Key Points
-
Centrosome duplication is tightly regulated to ensure that centrioles duplicate only once per cell cycle and that only one new centriole is produced per pre-existing centriole
-
Phosphorylation events have an important role in controlling centriole number. Polo-like kinase 1 (PLK1) has a key function in cell cycle control of centriole duplication, whereas PLK4 takes centre stage in controlling centriole copy number
-
Recent work has uncovered the existence of distinct signalling pathways that limit the proliferation of cells with an increase or decrease in centrosome number
-
The presence of extra centrosomes can endow cells with oncogenic properties. However, overcoming the inhibitory effect of extra centrosomes on cell proliferation is necessary to allow cells with extra centrosomes to sustain the cell divisions required for tumour development
-
Primary microcephaly may be caused by deregulation of centriole numbers and, potentially, by pathological activation of the mitotic surveillance pathway, and in consequence cell cycle arrest or apoptosis, in the developing brain
Abstract
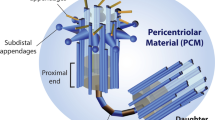
Centrioles are conserved microtubule-based organelles that form the core of the centrosome and act as templates for the formation of cilia and flagella. Centrioles have important roles in most microtubule-related processes, including motility, cell division and cell signalling. To coordinate these diverse cellular processes, centriole number must be tightly controlled. In cycling cells, one new centriole is formed next to each pre-existing centriole in every cell cycle. Advances in imaging, proteomics, structural biology and genome editing have revealed new insights into centriole biogenesis, how centriole numbers are controlled and how alterations in these processes contribute to diseases such as cancer and neurodevelopmental disorders. Moreover, recent work has uncovered the existence of surveillance pathways that limit the proliferation of cells with numerical centriole aberrations. Owing to this progress, we now have a better understanding of the molecular mechanisms governing centriole biogenesis, opening up new possibilities for targeting these pathways in the context of human disease.




Similar content being viewed by others
References
Nigg, E. A. & Raff, J. W. Centrioles, centrosomes, and cilia in health and disease. Cell 139, 663–678 (2009).
Bornens, M. The centrosome in cells and organisms. Science 335, 422–426 (2012).
Conduit, P. T., Wainman, A. & Raff, J. W. Centrosome function and assembly in animal cells. Nat. Rev. Mol. Cell Biol. 16, 611–624 (2015).
Fu, J., Hagan, I. M. & Glover, D. M. The centrosome and its duplication cycle. Cold Spring Harb. Perspect. Biol. 7, a015800 (2015).
Woodruff, J. B., Wueseke, O. & Hyman, A. A. Pericentriolar material structure and dynamics. Philos. Trans. R. Soc. Lond. B Biol Sci. 369, 20130459 (2014).
Arquint, C., Gabryjonczyk, A. M. & Nigg, E. A. Centrosomes as signalling centres. Philos. Trans. R. Soc. Lond. B Biol Sci. 369, 20130464 (2014).
Sanchez, I. & Dynlacht, B. D. Cilium assembly and disassembly. Nat. Cell Biol. 18, 711–717 (2016).
Braun, D. A. & Hildebrandt, F. Ciliopathies. Cold Spring Harb. Perspect. Biol. 9, a028191 (2016).
Bettencourt-Dias, M., Hildebrandt, F., Pellman, D., Woods, G. & Godinho, S. A. Centrosomes and cilia in human disease. Trends Genet. 27, 307–315 (2011).
Gonczy, P. Centrosomes and cancer: revisiting a long-standing relationship. Nat. Rev. Cancer 15, 639–652 (2015).
Azimzadeh, J. & Marshall, W. F. Building the centriole. Curr. Biol. 20, R816–R825 (2010).
Gonczy, P. Towards a molecular architecture of centriole assembly. Nat. Rev. Mol. Cell Biol. 13, 425–435 (2012).
Gupta, G. D. et al. A Dynamic protein interaction landscape of the human centrosome-cilium interface. Cell 163, 1484–1499 (2015). This study is a large proteomics effort to probe protein interactions at the centrosome–cilium interface.
Garcia, G. 3rd & Reiter, J. F. A primer on the mouse basal body. Cilia 5, 17 (2016).
Hirono, M. Cartwheel assembly. Philos. Trans. R. Soc. Lond. B Biol Sci. 369, 20130458 (2014).
Keller, D. et al. Mechanisms of HsSAS-6 assembly promoting centriole formation in human cells. J. Cell Biol. 204, 697–712 (2014).
Guichard, P., Chretien, D., Marco, S. & Tassin, A. M. Procentriole assembly revealed by cryo-electron tomography. EMBO J. 29, 1565–1572 (2010).
Guichard, P. et al. Cartwheel architecture of Trichonympha basal body. Science 337, 553 (2012).
Guichard, P. et al. Native architecture of the centriole proximal region reveals features underlying its 9-fold radial symmetry. Curr. Biol. 23, 1620–1628 (2013).
