Abstract
Cancer is characterised by DNA hypermethylation and gene silencing of CpG island-associated promoters, including tumour-suppressor genes. The methyl-CpG-binding domain (MBD) family of proteins bind to methylated DNA and can aid in the mediation of gene silencing through interaction with histone deacetylases and histone methyltransferases. However, the mechanisms responsible for eliciting CpG island hypermethylation in cancer, and the potential role that MBD proteins play in modulation of the methylome remain unclear. Our previous work demonstrated that MBD2 preferentially binds to the hypermethylated GSTP1 promoter CpG island in prostate cancer cells. Here, we use functional genetic approaches to investigate if MBD2 plays an active role in reshaping the DNA methylation landscape at this locus and genome-wide. First, we show that loss of MBD2 results in inhibition of both maintenance and spread of de novo methylation of a transfected construct containing the GSTP1 promoter CpG island in prostate cancer cells and Mbd2−/− mouse fibroblasts. De novo methylation was rescued by transient expression of Mbd2 in Mbd2−/− cells. Second, we show that MBD2 depletion triggers significant hypomethylation genome-wide in prostate cancer cells with concomitant loss of MBD2 binding at promoter and enhancer regulatory regions. Finally, CpG islands and shores that become hypomethylated after MBD2 depletion in LNCaP cancer cells show significant hypermethylation in clinical prostate cancer samples, highlighting a potential active role of MBD2 in promoting cancer-specific hypermethylation. Importantly, co-immunoprecipiation of MBD2 shows that MBD2 associates with DNA methyltransferase enzymes 1 and 3A. Together our results demonstrate that MBD2 has a critical role in ‘rewriting’ the cancer methylome at specific regulatory regions.
Similar content being viewed by others
Introduction
DNA methylation at the carbon-5 position of cytosine residues in CpG dinucleotides is an important epigenetic mechanism involved in normal biological regulatory processes including cellular differentiation, genomic imprinting and X-inactivation.1, 2, 3 In development, initiation and maintenance of CpG methylation occurs primarily during DNA replication through the action of DNA methyltransferase enzymes (DNMTs), which catalyse the DNA methylation reaction whereby the methyl donor S-adenosyl-methionine is covalently added to the 5’-position of the cytosine molecule.4 In the normal mammalian genome ~70% of all CpG sites are methylated, except for CpG islands, regions of high CpG density, that largely remain unmethylated.5 CpG island promoters and CpG island ‘shores’, located ~2 kb from CpG islands can acquire cell type-specific methylation during differentiation6 or can become abnormally methylated in cancer.6, 7 Hypermethylation of CpG islands is primarily associated with gene repression, in carcinogenesis;3, 8 however, the precise mechanisms that result in aberrant DNA methylation of typically unmethylated CpG island and CpG island shores remain largely unknown.
One of the proposed mechanisms of transcriptional repression mediated by CpG methylation involves the binding of methyl-CpG-binding domain (MBD) proteins9 and the recruitment of co-repressor complexes.10 The MBD family members MBD1, MBD2, MBD4 and MeCP2 all bind to methylated DNA via a methyl-binding domain.11, 12 MBD proteins are therefore often described as ‘readers’ of DNA methylation and evidence suggests that they also have a role in epigenetic remodelling and gene repression.13 For example, protein interaction studies have shown that the methyl-binding domain proteins recruit binding partners such as chromatin remodellers, histone deacetylases and histone methylases to methylated DNA leading to transcriptional deregulation.14, 15, 16 MBD2 is a subunit of the Mi2-NuRD complex and has been shown to mediate gene repression via recruitment of the complex to methylated promoters.11, 17 In particular, there is mounting evidence linking MBD2 function and the aberrant hypermethylation of CpG islands in cancer.18, 19, 20
The CpG island associated with the GSTP1 gene is a remarkable example of a CpG rich promoter, which is susceptible to hypermethylation in the majority of prostate tumours yet remains unmethylated in the normal prostate cell.21 We previously reported that hypermethylation of a GSTP1 CpG island promoter construct can be triggered by a combination of transcriptional gene silencing and ‘seeds’ of CpG methylation that potentially act as a catalyst for the spread of de novo methylation to neighbouring CpG sites.18 Notably, we showed that MBD2 was preferentially bound to the ‘seeds’ of CpG methylation at the GSTP1 CpG island promoter.18 In this study we address if MBD2 binding has an active role in promoting CpG island hypermethylation in cancer. Using a functional genetics approach we now demonstrate that loss of MBD2 results in the inhibition of both maintenance and spread of de novo methylation and triggers significant hypomethylation genome-wide in prostate cancer cells, particularly at cancer-associated CpG islands and CpG island shores. Our studies suggest that MBD2 is not just a ‘reader’ of methylation but is a key player in ‘rewriting’ methylation in cancer potentially through its interaction with the DNMT machinery.
Results
Knockdown of MBD2 inhibits the maintenance and spread of DNA hypermethylation of a GSTP1 CpG island promoter transfected construct
To investigate a potential causal role for MBD2 in promoting DNA hypermethylation, we first established LNCaP cell clones with reduced MBD2 expression using shRNA to MBD2 and a scrambled control shRNA. ShRNAs were transfected into LNCaP cells and multiple single colonies were isolated under neomycin selection and examined for MBD2 expression. Real-time quantitative PCR (qRT-PCR) and western blot analysis showed that MBD2 mRNA and protein expression was reduced by 60–80% in a number of the independent stable clones (examples shown in Supplementary Figure 1). Next we determined the effect of MBD2 knockdown (MBD2-KD) on triggering DNA hypermethylation of GSTP1 transfected gene constructs that were inactive and ‘seeded’ with or without a low level of methylation. Previously we had cloned a 2.3 kb XbaI/SnaBI fragment containing the GSTP1 gene, including 1.2 kb promoter upstream from the start of transcription and exons 1–4 and inserted a polyadenylation signal 3’ to ensure transcript stability.22 We modified the construct by deleting CpG sites −2 to −8 spanning the SpI sites and minimal promoter region to silence GSTP1 expression (GSTP1-Sp1). In addition, we created a GSTP1 construct using HpaII methylase to ‘pre-seed’ the GSTP1 construct (GSTP1-Sp1-M) (Figure 1a and Supplementary Figure 2). The shRNA-MBD2 plasmid and the GSTP1 constructs (GSTP1-Sp1 and GSTP1-Sp1-M) were co-transfected into LNCaP cells and single resistant colonies were selected and DNA and RNA was isolated after ~22 doublings (early passage). RNA was analysed for stable knockdown of MBD2 by qRT-PCR and clones that had achieved the highest level of knockdown were evaluated for DNA methylation status of the transfected GSTP1 CpG island using clonal bisulphite sequencing (Figures 1b and c).
