiquitin ligase trim13, which in turn leads to the loss of Nur77’s transcriptional inhibition of CD36 and FABP4, thereby promoting fatty acid uptake and cell proliferation [56]. In addition, our newly published research showed that miR-143 knockout (KO) inhibited the differentiation of primary white adipocytes, but did not affect the differentiation of primary brown adipocytes [57].
Although the target genes of miR-143 and the transcription factors regulated by these target genes have such a great influence, a large number of reports have focused on the regulation of miR-143 on adipogenesis of white adipocytes and rarely on other aspects of lipid metabolism. The studies of the functions of miR-143 in lipid metabolism, especially lipolysis and fatty acid transport, need to be supplemented and improved, and the different roles of miR-143 in the differentiation of brown and white adipocytes need further study.
MiR-143 regulates the glycolysis and gluconeogenesis
Glucose and fatty acids are predominant sources of energy for the human and animal body. In addition to adipogenesis, numerous studies have shown that miR-143 targets hexokinase 2 (HK2) to inhibit glycolysis [19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34]. MiR-143 is regarded as a novel type 2 diabetes regulator [18, 26, 58] that regulates insulin resistance through the oxysterol-binding protein-related protein 8 (ORP8)-AKT signaling pathway, and overexpression of miR-143 in the liver impairs glucose metabolism under physiological conditions [18]. Interestingly, ORP8 also inhibits sustained phosphorylation of glycogen synthase kinase 3α/β (GSK3α/β) via insulin-stimulated AKT activation [18]. Recently, our research showed that miR-143KO significantly inhibited gluconeogenesis in the liver of mice induced by high-fat diet (HFD) [59]. It was found that miR-143 targets mitogen-activated protein kinase 11 (MAPK11) [59], thereby inhibits gluconeogenesis by repressing the activity of phosphoenolpyruvate carboxykinase 1 (PCK1) and glucose-6-phosphatase catalytic (G6PC) [60]. Changes in glucose metabolism of mice with obesity after miR-143KO require a deeper assessment, including glycolysis, glycogen synthesis and glycogenolysis. Specific liver miR-143KO mice are also an important model for assessing the contribution of miR-143-regulated hepatic glycolysis and gluconeogenesis to obesity.
MiR-143 is involved in the progression of obesity
Obesity leads to dysregulation of miR-143 expression
Since the first demonstration of the link between obesity and energy metabolism of a miRNA in drosophila melanogaster [61], numerous studies have been performed in this field, including miR-143 (Fig. 1). Intriguingly, the effect of obesity on the expression of miR-143 in WAT remains controversial. A study suggests that miR-143 increased adipogenesis in adipocytes by affecting several key lipid metabolism genes, but the expression of miR-143 in isolated mature adipocytes from ob/ob mice is downregulated compared with adipocytes from wild-type (WT) mice [14]. Furthermore, the expression level of miR-143 in WAT is significantly reduced in both the diet-induced mice [18, 62] and the db/db mice [18]. Similarly, miR-143 is lower in the subcutaneous white adipose tissue (scWAT) of subjects with obesity and insulin-resistant than in scWAT of subjects with obesity and insulin-sensitive, and both subjects with obesity and resistant and obesity and sensitive were lower than lean subjects [63]. However, other studies have reported that the miR-143 is highly expressed in WAT of HFD-induced mice compared with the mice fed ND [64] and it was positively related to the body weight of mice or adipocyte volume in swine [64,65,66,67]. Meanwhile, our latest research found that the expression of miR-143 first rose and then decreased in WAT of HFD-induced mice [57]. This reverse pattern may be similar to the role of miR-143 in adipocyte differentiation [17]. In the early stage of HFD-induced, miR-143 increased in WAT, which promoted adipocytes expansion and stored energy. In the middle and late stages of HFD-induced, excessive obesity may inhibit the expression of miR-143 in WAT through feedback regulation, and reduced adipogenesis. This view is supported by the increased expression of adipogenic genes during adipogenesis whose expression is decreased in obesity and diabetes mellitus [68]. More importantly, we also found that miR-143 in brown adipose tissue (BAT) was significantly reduced in the process of HFD-induced obesity in mice, and the reduced miR-143 could resist obesity by promoting thermogenesis [57].
The upregulation of miR-143 in the liver of patients with obesity is contrary to the expression of miR-143 in BAT [18, 62]. The liver is the central metabolic organ that regulates glucose and lipid metabolism [69]. The upregulated miR-143 specifically inhibits the insulin AKT pathway by targeting the ORP8, thereby inhibiting AKT phosphorylation, and contributing to the development of obesity-induced insulin resistance [18]. In addition to the liver, miR-143 is also upregulated in the heart, muscle and pancreas of mice with obesity [18, 62]. The upregulation of miR-143 in the heart may be related to impaired exercise ability, as studies have reported that exercise training decreased the expression of miR-143 in the heart and serum [70, 71]. Previous studies have reported that miR-143KO promotes both proliferation and chemotaxis and inhibits differentiation of vascular smooth muscle cells (VSMCs), and the downregulation of miR-143 expression is associated with increased hemodynamic stress in the aorta of mice [35, 41, 44]. So, the upregulated of miR-143 in the heart and muscle induced by obesity may be to inhibit the coarctation and altered vessel histology, such as that occurring with atherosclerosis. Moreover, the upregulated expression of miR-143 in the pancreas does not alter the glucose-stimulated insulin secretion and morphology of pancreatic β-cell [18], and the function of miR-143 in the pancreas needs further study. To sum up, miR-143 is tissue-specifically expressed with multiple functions.
