Abstract
Although modulation of the cellular tumor-suppressor p53 is considered to have the major role in E1A/E1B-55K-mediated tumorigenesis, other promyelocytic leukemia nuclear body (PML-NB)/PML oncogenic domain (POD)-associated factors including SUMO, Mre11, Daxx, as well as the integrity of these nuclear bodies contribute to the transformation process. However, the biochemical consequences and oncogenic alterations of PML-associated E1B-55K by SUMO-dependent PML-IV and PML-V interaction have so far remained elusive. We performed mutational analysis to define a PML interaction motif within the E1B-55K polypeptide. Our results showed that E1B-55K/PML binding is not required for p53, Mre11 and Daxx interaction. We also observed that E1B-55K lacking subnuclear PML localization because of either PML-IV or PML-V-binding deficiency was no longer capable of mediating E1B-55K-dependent SUMOylation of p53, inhibition of p53-mediated transactivation or efficiently transforming primary rodent cells. These results together with the observation that E1B-55K-dependent SUMOylation of p53 is required for efficient cell transformation, provides evidence for the idea that the SUMO ligase activity of the E1B-55K viral oncoprotein is intimately linked to its growth-promoting oncogenic activities.
Similar content being viewed by others
Introduction
Previous reports demonstrated that human adenovirus (HAdV) genomes preferentially target subnuclear host cell structures called promyelocytic leukemia nuclear bodies (PML-NBs) or PML oncogenic domains (PODs) during initial stages of infection.1, 2, 3 Intriguingly, PML-NBs exhibit ambivalent characteristics, either acting as an intracellular antiviral defense barrier4, 5, 6 or having an important role in posttranslational modification and modulation of cellular proteins (for example, p53, Daxx and Mre11) during productive viral infection and/or oncogenic transformation.7, 8, 9, 10 In fact, dozens of viruses have apparently evolved multiple mechanisms to counteract this cellular antiviral defense barrier or modulate PML/PML-associated proteins.11, 12, 13, 14
The PML protein itself was first described as the causal agent in acute promyelocytic leukemia, primarily as a fusion with the retinoic acid receptor alpha receptor generated by the chromosomal translocation t(15;17).15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26 Subsequently, it became evident that PML is a general tumor suppressor frequently deregulated in various tumor types,27 as well as being involved in multiple cellular pathways such as protein degradation,28 transcriptional regulation,29, 30 cellular senescence,31, 32, 33, 34 tumor suppression,35, 36 DNA repair,37, 38 apoptosis39, 40 and epigenetic regulation.41
In humans, at least seven PML isoforms (PML-I to PML-VII) are expressed by alternative splicing of a single pml gene.42, 43 Transcription of this gene is tightly controlled by interferons and p53.44, 45 All PML isoforms possess the same N-terminal RBCC motif, also known as the tripartite motif (TRIM), comprising the sequential organization of a RING-finger (R), two B-Boxes (B) and a coiled-coil domain (CC); hence these isoforms are grouped as TRIM19 in the TRIM protein family.42 This motif is particularly important for dimerization and localization to the PML-NBs.46 The C-terminal region of human PML shows remarkable variability among the isoforms and accounts for isoform-specific functions.47
In addition, PML function is modulated by SUMO posttranslational modifications,42, 48, 49, 50, 51, 52 as these regulate PML localization53 and nuclear body formation.50 Mammalian cells encode four different isoforms, SUMO-1, -2, -3 and -4, although it is not clear whether SUMO-4 is conjugated in vivo.54 Analogous to ubiquitin, all SUMO proteins are covalently conjugated to substrates via a three-step enzymatic pathway. Considering the impressive number of cellular processes regulated by this posttranslational modification, it is not surprising that many viruses target and exploit the host SUMOylation system.12, 13, 55
HAdV5, a well-analyzed member of the small DNA viruses, is frequently used to define basic molecular mechanisms in virus/host cell interactions to understand molecular properties of viral oncoproteins and basic principles of virus-induced tumorigenesis. HAdV5 E1A polypeptides mediate the most critical step in cell transformation and are sufficient to promote immortalization and partial transformation of primary rodent cells.56, 57 However, this unprogrammed cell proliferation induces growth arrest and apoptosis through stabilizing the p53 tumor-suppressor protein.58 Complete cell transformation requires HAdV5 E1B-55K, the Bcl2-like viral protein E1B-19K59, 60 or other cooperating oncogenes such as activated Ha-ras.61 E1B-55K-dependent inactivation of p53-mediated processes induces complete manifestation of a transformed phenotype9, 62, 63 by deregulating p53-dependent tumor-suppressive functions.64, 65, 66, 67 To do this, the viral polypeptide binds to the amino terminal transactivation domain of p53, blocks its ability to activate transcription of responsive promoters and sequesters the tumor suppressor into large cytoplasmic structures.64, 66, 68
Besides E1B-55K's role in repressing p53 functions, transformation requires additional anti-apoptotic activities of E1B-55K involving various other protein interactions, especially with PML-NB-associated factors such as Mre11 and Daxx.7, 10, 69 Interestingly, functional inactivation of the E1B-55K leucine-rich nuclear export sequence (NES) not only induces enhanced posttranslational modification of the adenoviral oncogene by SUMO at lysine 104 (SUMO-1 conjugation motif (SCM)) but also elevates transformation of primary rodent cells by mediating accumulation of p53 in subnuclear aggregates.70, 71 This augments the argument that loss of SUMO conjugation abrogates the ability of E1B-55K to transform in combination with E1A.71
Recently, Penella et al.9 reported that E1B-55K is a p53-SUMO1 E3 ligase that mediates SUMOylation of p53, inducing retention of the tumor-suppressor protein in PML-positive nuclear aggregates and thus repressing its functions. In that context, we have previously found that E1B-55K interacts with PML, more specifically with the PML isoforms IV and V.8
Here, we analyzed E1B-55K mutants to define a PML interaction motif within the E1B-55K polypeptide. Biochemical analysis revealed that abolished binding of E1B-55K to either PML-IV or PML-V does not affect p53, Mre11 and Daxx interactions, but does restrict efficient E1B-55K SUMO conjugation. Such E1B-55K variants that are unable to bind either PML-IV or PML-V isoforms lost subnuclear PML-NB localization and hence SUMO ligase activity acting on p53. In addition, PML-IV and PML-V non-binding mutants of the viral E1B-55K oncogene lost the ability to transform primary rodent cells together with E1A.
Results
C-terminal mutations in E1B-55K abolish efficient isoform-specific PML-V interaction
We recently showed that E1B-55K physically binds to PML isoforms IV and V in an SCM/K104-dependent manner, and that interaction with PML-IV promotes targeting of E1B-55K to PML-NBs.8 To examine this further, we transfected cells with plasmids encoding E1B-55K-wt and different variants of the viral protein (E1B-55K-SCM, E1B-55K-NES, E1B-55K-R443A and E1B-55K-RTR), the latter two affecting isoform-specific PML interactions (Figure 1a). We then screened them via immunoprecipitation analysis for their ability to interact with PML-IV and PML-V (Figure 1b). Consistent with previous results indicating that PML-IV/E1B-55K interactions depend on SUMO, significantly less PML-IV (but not PML-V) co-precipitated with E1B-55K-SCM (Figure 1b, lane 7), whereas E1B-55K-NES shows interaction with equal amounts of both PML isoforms (Figure 1b, lanes 8 and 9). Interestingly, E1B-55K-R443A shows a tendency to interact with higher amounts of PML-V than PML-IV (Figure 1b, lanes 11 and 12) and the C-terminal E1B-55K-RTR mutant was significantly impaired in PML-V precipitation (Figure 1b, lane 14). This indicates that in addition to the SCM of E1B-55K, the C-terminus of the viral oncoprotein has an essential role in PML isoform-specific association (Figure 1a). Interestingly, the pattern of relative PML co-precipitation by the E1B-55K variants illustrates that the two PML isoforms do not seem to require the same interaction region within the E1B-55K primary sequence (Figure 1a).
PML-IV/-V binding to E1B-55K oncoprotein. (a) The schematic overview shows the coding sequence of E1B-55K including the specific C-terminal mutations influencing isoform-specific PML interactions. Mutations are illustrated according to original aa, position and substituted aa, including abbreviated names in brackets. Amino-acid positions are indicated relative to the E1B-55K sequence and mutations are highlighted. C/H-rich region, cysteine/histidine-rich region. (b) Subconfluent H1299 cells were transfected with 10 μg of pE1B-55K variants plus 10 μg of different constructs encoding N-terminal flag-tagged human PML isoforms IV or V and harvested after 48 h before preparing total cell extracts. Immunoprecipitation was performed using mAB 2A6 (α-E1B-55K), resolved by 10% SDS–PAGE and visualized by immunoblotting. Co-precipitated proteins (top panel) and input levels of total cell lysates were detected using mAb 2A6 (α-E1B-55K), mAb flag-M2 (α-flag) and mAB AC-15 (α-β-actin). Molecular weights in kDa are indicated on the left, whereas corresponding proteins are labeled on the right.
