Abstract
Recently, germline and somatic heterozygous mutations in the platelet-derived growth factor receptor β (PDGFRB) have been associated with familial infantile myofibromatosis (IM), which is characterized by soft tissue tumors, and overgrowth syndrome, a disease that predisposes to cancer. These mutations have not been functionally characterized. In the present study, the activity of three PDGFRB mutants associated with familial IM (R561C, P660T and N666K) and one PDGFRB mutant found in patients with overgrowth syndrome (P584R) was tested in various models. The P660T mutant showed no difference with the wild-type receptor, suggesting that it might represent a polymorphic variant unrelated to the disease. By contrast, the three other mutants were constitutively active and able to transform NIH3T3 and Ba/F3 cells to different extents. In particular, the germline mutant identified in overgrowth syndrome, P584R, was a stronger oncogene than the germline R561C mutant associated with myofibromatosis. The distinct phenotypes associated with these two mutations could be related to this difference of potency. Importantly, all activated mutants were sensitive to tyrosine kinase inhibitors such as imatinib, nilotinib and ponatinib. In conclusion, the PDGFRB mutations previously identified in familial IM and overgrowth syndrome activate the receptor in the absence of ligand, supporting the hypothesis that these mutations cause the diseases. Moreover, imatinib seems to be a promising treatment for patients carrying these mutations. To our knowledge, these are the first confirmed gain-of-function point mutations of PDGFRB in human cancer.
Similar content being viewed by others
Introduction
Infantile myofibromatosis (IM), although rare, is one of the most prevalent tumors of soft tissue in childhood.1, 2 It is characterized by firm nodules located in the skin, subcutaneous soft tissues, bones, muscles and viscera,3, 4 which generally develop before the age of 2 years.5, 6 The tumor cell morphology is intermediate between smooth muscle cells and fibroblasts.1, 3, 6 There are two main forms of IM: the most common is the solitary form with a single nodule whereas a second form with multiple lesions in the bones and soft tissues also exists.2, 3, 5 IM has a good prognosis and regression often occurs, except in the rare cases of visceral involvement, which is lethal in up to 70% of the cases.3, 6 Familial cases of IM have been reported with both recessive and dominant modes of inheritance.1, 7, 8 Currently, definitive guidelines for treatment of IM with visceral involvement do not exist.2
Recently, point mutations in platelet-derived growth factor (PDGF) receptor β (PDGFRβ) were associated with familial IM.8, 9 PDGF receptors, PDGFRα and PDGFRβ, are receptor tyrosine kinases that stimulate cell survival, proliferation and motility.10 They are encoded by two highly related genes, PDGFRA and PDGFRB, and are expressed in various cell types, including fibroblasts.11 Both genes are essential for proper embryo development in mice.12, 13 PDGFR proteins are divided into three different parts: the extracellular ligand binding domain, the transmembrane α-helix and the intracellular kinase domain. The activity of the kinase domain is controlled by the juxtamembrane domain, the activation loop and the C-terminal tail, which keep the receptor silent in the absence of ligand.14, 15, 16 Upon PDGF binding, the receptor dimerizes and undergoes a conformational change, which activates the kinase domain. PDGFRs trans-autophosphorylate on tyrosine residues, leading to the binding of Src homology 2-domain-containing proteins, and subsequently, to the activation of signaling pathways, such as mitogen-activated protein (MAP) kinases, PLCγ (phospholipase Cγ), signal transducers and activators of transcription (STAT) and PI3K (phosphatidylinositol-3 kinase).10, 14, 17
Martignetti et al.8 identified two germline missense mutations in PDGFRβ in eight families with members diagnosed with familial IM: the R561C substitution was present in seven families whereas the P660T was present in only one family.8 The R561 residue is located in the juxtamembrane domain of the receptor while the P660 residue is found in the kinase domain (Figure 1a). The R561C mutation was found in four additional families with IM in a second study.9 Interestingly, in one of the affected individuals, a somatic mutation located in the kinase domain, N666K, was found in addition to the germline one.9 All these mutations associated with familial IM are heterozygous.8, 9 Their impact on the receptor function has not been demonstrated experimentally.
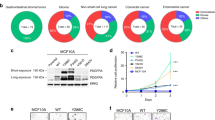
The R561C and N666K mutants, but not P660T, transform cells. (a) Schematic representation of PDGFRβ and localization of the mutations identified in familial IM (black) and overgrowth syndrome (gray). Ig-like, immunoglobulin-like; TM, transmembrane domain; JM, juxtamembrane domain; N-lobe, N-terminal lobe; C-lobe, C-terminal lobe; Ct, C-terminal tail. (b) Ba/F3 cells stably expressing wild-type (WT) PDGFRβ, P660T, R561C or N666K were washed twice and cultured without IL-3 for 2 days. Living cells were counted in a Bürker chamber. The histogram represents the relative number of living cells on day 2 compared with day 0. As a control, cells were also seeded in the presence of IL-3 and the number of living cells on day 2 was similar in each condition (data not shown). (c) Ba/F3 cells stably expressing WT PDGFRβ or P660T and selected cells expressing R561C or N666K were washed twice and treated with control medium or PDGF-BB. Cell proliferation was assessed after 24 h by measuring the incorporation of [3H]-thymidine. As a control, [3H]-thymidine incorporation was similar in every cell lines seeded with IL-3 (data not shown). (b, c) One representative experiment out of four is shown with s.d. (d) NIH3T3 cells were transfected with WT PDGFRβ, D850V, P660T, R561C or N666K in triplicate and processed as described.27 Three weeks after transfection, cells were fixed in ice-cold methanol for 20 min and foci were stained with 0.2% crystal violet (Sigma-Aldrich, Saint-Louis, MO, USA) in 20% ethanol for 20 min. (e) Foci density was quantified by using the Quantity One software (Bio-Rad) and normalized to 1 for the wild-type receptor. The average of three independent experiments is represented with s.e.m. (one-sided Student's t-test). *P<0.05; **P<0.01; ***P<0.001.
