Abstract
Although t-loops protect telomeres, they are at risk of cleavage by Holliday junction (HJ) resolvases if branch migration converts the three-way t-loop junction into four-way HJs. T-loop cleavage is repressed by the TRF2 basic domain, which binds three- and four-way junctions and protects HJs in vitro. By replacing the basic domain with bacterial-protein domains binding three- and four-way junctions, we demonstrated the in vivo relevance of branched-DNA binding. Branched-DNA binding also repressed PARP1, presumably by masking the PARP1 site in the t-loop junction. Although PARP1 recruits HJ resolvases and promotes t-loop cleavage, PARP1 activation alone did not result in t-loop cleavage, thus suggesting that the basic domain also prevents formation of HJs. Concordantly, removal of HJs by BLM helicase mitigated t-loop cleavage in response to loss of the basic domain. We propose that TRF2 masks and stabilizes the t-loop three-way junction, thereby protecting telomeres from detrimental deletions and PARP1 activation.






Similar content being viewed by others
References
de Lange, T. Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev. 19, 2100–2110 (2005).
Lazzerini-Denchi, E. & Sfeir, A. Stop pulling my strings: what telomeres taught us about the DNA damage response. Nat. Rev. Mol. Cell Biol. 17, 364–378 (2016).
Palm, W. & de Lange, T. How shelterin protects mammalian telomeres. Annu. Rev. Genet. 42, 301–334 (2008).
Celli, G.B. & de Lange, T. DNA processing is not required for ATM-mediated telomere damage response after TRF2 deletion. Nat. Cell Biol. 7, 712–718 (2005).
Karlseder, J., Broccoli, D., Dai, Y., Hardy, S. & de Lange, T. p53- and ATM-dependent apoptosis induced by telomeres lacking TRF2. Science 283, 1321–1325 (1999).
Smogorzewska, A., Karlseder, J., Holtgreve-Grez, H., Jauch, A. & de Lange, T. DNA ligase IV-dependent NHEJ of deprotected mammalian telomeres in G1 and G2. Curr. Biol. 12, 1635–1644 (2002).
van Steensel, B., Smogorzewska, A. & de Lange, T. TRF2 protects human telomeres from end-to-end fusions. Cell 92, 401–413 (1998).
Doksani, Y., Wu, J.Y., de Lange, T. & Zhuang, X. Super-resolution fluorescence imaging of telomeres reveals TRF2-dependent T-loop formation. Cell 155, 345–356 (2013).
Griffith, J.D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999).
Chasovskikh, S., Dimtchev, A., Smulson, M. & Dritschilo, A. DNA transitions induced by binding of PARP-1 to cruciform structures in supercoiled plasmids. Cytometry A 68, 21–27 (2005).
Pion, E. et al. Poly(ADP-ribose) polymerase-1 dimerizes at a 5′ recessed DNA end in vitro: a fluorescence study. Biochemistry 42, 12409–12417 (2003).
Potaman, V.N., Shlyakhtenko, L.S., Oussatcheva, E.A., Lyubchenko, Y.L. & Soldatenkov, V.A. Specific binding of poly(ADP-ribose) polymerase-1 to cruciform hairpins. J. Mol. Biol. 348, 609–615 (2005).
Saint-Léger, A. et al. The basic N-terminal domain of TRF2 limits recombination endonuclease action at human telomeres. Cell Cycle 13, 2469–2474 (2014).
Wang, R.C., Smogorzewska, A. & de Lange, T. Homologous recombination generates T-loop-sized deletions at human telomeres. Cell 119, 355–368 (2004).
Fouché, N. et al. The basic domain of TRF2 directs binding to DNA junctions irrespective of the presence of TTAGGG repeats. J. Biol. Chem. 281, 37486–37495 (2006).
Poulet, A. et al. TRF2 promotes, remodels and protects telomeric Holliday junctions. EMBO J. 28, 641–651 (2009).
Bianchi, A., Smith, S., Chong, L., Elias, P. & de Lange, T. TRF1 is a dimer and bends telomeric DNA. EMBO J. 16, 1785–1794 (1997).