Bauer, M., Cubizolles, F., Schmidt, A. & Nigg, E. A. Quantitative analysis of human centrosome architecture by targeted proteomics and fluorescence imaging. EMBO J. 35, 2152–2166 (2016). This study is an initial effort at obtaining quantitative information about the abundance of centrosomal proteins within cells and isolated organelles.
Kitagawa, D. et al. Structural basis of the 9-fold symmetry of centrioles. Cell 144, 364–375 (2011).
van Breugel, M. et al. Structures of SAS-6 suggest its organization in centrioles. Science 331, 1196–1199 (2011).
Guichard, P. et al. Cell-free reconstitution reveals centriole cartwheel assembly mechanisms. Nat. Commun. 8, 14813 (2017). This is a pioneering study demonstrating successful in vitro reconstitution of early steps of centriole assembly.
Wang, W. J. et al. De novo centriole formation in human cells is error-prone and does not require SAS-6 self-assembly. eLife 4, e10586 (2015).
Hilbert, M. et al. SAS-6 engineering reveals interdependence between cartwheel and microtubules in determining centriole architecture. Nat. Cell Biol. 18, 393–403 (2016).
Ohta, M. et al. Direct interaction of Plk4 with STIL ensures formation of a single procentriole per parental centriole. Nat. Commun. 5, 5267 (2014).
Dzhindzhev, N. S. et al. Plk4 phosphorylates Ana2 to trigger Sas6 recruitment and procentriole formation. Curr. Biol. 24, 2526–2532 (2014).
Moyer, T. C., Clutario, K. M., Lambrus, B. G., Daggubati, V. & Holland, A. J. Binding of STIL to Plk4 activates kinase activity to promote centriole assembly. J. Cell Biol. 209, 863–878 (2015).
Kratz, A. S., Barenz, F., Richter, K. T. & Hoffmann, I. Plk4-dependent phosphorylation of STIL is required for centriole duplication. Biol. Open 4, 370–377 (2015).
Stevens, N. R., Dobbelaere, J., Brunk, K., Franz, A. & Raff, J. W. Drosophila Ana2 is a conserved centriole duplication factor. J. Cell Biol. 188, 313–323 (2010).
Arquint, C., Sonnen, K. F., Stierhof, Y. D. & Nigg, E. A. Cell-cycle-regulated expression of STIL controls centriole number in human cells. J. Cell Sci. 125, 1342–1352 (2012).
Tang, C. J. et al. The human microcephaly protein STIL interacts with CPAP and is required for procentriole formation. EMBO J. 30, 4790–4804 (2011).
Hiraki, M., Nakazawa, Y., Kamiya, R. & Hirono, M. Bld10p constitutes the cartwheel-spoke tip and stabilizes the 9-fold symmetry of the centriole. Curr. Biol. 17, 1778–1783 (2007).
Lin, Y. C. et al. Human microcephaly protein CEP135 binds to hSAS-6 and CPAP, and is required for centriole assembly. EMBO J. 32, 1141–1154 (2013).
Sonnen, K. F., Schermelleh, L., Leonhardt, H. & Nigg, E. A. 3D-structured illumination microscopy provides novel insight into architecture of human centrosomes. Biol. Open 1, 965–976 (2012).
Kleylein-Sohn, J. et al. Plk4-induced centriole biogenesis in human cells. Dev. Cell 13, 190–202 (2007).
Pelletier, L., O'Toole, E., Schwager, A., Hyman, A. A. & Muller-Reichert, T. Centriole assembly in Caenorhabditis elegans. Nature 444, 619–623 (2006).
Sharma, A. et al. Centriolar CPAP/SAS-4 imparts slow processive microtubule growth. Dev. Cell 37, 362–376 (2016).
Zheng, X. et al. Molecular basis for CPAP-tubulin interaction in controlling centriolar and ciliary length. Nat. Commun. 7, 11874 (2016).
Galletta, B. J., Jacobs, K. C., Fagerstrom, C. J. & Rusan, N. M. Asterless is required for centriole length control and sperm development. J. Cell Biol. 213, 435–450 (2016).
Marshall, W. F. Cell geometry: how cells count and measure size. Annu. Rev. Biophys. 45, 49–64 (2016).
Delgehyr, N. et al. Klp10A, a microtubule-depolymerizing kinesin-13, cooperates with CP110 to control Drosophila centriole length. Curr. Biol. 22, 502–509 (2012).
Kobayashi, T., Tsang, W. Y., Li, J., Lane, W. & Dynlacht, B. D. Centriolar kinesin Kif24 interacts with CP110 to remodel microtubules and regulate ciliogenesis. Cell 145, 914–925 (2011).
Franz, A., Roque, H., Saurya, S., Dobbelaere, J. & Raff, J. W. CP110 exhibits novel regulatory activities during centriole assembly in Drosophila. J. Cell Biol. 203, 785–799 (2013).