PML-IV/-V interaction promotes efficient subnuclear targeting of E1B-55K to PML-NBs
To study whether PML-IV/-V-binding-deficient E1B-55K mutants interfere with the intracellular localization of the HAdV5 protein, plasmids encoding wild-type E1B-55K and mutant variants were transfected into human cells and subjected to indirect immunofluorescence analysis. Consistent with previous findings, wild-type E1B-55K protein localizes in the cytoplasm, mostly concentrated in a perinuclear body in all cells examined (n>50; Figure 2, panel a).68, 72, 73, 74, 75, 76 In nearly 90% of the cells, localization of E1B-55K-NES mutant protein differ from wild-type E1B-55K, and numerous E1B-55K containing aggregates colocalizing with PML bodies were distributed throughout the nucleus (Figure 2, panel c). In addition, the observed intracellular distribution of the E1B-55K-R443A polypeptide is similar to the wild-type localization (Figure 2, panel i). However, E1B-55K proteins that poorly interact with either PML-IV (E1B-55K-SCM; Figure 2, panel e) or PML-V (E1B-55K-RTR; Figure 2, panel g) are located exclusively in the cytoplasm forming several aggregates.
PML-IV/-V-binding targets E1B-55K to PML bodies. H1299 cells were transfected with constructs encoding E1B-55K variants, sometimes treated with +LMB to block E1B-55K nuclear export, and fixed with paraformaldehyde after 24 h, before labeling with mouse mab 2A6 (E1B-55K) and PML rabbit pAb NB100–59787. Primary Abs were detected with Alexa Fluor 594- and Alexa Fluor 488-conjugated secondary Abs. Representative α-E1B-55K (red) and α-PML (green) staining patterns of at least 50 analyzed cells and overlays of the single images (merge; a–j) are shown. E1B-55K and PML-positive nuclear accumulations are indicated by white arrows.
Previous experiments have shown that after blocking the CRM1-dependent nuclear export of E1B-55K by treatment with leptomycin B (LMB), the viral protein accumulates within dense nuclear aggregates.75, 77 We observed in almost all cells (90%) accumulation of E1B-55K wild-type (Figure 2, panel b) and PML-binding E1B-55K variants (E1B-55K-NES, Figure 2, panel d; E1B-55K-R443A, Figure 2, panel j) in nuclear aggregates, colocalizing with PML nuclear bodies. In contrast, PML-binding-deficient mutants E1B-55K-SCM (Figure 2, panel f) and E1B-55K-RTR (Figure 2, panel h) lost the ability to localize to PML-NBs, when artificially restricted to the cell nucleus.
E1B-55K interaction with PML-IV/-V is not required for p53, Mre11 and Daxx binding
As cellular transformation of primary rodent cells by the adenoviral oncogenes E1A and E1B-55K is a complex multi-step process predominantly involving modulation of the cellular tumor-suppressor protein p53, we first tested both PML-binding-deficient E1B-55K mutants for their ability to interact with p53 (Figure 3a) in plasmid-transfected human p53-negative cells. In agreement with previously published data, immunoprecipitation of exogenous p53 resulted in appropriate E1B-55K and p53 signals (Figure 3a, lanes 3–7). In summary, PML binding seems not to be a prerequisite for p53 binding.
PML-IV/-V-binding E1B-55K induces p53 SUMO conjugation. Subconfluent H1299 cells were transfected with 5 μg of pE1B-55K variants plus 3 μg of human p53 (pc53SN3) and harvested after 48 h before preparing total cell extracts. Following immunoprecipitation of p53 using mAb DO-I (α-p53), proteins were resolved by 10% SDS–PAGE and visualized by immunoblotting. Co-precipitated proteins (a) and input levels (b) of total cell lysates were detected using mAb 2A6 (α-E1B-55K), mAb DO-I (α-p53) and mAB AC-15 (α-β-actin). Molecular weights in kDa are indicated on the left, whereas corresponding proteins are labeled on the right. (c) Graphs showing the relative amount of SUMOylated E1B-55K fraction normalized to unmodified E1B-55K protein (left panel) and the relative amount of SUMOylated p53 fraction normalized to unmodified E1B-55K protein (right panel). Data were obtained from Figure 3a.
Next, the E1B-55K mutants were tested for their ability to interact with already known E1B-55K-associated proteins reportedly having a role in mediating E1B-55K oncogenicity. As expected, immunoprecipitation of Mre11 or Daxx resulted in co-precipitation of E1B-55K protein (Figure 4, lanes 2–6). Taken together, our observations indicated that E1B-55K mutants that lost binding to either PML-IV or PML-V do not restrict Mre11 or Daxx interaction with the viral polypeptide in transfected human cells.
PML-IV/-V does not affect E1B-55K binding to p53, Mre11 and Daxx. H1299 cells were transfected with 5 μg of pE1B-55K variants and harvested after 48 h before preparing total cell extracts. After immunoprecipitation of endogenous Mre11 and Daxx using Mre11 rabbit pAb pNB 100–142 and rabbit pAb α-Daxx, proteins were resolved by 10% SDS–PAGE and visualized by immunoblotting. Co-precipitated proteins (left panel) and input levels (right panel) of total cell lysates were detected using mAb 2A6 (α-E1B-55K), Mre11 rabbit pAb pNB 100–142, rabbit pAb α-Daxx and mAB AC-15 (α-β-actin). Molecular weights in kDa are indicated on the left, whereas corresponding proteins are labeled on the right.
PML-IV/-V interaction is necessary for SUMOylation of p53
Recently, p53 SUMOylation was shown to be regulated by adenoviral E1B-55K,78 revealing a p53-SUMO1 E3 ligase function for E1B-55K that mediates SUMOylation of p53 and induces functional inhibition of this pro-apoptotic cellular regulator.9 In agreement, we demonstrate here that p53 shows strong SUMOylation after coexpression of E1B-55K wt and E1B-55K-NES (Figure 3b, right panel, lanes 3 and 5). As expected, E1B-55K-SCM showed drastically reduced SUMO-modified p53 protein (Figure 3b, right panel, lane 4). Intriguingly, E1B-55K-RTR, which failed to bind PML-V, was also unable to promote p53 SUMOylation, because we could not detect SUMO-conjugated p53 moieties (Figure 3b, right panel, lane 7). E1B-55K-R443A was able to maintain SUMO ligase activity toward p53 (Figure 3a, right panel, lane 6). This indicates that although the binding of R443A to PML-IV is impaired (Figure 1b, lane 11), it is adequate enough for efficient PML-NB localization (Figure 2, panel j) and E1B-55K-R443A SUMO ligase capacity toward p53. We also observed less SUMOylated E1B-55K-RTR compared with the wt viral protein or other E1B-55K-PML interacting variants (E1B-55K-NES, E1B-55K-R443A). This supports the idea that besides E1B-55K/PML-IV binding, the viral oncoprotein should interact with PML-V for subnuclear PML-NB localization and posttranslational SUMO modification of E1B-55K, and subsequent efficient SUMOylation of p53. These results are fascinating because interaction with specific PML isoforms has until now not been linked to SUMOylation of the adenoviral oncoprotein.
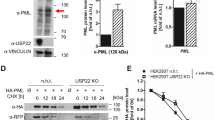
Finally, to further confirm a direct correlation between PML-IV/-V binding and E1B-55K SUMOylation, we performed SUMOylation analysis in vitro. We used Escherichia coli-produced, purified human SUMO2, SAE1/SAE2 (human SUMO E1), ubc9 (human SUMO E2) and in vitro translated E1B-55K variants. Addition of ubc9 stimulated ligation of SUMO2 to E1B-55K (Figure 5a). As anticipated, E1B-55K-SCM's mutation in the SCM completely abolished posttranslational modification with SUMO2 moieties (Figure 5a). Intriguingly, E1B-55K-RTR, which was not SUMOylated in vivo, showed poly-SUMO chains attached to the viral protein in vitro. This suggests that the inability of the PML-V-binding-deficient mutant E1B-55K-RTR to localize with PML-NBs is what prevents SUMO conjugation of the viral oncoprotein in vivo, not some intrinsic property of the protein, because this mutant can still be posttranslationally modified with SUMO protein as shown in vitro (Figure 5a, lane 9). Thus, E1B-55K is apparently SUMOylated because of specifically interacting with a PML isoform and hence localizing with PML-NBs.
PML-NB association is a prerequisite for E1B-55K SUMO conjugation. (a) E1B-55K wt and mutants were produced using the wheat germ in vitro translation system (TNT) as described in Materials and methods section. The reactions were incubated 4 h at 37 °C, then stopped by adding 2x SDS buffer, boiled and separated by SDS–PAGE. 35S-labeled proteins were visualized by autoradiography. (b) Subconfluent H1299 cells were transfected with 5 μg of pE1B-55K variants and harvested after 48 h before preparing total cell extracts. Immunoprecipitation of endogenous Ubc9 or PML was performed using mAb α-Ubc9 or rabbit pAb α-PML, resolved by 10% SDS–PAGE and visualized by immunoblotting. Co-precipitated proteins (left panel) and input levels (right panel) of total cell lysates were detected using mAb 2A6 (α-E1B-55K), mAb α-Ubc9, rabbit pAb α-PML and mAB AC-15 (α-β-actin). Molecular weights in kDa are indicated on the left, whereas corresponding proteins are labeled on the right.