Knockdown of MBD2 inhibits the maintenance and spread of DNA hypermethylation of the GSTPI CpG island promoter in LNCaP cells. (a) The GSTP1 vector constructs showing the GSTP1 CpG island and start of transcription of GSTP1 gene, the SpI deletion construct GSTP1-Sp1 (with deleted SpI binding sites: Δ denotes deletion of CpG sites −8 to −2), and the GSTP1-Sp1-M with red dots marking HpaII methylated CCGG sites within the GSTP1 CpG island map. (b) The methylation profile by clonal bisulphite sequencing of the GSTP1-Sp1 vector transfected into wild-type LNCaP cells and two independent LNCaP cells with MBD2 shRNA knockdown. The MBD2 expression relative to 18s is shown relative to LNCaP. Closed circles indicate a methylated CpG and open circles indicate an unmethylated CpG. (c) The methylation profile of the GSTP1-Sp1-M vector after transfection into wild-type LNCaP cells (control), and four independent transfected cell lines with MBD2 knockdown at early passage (22 doublings). Red dots mark the HpaII methylated CCGG sites on the vector construct. The level of MBD2 knockdown relative to 18s is shown expressed relative to LNCaP wild-type expression. (d) Student’s t-test showing statistically significant reduction in maintenance and de novo methylation in the MBD2 KD clones relative to wild-type LNCaP cells. Average number of methylated sites in wild-type LNCaP cells (n=11) and all LNCaP cell clones with MBD2 knockdown (n=48). Error bars represent s.e.m. P-values were calculated using two-tailed Student’s paired t-tests.
The unmethylated GSTP1 promoter construct (GSTP1-Sp1) remained essentially unmethylated after transfection into wild-type LNCaP cells or MBD2 knockdown clones (85 and 95%) (Figure 1b). In contrast, the GSTP1-Sp1-M construct was extensively methylated, both at the HpaII methylated ‘seeded’ sites (maintenance methylation) and at surrounding CpG sites (de novo methylation) after transfection into wild-type LNCaP cells (Figure 1c). However, we observed in the clones with MBD2 knockdown (Figure 1c) a significant lack of maintenance methylation of the HpaII sites (P<.00001), as well as a significant reduction of de novo DNA methylation at surrounding CpG sites (P=0.0007) (Figure 1d). After further passaging (~24 doublings: late passage), LNCaP cells transfected with GSTP-Sp1-M showed more extensive de novo methylation (Supplementary Figure 3a), while the MBD2-KD cells (clones #15 and #12), showed a significant decrease in both de novo and HpaII maintenance methylation (Supplementary Figure 3b).
DNA de novo methylation of a GSTP1 CpG island promoter transfected construct is inhibited in Mbd2-/- fibroblast cells
To further interrogate the role of MBD2 in promoting DNA hypermethylation, we performed a similar transfection experiment with mouse wild-type and Mbd2−/− knockout fibroblast cells.23 No detectable mRNA or protein expression was observed in the Mbd2−/− fibroblast cells compared with the WT fibroblast cells (Figures 2a and b). After co-transfection of GSTP1-Sp1 and the ‘empty’ expression vector pQCXIP into wild-type and Mbd2−/− fibroblast cells, stably transfected colonies were isolated and passaged (early (P1), mid (P5) and late passages (P18)) and the transfected GSTP1 CpG island was analysed for methylation by clonal bisulphite sequencing. We observed minimal methylation at early passage (P1) in the transfected wild-type cells (Figure 2c). However in later passages (P5 and P18) there was a notable increase of de novo methylation (Figure 2c). In contrast, in the Mbd2−/− cells, there was essentially no de novo methylation observed even after 18 passages (Figure 2d). Similarly, after transfection of GSTP1-Sp1-M into wild-type and Mbd2−/− cells we observed significantly reduced maintenance methylation (P=0.017) in Mbd2−/− cells at both early and late passages (Supplementary Figure 4).
DNA de novo methylation of a GSTP1 CpG island promoter transfected construct is inhibited in Mbd2−/− fibroblast cells. (a) RT-qPCR of Mbd2 expression in wild-type, Mbd2−/− knockout and Mbd2 knock-in (Mbd2−/− +Mbd2) mouse fibroblast cells normalised to 18s. (b) Western blot showing Mbd2 expression in wild-type, Mbd2−/− knockout and Mbd2 knock-in (Mbd2−/− +Mbd2) mouse fibroblast cells. (c and d) The methylation profile of the GSTP1-Sp1 vector after transfection into wild-type mouse fibroblast cells (c) and Mbd2−/− cells (d) at early (passage 1), mid (passage 5) and late (passage 18) passages. Closed circles indicate a methylated CpG and open circles indicate an unmethylated CpG. (e) The methylation profile of the GSTP1-Sp1 vector transfected in Mbd2−/− +Mbd2 knock-in cells at early (passage 1) and mid (passage 5). (f) Student’s t-test showing statistically significant reduction in methylation in the Mbd2−/− cells and gain in methylation in Mbd2−/− +Mbd2 knock-in cells. Average number of methylated sites in each cell line. Error bars represent s.e.m. P-values were calculated using two-tailed Student’s paired t-tests between wild-type Mbd2 (n=33) and Mbd2-/- cells (n=30) at P1, P5 and P18, and between Mbd2−/− (n=20) and Mbd2−/− +Mbd2 cells (n=25) at P1 and P5. RT-qPCR, real-time quantitative PCR.
To investigate if Mbd2 has an active role in the maintenance and de novo methylation, we next re-expressed Mbd2 in the knockout cells (Mbd2−/− +Mbd2). The Mbd2-pQCXIP expression vector was co-transfected with GSTP1-Sp1 vector into mouse Mbd2−/− cells and resistant colonies were pooled and passaged. Re-expression of Mbd2 was confirmed by qRT-PCR and western blot analysis (Figures 2a and b). DNA was analysed for methylation of the transfected GSTP1 CpG island by genomic bisulphite clonal sequencing. Although no methylation was observed in Mbd2−/− cells (Figure 2d), we found significant increase in levels of de novo methylation at individual CpG sites in Mbd2−/− +Mbd2 cells, and this methylation was maintained upon further passaging (Figures 2e and f; P<0.000001).
