Abstract
Background
Obesity is related to a myriad of cardiometabolic outcomes, each of which may have a specific metabolomic signature and a genetic basis. We identified plasma metabolites associating with different cardiometabolic risk factors (adiposity, cholesterol, insulin resistance, and inflammation) in monozygotic (MZ) twins. Additionally, we assessed if metabolite profiling can identify subgroups differing by cardiometabolic risk factors.
Methods
We quantified 111 plasma metabolites (Acquity UPLC—triple quadrupole mass spectrometry), and measured blood lipids, HOMA index, CRP, and adiposity (BMI, %bodyfat by DEXA, fat distribution by MRI) in 40 MZ twin pairs (mean BMI 27.9 kg/m2, age 30.7). We determined associations among individuals (via linear regression) between metabolites and clinical phenotypes, and assessed, with within-twin pair analysis, if these associations were free from genetic confounding. We also performed cluster analysis to identify distinct subgroups based on subjects’ metabolite profiles.
Results
We identified 42 metabolite-phenotype associations (FDR < 0.05), 19 remained significant after controlling for shared factors within the twin pairs. Aspartate, propionylcarnitine, tyrosine hexanoylcarnitine, and deoxycytidine associated positively with two or more adiposity measures. HDL cholesterol (HDL-C) associated negatively and BMI positively with the most numbers of metabolites; 12 were unique for HDL-C and 3 for BMI. Metabolites associating with HDL-C had the strongest effect size. Metabolite profiling revealed two distinct subgroups of individuals, differing by 32 metabolites (p < 0.05), and by total and LDL cholesterol (LDL-C). Forty-two metabolites predicted subgroup membership in correlation with total cholesterol and 45 metabolites predicted subgroup membership in correlation with LDL-C.
Conclusions
Different fat depots share metabolites associating with general adiposity. BMI and HDL-C associated with the most pronounced and specific metabolomic signature. Metabolomics profiling can be used to identify distinct subgroups of individuals that differ by cholesterol measures. Most of the observed metabolite-phenotype associations are free of confounding by genetics and environmental factors shared by the co-twins.
Similar content being viewed by others
Introduction
Metabolomics is increasingly used to obtain a detailed molecular view of normal physiology as well as of the pathology of metabolic diseases like obesity [1,2,3,4]. This may allow identification of metabolites as biomarkers capable of serving alongside traditional clinical markers for predicting metabolic health [5, 6].
Some studies indicate that body composition, rather than body weight [7] or body mass index (BMI) [8, 9], predicts metabolic health, while others have shown that BMI outperforms [10, 11] or is equal to more detailed adiposity measures in identifying metabolic risks [12]. It may well be that the effectiveness of BMI in describing metabolic health leans heavily on the premise that anthropometric measures are correlated with more direct measures of adiposity such as body fat percentage (%bodyfat) [13], subcutaneous adipose tissue (SAT), visceral adipose tissue (VAT) [14], or liver fat (LF) [15]. However, it is recognized that BMI does not capture subtle differences in body composition and cannot locate fat distribution [14, 16].
In addition to BMI, the distribution of fat in obesity is an important correlate for metabolic health; VAT [17, 18] or LF [19], rather than overall adiposity, contribute to unhealthy circulating metabolite profiles. In particular, VAT associates with elevations in branched chain amino acids (BCAA) [20, 21] and LF with increases in low-density lipoprotein cholesterol (LDL-C), fatty acids (FA), inflammatory markers, and BCAA [19]. VAT has also been independently linked to insulin resistance (IR) [5, 6] and dyslipidemia [6, 16]. Viscerally obese subjects with hyperinsulinemia also present with low high-density lipoprotein cholesterol (HDL-C) [22]. Consequently, the question remains as to whether BMI, as an adiposity measure, may not be enough to accurately gauge metabolic health of an individual, thereby requiring detailed body composition measures (SAT, VAT, and LF).
Monozygotic (MZ) co-twins share 100% of their genomic sequence and are perfectly or closely matched for many environmental factors. Studies on within-twin pair differences are free from genetic and shared environment confounding and highlight environmental effects not common to both twins in a pair (e.g., aspects of diet, exercise, and lifestyle) as a basis to explain individual differences within MZ twin pairs [23]. This study design is particularly useful when studying complex traits. Previous studies comparing fat distributions and cardiometabolic health measures between unrelated individuals are hampered by the fact that deposition of fat is at least moderately heritable, as are blood biochemistry measures. The heritability estimates for VAT range from 42 to 57%, for SAT from 36 to 42% [24, 25], and for LF around 37% [15]. To this end, the metabolic processes that result from differences in fat distribution between individuals may also be heritable. A genome-wide association study of 8330 Finnish individuals, including 561 twin pairs, has shown substantial twin-based heritability estimates of circulating metabolites, varying between 23 and 76% depending on the compound [26]. Genetic factors also strongly determine cardiometabolic health risk factors such as cholesterol values with heritability estimates of plasma cholesterol and triglycerides (TG) between 56 and 77% [27, 28].