To exclude any variations in ubc9 (human SUMO E2) association with E1B-55K, we validated binding of the viral proteins to the human SUMO E2 enzyme in plasmid-transfected human p53-negative H1299 cells (Figure 5b); immunoprecipitation of ubc9 resulted in signals for all E1B-55K variant proteins (Figure 5b, upper panel). However, less E1B-55K-SCM mutant viral protein was co-immunoprecipitated (Figure 5b, lane 3, upper panel). In addition, we also observed interaction of all mutants with PML co-immunoprecipitated with an endogenous PML antibody recognizing both PML-IV and PML-V (Figure 5b, lower panel).
PML isoform binding of E1B-55K is essential for p53 subcellular localization
To study whether PML-IV/-V-binding-deficient E1B-55K mutants interfere with the intracellular localization of the viral protein, plasmids encoding wild-type E1B-55K and mutant variants were transfected into human cells together with human p53 (Figure 6). Consistent with the findings in Figure 2, E1B-55K was mainly detected in the cytoplasm, mostly concentrated in a perinuclear body (n>50; Figure 6, panel a). E1B-55K localization in these subnuclear aggregates, and thus sequestering p53 (Figure 6, panel a), is known to be directly linked to the transforming potential of the viral oncogene in combination with E1A.68, 70, 71
PML-IV/-V-binding affects E1B-55K-dependent p53 subcellular localization. H1299 cells were transfected as described in Figure 2 before labeling with mouse mab 2A6 (E1B-55K) and DO-1 (α-p53). Primary Abs were detected with Alexa Fluor 594- and Alexa Fluor 488-conjugated secondary Abs. Representative α-E1B-55K (red) and α-p53 (green) staining patterns from at least 50 analyzed cells and overlays of the single images (merge; a–j) are shown. E1B-55K and p53-positive nuclear accumulations are indicated by white arrows.
After blocking the CRM1-dependent nuclear export of E1B-55K with +LMB, the wt viral protein accumulates with the cellular interaction partner p53 in dense nuclear aggregates (Figure 6, panel b).75, 77 Similarly, nearly 80% of the cells (n>50) transfected with E1B-NES mutant protein defective in efficient cytoplasmic export also accumulated numerous E1B-55K/p53 aggregates distributed throughout the nucleus in the absence and presence of LMB (Figure 6, panels c and d). In contrast, we observed an exclusively cytoplasmic localization pattern of both the E1B-55K-SCM and E1B-55K-RTR mutants, in the absence or presence of LMB, whereas p53 is mostly diffusely distributed in the nucleus (Figure 6, panels e, j, g and h). However, these two PML-binding-deficient E1B-55K variants did sequester some p53 into random cytoplasmic aggregates (Figure 6, panels e, g, h and j). In the presence of LMB, which blocks nuclear export in almost all cells (90%), we observed no colocalization of p53 with E1B-55K-SCM and E1B-55K-RTR in nuclear aggregates (Figure 6, panels h and f). Taken together, one can conclude that inactivating the E1B-55K/PML interaction abolishes the capacity of the viral oncogene to efficiently or specifically relocalize human p53.
PML-IV/-V interaction promotes p53 repression by the viral oncoprotein E1B-55K
Given the above findings, we performed a transient reporter gene assay to evaluate the PML-binding-deficient E1B-55K variants' ability to repress p53-mediated transcriptional activation (Figure 7a). H1299 cells were transfected with plasmids encoding E1B-55K-wt, E1B-55K-SCM, E1B-55K-NES and E1B-55K-RTR plus a p53 expression plasmid (pC53SN3) and p53-independent, as well as reporter p53-dependent luciferase constructs for a normalized system read-out. The reporter construct contains five artificial p53-binding sites (pRE-Luc) and therefore Firefly luciferase expression directly correlates with transcriptional activation by p53, which in turn is directly influenced by the adenoviral E1B-55K protein.66, 67, 79
PML-IV/-V interaction promotes p53 repression by E1B-55K in transiently transfected H1299 cells. (a) Subconfluent p53-negative H1299 cells were transfected with 1.0 μg of pRenilla-Luc, 0.25 μg of pRE-Luc, 0.02 μg of pc53SN3, pE1B-55K-wt/-SCM/-NES/-RTR plus empty vector pcDNA3 to a total amount of 2.5 μg DNA. The amount of transfected E1B-55K expression plasmid is shown in μg. The cells were harvested after 48 h according to the manufacturer's instructions using the Dual-Luciferase Reporter Assay System (Promega) and subsequently measured in a Lumat LB 9507 luminometer (Berthold Technologies, Bad Wildbad, Germany). Luciferase activity was determined by correlating p53-dependent Firefly luciferase activity with p53-independent Renilla luciferase activity to account for transfection efficiency. The luciferase activities were plotted relative to the positive control (mix 3) and are based on the average of three independent experiments. (b) Model of the interplay between E1B-55K and p53-dependent transcriptional repression.
As expected, neither transfection of pRenilla-Luc (Figure 7a, lane 1) nor pRenilla-Luc plus pRE-Luc (Figure 7a, lane 2) shows detectable luminescence because of the lack of endogenous p53 expression in H1299. However, additional co-transfection of pC53SN3 led to maximal activation of p53-dependent luciferase transcription/activity (Figure 7a, lane 3). Gradual addition of E1B-55K-wt drives a progressive drop in luciferase activity to a residual level of ~10% of the positive control (Figure 7a, lanes 4 to 7). Consistent with previous results,80 it is interesting that relatively low amounts of E1B-55K (1.0–3.0 μg) are sufficient to nearly completely repress p53 transcriptional activation.
As anticipated, SUMO-less pE1B-55K-SCM coexpression fails to efficiently repress luciferase activity (Figure 7a, lanes 8 and 9), whereas hyper-SUMOylated E1B-55K-NES represses p53 transactivation even more efficiently than the wt viral oncoprotein (Figure 7a, lanes 10 and 11).8, 70, 81 Intriguingly, PML-IV/-V binding seems to be directly linked to the p53 repression abilities of the different E1B-55K mutants, because E1B-55K-RTR (PML-V-binding deficiency) did not efficiently repress p53 transcriptional activation as E1B-55K-SCM (Figure 7a, lanes 12 and 13; schematically depicted in Figure 7b).
E1B-55K association with PML-IV/-V is essential for efficient transforming potential in combination with E1A
To more closely evaluate whether modulating endogenous PML-IV/-V is necessary for the transformation potential of E1B-55K, freshly prepared primary baby rat kidney cells were transfected simultaneously with plasmids encoding E1A and the newly identified PML-binding mutants. After incubation under standard conditions, focus formation was visualized by crystal violet staining and quantified relative to E1A plus E1B-55K-wt (Figure 8a). As expected, expression of E1A alone resulted in low focus formation activity (Figure 8a, lane 2), whereas simultaneous co-transduction with E1B-55K-wt significantly enhanced cellular transformation in combination with E1A (Figure 8a, mix 3). Interestingly, all combinations containing the PML-binding mutants pE1B-55K-SCM and pE1B-55K-RTR (Figure 8a, lanes 4 and 6) show a significant reduction in focus-forming activity compared with the positive control, suggesting that PML-IV/-V interaction with E1B-55K is important for efficient cellular transformation.
E1B-55K binding to PML-IV/-V promotes efficient transformation in combination with E1A. Freshly prepared primary (a) baby rat kidney (pBRK) and (b) baby mouse kidney cells (BMK/PML +/+, PML −/−) were simultaneously transfected with a combination of two plasmids encoding E1A and the individual E1B-55K variants. Cells were split after 48 h of absorption and cultivated for 6–8 weeks under standard conditions. Focus formation was visualized by crystal violet staining and absolute numbers were counted. Representative results normalized to the positive control (pE1A plus pE1B-55K-wt) were plotted based on the average of three independent experiments. (c) Model of the interplay between E1B-55K and the functional inhibition of the tumor-suppressor p53 mediated by the viral oncoprotein.
To validate this hypothesis in another rodent system, we also performed transformation assays in primary baby mouse kidney cells with a stable genetic knock-out of the PML gene (PML −/−).82 Consistent with the rat model system, we observed significantly lower cellular transformation levels with E1B-55K mutants defective in PML-IV/-V binding in combination with E1A (Figure 8b, lanes 4 and 6).
Strikingly, the focus-forming activity of all E1B-55K variants in combination with E1A was drastically reduced in primary rodent cells genetically depleted for PML expression, implying that E1B-55K/PML cooperation is a prerequisite for efficient cell transformation (Figure 8b). As our results above show that E1B-dependent SUMOylation of p53 also depends on PML-IV/-V interactions and is required for efficient cell transformation, this supports the idea that the SUMO ligase activity of E1B-55K is intimately linked to its growth-promoting oncogenic activities (Figure 8c).
Discussion
Our results show that E1B-55K derivatives defective in binding either PML-IV or PML-V, and hence unable to localize with subnuclear PML, are no longer capable of mediating E1B-55K-dependent SUMOylation of p53. These mutated E1B-55K proteins are also defective in inhibiting p53-mediated transactivation and have greatly reduced transforming potential in primary rodent cells.
