Abstract
Dysregulation of ribosome biogenesis or translation can promote cancer, but the underlying mechanisms remain unclear. UTP18 is a component of the small subunit processome, a nucleolar multi-protein complex whose only known function is to cleave pre-ribosomal RNA to yield the 18S ribosomal RNA component of 40S ribosomal subunits. Here, we show that UTP18 also alters translation to promote stress resistance and growth, and that UTP18 is frequently gained and overexpressed in cancer. We observed that UTP18 localizes to the cytoplasm in a subset of cells, and that serum withdrawal increases cytoplasmic UTP18 localization. Cytoplasmic UTP18 associates with the translation complex and Hsp90 to upregulate the translation of IRES-containing transcripts such as HIF1a, Myc and VEGF, thereby inducing stress resistance. Hsp90 inhibition decreases cytoplasmic UTP18 and UTP18-induced increases in translation. Importantly, elevated UTP18 expression correlates with increased aggressiveness and decreased survival in numerous cancers. Enforced UTP18 overexpression promotes transformation and tumorigenesis, whereas UTP18 knockdown inhibits these processes. This stress adaptation mechanism is thus co-opted for growth by cancers, and its inhibition may represent a promising new therapeutic target.
Similar content being viewed by others
Introduction
Both normal and neoplastic cells must balance the synthesis of new biomolecules with the need for those biomolecules in activities that require energy consumption such as cell growth, migration or proliferation. Ribosome synthesis and protein translation are closely coordinated, but how this is achieved is poorly understood. Dysregulation of any of these processes can lead to tumor formation,1, 2, 3 presumably as compensatory adjustments are made that avoid cellular demise and lead instead to unrestrained growth. For example, overexpression of certain translation initiation-related proteins can transform cells.3 For ribosomal proteins, loss of function mutations or gene deletions can promote cancer.1, 2 In each instance, changes in ribosome function or translation must be integrated effectively with other cellular processes to sustain growth. Identification of the processes that enable such integration provides new insights into cellular homeostasis and may yield additional strategies for therapeutically manipulating cell survival or growth.
UTP18 (WDR50/CGI-48) is an essential component of the small subunit processome, a nucleolar multi-protein complex whose only known function is the sequential cleavage of pre-ribosomal RNA to form 18S ribosomal RNA, a key component of 40S ribosomal subunits.4 UTP18 is highly conserved from yeast to mammals, and the loss of UTP18 function prevents the production of 18S ribosomal RNA.4 In drosophila, UTP18 regulates stem cell size, survival and self-renewal, although the mechanisms by which this occurs are unclear.5
Here, we present the first evidence that UTP18 not only regulates pre-ribosomal RNA processing in the nucleolus, but it also localizes to the cytoplasm in response to stress. There, it preferentially upregulates internal ribosome entry site (IRES)-dependent translation, induces stress resistance and promotes growth. We also show that UTP18 is gained and overexpressed with remarkable frequency in a range of human cancers, and it promotes tumor aggressiveness and decreases patient survival by increasing stress resistance in tumor cells.
Results
UTP18 is gained in human cancers, and its overexpression promotes transformation
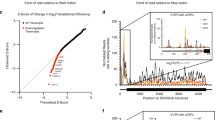
During a genome-wide, high-resolution single nucleotide polymorphism analysis of meningioma, we identified a focal 17q21.33 amplicon that was present exclusively in the more aggressive (but not low-grade) tumors, and that only contained UTP18 (Figure 1a). Studies of neuroblastoma, medulloblastoma and other cancers have previously suggested the existence of an unidentified oncogene in this region of chromosome 17q.6, 7, 8 We therefore analyzed publicly available array comparative genomic hybridization profiles from 257 neuroblastomas and found a common region of gain containing UTP18 at a 17q21.33 breakpoint (Figure 1b). Immunocytochemistry of paraffin-embedded sections from several cancer types and normal tissues revealed increased UTP18 expression in cancers with frequent gains of UTP18 such as neuroblastoma, melanoma and breast carcinoma when compared with cancers that have infrequent UTP18 gains such as ovarian cancer (Figure 1c). To examine the functional consequences of elevated UTP18 expression, we overexpressed human UTP18 in mouse NIH-3T3 fibroblasts. UTP18 overexpression transformed NIH-3T3 cells, as evidenced by subcutaneous tumor growth in nude mice (Figure 1d). Analysis of ribosomal RNA from these cells or from human NLF neuroblastoma cells overexpressing UTP18 indicated that UTP18 overexpression did not significantly increase the production of 18S ribosomal RNA (Supplementary Figure S1). In contrast, UTP18 depletion in IMR5 neuroblastoma cells led to the expected accumulation of incompletely processed pre-ribosomal RNA fragments (Supplementary Figure S1) and increased the accumulation and activation of p53 (Supplementary Figure S2), consistent with previous reports.4, 9 UTP18 overexpression, however, had no effect on p53 levels in neuroblastoma cells (Supplementary Figure S2). Taken together, these data indicate that UTP18 is frequently gained in neuroblastomas and aggressive meningiomas, and that enforced UTP18 overexpression promotes transformation via mechanisms that do not involve increased ribosomal RNA synthesis or p53 activation.
UTP18 is frequently gained in cancer and promotes transformation. (a) 500 K single nucleotide polymorphism analysis of 47 primary meningiomas and 12 matched blood samples. Data from WHO benign Grade I, recurrent Grade I (IR), Grade II and Grade III meningiomas. Arrow identifies amplicon containing UTP18/WDR50. (b) Array comparative genomic hybridization profiles from 257 human neuroblastomas displaying region of gain on chromosome 17 q. Vertical line indicates UTP18 location. (c) UTP18 immunohistochemistry in paraffin-embedded sections from neuroblastoma, hepatocellular carcinoma, breast carcinoma, ovarian carcinoma and the corresponding normal tissues. Some images obtained from www.proteinatlas.org. Scale=25 μm. (d) Subcutaneous transplants of control (C) or UTP18-overexpressing (U) NIH-3T3 cells were performed in nude mice and followed for 4 weeks. Only the UTP18-expressing cells formed tumors.