Although metabolite profiles in obesity have been investigated [1, 2], we know of only one study relating the metabolite profiles with all main fat compartments in young, healthy individuals [19]. Also, because most of these studies compare metabolites between unrelated obese and lean groups [1, 4] or focus on older individuals [3], very little information exists on early predictors which link to metabolic health in young, healthy individuals who have not yet developed the distinctions that drive these groupings of metabolic health. Little is also known about how different metabolic risk factors relate to metabolic health disturbances on the molecular level. Consequently, we sought to look at the metabolite levels from the viewpoint of different metabolic risk factors including accumulation of fat in different depots and traditional blood biochemistry markers. We employ a co-twin control study design using young adult MZ twin pairs to identify metabolites significantly associating with metabolic health. We first identified metabolite patterns that associate with different fat depots and selected cardiometabolic risk factors including cholesterol levels, IR, and inflammation markers. Because there is no consensus on which cardiometabolic risk factors are more important than the others [29], we compared a range of the phenotype-metabolite associations to reveal the relative importance of each of the phenotypes. In cases where associations with metabolites were shared across phenotypes, we compared the effect sizes of these associations to highlight those phenotypes that capture the largest metabolite level change in circulating blood. We also examined, via within-twin pair analysis, if these associations are independent of shared genetic and environmental influences. Lastly, we assessed if metabolite profiling can be used to identify subgroups that differ in cardiometabolic risk factors. The identification of subgroups would indicate that the heterogeneity in cardiometabolic risk factors among the twin individuals can be refined by metabolite screening.
Materials and methods
Subjects
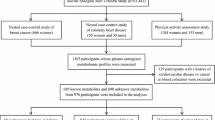
We studied 80 MZ twins (36 males, 44 females, 40 twin pairs) from two longitudinal population-based Finnish twin cohorts, FinnTwin16 (born between 1975–1979, n = 2839 pairs), and FinnTwin12 (born between 1983–1987, n = 2578 pairs) [30]. The participants have been described previously [31]. In brief, the mean BMI was 27.9 kg/m2, and mean age 30.7 years at the time of the study (Table 1). The twins were healthy, with the exception of one twin with inactive ulcerative colitis (treated with mesalazine and azathioprine), and another twin with type 2 diabetes (treated with metformin and insulin). All participants provided written informed consent. The study was performed according to the principles of the Helsinki Declaration with approval from the Ethics Committee of the Helsinki University Central Hospital.
Clinical measures were obtained as described previously [31]. We used dual-energy X-ray absorptiometry (DEXA) [32] to determine body composition including %bodyfat, magnetic resonance imaging (MRI) to measure body fat distribution to abdominal SAT and VAT, and proton magnetic resonance spectroscopy (1H-MRS) to determine LF [33]. We also measured fasting lipids (total cholesterol [TCHOL], HDL-C, LDL-C, and TG) and high-sensitivity CRP [31]. Additionally, we measured fasting glucose and insulin [31] to calculate IR by the homeostasis model assessment (HOMA). TCHOL, HDL-C, LDL-C, TG, CRP, fasting glucose, and insulin measures are collectively referred to as blood biochemistry measures in this article. The study participants disclosed their smoking behavior via questionnaire.
Metabolomic analysis
We collected plasma samples after 10–12 h fasting. Metabolites were analyzed on an Acquity Ultra (High) Performance Liquid Chromatography (UPLC)-Mass Spectrometry (MS) system coupled to a Xevo® TQ-S triple quadrupole mass spectrometer (Waters Corporation, Milford, MA, USA). Briefly, 10 µL of internal standard mixture was added to 100 µL of samples. Extraction solvent was added and the collected supernatant dispensed in OstroTM 96-well plate (Waters Corporation, Milford, USA). Then, 5 μL of vacuum-filtered sample extract was injected into UPLC-MS for metabolite separation and quantification. MassLynx 4.1 software was used for data acquisition, data handling, and instrument control. Data processing was done using TargetLynx software. Complete instrument parameters are available here [34]. The metabolites were selected such that they cover 15 different classes in a single analytical method with short running time.
We removed metabolites with missing values in any sample, and were left with 93 out of the initial 111 metabolites (Supplementary Table 1). The non-normally distributed metabolite data was rank transformed to a standard normal distribution with a mean of zero and variance of one (package GenABEL in R) [35].
Statistics and bioinformatics methods
First, we computed the associations between each phenotype (adiposity and blood biochemistry measures) and each metabolite across all individuals (n = 80). For this regression analysis, we used a linear mixed model (package lme4 in R) [35] to account for the family relationships. We identified the size of the effect of each phenotype on the metabolite by extracting the regression coefficient and p-value for each association. This regression coefficient represented the unit change in metabolite for each unit change in phenotype measure. For metabolites significantly associated with one or more phenotypes, we compared the standardized beta coefficients (in units of standard deviation) in order to see which associations had stronger effect sizes.
Next, to identify associations independent of genetic and shared environmental effects, we performed within-twin pair moderated t-tests separately for each phenotype (package limma in R) [35] to identify metabolites with significantly different concentration within-twin pairs. p-values were FDR-adjusted for multiple testing [36]. We adjusted both the regression and the moderated t-test models for age, sex, smoking, and experiment batch number.