HAdVs encode several multifunctional proteins that initiate transformation of primary rodent cells.83, 84, 85, 86 The oncogenic determinants of this viral model system are located within E1 and E4 region of the HAdV genome. The E1A and E1B proteins are absolutely required for oncogenic transformation,83, 84, 85, 86 because they simultanously accelerate cell cycle progression87 and inhibit tumor-suppressor proteins such as p53.66, 67, 79
As the initial description of adenoviral tumorigenicity88 it is still unclear why HAdVs are oncogenic in rodents, but not as far as we know, in humans. With very few exceptions,89, 90, 91, 92, 93, 94, 95, 96 attempts to transform primary human cells in culture using different types of HAdV subgenomic DNA fragments have proved extremely inefficient. In contrast, primary rat, hamster or mouse cells can be transformed efficiently.83, 84, 97, 98 Although purely speculative at this point, species-specific expression patterns of PML isoforms IV and V in humans and their orthologues in rodent tissue may represent the primate-specific factor as a prerequisite for efficient E1A/E1B-55K-mediated oncogenesis that mainly occurs in rodent cells, unlike cells of human origin. Although it is difficult to draw conclusions as yet, because no data are available on the expression of PML in rats, recent data on murine pml locus organization underline the possibility of species-specific PML expression with partially homologous, but nevertheless considerably different PML isoforms.52, 99 Moreover, as PML expression patterns also differ considerably because of extensive regulation in different mammalian tissues, oncogenic transformation may only occur in a specific subset of human cells.
Obviously, the cellular PML protein represents a key regulator of multiple cellular functions,100 because it has important implications in tumorigenesis.35, 36 It was described as a general tumor-suppressor protein frequently deregulated in human tumor types: 17% of colon adenocarcinomas, 21% of lung tumors, 27% of prostate adenocarcinomas, 31% of breast adenocarcinomas, 49% of CNS tumors (100% medulloblastomas, >90% oligodendroglial tumors), 49% of germ cell tumors and 68% of non-Hodgkin’s lymphomas. In our study, we demonstrate that cellular transformation by adenoviral oncoproteins E1B-55K and E1A orchestrates a novel pathway, as the data clearly show that PML expression (PML-IV, PML-V) is essential for sustaining the oncogenic potential of E1B-55K. Presumably, the anti-oncogenic functions of PML involve secondary effects of PML-NBs as sites of a huge PML-regulated network with over 150 different proteins involved in over 750 separate interactions and various biological processes.100
However, most of the molecular mechanisms, especially selective regulation of proteins by isoforms of PML (seven major isoforms so far), and any correlation with human tumorigenesis in different tissue types are still unknown. There are rare descriptions about the different functions of each isoform, or the molecular mechanisms of how they fulfill different functions despite close sequence similarities. In principle, the functions of all PML isoforms may differ because of the biochemical behavior of the isoform and the number/kind of specific cellular interaction partners. In addition, PML-NBs show high subnuclear mobility, coupled with continuous exchange of PML isoforms and associated components. Interestingly, PML-I/-II/-III/-IV/-VI exhibit a relatively high exchange rate, whereas the exchange rate for PML-V takes nearly an hour. Thus, it is assumed that PML-V represents the major scaffold protein of PML-NBs.101, 102, 103 Within our study, we showed for the first time that PML-IV and PML-V expression is highly implicated in oncogenic transformation of primary mammalian cells by adenoviral proteins E1A and E1B-55K.
Inconsistently, many reports illustrate the importance of PML deregulation and/or loss during tumorigenesis.27, 35, 36, 104, 105 On the contrary, PML is obviously stabilized by simultaneous expression of viral protein E1A,106 as well as E1B-55K in stably HAdV transformed rodent cells,8 implying some function during adenoviral cell transformation. Similarly, p53 is stabilized by E1A,58, 107, 108, 109, 110, 111, 112 whereas the pro-tumorigenic functions of adenoviral E1B-55K are primarily linked to repression of the tumor-suppressor p53. Subsequent direct interactions,64, 65 transcriptional repression 66, 67, 79 and nuclear-cytoplasmic relocalization70, 71 induce the complete silencing of p53-dependent tumor-suppressive functions.
Here, we show that the oncogenic functions of E1B-55K are tightly linked to binding of particular PML isoforms and efficient recruitment to PML-NBs. SUMOylation of E1B-55K at PML-NBs is a prerequisite for E1B-dependent modulation of p53, and thus closely linked to the growth-promoting activities of this viral oncoprotein. Although clearly speculative at this point, it appears that some of the other factors such as TERT and TRF1 described to interact with PML-IV are implicated in telomere maintenance. This then may represent one of the previously suggested alternative pathways of E1B-55K-mediated cellular transformation.7
Neither PML-IV nor PML-V harbor identical protein sequences that are unique compared with the remaining isoforms, thereby ruling out a specific E1B-55K-binding region. Both harbor the exons 1–6, which are present in all PML proteins except PML-VII, exon 7a, which again is also present in PML-I/-II/-III and their own unique C-termini. Interestingly, PML-IV is characterized by the unique amino-acid sequence of exon 8b and although sequence analysis has so far identified no defined domain of highly similar sequence motifs, this exon may contain an essential determinant for SUMOylation-dependent E1B-55K PML-IV interaction.
Conversely, efficient interaction with PML-V requires a highly conserved motif in the C-terminal region within E1B-55K. The sequence organization of this part of the viral oncoprotein seems to resemble some degree of organized tertiary structure comparable to a zinc-finger and/or RING domain. However, owing to the immense number of PML-NB-associated proteins100 and interactions of E1A and E1B-55K with various cellular components, it is also important to consider that specific binding of certain PML isoforms possibly occurs indirectly.
Although the molecular mechanisms have so far remained elusive, it appears that posttranslational modification of the viral E1B-55K protein is an essential determinant in the pro-tumorigenic functions of E1B-55K through modulating p53. Indeed, functional inactivation of the E1B-55K-NES induces enhanced posttranslational modification of E1B-55K by SUMO at lysine 104, in addition to transcriptional repression of the tumor-suppressor protein p53 and augmented transformation of primary rodent cells through the accumulation of p53, E1B-55K and PML in subnuclear aggregates.70, 71, 113
In contrast, mutational inactivation of the SCM domain abrogates E1B-55K SUMOylation, PML-IV interaction, p53 transcriptional repression and the oncogenic potential of E1B-55K in combination with E1A.78, 114 Certainly, inactivation of PML-V binding to E1B-55K adds another layer of complexity to adenoviral-mediated p53 modulation; such a mutation also abrogates E1B-55K SUMOylation, p53 inhibition and the transforming capacity of E1B-55K in combination with E1A.78
p53 itself is SUMOylated at an apparently unique C-terminal lysine residue (K386), resulting in modulated transcriptional activity in vivo.115, 116, 117, 118, 119 Although the modulation of p53 activity by SUMO conjugation remains controversial, this observation again underlines the difficulty in assigning particular functions to SUMOylation. However, simultaneous expression of E1B-55K and p53 leads to enhanced posttranslational modification of the tumor-suppressor protein in an E1B-55K-dependent manner, and here SUMO modification of E1B-55K itself is absolutely essential.9, 78
Besides the hypothesis that E1B-55K serves as an E3 SUMO ligase,9 a so far unknown factor may also facilitate p53 SUMOylation in the presence of SUMOylated E1B-55K. In this context, our results in general offer a new perspective to not only unravel the complexity of E1B-55K/PML cooperation, but also HAdV-mediated transcriptional repression of the tumor-suppressor protein p53.66, 67, 79 In addition, E1B-55K's oncogenic potential may also involve the proper localization of E1B-55K in subnuclear aggregates sequestering p53 and PML, which is regulated at least in part by the viral protein's binding capacity to PML-IV and PML-V.
In summary, a growing body of evidence suggests that repression of p53 by E1B-55K in isolation during transient transfection, involves additional determinants such as posttranslational modification, which mainly occur at PML-NBs.31, 70, 71, 113, 120, 121, 122, 123, 124 E1B-55K exploits certain PML isoforms and associated proteins to mediate its multiple functions during oncogenic processes, in particular conditional transcriptional repression of p53 (refs 65, 125, 126, 127, 128, 129, 130, 131, 132, 133) and control proteins involved in transcriptional regulation, tumor suppression, DNA repair and apoptosis.6, 7, 10, 69
Our study provides novel insight into the molecular mechanism regulating subnuclear localization of E1B-55K via PML-IV/PML-V interactions and PML-NB recruitment. Furthermore, isoform-specific interaction of E1B-55K and/or the remaining adenoviral oncoproteins represents a general mechanism of HAdV5 to gain access to the PML regulatory network and modulate the functions of specific PML-associated proteins. Identifying new interaction partners of the adenoviral oncoproteins may not only provide insights into the fundamental differences between human and rodent cells, but also into the basic principles of tumorigenesis, and potentially illuminate oncogenic processes in human tissue.
Materials and methods
Cell lines and culture conditions
H1299,134 HEK29389 and primary baby rat kidney cells135 were grown in Dulbecco’s modified Eagle’s medium supplemented with 10% fetal calf serum, 100 U of penicillin, 100 μg of streptomycin per ml in a 5% CO2 atmosphere at 37 °C.
