Abstract
The photosystem II protein PsbS has an essential role in qE-type nonphotochemical quenching, which protects plants from photodamage under excess light conditions. qE is initiated by activation of PsbS by low pH, but the mechanism of PsbS action remains elusive. Here we report the low-pH crystal structures of PsbS from spinach in its free form and in complex with the qE inhibitor N,N′-dicyclohexylcarbodiimide (DCCD), revealing that PsbS adopts a unique folding pattern, and, unlike other members of the light-harvesting-complex superfamily, it is a noncanonical pigment-binding protein. Structural and biochemical evidence shows that both active and inactive PsbS form homodimers in the thylakoid membranes, and DCCD binding disrupts the lumenal intermolecular hydrogen bonds of the active PsbS dimer. Activation of PsbS by low pH during qE may involve a conformational change associated with altered lumenal intermolecular interactions of the PsbS dimer.







Similar content being viewed by others
References
Caffarri, S., Kouřil, R., Kereïche, S., Boekema, E.J. & Croce, R. Functional architecture of higher plant photosystem II supercomplexes. EMBO J. 28, 3052–3063 (2009).
Demmig-Adams, B. & Adams, W.W. III. Photoprotection and other responses of plants to high light stress. Annu. Rev. Plant Physiol. Plant Mol. Biol. 43, 599–626 (1992).
Apel, K. & Hirt, H. Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 55, 373–399 (2004).
Horton, P., Ruban, A.V. & Walters, R.G. Regulation of light harvesting in green plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 655–684 (1996).
Müller, P., Li, X.-P. & Niyogi, K.K. Non-photochemical quenching: a response to excess light energy. Plant Physiol. 125, 1558–1566 (2001).
de Bianchi, S., Ballottari, M., Dall'osto, L. & Bassi, R. Regulation of plant light harvesting by thermal dissipation of excess energy. Biochem. Soc. Trans. 38, 651–660 (2010).
Külheim, C., Ågren, J. & Jansson, S. Rapid regulation of light harvesting and plant fitness in the field. Science 297, 91–93 (2002).
Ruban, A.V., Johnson, M.P. & Duffy, C.D.P. The photoprotective molecular switch in the photosystem II antenna. Biochim. Biophys. Acta 1817, 167–181 (2012).
Rochaix, J.D. Regulation and dynamics of the light-harvesting system. Annu. Rev. Plant Biol. 65, 287–309 (2014).
Kramer, D.M., Sacksteder, C.A. & Cruz, J.A. How acidic is the lumen? Photosynth. Res. 60, 151–163 (1999).
Demmig-Adams, B. Carotenoids and photoprotection in plants: a role for the xanthophyll zeaxanthin. Biochim. Biophys. Acta 1020, 1–24 (1990).
Ruban, A.V. et al. Identification of a mechanism of photoprotective energy dissipation in higher plants. Nature 450, 575–578 (2007).
Ahn, T.K. et al. Architecture of a charge-transfer state regulating light harvesting in a plant antenna protein. Science 320, 794–797 (2008).
Bode, S. et al. On the regulation of photosynthesis by excitonic interactions between carotenoids and chlorophylls. Proc. Natl. Acad. Sci. USA 106, 12311–12316 (2009).
Li, X.-P. et al. A pigment-binding protein essential for regulation of photosynthetic light harvesting. Nature 403, 391–395 (2000).
Kasajima, I. et al. Molecular distinction in genetic regulation of nonphotochemical quenching in rice. Proc. Natl. Acad. Sci. USA 108, 13835–13840 (2011).
Brooks, M.D., Jansson, S. & Niyogi, K.K. in Non-Photochemical Quenching and Energy Dissipation in Plants, Algae and Cyanobacteria Vol. 40 (eds. Demmig-Adams, B., Garab, G., Adams, W.W. III & Govindjee, U.o.I.) 297–314 (Springer, 2014).
Li, X.-P., Gilmore, A.M. & Niyogi, K.K. Molecular and global time-resolved analysis of a psbS gene dosage effect on pH- and xanthophyll cycle-dependent nonphotochemical quenching in photosystem II. J. Biol. Chem. 277, 33590–33597 (2002).