Schmidt, T. I. et al. Control of centriole length by CPAP and CP110. Curr. Biol. 19, 1005–1011 (2009).
Tsang, W. Y. et al. CP110 suppresses primary cilia formation through its interaction with CEP290, a protein deficient in human ciliary disease. Dev. Cell 15, 187–197 (2008).
Al-Hakim, A. K., Bashkurov, M., Gingras, A. C., Durocher, D. & Pelletier, L. Interaction proteomics identify NEURL4 and the HECT E3 ligase HERC2 as novel modulators of centrosome architecture. Mol. Cell. Proteomics 11, M111.014233 (2012).
Li, J. et al. USP33 regulates centrosome biogenesis via deubiquitination of the centriolar protein CP110. Nature 495, 255–259 (2013).
Li, J. et al. Neurl4, a novel daughter centriole protein, prevents formation of ectopic microtubule organizing centres. EMBO Rep. 13, 547–553 (2012).
Cao, J. et al. miR-129-3p controls cilia assembly by regulating CP110 and actin dynamics. Nat. Cell Biol. 14, 697–706 (2012).
Kohlmaier, G. et al. Overly long centrioles and defective cell division upon excess of the SAS-4-related protein CPAP. Curr. Biol. 19, 1012–1018 (2009).
Tang, C. J., Fu, R. H., Wu, K. S., Hsu, W. B. & Tang, T. K. CPAP is a cell-cycle regulated protein that controls centriole length. Nat. Cell Biol. 11, 825–831 (2009).
Comartin, D. et al. CEP120 and SPICE1 cooperate with CPAP in centriole elongation. Curr. Biol. 23, 1360–1366 (2013).
Lin, Y. N. et al. CEP120 interacts with CPAP and positively regulates centriole elongation. J. Cell Biol. 202, 211–219 (2013).
Keller, L. C. et al. Molecular architecture of the centriole proteome: the conserved WD40 domain protein POC1 is required for centriole duplication and length control. Mol. Biol. Cell 20, 1150–1166 (2009).
Azimzadeh, J. et al. hPOC5 is a centrin-binding protein required for assembly of full-length centrioles. J. Cell Biol. 185, 101–114 (2009).
Chang, C. W., Hsu, W. B., Tsai, J. J., Tang, C. J. & Tang, T. K. CEP295 interacts with microtubules and is required for centriole elongation. J. Cell Sci. 129, 2501–2513 (2016).
Saurya, S. et al. Drosophila Ana1 is required for centrosome assembly and centriole elongation. J. Cell Sci. 129, 2514–2525 (2016).
Bobinnec, Y. et al. Centriole disassembly in vivo and its effect on centrosome structure and function in vertebrate cells. J. Cell Biol. 143, 1575–1589 (1998).
Janke, C. & Bulinski, J. C. Post-translational regulation of the microtubule cytoskeleton: mechanisms and functions. Nat. Rev. Mol. Cell Biol. 12, 773–786 (2011).
Andersen, J. S. et al. Proteomic characterization of the human centrosome by protein correlation profiling. Nature 426, 570–574 (2003).
Jakobsen, L. et al. Novel asymmetrically localizing components of human centrosomes identified by complementary proteomics methods. EMBO J. 30, 1520–1535 (2011).
Prosser, S. L. & Pelletier, L. Mitotic spindle assembly in animal cells: a fine balancing act. Nat. Rev. Mol. Cell Biol. 18, 187–201 (2017).
Hori, A. & Toda, T. Regulation of centriolar satellite integrity and its physiology. Cell. Mol. Life Sci. 74, 213–229 (2017).
Fu, J. & Glover, D. M. Structured illumination of the interface between centriole and peri-centriolar material. Open Biol. 2, 120104 (2012).
Lawo, S., Hasegan, M., Gupta, G. D. & Pelletier, L. Subdiffraction imaging of centrosomes reveals higher-order organizational features of pericentriolar material. Nat. Cell Biol. 14, 1148–1158 (2012).
Mennella, V. et al. Subdiffraction-resolution fluorescence microscopy reveals a domain of the centrosome critical for pericentriolar material organization. Nat. Cell Biol. 14, 1159–1168 (2012).
Conduit, P. T. et al. The centrosome-specific phosphorylation of Cnn by Polo/Plk1 drives Cnn scaffold assembly and centrosome maturation. Dev. Cell 28, 659–669 (2014).
Conduit, P. T. et al. A molecular mechanism of mitotic centrosome assembly in Drosophila. eLife 3, e03399 (2014).
Feng, Z. et al. Structural basis for mitotic centrosome assembly in flies. Cell 169, 1078–1089.e13 (2017). This structural study focusing on the D. melanogaster PCM scaffolding protein Cnn provides insight into molecular interactions required for mitotic centrosome assembly.
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Zwicker, D., Decker, M., Jaensch, S., Hyman, A. A. & Julicher, F. Centrosomes are autocatalytic droplets of pericentriolar material organized by centrioles. Proc. Natl Acad. Sci. USA 111, E2636–2645 (2014).