Circulating miR-143 changes in obesity and its related disorders
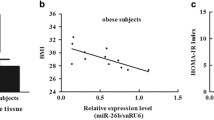
Evidence is accumulating that circulating miRNAs act as a new class of endocrine factors [72, 73]. Circulating miRNAs, which are released by many types of tissues including the adipose tissues, are valuable biomarkers yielding important insights into the pathogenesis of obesity. It is found that the level of circulating miR-143 is closely related to the occurrence of obesity. However, there are still great doubts about the expression pattern of circulating miR-143 in obesity and its related disorders (Table 3). On the one hand, circulating miR-143 has been shown to significantly increase morbidly adolescents with obesity and was positively correlated with body mass index (BMI), waist-to-hip ratio (WHR), diabetic control, and lipid profile parameters [74]. It was also significantly elevated in metabolic syndrome patients (MetS, defined according to standards generated by the joint committee for developing Chinese guidelines on the prevention and treatment of dyslipidemia in adults) compared with controls both in serum and urine samples [62], and was accompanied by an increasing trend in the homeostasis model assessment of insulin resistance (HOMA-IR), high-density lipoprotein-cholesterol (HDL-c) ratio and percentage of body fat [62]. miR-143 was also elevated in PBMCs of HFD/streptozotocin-treated type 2 diabetes mellitus rats [75]. On the other hand, in other studies, circulating miR-143 in children with obesity was significantly lower compared to those of controls, and miR-143 was correlated negatively with leptin, HOMA-IR, triglycerides (TG), BMI, and waist circumference (WC) [76]. In another report, circulating miR-143 levels were significantly lower in subjects with morbidly obesity and obesity than the normal or overweight subjects, but it did not yield any correlation between miR-143 levels and BMI and WHR [77]. Therefore, those conflicting data hints that the effect of obesity and its complications on circulating miR-143 needs to be verified by more studies.
More importantly, antagomiR-143 treated (downregulation of circulating miR-143) mice with obesity showed significant amelioration in insulin tolerance test, glucose tolerance test and the size of adipocytes in scWAT and epididymal white adipose tissue (eWAT), compared with control, but the body weight and the weight of WAT did not change [62], which may be due to the limited inhibitory effect of antagomiR-143 in the tissue. Extracellular vesicles (EVs), a type of nano-scale vesicles secreted by cells, mediate intercellular communications by transferring biological molecules, such as mRNAs, non-coding RNAs, lipids, and proteins [78,79,80,81]. miRNAs of EVs from the adipose tissue might act as novel forms of adipokine to facilitate diverse intercellular communication [82, 83]. Obesity and obesity-associated comorbidities may be due to the adipose tissue and liver cross-talk dysregulation mediated by EVs [84]. For example, EV-associated miRNAs from BAT are able to regulate gene expression in the liver [85]. Lean mice exhibit glucose intolerance and insulin resistance after the injection of adipose tissue-derived EVs from mice with obesity [86]. Researchers have found that BAT-EVs preferentially accumulate in the liver after intravenous injection [87]. Furthermore, BAT-EVs target liver metabolism more specifically and efficiently than WAT-EVs [85]. Thus, reducing the expression of miR-143 in adipose tissue with obesity, especially in BAT, may be a more promising target for the treatment of obesity with miR-143.
MiR-143KO is against diet-induced obesity by promoting BAT thermogenesis and inhibiting WAT adipogenesis
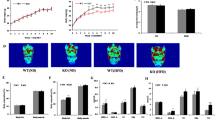
For a long time, the primary function of miR-143 was thought to promote adipogenesis of white adipocytes [12, 14, 15, 17, 64, 88, 89]. However, most of the findings are cross-sectional in nature, which do not show the impact of miR-143 on obesity. Our latest research fills the gap in the role of miR-143 in the development of obesity [57]. We construct global miR-143KO mice by using CRISPR/Cas9 technology. Although miR-143KO showed neither phenotypic abnormalities nor energy imbalance, and changes in BAT thermogenesis in mice fed an ND under normal conditions. As cold exposure stimulates stronger body temperature in miR-143KO mice [59], we speculate that miR-143 in adipose tissue acts as a stress miRNA upon metabolic challenge just like miR-21 [90]. Then, we evaluated the role of miR-143 in an HFD-induced obesity model. Surprisingly, the increase in body weight, fat mass, rectal temperature and energy expenditure of KO mice was significantly decreased during HFD feeding compared with WT mice. In addition, improvements in insulin sensitivity and glucose tolerance were also observed in miR-143KO mice. Activating BAT thermogenesis is an important strategy of against obesity, which is mediated by Uncoupling protein 1 (UCP1) protein located in mitochondria [91, 92]. We found that AC9 is a new target of miR-143 in brown adipocytes, and miR-143 promotes the expression of UCP1 by inhibiting adenylate cyclase 9 (AC9) [93], and improves the number and function of mitochondria in BAT, while the mechanism needs further study [59]. AC9 is a physiologically and clinically relevant effector of β2-adrenergic receptor (AR) signaling in particular [94, 95], and it also regulates cAMP production [95, 96]. More recent research found that BAT thermogenesis occurs through β2-AR signaling in humans [97], and AC9 inactivation increases weight gain and adipose tissue volume in mice with the treatment of an atherogenic diet [98]. Whether AC9 is sufficient and/or necessary for UCP1 induction thermogenesis needs more research.
In addition, our study confirmed that miR-143KO inhibit the differentiation of white adipocytes and fat synthesis and promote lipolysis in WAT [59]. The inhibition of miR-143KO on fat synthesis may be through the inhibition of fatty acid synthase protein expression through AMPK signaling pathway [99,100,101]. And miR-143KO significantly inhibited the differentiation of white adipocytes by promoting MEK5-PPARγ signaling pathway, which further verified previous studies [17]. The target gene of miR-143 regulating fat synthesis and lipolysis has not been found yet, which needs further research. At the same time, adipose tissue-specific knockout mice are the focus of the next study on the function of miR-143.