Li, X.-P., Müller-Moulé, P., Gilmore, A.M. & Niyogi, K.K. PsbS-dependent enhancement of feedback de-excitation protects photosystem II from photoinhibition. Proc. Natl. Acad. Sci. USA 99, 15222–15227 (2002).
Barros, T. & Kühlbrandt, W. Crystallisation, structure and function of plant light-harvesting complex II. Biochim. Biophys. Acta 1787, 753–772 (2009).
Funk, C., Schroder, W.P., Green, B.R., Renger, G. & Andersson, B. The intrinsic 22 kDa protein is a chlorophyll-binding subunit of photosystem II. FEBS Lett. 342, 261–266 (1994).
Funk, C. et al. The PSII-S protein of higher plants: a new type of pigment-binding protein. Biochemistry 34, 11133–11141 (1995).
Aspinall-O'Dea, M. et al. In vitro reconstitution of the activated zeaxanthin state associated with energy dissipation in plants. Proc. Natl. Acad. Sci. USA 99, 16331–16335 (2002).
Niyogi, K.K., Li, X.-P., Rosenberg, V. & Jung, H.S. Is PsbS the site of non-photochemical quenching in photosynthesis? J. Exp. Bot. 56, 375–382 (2005).
Dominici, P. et al. Biochemical properties of the PsbS subunit of photosystem II either purified from chloroplast or recombinant. J. Biol. Chem. 277, 22750–22758 (2002).
Bonente, G., Howes, B.D., Caffarri, S., Smulevich, G. & Bassi, R. Interactions between the photosystem II subunit PsbS and xanthophylls studied in vivo and in vitro. J. Biol. Chem. 283, 8434–8445 (2008).
Li, X.-P. et al. Regulation of photosynthetic light harvesting involves intrathylakoid lumen pH sensing by the PsbS protein. J. Biol. Chem. 279, 22866–22874 (2004).
Bergantino, E. et al. Light- and pH-dependent structural changes in the PsbS subunit of photosystem II. Proc. Natl. Acad. Sci. USA 100, 15265–15270 (2003).
Johnson, M.P. & Ruban, A.V. Arabidopsis plants lacking PsbS protein possess photoprotective energy dissipation. Plant J. 61, 283–289 (2010).
Johnson, M.P. & Ruban, A.V. Restoration of rapidly reversible photoprotective energy dissipation in the absence of PsbS protein by enhanced ΔpH. J. Biol. Chem. 286, 19973–19981 (2011).
Horton, P. et al. PS2001 Proc. 12th International Congress on Photosynthesis PL-003 (CSIRO Publishing, Melbourne, Australia, 2001).
Li, X.-P., Phippard, A., Pasari, J. & Niyogi, K.K. Structure-function analysis of photosystem II subunit S (PsbS) in vivo. Funct. Plant Biol. 29, 1131–1139 (2002).
Schultes, N.P. & Peterson, R.B. Phylogeny-directed structural analysis of the Arabidopsis PsbS protein. Biochem. Biophys. Res. Commun. 355, 464–470 (2007).
Wedel, N., Klein, R., Ljungberg, U., Andersson, B. & Herrmann, R.G. The single-copy gene psbS codes for a phylogenetically intriguing 22 kDa polypeptide of photosystem II. FEBS Lett. 314, 61–66 (1992).
Kim, S. et al. Characterization of a spinach psbS cDNA encoding the 22 kDa protein of photosystem II. FEBS Lett. 314, 67–71 (1992).
Jansson, S. A guide to the Lhc genes and their relatives in Arabidopsis. Trends Plant Sci. 4, 236–240 (1999).
Liu, Z. et al. Crystal structure of spinach major light-harvesting complex at 2.72 Å resolution. Nature 428, 287–292 (2004).
Pan, X. et al. Structural insights into energy regulation of light-harvesting complex CP29 from spinach. Nat. Struct. Mol. Biol. 18, 309–315 (2011).
Amunts, A., Toporik, H., Borovikova, A. & Nelson, N. Structure determination and improved model of plant photosystem I. J. Biol. Chem. 285, 3478–3486 (2010).
