Abstract
The human APOBEC3G (A3G) DNA cytosine deaminase restricts and hypermutates DNA-based parasites including HIV-1. The viral infectivity factor (Vif) prevents restriction by triggering A3G degradation. Although the structure of the A3G catalytic domain is known, the structure of the N-terminal Vif-binding domain has proven more elusive. Here, we used evolution- and structure-guided mutagenesis to solubilize the Vif-binding domain of A3G, thus permitting structural determination by NMR spectroscopy. A smaller zinc-coordinating pocket and altered helical packing distinguish the structure from previous catalytic-domain structures and help to explain the reported inactivity of this domain. This soluble A3G N-terminal domain is bound by Vif; this enabled mutagenesis and biochemical experiments, which identified a unique Vif-interacting surface formed by the α1-β1, β2-α2 and β4-α4 loops. This structure sheds new light on the Vif-A3G interaction and provides critical information for future drug development.





Similar content being viewed by others
References
Malim, M.H. & Emerman, M. HIV-1 accessory proteins: ensuring viral survival in a hostile environment. Cell Host Microbe 3, 388–398 (2008).
Harris, R.S., Hultquist, J.F. & Evans, D.T. The restriction factors of human immunodeficiency virus. J. Biol. Chem. 287, 40875–40883 (2012).
Malim, M.H. & Bieniasz, P.D. HIV restriction factors and mechanisms of evasion. Cold Spring Harb. Perspect. Med. 2, a006940 (2012).
Refsland, E.W. et al. Quantitative profiling of the full APOBEC3 mRNA repertoire in lymphocytes and tissues: implications for HIV-1 restriction. Nucleic Acids Res. 38, 4274–4284 (2010).
Refsland, E.W., Hultquist, J.F. & Harris, R.S. Endogenous origins of HIV-1 G-to-A hypermutation and restriction in the nonpermissive T cell line CEM2n. PLoS Pathog. 8, e1002800 (2012).
Ooms, M. et al. HIV-1 Vif adaptation to human APOBEC3H haplotypes. Cell Host Microbe 14, 411–421 (2013).
Sato, K. et al. APOBEC3D and APOBEC3F potently promote HIV-1 diversification and evolution in humanized mouse model. PLoS Pathog. 10, e1004453 (2014).
Haché, G., Liddament, M.T. & Harris, R.S. The retroviral hypermutation specificity of APOBEC3F and APOBEC3G is governed by the C-terminal DNA cytosine deaminase domain. J. Biol. Chem. 280, 10920–10924 (2005).
Navarro, F. et al. Complementary function of the two catalytic domains of APOBEC3G. Virology 333, 374–386 (2005).
Rathore, A. et al. The local dinucleotide preference of APOBEC3G can be altered from 5′-CC to 5′-TC by a single amino acid substitution. J. Mol. Biol. 425, 4442–4454 (2013).
Harjes, S. et al. Impact of H216 on the DNA binding and catalytic activities of the HIV restriction factor APOBEC3G. J. Virol. 87, 7008–7014 (2013).
Furukawa, A. et al. Structure, interaction and real-time monitoring of the enzymatic reaction of wild-type APOBEC3G. EMBO J. 28, 440–451 (2009).
Khan, M.A. et al. Analysis of the contribution of cellular and viral RNA to the packaging of APOBEC3G into HIV-1 virions. Retrovirology 4, 48 (2007).
Burnett, A. & Spearman, P. APOBEC3G multimers are recruited to the plasma membrane for packaging into human immunodeficiency virus type 1 virus-like particles in an RNA-dependent process requiring the NC basic linker. J. Virol. 81, 5000–5013 (2007).
Svarovskaia, E.S. et al. Human apolipoprotein B mRNA-editing enzyme-catalytic polypeptide-like 3G (APOBEC3G) is incorporated into HIV-1 virions through interactions with viral and nonviral RNAs. J. Biol. Chem. 279, 35822–35828 (2004).
Zennou, V., Perez-Caballero, D., Gottlinger, H. & Bieniasz, P.D. APOBEC3G incorporation into human immunodeficiency virus type 1 particles. J. Virol. 78, 12058–12061 (2004).
Douaisi, M. et al. HIV-1 and MLV Gag proteins are sufficient to recruit APOBEC3G into virus-like particles. Biochem. Biophys. Res. Commun. 321, 566–573 (2004).
Desimmie, B.A. et al. Multiple APOBEC3 restriction factors for HIV-1 and one Vif to rule them all. J. Mol. Biol. 426, 1220–1245 (2014).