Obesity affects the expression of miR-143 in adipocytes through immune cytokines
The immune system is a critical regulator of metabolic homeostasis. Adipose tissue is heterogeneous, and the function of adipose tissue is dynamically regulated by communication between adipocytes and other cell types, like macrophage [92, 102]. The enlargement of the adipose tissue during obesity is accompanied by the development of chronic low-grade inflammation of the adipose tissue, which includes infiltration of macrophages into the adipose tissue and increased levels of cytokines [103, 104]. The miR-143 expression in the BAT of mice (a population including ND-fed and HFD-fed mice) exhibited a statistically significant inverse relationship with tumor necrosis factor α (TNFα) and interleukin 6 (IL-6) expression in the BAT [93]. In adipocytes, miR-143 may be involved in the response of adipocytes to obesity-induced macrophage infiltration and cytokines secretion, as treatment of TNF-α [14] and LPS or IL-6 [93] to differentiated 3T3-L1 adipocytes or primary brown adipocytes reduced the expression of miR-143. Further details of the effect of TNF-α treatment on the decline of miR-143 in white adipocytes have not been clearly studied. IL-6 is a well-known cytokine activator of thermogenesis [105, 106]. More importantly, IL-6 treatment reduced the expression of miR-143 and increased the expression of Ucp1 mRNA in brown adipocytes, while miR-143KO also enhanced the levels of IL-6 [93]. The above results indicate that there is a mutual inhibition between miR-143 and IL-6 in brown adipocytes.
Obesity-associated pro-inflammatory changes in WAT can contribute to the development of metabolic diseases. WAT with obesity is characterized by increased accumulation of CD4+ T helper type 1 (Th1) cells, cytotoxic CD8+ T cells, pro-inflammatory classically activated macrophages, and decreased abundance of regulatory T cell (Treg), Group 2 innate lymphoid cell (ILC2), and eosinophils (Eos) [104]. miR-143 is found to affect the secretion of immune cytokines in some immune cells. For example, miR-143 overexpression increases the differentiation of central memory T CD8+ cells and pro-inflammatory cytokine secretion [107], while miR-143 is downregulated during T helper 9 (Th9) cell differentiation, and its overexpression inhibited Th9 differentiation, proliferation, and IL-9 production [108]. However, up to now, the function of miR-143 in characterized immune cells of WAT with obesity has not been reported. Therefore, the role of miR-143 in immune cells and how immune cells regulate the expression of miR-143 in adipocytes are the focuses of further research.
MiR-143 is involved in the regulation of body temperature
In recent years, brown or beige adipocyte has attracted extensive attention because of their unique thermogenic function as an important target for the prevention and treatment of obesity and its complications [109]. There is increasing evidence to suggest that specific miRNAs in brown or beige adipocytes can regulate thermogenesis and confront against obesity [110, 111]. miR-143 is highly expressed in adipose tissue, and the expression of miR-143 in BAT is even higher than that in WAT [18]. It has been reported that cold exposure and CL316,243 (CL) treatment, which are classic stimuli known to induce thermogenesis of BAT, significantly reduce the expression level of circulating miR-143 [112]. We confirmed that low-dose LPS can also promote body temperature and thermogenesis of BAT by reducing the level of miR-143 [93]. However, the mechanism of miR-143 in regulating thermogenesis has not been fully exploited. Moreover, the upregulated miR-143 is found in THP-1-derived macrophages and peripheral blood mononuclear cells (PBMCs) after heat exposure and in turn targets endogenous pyrogens including IL-6, IL6ST, TLR2, PGE2, and TNF to complete a negative feedback mechanism, which may be crucial to prevent pathological hyperthermia [113]. In this report, they also defined miR-143 as a temperature-sensitive miRNA [113]. Recent studies have found that IL-27–IL-27Rα signaling plays a critical role in inducing thermogenesis, protecting against diet-induced obesity, and ameliorating insulin resistance [114]. The regulatory effect of immune cytokines on thermogenesis is further emphasized. Thus, under physiological and pathophysiological conditions, the effect of miR-143 on the cross-talk of adipocytes and immune cells is an important direction for future research.
The current gap between miR-143 and obesity and the clinical significance of miR-143
Although previous studies have reported a strong correlation between obesity and miR-143, there is a lack of evidence that miR-143 directly controlled the development of obesity. Interestingly, our latest study found that global miR-143KO mice showed a significant reduction in increased body weight, fat mass, liver, and adipose tissue weight (including BAT and WAT), and increased glucose utilization and energy expenditure during HFD-induced obesity [57, 59]. Our study demonstrated that miR-143KO can resist diet-induced obesity in vivo. We summarize the functions and regulatory pathways of miR-143 in white adipocytes, brown adipocytes, hepatocytes, and smooth muscle cells (Fig. 2). Unfortunately, to date, the mechanism of miR-143 in BAT has rarely been illustrated, though BAT is a promising anti-obesity target. And in the current studies, obesity-induced changes in miR-143 in tissues were described, but it still remains unclear how miR-143 work in fat dysfunction and by which regulatory pathways. Furthermore, tissue-specific miR-143KO mice are urgently needed to validate the contribution of miR-143 functions in different tissues (liver, muscle, and adipose tissue) to obesity.