Plasmids, transient transfections and reporter gene assays
HAdV5 E1B-55K-wt, the E1B mutant proteins E1B-55K-NES (L83/87/91 A), E1B-55K-SCM (K104R),71 E1B-55K-R443A (R443A),10 E1B-55K-RTR (RTR448/449/450AAA)10 and human p53136 were expressed from pcDNA3. Subconfluent cells were transfected using 25 kDa polyethylenimin (Polysciences, Eppelheim, Germany) as described before.8 For luciferase assays, H1299 cells were transfected with indicated amounts of reporter and effector plasmids (see figure legend) and 1.0 μg of pRL-TK (Promega, Madison, WI, USA) expressing Renilla luciferase under control of the HSV-TK promoter. The p53-dependent (pRE-Luc) reporter constructs for expressing Firefly luciferase and p53 have been described previously.71, 137 Total cell extracts were prepared 36 h after transfection and Firefly activity was assayed in an automated luminometer measuring relative light units (luminescence) using Dual-Luciferase Reporter Assay (Promega). All samples were normalized for transfection efficiency by measuring Renilla activity.
Transformation assays
To test the oncogenic potential of E1B-55K in combination with E1A, freshly prepared primary baby rat kidney or baby mouse kidney cells were prepared from 3 to 5-day-old CD rats (Charles River, Wilmington, MA, USA)137 or mice.82 Subsequently, primary cells were transfected with the respective plasmid138 or altered versions with the respective mutations in the E1B-55K. Cells were expanded after 48 h, cultivated for 4–6 weeks changing the medium every 3–4 days, and multilayered cell accumulations (foci) were visualized by crystal violet staining.
Antibodies and protein analysis
Primary Abs for HAdV5 proteins included E1B-55K mouse mAb 2A6.139 Primary Abs for detecting of cellular and etopically expressed proteins used, included p53 mouse mAb DO-I (Sigma-Aldrich, Inc., St Louis, MO, USA), PML rabbit pAb NB100–59787 (Novus Biologicals, Inc., Littleton, CO, USA), Mre11 rabbit polyclonal antibody pNB 100–142 (Novus Biologicals, Inc.), PML rabbit pAb NB100–59787 (Novus Biologicals, Inc.), mAb α-Ubc9 (BD Transduction, Heidelberg, Germany), rabbit pAb α-Daxx (Millipore, Darmstadt, Germany), ß-actin mouse mAb AC-15 (Sigma-Aldrich, Inc.) and mouse mAb Flag-M2 (Sigma-Aldrich, Inc.). All protein extracts were prepared in RIPA buffer (50 mM Tris-HCl/pH 8.0, 150 mM NaCl, 5 mM EDTA, 1 mM DTT, 0.1% SDS, 1% NP-40, 0.1% Triton X-100, 0.5% sodium deoxycholate) containing 1% (v/v) phenylmethylsulfonyl fluoride, 0.1% (v/v) aprotinin, 1 μg/ml leupeptin, 1 μg/ml pepstatin, 1 mM DTT, 25 mM iodacetamide and 25 mM N-ethylmaleimide. For IP, proteinA-sepharose was coupled with primary Ab for 1 h at 4 °C. The Ab-coupled beads were added to precleared extracts and rotated for 2 h at 4 °C. Proteins bound to the beads were precipitated by centrifugation, washed three times and boiled for 3 min at 95 °C in 2xLaemmli buffer. For immunoblotting of protein, input levels/IP solutions equal amounts were separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE), transferred to nitrocellulose (Schleicher & Schüll/Whatman, Freiburg, Germany) and incubated as described previously.106 Bands were visualized by ECL as recommended by the manufacturer (Pierce, Rockford, IL, USA) on X-ray films (CEA RP, Hamburg, Germany; medical X-ray film). Autoradiograms were scanned, cropped and prepared using Adobe PhotoshopCS6 and Adobe IllustratorCS6.
Indirect immunofluorescence
Cells grown on glass coverslips in six-well dishes were transfected for 24 h with up to 1 μg of plasmid DNAs expressing E1B-55K or mutants. Some wells were treated with 20 nM LMB for 6 h. Cells were then fixed for 15 min with 4% paraformaldehyde (Electron Microscopy Science, Hatfield, PA, USA), and permeabilized for 15 min with 0.5% Triton X-100 in phosphate-buffered saline. Following washes in phosphate-buffered saline, immunofluorescence was performed with appropriate antibodies for 2 h at room temperature in a humidity chamber. The secondary antibodies used were antibodies conjugated to Alexa 594 and Alexa 488 dyes (Molecular Probes, Darmstadt, Germany). 4,6-Diamidino-2-phenylindole dye was added with the secondary antibodies. Coverslips were washed and mounted in Immu-mount (Thermo Scientific, Darmstadt, Germany). In all cases, we have analyzed cells expressing low-to-intermediate levels of protein to avoid potential localization artifacts related to overexpression. Images were obtained using an LSM4 confocal microscope (Zeiss, Jena, Germany) with a 63x objective using the Zen 2011 Image browser software (Jena, Germany). Images were cropped using Adobe PhotoshopCS6 and assembled with Adobe IllustratorCS6.
In vitro translation and in vitro SUMOylation
E1B-55K wt and mutants were produced using the wheat germ in vitro translation system (TNT, Promega) and 5 μCi L-[35S]methionine (43 TBq/mMol, 10 μCi/μl; Perkin-Elmer, Waltham, MA, USA) according to the supplier's protocol. In vitro SUMOylation reactions contained 1.5 μl 35S-labeled substrate, 1.5 μg SUMO-1, 10 ng Ubc9, 100 ng SAE1/2, 50 mM Tris-HCl, pH 7.6, 5 mM MgCl2, 2 mM ATP, 10 mM creatine phosphate, 3.5 U/ml creatine kinase and 0.6 U/ml inorganic pyrophosphatase. All in vitro SUMOylation reagents were kindly provided by Ron Hay. The reactions were incubated 4 h at 37 °C, then stopped by adding 2x SDS buffer, boiled and separated by SDS–PAGE. 35S-labeled proteins were visualized by autoradiography.
References
Carvalho T, Seeler JS, Öhman K, Jordan P, Pettersson U, Akusjärvi G et al. Targeting of adenovirus E1A and E4-ORF3 proteins to nuclear matrix-associated PML bodies. J Cell Biol 1995; 131: 45–56.
Maul GG . Nuclear domain 10, the site of DNA virus transcription and replication. Bioessays 1998; 20: 660–667.
Ishov AM, Maul GG . The periphery of nuclear domain 10 (ND10) as site of DNA virus deposition. J Cell Biol 1996; 134: 815–826.
Ullman AJ, Hearing P . Cellular proteins PML and Daxx mediate an innate antiviral defense antagonized by the adenovirus E4 ORF3 protein. J Virol 2008; 82: 7325–7335.
Ullman AJ, Reich NC, Hearing P . Adenovirus E4 ORF3 protein inhibits the interferon-mediated antiviral response. J Virol 2007; 81: 4744–4752.
Schreiner S, Wimmer P, Sirma H, Everett RD, Blanchette P, Groitl P et al. Proteasome-dependent degradation of Daxx by the viral E1B-55K protein in human adenovirus-infected cells. J Virol 2010; 84: 7029–7038.
Hartl B, Zeller T, Blanchette P, Kremmer E, Dobner T . Adenovirus type 5 early region 1B 55-kDa oncoprotein can promote cell transformation by a mechanism independent from blocking p53-activated transcription. Oncogene 2008; 27: 3673–3684.
Wimmer P, Schreiner S, Everett RD, Sirma H, Groitl P, Dobner T . SUMO modification of E1B-55K oncoprotein regulates isoform-specific binding to the tumour suppressor protein PML. Oncogene 2010; 29: 5511–5522.
Pennella MA, Liu Y, Woo JL, Kim CA, Berk AJ . Adenovirus E1B 55-kilodalton protein is a p53-SUMO1 E3 ligase that represses p53 and stimulates its nuclear export through interactions with promyelocytic leukemia nuclear bodies. J Virol 2010; 84: 12210–12225.
Schreiner S, Wimmer P, Groitl P, Chen SY, Blanchette P, Branton PE et al. Adenovirus type 5 early region 1B 55K oncoprotein-dependent degradation of cellular factor Daxx is required for efficient transformation of primary rodent cells. J Virol 2011; 85: 8752–8765.
Everett RD . DNA viruses and viral proteins that interact with PML nuclear bodies. Oncogene 2001; 20: 7266–7273.
Everett RD, Chelbi-Alix MK . PML and PML nuclear bodies: implications in antiviral defence. Biochimie 2007; 89: 819–830.
Tavalai N, Stamminger T . New insights into the role of the subnuclear structure ND10 for viral infection. Biochim Biophys Acta 2008; 1783: 2207–2221.
Van Damme E, Van Ostade X . Crosstalk between viruses and PML nuclear bodies: a network-based approach. Front Biosci 2011; 17: 2910–2920.
Ascoli CA, Maul GG . Identification of a novel nuclear domain. J Cell Biol 1991; 112: 785–795.
Dyck JA, Maul GG, Miller WH Jr, Chen JD, Kakizuka A, Evans RM . A novel macromolecular structure is a target of the promyelocyte-retinoic acid receptor oncoprotein. Cell 1994; 76: 333–343.
Koken MH, Puvion-Dutilleul F, Guillemin MC, Viron A, Linares-Cruz G, Stuurman N et al. The t(15;17) translocation alters a nuclear body in a retinoic acid-reversible fashion. EMBO J 1994; 13: 1073–1083.