UTP18 localizes to the cytoplasm as well as the nucleolus in human cells
UTP18 has classically been described as a nucleolar protein.4 However, we observed endogenous UTP18 immunoreactivity in both the nucleolus and the cytoplasm of a subpopulation of human neuroblastoma, hepatocellular carcinoma, melanoma and breast carcinoma cells in paraffin-embedded specimens (Figures 1c and 2a). A similar pattern of UTP18 immunoreactivity was seen in cultured human IMR5 neuroblastoma cells (Figure 2b). Confocal microscopy of UTP18 immunoreactivity in HEK 293 cells clearly identified UTP18 immunoreactivity in the cytoplasm of some cells but not others (Figure 2c). Overexpression of a human UTP18-GFP fusion protein in HEK 293 cells confirmed that UTP18 localized to both the nucleolus and the cytoplasm in a subpopulation of cells, and to the nucleus and nucleolus in a separate subpopulation (Figure 2d and Supplementary Figure S3). Furthermore, separation of HeLa cells into nuclear and cytoplasmic fractions confirmed that endogenous UTP18 resides in both the cytoplasm and the nucleus in mammalian cells (Figure 2e). This finding raised the possibility that, in addition to its nucleolar role as a small subunit processome component, UTP18 may have functions outside the nucleolus.
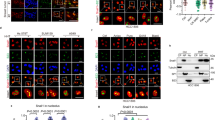
UTP18 localizes to the cytoplasm and the nucleolus. (a) UTP18 immunohistochemistry in paraffin-embedded section from neuroblastoma, hepatocellular carcinoma, melanoma and breast carcinoma. Arrows identify cells with cytoplasmic UTP18 immunoreactivity. Scale ~15 μm. (b) Cytoplasmic (arrows) and nucleolar UTP18 immunoreactivity in cultured human IMR5 neuroblastoma cells. Nuclei stained with DAPI (blue). Scale ~10 μm. (c) Confocal images obtained from the Human Protein Atlas illustrating subcellular localization of UTP18 immunoreactivity in cultured HEK 293 cells. Closed arrows identify cells with cytoplasmic UTP18. Open arrows identify a cell with no cytoplasmic UTP18. UTP18 fluorescence intensity map is shown for comparison. Endoplasmic reticulum stain (calreticulin, yellow) and microtubule stain (tubulin, red) are also shown. (d) Cytoplasmic and nucleolar (closed arrow) versus nuclear (open arrow) UTP18-GFP localization in HEK 293 cells. Scale ~8 μm. (e) Western blot identification of UTP18 after subcellular fractionation of human IMR5 neuroblastoma cells. Cytoplasmic (p-FOXO3a, p53) and nuclear (c-Myc, p53) proteins indicate efficiency of separation.
The variation in UTP18 localization observed from cell to cell in paraffin-embedded tumor specimens and in cultured cells suggested that the subcellular localization of UTP18 may be regulated by extracellular or intracellular factors. Indeed, we observed that serum withdrawal for 24 h increased the cytoplasmic localization of UTP18 in HEK 293 cells (Figure 3a). Western blot analysis of nuclear and cytoplasmic fractions from these cells after serum withdrawal confirmed that the ratio of cytoplasmic to nuclear UTP18 increased under these conditions (Figure 3b).
Dynamic regulation of UTP18 localization. (a) Fluorescence immunocytochemistry illustrating the effect of serum-free medium on the localization of UTP18 immunoreactivity in HEK 293 cells. Nuclei stained with DAPI (blue). Scale=15 μm. (b) Western blot illustrating the effect of serum withdrawal on the subcellular localization of UTP18. Cells were exposed to serum-free medium for 24 h prior to collecting the cells and fractionating them into nuclear and cytoplasmic fractions. Protein lysates were prepared for western blots. Tubulin and fibrillarin were used for cytoplasmic or nuclear normalization, respectively, and to determine the efficiency of fractionation. Densitometry was performed for quantitation. Densitometry ratios shown are the ratio of cytoplasmic UTP18 to tubulin divided by the ratio of nuclear UTP18 to fibrillarin.
UTP18 increases IRES-dependent translation
The lack of effect of UTP18 overexpression on ribosomal RNA processing and p53 levels, combined with the unexpected observation that a significant amount of UTP18 can be found in the cytoplasm, prompted a search for other mechanisms that might underly UTP18’s ability to transform cells. We overexpressed UTP18-FLAG in HEK 293 cells and used immunoprecipitation to identify UTP18-interacting proteins. Mass spectrometry analysis identified several nucleolar proteins involved in ribosome synthesis.4, 9 However, many of the proteins identified were cytoplasmic proteins that participate in translation initiation or mRNA splicing, including eIF2B, eIF3B, eIF3E, eIF3F, eIF4A2, PTBP1 and others (Supplementary Table 1). Immunoprecipitation of endogenous UTP18 followed by western blot for eIF3B and PTBP1 confirmed this finding (Figure 4a).
UTP18 associates with the translation complex. (a) Western blot validation of UTP18-interacting proteins in HEK 293 cells. Left panel, UTP18 immunoprecipitation and eIF3B western blot. Right panels, PTBP1 or UTP18 immunoprecipitation and UTP18 or PTBP1 western blot. (b) Upper panels, ribosomal proteins and RNA in sucrose gradient centrifugation fractions from IMR5 cells. We used antibodies to identify proteins associated with specific stages of ribosome biogenesis. RIO2 (pre-40S), RPS7 (40S), RPL13 (60S), MPP10 (pre-18S, 60 s-80S), eIF4G (translation initiation complex), eIF3b (translation initiation complex). Lower panel, UTP18 in IMR5 sucrose gradient fractions after adding RNAse before centrifugation. (c) UTP18 effect on ribosomes and polysomes after exposure of IMR5 cells to vehicle (left panel) or the translation inhibitor, 4EGI-1 (right panel). Note that UTP18 increases polysome loading (increased optical density indicated by orange line, left panel) when compared with control. Blockade of translation initiation using 4EGI-1 (right panel) abolishes this difference.