Lastly, we assessed whether individuals form separate subgroups based on their metabolite profiles. We first obtained residuals from metabolites modeled using a linear mixed model. Using the resultant residuals after controlling for effects by age, sex, smoking, family, and batch number, we performed principal component analysis to capture variance in the adjusted data while accounting for collinearity between the metabolites. Using the 20 principal components that captured 80% of the variance in the data, we performed k-means clustering to cluster the individuals according to their metabolite profiles. We compared the metabolite concentrations between the subgroups using moderated t-tests (package limma in R). We also performed Welch t-test (for comparison of means in groups of unequal sample sizes, package psych in R) to determine if the phenotypes of these groups were significantly different. Normalized phenotype data was used and the model adjusted for age, sex, smoking, family, and the experiment batch effects. Lastly, we used partial least squares discriminate analysis (PLSDA) to simultaneously model the metabolites and each phenotype such that the linear combinations of the metabolite variables are in maximum covariance with the phenotypes [37]. We performed the PLSDA analysis using, as outcome variables, phenotypes that were significantly different between groups.
Metabolite functions were queried from the Human Metabolome DataBase (HMDB) database. We used Ingenuity Pathway Analysis (QIAGEN, Redwood City, California) to identify pathways represented by the metabolites that differed between the subgroups. Pathways with Fisher’s exact test p-value < 0.05 were considered significant.
Results
Clinical characteristics including adiposity and blood biochemistry measures for all the 80 twins are presented in Table 1. The 93 metabolites and their mean concentrations among the twin sample are listed in Supplementary Table 1.
Associations between metabolites and adiposity measures
We first regressed 93 metabolites against adiposity measures (BMI, %bodyfat, SAT, VAT, and LF). We identified 91 associations (p < 0.05) between metabolites and adiposity measures. Of these, 18 remained significant (p < 0.05) after FDR adjustment (Fig. 1 and Table 2). The 18 associations were formed between nine metabolites and the adiposity measures. Aspartate associated significantly with all the adiposity measures, propionylcarnitine with all measures but LF, deoxycytidine with BMI, SAT and VAT, and tyrosine with BMI and VAT. In addition to these four metabolites, BMI showed unique significant associations with isobutyryl carnitine, cysteine, tyrosine, and valine. SAT associated with three metabolites that also associated with VAT; VAT associated with one additional metabolite. All metabolites uniquely associating with each phenotype and shared between phenotypes are shown in Fig. 1.
Metabolites significantly (FDR < 0.05) associated with one or more phenotypes. The color intensity of the individual rectangles shows the value of the standardized regression coefficient (effect size) while the asterisks indicate if the associations are free from genetic and shared environmental confounding. The gray squares indicate non-significant associations. CRP high-sensitivity C-reactive protein, HDL-C high-density lipoprotein cholesterol, HOMA homeostatic model assessment, LDL-C low-density lipoprotein cholesterol, LF liver fat, SAT subcutaneous adipose tissue, TCHOL total cholesterol, TG triglycerides, VAT visceral adipose tissue
Among the three phenotype-associating metabolites shared between the adiposity measures, propionylcarnitine and deoxycytidine showed the largest effect sizes for BMI, while among the adiposity measures, aspartate showed the largest effect size for LF (Table 2, Fig. 1).
To assess if any of the significant associations between the metabolites and adiposity measures were confounded by genetic or shared environmental factors, we performed within-twin pair comparisons using all 40 twin pairs. Half of the significant associations remained, implying that these associations are independent of confounding by factors shared by the co-twins, and are rather due to unique environmental factors experienced by the individual twins (Table 2).
Associations between metabolites and blood biochemistry measures
Regression analysis of the 93 metabolites against blood biochemistry measures (TCHOL, HDL-C, LDL-C, TG, HOMA, and CRP) revealed 106 associations (p < 0.05), of which 24 metabolite-phenotype associations remained significant (p < 0.05) after FDR adjustment. HDL-C associated negatively with 15 metabolites while TG, CRP, and HOMA associated positively with two or more metabolites (Table 2, Fig. 1). A total of 14 of the 15 HDL-C associated metabolites were unique to HDL-C, and not shared with any other blood biochemistry measure. HOMA, HDL-C, TG, and CRP mostly had unique associations with metabolites except for S-Adenosyl-L-Homocysteine which associated with both HOMA and TG and propionylcarnitine which associated with both HOMA and HDL-C.
In the within-twin pair analyses (Table 2), 10 of the 24 associations (all with HDL-C) remained significant (FDR corrected p < 0.05), demonstrating that these observed associations exist after controlling for genetic influences and shared environmental factors.
Shared metabolite associations between adiposity and blood biochemistry measures
Six metabolites, namely, aspartate, tyrosine, deoxycytidine, hexanoylcarnitine, propionylcarnitine, S-Adenosyl-L-Homocysteine were significantly associated with two or more phenotypes (Table 2 and Fig. 1). All the shared metabolite associations were negative for HDL-C, and positive for HOMA, TG and all adiposity measures, as expected due to the opposing effects of HDL-C and the other clinical measures on metabolic health. Propionlycarnitine and deoxycytidine, found to be associated with BMI, SAT, VAT, and HDL-C, were highly correlated with each other (r > 0.7) (Supplementary Fig. 1).
In general, when a metabolite associated with adiposity and blood biochemistry measures, the association was strongest (based on the unstandardized effect size) for the blood biochemistry measures. The same was true when assessing the significant (FDR < 0.05) log fold changes in the within-twin pair analyses.
Out of the 42 significant associations between the metabolites and the phenotypes, for 13 associations the absolute effect size exceeded 0.6. All these metabolites associated with HDL-C. Out of the three metabolites associating with HDL-C that also associated with at least one adiposity phenotype, the effect size (standardized) with HDL-C was always the largest (Table 2, Fig. 1) except for propionylcarnitine. HDL-C, followed by BMI, also had the most number of uniquely associated metabolites (Fig. 2).