Yoshimura, S.H., Maruyama, H., Ishikawa, F., Ohki, R. & Takeyasu, K. Molecular mechanisms of DNA end-loop formation by TRF2. Genes Cells 9, 205–218 (2004).
Wyatt, H.D. & West, S.C. Holliday junction resolvases. Cold Spring Harb. Perspect. Biol. 6, a023192 (2014).
Benson, F.E. & West, S.C. Substrate specificity of the Escherichia coli RuvC protein: resolution of three- and four-stranded recombination intermediates. J. Biol. Chem. 269, 5195–5201 (1994).
Hiom, K. & West, S.C. Branch migration during homologous recombination: assembly of a RuvAB-Holliday junction complex in vitro. Cell 80, 787–793 (1995).
Hiom, K., Tsaneva, I.R. & West, S.C. The directionality of RuvAB-mediated branch migration: in vitro studies with three-armed junctions. Genes Cells 1, 443–451 (1996).
Iwasaki, H., Takahagi, M., Nakata, A. & Shinagawa, H. Escherichia coli RuvA and RuvB proteins specifically interact with Holliday junctions and promote branch migration. Genes Dev. 6, 2214–2220 (1992).
Parsons, C.A. & West, S.C. Formation of a RuvAB-Holliday junction complex in vitro. J. Mol. Biol. 232, 397–405 (1993).
Parsons, C.A., Stasiak, A., Bennett, R.J. & West, S.C. Structure of a multisubunit complex that promotes DNA branch migration. Nature 374, 375–378 (1995).
Lloyd, R.G. & Sharples, G.J. Processing of recombination intermediates by the RecG and RuvAB proteins of Escherichia coli. Nucleic Acids Res. 21, 1719–1725 (1993).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Martínez, P. et al. Increased telomere fragility and fusions resulting from TRF1 deficiency lead to degenerative pathologies and increased cancer in mice. Genes Dev. 23, 2060–2075 (2009).
Saito, A., Iwasaki, H., Ariyoshi, M., Morikawa, K. & Shinagawa, H. Identification of four acidic amino acids that constitute the catalytic center of the RuvC Holliday junction resolvase. Proc. Natl. Acad. Sci. USA 92, 7470–7474 (1995).
Shah, R., Bennett, R.J. & West, S.C. Genetic recombination in E. coli: RuvC protein cleaves Holliday junctions at resolution hotspots in vitro. Cell 79, 853–864 (1994).
Rai, R., Chen, Y., Lei, M. & Chang, S. TRF2-RAP1 is required to protect telomeres from engaging in homologous recombination-mediated deletions and fusions. Nat. Commun. 7, 10881 (2016).
Mateos-Gomez, P.A. et al. Mammalian polymerase θ promotes alternative NHEJ and suppresses recombination. Nature 518, 254–257 (2015).
Sfeir, A. & de Lange, T. Removal of shelterin reveals the telomere end-protection problem. Science 336, 593–597 (2012).
Wang, M. et al. PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res. 34, 6170–6182 (2006).
Bizard, A.H. & Hickson, I.D. The dissolution of double Holliday junctions. Cold Spring Harb. Perspect. Biol. 6, a016477 (2014).
Vannier, J.B., Pavicic-Kaltenbrunner, V., Petalcorin, M.I., Ding, H. & Boulton, S.J. RTEL1 dismantles T-loops and counteracts telomeric G4-DNA to maintain telomere integrity. Cell 149, 795–806 (2012).
Sfeir, A., Kabir, S., van Overbeek, M., Celli, G.B. & de Lange, T. Loss of Rap1 induces telomere recombination in the absence of NHEJ or a DNA damage signal. Science 327, 1657–1661 (2010).
Konishi, A., Izumi, T. & Shimizu, S. TRF2 protein interacts with core histones to stabilize chromosome ends. J. Biol. Chem. 291, 20798–20810 (2016).
Grigoriev, M. & Hsieh, P. A histone octamer blocks branch migration of a Holliday junction. Mol. Cell. Biol. 17, 7139–7150 (1997).