To further investigate the interaction between UTP18 and the translation machinery, we performed sucrose gradient centrifugation of neuroblastoma cell lysates. UTP18 was associated primarily with the pre-40S, 40S and, to a lesser extent, the 80S and polysomal fractions (Figure 4b). The addition of RNAse prior to centrifugation disrupted this pattern of UTP18/ribosomal protein association, revealing that this pattern depends upon the formation of protein/RNA complexes (Figure 4b). Whereas UTP18 overexpression did not increase the abundance of 40S ribosomes, it did increase polysome loading (Figure 4c). This effect was antagonized by the translation inhibitor, 4EGI-1.10
We next examined the effect of UTP18 on protein translation in HeLa cells expressing a bicistronic vector that encoded renilla luciferase under the control of the cytomegalovirus (CMV) promoter and firefly luciferase under the control of mammalian IRES sequences from c-Myc, VEGF or HIF1a.11, 12 A vector containing a stop codon and no IRES sequence (pRF) was used for comparison. The cells were then transfected to express full length UTP18, a truncated form of UTP18 lacking the N-terminal 200 amino acids (which localizes diffusely to the cytoplasm and the nucleus) or an empty control vector. Whereas UTP18 overexpression increased cap-dependent translation by up to 50% (Figure 5a, P<0.0001, unpaired t-test), it increased IRES-dependent translation by more than 200% (P<0.0001, unpaired t-test). Moreover, UTP18 increased translation of the downstream firefly luciferase more than cap-dependent translation of the renilla luciferase only when an IRES sequence was present (P<0.0001, unpaired t-test). In contrast to UTP18, overexpression of an N-terminal deleted form of UTP18 that failed to transform NIH-3T3 cells (data not shown) was significantly less effective in increasing translation (Figure 5a). When HEK 293 cells overexpressing UTP18 or a control vector were transfected with a separate bicistronic vector containing red fluorescent protein under the control of the CMV promoter and green fluorescent protein (GFP) under the control of the encephalomyocarditis virus IRES sequence, UTP18 increased the ratio of GFP to red fluorescent protein, again indicating that UTP18 induces a relative increase in IRES-dependent translation (Supplementary Figure S4).
UTP18 upregulates IRES-dependent translation. (a) Effect of overexpressing vectors encoding human UTP18, a truncated form of UTP18 lacking the N-terminal 200 amino acids (del-UTP), or control on the translation of bicistronic CMV-renilla/IRES-firefly luciferase vectors in HeLa cells. Mammalian cellular IRES sequences for VEGF (pRvegfF), Myc (pRmycF) and Hif1a (pRhif1F) were used. A control vector lacking an IRES sequence (pRF) was also used. Data are normalized to control cap-renilla luciferase activity for each vector. Data shown are mean±s.e.m. of three replicates. Note that UTP18 significantly increased cap-dependent and IRES-dependent translation when compared to control for all four vectors (P<0.0001, unpaired t-test), but increased IRES-dependent translation greater than cap-dependent translation only when an IRES sequence was present (P<0.0001, unpaired t-test). (b) Effect of in vitro translated UTP18 or a control (NF2) on the in vitro translation of encephalomyocarditis virus-IRES-luciferase mRNA. Data shown are mean±s.e.m. of six replicates (P<0.0001, unpaired t-test). (c) Left upper panel, western blot illustrating UTP18 co-immunoprecipitation with 5’- mRNA cap using IMR5 neuroblastoma lysates. Left lower panel, UTP18 overexpression induced co-immunoprecipitation of c-Myc IRES mRNA with eIF4G. Upper blot shows RNA gel electrophoresis, lower blot shows protein (western blot). Right lower panel, western blot illustrating effect of UTP18 overexpression on c-Myc expression. (d) Polysome profiles illustrating effect of UTP18 overexpression on IRES-containing (c-MYC, eIF4G and LEF1) and non-IRES-containing (MTOR, DICER1, ALK3) mRNAs. mRNA levels determined by real-time PCR. (e) UTP18-induced increase in proteins derived from IRES-containing mRNAs. (f) Flow cytometry illustrating effect of UTP18 overexpression on human neuroblastoma cell apoptosis induced by exposure to H2O2, hypoxia or low glucose medium. At least 3000 cells were counted for each condition, and the percentage of apoptotic cells was determined (*P<0.01, proportion test). (g) Effect of UTP18 overexpression (left panel) or UTP18 knockdown (right panel) on glutaminase (GLS) or pyruvate kinase (PKM1/PKM2) expression in HEK 293 or SH-SY5Y neuroblastoma cells. (h) Effect of UTP18 overexpression on lactate and ATP production in human neuroblastoma cells. Data shown are the mean±s.e.m. of six replicates (*P<0.05, unpaired t-test).
To determine whether UTP18 regulates translation directly, we used a mammalian cell-free in vitro translation assay. An mRNA containing the T7 promoter and the encephalomyocarditis virus-IRES sequence upstream of luciferase was transcribed in vitro and added as a reporter template. To generate UTP18, we transcribed UTP18 mRNA in vitro and added this to the translation reaction mixture. As a control, an equivalent amount of a similarly sized in vitro transcribed control mRNA (NF2) was used. Compared with the control, UTP18 increased IRES-dependent translation in this cell-free system more than 250% (Figure 5b, P<0.0001, unpaired t-test).
Using IMR5 neuroblastoma cell lysates, we observed that endogenous UTP18 co-immunoprecipitated with the 5’-mRNA cap (Figure 5c). In HeLa cells, UTP18 overexpression increased association of the translation initiation factor, eIF4G, with the c-Myc IRES RNA (Figure 5c). UTP18 also increased c-Myc protein levels (Figure 5c), but no significant increase in c-Myc mRNA was detected (Supplementary Figure S5).
Polysome profiling revealed that UTP18 shifted several IRES-containing transcripts to heavier polysome fractions while shifting non-IRES-containing transcripts to lighter fractions, consistent with our earlier finding that UTP18 preferentially increases the translation of IRES-containing mRNAs (Figure 5d). Accordingly, UTP18 increased the expression of several proteins derived from IRES-containing transcripts, including VEGF, Bcl2 and HIF1a (Figure 5e) as well as c-Myc (Figure 5c).11, 12 RNA-Seq analysis of mRNA from NIH-3T3 cells overexpressing UTP18 (Supplementary Table 2) or real-time PCR of c-Myc mRNA levels in neuroblastoma cells overexpressing UTP18 (Supplementary Figure S5) indicated that UTP18 overexpression caused little change in the mRNA levels for these proteins.