Explained variation in metabolites and outcome variables by the (Orthogonal Partial Least Squares) OPLS components. a PLSDA plot showing separation between both the two subgroup as well as TCHOL values. Here, six OPLS components explain 53% of the variation in the metabolite concentrations and 80% of the variation in both the subgroup membership and TCHOL values. b PLSDA plot showing separation between both the two subgroup as well as LDL-C values. Here, six orthogonal PLS components explain 53% of the variation in the metabolite concentrations and 80% of the variation in both the subgroup membership and LDL-C values. LDL-C low-density lipoprotein cholesterol, PLSDA Partial Least Square Discriminate Analysis, TCHOL total cholesterol
Clustering based on the metabolite profiling
Next, we explored the variances of each metabolite across the samples in order to cluster the individuals according to their metabolite profiles. We found two subgroups (clusters) of individuals differing by 32 metabolites (p < 0.05) (Supplementary Table 2). The concentrations of all these metabolites were higher in Subgroup 2 compared with Subgroup 1. Acylcarnitines and amino acids valine, leucine, isoleucine, and phenylalanine were the most prevalent metabolites differing between the subgroups. Pathway analysis for the differing metabolites revealed 14 significant pathways responsible for amino acid metabolism and cell signaling (Supplementary Fig. 2). We then performed group comparison analysis to determine if the two groups differ by their clinical phenotypes. We found that Subgroup 2 had significantly higher TCHOL and LDL-C compared to Subgroup 1 (Table 3). Additionally, we performed PLSDA analysis to identify metabolites that were important predictors for the subgroup membership and clinical phenotype. Using PLSDA, 10 PLS components captured 50% of the variation in metabolite concentration correlating with 80% of the variation in subgroup membership and either TCHOL or LDL-C (Fig. 2a, b). A total of 42 metabolites predicted subgroup membership correlating with TCHOL values and 45 metabolites predicted subgroup membership correlating with LDL-C values. In both groups of metabolites, amino acids and acylcarnitines featured prominently (Table 4).
Discussion
BMI associated positively with more metabolites than any of the other studied adiposity measures (%bodyfat, SAT, VAT, and LF) in this study. HDL-C was negatively associated with the most number of metabolites, and showed the strongest effect sizes. Half of the associations remained significant after controlling for genetic and shared environmental influences within the twin pairs. Metabolic profiling clustered the subjects into two distinct subgroups, which differed by 32 metabolites, as well as by total and LDL-C.
We first identified metabolites that significantly associated with different adiposity measures in individual twins. Many of these adiposity-associated metabolites were amino acids followed by acylcarnitines. BMI is often used as a measure of obesity and an indirect measure of body fat. In accordance with this, we found that two out of the three metabolites associating with body fat percentage also associated with BMI, with higher effect sizes. There were overlaps in the metabolites associated with each of the fat depots and BMI, reflecting known high correlations between the different adiposity measures [38]. Our findings of several phenotype-associating metabolites commonly shared among the adiposity measures are consistent with these observations. Several studies on plasma metabolites have identified positive associations between BCAA and aromatic amino acids (AAA) with BMI [39, 40], being in line with our findings of BCAA valine and AAA tyrosine associating with BMI.
We also identified metabolites significantly associated with different blood biochemistry measures. Out of six blood biochemistry measures, HOMA, HDL-C, TG, and CRP associated with one or more metabolites. HDL-C had associations with the most number of metabolites; whenever these metabolites also associated with other phenotypes (both adiposity and blood biochemistry measures), the effect size of the association with HDL-C was always higher. HDL-C also associated with the most number of metabolites not associated with any other measure. Low serum HDL-C has previously been shown to associate with obesity and high risk for metabolic disease [41]. We propose that HDL-C is an important measure because of its ability to indirectly measure the most comprehensive metabolite profile changes, even when the differences in metabolite concentrations are small. Previous studies have also found strong negative correlation between HDL-C and central obesity characterized by VAT [42, 43]. While our findings agree with a previous study [44] and highlight HDL-C as an important predictor of metabolic health, further studies are still needed to assess if metabolites associated with HDL-C are independent or somewhat related to the increase in VAT in obesity. HOMA and TG associated positively with S-Adenosyl-L-Homocysteine, a metabolite found to be increased in vitamin B12 deficiencies and to alter adipocyte functionality through epigenetic mechanisms [45]. S-Adenosyl-L-Homocysteine treatment for cultured 3T3-L1 adipocytes resulted in reduced glucose uptake and lipolysis for the cells [45], but how plasma levels of this metabolite relate to adipose tissue function in vivo remains to be studied.
The highest number of significant metabolite associations were found for BMI and HDL-C. BMI and HDL-C associated with several amino acids (both AAA and BCAA) and acylcarnitines in the current study. Associations of BCAAs, AAAs, and acylcarnitines with body weight and IR have already been established in several studies [46, 47]. Newgard et al. [4] found a metabolic signature made out of AAA, BCAA, and acylcarnitines associating with metabolic disease [46]. More specifically, downregulation of adipose tissue mitochondrial catabolism of BCAAs in obesity is a consistent finding [31, 48]. Low mitochondrial oxidation in the tissues could explain why the BCAA levels are increased in the plasma of obese individuals. BCAA catabolism is mainly a mitochondrial event, and it is known to decrease in muscle [49], liver [50], and adipose tissue [31, 48] in obesity-related conditions, therefore resulting in accumulation of the BCAAs in the bloodstream. In obesity, the incompletely oxidized degradation products from BCAAs, the short-chain acylcarnitines, are also elevated in plasma [4, 40]. In other words, both the increase in circulating acylcarnitine levels and amino acids observed in our study may be indicative of the incomplete BCAA oxidation process.