González-Prieto, R., Cuijpers, S.A., Luijsterburg, M.S., van Attikum, H. & Vertegaal, A.C. SUMOylation and PARylation cooperate to recruit and stabilize SLX4 at DNA damage sites. EMBO Rep. 16, 512–519 (2015).
Doksani, Y. & de Lange, T. The role of double-strand break repair pathways at functional and dysfunctional telomeres. Cold Spring Harb. Perspect. Biol. 6, a016576 (2014).
Zimmermann, M., Kibe, T., Kabir, S. & de Lange, T. TRF1 negotiates TTAGGG repeat-associated replication problems by recruiting the BLM helicase and the TPP1/POT1 repressor of ATR signaling. Genes Dev. 28, 2477–2491 (2014).
Pickett, H.A., Cesare, A.J., Johnston, R.L., Neumann, A.A. & Reddel, R.R. Control of telomere length by a trimming mechanism that involves generation of t-circles. EMBO J. 28, 799–809 (2009).
Pickett, H.A., Henson, J.D., Au, A.Y., Neumann, A.A. & Reddel, R.R. Normal mammalian cells negatively regulate telomere length by telomere trimming. Hum. Mol. Genet. 20, 4684–4692 (2011).
Rivera, T., Haggblom, C., Cosconati, S. & Karlseder, J. A balance between elongation and trimming regulates telomere stability in stem cells. Nat. Struct. Mol. Biol. 24, 30–39 (2017).
Li, B. & Lustig, A.J. A novel mechanism for telomere size control in Saccharomyces cerevisiae. Genes Dev. 10, 1310–1326 (1996).
Li, J.S. et al. TZAP: a telomere-associated protein involved in telomere length control. Science 355, 638–641 (2017).
Karlseder, J., Smogorzewska, A. & de Lange, T. Senescence induced by altered telomere state, not telomere loss. Science 295, 2446–2449 (2002).
Broccoli, D., Smogorzewska, A., Chong, L. & de Lange, T. Human telomeres contain two distinct Myb-related proteins, TRF1 and TRF2. Nat. Genet. 17, 231–235 (1997).
Dunderdale, H.J., Sharples, G.J., Lloyd, R.G. & West, S.C. Cloning, overexpression, purification, and characterization of the Escherichia coli RuvC Holliday junction resolvase. J. Biol. Chem. 269, 5187–5194 (1994).
Frank, K.M. et al. Late embryonic lethality and impaired V(D)J recombination in mice lacking DNA ligase IV. Nature 396, 173–177 (1998).
Gu, Y. et al. Growth retardation and leaky SCID phenotype of Ku70-deficient mice. Immunity 7, 653–665 (1997).
Hockemeyer, D., Daniels, J.P., Takai, H. & de Lange, T. Recent expansion of the telomeric complex in rodents: two distinct POT1 proteins protect mouse telomeres. Cell 126, 63–77 (2006).
Jacks, T. et al. Tumor spectrum analysis in p53-mutant mice. Curr. Biol. 4, 1–7 (1994).
Kibe, T., Osawa, G.A., Keegan, C.E. & de Lange, T. Telomere protection by TPP1 is mediated by POT1a and POT1b. Mol. Cell. Biol. 30, 1059–1066 (2010).
Nussenzweig, A. et al. Requirement for Ku80 in growth and immunoglobulin V(D)J recombination. Nature 382, 551–555 (1996).
Takai, K.K., Kibe, T., Donigian, J.R., Frescas, D. & de Lange, T. Telomere protection by TPP1/POT1 requires tethering to TIN2. Mol. Cell 44, 647–659 (2011).
Ventura, A. et al. Restoration of p53 function leads to tumour regression in vivo. Nature 445, 661–665 (2007).
Takai, K.K., Hooper, S., Blackwood, S., Gandhi, R. & de Lange, T. In vivo stoichiometry of shelterin components. J. Biol. Chem. 285, 1457–1467 (2010).
Takai, H., Smogorzewska, A. & de Lange, T. DNA damage foci at dysfunctional telomeres. Curr. Biol. 13, 1549–1556 (2003).