Feng, Y., Baig, T.T., Love, R.P. & Chelico, L. Suppression of APOBEC3-mediated restriction of HIV-1 by Vif. Front. Microbiol. 5, 450 (2014).
Jäger, S. et al. Vif hijacks CBF-β to degrade APOBEC3G and promote HIV-1 infection. Nature 481, 371–375 (2012).
Zhang, W., Du, J., Evans, S.L., Yu, Y. & Yu, X.F. T-cell differentiation factor CBF-β regulates HIV-1 Vif-mediated evasion of host restriction. Nature 481, 376–379 (2012).
Kao, S. et al. The human immunodeficiency virus type 1 Vif protein reduces intracellular expression and inhibits packaging of APOBEC3G (CEM15), a cellular inhibitor of virus infectivity. J. Virol. 77, 11398–11407 (2003).
Marin, M., Rose, K.M., Kozak, S.L. & Kabat, D. HIV-1 Vif protein binds the editing enzyme APOBEC3G and induces its degradation. Nat. Med. 9, 1398–1403 (2003).
Sheehy, A.M., Gaddis, N.C. & Malim, M.H. The antiretroviral enzyme APOBEC3G is degraded by the proteasome in response to HIV-1 Vif. Nat. Med. 9, 1404–1407 (2003).
Conticello, S.G., Harris, R.S. & Neuberger, M.S. The Vif protein of HIV triggers degradation of the human antiretroviral DNA deaminase APOBEC3G. Curr. Biol. 13, 2009–2013 (2003).
Russell, R.A. & Pathak, V.K. Identification of two distinct human immunodeficiency virus type 1 Vif determinants critical for interactions with human APOBEC3G and APOBEC3F. J. Virol. 81, 8201–8210 (2007).
Pery, E., Rajendran, K.S., Brazier, A.J. & Gabuzda, D. Regulation of APOBEC3 proteins by a novel YXXL motif in human immunodeficiency virus type 1 Vif and simian immunodeficiency virus SIVagm Vif. J. Virol. 83, 2374–2381 (2009).
Chen, G., He, Z., Wang, T., Xu, R. & Yu, X.F. A patch of positively charged amino acids surrounding the human immunodeficiency virus type 1 Vif SLVx4Yx9Y motif influences its interaction with APOBEC3G. J. Virol. 83, 8674–8682 (2009).
Dang, Y., Wang, X., Zhou, T., York, I.A. & Zheng, Y.H. Identification of a novel WxSLVK motif in the N terminus of human immunodeficiency virus and simian immunodeficiency virus Vif that is critical for APOBEC3G and APOBEC3F neutralization. J. Virol. 83, 8544–8552 (2009).
Guo, Y. et al. Structural basis for hijacking CBF-β and CUL5 E3 ligase complex by HIV-1 Vif. Nature 505, 229–233 (2014).
Chen, K.M. et al. Structure of the DNA deaminase domain of the HIV-1 restriction factor APOBEC3G. Nature 452, 116–119 (2008).
Harjes, E. et al. An extended structure of the APOBEC3G catalytic domain suggests a unique holoenzyme model. J. Mol. Biol. 389, 819–832 (2009).
Holden, L.G. et al. Crystal structure of the anti-viral APOBEC3G catalytic domain and functional implications. Nature 456, 121–124 (2008).
Shandilya, S.M. et al. Crystal structure of the APOBEC3G catalytic domain reveals potential oligomerization interfaces. Structure 18, 28–38 (2010).
Bohn, M.F. et al. Crystal structure of the DNA cytosine deaminase APOBEC3F: the catalytically active and HIV-1 Vif-binding domain. Structure 21, 1042–1050 (2013).
Siu, K.K., Sultana, A., Azimi, F.C. & Lee, J.E. Structural determinants of HIV-1 Vif susceptibility and DNA binding in APOBEC3F. Nat. Commun. 4, 2593 (2013).
Byeon, I.J. et al. NMR structure of human restriction factor APOBEC3A reveals substrate binding and enzyme specificity. Nat. Commun. 4, 1890 (2013).
Kitamura, S. et al. The APOBEC3C crystal structure and the interface for HIV-1 Vif binding. Nat. Struct. Mol. Biol. 19, 1005–1010 (2012).
LaRue, R.S. et al. Guidelines for naming nonprimate APOBEC3 genes and proteins. J. Virol. 83, 494–497 (2009).
Salzmann, M., Pervushin, K., Wider, G., Senn, H. & Wuthrich, K. TROSY in triple-resonance experiments: new perspectives for sequential NMR assignment of large proteins. Proc. Natl. Acad. Sci. USA 95, 13585–13590 (1998).