Across all the measured metabolites, propionylcarnitine, aspartate, tyrosine, deoxycytidine, S-Adenosyl-L-Homocysteine, and Hexanoylcarnitine associated with two or more adiposity or blood biochemistry phenotypes. Propionylcarnitine associated negatively with HDL-C and positively with HOMA and all the adiposity measures, being in line with previously reported associations with BMI [4, 40]. Propionylcarnitine is generated in order to decrease plasma propionyl CoA produced from the catabolism of BCAA and odd-chain FAs [46]. Thus, the increased propionylcarnitine levels observed in the current study may indicate the body’s attempt to lower the circulating BCAA and FAs in plasma [51]. Aspartate was found to positively associate with TG and all adiposity measures in our study. While not much is known about aspartate in metabolic health, it does play a major role in nucleotide and protein biosynthesis [52] as well as oxidative phosphorylation in mitochondria which was previously shown to be impaired in obesity [53]. The positive association between tyrosine and BMI, and negative association between tyrosine and HDL-C observed in the current study are in line with a previous study linking tyrosine to IR and obesity [4]. Deoxycytidine was positively associated with BMI, SAT, and VAT, negatively with HDL-C. Deoxycytidine has multiple functions in DNA synthesis and repair, as well as in cellular growth and proliferation [54].
HOMA associated with three metabolites, two were acylcarnitines. Acylcarnitines in mitochondria have previously been identified as biomarkers of IR [47]. Whether impaired insulin or glucose function drives the increased levels of acylcarnitines or vice versa, is unknown [55, 56]. In the within-twin pair analysis, HOMA-metabolite associations disappeared, suggesting that HOMA is highly heritable. Heritability for HOMA has previously been estimated around 0.78 [57].
It is noteworthy that there were no metabolites that significantly associated with TCHOL and LDL-C, which are important measures related to metabolic health. Especially high LDL-C is often seen as an important independent risk factor for atherosclerosis and cardiovascular disease [58]. The absence of significant metabolite associations for these metabolic health-related phenotypes could be because our twins were relatively healthy, or because of low statistical power. Despite the lack of associations between the individual metabolites and these metabolic health measures, we show that clustering by metabolite profiling can identify subgroups of individuals that differ for their metabolic health. We identified two distinct groups that differ by LDL-C and TCHOL values. The group with higher LDL-C and total cholesterol values also showed higher concentrations of 32 metabolites compared with the other group. Circulating acylcarnitines made up one third of these 32 metabolites. There were also higher amounts of all three BCAAs and AAA phenylalanine, which were also captured as pathways responsible for amino acid metabolism and cell signaling in this group. Lastly, we identified 42 metabolites that were predictors for classifying individuals into subgroups according to the TCHOL values and 45 metabolites that were predictors for classifying individuals into subgroups according to the LDL-C values. While our sample size is small, this is an important step in identifying metabolites as classifiers for our two subgroups of differing metabolic health.
In the within-twin pair analysis, most of the relationships between the phenotypes and metabolites remained. Because MZ co-twins are genetically identical at the sequence level and share many environmental factors, significant within-twin pair differences in metabolite levels point to environmental factors for which the twin pair is discordant as an underlying contributor for the differing metabolite levels. It has been previously shown that BMI, HDL-C, TG, as well as BCAA, AAA, and acylcarnitines change by the effect of nutrition and exercise [59]. The present study adds to this by showing a fairly large number of specific phenotype-associated metabolites differing within the twin pairs, suggesting that these associations are not driven by genetic factors but rather by the unique environment, such as diet. However, in a prior study including a subset of these twins, we did not identify any significant within-twin pair differences in food intake or physical activity [31]. Further research is needed to identify the specific environmental component driving these differences. Metabolite differences between individuals with the same genotype could well be because of differences in their epigenetic profiles. Epigenetic marks react to environmental effects and thereby mediate their effects on the function of the genome. Therefore differing environments within MZ twin pairs may have produced epigenetic differences, which may have resulted in the differing clinical phenotypes or metabolite levels observed in this study.
MZ co-twin studies provide perhaps the best controlled study design available in humans owing to the complete or close match for genes, age, gender, and intrauterine and childhood environment. The germline genetic background is identical in each pair, hence acquired factors must account for the phenotype discordance [60]. However, while we can rule out genetic and shared environmental factors from the observed associations in this study, the cross-sectional design does not allow us to determine cause and effect. Another limitation of our study is the limited number of metabolites assessed. However, the targeted platform we used allows (semi) quantification of the identified metabolites while platforms with untargeted approach yield only qualitative data.
To summarize, we established that associations with HDL-C had stronger effect sizes; HDL-C also associated with the most number of metabolites. Measuring HDL-C, the most comprehensive and sensitive phenotype in our study, is sufficient for capturing even small changes in metabolite levels and is hence a suitable detector of early disturbances in metabolic health. Metabolite profiles consisting of several metabolites also proved to be capable of discerning different categories of metabolic health. To conclude, metabolites, either individually or as part of a larger profile, are useful biomarkers for metabolic health.