Germline mutations in PDGFRβ have also been associated with overgrowth syndromes, which are characterized by accelerated linear growth with increased risks of cancer and cognitive disorders.18 In a recent study, a germline mutation in the juxtamembrane domain of PDGFRβ (Figure 1a), P584R, was identified in two girls presenting an overgrowth phenotype with severe neurological problems.19 Interestingly, one of these two patients presented a myofibroma on her mandibula.19
Although PDGFRA mutations have been associated with various types of cancers such as gastrointestinal stromal tumors (GIST), glioblastoma and inflammatory fibroid polyps,10, 16, 20 the occurrence of PDGFRB activating point mutations have not been confirmed yet. Furthermore, mice carrying a weak activating point mutation in PDGFRβ, D849N, are viable and show no sign of tumor development.21 Only rare PDGFRB fusions with different partner genes were reported in myeloid neoplasms associated with hypereosinophilia.16, 20, 22, 23 These patients can be efficiently treated with imatinib, a tyrosine kinase inhibitor, which was initially developed for BCR-ABL-positive chronic myeloid leukemia.20 By contrast, loss-of-function mutations in PDGFRβ have a role in idiopathic basal ganglia calcification (or Fahr disease).24, 25, 26
The aim of the present work was to characterize the PDGFRβ mutations associated with familial IM and overgrowth syndrome. We demonstrate that the somatic N666K mutation and, to a lesser extent, the germline R561C substitution activate PDGFRβ and allow cell transformation. In contrast, PDGFRβ P660T behaves like the wild-type receptor. The germline mutation P584R associated with overgrowth elicits a stronger response than the R561C substitution. Finally, activated mutants are sensitive to tyrosine kinase inhibitors such as imatinib.
Results
The N666K mutant and, to a lesser extent, the R561C mutant transform cells
We first assessed the transforming potential of three familial IM-associated PDGFRβ mutations, namely P660T, R561C and N666K (Figure 1a), in two different cell lines.
First, we tested their ability to transform Ba/F3 cells, which need interleukin (IL)-3 in the culture medium to survive and proliferate, except if they express an oncogene. This is a widely used and very sensitive model of cell transformation.27, 28 We stably expressed the empty vector, wild-type PDGFRβ or its mutants (P660T, R561C or N666K). The expression of the different receptors at the cell surface was verified by flow cytometry (Supplementary Figure 1). Then, cells were washed and cultured in a medium without IL-3. We determined the number of cells that were still alive 2 days after IL-3 removal (Figure 1b). We observed that the relative number of living Ba/F3 cells expressing the R561C and N666K mutants was higher than that of cells expressing the wild-type receptor and the P660T mutant. About 2 weeks after IL-3 removal, we obtained selected cell lines steadily growing without IL-3 for the N666K mutant but not for the wild-type receptor or the P660T mutant. Regarding R561C, we also obtained IL-3-independent selected cell lines, but the process was slower: it took more than 4 weeks. The proliferation of these cells was assessed by quantifying the incorporation of [3H]-thymidine (Figure 1c). Selected cells expressing the R561C and N666K mutants proliferated in the absence of exogenous stimulation, whereas cells expressing the wild-type receptor and the P660T mutant only proliferated when they were stimulated with PDGF. These results were confirmed by a colony formation assay in the absence of cytokines (Supplementary Figure 2). Selected cells expressing the N666K mutant gave rise to more colonies than those expressing the R561C mutant, while the wild-type receptor and the P660T mutant did not allow colony formation.
To further establish the transforming potential of the mutants, we performed a foci formation assay in NIH3T3 fibroblasts that were transfected with the wild-type receptor or one of the mutants. PDGFRβ D850V, which corresponds to the gastrointestinal stromal tumor-associated D842V mutation in PDGFRα29 and to the activating D849V substitution in murine PDGFRβ,30 was used as a positive control. As expected, the wild-type receptor did not give rise to many foci whereas cells transfected with the D850V mutant formed robust foci (Figure 1d). Interestingly, the N666K mutant gave rise to a large number of foci, even more than the D850V mutant. Regarding the two other substitutions, R561C and P660T, it was difficult to assess whether they were different from the wild-type receptor by naked eye (Figure 1d). Nevertheless, the quantification of three independent experiments showed that the increase in foci density was significant for the R561C mutant, as well as D850V and N666K, compared with the wild-type receptor (Figure 1e).
Altogether, these results show that the N666K mutant transforms Ba/F3 and NIH3T3 cells while R561C is a weaker oncogene. On the contrary, the P660T mutant does not induce cell transformation in our assays.
The R561C and N666K mutants are constitutively activated
We next tested whether the mutations constitutively activated the receptor kinase activity. Receptors from stably transfected Ba/F3 cells were immunoprecipitated and their expression and phosphorylation levels were assessed by western blot (Figure 2a).
PDGFRβ R561C and N666K are constitutively phosphorylated and constitutively activate signaling pathways. (a) Analysis of the phosphorylation and expression levels of the wild-type and mutated receptors by western blot. Ba/F3 cells stably expressing WT PDGFRβ or P660T and selected cells expressing R561C or N666K were starved for 4 h and stimulated with PDGF-BB for 15 min or left untreated. PDGFRβ was immunoprecipitated and analyzed by western blot experiments using an anti-phosphotyrosine antibody. Membranes were re-probed with an anti-PDGFRβ antibody. As a negative control, cells expressing the empty vector were used. Representative blots are shown (n=4). (b) Analysis of the phosphorylation of ERK1/2, PLCγ, STAT3, STAT5 and Akt by western blot. Ba/F3 cells stably expressing WT PDGFRβ or P660T and selected cells expressing R561C or N666K were starved for 4 h and stimulated with PDGF-BB for 15 min or left untreated. The phosphorylated protein was first analyzed and the membrane was re-probed with an antibody targeting the corresponding protein. Similar results were obtained in four independent experiments. (c) MCF7 cells were co-transfected with WT or mutated PDGFRβ, β-galactosidase and an SRE promoter controlling luciferase expression. Cells were washed 4 h after transfection and left in the incubator for 24 h. The results are expressed as a ratio between luciferase and β-galactosidase activity. One representative experiment out of four is shown with s.d. (d) HT-1080 cells were processed as in (c), except that the luciferase expression was controlled by a STAT-driven promoter (pGRR5). **P<0.01; ***P<0.001.