Woodruff, J. B. et al. Regulated assembly of a supramolecular centrosome scaffold in vitro. Science 348, 808–812 (2015).
Woodruff, J. B. et al. The centrosome is a selective condensate that nucleates microtubules by concentrating tubulin. Cell 169, 1066–1077.e10 (2017). This study shows that the C. elegans PCM protein SPD-5 forms a selective condensate able to nucleate microtubules, suggesting that PCM formation in vivo involves a phase separation process (compare with reference 70).
Nigg, E. A. Centrosome duplication: of rules and licenses. Trends Cell Biol. 17, 215–221 (2007).
Tsou, M. F. & Stearns, T. Mechanism limiting centrosome duplication to once per cell cycle. Nature 442, 947–951 (2006).
Tsou, M. F. et al. Polo kinase and separase regulate the mitotic licensing of centriole duplication in human cells. Dev. Cell 17, 344–354 (2009).
Loncarek, J., Hergert, P., Magidson, V. & Khodjakov, A. Control of daughter centriole formation by the pericentriolar material. Nat. Cell Biol. 10, 322–328 (2008).
Matsuo, K. et al. Kendrin is a novel substrate for separase involved in the licensing of centriole duplication. Curr. Biol. 22, 915–921 (2012).
Lee, K. & Rhee, K. Separase-dependent cleavage of pericentrin B is necessary and sufficient for centriole disengagement during mitosis. Cell Cycle 11, 2476–2485 (2012).
Kim, J., Lee, K. & Rhee, K. PLK1 regulation of PCNT cleavage ensures fidelity of centriole separation during mitotic exit. Nat. Commun. 6, 10076 (2015). This study, along with reference 79, identifies pericentrin as a key substrate of separase, which is important for licensing of centriole duplication.
Schockel, L., Mockel, M., Mayer, B., Boos, D. & Stemmann, O. Cleavage of cohesin rings coordinates the separation of centrioles and chromatids. Nat. Cell Biol. 13, 966–972 (2011).
Oliveira, R. A. & Nasmyth, K. Cohesin cleavage is insufficient for centriole disengagement in Drosophila. Curr. Biol. 23, R601–603 (2013).
Kuriyama, R. & Borisy, G. G. Centriole cycle in Chinese hamster ovary cells as determined by whole-mount electron microscopy. J. Cell Biol. 91, 814–821 (1981).
Shukla, A., Kong, D., Sharma, M., Magidson, V. & Loncarek, J. Plk1 relieves centriole block to reduplication by promoting daughter centriole maturation. Nat. Commun. 6, 8077 (2015). This is a correlative live and electron microscopy study emphasizing the role of PLK1 in triggering early licensing events important for a new round of centriole duplication.
Arquint, C. & Nigg, E. A. STIL microcephaly mutations interfere with APC/C-mediated degradation and cause centriole amplification. Curr. Biol. 24, 351–360 (2014).
Kim, M. et al. Promotion and suppression of centriole duplication are catalytically coupled through PLK4 to ensure centriole homeostasis. Cell Rep. 16, 1195–1203 (2016).
Izquierdo, D., Wang, W. J., Uryu, K. & Tsou, M. F. Stabilization of cartwheel-less centrioles for duplication requires CEP295-mediated centriole-to-centrosome conversion. Cell Rep. 8, 957–965 (2014).
Fu, J. et al. Conserved molecular interactions in centriole-to-centrosome conversion. Nat. Cell Biol. 18, 87–99 (2016). This study illuminates an evolutionarily conserved mechanism underlying centriole-to-centrosome conversion.
Wang, W. J., Soni, R. K., Uryu, K. & Tsou, M. F. The conversion of centrioles to centrosomes: essential coupling of duplication with segregation. J. Cell Biol. 193, 727–739 (2011). This study emphasizes a key role for PLK1 in both centriole disengagement and PCM assembly on procentrioles, two mitotic events critical for licensing centrioles for the next round of duplication.
Novak, Z. A., Wainman, A., Gartenmann, L. & Raff, J. W. Cdk1 Phosphorylates Drosophila Sas-4 to recruit Polo to daughter centrioles and convert them to centrosomes. Dev. Cell 37, 545–557 (2016). This study describes an early key step leading to centrosomal recruitment of PLK1, which is important for centriole-to-centrosome conversion (see also references 85, 89 and 90).
Novak, Z. A., Conduit, P. T., Wainman, A. & Raff, J. W. Asterless licenses daughter centrioles to duplicate for the first time in Drosophila embryos. Curr. Biol. 24, 1276–1282 (2014).
Tsuchiya, Y., Yoshiba, S., Gupta, A., Watanabe, K. & Kitagawa, D. Cep295 is a conserved scaffold protein required for generation of a bona fide mother centriole. Nat. Commun. 7, 12567 (2016).