MBD2 protein is coupled with DNA methyltransferases
Since MBD2-KD is associated with poor maintenance methylation and a decrease in de novo methylation, we next determined if MBD2 is associated with DNA methyltransferase enzymes (DNMTs). Whole-cell lysates from LNCaP cells were immunoprecipitated with MBD2 antibody and the eluted immunoprecipitates were immunoblotted using DNMT1 and DNMT3a antibodies (Figure 3). Notably we found that MBD2 co-immunoprecipitated with not only the histone deacetyltransferase HDAC1 as previously reported24 but also DNMT1 and DNMT3a (Figures 3a and b). Conversely, DNMT1 and DNMT3a co-immunoprecipited with HDAC1 and MBD2 (Figures 3c and d). Association of DNMT1 and DNMT3a with MBD2 supports a role for MBD2 in maintenance and de novo DNA methylation.
MBD2 is coupled with DNA methyltransferases DNMT1 and DNMT3A. (a) LNCaP whole-cell extracts were immunoprecipitated with an antibody against MBD2 (Sigma M7218). Western blot analysis with HDAC1 and DNMT1 antibodies shows physical interaction between MBD2 and HDAC1 and DNMT1. (b) Formaldehyde-fixed (1.5%) LNCaP nuclear extracts were immunoprecipitated with MBD2 antibody and then immunoblotted with DNMT3A showing MBD2 in complex with DNMT3A. (c) Reciprocal immunoprecipitation of LNCaP whole-cell extract with DNMT1 antibody shows reciprocal interactions with MBD2 and HDAC1. (d) Reciprocal immunoprecipitation of LNCaP formaldehyde-fixed nuclear extract (1.0%) with DNMT3A shows reciprocal interaction with MBD2. Input, unprecipitated extracts; IgG IP, control IP; IP, immunoprecipitation; * denotes formaldehyde-fixed nuclear extract.
MBD2 knockdown causes genome-wide hypomethylation
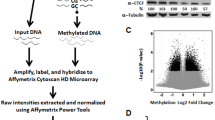
We next asked if there was a genome-wide change in the DNA methylation landscape as a consequence of decreased MBD2 expression. We performed Infinium HumanMethylation450 BeadChip (HM450K) arrays on three independent MBD2-KD clones and three scrambled controls (Supplementary Figure S1). Differential methylation analysis of MBD2 knockdown and scrambled controls showed that a depletion of MBD2 predominantly results in a loss of DNA methylation. We identified 20 630 hypomethylated probes in the MBD2-KD clones and only 2248 hypermethylated probes (β-value difference >10%, and unadjusted P-value cutoff of 0.05). Using a more stringent cutoff of 20%, we identified 4493 hypomethylated probes and 432 hypermethylated probes, while a 30% cutoff identified 745 hypo and 60 hypermethylated probes (Supplementary Table 1). Notably, using MiSeq bisulphite amplicon sequencing of six regions that encompassed hypomethylated probes, we found that methylation loss was not just limited to the individual CpG site identified on the array, but extended across neighbouring CpG sites (Figure 4a). The average methylation levels detected in the MBD2-KD clones were significantly reduced compared with the scrambled controls (ASS1, P=0.0001; BARHL2, P=0.00013; SGK2, P=0.0064 (Figure 4a); and LHX8, P=0.0001; TESK2, P=0.036; ZNF814, P=0.00043) (Supplementary Figure S5). Next, we showed that there was a reduction in direct binding of MBD2 to the hypomethylated loci (ASS1, SGK2, BARHL2, LHX8, TESK2, CDC42EP3 and OAT) in the MBD2-KD clones in contrast to the scrambled controls (Figure 4b). Thus confirming that MBD2 was integral for maintaining DNA methylation at these sites.
MBD2 Knockdown causes genome-wide hypomethylation. (a) HM450K screen shots of candidate regions ASS1, BARHL2 and SGK2 showing three scrambled controls and three MBD2-KD clones. Hypo-probes show individual significantly hypomethylated probes with a β-value difference in methylation of >20% between MBD2-KD and scrambled clones. The shaded grey denotes the region that was validated by MiSeq amplicon bisulphite sequencing. The regions were PCR amplified using bisulphite treated DNA from three MBD2-KD and three scrambled clones, and sequenced on the Illumina MiSeq platform. The scatter plots show the % methylation of CpG sites across the regions of interest: ASS1 (chr9: 133320721-133321061); BARHL2 (chr1: 91182069-91182442) and SGK2 (chr20: 42187225-42187451). Each dot represents the % methylation at an individual CpG site for a single clone and the lines represent the average methylation for the region in MBD2-KD (red) and scrambled controls (blue). Filled black triangle denotes CpG sites represented on the array. The bar charts on the right show the average methylation of scrambled control and MBD2-KD; P-values were calculated using two-tailed Student’s paired t-tests comparing average methylation percentage of all CpG sites; error bars represent s.e.m. (b) MBD2 chromatin immunoprecipitation was performed on wild-type and MBD2-KD LNCaP cells. Quantitative PCR of candidate hypomethylated regions (ASS1, BARHL2, SGK2, LHX8, CDC42EP3, OAT and TESK2) shows enrichment of MBD2 binding (ChIP/input) at these genomic loci and reduction in MBD2 binding in MBD2-KD. Error bars represent s.e.m. (c) Pie diagram showing the % of hypomethylated probes across CpG islands and CpG island shores. (d) Pie chart showing the percentage of hypomethylated probes across ChromHMM functionally annotated regions of the genome.
To determine whether MBD2-KD-associated hypomethylation is localised to specific regulatory regions of the genome, we overlapped regional hypomethylated CpG probes with CpG islands, CpG island shores and computationally derived chromatin segmentation (ChromHMM) of the LNCaP genome.25 Hypomethylated regions were defined by at least three hypomethylated probes with the maximal distance between probes of 500 bp resulting in 353 hypomethylated regions containing 1263 hypomethylated probes (Supplementary Table 1). We found that 21 and 27% of hypomethylated regions, respectively, occurred at CpG islands and CpG island shores (Figure 4c). Using the ChromHMM annotation of the LNCaP genome the largest fraction of hypomethylated regions (25%) fell within Polycomb repressed regions. In addition, active and poised enhancer regions (~28%) also showed hypomethylation (Figure 4d). Interestingly, we found that the hypomethylated probes are located in regions of decreased methylated CpG density (Supplementary Figure S6).