Wild-type PDGFRβ and the mutants were expressed at a similar level, except the N666K mutant whose expression was higher. As expected, two major bands were present for the different receptors: the upper one represents the mature glycosylated form whereas the lower one is the immature form of the receptor located in the endoplasmic reticulum.31 The wild-type and the P660T receptors were phosphorylated only in response to PDGF. In contrast, the R561C and N666K mutants were constitutively phosphorylated (Figure 2a). Noticeably, the phosphorylation of R561C was not detectable in some of the selected cell lines used in replicate experiments.
We next analyzed the activation of different signaling pathways by western blot (Figure 2b). In agreement with their ability to transform cells and their constitutive activation, we observed that the R561C and N666K mutants induced the phosphorylation of ERK1/2, PLCγ, STAT3, STAT5 and Akt in the absence of PDGF stimulation. STAT phosphorylation by the R561C mutant was weaker than that induced by the N666K. The P660T mutant behaved as the wild-type receptor. Interestingly, phosphorylated STAT3 and STAT5 were barely detectable in stimulated cells expressing the wild-type receptor or the P660T mutant (Figure 2b). This was consistent in all independent selected cell lines.
We confirmed these results in luciferase reporter assays. MCF7 and HT-1080 cells were transfected with wild-type or mutant PDGFRβ and with a reporter construct sensitive to MAP kinases (Figure 2c) or to STAT transcription factors (Figure 2d). In line with western blot experiments, the R561C and N666K mutants constitutively activated MAP kinases and STAT, whereas the P660T mutant showed no significant difference with the wild-type receptor (Figures 2c and d).
Taken together, our results demonstrate that the R561C and N666K mutants, in the absence of PDGF, activate signaling pathways that are normally activated by the stimulated wild-type receptor. Furthermore, these two mutants activate STATs to a greater extent than the stimulated wild-type and P660T receptors.
The overgrowth-associated P584R mutant is more active than R561C
In the course of the present project, the overgrowth-associated P584R substitution was described. It is a germline mutation, like the familial IM-associated R561C mutation. We compared the R561C and P584R substitutions to understand how two mutations located in the same part of PDGFRβ, the juxtamembrane domain, could give rise to different phenotypes.
We first verified whether the P584R substitution had an impact on PDGFRβ activity by performing luciferase reporter assays. We observed that the P584R mutant was able to activate MAP kinases and STAT reporters in the absence of PDGF to a greater extent than R561C (Figures 3a and b). To confirm this difference, we performed a foci formation assay in NIH3T3 cells that showed that the P584R mutant gave rise to more foci than the R561C mutant (Figure 3c). We also expressed P584R in Ba/F3 cells. After verifying the cell surface expression of the mutant by flow cytometry (Supplementary Figure 1), we counted the number of living cells after IL-3 removal (Figure 3d). Surprisingly, 3 days after IL-3 removal, the P584R mutation conferred a modest survival advantage compared with the wild-type receptor. In contrast, the relative number of cells expressing the R561C mutant was much higher, in agreement with Figure 1b. However, about 3 weeks after IL-3 removal, we obtained selected cells for the P584R mutant, while it took more than 4 weeks to obtain IL-3-independent selected cell lines for the R561C mutant. These cells were able to proliferate in the absence of PDGF stimulation (data not shown). Thus, both mutations stimulated Ba/F3 cells proliferation with different kinetics. Furthermore, in a colony formation assay, selected cells expressing the P584R mutant gave rise to a higher number of colonies, which were also more spread, compared with those expressing the R561C mutant (Supplementary Figure 2).
The P584R mutant found in overgrowth syndrome is more active than the R561C. (a) MCF7 cells were co-transfected with WT PDGFRβ, R561C or P584R, β-galactosidase and an SRE promoter controlling luciferase expression and processed as described in (b). One representative experiment out of three is shown with s.d. (b) HT-1080 cells were processed as in (a), except that the luciferase expression was controlled by a STAT-driven promoter (pGRR5). (c) NIH3T3 cells were transfected with the empty vector, R561C or P584R in triplicate and processed as described.27 Foci were stained as described in Figure 1d. Foci density was quantified as described in Figure 1e and normalized to 1 for the R561C mutant. The average of three independent experiments is represented. (d) Ba/F3 cells stably expressing WT PDGFRβ, R561C or P584R were processed as in Figure 1b except that they were cultured for 3 days. The histogram represents the relative number of living cells on day 3 compared with day 0. One representative experiment out of two is shown with s.d. *P<0.05; **P<0.01; ***P<0.001.
Taken together, our results show that the overgrowth-associated P584R mutation activates PDGFRβ to a greater extent than the R561C substitution.