Pan, X., Liu, Z.F., Li, M. & Chang, W.R. Architecture and function of plant light-harvesting complexes II. Curr. Opin. Struct. Biol. 23, 515–525 (2013).
Funk, C., Adamska, I., Green, B.R., Andersson, B. & Renger, G. The nuclear-encoded chlorophyll-binding photosystem II-S protein is stable in the absence of pigments. J. Biol. Chem. 270, 30141–30147 (1995).
Wilk, L., Grunwald, M., Liao, P.N., Walla, P.J. & Kühlbrandt, W. Direct interaction of the major light-harvesting complex II and PsbS in nonphotochemical quenching. Proc. Natl. Acad. Sci. USA 110, 5452–5456 (2013).
Plumley, F.G. & Schmidt, G.W. Reconstitution of chlorophyll a/b light-harvesting complexes: xanthophyll-dependent assembly and energy transfer. Proc. Natl. Acad. Sci. USA 84, 146–150 (1987).
Magalhaes, A., Maigret, B., Hoflack, J., Gomes, J.A.N.F. & Scheraga, H.A. Contribution of unusual arginine-arginine short-range interactions to stabilization and recognition in proteins. J. Protein Chem. 13, 195–215 (1994).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies form crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Goss, R., Opitz, C., Lepetit, B. & Wilhelm, C. The synthesis of NPQ-effective zeaxanthin depends on the presence of a transmembrane proton gradient and a slightly basic stromal side of the thylakoid membrane. Planta 228, 999–1009 (2008).
Zaks, J., Amarnath, K., Sylak-Glassman, E.J. & Fleming, G.R. Models and measurements of energy-dependent quenching. Photosynth. Res. 116, 389–409 (2013).
Azzi, A., Casey, R.P. & Nalecz, M.J. The effect of N,N′-dicyclohexylcarbodiimide on enzymes of bioenergetic relevance. Biochim. Biophys. Acta 768, 209–226 (1984).
Mizutani, K. et al. Structure of the rotor ring modified with N,N′-dicyclohexylcarbodiimide of the Na+-transporting vacuolar ATPase. Proc. Natl. Acad. Sci. USA 108, 13474–13479 (2011).
Pogoryelov, D. et al. Microscopic rotary mechanism of ion translocation in the F0 complex of ATP synthases. Nat. Chem. Biol. 6, 891–899 (2010).
Teardo, E. et al. Evidences for interaction of PsbS with photosynthetic complexes in maize thylakoids. Biochim. Biophys. Acta 1767, 703–711 (2007).
Johnson, M.P. et al. Photoprotective energy dissipation involves the reorganization of photosystem II light-harvesting complexes in the grana membranes of spinach chloroplasts. Plant Cell 23, 1468–1479 (2011).
Betterle, N. et al. Light-induced dissociation of an antenna hetero-oligomer is needed for non-photochemical quenching induction. J. Biol. Chem. 284, 15255–15266 (2009).
Kiss, A.Z., Ruban, A.V. & Horton, P. The PsbS protein controls the organization of the photosystem II antenna in higher plant thylakoid membranes. J. Biol. Chem. 283, 3972–3978 (2008).
Kereïche, S., Kiss, A.Z., Kouril, R., Boekema, E.J. & Horton, P. The PsbS protein controls the macro-organisation of photosystem II complexes in the grana membranes of higher plant chloroplasts. FEBS Lett. 584, 759–764 (2010).
Goral, T.K. et al. Light-harvesting antenna composition controls the macrostructure and dynamics of thylakoid membranes in Arabidopsis. Plant J. 69, 289–301 (2012).
Sylak-Glassman, E.J. et al. Distinct roles of the photosystem II protein PsbS and zeaxanthin in the regulation of light harvesting in plants revealed by fluorescence lifetime snapshots. Proc. Natl. Acad. Sci. USA 111, 17498–17503 (2014).
Berthold, D.A., Babcock, G.T. & Yocum, C.F. A highly resolved, oxygen-evolving photosystem II preparation from spinach thylakoid membranes. FEBS Lett. 134, 231–234 (1981).