UTP18 associates with Hsp90 to alter translation and induce stress resistance
Cells increase IRES-dependent translation as part of an adaptive response to environmental stress,13 and our earlier observations indicated that serum withdrawal alters the subcellular localization of UTP18. We therefore examined whether UTP18 participates in the cellular stress response. UTP18 overexpression significantly decreased neuroblastoma cell death occurring after exposure to hydrogen peroxide, hypoxia or glucose deprivation (Figure 5f). In addition, UTP18 overexpression increased (and UTP18 depletion decreased) the expression of glutaminase and expression of the PKM2 isoform of pyruvate kinase relative to PKM1 (Figure 5g), key enzymes in glutaminolysis and aerobic glycolysis, respectively.14 Knockdown of UTP18 in human SH-SY5Y neuroblastoma cells decreased the expression of these enzymes. Consistent with these observations, UTP18 increased the cellular production of lactate, an end product of aerobic glycolysis (Figure 5h). Of note, aerobic glycolysis and glutaminolysis often increase during cell transformation and have been implicated in cancer metabolism.14
We further tested the hypothesis that cytoplasmic localization of UTP18 is required for UTP18’s effect on translation. We identified a putative nuclear localization sequence within the first 24 amino acids of UTP18. Deletion of this short sequence abolished UTP18-GFP nuclear localization (Figure 6). Next, we serially deleted each of the six C-terminal WD40 domains of UTP18. Both nuclear and cytoplasmic localization was retained after co-deletion of WD40 domains 4, 5 and 6. However, co-deletion of WD40 domains 3, 4, 5 and 6 abolished cytoplasmic localization of UTP18 (Figure 6). When tested in a cell-based translation reporter assay, overexpression of the cytoplasmic UTP18-GFP mutant lacking the N-terminal nuclear localization sequence nevertheless promoted IRES-dependent translation. In general, UTP18-GFP mutants that displayed cytoplasmic localization tended to increase translation, whereas those with nuclear localization inhibited translation. The mechanism of translation inhibition by UTP18 deletion mutants displaying exclusive nuclear localization is unclear, although one possibility is that they interfere with 18S ribosomal RNA processing.
UTP18-mediated increase in translation is associated with UTP18 cytoplasmic localization. Left, diagram depicting the structures of serial UTP18-GFP deletion mutants and corresponding fluorescence micrographs illustrating the subcellular localization of each UTP18-GFP deletion mutant in HEK 293 cells. Lower right, color-coded bar graph quantitating the effect of each UTP18-GFP deletion mutant on the translation of a Myc-IRES-luciferase vector in HEK 293 cells. Mean±s.e.m. of six replicates. *P<0.05, t-test. Statistical significance calculated relative to control. n.s.=not significant.
Our earlier findings indicated that serum withdrawal increases cytoplasmic UTP18 localization (Figure 3). This raised the possibility that stress signals may alter the cytoplasmic localization of UTP18, thereby regulating its ability to interact with the translation machinery and activate a stress response program. To investigate this possibility, we used mass spectrometry analysis of UTP18 immunoprecipitates to look for UTP18-interacting proteins that have been implicated in the cellular response to stress. We identified the 90 kDa heat shock protein (Hsp90) as one of the most abundant UTP18-interacting proteins (Supplementary Table S1). Hsp90 is a stress-induced molecular chaperone that regulates the conformation, stability and/or activity of dozens of cellular proteins via its intrinsic ATPase activity.15 Cellular subfractionation indicated that both UTP18 and Hsp90 were present together in the cytoplasm (Figure 7a), and immunoprecipitation of endogenous UTP18 followed by western blot confirmed the association of Hsp90 with UTP18 (Figure 7b). Importantly, the small molecule Hsp90 inhibitor, 17-N-allylamino-17-demethoxygeldanamycin (250 nM), increased the nuclear localization of UTP18 (Figure 7c) and antagonized the ability of UTP18 to preferentially increase translation from the HIF1a IRES sequence (Figure 7d, P<0.0016, unpaired t-test). In addition, Hsp90 inhibition abrogated the increase in IRES-dependent translation caused by the MTOR inhibitor, rapamycin (Figure 7e, P<0.0025, unpaired t-test),16 and it worked synergistically with rapamycin (50 nM) to inhibit the growth of human CH157 malignant meningioma cells, even though neither inhibitor alone had a statistically significant effect on growth (Figure 7f, P<0.0001, unpaired t-test).
Hsp90 regulates the localization of UTP18 and its effect on translation. (a) Western blot demonstrating expression of Hsp90 and UTP18 in nuclear and cytoplasmic lysates after subcellular fractionation of HEK 293 cells. (b) Immunoprecipitation of endogenous UTP18 from total cell lysates derived from human IMR5 neuroblastoma cells, followed by western blot for UTP18 and Hsp90. (c) Fluorescence immunocytochemistry illustrating the effect of 17-N-allylamino-17-demethoxygeldanamycin (10 nM, 6 h exposure) on nuclear localization of UTP18 in HEK 293 cells in vitro. Nuclei stained with DAPI (blue). Scale is approximately 20 μm. (d) Effect of 17-N-allylamino-17-demethoxygeldanamycin (250 nM) on the UTP18-induced increase in IRES-dependent translation of a bicistronic CMV-renilla/HIF1-IRES/firefly luciferase vector in HEK 293 cells. Data shown are mean±s.e.m. of three replicates, *P<0.0016, unpaired t-test. (e) Effect of 17-N-allylamino-17-demethoxygeldanamycin (250 nM) and the MTOR inhibitor, rapamycin (50 nM) on IRES-dependent translation of a bicistronic CMV-renilla/VEGF IRES-firefly luciferase vector in HEK 293 cells. Data shown are mean±s.e.m. of four replicates, *P<0.0025, unpaired t-test. (f) MTT cell growth assay illustrating effect of 17-N-allylamino-17-demethoxygeldanamycin (250 nM) and rapamycin (50 nM) on the growth of human CH157 meningioma cells. Data shown are mean±s.e.m. of six replicates. *P<0.0001, unpaired t-test when compared with control.
UTP18 overexpression correlates with tumor aggressiveness and promotes tumorigenesis
Our earlier observation that UTP18 is frequently gained in aggressive meningiomas, neuroblastomas and other cancers, combined with our finding that UTP18 overexpression promotes transformation and cell survival, suggested that UTP18 promotes growth in human cancers. Indeed, when overexpressed in human NLF neuroblastoma cells transplanted subcutaneously into nude mice, UTP18 increased neuroblastoma growth (Figure 8a and Supplementary Figure S6, P=0.05, unpaired t-test). Conversely, shRNA-mediated UTP18 knockdown in human SH-SY5Y neuroblastoma cells (which harbor a gain of UTP18)17 caused cell differentiation in vitro and significantly inhibited neuroblastoma growth in mice (Figure 8b, P<0.001, unpaired t-test). UTP18 knockdown in SK-N-AS neuroblastoma cells also inhibited cell growth in vitro and decreased subcutaneous tumor growth in vivo (Supplementary Figure S6).