Pervushin, K., Riek, R., Wider, G. & Wuthrich, K. Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl. Acad. Sci. USA 94, 12366–12371 (1997).
Newman, E.N. et al. Antiviral function of APOBEC3G can be dissociated from cytidine deaminase activity. Curr. Biol. 15, 166–170 (2005).
Russell, R.A., Smith, J., Barr, R., Bhattacharyya, D. & Pathak, V.K. Distinct domains within APOBEC3G and APOBEC3F interact with separate regions of human immunodeficiency virus type 1 Vif. J. Virol. 83, 1992–2003 (2009).
Bogerd, H.P., Doehle, B.P., Wiegand, H.L. & Cullen, B.R. A single amino acid difference in the host APOBEC3G protein controls the primate species specificity of HIV type 1 virion infectivity factor. Proc. Natl. Acad. Sci. USA 101, 3770–3774 (2004).
Mangeat, B., Turelli, P., Liao, S. & Trono, D. A single amino acid determinant governs the species-specific sensitivity of APOBEC3G to Vif action. J. Biol. Chem. 279, 14481–14483 (2004).
Schröfelbauer, B., Chen, D. & Landau, N.R. A single amino acid of APOBEC3G controls its species-specific interaction with virion infectivity factor (Vif). Proc. Natl. Acad. Sci. USA 101, 3927–3932 (2004).
Xu, H. et al. A single amino acid substitution in human APOBEC3G antiretroviral enzyme confers resistance to HIV-1 virion infectivity factor-induced depletion. Proc. Natl. Acad. Sci. USA 101, 5652–5657 (2004).
Yamashita, T., Kamada, K., Hatcho, K., Adachi, A. & Nomaguchi, M. Identification of amino acid residues in HIV-1 Vif critical for binding and exclusion of APOBEC3G/F. Microbes Infect. 10, 1142–1149 (2008).
Lavens, D. et al. Definition of the interacting interfaces of Apobec3G and HIV-1 Vif using MAPPIT mutagenesis analysis. Nucleic Acids Res. 38, 1902–1912 (2010).
Shandilya, S.M., Bohn, M.F. & Schiffer, C.A. A computational analysis of the structural determinants of APOBEC3's catalytic activity and vulnerability to HIV-1 Vif. Virology 471–473, 105–116 (2014).
Carlow, D.C., Short, S.A. & Wolfenden, R. Role of glutamate-104 in generating a transition state analogue inhibitor at the active site of cytidine deaminase. Biochemistry 35, 948–954 (1996).
Chen, K.M. et al. Extensive mutagenesis experiments corroborate a structural model for the DNA deaminase domain of APOBEC3G. FEBS Lett. 581, 4761–4766 (2007).
Bach, D. et al. Characterization of APOBEC3G binding to 7SL RNA. Retrovirology 5, 54 (2008).
Bogerd, H.P. & Cullen, B.R. Single-stranded RNA facilitates nucleocapsid: APOBEC3G complex formation. RNA 14, 1228–1236 (2008).
Wang, T. et al. 7SL RNA mediates virion packaging of the antiviral cytidine deaminase APOBEC3G. J. Virol. 81, 13112–13124 (2007).
Zhang, L. et al. Function analysis of sequences in human APOBEC3G involved in Vif-mediated degradation. Virology 370, 113–121 (2008).
Reingewertz, T.H. et al. Mapping the Vif-A3G interaction using peptide arrays: a basis for anti-HIV lead peptides. Bioorg. Med. Chem. 21, 3523–3532 (2013).
Huthoff, H. & Malim, M.H. Identification of amino acid residues in APOBEC3G required for regulation by human immunodeficiency virus type 1 Vif and Virion encapsidation. J. Virol. 81, 3807–3815 (2007).
Albin, J.S. et al. A single amino acid in human APOBEC3F alters susceptibility to HIV-1 Vif. J. Biol. Chem. 285, 40785–40792 (2010).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Hayashi, K. & Kojima, C. pCold-GST vector: a novel cold-shock vector containing GST tag for soluble protein production. Protein Expr. Purif. 62, 120–127 (2008).
Ikura, M., Kay, L.E. & Bax, A. A novel approach for sequential assignment of 1H, 13C, and 15N spectra of proteins: heteronuclear triple-resonance three-dimensional NMR spectroscopy. Application to calmodulin. Biochemistry 29, 4659–4667 (1990).
Salzmann, M., Pervushin, K., Wider, G., Senn, H. & Wuthrich, K. TROSY in triple-resonance experiments: new perspectives for sequential NMR assignment of large proteins. Proc. Natl. Acad. Sci. USA 95, 13585–13590 (1998).