Functional characterisation of MBD2-regulated genomic loci
To interrogate the functional role of regions that were associated with MBD2-KD hypomethylation we profiled RNA expression of three MBD2-KD and three scrambled clones on Affymetrix HuGene 1.0ST arrays that were profiled for DNA methylation. Differential expression analysis identified 134 upregulated and 103 downregulated genes in MBD2-KD clones as compared with scrambled controls (Figure 5a and Supplementary Table 2). GO term analysis of upregulated genes revealed significant enrichment of genes associated with chromatin and nucleosome assembly and remodelling (Benjamin P-value<0.05) (Figure 5b). Validation by qRT-PCR confirmed decreased MBD2 expression (Supplementary Figure S1) and increased expression of upregulated candidate genes ASS1, TESK2 and ELF5 (Figure 5c and Supplementary Figure S7) in MBD2-KD clones. Next, we asked if MBD2-KD associated hypomethylation results in increased expression of corresponding target genes. Given the enrichment of hypomethylated regions in LNCaP ChromHMM active and poised enhancers we mapped these to putative target genes using a distance threshold of ±20 kb. Bootstrapping analysis revealed that MBD2-KD hypomethylation of enhancers was significantly associated with increased (P<0.001) but not decreased expression (P=0.13) of the target genes. Similar analysis revealed significant association between MBD2-KD hypomethylation of promoter regions and gene activation (P=0.03), but not repression (P=0.89) of the corresponding genes. We conclude that MBD2-KD hypomethylation of both enhancer and promoter regions results in upregulation of target genes. Cell proliferation experiments also showed that MBD2 KD decreases cell viability (Supplementary Figure S8). To predict the functional significance of the hypomethylated regions we performed gene set enrichment using GREAT analysis. This analysis identified gene sets significantly enriched in the MSigDB Genetic Perturbation database associated with prostate cancer, prostate carcinogenesis, methylated in cancer, as well as other cancers such as kidney cancer, glioblastoma and multiple myeloma (Supplementary Figure S9a). This suggests that MBD2 has an important role in promoting DNA hypermethylation in cancer-related genes.
Gene expression changes associated with decreased MBD2 expression. (a) Affymetrix HuGene 1.0ST expression arrays were performed on RNA isolated from three scrambled controls and three independent MBD2-KD clones, #3, #9 and #33. Heatmaps showing the top 50 genes of up- and downregulated genes (filtered so that logFC>1.25 and P< 0.005 — 50 genes). Scale indicates level of gene expression. Genes highlighted in yellow were validated by RT-qPCR. (b) GO term analysis of upregulated genes showing significantly enriched GO Terms (Benjamin P-value<0.05). (c) RT-qPCR was performed on RNA isolated from three scrambled controls and three independent MBD2-KD clones, #3, #9 and #33 of candidate upregulated genes ASS1, TESK2 and ELF5. Bars represent the mean and error bars represent s.e.m. * indicates a significant difference (0.01<P<0.05) and ** indicates a very significant difference (0.001<P<0.01) using GraphPad Prism (Prism6 for MacOSX, GraphPad software, La Jolla, CA, USA), unpaired t-test, two-tailed, with equal s.d. RT-qPCR, real-time quantitative PCR.
Cancer-specific CpG island and shore hypermethylation is linked to MBD2 binding
To determine if MBD2 associated gain of methylation is likely to have a role in cancer, we asked if the regions that become hypomethylated upon MBD2-KD are differentially methylated in normal and cancer (i) cell lines and (ii) tissue samples. First we tested whether there was a difference in the distribution of methylation β-values across probes that become hypomethylated upon MBD2-KD in the normal prostate epithelium cell line PrEC as compared with prostate cancer cell line LNCaP. Our results showed that DNA methylation was significantly lower in the PrEC cells (Figure 6a). Moreover, elevated methylation levels in LNCaP cells were reduced to levels comparable to those in the normal PrEC cells upon MBD2-KD and this shift was strongest at CpG islands and CpG island shores (Figure 6b). Since CpG islands and CpG island shores are focal points of aberrant hypermethylation in cancer we asked whether MBD2-KD hypomethylated CpG sites are more prone to aberrant hypermethylation. We performed log-odds statistical testing against probes that are statistically significantly hypermethylated in LNCaP as compared with PrEC cells (false discovery rate<0.05; avg.meth.diff>10%) and found a statistically significant association between cancer-specific aberrant hypermethylated and MBD2-KD hypomethylation (log-odds ratio 0.55; P<2e−16). The association remained significant even when probes were stratified according to their functional annotation: CpG islands (log-odds ratio 0.74; P=2.26e−07), CpG island shores (log-odds ratio 0.71; P=2.61e−10) (Figure 6c).
Cancer-specific hypermethylation is linked to MBD2 binding. (a) Distribution of Methylation β-values of all 353 hypomethylated regions (n=1,263 probes) in normal prostate epithelial cells PrEC, prostate cancer LNCaP cells and MBD2-KD LNCaP cells visualised as boxplots. (b) Distribution of Methylation β-values of 353 hypomethylated regions in normal prostate epithelial cells PrEC cells, LNCaP cancer cell line, and MBD2-KD cells, sub-setting the regions into CpG islands and CpG shores. (c) Log-odds ratios with corresponding 95% confidence intervals quantifying positive association between MBD2-KD hypomethylation and LNCaP hypermethylation events across all probes, CpG island probes, and CpG island shores probes. (d) Distribution of Methylation β-values of 353 hypomethylated regions in TCGA tumour (n=339) and TCGA normal (n=49) samples, visualised as boxplots. (e) Distribution of Methylation β-values of 353 regions, sub-setting the regions as CpG islands and CpG island shores. (f) Log-odds test for positive association between MBD2-KD hypomethylation and prostate tumour hypermethylation events across all probes, CpG island probes, and CpG island shores probes. In a, b, d and e), Wilcoxon rank-sum test was used to assess statistical significance of difference in distributions (***P-value<0.001; **P-value<0.01; *P-value<0.05). In (c and f) log-odds test was used to assess statistical significance of positive association (***P-value<0.001; **P-value<0.01; *P-value<0.05).
Next, we extended our analysis to clinical samples of normal and tumour prostate tissues for which HM450K data is available from The Cancer Genome Atlas (TCGA) consortium. Again, we observed that regions that become hypomethylated upon MBD2-KD showed significantly lower methylation in normal prostate as compared to prostate tumour tissue samples (P=1.02e−03) and the differences are more pronounced at CpG islands (P=2.51e−06) and CpG island shores (P=1.23e−02) (Figures 6d and e). Log-odds test confirmed statistically significant positive associations between MBD2 KD driven hypomethylation and aberrant hypermethylation in prostate cancer at all probes (log-odds ratio 0.28; P=2.97e−05), CpG islands (log-odds ratio 0.32; P=0.01), and CpG island shores (log-odds ratio 0.35; P=0.004) (Figure 6f). Notably the CpG islands and shores that show cancer-specific hypermethylation in prostate cancer are associated with gene sets significantly enriched in the MolSigDB Genetic Perturbation database, affected by epigenetic aberrations in prostate cancer, downregulated in prostate cancer, genes methylated in colon cancer, primary tumours and across a panel of cancer cell lines (Supplementary Figure S9b and c). GO Molecular Function also showed significant enrichment of peptide, hormone, enhancer and regulatory region sequence-specific DNA binding of these regions. Together these data support a key role for MBD2 in the spread of hypermethylation at CpG islands and shores in prostate cancer.