The R561C, N666K and P584R mutants can induce cancer development in vivo
Our data indicate that two familial IM-associated mutations, R561C and N666K, and the overgrowth-associated mutation P584R constitutively activate PDGFRβ. To further assess the oncogenic properties of these mutations, we used a previously described in vivo model of cancer development.32 Selected Ba/F3 cells expressing the R561C, N666K or P584R mutant were tumorigenic in vivo when intravenously injected into immunodeficient BALB/c rag2−/− mice, as shown by monitoring survival (Figure 4a). The mice that received cells expressing the N666K and P584R mutants developed the disease faster than those injected with cells expressing the R561C mutant, whereas the wild-type receptor and the P660T mutant had no effect. As expected, increased spleen weight correlated with poor survival and constitutive activity of PDGFRβ mutants (Figure 4b). Splenomegaly was not observed in mice injected with cells expressing the wild-type receptor and the P660T mutant.
The R561C, N666K and P584R mutants induce cancer development in vivo. Two million Ba/F3 cells stably expressing the wild-type receptor or the P660T mutant or selected Ba/F3 cells expressing the R561C, N666K or P584R mutant were intravenously injected into BALB/c rag2−/− mice (5 per group). (a) Mice survival was followed for 50 days. (b) Spleens were isolated at the time of killing and weighed. The black line represents the median. **P<0.01; ***P<0.001.
In conclusion, our in vivo data support the in vitro results demonstrating that the R561C, N666K and P584R mutants are active and could cause the disease. Furthermore, this assay confirms that the R561C mutant is less active than N666K and P584R.
R561C, N666K and P584R are sensitive to tyrosine kinase inhibitors
Our results suggest that blocking the PDGF receptor activity might offer a therapeutic option for myofibromatosis and overgrowth. Therefore, we tested whether activated mutants were sensitive to imatinib, a first-generation tyrosine kinase inhibitor.
Firstly, we tested the effect of imatinib in a luciferase reporter assay responsive to MAP kinases (Figure 5). The luciferase activity induced by the activated mutants R561C, N666K and P584R was much reduced upon imatinib treatment. As expected, the D850V mutation was resistant to this tyrosine kinase inhibitor, in agreement with published reports regarding the corresponding mutation in PDGFRA.33, 34
The R561C and N666K mutants are sensitive to imatinib. MCF7 cells were co-transfected with different vectors containing WT PDGFRβ or PDGFRβ mutants, β-galactosidase and an SRE promoter controlling luciferase expression. Cells were washed 4 h after transfection and treated or not with 500 nM imatinib for 24 h. In addition, cells expressing the wild-type receptor were stimulated with PDGF-BB. The histogram represents the mean of the ratio between luciferase and β-galactosidase activity of three independent experiments with s.e.m. The condition without imatinib was used as a reference. **P<0.01; ***P<0.001.
To better characterize the sensitivity of the three activated mutants to imatinib, we performed a [3H]-thymidine incorporation assay with increasing doses of imatinib. At a concentration of 1 μM, a concentration corresponding to the plasma level of imatinib in patients,35, 36 cell proliferation induced by the stimulated wild-type receptor or by the activated mutants R561C, N666K and P584R was abolished (Figure 6a), in line with the decrease in signaling observed in luciferase experiments (Figure 5). Interestingly, the R561C and P584R mutants were consistently more sensitive to imatinib than the stimulated wild-type receptor, whereas the N666K mutant was slightly less sensitive (Figure 6a). This was confirmed by the calculation of the half maximal inhibitory concentration (IC50) (Figure 6d). Importantly, IL-3-dependent cell proliferation was not affected by imatinib, which selectively blocks PDGFRs and a few other kinases such as ABL (Figure 6a, gray line).
PDGFRβ mutations affect sensitivity to tyrosine kinase inhibitors. (a–c) Ba/F3 cells stably expressing WT PDGFRβ and selected cells expressing R561C, N666K or P584R were washed twice and treated with increasing concentrations of imatinib (a), nilotinib (b) or ponatinib (c). In addition, cells expressing the wild-type receptor were stimulated with PDGF-BB. As a negative control, cells expressing the wild-type receptor were also seeded in the presence of IL-3 (gray line). Cell proliferation was assessed after 24 h by measuring the incorporation of [3H]-thymidine. One representative experiment is shown with s.d. (n⩾3). Treatment with corresponding concentrations of vehicles had no effect on cell proliferation (data not shown). (d) IC50 (nM) was calculated for each inhibitor and each mutant. The table represents the mean of at least three independent experiments.
Since N666K was less sensitive to imatinib, we tested two other inhibitors approved for chronic myeloid leukemia: nilotinib, a second-generation inhibitor, and ponatinib, a third-generation inhibitor. In both cases, the R561C and P584R mutants were more sensitive than the stimulated wild-type receptor (Figures 6b and c). This was confirmed by a lower IC50 (Figure 6d). According to the dose–response curve and the calculation of the average IC50 (Figures 6b and d), the sensitivity of the N666K mutant to nilotinib did not differ significantly from that of the wild-type receptor. However, N666K was less sensitive to ponatinib than the wild-type receptor (Figure 6c). Nevertheless, ponatinib inhibited N666K at lower concentrations compared with the two other compounds (Figure 6d).
These results were confirmed by the analysis of the phosphorylation level of receptors and downstream signaling mediators by western blot (Figure 7). As expected, we observed that phosphorylation of the wild-type receptor in response to PDGF and phosphorylation of ERK1/2 were progressively decreased with increasing doses of imatinib (Figure 7a), nilotinib (Figure 7b) and ponatinib (Figure 7c). In agreement with the results obtained in proliferation assays, the constitutive phosphorylation of R561C and P584R was abolished with lower doses of inhibitors, in comparison with the wild-type receptor. In contrast, the constitutive phosphorylation of N666K was less sensitive to all inhibitors than the PDGF-induced phosphorylation of the wild-type receptor. For the three mutants, the decrease in constitutive signaling following the treatment with inhibitors was in line with the decrease in receptor phosphorylation. None of the inhibitors affected phosphorylation of ERK1/2 and STAT5 induced by IL-3, demonstrating the specificity of the treatment (Figure 7d).