Celli, G.B., Denchi, E.L. & de Lange, T. Ku70 stimulates fusion of dysfunctional telomeres yet protects chromosome ends from homologous recombination. Nat. Cell Biol. 8, 885–890 (2006).
Poon, S.S. & Lansdorp, P.M. Quantitative fluorescence in situ hybridization (Q-FISH). Curr. Protoc. Cell Biol. 12, 18.4 (2001).
Loayza, D. & De Lange, T. POT1 as a terminal transducer of TRF1 telomere length control. Nature 423, 1013–1018 (2003).
de Lange, T. et al. Structure and variability of human chromosome ends. Mol. Cell. Biol. 10, 518–527 (1990).
Kabir, S., Hockemeyer, D. & de Lange, T. TALEN gene knockouts reveal no requirement for the conserved human shelterin protein Rap1 in telomere protection and length regulation. Cell Rep. 9, 1273–1280 (2014).
de Lange, T. Human telomeres are attached to the nuclear matrix. EMBO J. 11, 717–724 (1992).
Bayani, J. & Squire, J.A. Sister chromatid exchange. Curr. Protoc. Cell Biol. 25, 22.7 (2005).
Acknowledgements
We thank D. White for expert mouse husbandry and the members of the laboratory of T.d.L. for sharing cell lines, for discussion and for comments on the manuscript. I.S. was supported by funds from a Women & Science Irene Diamond Fellowship in Breast Cancer Research at the Rockefeller University and by Mobility fellowships from the Swiss National Science foundation (Early Mobility fellowships PBFRP3-137230 and PBFRP3_142824, and Advanced Mobility fellowship P300PA_161006). L.T. was supported by an Anderson Center for Cancer Research Graduate Fellowship (contract FA9550-11-C-0028, awarded by the Department of Defense, Air Force Office of Scientific Research, National Defense Science and Engineering Graduate (NDSEG) Fellowship 32 CFR 168a). This work was supported by grants from the NIH (AG016642) to T.d.L. and (GM104962) to D.J.P., and by a Memorial Sloan-Kettering Cancer Center NIH Core Grant (P30 CA008748). T.d.L. is supported as an American Cancer Society Rose Zarucki Trust Research Professor. SPR experiments were performed at the RU High-Throughput and Spectroscopy Resource Center with expert assistance from A. Alcaino. OMX-SI imaging experiments were supported by a grant (S10RR031855) from the National Center For Research Resources to the Bio-Imaging Center at RU. The content of this manuscript is solely the responsibility of the authors and does not necessarily represent the official views of the National Center For Research Resources or the National Institutes of Health. We thank G. Hannon (Cancer Research UK Cambridge Institute) for reagents.
Author information
Authors and Affiliations
Contributions
I.S. and T.d.L. designed the experiments and wrote the manuscript. I.S. performed all in vivo experiments, except for the t-loop assays, which were performed by L.T. W.X. and D.J.P. performed the in vitro binding studies.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Functional analysis of TRF2 alleles.
(a) Schematic of RuvC-TRF2ΔB fusion proteins and the TRF2H31A construct. RuvCm contains mutations in the active site (D138N/D140N) and RuvCwt represents the wild type protein. The mutations in the active site abolish resolvase activity of RuvC but do not affect its binding to branched DNA structures29. (b) Immunoblot for TRF2 and γ-tubulin in NIH3T3 cells. (c) IF-FISH for TRF2 (red) and telomeres (green) in NIH3T3 cells expressing the indicated alleles. Blue, DAPI; Scale bar, 5 μm. (d) Telomeric ChIP analysis with TRF2 Ab in NIH3T3 cells expressing the different TRF2 alleles. (e) Quantification of telomeric DNA recovered with the TRF2 Ab (average % telomeric DNA recovered in two independent experiments). (f) IF-FISH for TRF1 (red) and telomeres (green) in NIH3T3 cells expressing the indicated alleles. Blue, DAPI; Scale bar, 5 μm. (g) Analysis of fragile telomeres, a proxy for TRF1 dysfunction, in the indicated NIH3T3 cells. Metaphase chromosomes were prepared five days after introduction of the TRF2 alleles. Data points represent individual metaphases from 3 independent experiments; bars show the median. Significance was calculated by One-way ANOVA with Tukey post-test, ns, not significant. (h) Immunoblot showing expression of TRF2 in SV40-LT TRF2F/F MEFs complemented with TRF2 alleles 96 h after Cre. Cre minus cells were harvested at the same time-point. (i) 53BP1 TIF analysis in cells as in (h); means±SDs from 3 independent experiments. (j) Quantification of telomere fusions in cells as in (h); median fusion frequency in one experiment (13-15 metaphases).