In brown adipocytes, miR-143 targets AC9 to inhibit thermogenesis by decreased the expression of UCP1. In white adipocytes, miR-143 targets MEK5 and ERK5 to inhibit adipogenesis by inhibiting the expression of PPARγ and C/EBPα. PPARγ promotes the ubiquitination and degradation of Nur77, which in turn leads to the loss of Nur77’s transcriptional inhibition of CD36 and FABP4, thereby promoting fatty acid uptake and cell proliferation. In hepatocytes, miR-143 targets ORP8 and contributes to the development of obesity-induced insulin resistance. MiR-143 can also target MAPK11, and inhibits gluconeogenesis by repressing the activity of PCK1 and G6PC. In smooth muscle cells, miR-143 targets IGFBP5 and Elk-1 to affect cell proliferation and differentiation.
Obesity-related circulating miRNAs, an endocrine factor that promotes communication between metabolic organs and tissues [85] while having good stability [115], may be the most important indicators of potential non-invasive biomarkers for the management of obesity and related metabolic disorders [73]. Several specific miRNAs, such as miR-132 and miR-17-5p [116], are known to be differentially expressed in patients with obesity as compared with healthy individuals and are potential biomarker candidates. According to the current data, circulating miR-143 may not be suitable as a marker for obesity, due to inconsistent results from different research works, and that the downregulation of circulating miR-143 was not effective in reducing obesity in mice, although it alleviated insulin resistance [62].
Since the discovery of miRNA in the 1990s [117], miRNAs have emerged as attractive therapeutic agents for various diseases, including obesity. Two miRNA-based therapeutic tools have shown promise in preclinical studies and clinical trials: miRNA mimics and anti-miRNA oligonucleotides (inhibitors). To date, the miRNA-targeted therapeutics for metabolic disorders that have reached phase I clinical trials are antimiR-103 and anti-miR-107 (known as RG-125/AZD4076), which are intended to treat type 2 diabetes mellitus with non-alcoholic fatty liver disease (NAFLD) [118] and non-alcoholic steatohepatitis (NASH) [119]. miR-143 inhibitors may be a potential way in the treatment of obesity, since knock out of miR-143 reduced HFD-induced obesity, but it still has a long way off Another important question is how to deliver miR-143 inhibitors more efficiently and specifically to adipose tissue.
Concluding remarks and prospective
Studies have shown that miR-143 is stably and highly expressed in WAT and BAT and is downregulated as obesity occurs. This could be due to the intrinsic homeostasis regulation mechanism, which inhibits adipogenesis and promotes thermogenesis by reducing miR-143 to resist obesity. Circulating miRNAs derived from adipose tissues have been proposed as important endocrine factors in whole-body metabolic control, providing new insight into cross-organ communications. Although the expression of WAT-derived and circulating miR-143 in patients and mice with obesity is still controversial, it is confirmed that miR-143 is downregulated in the BAT of mice with obesity, indicating the important role of miR-143 in the BAT in anti-obesity. Throughout the research history of miR-143, it is not difficult to find that, in addition to our recent reports, there is almost no information on the regulatory role of miR-143 in fat mobilization, including lipid transport, lipolysis, and thermogenesis. Further study in this field and a complete description of miR-143 regulatory function in adipose tissue is an important theoretical basis for guiding miR-143 to obesity treatment.
References
Bluher M. Obesity: global epidemiology and pathogenesis. Nat Rev Endocrinol. 2019;15:288–98.
Kahn SE, Hull RL, Utzschneider KM. Mechanisms linking obesity to insulin resistance and type 2 diabetes. Nature. 2006;444:840–6.
Yancy WS Jr, Olsen MK, Guyton JR, Bakst RP, Westman EC. A low-carbohydrate, ketogenic diet versus a low-fat diet to treat obesity and hyperlipidemia: a randomized, controlled trial. Ann Intern Med. 2004;140:769–77.
Piche ME, Tchernof A, Despres JP. Obesity phenotypes, diabetes, and cardiovascular diseases. Circ Res. 2020;126:1477–500.
Brown KA. Metabolic pathways in obesity-related breast cancer. Nat Rev Endocrinol. 2021;17:350–63.
Wang H, Deng Q, Lv Z, Ling Y, Hou X, Chen Z, et al. N6-methyladenosine induced miR-143-3p promotes the brain metastasis of lung cancer via regulation of VASH1. Mol Cancer. 2019;18:181.
Cypess AM, Lehman S, Williams G, Tal I, Rodman D, Goldfine AB, et al. Identification and importance of brown adipose tissue in adult humans. N Engl J Med. 2009;360:1509–17.
van Marken Lichtenbelt WD, Vanhommerig JW, Smulders NM, Drossaerts JM, Kemerink GJ, Bouvy ND, et al. Cold-activated brown adipose tissue in healthy men. N Engl J Med. 2009;360:1500–8.
Wu J, Bostrom P, Sparks LM, Ye L, Choi JH, Giang AH, et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell. 2012;150:366–76.
Chen Y, Peng H, Zhou S, Zhuang Y. ADAR1 is targeted by miR-143 to regulate IL-1beta-induced endothelial activation through the NFkappaB pathway. Int J Biochem Cell Biol. 2017;89:25–33.
Pfeifer A, Hoffmann LS. Brown, beige, and white: the new color code of fat and its pharmacological implications. Annu Rev Pharmacol Toxicol. 2015;55:207–27.
Esau C, Kang X, Peralta E, Hanson E, Marcusson EG, Ravichandran LV, et al. MicroRNA-143 regulates adipocyte differentiation. J Biol Chem. 2004;279:52361–5.
Yi C, Xie WD, Li F, Lv Q, He J, Wu J, et al. MiR-143 enhances adipogenic differentiation of 3T3-L1 cells through targeting the coding region of mouse pleiotrophin. FEBS Lett. 2011;585:3303–9.
Xie H, Lim B, Lodish HF. MicroRNAs induced during adipogenesis that accelerate fat cell development are downregulated in obesity. Diabetes. 2009;58:1050–7.