Hatch, E. M., Kulukian, A., Holland, A. J., Cleveland, D. W. & Stearns, T. Cep152 interacts with Plk4 and is required for centriole duplication. J. Cell Biol. 191, 721–729 (2010).
Cizmecioglu, O. et al. Cep152 acts as a scaffold for recruitment of Plk4 and CPAP to the centrosome. J. Cell Biol. 191, 731–739 (2010).
Kim, T. S. et al. Hierarchical recruitment of Plk4 and regulation of centriole biogenesis by two centrosomal scaffolds, Cep192 and Cep152. Proc. Natl Acad. Sci. USA 110, E4849–4857 (2013).
Sonnen, K. F., Gabryjonczyk, A. M., Anselm, E., Stierhof, Y. D. & Nigg, E. A. Human Cep192 and Cep152 cooperate in Plk4 recruitment and centriole duplication. J. Cell Sci. 126, 3223–3233 (2013).
Park, S. Y. et al. Molecular basis for unidirectional scaffold switching of human Plk4 in centriole biogenesis. Nat. Struct. Mol. Biol. 21, 696–703 (2014).
Sugioka, K. et al. Centriolar SAS-7 acts upstream of SPD-2 to regulate centriole assembly and pericentriolar material formation. eLife 6, e20353 (2017).
Habedanck, R., Stierhof, Y. D., Wilkinson, C. J. & Nigg, E. A. The Polo kinase Plk4 functions in centriole duplication. Nat. Cell Biol. 7, 1140–1146 (2005).
Bettencourt-Dias, M. et al. SAK/PLK4 is required for centriole duplication and flagella development. Curr. Biol. 15, 2199–2207 (2005).
Cunha-Ferreira, I. et al. Regulation of autophosphorylation controls PLK4 self-destruction and centriole number. Curr. Biol. 23, 2245–2254 (2013).
Holland, A. J., Lan, W., Niessen, S., Hoover, H. & Cleveland, D. W. Polo-like kinase 4 kinase activity limits centrosome overduplication by autoregulating its own stability. J. Cell Biol. 188, 191–198 (2010).
Klebba, J. E. et al. Polo-like kinase 4 autodestructs by generating its Slimb-binding phosphodegron. Curr. Biol. 23, 2255–2261 (2013).
Guderian, G., Westendorf, J., Uldschmid, A. & Nigg, E. A. Plk4 trans-autophosphorylation regulates centriole number by controlling betaTrCP-mediated degradation. J. Cell Sci. 123, 2163–2169 (2010).
Holland, A. J. et al. The autoregulated instability of Polo-like kinase 4 limits centrosome duplication to once per cell cycle. Genes Dev. 26, 2684–2689 (2012).
Lopes, C. A. et al. PLK4 trans-autoactivation controls centriole biogenesis in space. Dev. Cell 35, 222–235 (2015).
Arquint, C. et al. STIL binding to Polo-box 3 of PLK4 regulates centriole duplication. eLife 4, e07888 (2015).
Lettman, M. M. et al. Direct binding of SAS-6 to ZYG-1 recruits SAS-6 to the mother centriole for cartwheel assembly. Dev. Cell 25, 284–298 (2013).
Cottee, M. A. et al. Crystal structures of the CPAP/STIL complex reveal its role in centriole assembly and human microcephaly. eLife 2, e01071 (2013).
Hatzopoulos, G. N. et al. Structural analysis of the G-box domain of the microcephaly protein CPAP suggests a role in centriole architecture. Structure 21, 2069–2077 (2013).
Zheng, X. et al. Conserved TCP domain of Sas-4/CPAP is essential for pericentriolar material tethering during centrosome biogenesis. Proc. Natl Acad. Sci. USA 111, E354–E363 (2014).
Goryachev, A. B. & Leda, M. Many roads to symmetry breaking: molecular mechanisms and theoretical models of yeast cell polarity. Mol. Biol. Cell 28, 370–380 (2017).
Fong, C. S., Kim, M., Yang, T. T., Liao, J. C. & Tsou, M. F. SAS-6 assembly templated by the lumen of cartwheel-less centrioles precedes centriole duplication. Dev. Cell 30, 238–245 (2014).
Zitouni, S. et al. CDK1 prevents unscheduled PLK4-STIL complex assembly in centriole biogenesis. Curr. Biol. 26, 1127–1137 (2016).
Peel, N. et al. Protein phosphatase 1 down regulates ZYG-1 levels to limit centriole duplication. PLoS Genet. 13, e1006543 (2017).
Agircan, F. G., Schiebel, E. & Mardin, B. R. Separate to operate: control of centrosome positioning and separation. Philos. Trans. R. Soc. Lond. B Biol Sci. 369, 20130461 (2014).
Mayor, T., Stierhof, Y. D., Tanaka, K., Fry, A. M. & Nigg, E. A. The centrosomal protein C-Nap1 is required for cell cycle-regulated centrosome cohesion. J. Cell Biol. 151, 837–846 (2000).