UTP18 promotes tumorigenesis and correlates with decreased survival in numerous cancers. (a) Effect of UTP18 overexpression on the subcutaneous growth of human NLF neuroblastoma xenografts in nude mice after 4 weeks (1356±564 versus 4690±1387 mm3, mean±s.e.m., n=6, P<0.05, unpaired t-test). (b) Upper panels, SH-SY5Y neuroblastoma cells overexpressing control shRNA or a UTP18-shRNA in vitro. Scale is approximately 20 μm. Lower panel, subcutaneous tumor formation by transplanted SH-SY5Y neuroblastoma cells overexpressing control shRNA or UTP18 shRNA in mice (P<0.001, t-test). (c) Upper panel, UTP18 gain frequency in 3796 array comparative genomic hybridization profiles from 16 cancer types. Lower panel, real-time PCR of UTP18 mRNA expression in 38 primary tumor or matched normal specimens. (d) Kaplan–Meier analyses of UTP18 mRNA expression (203721_s_at) and survival in 88 neuroblastoma patients (left, P=0.008, Logrank) or 236 breast cancer patients (right, P=0.022, Logrank). Neuroblastoma (Versteeg) or breast cancer (Zhang) datasets obtained from http://hgserver1.amc.nl/cgi-bin/r2/main.cgi. (e) Correlation between UTP18 mRNA expression and relapse in 204 medulloblastoma patients. Samples are ordered based upon UTP18 expression. Clinical parameters shown include clinical risk, gender, presence or absence of metastases, molecular subgroup as defined by non-negative matrix factorization analysis and relapse. Color code key for each parameter is provided. Data and analysis obtained from http://hgserver1.amc.nl/cgi-bin/r2/main.cgi. When the upper and lower deciles of UTP18 expression were compared, increased UTP18 expression correlated with increased relapse (P=0.007, proportion test).
Analysis of 3796 genome-wide array comparative genomic hybridization profiles from multiple cancer types revealed a remarkably high frequency of gains of UTP18 in neuroblastoma (80%), medulloblastoma (45%), melanoma (38%), hepatocellular carcinoma (25%), breast carcinoma (18%) and several other cancers (Figure 8c). Real-Time PCR using mRNA obtained from primary tumors indicated that increased UTP18 mRNA expression was associated with increasing histologic grade in meningioma and breast carcinoma where UTP18 gain is frequent, but not in ovarian carcinoma which generally lacks gains of UTP18 (Figure 8c). Furthermore, analysis of publicly available gene expression datasets18, 19 revealed a significant correlation between increased UTP18 mRNA expression and decreased survival of neuroblastoma and breast cancer patients (Figure 8d). In medulloblastoma, elevated UTP18 mRNA expression (upper decile) was significantly associated with increased risk of relapse when compared with the lowest decile (Figure 8e).19 There were also twice as many patients with metastases in the upper decile when compared with the lower decile, and this pattern persisted when the upper 20th percentile (in terms of UTP18 mRNA expression) was compared with the lowest 20th percentile (Figure 8e). Taken together, these findings reveal a role for UTP18 in promoting aggressiveness in a range of human cancers.
Discussion
These data reveal a new adaptive mechanism that couples ribosomal RNA processing to translation to maintain cell growth and survival under stressful conditions. Whereas decreased UTP18 expression inhibits ribosomal RNA maturation and activates p53 to inhibit growth,4, 9 elevated UTP18 preferentially increases IRES-dependent translation to promote cell survival.12, 20, 21 Our findings that UTP18 associates with proteins of the translation pre-initiation complex and regulates translation in intact cells and in a cell-free system indicate a direct effect of UTP18 on translation. This represents the first evidence that a nucleolar small subunit processome component participates directly in translation, and it reveals a novel mechanism by which cells coordinate 40S ribosomal subunit production and protein synthesis.
Our data suggest that cellular stress enables the effect of UTP18 on translation by increasing its cytosolic localization in an Hsp90-dependent manner. Small molecule inhibition of the Hsp90 molecular chaperone function promotes re-localization of UTP18 to the nucleus, and it antagonizes the ability of UTP18 to increase IRES-dependent translation. Hsp90 plays a key role in the stress response via myriad sites of action. Like UTP18, Hsp90 has been reported to associate with the 5’mRNA cap, to regulate cap-dependent translation and to interact with eIF4G.22, 23 In addition, an interaction between Hsp90 and eIF3 is said to be necessary for translation from the hepatitis C virus IRES site.24 Thus, we propose that UTP18 and Hsp90 cooperate to preferentially upregulate the translation of IRES-containing transcripts such as HIF1a, Myc, VEGF, Bcl2 and others.
Remarkably, elevated UTP18 expression reversibly triggers alternative splicing of pyruvate kinase and increases the expression of glutaminase. Because previous studies have indicated that these changes may play an important role in sustaining cancer cells,14 manipulation of UTP18 may represent a new approach to altering the expression of pyruvate kinase isoforms or glutaminase expression. UTP18 knockdown also decreases the production of 18 s ribosomal RNA and upregulates p53 which, in turn, promotes growth arrest or apoptosis.9 Together, these effects likely account for our observation that UTP18 depletion induces neuroblastoma cell differentiation in vitro and inhibits neuroblastoma xenograft growth in mice. They may also help to explain a previous report that the loss of UTP18 function induces premature differentiation and decreases germline stem cell number in drosophila.5
This adaptive UTP18-dependent mechanism for sustaining cell growth and survival during cellular stress is co-opted in human cancers. Focal gains of UTP18 in aggressive meningiomas (and broader gains on 17q involving UTP18 in medulloblastomas,25 neuroblastomas6, 18 and other tumor types) identify UTP18 as a frequent target of copy number alteration in numerous cancers. The frequency of UTP18 gain is remarkably high in some cancer types, exceeding 80% in neuroblastoma and 40% in medulloblastoma. In general, these are low-level gains that result in one additional copy of UTP18. This is consistent with our observation that very high levels of UTP18 protein expression are toxic to cells (see Figure 5f), whereas slightly elevated UTP18 levels protect cells from stressful stimuli. Elevated UTP18 mRNA is associated with higher histologic grade and decreased patient survival in meningioma, neuroblastoma and breast carcinoma. Likewise, the increased risk of relapse associated with high UTP18 expression in medulloblastoma may reflect the increased ability of UTP18-expressing cells to survive stressful stimuli such as chemotherapeutic agents.