Bae IS, Park PJ, Lee JH, Cho EG, Lee TR, Kim SH. PPARgamma-mediated G-protein coupled receptor 120 signaling pathway promotes transcriptional activation of miR-143 in adipocytes. Gene. 2017;626:64–9.
Kajimoto K, Naraba H, Iwai N. MicroRNA and 3T3-L1 pre-adipocyte differentiation. RNA. 2006;12:1626–32.
Chen L, Hou J, Ye L, Chen Y, Cui J, Tian W, et al. MicroRNA-143 regulates adipogenesis by modulating the MAP2K5-ERK5 signaling. Sci Rep. 2014;4:3819.
Jordan SD, Kruger M, Willmes DM, Redemann N, Wunderlich FT, Bronneke HS, et al. Obesity-induced overexpression of miRNA-143 inhibits insulin-stimulated AKT activation and impairs glucose metabolism. Nat Cell Biol. 2011;13:434–46.
Jiang S, Zhang LF, Zhang HW, Hu S, Lu MH, Liang S, et al. A novel miR-155/miR-143 cascade controls glycolysis by regulating hexokinase 2 in breast cancer cells. EMBO J. 2012;31:1985–98.
Fang R, Xiao T, Fang Z, Sun Y, Li F, Gao Y, et al. MicroRNA-143 (miR-143) regulates cancer glycolysis via targeting hexokinase 2 gene. J Biol Chem. 2012;287:23227–35.
Peschiaroli A, Giacobbe A, Formosa A, Markert EK, Bongiorno-Borbone L, Levine AJ, et al. miR-143 regulates hexokinase 2 expression in cancer cells. Oncogene. 2013;32:797–802.
Miao Y, Zhang LF, Guo R, Liang S, Zhang M, Shi S. et al. (18)F-FDG PET/CT for monitoring the response of breast cancer to miR-143-based therapeutics by targeting tumor glycolysis. Mol Ther Nucleic Acids. 2016;5:e357
Chen J, Yu Y, Li H, Hu Q, Chen X, He Y, et al. Long non-coding RNA PVT1 promotes tumor progression by regulating the miR-143/HK2 axis in gallbladder cancer. Mol Cancer. 2019;18:33.
Wang Y, Li G, Zhao L, Lv J. Long noncoding RNA HOTTIP alleviates oxygen-glucose deprivation-induced neuronal injury via modulating miR-143/hexokinase 2 pathway. J Cell Biochem. 2018;119:10107–17.
Zeng XZ, Liu N, Zhang J, Wang L, Zhang ZC, Zhu J, et al. Inhibition of miR-143 during ischemia cerebral injury protects neurones through recovery of the hexokinase 2-mediated glucose uptake. Biosci Rep. 2017;37:BSR20170216.
Muralimanoharan S, Maloyan A, Myatt L. Mitochondrial function and glucose metabolism in the placenta with gestational diabetes mellitus: role of miR-143. Clin Sci (Lond). 2016;130:931–41.
Fong LY, Jing R, Smalley KJ, Taccioli C, Fahrmann J, Barupal DK, et al. Integration of metabolomics, transcriptomics, and microRNA expression profiling reveals a miR-143-HK2-glucose network underlying zinc-deficiency-associated esophageal neoplasia. Oncotarget. 2017;8:81910–25.
Xu RH, Liu B, Wu JD, Yan YY, Wang JN. miR-143 is involved in endothelial cell dysfunction through suppression of glycolysis and correlated with atherosclerotic plaques formation. Eur Rev Med Pharmacol Sci. 2016;20:4063–71.
Miao Y, Zhang LF, Zhang M, Guo R, Liu MF, Li B. Therapeutic delivery of miR-143 targeting tumor metabolism in poorly differentiated thyroid cancer xenografts and efficacy evaluation using F-18-FDG microPET-CT. Hum Gene Ther. 2019;30:882–92.
Yao MY, Wang XH, Tang Y, Zhang WH, Cui B, Liu QH, et al. Dicer mediating the expression of miR-143 and miR-155 regulates hexokinase II associated cellular response to hypoxia. Am J Physiol Lung C. 2014;307:L829–L37.
Sun XH, Zhang L. MicroRNA-143 suppresses oral squamous cell carcinoma cell growth, invasion and glucose metabolism through targeting hexokinase 2. Biosci Rep. 2017;37:BSR20160404.
Guo Y, Liang F, Zhao FL, Zhao J. Resibufogenin suppresses tumor growth and Warburg effect through regulating miR-143-3p/HK2 axis in breast cancer. Mol Cell Biochem. 2020;466:103–15.
Gregersen LH, Jacobsen A, Frankel LB, Wen JY, Krogh A, Lund AH. MicroRNA-143 down-regulates Hexokinase 2 in colon cancer cells. BMC Cancer. 2012;12:232.
Zhou P, Chen WG, Li XW. MicroRNA-143 acts as a tumor suppressor by targeting hexokinase 2 in human prostate cancer. Am J Cancer Res. 2015;5:2056–63.
Cordes KR, Sheehy NT, White MP, Berry EC, Morton SU, Muth AN, et al. miR-145 and miR-143 regulate smooth muscle cell fate and plasticity. Nature. 2009;460:705–10.
Chivukula RR, Shi G, Acharya A, Mills EW, Zeitels LR, Anandam JL, et al. An essential mesenchymal function for miR-143/145 in intestinal epithelial regeneration. Cell. 2014;157:1104–16.
Yang JS, Lai EC. Alternative miRNA biogenesis pathways and the interpretation of core miRNA pathway mutants. Mol Cell. 2011;43:892–903.
Czech B, Hannon GJ. Small RNA sorting: matchmaking for Argonautes. Nat Rev Genet. 2011;12:19–31.
Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, Tuschl T. Identification of tissue-specific microRNAs from mouse. Curr Biol. 2002;12:735–9.
Boettger T, Beetz N, Kostin S, Schneider J, Kruger M, Hein L, et al. Acquisition of the contractile phenotype by murine arterial smooth muscle cells depends on the Mir143/145 gene cluster. J Clin Invest. 2009;119:2634–47.
Elia L, Quintavalle M, Zhang J, Contu R, Cossu L, Latronico MV, et al. The knockout of miR-143 and -145 alters smooth muscle cell maintenance and vascular homeostasis in mice: correlates with human disease. Cell Death Differ. 2009;16:1590–8.
Kent OA, McCall MN, Cornish TC, Halushka MK. Lessons from miR-143/145: the importance of cell-type localization of miRNAs. Nucleic Acids Res. 2014;42:7528–38.
Zhong Y, Li L, Chen Z, Diao S, He Y, Zhang Z, et al. MIR143 inhibits steroidogenesis and induces apoptosis repressed by H3K27me3 in granulosa cells. Front Cell Dev Biol. 2020;8:565261.
Xin M, Small EM, Sutherland LB, Qi X, McAnally J, Plato CF, et al. MicroRNAs miR-143 and miR-145 modulate cytoskeletal dynamics and responsiveness of smooth muscle cells to injury. Genes Dev. 2009;23:2166–78.
Sun K, Kusminski CM, Scherer PE. Adipose tissue remodeling and obesity. J Clin Invest. 2011;121:2094–101.
Vishvanath L, Gupta RK. Contribution of adipogenesis to healthy adipose tissue expansion in obesity. J Clin Invest. 2019;129:4022–31.
Zhang L, Wu ZQ, Wang YJ, Wang M, Yang WC. MiR-143 regulates milk fat synthesis by targeting Smad3 in bovine mammary epithelial cells. Animals (Basel). 2020;10:1453.
Cargnello M, Roux PP. Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol Mol Biol R. 2011;75:50–83.
Hu E, Kim JB, Sarraf P, Spiegelman BM. Inhibition of adipogenesis through MAP kinase-mediated phosphorylation of PPARγ. Science. 1996;274:100–3.
Akaike M, Che W, Marmarosh NL, Ohta S, Osawa M, Ding B, et al. The hinge-helix 1 region of peroxisome proliferator-activated receptor gamma1 (PPARgamma1) mediates interaction with extracellular signal-regulated kinase 5 and PPARgamma1 transcriptional activation: involvement in flow-induced PPARgamma activation in endothelial cells. Mol Cell Biol. 2004;24:8691–704.
Erding Hu JBK, Sarraf P, Spiegelman BM. Inhibition of adipogenesis through MAP kinase-mediated phosphorylation of PPARgamma. Science. 1996;274:2100–3.
Festuccia WT, Laplante M, Berthiaume M, Gelinas Y, Deshaies Y. PPARgamma agonism increases rat adipose tissue lipolysis, expression of glyceride lipases, and the response of lipolysis to hormonal control. Diabetologia. 2006;49:2427–36.
Hua TNM, Kim MK, Vo VTA, Choi JW, Choi JH, Kim HW, et al. Inhibition of oncogenic Src induces FABP4-mediated lipolysis via PPARgamma activation exerting cancer growth suppression. EBioMedicine. 2019;41:134–45.
Ahmadian M, Suh JM, Hah N, Liddle C, Atkins AR, Downes M, et al. PPAR gamma signaling and metabolism: the good, the bad and the future. Nat Med. 2013;19:557–66.
Camp HS, Tafuri SR. Regulation of peroxisome proliferator-activated receptor gamma activity by mitogen-activated protein kinase. J Biol Chem. 1997;272:10811–6.
Yang PB, Hou PP, Liu FY, Hong WB, Chen HZ, Sun XY, et al. Blocking PPARgamma interaction facilitates Nur77 interdiction of fatty acid uptake and suppresses breast cancer progression. Proc Natl Acad Sci USA. 2020;117:27412–22.
Liu J, Liu J, Zeng D, Wang H, Wang Y, Xiong J, et al. MiR-143-null is against diet-induced obesity by promoting BAT thermogenesis and inhibiting WAT adipogenesis. Int J Mol Sci. 2022;23:13058.
Li B, Fan J, Chen N. A novel regulator of type II diabetes: microRNA-143. Trends Endocrinol Metab. 2018;29:380–8.
Chen X, Luo J, Yang L, Guo Y, Fan Y, Liu J, et al. miR-143-mediated responses to betaine supplement repress lipogenesis and hepatic gluconeogenesis by targeting MAT1a and MAPK11. J Agric Food Chem. 2022;70:7981–992.
Ji YX, Wang YT, Li PL, Cai L, Wang XM, Bai L, et al. A kinome screen reveals that Nemo-like kinase is a key suppressor of hepatic gluconeogenesis. Cell Metab. 2021;33:1171.
Xu PZ, Vernooy SY, Guo M, Hay BA. The Drosophila microRNA mir-14 suppresses cell death and is required for normal fat metabolism. Curr Biol. 2003;13:790–5.
Xihua L, Shengjie T, Weiwei G, Matro E, Tingting T, Lin L, et al. Circulating miR-143-3p inhibition protects against insulin resistance in metabolic syndrome via targeting of the insulin-like growth factor 2 receptor. Transl Res. 2019;205:33–43.
Dahlman I, Belarbi Y, Laurencikiene J, Pettersson AM, Arner P, Kulyte A. Comprehensive functional screening of miRNAs involved in fat cell insulin sensitivity among women. Am J Physiol Endocrinol Metab. 2017;312:E482–94.