Nigg, E. A. & Stearns, T. The centrosome cycle: Centriole biogenesis, duplication and inherent asymmetries. Nat. Cell Biol. 13, 1154–1160 (2011).
Kong, D. et al. Centriole maturation requires regulated Plk1 activity during two consecutive cell cycles. J. Cell Biol. 206, 855–865 (2014).
Lane, H. A. & Nigg, E. A. Antibody microinjection reveals an essential role for human polo-like kinase 1 (Plk1) in the functional maturation of mitotic centrosomes. J. Cell Biol. 135, 1701–1713 (1996).
Lee, K. & Rhee, K. PLK1 phosphorylation of pericentrin initiates centrosome maturation at the onset of mitosis. J. Cell Biol. 195, 1093–1101 (2011).
Joukov, V., Walter, J. C. & De Nicolo, A. The Cep192-organized aurora A-Plk1 cascade is essential for centrosome cycle and bipolar spindle assembly. Mol. Cell 55, 578–591 (2014).
Laos, T., Cabral, G. & Dammermann, A. Isotropic incorporation of SPD-5 underlies centrosome assembly in C. elegans. Curr. Biol. 25, R648–649 (2015).
Spassky, N. & Meunier, A. The development and functions of multiciliated epithelia. Nat. Rev. Mol. Cell Biol. 18, 423–436 (2017).
Azimzadeh, J., Wong, M. L., Downhour, D. M., Sanchez Alvarado, A. & Marshall, W. F. Centrosome loss in the evolution of planarians. Science 335, 461–463 (2012). By showing that in planarians centrioles assemble only in terminally differentiating ciliated cells but are not required for mitotic cell divisions, this study has important implications for the understanding of the evolution of the animal centrosome and its role in development.
Basto, R. et al. Flies without centrioles. Cell 125, 1375–1386 (2006).
Debec, A., Sullivan, W. & Bettencourt-Dias, M. Centrioles: active players or passengers during mitosis? Cell. Mol. Life Sci. 67, 2173–2194 (2010).
Szollosi, D., Calarco, P. & Donahue, R. P. Absence of centrioles in the first and second meiotic spindles of mouse oocytes. J. Cell Sci. 11, 521–541 (1972).
Howe, K. & FitzHarris, G. A non-canonical mode of microtubule organization operates throughout pre-implantation development in mouse. Cell Cycle 12, 1616–1624 (2013).
Coelho, P. A. et al. Spindle formation in the mouse embryo requires Plk4 in the absence of centrioles. Dev. Cell 27, 586–597 (2013).
Bazzi, H. & Anderson, K. V. Acentriolar mitosis activates a p53-dependent apoptosis pathway in the mouse embryo. Proc. Natl Acad. Sci. USA 111, E1491–1500 (2014).
Wong, Y. L. et al. Reversible centriole depletion with an inhibitor of Polo-like kinase 4. Science 348, 1155–1160 (2015).
Lambrus, B. G. et al. p53 protects against genome instability following centriole duplication failure. J. Cell Biol. 210, 63–77 (2015).
Lambrus, B. G. et al. A USP28-53BP1-p53-p21 signaling axis arrests growth after centrosome loss or prolonged mitosis. J. Cell Biol. 214, 143–153 (2016). This study uses genome-wide screening to uncover a surveillance pathway involving 53BP1 and USP28 that acts to activate p53 in response to loss of centrosomes or extended duration of mitosis.
Fong, C. S. et al. 53BP1 and USP28 mediate p53-dependent cell cycle arrest in response to centrosome loss and prolonged mitosis. Elife 5 (2016). This study uses CRISPR screening to identify 53BP1 and USP28 as components acting upstream of p53 to arrest the cell cycle in response to centrosome loss or an increased duration of mitosis.
Meitinger, F. et al. 53BP1 and USP28 mediate p53 activation and G1 arrest after centrosome loss or extended mitotic duration. J. Cell Biol. 214, 155–166 (2016). This study presents results in line with those from references 135 and 136 and additionally shows that loss of the ubiquitin ligase TRIM37 enables the formation of extra-centrosomal microtubule organizing centres, thereby allowing the cell to bypass cell cycle arrest caused by centrosome loss.
Knobel, P. A. et al. USP28 is recruited to sites of DNA damage by the tandem BRCT domains of 53BP1 but plays a minor role in double-strand break metabolism. Mol. Cell. Biol. 34, 2062–2074 (2014).
Zhang, D., Zaugg, K., Mak, T. W. & Elledge, S. J. A role for the deubiquitinating enzyme USP28 in control of the DNA-damage response. Cell 126, 529–542 (2006).
Zimmermann, M. & de Lange, T. 53BP1: pro choice in DNA repair. Trends Cell Biol. 24, 108–117 (2014).
Cuella-Martin, R. et al. 53BP1 integrates DNA repair and p53-dependent cell fate decisions via distinct mechanisms. Mol. Cell 64, 51–64 (2016).