Tyrosine kinase inhibitors decrease receptor phosphorylation and signaling downstream of PDGFRβ mutants. (a–c) Analysis of the phosphorylation of the activated PDGFRβ mutants, as well as the activation of ERK1/2 and STAT5, in response to imatinib (a), nilotinib (b) or ponatinib (c) treatment. Western blot experiments on Ba/F3 cells expressing WT PDGFRβ and selected cells expressing R561C, N666K or P584R were performed as described in Figure 2, except that cells were treated with increasing doses of imatinib, nilotinib or ponatinib during starvation. We were not able to observe any phosphorylation of STAT5 downstream of the PDGF-stimulated wild-type receptor (ND, not detectable). The maximal concentration used for ponatinib was 1 μM because higher doses were toxic (data not shown). (d) As a control, the IL-3-induced phosphorylation of ERK1/2 and STAT5 in Ba/F3 cells transfected with the empty vector and incubated with the three inhibitors was also analyzed by western blot. (a–d) Two experiments were performed with similar results.
Taken together, these results indicated that R561C, N666K and P584R are sensitive to specific tyrosine kinase inhibitors.
Discussion
In this study, we show for the first time that PDGFRB mutations found in patients with familial IM or overgrowth syndrome are of an activating character and sensitive to appropriate tyrosine kinase inhibitors.
Our results demonstrate that both the R561C and N666K mutations promote the PDGFR active conformation in the absence of ligand. Indeed, they allowed the transformation of two different cell lines (Ba/F3 and NIH3T3 cells), constitutively activated signaling pathways that are normally induced by the wild-type receptor in response to PDGF and promoted tumorigenesis in vivo. Furthermore, N666K seemed to be a stronger activating mutation compared with R561C. In agreement with the observation that the N666K mutant generated selected Ba/F3 cell lines faster than the R561C mutant, N666K induced a stronger activation of STAT and had a greater ability to transform NIH3T3 cells. This may be related to the fact that N666K is a somatic mutation while R561C is a germline one, which must be compatible with the embryo development. In this respect, Linhares et al.37 described an asymptomatic R561C carrier and suggested that an additional mutation was required for a full phenotypic penetrance.
In contrast, we were not able to show any impact of the P660T substitution on PDGFRβ activity, even if it was predicted to be damaging by different prediction algorithms.8 However, we cannot exclude the possibility that the techniques that we used were not sensitive enough to detect a difference between the wild-type receptor and P660T. Our results correlate with the fact that P660T has been described as a polymorphism in one individual. The residue P660, located in the kinase domain, is not well conserved among PDGFR family members, by contrast to N666. Altogether, these observations do not support the involvement of the P660T mutation in the disease.
Regarding the germline PDGFRB mutation associated with overgrowth, our results show that the P584R mutation also confers a gain of function. P584R presented a higher activity than the germline familial IM-associated R561C mutation in luciferase, cell transformation and in vivo assays. This difference in the level of receptor activation could explain how two activating germline mutations located in the same domain of the receptor (juxtamembrane domain) could lead to different phenotypes. In this respect, P584R was associated with one case of myofibroma in one of the reported overgrowth patients. Future studies will have to decipher whether the P584R mutation could also specifically impact a particular signaling pathway downstream of PDGFRβ, compared with R561C. Indeed, different studies demonstrated involvement of the PI3K in overgrowth syndromes.38, 39 The discovery of a link between an activating PDGFRβ mutation and overgrowth is also reminiscent of the growth retardation effect of imatinib in children with chronic myeloid leukemia.40
Finally, we demonstrated that PDGFRB activating mutations associated with familial IM and severe overgrowth syndrome, which is associated with various neurological symptoms, are sensitive to tyrosine kinase inhibitors such as imatinib, nilotinib and ponatinib. Noticeably, their sensitivity differs slightly from the wild-type receptor. It could be due to the different localization of the mutations in the receptor. In particular, the R561C and P584R mutations may confer a higher sensitivity to imatinib, which could be given at a lower dose, similar to that given to patients with hypereosinophilia associated with PDGFRA fusion, with fewer side effects. Nevertheless, potential benefits must be carefully balanced with the side effects of a long-term imatinib treatment in children.40
To conclude, we show that PDGFRB mutations associated with familial IM and overgrowth syndromes are activating and sensitive to tyrosine kinase inhibitors. Our data support the hypothesis that these mutations cause these two diseases and suggest a potential treatment for severe cases. PDGFRB should thus be systematically sequenced in suspected cases of familial IM and overgrowth. The difference between the two phenotypes could be related to the mutation potency. To our knowledge, these are the first confirmed gain-of-function point mutations of PDGFRB in a human cancer.
Materials and methods
Reagents and antibodies
PDGF-BB was purchased from PeproTech (Rocky Hill, NJ, USA). Imatinib and nilotinib were purchased from LC Laboratories (Woburn, MA, USA). Ponatinib was purchased from Selleck Chemicals (Munich, Germany). IL-3 was produced as described.32, 41 See Supplementary Materials and Methods for detailed descriptions of antibodies.
Cell culture
The NIH3T3 (All) cell line was purchased from ATCC (Manassas, VA, USA) and cultured on collagen-coated plates (30 μg/ml, Purecol, Advanced Biomatrix, San Diego, CA, USA) in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco, Life Technologies, Grand Island, NY, USA) supplemented with 10% decomplemented fetal calf serum (FCS), 1 mM sodium pyruvate and antibiotics (50 U/ml penicillin and 50 μg/ml streptomycin). HT-1080 cells, derived from a human fibrosarcoma, were purchased from ATCC and were grown in Iscove’s Modified Dulbecco’s Medium (Gibco, Life Technologies) supplemented with 10% FCS.26 MCF7 cells33 derived from a human breast adenocarcinoma were cultured in DMEM supplemented with 10% FCS. IL-3-dependent Ba/F3 cells,33 a murine bone marrow-derived pro-B cell line, were grown in DMEM supplemented with 10% FCS and IL-3 (500 U/ml).