Discussion
Even though it has long been established that cancer is characterised by global hypomethylation and focal hypermethylation of CpG island gene promoters the mechanisms responsible for the alteration of DNA methylation patterns in cancer cells are still unclear. We previously reported that in prostate cancer hypermethylation of the GSTP1 CpG island promoter can be initiated by a combination of transcriptional gene silencing and ‘seeds’ of methylation that potentially act as a catalyst for the spread of methylation to neighbouring CpG sites.22 We showed that the methyl-CpG-binding protein MBD2 preferentially binds to the ‘seeds’ of methylation18 and proposed that MBD2 recruitment may have a role in de novo methylation of the GSTP1 CpG island promoter. In this present study, we show that MBD2 has a casual role in reshaping the DNA methylome at both this locus and other cancer-associated CpG islands and CpG island shores.
The MBD family proteins are critical players in determining the transcriptional state of the epigenome.13, 26 MBD proteins interact with other protein partners,14, 15, 16 including SIN3A, CoREST, SWI-SNF and NuRD (nucleosome remodelling and deacetylase complex), and have a key role in guiding protein complexes with chromatin remodelling and/or histone modifying activity to specific sequences in the genome. Although the role of MBD proteins as repressive chromatin remodellers is well described, the potential role of the MBD proteins in guiding the ‘writing’ of the DNA methylation landscape is less clear. Here, we show that MBD2 not only binds to histone deacetylase HDAC1 (as also reported by others),11, 27 but also binds to the maintenance methyltransferase (DNMT1) and de novo methyltransferase enzyme (DNMT3a). The DNMTs have also been reported to associate with the NuRD complex,11, 24 and since MBD2 recruits NuRD to methylated DNA, this suggested that DNA methylation is working in conjunction with chromatin remodelling.11 We asked if MBD2 could by playing a similar role in directly guiding ‘writing’ of the DNA methylation landscape. Using functional genetic approaches we showed that loss of MBD2 inhibited both the maintenance and spread of de novo methylation of a transfected GSTP1 CpG island promoter construct. Interestingly, re-expression of MBD2 rescued the de novo methylation capacity of the cells suggesting that MBD2 can recruit DNMTs to specific CpG sites to promote methylation.
We next asked if loss of MBD2 affects the methylation landscape genome-wide. Remarkably we found that MBD2 depletion led to a prominent 10-fold increase of hypomethylation compared with hypermethylation events, in contrast to others reporting hypermethylation of candidate genes.28, 29 Interestingly, we found that CpG islands and CpG island shores were enriched in the regions that became demethylated in the MBD2-KD cells. This is in keeping with previous studies showing a preference for MBD2 binding at methylated CpG island containing promoters.30, 31, 32 In addition, we found enrichment of hypomethylation at other gene-regulatory genomic regions, including polycomb-marked regions and enhancers. These regions are less dense in CpG content and methylation, which is consistent with previous work showing that MBD2 directly binds at promoters with intermediate CpG content.26 Interestingly, some CpG island regions, that harbour a high density of CpG methylation, show no discernible demethylation upon MBD2 loss, suggesting a threshold of CpG methylation that is not amenable to methylation remodelling.33, 34
The functional consequence of MBD2 depletion is genome-wide changes in gene expression, including both gene activation and gene repression, in agreement with other studies.31, 33 Activation of genes supports depletion of the direct repressive effect of MBD2,17 whereas gene repression may be a result either of an indirect effect or potential loss of reported MBD2 binding to active genes and unmethylated promoters.26, 31, 35 Notably, we found that a number of the activated genes are accompanied by promoter and enhancer demethylation. Gene activation in conjunction with demethylation upon MBD2 depletion supports a direct role of MBD2 in the facilitation of DNA methylation. Interestingly these activated gene sets are enriched with ontology database terms associated prostate cancer, prostate carcinogenesis, methylated in cancer. This agrees with previous evidence showing that the removal of MBD2 expression alleviates the repression of tumour-suppressor genes p16INK4a and p14ARF, as well as candidate tumour suppressor genes in HeLa cells,33 pTERT20 and BRAC1-NBR236 in MBD2 knockdown cells and pS2 expression in MCF-7 cells,37 consistent with the findings from our data.
Accumulating evidence has supported a significant role for MBD2 in the hypermethylation of candidate CpG islands in cancer.18, 20, 38, 39, 40, 41, 42 In this study, we demonstrated a causal role for MBD2 in facilitation of DNA methylation principally at aberrantly hypermethylated CpG islands and CpG shores and regulatory regions in LNCaP cells, and importantly we show that these same regions are also significantly hypermethylated in clinical prostate cancer. MBD proteins have been implicated in several cancers13 and MBD2-deficient mice crossed onto tumour-prone ApcMin/+ background, develop significantly smaller and few tumours.43 With the increase in cancer genome sequencing the mutational frequency of MBD protein family members is now being better illuminated in different cancer types. Mutations in MBD2 commonly lie in the methyl-binding domain and transcriptional repressor domain, which potentially disrupts crucial MBD2 functions in cancer.13 Together our results support a model whereby the methyl-binding domain protein MBD2 has a key role in the maintenance and spread of DNA methylation of CpG islands and shores in cancer potentially through its interactions with the DNMT machinery. Importantly we show that MBD2 is not only just a ‘reader’ of the methylation landscape but also can have a direct role in promoting the spread of cancer-specific DNA hypermethylation.
Materials and methods
Cell culture
LNCaP prostate cancer cells were cultured and transfected as described previously.22 Tailtip fibroblasts from Mbd2−/− (2-7 S) and wild-type (2-3 S) mice were a kind gift from Adrian Bird’s Laboratory, Edinburgh and cultured as described in.17, 23 The identity of each cell line was authenticated and cells were routinely tested for mycoplasma contamination.
MBD2 knockdown and Mbd2-/- transfections
ShRNA directed against MBD2 (Acc. No. AF072242.1) (5′-GATGATGCCTAGTAAATTA-3′ (865-894 bp)) and scrambled control (5′-GTGAATACAGGCTTTAAATAG-3) were cloned into the siLentGene-2 Vector (Promega, Madison, WI, USA; Cat.#C8070) according to the manufacturer’s instructions. The GSTP1 vector constructs were prepared and co-transfected with Promega shRNA MBD2 vector (10:1 ratio) into LNCaP cells (as detailed in Song et al.22). Stable transfectants were selected by GeneticinR (G418) (Gibco BRL). The 2-3 S and Mbd2−/− cells were transfected using JetPEI transfectant reagent (Qbiogene, Montreal, QC, Canada) and a co-transfection (1:10 ratio) of selection plasmid pQCXIP (Clontech Laboratories, Clayton, VIC, Australia) to GSTP1 vector was used. Stable clones were selected with puromycin (ThermoFisher Scientific, Waltham, MA, USA).