Supplementary Figure 2 RuvA and RuvC protect telomeres in place of the basic domain.
(a) Top: Illustration of the approach for measuring unequal sister telomeres. Middle: Representative image showing metaphase chromosomes of TRF2ΔB expressing NIH3T3 cells one day after introduction by retroviral infection. Scale bar, 10 μm. Green: Telomeric FISH; red, DAPI. Arrows: unequal sister telomere signals. Bottom: Enlarged examples of chromosomes with unequal sister telomeres. Right: a scatter plot of the raw sister telomere ratios from one representative experiment is shown. This experiment is part of the analysis of unequal sister telomeres shown in Fig. 2e. Data points represent individual sister ratios (from 330 chromosome ends on 10 metaphase spreads). The dotted line depicts the cut-off used in this experiment to calculate the % of ends with unequal sister telomeres. The cut-off represents the 80th percentile sister ratio in control cells (vec). The scatter plot shows more chromosome ends with a ratio above the threshold in cells expressing TRF2ΔB. The % of ratios above the threshold is indicated. All unequal sister ratios were determined as indicated in (a). (b) Scatter plot of raw telomere fluorescence units (TFU in arbitrary units) from Q-FISH analysis in NIH3T3 cells expressing the indicated TRF2 alleles as in Fig. 2f. Median TFU units from 3 independent experiments (medians±SEMs) are indicated (white bars). (c) Telomere blot of HindIII-digested DNA from NIH3T3 cells (day 5 post-infection). The membrane was hybridized with a chromosome-internal probe for normalization, stripped, and hybridized to the telomere-specific Sty11 probe to detect telomeric repeats. (d) Quantification of telomeric signal intensity, as determined in (c). Telomeric signals were normalized to the signal from the chromosome-internal probe and are shown relative to wt TRF2 (100%). Means±SDs from 6 independent experiments; p-values from One-way Anova with Dunnett’s test. (e) Quantification of chromosome ends with unequal sister telomeres in NIH3T3 cells. Scatter plots with medians are shown (≥25 metaphases from four independent experiments; p values from One-way ANOVA with Tukey post-test). The same data sets for vec, WT, and ΔB are shown Fig. 2l. (f) Q-FISH analysis of telomeric DNA signals in NIH3T3 cells expressing the indicated alleles (% telomeric DNA relative to empty vector; means±SDs from four independent experiments. The same data sets for vec, WT, and ΔB are shown in Fig. 2m. (g) Scatter plot of sister telomere ratios from one representative experiment as in Fig. 2l. Data points represent individual sister ratios. This experiment is part of the analysis of unequal sister telomeres shown in Fig. 2l. (h) Scatter plot of raw TFUs from Q-FISH analysis in NIH3T3 cells expressing the indicated TRF2 alleles as in Fig. 2m. Median TFU units from 5 independent experiments (medians±SEMs) are indicated (white bars).
Supplementary Figure 3 Control experiments on HeLa cells.
(a) IF-FISH to detect TRF2 (red) and telomeres (green) in HeLa1.2.11 cells (Day 8 post-infection). Blue, DAPI; Scale bar, 5 μm. (b) IF-FISH to detect TRF1 (red) and telomeres (green) in HeLa1.2.11 cells as in (a). TRF1 localization to telomeres is not affected by overexpression of TRF2 alleles. Blue, DAPI; Scale bar, 5 μm. (c) Quantification of chromosome ends with fragile telomeres, a proxy for TRF1 function in HeLa1.2.11 cells as in (a). The analysis shows that similar to mouse telomeres, expression of TRF2 alleles does not induce fragile telomeres. Data points represent individual metaphases from 3 independent experiments and significance is based on One-way ANOVA with Tukey post-test as in Fig. 1. Bars represent the median.