Site-directed mutagenesis
After cloning of wild-type PDGFRβ in pcDNA3.1 (Invitrogen, Life Technologies) and pEF/myc/cyto (Invitrogen, Life Technologies), the point mutations D850V, P660T, R561C, N666K and P584R were introduced in each vector by site-directed mutagenesis according to the QuickChange XL-II kit protocol (Stratagene, La Jolla, CA, USA). In all cases, the whole insert was sequenced.
Production of Ba/F3 stable cell lines expressing PDGFRβ
Ten million Ba/F3 cells were electroporated (200 V, 75 Ohm, 1300 μF) with 50 μg pEF/myc/cyto-PDGFRβ wild-type, R561C, P660T, N666K or P584R and seeded in 15-cm dishes. Twenty-four hours after transfection, 500 000 electroporated cells were transferred to 10-cm dishes and selected with 3 mg/ml G418. After approximately 2 weeks of selection, Ba/F3 cells stably expressing the receptors were sorted by flow cytometry. Briefly, cells were stained with a primary anti-PDGFRβ antibody (AH 17.2, 5 μg/ml) for 40 min at 4 °C, washed and incubated with a secondary antibody conjugated to phycoerythrin for 40 min at 4 °C in the dark. Finally, stained cells were sorted by FACSAria III (BD Biosciences, Erembodegem, Belgium).
Sorted Ba/F3 cells stably expressing the R561C, N666K and P584R mutants were then selected in DMEM supplemented with 10% FCS without IL-3.
Living cell count
Ba/F3 cells stably expressing wild-type PDGFRβ or one of its mutants were washed twice and seeded at 400 000 cells/well in 6-well plates in a medium without IL-3 in triplicate. On days 2 and 3, the number of living cells was assessed in a Bürker chamber. As a control, cells were seeded in a medium with IL-3.
[3H]-thymidine incorporation assay
Ba/F3 cells stably expressing wild-type PDGFRβ or PDGFRβ P660T and selected Ba/F3 cells expressing PDGFRβ R561C, N666K or P584R were washed twice with DMEM supplemented with 10% FCS to remove IL-3 and were seeded at 10 000 cells/well in 96-well plates. Then, 20 ng/ml PDGF-BB, imatinib, nilotinib, ponatinib, IL-3 or vehicle was added in the appropriate wells. Twenty hours later, 0.5 μCi [3H]-thymidine (GE Healthcare, Diegem, Belgium) was added in each well for 4 h and [3H]-thymidine incorporation was measured using a TopCount instrument (Perkin Elmer, Waltham, MA, USA) as described.33
Foci formation assay
NIH3T3 cells were transfected with wild-type PDGFRβ or one of its mutants (0.55 μg) with Lipofectamine 2000 as recommended by the manufacturer (Invitrogen, Life Technologies) and processed as described.27
Western blot
Transfected Ba/F3 cells were washed twice with DMEM supplemented with 10% FCS to remove IL-3 and were seeded in 6-well plates (2.106–4.106 cells/well), with or without increasing doses of imatinib, nilotinib or ponatinib. Four hours after IL-3 withdrawal, cells were stimulated with IL-3 or 25 ng/ml PDGF-BB or left untreated. After 15 min of stimulation, cells were washed with PBS. Cell lysates and western blots were performed as previously described.26
For immunoprecipitation experiments, cell lysates were produced as described above. Then, they were first incubated with an anti-PDGFRβ antibody (Santa Cruz Biotechnology) overnight on wheel at 4 °C, and Protein A/G UltraLink Resin (Thermo Scientific, Thermo Fisher Scientific, Waltham, MA, USA) for 1 h 30 min on wheel at 4 °C. Beads were washed twice with lysis buffer and once with PBS, and were resuspended in 2 × Laemmli buffer for western blot analysis.
Luciferase assays
MCF7 cells were seeded at 150 000 cells/well in 12-well plates. The day after, cells were co-transfected by using TurboFect Transfection Reagent (Thermo Scientific, Thermo Fisher Scientific) or FuGENE HD Transfection Reagent (Promega, Leiden, The Netherlands), as recommended by the manufacturers, with different vectors: 0.5 μg pcDNA3.1-PDGFRβ (wild-type or mutant), 0.5 μg of a plasmid in which the luciferase expression is controlled by serum-response elements (pSRE)42 and 0.5 μg pEF1-β-galactosidase (Invitrogen, Life Technologies). Four hours after transfection, cells were washed with PBS, fresh medium was added and 20 ng/ml PDGF-BB and/or 500 nM imatinib were added in the appropriated wells or cells were left untreated.
HT-1080 cells were seeded at 175 000 cells/well in 12-well plates. The day after, cells were co-transfected with different vectors: 0.5 μg pcDNA3.1-PDGFRβ (wild-type or mutant), 0.5 μg of a plasmid containing a luciferase reporter controlled by a STAT-sensitive promoter (pGRR5)43 and 0.5 μg pEF1-β-galactosidase (Invitrogen, Life Technologies), as described above. Four hours after transfection, cells were washed with PBS and starved in serum-free medium.
Cells were lysed 24 h after transfection and the luciferase activity was assessed as described33 by using GloMax (Promega). The β-galactosidase activity was assessed as described.26
Tumorigenicity assay in BALB/c rag2−/− mice
Two million Ba/F3 cells stably expressing the wild-type receptor or the P660T mutant or selected Ba/F3 cells expressing the R561C, N666K or P584R mutant were intravenously injected in 9-week-old female BALB/c rag2−/− mice (Taconic Biosciences Inc., Hudson, NY, USA). Five mice per condition were tested. Survival was followed and isolated spleens were weighed after death. Mice that received cells expressing the wild-type receptor or the P660T mutant were still alive 50 days after injection and were killed on that day. Animal procedures were performed in accordance with the guidelines of the Ethics Committee on Animal Experimentation of the UCL Faculty of Medicine.