Uetake, Y. & Sluder, G. Prolonged prometaphase blocks daughter cell proliferation despite normal completion of mitosis. Curr. Biol. 20, 1666–1671 (2010).
Schulein-Volk, C. et al. Dual regulation of Fbw7 function and oncogenic transformation by Usp28. Cell Rep. 9, 1099–1109 (2014).
Diefenbacher, M. E. et al. The deubiquitinase USP28 controls intestinal homeostasis and promotes colorectal cancer. J. Clin. Invest. 124, 3407–3418 (2014).
Lambrus, B. G. & Holland, A. J. A new mode of mitotic surveillance. Trends Cell Biol. 27, 314–321 (2017).
Levine, M. S. et al. Centrosome amplification is sufficient to promote spontaneous tumorigenesis in mammals. Dev. Cell 40, 313–322.e5 (2017). Using a mouse model allowing moderately increased expression of PLK4, this study provides long-awaited evidence for the hypothesis that centrosome amplification is sufficient to promote spontaneous tumorigenesis.
Ganem, N. J. et al. Cytokinesis failure triggers hippo tumor suppressor pathway activation. Cell 158, 833–848 (2014). This study reports that cytokinesis failure and concomitant centrosome amplification result in the activation of the Hippo pathway.
Fava, L. L. et al. The PIDDosome activates p53 in response to supernumerary centrosomes. Genes Dev. 31, 34–45 (2017). This study reports that PIDDosome–caspase 2 axis activation has a key role in stabilizing p53 and provides a means to halt cell proliferation in response to centrosome amplification (see also reference 147).
Tinel, A. & Tschopp, J. The PIDDosome, a protein complex implicated in activation of caspase-2 in response to genotoxic stress. Science 304, 843–846 (2004).
Chan, J. Y. A clinical overview of centrosome amplification in human cancers. Int. J. Biol. Sci. 7, 1122–1144 (2011).
Schnerch, D. & Nigg, E. A. Structural centrosome aberrations favor proliferation by abrogating microtubule-dependent tissue integrity of breast epithelial mammospheres. Oncogene 35, 2711–2722 (2016).
Basto, R. et al. Centrosome amplification can initiate tumorigenesis in flies. Cell 133, 1032–1042 (2008).
Sabino, D. et al. Moesin is a major regulator of centrosome behavior in epithelial cells with extra centrosomes. Curr. Biol. 25, 879–889 (2015).
Marthiens, V. et al. Centrosome amplification causes microcephaly. Nat. Cell Biol. 15, 731–740 (2013).
Vitre, B. et al. Chronic centrosome amplification without tumorigenesis. Proc. Natl Acad. Sci. USA 112, E6321–E6330 (2015).
Sercin, O. et al. Transient PLK4 overexpression accelerates tumorigenesis in p53-deficient epidermis. Nat. Cell Biol. 18, 100–110 (2016).
Coelho, P. A. et al. Over-expression of Plk4 induces centrosome amplification, loss of primary cilia and associated tissue hyperplasia in the mouse. Open Biol. 5, 150209 (2015).
Kulukian, A. et al. Epidermal development, growth control, and homeostasis in the face of centrosome amplification. Proc. Natl Acad. Sci. USA 112, E6311–E6320 (2015).
Loncarek, J., Hergert, P. & Khodjakov, A. Centriole reduplication during prolonged interphase requires procentriole maturation governed by Plk1. Curr. Biol. 20, 1277–1282 (2010).
Douthwright, S. & Sluder, G. Link between DNA damage and centriole disengagement/reduplication in untransformed human cells. J. Cell. Physiol. 229, 1427–1436 (2014).
Inanc, B., Dodson, H. & Morrison, C. G. A centrosome-autonomous signal that involves centriole disengagement permits centrosome duplication in G2 phase after DNA damage. Mol. Biol. Cell 21, 3866–3877 (2010).
Ganem, N. J., Storchova, Z. & Pellman, D. Tetraploidy, aneuploidy and cancer. Curr. Opin. Genet. Dev. 17, 157–162 (2007).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Fujiwara, T. et al. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature 437, 1043–1047 (2005).
Krzywicka-Racka, A. & Sluder, G. Repeated cleavage failure does not establish centrosome amplification in untransformed human cells. J. Cell Biol. 194, 199–207 (2011).
Fan, G. et al. Loss of KLF14 triggers centrosome amplification and tumorigenesis. Nat. Commun. 6, 8450 (2015).
Li, J. et al. SAK, a new polo-like kinase, is transcriptionally repressed by p53 and induces apoptosis upon RNAi silencing. Neoplasia 7, 312–323 (2005).
Fukasawa, K., Choi, T., Kuriyama, R., Rulong, S. & Vande Woude, G. F. Abnormal centrosome amplification in the absence of p53. Science 271, 1744–1747 (1996).