Supplementary Figure 4 TRF2ΔB-induced telomere loss depends on PARP1.
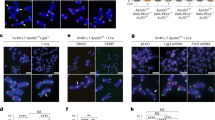
(a) Detection of PARP1 and telomeres in TRF1F/FTRF2F/FKu80-/-p53-/- MEFs with and without Cre (96 h). Red, IF for PARP1; green, telomeric FISH; blue, DAPI. Scale bar, 5 μm. (b) Quantification of PARP1 signals co-localizing with telomeres as assessed by IF-FISH in TRF1F/FTRF2F/FKu80-/-p53-/- MEFs with and without Cre treatment as in (a) (means±SDs from 3 independent experiments; Colocalization was assessed in 40-50 nuclei per experiment and experimental condition; p values from One-way ANOVA with Tukey post test). (c) Immunoblot showing TRF2 and TRF1 loss, and induction of PARsylation upon Cre treatment of TRF1F/FTRF2F/FKu80-/-p53-/- cells. Treatment with 5 μM Olaparib (PARPi) verifies PARP1 dependent PARsylation. (d) IF for PAR (green) in TRF1F/FTRF2F/FKu80-/-p53-/- cells. Dysfunctional telomeres are visualized by 53BP1 IF (red). Blue, DAPI; Scale bar, 5 μm. (e) Immunoblot showing TRF2 in control and TRF2ΔB-expressing NIH3T3 cells that were mock treated (DMSO) or treated with 2 μM Olaparib (PARPi). (f) Immunoblot showing TRF2ΔB expression in PARP1-proficient and -deficient MEFs infected with empty vector or TRF2ΔB. Control cell lines were PARP1wt/- (ctrl1) and PARP1wt/wt (ctrl2). (g) Scatter plot of telomere fluorescence units (TFU) from Q-FISH analysis in PARP1-proficient and -deficient MEFs infected with vec or TRF2ΔB as in (f). TFU units from 2 independent experiments (medians±SEMs) are shown (white bars).
Supplementary Figure 5 TRF2 and TIN2 independently repress telomeric PARP1 signaling.
(a) Quantification of cells with ≥5 53BP1 TIFs in the indicated conditional KO MEFs (Cre minus and plus; means±SDs from ≥3 independent experiments). 53BP1 TIF analysis serves as a control for efficient deletion of shelterin components in the survey of MEF cell lines for PARP1 activation. Note that deletion of Rap1 does not induce TIFs in Ku-proficient cells and only to a minor level in Ku-deficient cells. Efficient deletion was also verified by immunoblot analysis using specific antibodies for the individual shelterin subunits. (b) Quantification of cells with ≥5 53BP1 TIFs in cells as in Fig. 4b (Cre minus and plus; means±SDs from 3 independent experiments). 53BP1 TIF analysis was used to control for efficient deletion of shelterin components. Efficient deletion was also verified by immunoblot analysis using specific antibodies for the individual shelterin subunits (c) Immunoblot showing PAR and TRF2 in TIN2F/FKu70-/- MEFs. Note that PARsylation was rescued by expression of TIN2, but not expression of TRF2. (d) Immunoblot showing TRF2 overexpression in TIN2F/FKu70-/- MEFs. (e) Telomeric ChIP for TRF2 and TIN2 in TIN2F/FKu70-/- MEFs. The quantification shows that TRF2 overexpression re-establishes the telomeric TRF2 levels in TIN2/Ku70 DKO cells. (f) IF-FISH for PARP1 (red) and telomeres (green) in cells as in (d); DAPI (blue); Scale bar, 5 μm. (g) Quantification of PARP1 co-localization with telomeres as in (f) (means±SDs from 3 independent experiments). All experiments were performed at 96 h after Cre. For all experiments, P values are as in Fig. 1 based on One-way ANOVA with Tukey post-test.