Weis K, Rambaud S, Lavau C, Jansen J, Carvalho T, Carmo-Fonseca M et al. Retinoic acid regulates aberrant nuclear localization of PML-RAR alpha in acute promyelocytic leukemia cells. Cell 1994; 76: 345–356.
de The H, Lavau C, Marchio A, Chomienne C, Degos L, Dejean A . The PML-RAR alpha fusion mRNA generated by the t(15;17) translocation in acute promyelocytic leukemia encodes a functionally altered RAR. Cell 1991; 66: 675–684.
Kakizuka A, Miller WH Jr, Umesono K, Warrell RP Jr, Frankel SR, Murty VV et al. Chromosomal translocation t(15;17) in human acute promyelocytic leukemia fuses RAR alpha with a novel putative transcription factor, PML. Cell 1991; 66: 663–674.
Pandolfi PP, Alcalay M, Fagioli M, Zangrilli D, Mencarelli A, Diverio D et al. Genomic variability and alternative splicing generate multiple PML/RAR alpha transcripts that encode aberrant PML proteins and PML/RAR alpha isoforms in acute promyelocytic leukaemia. EMBO J 1992; 11: 1397–1407.
Kastner P, Perez A, Lutz Y, Rochette Egly C, Gaub MP, Durand B et al. Structure, localization and transcriptional properties of two classes of retinoic acid receptor alpha fusion proteins in acute promyelocytic leukemia (APL): structural similarities with a new family of oncoproteins. EMBO J 1992; 11: 629–642.
Goddard AD, Borrow J, Solomon E . A previously uncharacterized gene, PML, is fused to the retinoic acid receptor alpha gene in acute promyelocytic leukaemia. Leukemia 1992; 6 (Suppl 3): 117S–119SS.
Melnick A, Fruchtman S, Zelent A, Liu M, Huang Q, Boczkowska B et al. Identification of novel chromosomal rearrangements in acute myelogenous leukemia involving loci on chromosome 2p23, 15q22 and 17q21. Leukemia 1999; 13: 1534–1538.
Melnick A, Licht JD . Deconstructing a disease: RARalpha, its fusion partners, and their roles in the pathogenesis of acute promyelocytic leukemia. Blood 1999; 93: 3167–3215.
Chang KS, Stass SA, Chu DT, Deaven LL, Trujillo JM, Freireich EJ . Characterization of a fusion cDNA (RARA/myl) transcribed from the t(15;17) translocation breakpoint in acute promyelocytic leukemia. Mol Cell Biol 1992; 12: 800–810.
Gurrieri C, Capodieci P, Bernardi R, Scaglioni PP, Nafa K, Rush LJ et al. Loss of the tumor suppressor PML in human cancers of multiple histologic origins. J Natl Cancer Inst 2004; 96: 269–279.
Lallemand-Breitenbach V, Zhu J, Puvion F, Koken M, Honore N, Doubeikovsky A et al. Role of promyelocytic leukemia (PML) sumolation in nuclear body formation, 11 S proteasome recruitment, and As2O3-induced PML or PML/retinoic acid receptor alpha degradation. J Exp Med 2001; 193: 1361–1371.
Li H, Leo C, Zhu J, Wu X, O'Neil J, Park EJ et al. Sequestration and inhibition of Daxx-mediated transcriptional repression by PML. Mol Cell Biol 2000; 20: 1784–1796.
Zhong S, Salomoni P, Pandolfi PP . The transcriptional role of PML and the nuclear body. Nat Cell Biol 2000; 2: E85–E90.
Pearson M, Carbone R, Sebastiani C, Cioce M, Fagioli M, Saito S et al. PML regulates p53 acetylation and premature senescence induced by oncogenic Ras. Nature 2000; 406: 207–210.
Ferbeyre G, de Stanchina E, Querido E, Baptiste N, Prives C, Lowe SW . PML is induced by oncogenic ras and promotes premature senescence. Genes Dev 2000; 14: 2015–2027.
Bischof O, Kirsh O, Pearson M, Itahana K, Pelicci PG, Dejean A . Deconstructing PML-induced premature senescence. Embo J 2002; 21: 3358–3369.
Langley E, Pearson M, Faretta M, Bauer UM, Frye RA, Minucci S et al. Human SIR2 deacetylates p53 and antagonizes PML/p53-induced cellular senescence. Embo J 2002; 21: 2383–2396.
Salomoni P, Ferguson BJ, Wyllie AH, Rich T . New insights into the role of PML in tumour suppression. Cell Res 2008; 18: 622–640.
Salomoni P, Pandolfi PP . The role of PML in tumor suppression. Cell 2002; 108: 165–170.
Bischof O, Kim SH, Irving J, Beresten S, Ellis NA, Campisi J . Regulation and localization of the Bloom syndrome protein in response to DNA damage. J Cell Biol 2001; 153: 367–380.
Carbone R, Pearson M, Minucci S, Pelicci PG . PML NBs associate with the hMre11 complex and p53 at sites of irradiation induced DNA damage. Oncogene 2002; 21: 1633–1640.
Takahashi Y, Lallemand-Breitenbach V, Zhu J, de The H . PML nuclear bodies and apoptosis. Oncogene 2004; 23: 2819–2824.
Hofmann TG, Will H . Body language: the function of PML nuclear bodies in apoptosis regulation. Cell Death Differ 2003; 10: 1290–1299.
Torok D, Ching RW, Bazett-Jones DP . PML nuclear bodies as sites of epigenetic regulation. Front Biosci 2009; 14: 1325–1336.
Jensen K, Shiels C, Freemont PS . PML protein isoforms and the RBCC/TRIM motif. Oncogene 2001; 20: 7223–7233.
Fagioli M, Alcalay M, Pandolfi PP, Venturini L, Mencarelli A, Simeone A et al. Alternative splicing of PML transcripts predicts coexpression of several carboxy-terminally different protein isoforms. Oncogene 1992; 7: 1083–1091.
Stadler M, Chelbi-Alix MK, Koken MH, Venturini L, Lee C, Saib A et al. Transcriptional induction of the PML growth suppressor gene by interferons is mediated through an ISRE and a GAS element. Oncogene 1995; 11: 2565–2573.
de Stanchina E, Querido E, Narita M, Davuluri RV, Pandolfi PP, Ferbeyre G et al. PML is a direct p53 target that modulates p53 effector functions. Mol Cell 2004; 13: 523–535.
Kentsis A, Gordon RE, Borden KL . Self-assembly properties of a model RING domain. Proc Natl Acad Sci USA 2002; 99: 667–672.
Nisole S, Maroui MA, Mascle XH, Aubry M, Chelbi-Alix MK . Differential Roles of PML Isoforms. Front Oncol 2013; 3: 125.
Muller S, Matunis MJ, Dejean A . Conjugation with the ubiquitin-related modifier SUMO-1 regulates the partitioning of PML within the nucleus. Embo J 1998; 17: 61–70.
Everett RD, Lomonte P, Sternsdorf T, van Driel R, Orr A . Cell cycle regulation of PML modification and ND10 composition. J Cell Sci 1999; 112 (Pt 24): 4581–4588.
Zhong S, Muller S, Ronchetti S, Freemont PS, Dejean A, Pandolfi PP . Role of SUMO-1-modified PML in nuclear body formation. Blood 2000; 95: 2748–2752.
Scaglioni PP, Yung TM, Cai LF, Erdjument-Bromage H, Kaufman AJ, Singh B et al. A CK2-dependent mechanism for degradation of the PML tumor suppressor. Cell 2006; 126: 269–283.
Condemine W, Takahashi Y, Zhu J, Puvion-Dutilleul F, Guegan S, Janin A et al. Characterization of endogenous human promyelocytic leukemia isoforms. Cancer Res 2006; 66: 6192–6198.
Duprez E, Saurin AJ, Desterro JM, Lallemand-Breitenbach V, Howe K, Boddy MN et al. SUMO-1 modification of the acute promyelocytic leukaemia protein PML: implications for nuclear localisation. J Cell Sci 1999; 112 (Pt 3): 381–393.
Owerbach D, McKay EM, Yeh ET, Gabbay KH, Bohren KM . A proline-90 residue unique to SUMO-4 prevents maturation and sumoylation. Biochem Biophys Res Commun 2005; 337: 517–520.
Wimmer P, Schreiner S, Dobner T . Human pathogens and the host cell SUMOylation system. J Virol 2012; 86: 642–654.
Haley KP, Overhauser J, Babiss LE, Ginsberg HS, Jones NC . Transformation properties of type 5 adenovirus mutants that differentially express the E1 A gene products. Proc Natl Acad Sci USA 1984; 81: 5734–5738.
Moran E, Grodzicker T, Roberts RJ, Mathews MB, Zerler B . Lytic and transforming functions of individual products of the adenovirus E1A gene. J Virol 1986; 57: 765–775.
Lowe SW, Ruley HE . Stabilization of the p53 tumor suppressor is induced by adenovirus 5 E1A and accompanies apoptosis. Genes Dev 1993; 7: 535–545.
White E . Regulation of the cell cycle and apoptosis by the oncogenes of adenovirus. Oncogene 2001; 20: 7836–7846.
McLorie W, McGlade CJ, Takayesu D, Branton PE . Individual adenovirus E1B proteins induce transformation independently but by additive pathways. J Gen Virol 1991; 72: 1467–1471.
Ruley HE . Adenovirus early region 1A enables viral and cellular transforming genes to transform primary cells in culture. Nature 1983; 304: 602–606.