Mishra, R.K. & Ghanotakis, D.F. Selective extraction of CP26 and CP29 proteins without affecting the binding of the extrinsic protein (33, 23 and 17 kDa) and the DCMU sensitivity of a photosystem II core complex. Photosynth. Res. 42, 37–42 (1994).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Sheldrick, G.M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Laskowski, R.A., Macarthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Joly, D. & Carpentier, R. in Photosynthesis Research Protocols, Methods in Molecular Biology Vol. 684 (ed. Carpentier, R.) 321–325 (Springer, 2011).
Farber, A., Young, A.J., Ruban, A.V., Horton, P. & Jahns, P. Dynamics of xanthophyll-cycle activity in different antenna subcomplexes in the photosynthetic membranes of higher plants: the relationship between zeaxanthin conversion and nonphotochemical fluorescence quenching. Plant Physiol. 115, 1609–1618 (1997).
Acknowledgements
We thank R. Bassi (Dipartimento di Biotecnologie, Università di Verona) for discussion, manuscript reading and providing seeds of npq4-E122Q E226Q double-mutant Arabidopsis, and N. Isaacs, K.K. Niyogi and J. Barber for manuscript reading. We are grateful to the staff at the Shanghai Synchrotron Radiation Facility and the Photo Factory for technical support. This work was supported by grants 2011CBA00902 (to W.C.) and 2011CBA00903 (to Z.L.) from the National Key Basic Research Program of China; grant XDB08020302 (to W.C.) from the Strategic Priority Research Program of the Chinese Academy of Sciences; and grants 31021062 (to W.C.), 31270793 (to M.L.), 31170703 (to X.P.), and 31100534 (to P.C.) from the National Natural Science Foundation of China.
Author information
Authors and Affiliations
Contributions
M.F., M.L. and W.C. conceived the project. M.F. purified PsbS and performed the structural determination and the biochemical experiments with PsbS. P.C., M.L. and H.Z. assisted with data collection. X.Z. and J.Z. assisted with isolation of BBY membranes. X.P. assisted with HPLC experiments. M.F., M.L., Z.L. and W.C. discussed the results and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Sequence alignment of PsbS from different plants.
The species used for alignment are Spinacia oleracea, Arabidopsis thaliana, Oryza sativa, Zea mays, Hordeum vulgare, Populus trichocarpa, Pinus sitchensis, Selaginella moellendorffii, and Physcomitrella patens. The secondary structure of PsbS is shown above the sequence. Fully conserved residues are shaded in red. The residues mediating dimerization are marked with circles (hydrogen bond interactions) and triangles (hydrophobic interactions). The two pH-sensing glutamates are marked with squares.
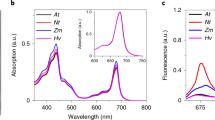
Supplementary Figure 2 The purified PsbS protein sample contains chlorophylls.
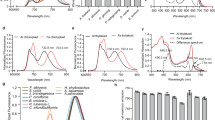
(a) Size exclusion chromatography result showed that the chlorophyll molecules co-elute with PsbS from a gel filtration column. PsbS and chlorophyll were monitored by the absorption at 280 nm (A280) and 663 nm (A663), respectively.
(b) HPLC analysis of the pigments in the purified PsbS protein sample. The identification of Chl a and Chl b was based on the absorption spectra of each peak fraction.
Supplementary Figure 3 The packing mode of PsbS crystal.
(a) The packing of PsbS molecules within one layer in the crystal is mediated by hydrophobic interactions between the transmembrane helices of PsbS.
(b) The packing of PsbS molecules between layers in the crystal is mediated by hydrophilic interactions between the lumenal part and the N-terminal part of PsbS. One of the contact sites is indicated by a red ellipse.
Supplementary Figure 4 Structural explanation of previous mutation studies on PsbS.
(a) In Arabidopsis, the two ethylmethane sulfonate (EMS)-induced mutations G84D and G150E (equivalent to Gly31 and Gly97 in spinach) were reported to significantly affect the stability of PsbS (Li, X.-P. et al., Funct. Plant Biol. 29, 1131–1139, 2002). In the structure, we find that both Gly31 and Gly97 are located in the region where extensive hydrogen bonds are formed and stabilize the stromal conformation of PsbS. Mutations of them to large and charged residues will disrupt these interactions.