Supplementary Figure 6 Rap1 deletion does not exacerbate phenotypes associated with telomeres containing TRF2ΔB.
(a) Immunoblot showing TRF2 alleles and Rap1 in Rap1F/F Ku70+/- p53-/- MEFs with and without Cre treatment. Cells were infected with TRF2 alleles and selected for integration of the TRF2 expression plasmid for 4-5 days. Then the cells were split, H&R Cre infected and harvested 120h post Cre treatment. This experimental setting allows the analysis of the effect of Rap1 deletion on TRF2ΔB-induced phenotypes in an isogenic background. (b) Quantification of PARP1 signals co-localizing with telomeres as assessed by IF-FISH in Rap1F/F Ku70+/- p53-/- MEFs as in (a) (means±SDs from 3 independent experiments, P values are based on One-way ANOVA with Tukey post-test). (c) Quantification of chromatids that do not have detectable telomere FISH signal in Rap1F/F Ku70+/- p53-/- MEFs as in (a). Scatter plots with medians are shown (≥40 metaphases from three independent experiments; p values from One-way ANOVA with Tukey post-test.) (d) Telomere blot analysis of MboI/AluI-digested DNA from Rap1F/F Ku70+/- p53-/- MEFs expressing the indicated TRF2 alleles with or without Cre treatment. Cells were harvested 120h (left panels) or 240h (right panels) post Cre treatment. The lower panels show the signal detected with the BamHI probe, which serves as loading control. The relative telomere abundance calculated from the telomere blot and normalized to the BamHI signal is indicated. The signal in Cre minus control cells was set to 100% for each time point. This panel shows a representative blot. The experiment was repeated 3 times. (e) Quantification of chromosome fusions that do not have detectable telomere signals at the fusion sites in Rap1F/F Ku70+/- p53-/- MEFs as in (a) (means±SDs from 3 independent experiments, ≥40 metaphase spreads, p-values from unpaired t-test). P value symbols are as in Fig.1.
Supplementary Figure 7 Generation of BLM-KO cells by CRISPR–Cas9 gene editing.
(a) Schematic of the BLM locus showing landmarks relevant to CRISPR/Cas9 gene editing. The guide RNA (gRNA) region and the PAM are indicated in the reference sequence and the changes in sequence are highlighted in the edited alleles of the two CRISPR clones. In clone BLM KO-2 only one allele variant (and no wt allele) was detected. The sequences were derived from TOPO-cloned PCR products amplified from the indicated primer pair. (b) Immunoblot showing BLM (arrow) in parental NIH3T3 cells but not in the KO clones. (c) SCE assay in parental NIH3T3 cells and the two BLM KO clones confirming BLM deficiency. (d) Quantification of chromosome fusions that do not have detectable telomere signals at the fusion sites in parental and BLM KO NIH3T3 cells on day 1 after infection. Dots represent the % of fused chromosomes in individual metaphase spreads and the mean is shown (40 metaphase spreads from 4 independent experiments. P values are as in Fig. 1 based on One-way ANOVA with Tukey post-test.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1. (PDF 2069 kb)
Rights and permissions
About this article
Cite this article
Schmutz, I., Timashev, L., Xie, W. et al. TRF2 binds branched DNA to safeguard telomere integrity. Nat Struct Mol Biol 24, 734–742 (2017). https://doi.org/10.1038/nsmb.3451
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3451
- Springer Nature America, Inc.
This article is cited by
-
The risky business of ADP-ribosylating telomeric DNA
Nature Structural & Molecular Biology (2024)
-
Deregulated DNA ADP-ribosylation impairs telomere replication
Nature Structural & Molecular Biology (2024)
-
Homology directed telomere clustering, ultrabright telomere formation and nuclear envelope rupture in cells lacking TRF2B and RAP1
Nature Communications (2023)
-
ZNF524 directly interacts with telomeric DNA and supports telomere integrity
Nature Communications (2023)
-
The impact of reproductive factors on DNA methylation-based telomere length in healthy breast tissue
npj Breast Cancer (2022)