Over expression of Mbd2
The full-length mouse Mbd2 cDNA clone (ID IRAVp968H0390D6; Accession no BC046607) was purchased from RZPD Deutsches Ressourcenzentrum fur Genomforschung GmbH, Berlin, Germany. The Mbd2 gene (1.8 kb) was excised from the pCMV-Sport6 vector by 5’Sal1 -3’Not1 digestion and ligated into NotI-SalI digested pENTRTM2B Gateway vector (Invitrogen Life Technologies, Carlsbad, CA, USA) and inserted into expression vector pQCXIP (Clontech Laboratories, Clayton, VIC, Australia). The sequence of the inserted Mbd2 was confirmed by sequencing. Transfections were performed as described above with a co-transfection ratio of 10:1 of MBD2-pQCXIP (Clontech Laboratories) to GSTP1 vector and stable clones selected with puromycin (ThermoFisher Scientific).
Quantitative Real-Time qRT-PCR analysis
RNA was extracted using Trizol reagent (Ambion, 15596-018 supplied by ThermoFisher Scientific) and cDNA was reverse transcribed as described previously.18 qRT-PCR primers listed in Supplementary Table 3.
Clonal bisulphite and MiSeq amplicon sequencing
Clonal bisulphite methylation sequencing analysis from triplicate PCRs were performed as described previously.18, 38 PCRs for MiSeq amplicon sequencing were performed in duplicate and pooled. Library prep was performed according to the Illumina TruSeq DNA sample prep kit (Cat No FC-121-2001) (Illumina, San Diego, CA, USA) and sequenced on the Illumina MiSeq sequencer. Primers listed in Supplementary Table 3.
Protein extraction and western blot analysis
Whole-cell protein lysates were prepared as previously described.44 Protein concentration was determined using a BCA assay (Pierce, 23225/23227 supplied by ThermoFisher Scientific). Western transfer protocol was carried out according to manufacturer’s instructions with the XCell SureLock system (ThermoFisher Scientific). Immunoprecipitation was performed according to the Abcam (Cambridge, UK) protocol (http://docs.abcam.com/pdf/protocols/Immunoprecipitation_protocol.pdf). For fixed protein immunoprecipitation, cells were fixed on the dish at room temperature for 10 mins in 1.5% formaldehyde.45, 46 Fixed cell pellets were resuspended in sonication buffer (10 mm EDTA pH 8.0, 50 mM Tris-HCL pH8.0, 1% SDS) and sonicated for five cycles of 10 pulses of 0.9 s ON and 0.1 s OFF (Branson Sonifier, Danbury, CT, USA). Unsonicated material was pelleted and supernatant diluted 1:10 with PBS before immunoprecipitation. GAPDH was used as a control for nonspecific binding. For western blot analysis following immunoprecipitation (IP), IP lysate was resolved as above or under non-reducing conditions (37 oC for 10 min, exclusion of reducing agent in the sample preparation). For all antibodies used see Supplementary Table 4.
Chromatin Immunoprecipitation
Chromatin immunoprecipitations (MBD2 antibody: Sigma M7318 Lot#111M4751) were carried out using the ChIP-IT High Sensitivity Kit (Active Motif Cat No. 53040) according to the manufacturer’s instructions. ChiP-qPCR of candidate regions were performed and normalised to input. ChiP-qPCR primers listed in Supplementary Table 3.
Gene Expression Array Analysis
RNA was extracted from cell lines using Trizol reagent (Invitrogen) according to the manufacturer's protocol. Affymetrix GeneChip human gene 1.0ST (Santa Clara, CA, USA) arrays were performed as described previously.47
Illumina HumanMethylation450 BeadChip (HM450K) analysis
Bisulphite converted DNA was hybridised to Illumina HumanMethylation450 BeadChip (HM450K) arrays48 and run on an Illumina HiScan System according to the manufacturer’s instructions. Three replicates per condition (MBD2-KD or scrambled controls) were profiled. Raw HM450K signal intensity data was processed and background-normalised using preprocessIllumina function from Bioconductor minfi package.49 Methylation beta-values (β) were used for visualisation and M-values for statistical analyses.49 Differentially methylated probes between MBD2-KD and scrambled controls were identified using linear models and empirical Bayes methods implemented in the Bioconductor limma package.50 We defined hypomethylated regions as regions with at least 3 hypomethylated probes within 500 bp.
Genomic location of hypomethylated probes
Genomic coordinates (hg19) of CpG islands were obtained from UCSC genome browser. Genomic coordinates of CpG island shores were derived by taking +/-2Kb flanking regions around CpG islands. ChromHMM segmentation25, 51 of LNCaP chromatin was performed then collapsed into seven distinct states and manually annotated by comparison to the published ChromHMM model for HMEC cells.51 We used hyper-geometric testing to determine statistical significance of overlap between regional hypomethylated probes and the above functional annotations of the genome. Regional hypomethylated probes (n=1263) were used as test set and regional background probes (n=145 397) as background-set. Methylated CpG density was calculated as the number of CpG sites with methylation above 0.5 in 100 bp region entered on the probe.
Affymetrix HuGene 1.0ST expression array analysis
Arrays were normalised within each batch using RMA function from Bioconductor oligo package.50 Only probes that were annotated in RefSeq or GenBank (according to the Affymetrix annotation release 32) were included in downstream analyses. Differentially expressed genes were determined using Bioconductor limma package,50 with the linear model including a covariate representing batch.
Correlation of MBD2-KD hypomethylation events with aberrant methylation in cell line model and clinical samples of prostate cancer
Differential methylation analysis HM450K data from three replicates of LNCaP/PrEC cell resulted in 62,346 (out of 145 397 regional background probes) as being hypermethylated in LNCaP cells (false discovery rate<0.05; Δβ>10%). Raw HM450K data from TCGA prostate cancer (PRAD) cohort52 was obtained in April 2014 comprising 49 normal and 340 tumour samples. Differential methylation analysis resulted in 28 140 (out of 145 397 regional background probes) as being hypermethylated in prostate tumour samples (false discovery rate<0.05; Δβ>10%). Statistical significance of positive association between MBD2-KD hypomethylation and LNCaP hypermethylation in prostate cancer (cell line model or clinical samples) was assessed with log-odds test as implemented in R vcd package.