References
Hanzu FA, Vinaixa M, Papageorgiou A, Parrizas M, Correig X, Delgado S, et al. Obesity rather than regional fat depots marks the metabolomic pattern of adipose tissue: an untargeted metabolomic approach. Obesity. 2014;22:698–704.
Jennings A, MacGregor A, Pallister T, Spector T, Cassidy A. Associations between branched chain amino acid intake and biomarkers of adiposity and cardiometabolic health independent of genetic factors: a twin study. Int J Cardiol. 2016;223:992–8.
Kraus WE, Pieper CF, Huffman KM, Thompson DK, Kraus VB, Morey MC, et al. Association of plasma small-molecule intermediate metabolites with age and body mass index across six diverse study populations. J Gerontol A Biol Sci Med Sci. 2016;71:1507–13.
Newgard CB, An J, Bain JR, Muehlbauer MJ, Stevens RD, Lien LF, et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab. 2009;9:311–26.
McKillop AM, Flatt PR. Emerging applications of metabolomic and genomic profiling in diabetic clinical medicine. Diabetes Care. 2011;34:2624–30.
Zhang A, Sun H, Wang X. Serum metabolomics as a novel diagnostic approach for disease: a systematic review. Anal Bioanal Chem. 2012;404:1239–45.
Segal KR, Dunaif A, Gutin B, Albu J, Nyman A, Pi-Sunyer FX. Body composition, not body weight, is related to cardiovascular disease risk factors and sex hormone levels in men. J Clin Invest. 1987;80:1050–5.
Lahmann PH, Lissner L, Gullberg B, Berglund G. A prospective study of adiposity and all-cause mortality: the Malmö diet and cancer study. Obes Res. 2002;10:361–9.
Carroll JF, Chiapa AL, Rodriquez M, Phelps DR, Cardarelli KM, Vishwanatha JK, et al. Visceral fat, waist circumference, and BMI: impact of race/ethnicity. Obesity. 2008;16:600–7.
Bosy-Westphal A, Geisler C, Onur S, Korth O, Selberg O, Schrezenmeir J, et al. Value of body fat mass vs anthropometric obesity indices in the assessment of metabolic risk factors. Int J Obes. 2006;30:475–83.
Lee K, Song YM, Sung J. Which obesity indicators are better predictors of metabolic risk?: healthy twin study. Obesity. 2008;16:834–40.
Paniagua L, Lohsoonthorn V, Lertmaharit S, Jiamjarasrangsi W, Williams MA. Comparison of waist circumference, body mass index, percent body fat and other measure of adiposity in identifying cardiovascular disease risks among Thai adults. Obes Res Clin Pract. 2008;2:215–23.
Ranasinghe C, Gamage P, Katulanda P, Andraweera N, Thilakarathne S, Tharanga P. Relationship between body mass index (BMI) and body fat percentage, estimated by bioelectrical impedance, in a group of Sri Lankan adults: a cross sectional study. BMC Public Health. 2013;13:797.
Camhi SM, Bray GA, Bouchard C, Greenway FL, Johnson WD, Newton RL, et al. The relationship of waist circumference and BMI to visceral, subcutaneous, and total body fat: sex and race differences. Obesity. 2011;19:402–8.
Schwimmer JB, Celedon MA, Lavine JE, Salem R, Campbell N, Schork NJ, et al. Heritability of nonalcoholic fatty liver disease. Gastroenterology. 2009;136:1585–92.
Frankenfield DC, Rowe WA, Cooney RN, Smith JS, Becker D. Limits of body mass index to detect obesity and predict body composition. Nutrition. 2001;17:26–30.
Gao X, Zhang W, Wang Y, Pedram P, Cahill F, Zhai G, et al. Serum metabolic biomarkers distinguish metabolically healthy peripherally obese from unhealthy centrally obese individuals. Nutr Metab. 2016;13:33.
Schlecht I, Gronwald W, Behrens G, Baumeister SE, Hertel J, Hochrein J, et al. Visceral adipose tissue but not subcutaneous adipose tissue is associated with urine and serum metabolites. PLoS ONE. 2017;12:e0175133.
Rämö JT, Kaye SM, Jukarainen S, Bogl LH, Hakkarainen A, Lundbom J, et al. Liver fat and insulin sensitivity define metabolite profiles during a glucose tolerance test in young adult twins. J Clin Endocrinol Metab. 2016;102:220-23.
Boulet MM, Chevrier G, Grenier-Larouche T, Pelletier M, Nadeau M, Scarpa J, et al. Alterations of plasma metabolite profiles related to adipose tissue distribution and cardiometabolic risk. Am J Physiol Endocrinol Metab. 2015;309:E736–46.
Bogl LH, Kaye SM, Ramo JT, Kangas AJ, Soininen P, Hakkarainen A, et al. Abdominal obesity and circulating metabolites: a twin study approach. Metab Clin Exp. 2016;65:111–21.
Pascot A, Lemieux I, Prud’homme D, Tremblay A, Nadeau A, Couillard C, et al. Reduced HDL particle size as an additional feature of the atherogenic dyslipidemia of abdominal obesity. J Lipid Res. 2001;42:2007–14.
van Dongen J, Slagboom PE, Draisma HHM, Martin NG, Boomsma DI. The continuing value of twin studies in the omics era. Nat Rev Genet. 2012;13:640–653.