Yamazaki, T., Lee, W., Arrowsmith, C.H., Muhandiram, D.R. & Kay, L.E. A suite of triple resonance NMR experiments for the backbone assignment of 15N, 13C, 2H labeled proteins with high sensitivity. J. Am. Chem. Soc. 116, 11655–11666 (1994).
Jaipuria, G., Krishnarjuna, B., Mondal, S., Dubey, A. & Atreya, H.S. Amino acid selective labeling and unlabeling for protein resonance assignments. Adv. Exp. Med. Biol. 992, 95–118 (2012).
Wuthrich, K. NMR of Proteins and Nucleic Acids (Wiley, New York, 1986).
Wishart, D.S. et al. 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J. Biomol. NMR 6, 135–140 (1995).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Schwieters, C.D., Kuszewski, J.J., Tjandra, N. & Clore, G.M. The Xplor-NIH NMR molecular structure determination package. J. Magn. Reson. 160, 65–73 (2003).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999).
Koradi, R., Billeter, M. & Wuthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55 (1996).
Laskowski, R.A., Rullmannn, J.A., MacArthur, M.W., Kaptein, R. & Thornton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Albin, J.S., Hache, G., Hultquist, J.F., Brown, W.L. & Harris, R.S. Long-term restriction by APOBEC3F selects human immunodeficiency virus type 1 variants with restored Vif function. J. Virol. 84, 10209–10219 (2010).
Haché, G., Shindo, K., Albin, J.S. & Harris, R.S. Evolution of HIV-1 isolates that use a novel Vif-independent mechanism to resist restriction by human APOBEC3G. Curr. Biol. 18, 819–824 (2008).
Acknowledgements
This work was supported by grants from the US National Institutes of Health (AI073167 to H.M. and GM091743 to R.S.H.). Salary support for T.K. was provided in part by a Toyobo Biotechnology Foundation Fellowship. The University of Minnesota Supercomputing and NMR center (US National Science Foundation, BIR-961477) provided NMR instrumentation. We thank C. Kojima (Institute for Protein Research, Osaka University) for the pCold vector, K. Strebel (US National Institutes of Health) for the pcDNA-hVif plasmid, M. Katahira for advice on structure calculations, Y. Iwatani for advice on degradation assays, Y. Xia for advice on NMR experiments, Y. Hong for supporting preparation of expression vectors, F. Liu for advice on human-cell experiments and K. Walters for editing the manuscript.
Author information
Authors and Affiliations
Contributions
T.K. designed the strategy for searching for soluble A3G NTD mutants and all A3G NTD mutants; performed harvest assays, Vif binding assays in E. coli, Vif-dependent degradation assays and deamination assays; prepared NMR samples; and determined the solution structure. E.M.L. performed single-cycle infectivity assays. M.S. optimized human-cell experiments. S.M.D.S. analyzed surface-charge distribution and binding-pocket volumes of the NTD structure. J.Z. provided support in harvest assays. L.C. and M.H. provided support with preparation of expression plasmids and with the Vif binding assay in E. coli. T.K., E.M.L., S.M.D.S., C.A.S., R.S.H. and H.M. contributed to writing and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Amino acid substitutions for increasing solubility of the A3G N-terminal domain.
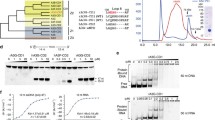
(a) Amino acid sequence alignments of A3G NTD (NP_068594), sNTD (this study), consensus (this study), A3G CTD (NP_068594), A3B CTD (NP_004891) and A3A (NP_663745). Identical and mismatched amino acid residues between wild-type NTD and sNTD are shown in red and gray, respectively. Residue numbers correspond to wild-type A3G NTD. (b) Schematic presentation of amino acid sequences of sNTD and its variants. Residues identical to wild-type NTD are illustrated in red, whereas those that differ are indicated in gray. Black triangles indicate the location of residues that were substituted back to their endogenous amino acid identity with sNTD as a template. (c) Results of protein harvest assays showing solubility of sNTD variants. The band intensities of GST fused sNTD (lane 1), wild-type A3G NTD (lane 2), A3G CTD-2K3A (lane 3), and sNTD variants, including #1 - #12 of figure 1b, in a Coomassie gel were analyzed and normalized using the value of sNTD. The mean values ± s. d. for n = 3 independent cell cultures is plotted.
Supplementary Figure 2 Cα region of 1H, 13C HSQC spectra acquired on sNTD samples with selective 1H- and 13C-labeling by amino acid type.