Gene set enrichment analysis of MBD2 hypomethylation events
GREAT (Genomic Regions Enrichment of Annotations Tool) analysis was applied53 to regional MBD2-KD hypomethylated probes to assess enrichment of gene sets in GO Molecular Function and MolSigDB Genetic Perturbation categories. Analysis was repeated with a subset of probes restricted to CpG islands and CpG island shores. In both analyses the distal extension parameter was set to 20Kb. Gene Ontology enrichment analysis was performed using the David Bioinformatics resources54, 55 on differentially expressed genes.
Data Access
Affymetrix expression array and HM450K methylation data have been deposited to GEO (GSE77206).
References
Mohandas T, Sparkes RS, Shapiro LJ . Reactivation of an inactive human X chromosome: evidence for X inactivation by DNA methylation. Science 1981; 211: 393–396.
Gartler SM, Riggs AD . Mammalian X-chromosome inactivation. Annu Rev Genet 1983; 17: 155–190.
Baylin SB, Jones PA . A decade of exploring the cancer epigenome - biological and translational implications. Nat Rev Cancer 2011; 11: 726–734.
Bestor TH . Cloning of a mammalian DNA methyltransferase. Gene 1988; 74: 9–12.
Stirzaker C, Taberlay PC, Statham AL, Clark SJ . Mining cancer methylomes: prospects and challenges. Trends Genet 2014; 30: 75–84.
Doi A, Park IH, Wen B, Murakami P, Aryee MJ, Irizarry R et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat Genet 2009; 41: 1350–1353.
Irizarry RA, Ladd-Acosta C, Wen B, Wu Z, Montano C, Onyango P et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat Genet 2009; 41: 178–186.
Bert SA, Robinson MD, Strbenac D, Statham AL, Song JZ, Hulf T et al. Regional activation of the cancer genome by long-range epigenetic remodeling. Cancer Cell 2013; 23: 9–22.
Hendrich B, Bird A . Identification and characterization of a family of mammalian methyl-CpG binding proteins. Mol Cell Biol 1998; 18: 6538–6547.
Klose RJ, Bird AP, Genomic DNA . methylation: the mark and its mediators. Trends Biochem Sci 2006; 31: 89–97.
Zhang Y, Ng HH, Erdjument-Bromage H, Tempst P, Bird A, Reinberg D . Analysis of the NuRD subunits reveals a histone deacetylase core complex and a connection with DNA methylation. Genes Dev 1999; 13: 1924–1935.
Saito M, Ishikawa F . The mCpG-binding domain of human MBD3 does not bind to mCpG but interacts with NuRD/Mi2 components HDAC1 and MTA2. J Biol Chem 2002; 277: 35434–35439.
Du Q, Luu PL, Stirzaker C, Clark SJ . Methyl-CpG-binding domain proteins: readers of the epigenome. Epigenomics 2015; 7: 1051–1073.
Boeke J, Ammerpohl O, Kegel S, Moehren U, Renkawitz R . The minimal repression domain of MBD2b overlaps with the methyl-CpG-binding domain and binds directly to Sin3A. J Biol Chem 2000; 275: 34963–34967.
Wade PA, Gegonne A, Jones PL, Ballestar E, Aubry F, Wolffe AP . Mi-2 complex couples DNA methylation to chromatin remodelling and histone deacetylation. Nat Genet 1999; 23: 62–66.
Ng HH, Jeppesen P, Bird A . Active repression of methylated genes by the chromosomal protein MBD1. Mol Cell Biol 2000; 20: 1394–1406.
Barr H, Hermann A, Berger J, Tsai HH, Adie K, Prokhortchouk A et al. Mbd2 contributes to DNA methylation-directed repression of the Xist gene. Mol Cell Biol 2007; 27: 3750–3757.
Stirzaker C, Song JZ, Davidson B, Clark SJ . Transcriptional gene silencing promotes DNA hypermethylation through a sequential change in chromatin modifications in cancer cells. Cancer Res 2004; 64: 3871–3877.
Pulukuri SM, Rao JS . CpG island promoter methylation and silencing of 14-3-3sigma gene expression in LNCaP and Tramp-C1 prostate cancer cell lines is associated with methyl-CpG-binding protein MBD2. Oncogene 2006; 25: 4559–4572.
Chatagnon A, Bougel S, Perriaud L, Lachuer J, Benhattar J, Dante R . Specific association between the methyl-CpG-binding domain protein 2 and the hypermethylated region of the human telomerase reverse transcriptase promoter in cancer cells. Carcinogenesis 2009; 30: 28–34.
Millar DS, Ow KK, Paul CL, Russell PJ, Molloy PL, Clark SJ . Detailed methylation analysis of the glutathione S-transferase pi (GSTP1) gene in prostate cancer. Oncogene 1999; 18: 1313–1324.
Song JZ, Stirzaker C, Harrison J, Melki JR, Clark SJ . Hypermethylation trigger of the glutathione-S-transferase gene (GSTP1) in prostate cancer cells. Oncogene 2002; 21: 1048–1061.
Hendrich B, Guy J, Ramsahoye B, Wilson VA, Bird A . Closely related proteins MBD2 and MBD3 play distinctive but interacting roles in mouse development. Genes Dev 2001; 15: 710–723.
Cai Y, Geutjes EJ, de Lint K, Roepman P, Bruurs L, Yu LR et al. The NuRD complex cooperates with DNMTs to maintain silencing of key colorectal tumor suppressor genes. Oncogene 2014; 33: 2157–2168.
Ernst J, Kellis M . ChromHMM: automating chromatin-state discovery and characterization. Nat Methods 2012; 9: 215–216.
Baubec T, Ivanek R, Lienert F, Schubeler D . Methylation-dependent and -independent genomic targeting principles of the MBD protein family. Cell 2013; 153: 480–492.
Ng HH, Zhang Y, Hendrich B, Johnson CA, Turner BM, Erdjument-Bromage H et al. MBD2 is a transcriptional repressor belonging to the MeCP1 histone deacetylase complex. Nat Genet 1999; 23: 58–61.
Stefanska B, Huang J, Bhattacharyya B, Suderman M, Hallett M, Han ZG et al. Definition of the landscape of promoter DNA hypomethylation in liver cancer. Cancer Res 2011; 71: 5891–5903.
Wang L, Liu Y, Han R, Beier UH, Thomas RM, Wells AD et al. Mbd2 promotes foxp3 demethylation and T-regulatory-cell function. Mol Cell Biol 2013; 33: 4106–4115.