Lowe SW, Jacks T, Housman DE, Ruley HE . Abrogation of oncogene-associated apoptosis allows transformation of p53-deficient cells. Proc Natl Acad Sci USA 1994; 91: 2026–2030.
White E . Regulation of apoptosis by the transforming genes of the DNA tumor virus adenovirus. Proc Soc Exp Biol Med 1993; 204: 30–39.
Kao CC, Yew PR, Berk AJ . Domains required for in vitro association between the cellular p53 and the adenovirus 2 E1B 55K proteins. Virology 1990; 179: 806–814.
Sarnow P, Ho YS, Williams J, Levine AJ . Adenovirus E1b-58kd tumor antigen and SV40 large tumor antigen are physically associated with the same 54 kd cellular protein in transformed cells. Cell 1982; 28: 387–394.
Yew PR, Liu X, Berk AJ . Adenovirus E1B oncoprotein tethers a transcriptional repression domain to p53. Genes Dev 1994; 8: 190–202.
Martin ME, Berk AJ . Corepressor required for adenovirus E1B 55,000-molecular-weight protein repression of basal transcription. Mol Cell Biol 1999; 19: 3403–3414.
Zantema A, Fransen JA, Davis OA, Ramaekers FC, Vooijs GP, DeLeys B et al. Localization of the E1B proteins of adenovirus 5 in transformed cells, as revealed by interaction with monoclonal antibodies. Virology 1985; 142: 44–58.
Sieber T, Dobner T . Adenovirus type 5 early region 1B 156 R protein promotes cell transformation independently of repression of p53-stimulated transcription. J Virol 2007; 81: 95–105.
Endter C, Hartl B, Spruss T, Hauber J, Dobner T . Blockage of CRM1-dependent nuclear export of the adenovirus type 5 early region 1B 55-kDa protein augments oncogenic transformation of primary rat cells. Oncogene 2005; 24: 55–64.
Endter C, Kzhyshkowska J, Stauber R, Dobner T . SUMO-1 modification required for transformation by adenovirus type 5 early region 1B 55-kDa oncoprotein. Proc Natl Acad Sci USA 2001; 98: 11312–11317.
Zantema A, Schrier PI, Davis OA, van Laar T, Vaessen RT, van der Eb AJ . Adenovirus serotype determines association and localization of the large E1B tumor antigen with cellular tumor antigen p53 in transformed cells. Mol Cell Biol 1985; 5: 3084–3091.
Goodrum FD, Shenk T, Ornelles DA . Adenovirus early region 4 34-kilodalton protein directs the nuclear localization of the early region 1B 55-kilodalton protein in primate cells. J Virol 1996; 70: 6323–6335.
König C, Roth J, Dobbelstein M . Adenovirus type 5 E4orf3 protein relieves p53 inhibition by E1B-55-kilodalton protein. J Virol 1999; 73: 2253–2262.
Krätzer F, Rosorius O, Heger P, Hirschmann N, Dobner T, Hauber J et al. The adenovirus type 5 E1B-55k oncoprotein is a highly active shuttle protein and shuttling is independent of E4orf6, p53 and Mdm2. Oncogene 2000; 19: 850–857.
Wienzek S, Roth J, Dobbelstein M . E1B 55-kilodalton oncoproteins of adenovirus types 5 and 12 inactivate and relocalize p53, but not p51 or p73, and cooperate with E4orf6 proteins to destabilize p53. J Virol 2000; 74: 193–202.
Dosch T, Horn F, Schneider G, Krätzer F, Dobner T, Hauber J et al. The adenovirus type 5 E1B-55K oncoprotein actively shuttles in virus-infected cells, whereas transport of E4orf6 is mediated by a CRM1 independent-mechanism. J Virol 2001; 75: 5677–5683.
Muller S, Dobner T . The adenovirus E1B-55K oncoprotein induces SUMO modification of p53. Cell Cycle 2008; 7: 754–758.
Martin ME, Berk AJ . Adenovirus E1B 55K represses p53 activation in vitro. J Virol 1998; 72: 3146–3154.
Kindsmuller K, Schreiner S, Leinenkugel F, Groitl P, Kremmer E, Dobner T . A 49-kilodalton isoform of the adenovirus type 5 early region 1B 55-kilodalton protein is sufficient to support virus replication. J Virol 2009; 83: 9045–9056.
Wimmer P, Blanchette P, Schreiner S, Ching W, Groitl P, Berscheminski J et al. Cross-talk between phosphorylation and SUMOylation regulates transforming activities of an adenoviral oncoprotein. Oncogene 2013; 32: 1626–1637.
Wang ZG, Delva L, Gaboli M, Rivi R, Giorgio M, Cordon-Cardo C et al. Role of PML in cell growth and the retinoic acid pathway. Science 1998; 279: 1547–1551.
Branton PE, Bayley ST, Graham FL . Transformation by human adenoviruses. Biochim Biophys Acta 1985; 780: 67–94.
Graham FL . Transformation by and oncogenicity of human adenoviruses. In: Ginsberg HS (ed). The Adenoviruses. Plenum Press: New York, pp 339–398 1984.
Ricciardi RP . Transformation and tumorigenesis mediated by the adenovirus E1A and E1B oncogenes. In: Barbanti-Brodano G (ed). DNA Tumor Viruses: Oncogenic Mechanisms. Plenum Press: New York, pp 195–210 1995.
Nevins JR, Vogt PK . Cell transformation by viruses. In: Fields BN, Knipe DM, Howley PM (eds). Virology Fields Virology 1, Third edn. Lippincott-Raven: New York, pp, 301–343 1996.
Gallimore PH, Turnell AS . Adenovirus E1A: remodelling the host cell, a life or death experience. Oncogene 2001; 20: 7824–7835.
Trentin JJ, Yabe Y, Taylor G . The quest for human cancer viruses: a new approach to an old problem reveals cancer induction in hamster hy human adenoviruses. Science 1962; 137: 835–849.
Graham FL, Smiley J, Russel WC, Nairn R . Characteristics of a human cell line transformed by DNA from human adenovirus type 5. J Gen Virol 1977; 36: 59–72.
Whittaker JL, Byrd PJ, Grand RJ, Gallimore PH . Isolation and characterization of four adenovirus type 12-transformed human embryo kidney cell lines. Mol Cell Biol 1984; 4: 110–116.
van den Heuvel SJL, The SI, Klein B, Jochemsen AG, Zantema A, van der Eb AJ . p53 shares an antigenic determinant with proteins of 92 and 150 kilodaltons that may be involved in senescence of human cells. J Virol 1992; 66: 591–595.
Fallaux FJ, Bout A, van der Velde I, van den Wollenberg DJ, Hehir KM, Keegan J et al. New helper cells and matched early region 1-deleted adenovirus vectors prevent generation of replication-competent adenoviruses. Hum Gene Ther 1998; 9: 1909–1917.
Fallaux FJ, Kranenburg O, Cramer SJ, Houweling A, Van Ormondt H, Hoeben RC et al. Characterization of 911: a new helper cell line for the titration and propagation of early region 1-deleted adenoviral vectors. Hum Gene Ther 1996; 7: 215–222.
Gallimore PH, Grand RJ, Byrd PJ . Transformation of human embryo retinoblasts with simian virus 40, adenovirus and ras oncogenes. Anticancer Res 1986; 6 (3 Pt B): 499–508.
Schiedner G, Hertel S, Kochanek S . Efficient transformation of primary human amniocytes by E1 functions of ad5: generation of new cell lines for adenoviral vector production. Hum Gene Ther 2000; 11: 2105–2116.
Byrd P, Brown KW, Gallimore PH . Malignant transformation of human embryo retinoblasts by cloned adenovirus 12 DNA. Nature 1982; 298: 69–71.
Endter C, Dobner T . Cell transformation by human adenoviruses. Curr Top Microbiol Immunol 2004; 273: 163–214.
Graham FL, Rowe DT, McKinnon R, Bacchetti S, Ruben M, Branton PE . Transformation by human adenoviruses. J Cell Physiol Suppl 1984; 3: 151–163.
Goddard AD, Yuan JQ, Fairbairn L, Dexter M, Borrow J, Kozak C et al. Cloning of the murine homolog of the leukemia-associated PML gene. Mamm Genome 1995; 6: 732–737.
Van Damme E, Laukens K, Dang TH, Van Ostade X . A manually curated network of the PML nuclear body interactome reveals an important role for PML-NBs in SUMOylation dynamics. Int J Biol Sci 2010; 6: 51–67.
Brand P, Lenser T, Hemmerich P . Assembly dynamics of PML nuclear bodies in living cells. PMC Biophys 2010; 3: 3.
Weidtkamp-Peters S, Lenser T, Negorev D, Gerstner N, Hofmann TG, Schwanitz G et al. Dynamics of component exchange at PML nuclear bodies. J Cell Sci 2008; 121: 2731–2743.
Geng Y, Monajembashi S, Shao A, Cui D, He W, Chen Z et al. Contribution of the C-terminal regions of promyelocytic leukemia protein (PML) isoforms II and V to PML nuclear body formation. J Biol Chem 2012; 287: 30729–30742.