Zhang P, Du J, Wang L, Niu L, Zhao Y, Tang G, et al. MicroRNA-143a-3p modulates preadipocyte proliferation and differentiation by targeting MAPK7. Biomed Pharmacother. 2018;108:531–9.
Takanabe R, Ono K, Abe Y, Takaya T, Horie T, Wada H, et al. Up-regulated expression of microRNA-143 in association with obesity in adipose tissue of mice fed high-fat diet. Biochem Biophys Res Commun. 2008;376:728–32.
Li M, Wu H, Luo Z, Xia Y, Guan J, Wang T, et al. An atlas of DNA methylomes in porcine adipose and muscle tissues. Nat Commun. 2012;3:850.
Nazari M, Saberi A, Karandish M, Neisi N, Jalali MT, Makvandi M. Influence of L-carnitine on the expression level of adipose tissue miRNAs related to weight changes in obese rats. Pak J Biol Sci. 2016;19:227–32.
Nadler ST, Stoehr JP, Schueler KL, Tanimoto G, Yandell BS, Attie AD. The expression of adipogenic genes is decreased in obesity and diabetes mellitus. Proc Natl Acad Sci USA. 2000;97:11371–6.
van den Berghe G. The role of the liver in metabolic homeostasis: implications for inborn errors of metabolism. J Inherit Metab Dis. 1991;14:407–20.
Fernandes T, Hashimoto NY, Magalhaes FC, Fernandes FB, Casarini DE, Carmona AK, et al. Aerobic exercise training-induced left ventricular hypertrophy involves regulatory MicroRNAs, decreased angiotensin-converting enzyme-angiotensin ii, and synergistic regulation of angiotensin-converting enzyme 2-angiotensin (1-7). Hypertension. 2011;58:182–9.
de Gonzalo-Calvo D, Davalos A, Montero A, Garcia-Gonzalez A, Tyshkovska I, Gonzalez-Medina A, et al. Circulating inflammatory miRNA signature in response to different doses of aerobic exercise. J Appl Physiol (1985). 2015;119:124–34.
Mori MA, Ludwig RG, Garcia-Martin R, Brandao BB, Kahn CR. Extracellular miRNAs: from biomarkers to mediators of physiology and disease. Cell Metab. 2019;30:656–73.
Ji C, Guo X. The clinical potential of circulating microRNAs in obesity. Nat Rev Endocrinol. 2019;15:731–43.
Al-Rawaf HA. Circulating microRNAs and adipokines as markers of metabolic syndrome in adolescents with obesity. Clin Nutr. 2019;38:2231–8.
Vatandoost N, Amini M, Iraj B, Momenzadeh S, Salehi R. Dysregulated miR-103 and miR-143 expression in peripheral blood mononuclear cells from induced prediabetes and type 2 diabetes rats. Gene. 2015;572:95–100.
Can U, Buyukinan M, Yerlikaya FH. The investigation of circulating microRNAs associated with lipid metabolism in childhood obesity. Pediatr Obes. 2016;11:228–34.
Kilic ID, Dodurga Y, Uludag B, Alihanoglu YI, Yildiz BS, Enli Y, et al. MicroRNA -143 and -223 in obesity. Gene. 2015;560:140–2.
Kalluri R, LeBleu VS. The biology, function, and biomedical applications of exosomes. Science. 2020;367:6478.
van Niel G, D’Angelo G, Raposo G. Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol. 2018;19:213–28.
Keller S, Sanderson MP, Stoeck A, Altevogt P. Exosomes: from biogenesis and secretion to biological function. Immunol Lett. 2006;107:102–8.
Thery C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2:569–79.
Chen S, Tang Y, Liu Y, Zhang P, Lv L, Zhang X, et al. Exosomes derived from miR-375-overexpressing human adipose mesenchymal stem cells promote bone regeneration. Cell Prolif. 2019;52:e12669.
Ying W, Riopel M, Bandyopadhyay G, Dong Y, Olefsky JM. Adipose tissue macrophage-derived exosomal miRNAs can modulate invivo and invitro insulin sensitivity. Cell. 2017;171:372.
Isabel H-D, Chen-Yu Z, Antonio V-P. Extracellular vesicles: novel mediators of cell communication in metabolic disease. Trends Endocrinol Metab. 2017;28:3–18.
Thomou T, Mori MA, Dreyfuss JM, Konishi M, Sakaguchi M, Wolfrum C, et al. Adipose-derived circulating miRNAs regulate gene expression in other tissues. Nature. 2017;542:450–5.
De Ng ZB, Poliakov A, Hardy RW, Clements R, Zhang HG. Adipose tissue exosome-like vesicles mediate activation of macrophage-induced insulin resistance. Diabetes. 2009;58:2498–505.
Zhou X, Li Z, Qi M, Zhao P, Duan Y, Yang G, et al. Brown adipose tissue-derived exosomes mitigate the metabolic syndrome in high fat diet mice. Theranostics. 2020;10:8197.
Engin AB. MicroRNA and adipogenesis. Adv Exp Med Biol. 2017;960:489–509.
Lee MS, Kim Y. Mulberry fruit extract ameliorates adipogenesis via increasing AMPK activity and downregulating microRNA-21/143 in 3T3-L1 adipocytes. J Med Food. 2020;23:266–72.
Calo N, Ramadori P, Sobolewski C, Romero Y, Maeder C, Fournier M, et al. Stress-activated miR-21/miR-21* in hepatocytes promotes lipid and glucose metabolic disorders associated with high-fat diet consumption. Gut. 2016;65:1871–81.
Shamsi F, Wang CH, Tseng YH. The evolving view of thermogenic adipocytes – ontogeny, niche and function. Nat Rev Endocrinol. 2021;17:726–44.