Taken together, our findings suggest a model in which constitutive UTP18 overexpression due to genomic gain or cellular stress increases cytoplasmic UTP18. This upregulates IRES-dependent translation of transcripts that promote growth and stress resistance, thereby increasing tumor aggressiveness and decreasing patient survival (Supplementary Figure S7). This UTP18-activated stress resistance pathway may confer therapeutic resistance in tumors in which UTP18 is overexpressed. Interruption of this pathway via Hsp90 inhibition may thus represent a useful new therapeutic strategy in susceptible cancers. Indeed, Hsp90 inhibitors have been reported to decrease neuroblastoma and medulloblastoma growth in animal models.26, 27, 28 These findings predict that inhibition of the UTP18-activated stress response pathway will sensitize tumors harboring gains of UTP18 to other chemotherapeutic agents, as we observed with rapamycin in the current study.
Materials and methods
Genome-scale expression data and analysis
A genome-wide 500 K single nucleotide polymorphism analysis of 47 meningioma specimens was performed as described previously.29 Array comparative genomic hybridization data analysis of neuroblastoma genomes was performed as we have described previously.30, 31 Additional data for Kaplan–Meier survival analyses in neuroblastoma, breast cancer and medulloblastoma were obtained at http://hgserver1.amc.nl/cgi-bin/r2/main.cgi19 using publicly available software.
Cell lines and tumor specimens
Human SK-N-AS and SH-SY5Y neuroblastoma cell lines were kindly provided by Dr R George. Human IMR-5 and NLF neuroblastoma cell lines were kindly provided by Dr J Maris. Human CH157-MN meningioma cells were provided by Dr D Gutmann. HeLa, HEK 293 and mouse NIH-3T3 cells were obtained from the American Tissue Type Culture Collection (ATCC) (Manassas, VA, USA). The cells were maintained at 37 °C in a 5% CO2 atmosphere using growth medium supplemented with 10% serum. In some experiments, cells were briefly exposed to serum-free or low-glucose media. A tissue microarray containing multiple paraffin-embedded neuroblastoma specimens and other tissues was purchased from US Biomax (Rockville, MD, USA). A collection of cDNAs derived from total RNA isolated from a range of primary human cancers was purchased from Origene (Rockville, MD, USA).
Western blots, immunocytochemistry, immunohistochemistry and immunoprecipitation
Western blots were performed as described.30 Briefly, total protein was extracted from cells using RIPA buffer. In some cases, protein present in sucrose gradient fractions was used. The protein was separated by gel electrophoresis, transferred to nitrocellulose membranes and probed overnight using the appropriate, commercially available primary antibodies. β-actin was used as the loading reference. After incubation in the appropriate secondary antibodies, immunoreactive bands were visualized using chemiluminescence. Antibodies used in this study include: UTP18 and Rio2, Bethyl Laboratories (Montgomery, TX, USA); RSP7, Abnova (Walnut, CA, USA); RSP13, Sigma (St Louis, MO, USA); eIF3B, Novusbio (Littleton, CO, USA); VEGF and MPP10, Santa Cruz Biotechnology (Dallas, TX, USA); PKM1, PKM2 and GLS, Proteintech (Chicago, IL, USA); eIF4G and Bcl-2, Cell Signaling Technology (Danvers, MA, USA); PTBP1, Aviva Systems Biology (San Diego, CA, USA).
Immunocytochemistry of cultured cells was performed as we have described previously.32 For immunohistochemistry, a tissue microarray containing multiple paraffin-embedded neuroblastoma specimens and other tissues (US Biomax) was used. After de-paraffination, the sections were incubated overnight using a specific anti-UTP18 antibody (Bethyl Laboratories). After washing and incubation in the appropriate peroxidase-labeled secondary antibody, specific immunoreactivity was visualized using diaminobenzidine and peroxide. Sections were counterstained with eosin. Where indicated, images of paraffin-embedded human tissues or cultured cells stained for UTP18 immunoreactivity were obtained from a publicly available database (The Human Protein Atlas, www.proteinatlas.org). In such cases, a rabbit polyclonal anti-UTP18 antibody (HPA052378, Sigma Aldrich, St Louis, MO, USA) was used, and sections were counterstained with hematoxylin.
For immunoprecipitation experiments, whole-cell lysates from cultured cells overexpressing UTP18-FLAG were immunoprecipitated with 1 μg of anti-FLAG antibody or UTP18 antibody prebound to 30 μl of Protein A-Sepharose 4B (GE Healthcare, Piscataway, NJ, USA). Appropriate isotype antibodies were used as controls. The beads were washed and boiled in sodium dodecyl sulfate buffer, and the eluates were processed for western blot or coomassie blue gel electrophoresis and mass spectrometry.
RNA immunoprecipitation studies
For RNA immunoprecipitation experiments, whole-cell lysates from cultured control cells or cells overexpressing UTP18 were immunoprecipitated with 1 μg of EIF4G antibody prebound to 30 μl of Protein A-Sepharose 4B (GE Healthcare). The beads were washed with RIPA buffer and total RNA was isolated using trizol. RT–PCR for the cMyc IRES sequence was then performed.
Cloning and generation of expression vectors
Human UTP18 was cloned from HEK 293 cells and confirmed by direct sequencing. A form of UTP18 lacking the N-terminal 200 amino acids was also prepared. UTP18-GFP, UTP18-IRES-GFP, UTP18-Myc and UTP18-FLAG fusion proteins were generated by subcloning human UTP18 into pCMV6-AC-GFP, pCMV-Tag1 or pLenti-IRES-GFP vectors. Lentiviral control or UTP18 shRNA vectors were purchased (TRCN0000144750 and TRCN0000145055, Open Biosystems, Lafayette, CO, USA) and packaged into lentiviruses as we have described previously.30
Protein translation assays
Cell-based translation assays utilized bicistronic luciferase or red fluorescent protein/GFP reporter vectors containing an upstream CMV promoter and downstream encephalomyocarditis virus, c-Myc, HIF1a or VEGF IRES sequences as described previously.33, 34, 35, 36 We also used a commercially available, cell-free in vitro translation assay derived from human cell lysates according to the manufacturer’s protocol (Thermo Scientific, Rockford, IL, USA). In vitro transcribed encephalomyocarditis virus-luciferase mRNA and control (NF2) or UTP18 mRNA were added to this mixture. Luminescence from the activity of translated luciferase was measured using a PolarStar Omega luminometer (BMG LABTECH, Cary, NC, USA). Samples were assayed in triplicate. Statistical significance was determined by unpaired t-test.