Gurrieri C, Nafa K, Merghoub T, Bernardi R, Capodieci P, Biondi A et al. Mutations of the PML tumor suppressor gene in acute promyelocytic leukemia. Blood 2004; 103: 2358–2362.
Koken MH, Linares-Cruz G, Quignon F, Viron A, Chelbi-Alix MK, Sobczak-Thèpot J et al. The PML growth-suppressor has an altered expression in human oncognesis. Oncogene 1995; 10: 1315–1324.
Berscheminski J, Groitl P, Dobner T, Wimmer P, Schreiner S . The adenoviral oncogene E1A-13S interacts with a specific isoform of the tumor suppressor PML to enhance viral transcription. J Virol 2013; 87: 965–977.
Debbas M, White E . Wild-type p53 mediates apoptosis by E1A, which is inhibited by E1B. Genes Dev 1993; 7: 546–554.
Grand RJ, Grant ML, Gallimore PH . Enhanced expression of p53 in human cells infected with mutant adenoviruses. Virology 1994; 203: 229–240.
Sabbatini P, Lin J, Levine AJ, White E . Essential role for p53-mediated transcription in E1A-induced apoptosis. Genes Dev 1995; 9: 2184–2192.
Samuelson AV, Lowe SW . Selective induction of p53 and chemosensitivity in RB-deficient cells by E1A mutants unable to bind the RB-related proteins. Proc Natl Acad Sci USA 1997; 94: 12094–12099.
Turnell AS, Grand RJ, Gorbea C, Zhang X, Wang W, Mymryk JS et al. Regulation of the 26S proteasome by adenovirus E1A. Embo J 2000; 19: 4759–4773.
Querido E, Teodoro JG, Branton PE . Accumulation of p53 induced by the adenovirus E1A protein requires regions involved in the stimulation of DNA synthesis. J Virol 1997; 71: 3526–3533.
Kindsmüller K, Groitl P, Härtl B, Blanchette P, Hauber J, Dobner T . Intranuclear targeting and nuclear export of the adenovirus E1B-55K protein are regulated by SUMO1 conjugation. Proc Natl Acad Sci USA 2007; 104: 6684–6689.
Lethbridge KJ, Scott GE, Leppard KN . Nuclear matrix localization and SUMO-1 modification of adenovirus type 5 E1b 55K protein are controlled by E4 Orf6 protein. J Gen Virol 2003; 84: 259–268.
Müller S, Berger M, Lehembre F, Seeler JS, Haupt Y, Dejean A . c-Jun and p53 activity is modulated by SUMO-1 modification. J Biol Chem 2000; 275: 13321–13329.
Gostissa M, Hengstermann A, Fogal V, Sandy P, Schwarz SE, Scheffner M et al. Activation of p53 by conjugation to the ubiquitin-like protein SUMO-1. EMBO J 1999; 18: 6462–6471.
Kahyo T, Nishida T, Yasuda H . Involvement of PIAS1 in the sumoylation of tumor suppressor p53. Mol Cell 2001; 8: 713–718.
Rodriguez MS, Desterro JM, Lain S, Midgley CA, Lane DP, Hay RT . SUMO-1 modification activates the transcriptional response of p53. EMBO J 1999; 18: 6455–6461.
Schmidt D, Müller S . Members of the PIAS family act as SUMO ligases for c-Jun and p53 and repress p53 activity. Proc Natl Acad Sci USA 2002; 99: 2872–2877.
Bernardi R, Pandolfi PP . Role of PML and the PML-nuclear body in the control of programmed cell death. Oncogene 2003; 22: 9048–9057.
Fogal V, Gostissa M, Sandy P, Zacchi P, Sternsdorf T, Jensen K et al. Regulation of p53 activity in nuclear bodies by a specific PML isoform. EMBO J 2000; 19: 6185–6195.
Guo A, Salomoni P, Luo J, Shih A, Zhong S, Gu W et al. The function of PML in p53-dependent apoptosis. Nat Cell Biol 2000; 2: 730–736.
Gostissa M, Hofmann TG, Will H, del Sal G . Regulation of p53 functions: let's meet at the nuclear bodies. Curr Opin Cell Biol 2003; 15: 351–357.
Pearson MPP . PML interactions with p53 and its role in apoptosis and replicative senescence. Oncogene 2001; 20: 7250–7256.
Dobner T, Kzhyshkowska J . Nuclear export of adenovirus RNA. Curr Top Microbiol Immunol 2001; 259: 25–54.
Querido E, Morisson MR, Chu-Pham-Dang H, Thirlwell SW, Boivin D, Branton PE . Identification of three functions of the adenovirus E4orf6 protein that mediate p53 degradation by the E4orf6-E1B55K complex. J Virol 2001; 75: 699–709.
Stracker TH, Carson CT, Weitzman MD . Adenovirus oncoproteins inactivate the Mre11 Rad50 NBS1 DNA repair complex. Nature 2002; 418: 348–352.
Baker A, Rohleder KJ, Hanakahi LA, Ketner G . Adenovirus E4 34k and E1b 55k oncoproteins target host DNA ligase IV for proteasomal degradation. J Virol 2007; 81: 7034–7040.
Dallaire F, Blanchette P, Groitl P, Dobner T, Branton PE . Identification of integrin alpha3 as a new substrate of the adenovirus E4orf6/E1B 55-kilodalton E3 ubiquitin ligase complex. J Virol 2009; 83: 5329–5338.
Blanchette P, Cheng CY, Yan Q, Ketner G, Ornelles DA, Dobner T et al. Both BC-box motifs of adenovirus protein E4orf6 are required to efficiently assemble an E3 ligase complex that degrades p53. Mol Cell Biol 2004; 24: 9619–9629.
Querido E, Blanchette P, Yan Q, Kamura T, Morrison M, Boivin D et al. Degradation of p53 by adenovirus E4orf6 and E1B55K proteins occurs via a novel mechanism involving a Cullin-containing complex. Genes Dev 2001; 15: 3104–3117.
Querido E, Marcellus RC, Lai A, Charbonneau R, Teodoro JG, Ketner G et al. Regulation of p53 levels by the E1B 55-kilodalton protein and E4orf6 in adenovirus-infected cells. J Virol 1997; 71: 3788–3798.
Sarnow P, Hearing P, Anderson CW, Halbert DN, Shenk T, Levine AJ . Adenovirus early region 1B 58,000-dalton tumor antigen is physically associated with an early region 4 25,000-dalton protein in productively infected cells. J Virol 1984; 49: 692–700.
Mitsudomi T, Steinberg SM, Nau MM, Carbone D, D'Amico D, Bodner HK et al. p53 gene mutations in non-small-lung cell cancer cell lines and their correlation with the presence of ras mutations and clinical features. Oncogene 1992; 7: 171–180.
Nevels M, Dobner T . Determination of the transforming activities of adenovirus oncogenes. In: Wold WS, Tollefson AE (eds). Adenovirus Methods and Protocols Methods in Molecular Medicine 2 2nd edn. Humana Press Inc.: Totowa, NJ, pp 187–195 2006.
Dobner T, Horikoshi N, Rubenwolf S, Shenk T . Blockage by adenovirus E4orf6 of transcriptional activation by the p53 tumor suppressor. Science 1996; 272: 1470–1473.
Okamoto K, Beach D . Cyclin G is a transcriptional target of the p53 tumor suppressor protein. Embo J 1994; 13: 4816–4822.
Logan J, Pilder S, Shenk T . Functional analysis of adenovirus type 5 early region 1B. Cancer Cells 1984; 2: 527–532.
Sarnow P, Sullivan CA, Levine AJ . A monoclonal antibody detecting the adenovirus type 5-E1b-58Kd tumor antigen: characterization of the E1b-58Kd tumor antigen in adenovirus-infected and -transformed cells. Virology 1982; 120: 510–517.
Acknowledgements
We thank Roger D Everett for providing reagents, and greatly appreciate the critical comments and very helpful advise from Thomas Sternsdorf. We thank Ellis Jaffray, Gabriele Dobner and Thomas Speiseder for technical support and Hannah Staege for animal work. The Heinrich Pette Institute, Leibniz Institute for Experimental Virology is supported by the Freie und Hansestadt Hamburg and the Bundesministerium für Gesundheit (BMG). SS was supported by the Peter und Traudl Engelhorn Stiftung and additional grants from the Erich und Gertrud Roggenbuck Stiftung, the Else Kröner-Fresenius-Stiftung and the Deutsche Krebshilfe e. V. TD is supported by the Deutsche Forschungsgemeinschaft (DFG) and the Wilhelm Sander-Stiftung. Part of this work was supported by the B Braun Stiftung, the Dräger Stiftung and the Fonds der Chemischen Industrie. PB and PEB were supported by grants from the Canadian Institutes of Health Research. Image collection for the manuscript was performed in the McGill University Life Sciences Complex Imaging Facility. Purchase of equipment in the facility was made possible with funding from the Canadian Foundation for Innovation (CFI) and the Ministère du Développement économique, innovation et exportation Québec (MDEIE).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
PW is currently an employee of Novartis. The remaining authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Wimmer, P., Berscheminski, J., Blanchette, P. et al. PML isoforms IV and V contribute to adenovirus-mediated oncogenic transformation by functionally inhibiting the tumor-suppressor p53. Oncogene 35, 69–82 (2016). https://doi.org/10.1038/onc.2015.63
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2015.63
- Springer Nature Limited