Sun W, Modica S, Dong H, Wolfrum C. Plasticity and heterogeneity of thermogenic adipose tissue. Nat Metab. 2021;3:751–61.
Liu J, Zeng D, Luo J, Wang H, Xiong J, Chen X, et al. LPS-induced inhibition of miR-143 expression in brown adipocytes promotes thermogenesis and fever. Int J Mol Sci. 2022;23:13805.
Sunahara RK, Dessauer CW, Gilman AG. Complexity and diversity of mammalian adenylyl cyclases. Annu Rev Pharmacol Toxicol. 1996;36:461–80.
Lazar AM, Irannejad R, Baldwin TA, Sundaram AB, Gutkind JS, Inoue A, et al. G protein-regulated endocytic trafficking of adenylyl cyclase type 9. Elife. 2020;9:e58039–53.
Qi C, Sorrentino S, Medalia O, Korkhov VM. The structure of a membrane adenylyl cyclase bound to an activated stimulatory G protein. Science. 2019;364:389–94.
Blondin DP, Nielsen S, Kuipers EN, Severinsen MC, Jensen VH, Miard S, et al. Human brown adipocyte thermogenesis is driven by beta2-AR stimulation. Cell Metab. 2020;32:287–300.
Rautureau Y, Deschambault V, Higgins ME, Rivas D, Mecteau M, Geoffroy P, et al. ADCY9 (Adenylate Cyclase Type 9) inactivation protects from atherosclerosis only in the absence of CETP (Cholesteryl Ester Transfer Protein). Circulation. 2018;138:1677–92.
Herzig S, Shaw RJ. AMPK: guardian of metabolism and mitochondrial homeostasis. Nat Rev Mol Cell Biol. 2018;19:121–35.
Garcia D, Shaw RJ. AMPK: mechanisms of cellular energy sensing and restoration of metabolic balance. Mol Cell. 2017;66:789–800.
Trefts E, Shaw RJ. AMPK: restoring metabolic homeostasis over space and time. Mol Cell. 2021;81:3677–90.
Scheele C, Wolfrum C. Brown adipose crosstalk in tissue plasticity and human metabolism. Endocr Rev. 2020;41:53–65.
Osborn O, Olefsky JM. The cellular and signaling networks linking the immune system and metabolism in disease. Nat Med. 2012;18:363–74.
Brestoff JR, Artis D. Immune regulation of metabolic homeostasis in health and disease. Cell. 2015;161:146–60.
Villarroya F, Cereijo R, Villarroya J, Giralt M. Brown adipose tissue as a secretory organ. Nat Rev Endocrinol. 2017;13:26–35.
Yang FT, Stanford KI. Batokines: mediators of inter-tissue communication (a mini-review). Curr Obes Rep. 2022;11:1–9.
Zhang T, Zhang Z, Li F, Ping Y, Qin G, Zhang C, et al. miR-143 regulates memory T cell differentiation by reprogramming T cell metabolism. J Immunol. 2018;201:2165–75.
Qiu X, Shi Q, Huang Y, Jiang H, Qin S. miR-143/145 inhibits Th9 cell differentiation by targeting NFATc1. Mol Immunol. 2021;132:184–91.
Arany Z. Taking a BAT to the chains of diabetes. N Engl J Med. 2019;381:2270–2.
He L, Tang M, Xiao T, Liu H, Liu W, Li G, et al. Obesity-associated miR-199a/214 cluster inhibits adipose browning via PRDM16-PGC-1alpha transcriptional network. Diabetes. 2018;67:2585–600.
Fischer C, Seki T, Lim S, Nakamura M, Andersson P, Yang Y, et al. A miR-327-FGF10-FGFR2-mediated autocrine signaling mechanism controls white fat browning. Nat Commun. 2017;8:2079.
Chen Y, Buyel JJ, Hanssen MJ, Siegel F, Pan R, Naumann J, et al. Exosomal microRNA miR-92a concentration in serum reflects human brown fat activity. Nat Commun. 2016;7:11420.
Wong JJ, Au AY, Gao D, Pinello N, Kwok CT, Thoeng A, et al. RBM3 regulates temperature sensitive miR-142-5p and miR-143 (thermomiRs), which target immune genes and control fever. Nucleic Acids Res. 2016;44:2888–97.
Wang Q, Li D, Cao G, Shi Q, Zhu J, Zhang M, et al. IL-27 signalling promotes adipocyte thermogenesis and energy expenditure. Nature. 2021;600:314–8.
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18:997–1006.
Heneghan HM, Miller N, McAnena OJ, O’Brien T, Kerin MJ. Differential miRNA expression in omental adipose tissue and in the circulation of obese patients identifies novel metabolic biomarkers. J Clin Endocrinol Metab. 2011;96:E846–50.
Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–54.
US National Library of Medicine. ClinicalTrials.gov. 2016. https://clinicaltrials.gov/ct2/show/NCT02826525.
US National Library of Medicine. ClinicalTrials.gov. 2015. https://clinicaltrials.gov/ct2/show/NCT02612662.
Acknowledgements
The research was supported by grants from the National Natural Science Foundation of China (32072812, 31802156, 32072814), and the Key Project of Guangdong Provincial Nature Science Foundation (2021A1515011310, 2020A1515010062).
Author information
Authors and Affiliations
Contributions
JL, HW, and DZ drafted the manuscript. JX, JL, XC, TC, QX, and JS collected the data and organized the references. XR and YZ participated in the study design. All of the authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, J., Wang, H., Zeng, D. et al. The novel importance of miR-143 in obesity regulation. Int J Obes 47, 100–108 (2023). https://doi.org/10.1038/s41366-022-01245-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-022-01245-6
- Springer Nature Limited