Menafra R, Brinkman AB, Matarese F, Franci G, Bartels SJ, Nguyen L et al. Genome-wide binding of MBD2 reveals strong preference for highly methylated loci. PLoS One 2014; 9: e99603.
Gunther K, Rust M, Leers J, Boettger T, Scharfe M, Jarek M et al. Differential roles for MBD2 and MBD3 at methylated CpG islands, active promoters and binding to exon sequences. Nucleic Acids Res 2013; 41: 3010–3021.
Chatagnon A, Perriaud L, Nazaret N, Croze S, Benhattar J, Lachuer J et al. Preferential binding of the methyl-CpG binding domain protein 2 at methylated transcriptional start site regions. Epigenetics 2011; 6: 1295–1307.
Lopez-Serra L, Ballestar E, Ropero S, Setien F, Billard LM, Fraga MF et al. Unmasking of epigenetically silenced candidate tumor suppressor genes by removal of methyl-CpG-binding domain proteins. Oncogene 2008; 27: 3556–3566.
Mian OY, Wang SZ, Zhu SZ, Gnanapragasam MN, Graham L, Bear HD et al. Methyl-binding domain protein 2-dependent proliferation and survival of breast cancer cells. Mol Cancer Res 2011; 9: 1152–1162.
Baubec T, Schubeler D . Genomic patterns and context specific interpretation of DNA methylation. Curr Opin Genet Devel 2014; 25: 85–92.
Auriol E, Billard LM, Magdinier F, Dante R . Specific binding of the methyl binding domain protein 2 at the BRCA1-NBR2 locus. Nucleic Acids Res 2005; 33: 4243–4254.
Chatagnon A, Ballestar E, Esteller M, Dante R . A role for methyl-CpG binding domain protein 2 in the modulation of the estrogen response of pS2/TFF1 gene. PLoS One 2010; 5: e9665.
Lopez-Serra L, Ballestar E, Fraga MF, Alaminos M, Setien F, Esteller M . A profile of methyl-CpG binding domain protein occupancy of hypermethylated promoter CpG islands of tumor suppressor genes in human cancer. Cancer Res 2006; 66: 8342–8346.
Ballestar E, Esteller M . Methyl-CpG-binding proteins in cancer: blaming the DNA methylation messenger. Biochem Cell Biol 2005; 83: 374–384.
Berger J, Bird A . Role of MBD2 in gene regulation and tumorigenesis. Biochem Soc Trans 2005; 33: 1537–1540.
Fraga MF, Ballestar E, Montoya G, Taysavang P, Wade PA, Esteller M . The affinity of different MBD proteins for a specific methylated locus depends on their intrinsic binding properties. Nucleic Acids Res 2003; 31: 1765–1774.
Magdinier F, Wolffe AP . Selective association of the methyl-CpG binding protein MBD2 with the silent p14/p16 locus in human neoplasia. Proc Natl Acad Sci USA 2001; 98: 4990–4995.
Sansom OJ, Berger J, Bishop SM, Hendrich B, Bird A, Clarke AR . Deficiency of Mbd2 suppresses intestinal tumorigenesis. Nat Genet 2003; 34: 145–147.
Kalyuga M, Gallego-Ortega D, Lee HJ, Roden DL, Cowley MJ, Caldon CE et al. ELF5 suppresses estrogen sensitivity and underpins the acquisition of antiestrogen resistance in luminal breast cancer. PLoS Biol 2012; 10: e1001461.
Mohammed H, D'Santos C, Serandour AA, Ali HR, Brown GD, Atkins A et al. Endogenous purification reveals GREB1 as a key estrogen receptor regulatory factor. Cell Rep 2013; 3: 342–349.
Mohammed H, Taylor C, Brown GD, Papachristou EK, Carroll JS, D'Santos CS . Rapid immunoprecipitation mass spectrometry of endogenous proteins (RIME) for analysis of chromatin complexes. Nat Protoc 2016; 11: 316–326.
Coolen MW, Stirzaker C, Song JZ, Statham AL, Kassir Z, Moreno CS et al. Consolidation of the cancer genome into domains of repressive chromatin by long-range epigenetic silencing (LRES) reduces transcriptional plasticity. Nat Cell Biol 2010; 12: 235–246.
Bibikova M, Barnes B, Tsan C, Ho V, Klotzle B, Le JM et al. High density DNA methylation array with single CpG site resolution. Genomics 2011; 98: 288–295.
Du P, Zhang X, Huang CC, Jafari N, Kibbe WA, Hou L et al. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinformatics 2010; 11: 587.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015; 43: e47.
Ernst J, Kheradpour P, Mikkelsen TS, Shoresh N, Ward LD, Epstein CB et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 2011; 473: 43–49.
Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell 2015; 163: 1011–1025.
McLean CY, Bristor D, Hiller M, Clarke SL, Schaar BT, Lowe CB et al. GREAT improves functional interpretation of cis-regulatory regions. Nat Biotechnol 2010; 28: 495–501.
Huang, da W, Sherman BT, Lempicki RA . Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 2009; 37: 1–13.
Huang, da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature protocols 2009; 4: 44–57.
Acknowledgements
We thank Brigid O’Gorman for preparation of figures and reviewing the manuscript. Mouse fibroblast Mbd2−/− and wild-type cells were gratefully received as a gift from Adrian Bird. The results published here are in part based upon data generated by the TCGA Research Network: http://cancergenome.nih.gov/. We thank the Ramaciotti Centre, University of New South Wales (Sydney, Australia) for conducting the Affymetrix array experiments and the Australian Genome Research Facility for the HM450K arrays. This work is supported by National Health and Medical Research Council project grants (1029584 and 1088144) and NHMRC Fellowship to SJC (1063559) The contents of the published material are solely the responsibility of the administering institution and individual authors and do not reflect the views of the NHMRC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Stirzaker, C., Song, J., Ng, W. et al. Methyl-CpG-binding protein MBD2 plays a key role in maintenance and spread of DNA methylation at CpG islands and shores in cancer. Oncogene 36, 1328–1338 (2017). https://doi.org/10.1038/onc.2016.297
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.297
- Springer Nature Limited
This article is cited by
-
DiPRO1 distinctly reprograms muscle and mesenchymal cancer cells
EMBO Molecular Medicine (2024)
-
The roles of chromatin regulatory factors in endometriosis
Journal of Assisted Reproduction and Genetics (2024)
-
Computational discovery of novel inhibitory candidates targeting versatile transcriptional repressor MBD2
Journal of Molecular Modeling (2022)
-
Folic Acid Supplementation in Patients with Elevated Homocysteine Levels
Advances in Therapy (2020)
-
Integrated epigenomic analysis stratifies chromatin remodellers into distinct functional groups
Epigenetics & Chromatin (2019)