Spectrum acquired on sNTD that was (a) uniformly 13C-labeled, (b) [1H, 13C]Ala-labeled, (c) [1H, 13C]Phe-labeled, (d) [1H, 13C]Ile-labeled, (e) [1H, 13C]Lys-labeled, (f) [1H, 13C]Leu-labeled, (g) [1H, 13C]Met-labeled, (h) [1H, 13C]Arg-labeled, (i) [1H, 13C]Thr-labeled, (j) [1H, 13C]Tyr-labeled, and (k) [1H, 13C]Val-labeled. Assignments are provided in spectra.
Supplementary Figure 3 1H-15N TROSY spectrum of sNTD.
Assignments of backbone amide protons are presented next to signals. Signals labeled with asterisks seem to arise from minor conformations. The inlet figure shows that major and minor signals for E38, V39 and K40.
Supplementary Figure 4 β-sheet conformation found in sNTD.
Inter-strand NOE connectivities are shown by double-headed arrows, and hydrogen bonds identified using an H/D-exchange experiment are indicated with red oval rectangles.
Supplementary Figure 5 Deamination and Vif binding assays.
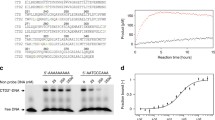
(a-f) Deamination assays. 10mer DNA fragment, d(5’-ATTCCCAATT-3’), was used as a substrate for the deamination reaction. (a-c) display spectra of the substrate (a), the first deamination product d(5’-ATTCCUAATT-3’) (b) and the second deamination product d(5’-ATTCUUAATT-3’) (c). (d-e) Spectra of the substrate incubated with A3G CTD-2K3A for 30 minutes (d) and 24 hours (e). (f) Spectrum of the substrate after incubation with sNTD for 24 hours. (g) Representative SDS-PAGE gel images of the E. coli co-expression and GST-pull down assays used to identify residues critical for sNTD binding to Vif. Substituted residues are indicated on top of each gel image. In each gel, P, S and B show the amount of protein in pellet, soluble fraction and GST resin, respectively. The position of each protein is shown on the right side of the figure.
Supplementary Figure 6 Structural features of the zinc-coordinating region of sNTD.
(a) Stereoview of the zinc coordinating region of sNTD. α2 and α3 helices are colored green, the zinc ion is colored purple, and side chain atoms of zinc-coordinating residues, including H65, C97 and C100, and E67, are shown in ball-and-stick representation. Side chain atoms of L35, W90, I92 and M104 are located close to the zinc ion (see main text). (b) Comparison of the substrate-binding pocket and surface properties of sNTD, the A3G-NTD homology model (A3G NTD), and A3G-CTD. Top panels: Substrate-binding pocket volumes are presented as orange blobs, with the protein molecular surface shown as dark-blue mesh. Bottom panels: Molecular surface electrostatic potential are shown in red and blue for negative and positive charges, respectively. (bottom). Orientation of molecule is shown on left by cartoon models (Zn2+ is shown in purple). Location of D128 residue is shown in NTD structures. (c-d) A hydrophobic core formed by β1-α2-β3-α3. (c) Amino acid sequence alignment of wild-type A3G NTD, sNTD and A3 domains with available structures. Residue numbers refer to A3G NTD. Residues involved in the hydrophobic core are colored red. (d) A stereoview of the hydrophobic core in the sNTD structure.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 765 kb)
Supplementary Data Set 1
Uncropped gel images (PDF 2079 kb)
Rights and permissions
About this article
Cite this article
Kouno, T., Luengas, E., Shigematsu, M. et al. Structure of the Vif-binding domain of the antiviral enzyme APOBEC3G. Nat Struct Mol Biol 22, 485–491 (2015). https://doi.org/10.1038/nsmb.3033
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3033
- Springer Nature America, Inc.
This article is cited by
-
HIV-1 Vif protein sequence variations in South African people living with HIV and their influence on Vif-APOBEC3G interaction
European Journal of Clinical Microbiology & Infectious Diseases (2024)
-
Structural basis of sequence-specific RNA recognition by the antiviral factor APOBEC3G
Nature Communications (2022)
-
Structural basis of antagonism of human APOBEC3F by HIV-1 Vif
Nature Structural & Molecular Biology (2019)
-
Crystal structure of the catalytic domain of HIV-1 restriction factor APOBEC3G in complex with ssDNA
Nature Communications (2018)
-
Identification of small molecule compounds targeting the interaction of HIV-1 Vif and human APOBEC3G by virtual screening and biological evaluation
Scientific Reports (2018)