Polysome profiling and RNA analysis
Sucrose gradient centrifugation and polysome profiling of neuroblastoma lysates were performed as described.36 In some experiments, protein was precipitated from sucrose gradient fractions and analyzed by western blot as we have described previously, and RNA was extracted from each sucrose gradient fraction for RT–PCR or real-time PCR. Exogenous luciferase mRNA was added to the lysates prior to centrifugation as a reference to normalize for centrifugation yield. The experiment was repeated three times and yielded similar results each time.
Total RNA was extracted from cell lines using Trizol or from fractions of sucrose gradient centrifugation using Trizol-LS (Life Technologies, Grand Island, NY, USA). In cDNA was then prepared using 1 ug of total RNA from each sample (SuperScriptRIII First-Strand Synthesis SuperMix, Invitrogen, Grand Island, NY, USA). In some cases, a collection of cDNAs derived from total RNA isolated from a range of human cancers of different histologic grades was purchased from Origene. Real-time PCR primers were purchased from Applied Biosystems (Carlsbad, CA, USA). Real-time PCR was performed as we have described previously.30 β-actin was used as the reference. The samples were analyzed in triplicate using an ABI 7300 Real-time PCR machine (Life Technologies), and statistical analysis was performed using the unpaired t-test. RNA-Seq analysis of mRNA obtained from control or UTP18-overexpressing NIH-3T3 cells was performed using an Illumina Hi-Seq sequencer (Illumina, San Diego, CA, USA) and analyzed using standard methods.
Lactate and ATP assays
L(+)-lactate production by control or UTP18-overexpressing IMR5 neuroblastoma cells was determined using a commercially available colorimetric assay according to the manufacturer’s protocol (Biovision, Milpitas, CA, USA). Soluble ATP production was measured using a commercially available colorimetric assay kit according to the manufacturer’s protocol (Biovision). There were six replicates for each condition, and each experiment was repeated three times. Statistical significance was determined using the unpaired t-test.
In vitro proliferation, growth and apoptosis assays
Cell growth or cell survival was assayed in vitro using the MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay according to the manufacturer’s protocol (Roche Applied Sciences, Indianapolis, IN, USA) as we have described previously.30 Cell proliferation was measured using bromodeoxy-uridine incorporation into DNA as we have described previously.30 For apoptosis assays, human neuroblastoma cells overexpressing UTP18 or an appropriate control vector were dissociated and filtered to yield single cells, and flow cytometry was then performed to measure the percentage of apoptotic cells using a FACScan Flow Cytometer (BD Biosciences, Franklin Lake, NJ, USA). At least 3000 cells were counted for each condition. Statistical significance was determined using the proportion test.
In vivo tumor growth assays
All animal studies were conducted under the auspices of an IACUC protocol approved by the Harvard Medical Area Standing Committee on Animals. Adult male athymic CD1 nude mice (Charles River, Cambridge, MA, USA) were used for these experiments.
For tumor growth assays, mouse NIH-3T3 fibroblasts or human NLF neuroblastoma cells were transfected with control or UTP18 vectors and stable cell lines were selected in puromycin. For UTP18 knockdown experiments, human SH-SY5Y neuroblastoma cells were transduced with control shRNA or UTP18 shRNA lentiviruses to generate stable cell lines. Approximately 1 × 107 cells overexpressing the desired sequences were then implanted subcutaneously into nude mice (n=6 animals/group), and tumor growth was monitored as described previously.30 Tumor volume was calculated using the formula for a spheroid. Significance was determined using the unpaired t-test.
References
Belin S, Beghin A, Solano-Gonzalez E, Bezin L, Brunet-Manquat S, Textoris J et al. Dysregulation of ribosome biogenesis and translational capacity is associated with tumor progression of human breast cancer cells. PLoS One 2009; 4: e7147.
Narla A, Ebert BL . Ribosomopathies: human disorders of ribosome dysfunction. Blood 2010; 115: 3196–3205.
Silvera D, Formenti SC, Schneider RJ . Translational control in cancer. Nat Rev Cancer 2010; 10: 254–266.
Bernstein KA, Gallagher JE, Mitchell BM, Granneman S, Baserga SJ . The small-subunit processome is a ribosome assembly intermediate. Eukaryot Cell 2004; 3: 1619–1626.
Fichelson P, Moch C, Ivanovitch K, Martin C, Sidor CM, Lepesant JA et al. Live-imaging of single stem cells within their niche reveals that a U3snoRNP component segregates asymmetrically and is required for self-renewal in Drosophila. Nat Cell Biol 2009; 11: 685–693.
De Preter K, Vandesompele J, Heimann P, Yigit N, Beckman S, Schramm A et al. Human fetal neuroblast and neuroblastoma transcriptome analysis confirms neuroblast origin and highlights neuroblastoma candidate genes. Genome Biol 2006; 7: R84.
Reardon DA, Michalkiewicz E, Boyett JM, Sublett JE, Entrekin RE, Ragsdale ST et al. Extensive genomic abnormalities in childhood medulloblastoma by comparative genomic hybridization. Cancer Res 1997; 57: 4042–4047.
Schleiermacher G, Michon J, Huon I, d'Enghien CD, Klijanienko J, Brisse H et al. Chromosomal CGH identifies patients with a higher risk of relapse in neuroblastoma without MYCN amplification. Br J Cancer 2007; 97: 238–246.
Holzel M, Orban M, Hochstatter J, Rohrmoser M, Harasim T, Malamoussi A et al. Defects in 18 S or 28 S rRNA processing activate the p53 pathway. J Biol Chem 2010; 285: 6364–6370.
Moerke NJ, Aktas H, Chen H, Cantel S, Reibarkh MY, Fahmy A et al. Small-molecule inhibition of the interaction between the translation initiation factors eIF4E and eIF4G. Cell 2007; 128: 257–267.