Hong Y, Rice T, Gagnon J, Després J-P, Nadeau A, Pérusse L, et al. Familial clustering of insulin and abdominal visceral fat: the HERITAGE Family Study 1. J Clin Endocrinol Metab. 1998;83:4239–45.
Pérusse L, Després JP, Lemieux S, Rice T, Rao DC, Bouchard C, et al. Familial aggregation of abdominal visceral fat level: results from the Quebec family study. Metabolism. 1996;45:378–82.
Kettunen J, Tukiainen T, Sarin AP, Ortega-Alonso A, Tikkanen E, Lyytikainen LP, et al. Genome-wide association study identifies multiple loci influencing human serum metabolite levels. Nat Genet. 2012;44:269–76.
Elder SJ, Lichtenstein AH, Pittas AG, Roberts SB, Fuss PJ, Greenberg AS, et al. Genetic and environmental influences on factors associated with cardiovascular disease and the metabolic syndrome. J Lipid Res. 2009;50:1917–26.
Teucher B, Skinner J, Skidmore PM, Cassidy A, Fairweather-Tait SJ, Hooper L, et al. Dietary patterns and heritability of food choice in a UK female twin cohort. Twin Res Hum Genet. 2007;10:734–48.
Gomez-Ambrosi J, Silva C, Galofre JC, Escalada J, Santos S, Millan D, et al. Body mass index classification misses subjects with increased cardiometabolic risk factors related to elevated adiposity. Int J Obes. 2012;36:286–94.
Kaprio J. Twin studies in Finland 2006. Twin Res Human Genet. 2006;9:772–7.
Muniandy M, Heinonen S, Yki-Jarvinen H, Hakkarainen A, Lundbom J, Lundbom N, et al. Gene expression profile of subcutaneous adipose tissue in BMI-discordant monozygotic twin pairs unravels molecular and clinical changes associated with sub-types of obesity. Int J Obes. 2017;41:1176-1184.
Pietrobelli A, Formica C, Wang Z, Heymsfield SB. Dual-energy X-ray absorptiometry body composition model: review of physical concepts. Am J Physiol. 1996;271:941.
Lundbom J, Hakkarainen A, Söderlund S, Westerbacka J, Lundbom N, Taskinen M-R, et al. Long-TE 1H MRS suggests that liver fat is more saturated than subcutaneous and visceral fat. NMR Biomed. 2011;24:238–45.
Roman-Garcia P, Quiros-Gonzalez I, Mottram L, Lieben L, Sharan K, Wangwiwatsin A, et al. Vitamin B12–dependent taurine synthesis regulates growth and bone mass. J Clin Invest. 2014;124:2988–3002.
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B. 1995;57:289–300.
Wold S, Ruhe A, Wold H, Dunn I WJ. The collinearity problem in linear regression. The partial least squares (PLS) approach to generalized inverses. SIAM J Sci Stat Comput. 1984;5:735–43.
Smith SR, Lovejoy JC, Greenway F, Ryan D, deJonge L, de la Bretonne J, et al. Contributions of total body fat, abdominal subcutaneous adipose tissue compartments, and visceral adipose tissue to the metabolic complications of obesity. Metab Clin Exp. 2001;50:425–35.
Ho JE, Larson MG, Ghorbani A, Cheng S, Chen MH, Keyes M, et al. Metabolomic profiles of body mass index in the Framingham heart study reveal distinct cardiometabolic phenotypes. PLoS ONE. 2016;11:e0148361.
Kim JY, Park JY, Kim OY, Ham BM, Kim H-J, Kwon DY, et al. Metabolic profiling of plasma in overweight/obese and lean men using ultra performance liquid chromatography and Q-TOF mass spectrometry (UPLC-Q-TOF MS). J Proteome Res. 2010;9:4368–75.
Assmann G, Schulte H, von Eckardstein A, Huang Y. High-density lipoprotein cholesterol as a predictor of coronary heart disease risk. The PROCAM experience and pathophysiological implications for reverse cholesterol transport. Atherosclerosis. 1996;124:S11–20.
Després JP, Moorjani S, Ferland M, Tremblay A, Lupien PJ, Nadeau A, et al. Adipose tissue distribution and plasma lipoprotein levels in obese women. Importance of intra-abdominal fat. Arterioscler Thromb Vasc Biol. 1989;9:203–10.
Nieves DJ, Cnop M, Retzlaff B, Walden CE, Brunzell JD, Knopp RH, et al. The atherogenic lipoprotein profile associated with obesity and insulin resistance is largely attributable to intra-abdominal fat. Diabetes. 2003;52:172–9.
Smith SC. Multiple risk factors for cardiovascular disease and diabetes mellitus. Am J Med. 2007;120:S3–11.
Ngo S, Li X, O’Neill R, Bhoothpur C, Gluckman P, Sheppard A, et al. Elevated S-adenosylhomocysteine alters adipocyte functionality with corresponding changes in gene expression and associated epigenetic marks. Diabetes. 2014;63:2273–83.
Newgard CB. Interplay between lipids and branched-chain amino acids in development of insulin resistance. Cell Metab. 2012;15:606–14.
Pallares-Mendez R, Aguilar-Salinas CA, Cruz-Bautista I, Del Bosque-Plata L. Metabolomics in diabetes, a review. Ann Med. 2016;48:89–102.