Statistics
Experiments were performed at least three times (unless otherwise stated) and produced similar results. When the average of the different experiments is represented, standard error of the mean (s.e.m) is indicated while if one representative experiment is shown, s.d. is specified. Normal distribution was tested and statistical analyses were performed using Student's t-test (*P<0.05; **P<0.01; ***P<0.001).
References
Zand DJ, Huff D, Everman D, Russell K, Saitta S, McDonald-McGinn D et al. Autosomal dominant inheritance of infantile myofibromatosis. Am J Med Genet A 2004; 126A: 261–266.
Mashiah J, Hadj-Rabia S, Dompmartin A, Harroche A, Laloum-Grynberg E, Wolter M et al. Infantile myofibromatosis: a series of 28 cases. J Am Acad Dermatol 2014; 71: 264–270.
Wiswell TE, Sakas EL, Stephenson SR, Lesica JJ, Reddoch SR . Infantile myofibromatosis. Pediatrics 1985; 76: 981–984.
Miklossy J, Mackenzie IR, Dorovini-Zis K, Calne DB, Wszolek ZK, Klegeris et al. Severe vascular disturbance in a case of familial brain calcinosis. Acta Neuropathol 2005; 109: 643–653.
Ang P, Tay YK, Walford NQ . Infantile myofibromatosis: a case report and review of the literature. Cutis 2004; 73: 229–231.
Chung EB, Enzinger FM . Infantile myofibromatosis. Cancer 1981; 48: 1807–1818.
Bracko M, Cindro L, Golouh R . Familial occurrence of infantile myofibromatosis. Cancer 1992; 69: 1294–1299.
Martignetti JA, Tian L, Li D, Ramirez MC, Camacho-Vanegas O, Camacho SC et al. Mutations in PDGFRB cause autosomal-dominant infantile myofibromatosis. Am J Hum Genet 2013; 92: 1001–1007.
Cheung YH, Gayden T, Campeau PM, Leduc CA, Russo D, Nguyen VH et al. A recurrent PDGFRB mutation causes familial infantile myofibromatosis. Am J Hum Genet 2013; 92: 996–1000.
Demoulin JB, Essaghir A . PDGF receptor signaling networks in normal and cancer cells. Cytokine Growth Factor Rev 2014; 25: 273–283.
Andrae J, Gallini R, Betsholtz C . Role of platelet-derived growth factors in physiology and medicine. Genes Dev 2008; 22: 1276–1312.
Soriano P . The PDGF alpha receptor is required for neural crest cell development and for normal patterning of the somites. Development 1997; 124: 2691–2700.
Soriano P . Abnormal kidney development and hematological disorders in PDGF beta-receptor mutant mice. Genes Dev 1994; 8: 1888–1896.
Verstraete K, Savvides SN . Extracellular assembly and activation principles of oncogenic class III receptor tyrosine kinases. Nat Rev Cancer 2012; 12: 753–766.
Chiara F, Bishayee S, Heldin CH, Demoulin JB . Autoinhibition of the platelet-derived growth factor beta-receptor tyrosine kinase by its C-terminal tail. J Biol Chem 2004; 279: 19732–19738.
Toffalini F, Demoulin JB . New insights into the mechanisms of hematopoietic cell transformation by activated receptor tyrosine kinases. Blood 2010; 116: 2429–2437.
Heldin CH, Ostman A, Ronnstrand L . Signal transduction via platelet-derived growth factor receptors. Biochim Biophys Acta 1998; 1378: F79–F113.
Lacerda Lda S, Alves UD, Zanier JF, Machado DC, Camilo GB, Lopes AJ . Differential diagnoses of overgrowth syndromes: the most important clinical and radiological disease manifestations. Radiol Res Pract 2014; 2014: 947451.
Takenouchi T, Yamaguchi Y, Tanikawa A, Kosaki R, Okano H, Kosaki K . Novel Overgrowth Syndrome Phenotype Due to Recurrent De Novo PDGFRB Mutation. J Pediatr 2015; 166: 483–486.
Heldin CH . Targeting the PDGF signaling pathway in tumor treatment. Cell Commun Signal 2013; 11: 97.
Chiara F, Goumans MJ, Forsberg H, Ahgren A, Rasola A, Aspenstrom P et al. A gain of function mutation in the activation loop of platelet-derived growth factor beta-receptor deregulates its kinase activity. J Biol Chem 2004; 279: 42516–42527.
Medves S, Duhoux FP, Ferrant A, Toffalini F, Ameye G, Libouton JM et al. KANK1, a candidate tumor suppressor gene, is fused to PDGFRB in an imatinib-responsive myeloid neoplasm with severe thrombocythemia. Leukemia 2010; 24: 1052–1055.
Cools J, DeAngelo DJ, Gotlib J, Stover EH, Legare RD, Cortes J et al. A tyrosine kinase created by fusion of the PDGFRA and FIP1L1 genes as a therapeutic target of imatinib in idiopathic hypereosinophilic syndrome. N Engl J Med 2003; 348: 1201–1214.
Nicolas G, Pottier C, Maltete D, Coutant S, Rovelet-Lecrux A, Legallic S et al. Mutation of the PDGFRB gene as a cause of idiopathic basal ganglia calcification. Neurology 2013; 80: 181–187.
Nicolas G, Pottier C, Charbonnier C, Guyant-Marechal L, Le Ber I, Pariente J et al. Phenotypic spectrum of probable and genetically-confirmed idiopathic basal ganglia calcification. Brain 2013; 136: 3395–3407.