(b,c) In Arabidopsis, the mutations of the two salt bridges connecting TM1 and TM3 were reported to significantly affect the function of PsbS in qE but not its expression without reasonable explanation (Schultes, N.P. & Peterson, R.B. Biochem. Biophys. Res. Comm. 355, 464–470, 2007). Our structural analysis have revealed that PsbS cannot bind chlorophyll molecules here; therefore, the most probable explanation is that these mutations (E to V and R to L) disrupt surrounding hydrogen bonds and affect local conformations. The more hydrogen bonds around the second salt bridge (Arg42–Glu141) are consistent with the bigger effect of its mutation. The above-mentioned residues are shown as yellow sticks, and the other involved residues are shown in white.
Supplementary Figure 5 The potential chlorophyll molecule bound to PsbS.
(a) A picture of green PsbS crystals.
(b) HPLC analysis of the pigments in the PsbS crystals.
(c) The Chl a molecule is bound at the lumenal dimerization interface of PsbS dimer. The 2Fo – Fc (0.8σ level) electron density of the Chl a molecule is shown. The two PsbS monomers are shown in limon and palecyan, respectively. The surrounding non-polar residues interacting with the Chl a molecule are shown in stick. For clarity, the phytyl chain of the Chl a molecule is not shown.
(d) Comparison of the absorption spectra of the purified PsbS protein and free Chl a.
(e) Comparison of the circular dichroism (CD) spectra of the purified PsbS and LHCII proteins.
Supplementary Figure 6 PsbS is dimeric in both its active and inactive states in Arabidopsis.
Crosslinking of PsbS using the thylakoids of wild-type (a) and pH-insensitive npq4-E122Q E226Q double mutant (b) of Arabidopsis at pH 5.0 with EDC. Under low pH or high light conditions, wild-type PsbS is in its active state, but pH-insensitive PsbS mutant is still in its inactive state.
Supplementary Figure 7 The DCCD-binding property of the purified PsbS protein.
(a) Mass spectrometric determination of the molecular weight of native PsbS. The subtilisin-treated PsbS (SU-PsbS) lacking N-terminal five residues (for crystallization) was used in mass spectrometric analysis. The experimental molecular weight (21923.36) of SU-PsbS is very close to the sequence-based molecular weight (21922.5) of SU-PsbS.
(b) Mass spectrometric analysis of DCCD-incubated SU-PsbS. Considering that DCCD (206.33) is covalently bound to SU-PsbS without molecular weight loss, the three peaks (22129.94, 22336.32, 22542.57) presented in the mass spectrometric result should correspond to SU-PsbS plus one DCCD, SU-PsbS plus two DCCD, and SU-PsbS plus three DCCD, respectively. The last peak is very small and may be because of the binding of DCCD to Glu69, Glu173 and another lumenal acidic residue of PsbS.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1 (PDF 1352 kb)
Supplementary Data Set 1
Original gel and immunoblot images (PDF 347 kb)
Rights and permissions
About this article
Cite this article
Fan, M., Li, M., Liu, Z. et al. Crystal structures of the PsbS protein essential for photoprotection in plants. Nat Struct Mol Biol 22, 729–735 (2015). https://doi.org/10.1038/nsmb.3068
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3068
- Springer Nature America, Inc.
This article is cited by
-
Proteomics Studies Indicate Differential Regulation of Key Metabolism, Energy, and Stress-Responsive Proteins in Azadirachta indica Callus Upon Exposure to Salinity and Jasmonic Acid Treatments
Journal of Plant Growth Regulation (2023)
-
Effects of high-temperature stress on photosynthesis of a new variety "SF-2" in Neoporphyra haitanensis
Journal of Applied Phycology (2022)
-
Macromolecular conformational changes in photosystem II: interaction between structure and function
Biophysical Reviews (2022)
-
The PsbS protein and low pH are necessary and sufficient to induce quenching in the light-harvesting complex of plants LHCII
Scientific Reports (2021)
-
The molecular pH-response mechanism of the plant light-stress sensor PsbS
Nature Communications (2021)