Bert AG, Grepin R, Vadas MA, Goodall GJ . Assessing IRES activity in the HIF-1alpha and other cellular 5′ UTRs. RNA 2006; 12: 1074–1083.
Willimott S, Wagner SD . Post-transcriptional and post-translational regulation of Bcl2. Biochem Soc Trans 2010; 38: 1571–1575.
Spriggs KA, Stoneley M, Bushell M, Willis AE . Re-programming of translation following cell stress allows IRES-mediated translation to predominate. Biol Cell 2008; 100: 27–38.
DeBerardinis RJ, Mancuso A, Daikhin E, Nissim I, Yudkoff M, Wehrli S et al. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc Natl Acad Sci U S A 2007; 104: 19345–19350.
Jackson SE . Hsp90: structure and function. Top Curr Chem 2013; 328: 155–240.
Shi Y, Sharma A, Wu H, Lichtenstein A, Gera J . Cyclin D1 and c-myc internal ribosome entry site (IRES)-dependent translation is regulated by AKT activity and enhanced by rapamycin through a p38 MAPK- and ERK-dependent pathway. J Biol Chem 2005; 280: 10964–10973.
Chen QR, Bilke S, Wei JS, Whiteford CC, Cenacchi N, Krasnoselsky AL et al. cDNA array-CGH profiling identifies genomic alterations specific to stage and MYCN-amplification in neuroblastoma. BMC Genomics 2004; 5: 70.
Molenaar JJ, Koster J, Zwijnenburg DA, van Sluis P, Valentijn LJ, van der Ploeg I et al. Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes. Nature 2012; 483: 589–593.
R2: Microarray analysis and visualization platform (http://r2.amc.nl).
Goda N, Kanai M . Hypoxia-inducible factors and their roles in energy metabolism. Int J Hematol 2012; 95: 457–463.
van Riggelen J, Yetil A, Felsher DW . MYC as a regulator of ribosome biogenesis and protein synthesis. Nat Rev Cancer 2010; 10: 301–309.
Pisa V, Cozzolino M, Gargiulo S, Ottone C, Piccioni F, Monti M et al. The molecular chaperone Hsp90 is a component of the cap-binding complex and interacts with the translational repressor Cup during Drosophila oogenesis. Gene 2009; 432: 67–74.
Suzuki Y, Minami M, Suzuki M, Abe K, Zenno S, Tsujimoto M et al. The Hsp90 inhibitor geldanamycin abrogates colocalization of eIF4E and eIF4E-transporter into stress granules and association of eIF4E with eIF4G. J Biol Chem 2009; 284: 35597–35604.
Ujino S, Nishitsuji H, Sugiyama R, Suzuki H, Hishiki T, Sugiyama K et al. The interaction between human initiation factor eIF3 subunit c and heat-shock protein 90: a necessary factor for translation mediated by the hepatitis C virus internal ribosome entry site. Virus Res 2012; 163: 390–395.
Pan E, Pellarin M, Holmes E, Smirnov I, Misra A, Eberhart CG et al. Isochromosome 17q is a negative prognostic factor in poor-risk childhood medulloblastoma patients. Clin Cancer Res 2005; 11: 4733–4740.
Calabrese C, Frank A, Maclean K, Gilbertson R . Medulloblastoma sensitivity to 17-allylamino-17-demethoxygeldanamycin requires MEK/ERKM. J Biol Chem 2003; 278: 24951–24959.
Kang J, Kamal A, Burrows FJ, Evers BM, Chung DH . Inhibition of neuroblastoma xenograft growth by Hsp90 inhibitors. Anticancer Res 2006; 26: 1903–1908.
Sasaki T, Okuda K, Zheng W, Butrynski J, Capelletti M, Wang L et al. The neuroblastoma-associated F1174L ALK mutation causes resistance to an ALK kinase inhibitor in ALK-translocated cancers. Cancer Res 2010; 70: 10038–10043.
Goutagny S, Yang HW, Zucman-Rossi J, Chan J, Dreyfuss JM, Park PJ et al. Genomic profiling reveals alternative genetic pathways of meningioma malignant progression dependent on the underlying NF2 status. Clin Cancer Res 2010; 16: 4155–4164.
Kim H, Huang W, Jiang X, Pennicooke B, Park PJ, Johnson MD . Integrative genome analysis reveals an oncomir/oncogene cluster regulating glioblastoma survivorship. Proc Natl Acad Sci U S A 2010; 107: 2183–2188.
Kim TM, Xi R, Luquette LJ, Park RW, Johnson MD, Park PJ . Functional genomic analysis of chromosomal aberrations in a compendium of 8000 cancer genomes. Genome Res 2012; 23: 217–227.
Kim TM, Huang W, Park R, Park PJ, Johnson MD . A developmental taxonomy of glioblastoma defined and maintained by MicroRNAs. Cancer Res 2011; 71: 3387–3399.
Lang KJ, Kappel A, Goodall GJ . Hypoxia-inducible factor-1alpha mRNA contains an internal ribosome entry site that allows efficient translation during normoxia and hypoxia. Mol Biol Cell 2002; 13: 1792–1801.
Stein I, Itin A, Einat P, Skaliter R, Grossman Z, Keshet E . Translation of vascular endothelial growth factor mRNA by internal ribosome entry: implications for translation under hypoxia. Mol Cell Biol 1998; 18: 3112–3119.
Nanbru C, Lafon I, Audigier S, Gensac MC, Vagner S, Huez G et al. Alternative translation of the proto-oncogene c-myc by an internal ribosome entry site. J Biol Chem 1997; 272: 32061–32066.
del Prete MJ, Vernal R, Dolznig H, Mullner EW, Garcia-Sanz JA . Isolation of polysome-bound mRNA from solid tissues amenable for RT-PCR and profiling experiments. RNA 2007; 13: 414–421.
Acknowledgements
This work was supported by a Brain Science Foundation Research Award, the Santos Family Foundation, a Hagerty Fund Research Award, R01 NS062219 from the National Institute of Neurological Disorders and Stroke and an NIH Director’s New Innovator Award (DP2 OD002319) to MDJ.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Yang, H., Kim, TM., Song, S. et al. A small subunit processome protein promotes cancer by altering translation. Oncogene 34, 4471–4481 (2015). https://doi.org/10.1038/onc.2014.376
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2014.376
- Springer Nature Limited