Arts FA, Velghe AI, Stevens M, Renauld JC, Essaghir A, Demoulin JB . Idiopathic basal ganglia calcification-associated PDGFRB mutations impair the receptor signalling. J Cell Mol Med 2015; 19: 239–248.
Schonherr C, Ruuth K, Yamazaki Y, Eriksson T, Christensen J, Palmer RH et al. Activating ALK mutations found in neuroblastoma are inhibited by Crizotinib and NVP-TAE684. Biochem J 2011; 440: 405–413.
Nakahara M, Isozaki K, Hirota S, Miyagawa J, Hase-Sawada N, Taniguchi M et al. A novel gain-of-function mutation of c-kit gene in gastrointestinal stromal tumors. Gastroenterology 1998; 115: 1090–1095.
Heinrich MC, Corless CL, Duensing A, McGreevey L, Chen CJ, Joseph N et al. PDGFRA activating mutations in gastrointestinal stromal tumors. Science 2003; 299: 708–710.
Looman C, Sun T, Yu Y, Zieba A, Ahgren A, Feinstein R et al. An activating mutation in the PDGF receptor-beta causes abnormal morphology in the mouse placenta. Int J Dev Biol 2007; 51: 361–370.
Keating MT, Harryman CC, Williams LT . Platelet-derived growth factor receptor inducibility is acquired immediately after translation and does not require glycosylation. J Biol Chem 1989; 264: 9129–9132.
Demoulin JB, Uyttenhove C, Lejeune D, Mui A, Groner B, Renauld JC . STAT5 activation is required for interleukin-9-dependent growth and transformation of lymphoid cells. Cancer Res 2000; 60: 3971–3977.
Velghe AI, Van Cauwenberghe S, Polyansky AA, Chand D, Montano-Almendras CP, Charni S et al. PDGFRA alterations in cancer: characterization of a gain-of-function V536E transmembrane mutant as well as loss-of-function and passenger mutations. Oncogene 2014; 33: 2568–2576.
Corless CL, Schroeder A, Griffith D, Town A, McGreevey L, Harrell P et al. PDGFRA mutations in gastrointestinal stromal tumors: frequency, spectrum and in vitro sensitivity to imatinib. J Clin Oncol 2005; 23: 5357–5364.
David M, Cross NC, Burgstaller S, Chase A, Curtis C, Dang R et al. Durable responses to imatinib in patients with PDGFRB fusion gene-positive and BCR-ABL-negative chronic myeloproliferative disorders. Blood 2007; 109: 61–64.
Li-Wan-Po A, Farndon P, Craddock C, Griffiths M . Integrating pharmacogenetics and therapeutic drug monitoring: optimal dosing of imatinib as a case-example. Eur J Clin Pharmacol 2010; 66: 369–374.
Linhares ND, Freire MC, Cardenas RG, Bahia M, Puzenat E, Aubin F et al. Modulation of expressivity in PDGFRB-related infantile myofibromatosis: a role for PTPRG? Genet Mol Res 2014; 13: 6287–6292.
Barker KT, Houlston RS . Overgrowth syndromes: is dysfunctional PI3-kinase signalling a unifying mechanism? Eur J Hum Genet 2003; 11: 665–670.
Lindhurst MJ, Sapp JC, Teer JK, Johnston JJ, Finn EM, Peters K et al. A mosaic activating mutation in AKT1 associated with the Proteus syndrome. N Engl J Med 2011; 365: 611–619.
Millot F, Guilhot J, Baruchel A, Petit A, Leblanc T, Bertrand Y et al. Growth deceleration in children treated with imatinib for chronic myeloid leukaemia. Eur J Cancer 2014; 50: 3206–3211.
Demoulin JB, Louahed J, Dumoutier L, Stevens M, Renauld JC . MAP kinase activation by interleukin-9 in lymphoid and mast cell lines. Oncogene 2003; 22: 1763–1770.
Noel LA, Arts FA, Montano-Almendras CP, Cox L, Gielen O, Toffalini F et al. The tyrosine phosphatase SHP2 is required for cell transformation by the receptor tyrosine kinase mutants FIP1L1-PDGFRalpha and PDGFRalpha D842V. Mol Oncol 2014; 8: 728–740.
Medves S, Noel LA, Montano-Almendras CP, Albu RI, Schoemans H, Constantinescu SN et al. Multiple oligomerization domains of KANK1-PDGFRbeta are required for JAK2-independent hematopoietic cell proliferation and signaling via STAT5 and ERK. Haematologica 2011; 96: 1406–1414.
Acknowledgements
We would like to thank Nicolas Dauguet for cell sorting, and Anabelle Decottignies, Guy Warnier and Laure Dumoutier for generous donation of reagents. We are grateful to Ahmed Essaghir for statistical analysis. FAA is the recipient of a fellowship from FRS-FNRS – Télévie (grant number 7.4501.13 F).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Arts, F., Chand, D., Pecquet, C. et al. PDGFRB mutants found in patients with familial infantile myofibromatosis or overgrowth syndrome are oncogenic and sensitive to imatinib. Oncogene 35, 3239–3248 (2016). https://doi.org/10.1038/onc.2015.383
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2015.383
- Springer Nature Limited
This article is cited by
-
PDGFRA K385 mutants in myxoid glioneuronal tumors promote receptor dimerization and oncogenic signaling
Scientific Reports (2024)
-
Prenatal genetic diagnosis of disseminated infantile myofibromatosis: a case report and literature review
BMC Medical Genomics (2023)
-
Giant intracranial infantile myofibromatosis of the skull base: report of two cases
Child's Nervous System (2022)
-
Aggressive infantile myofibromatosis with intestinal involvement
Molecular and Cellular Pediatrics (2021)
-
Significant Antitumor Activity of Pazopanib in a Patient with PDGFRB-Mutated Metastatic Phyllodes Tumor: a Case Report
SN Comprehensive Clinical Medicine (2021)