Pietilainen KH, Naukkarinen J, Rissanen A, Saharinen J, Ellonen P, Keranen H, et al. Global transcript profiles of fat in monozygotic twins discordant for BMI: pathways behind acquired obesity. PLoS Med. 2008;5:e51.
Leskinen T, Rinnankoski-Tuikka R, Rintala M, Seppanen-Laakso T, Pollanen E, Alen M, et al. Differences in muscle and adipose tissue gene expression and cardio-metabolic risk factors in the members of physical activity discordant twin pairs. PLoS ONE. 2010;5. pii: e12609.
She P, Van Horn C, Reid T, Hutson SM, Cooney RN, Lynch CJ, et al. Obesity-related elevations in plasma leucine are associated with alterations in enzymes involved in branched-chain amino acid metabolism. Am J Physiol Endocrinol Metab. 2007;293:1552.
Brass EP, Beyerinck RA. Effects of propionate and carnitine on the hepatic oxidation of short- and medium-chain-length fatty acids. Biochem J. 1988;250:819–25.
Sullivan Lucas B, Gui Dan Y, Hosios Aaron M, Bush Lauren N, Freinkman E, Vander Heiden Matthew G. Supporting aspartate biosynthesis is an essential function of respiration in proliferating cells. Cell. 2015;162:552–63.
Heinonen S, Buzkova J, Muniandy M, Kaksonen R, Ollikainen M, Ismail K, et al. Impaired mitochondrial biogenesis in adipose tissue in acquired obesity. Diabetes. 2015;64:3135–45.
Radu CG, Shu CJ, Nair-Gill E, Shelly SM, Barrio JR, Satyamurthy N, et al. Molecular imaging of lymphoid organs and immune activation using positron emission tomography with a new 18F-labeled 2′-deoxycytidine analog. Nat Med. 2008;14:783.
Mai M, Tonjes A, Kovacs P, Stumvoll M, Fiedler GM, Leichtle AB, et al. Serum levels of acylcarnitines are altered in prediabetic conditions. PLoS ONE. 2013;8:e82459.
Adams SH, Hoppel CL, Lok KH, Zhao L, Wong SW, Minkler PE, et al. Plasma acylcarnitine profiles suggest incomplete long-chain fatty acid beta-oxidation and altered tricarboxylic acid cycle activity in type 2 diabetic African-American women. J Nutr. 2009;139:1073–81.
Mills GW, Avery PJ, McCarthy MI, Hattersley AT, Levy JC, Hitman GA, et al. Heritability estimates for beta cell function and features of the insulin resistance syndrome in UK families with an increased susceptibility to type 2 diabetes. Diabetologia. 2004;47:732–8.
Sharrett AR, Ballantyne CM, Coady SA, Heiss G, Sorlie PD, Catellier D, et al. Coronary heart disease prediction from lipoprotein cholesterol levels, triglycerides, lipoprotein(a), apolipoproteins A-I and B, and HDL density subfractions. The Atherosclerosis Risk in Communities (ARIC) Study. Circulation. 2001;104:1108–13.
Brass EP. Supplemental carnitine and exercise. Am J Clin Nutr. 2000;72:618s–23s.
Martin N, Boomsma D, Machin G. A twin-pronged attack on complex traits. Nat Genet. 1997;17:387–92.
Acknowledgements
We thank the twins for their invaluable contribution to the study. The Obesity Research Unit team and the staff at the Finnish Twin Cohort Study and FIMM Metabolomics Unit are acknowledged for their help in the collection of the data.
Funding
This study was supported by The Academy of Finland (grant number 266286 to KHP, and 251316 and 297908 to MO, and 141054, 265240, 263278, and 264146 to JK); Centre of Excellence in Research on Mitochondria, Metabolism and Disease (FinMIT) to KHP (grant 272376); Helsinki University Central Hospital (NL, AH, AR, KHP); The University of Helsinki Research Funds (MO, KHP); grants from following Foundations: Novo Nordisk (KHP), Signe and Ane Gyllenberg (KHP), Jalmari and Rauha Ahokas (KHP), The Sigrid Juselius Foundation (MO and JK), Finnish Diabetes Research Foundation (KHP), Finnish Foundation for Cardiovascular Research (KHP). Data collection in FinnTwin16 and FinnTwin12 was supported by the National Institute of Alcohol Abuse and Alcoholism (grants AA-12502 and AA-09203 to RJ Rose).
Author contributions
M.O. and K.H.P. conceived and designed the study, supervised the work, and participated in the discussion and interpretation of the results. M.M. analyzed all the data and wrote the manuscript. K.H.P., J.K., and A.R. collected the study material. A.H., J.L., and N.L. participated in the imaging of the twins. V.V. supervised the metabolomics sample analyses. J.K. was responsible for the base cohorts from which the pairs were sampled. All authors read and approved the manuscript. M.O. and K.H.P. are the guarantors of this work and as such had full access to the data and take responsibility for the integrity of the data and the accuracy of the data analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Muniandy, M., Velagapudi, V., Hakkarainen, A. et al. Plasma metabolites reveal distinct profiles associating with different metabolic risk factors in monozygotic twin pairs. Int J Obes 43, 487–502 (2019). https://doi.org/10.1038/s41366-018-0132-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-018-0132-z
- Springer Nature Limited
This article is cited by
-
Maximizing the value of twin studies in health and behaviour
Nature Human Behaviour (2023)