Ganem, N. J., Godinho, S. A. & Pellman, D. A mechanism linking extra centrosomes to chromosomal instability. Nature 460, 278–282 (2009).
Quintyne, N. J., Reing, J. E., Hoffelder, D. R., Gollin, S. M. & Saunders, W. S. Spindle multipolarity is prevented by centrosomal clustering. Science 307, 127–129 (2005).
Leber, B. et al. Proteins required for centrosome clustering in cancer cells. Sci. Transl Med. 2, 33ra38 (2010).
Kwon, M. et al. Mechanisms to suppress multipolar divisions in cancer cells with extra centrosomes. Genes Dev. 22, 2189–2203 (2008).
Silkworth, W. T., Nardi, I. K., Scholl, L. M. & Cimini, D. Multipolar spindle pole coalescence is a major source of kinetochore mis-attachment and chromosome mis-segregation in cancer cells. PLoS ONE 4, e6564 (2009).
Zhang, Y. et al. USP44 regulates centrosome positioning to prevent aneuploidy and suppress tumorigenesis. J. Clin. Invest. 122, 4362–4374 (2012).
Silkworth, W. T., Nardi, I. K., Paul, R., Mogilner, A. & Cimini, D. Timing of centrosome separation is important for accurate chromosome segregation. Mol. Biol. Cell 23, 401–411 (2012).
van Ree, J. H., Nam, H. J., Jeganathan, K. B., Kanakkanthara, A. & van Deursen, J. M. Pten regulates spindle pole movement through Dlg1-mediated recruitment of Eg5 to centrosomes. Nat. Cell Biol. 18, 814–821 (2016).
Nam, H. J. & van Deursen, J. M. Cyclin B2 and p53 control proper timing of centrosome separation. Nat. Cell Biol. 16, 538–549 (2014).
Janssen, A., van der Burg, M., Szuhai, K., Kops, G. J. & Medema, R. H. Chromosome segregation errors as a cause of DNA damage and structural chromosome aberrations. Science 333, 1895–1898 (2011).
Crasta, K. et al. DNA breaks and chromosome pulverization from errors in mitosis. Nature 482, 53–58 (2012).
Zhang, C. Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015).
Castellanos, E., Dominguez, P. & Gonzalez, C. Centrosome dysfunction in Drosophila neural stem cells causes tumors that are not due to genome instability. Curr. Biol. 18, 1209–1214 (2008).
Godinho, S. A. & Pellman, D. Causes and consequences of centrosome abnormalities in cancer. Philos. Trans. R. Soc. Lond. B Biol Sci. 369, 20130467 (2014).
Godinho, S. A. et al. Oncogene-like induction of cellular invasion from centrosome amplification. Nature 510, 167–171 (2014).
Mahjoub, M. R. & Stearns, T. Supernumerary centrosomes nucleate extra cilia and compromise primary cilium signaling. Curr. Biol. 22, 1628–1634 (2012).
Han, Y. G. et al. Dual and opposing roles of primary cilia in medulloblastoma development. Nat. Med. 15, 1062–1065 (2009).
Wong, S. Y. et al. Primary cilia can both mediate and suppress Hedgehog pathway-dependent tumorigenesis. Nat. Med. 15, 1055–1061 (2009).
Nano, M. & Basto, R. Consequences of centrosome dysfunction during brain development. Adv. Exp. Med. Biol. 1002, 19–45 (2017).
Chavali, P. L., Putz, M. & Gergely, F. Small organelle, big responsibility: the role of centrosomes in development and disease. Philos. Trans. R. Soc. Lond. B Biol Sci. 369, 20130468 (2014).
Faheem, M. et al. Molecular genetics of human primary microcephaly: an overview. BMC Med. Genom. 8 (Suppl. 1), S4 (2015).
Jayaraman, D. et al. Microcephaly proteins Wdr62 and Aspm define a mother centriole complex regulating centriole biogenesis, apical complex, and cell fate. Neuron 92, 813–828 (2016).
Insolera, R., Bazzi, H., Shao, W., Anderson, K. V. & Shi, S. H. Cortical neurogenesis in the absence of centrioles. Nat. Neurosci. 17, 1528–1535 (2014).
Marjanovic, M. et al. CEP63 deficiency promotes p53-dependent microcephaly and reveals a role for the centrosome in meiotic recombination. Nat. Commun. 6, 7676 (2015).
Martin, C. A. et al. Mutations in PLK4, encoding a master regulator of centriole biogenesis, cause microcephaly, growth failure and retinopathy. Nat. Genet. 46, 1283–1292 (2014).
Morin, X. & Bellaiche, Y. Mitotic spindle orientation in asymmetric and symmetric cell divisions during animal development. Dev. Cell 21, 102–119 (2011).
Lizarraga, S. B. et al. Cdk5rap2 regulates centrosome function and chromosome segregation in neuronal progenitors. Development 137, 1907–1917 (2010).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Konno, D. et al. Neuroepithelial progenitors undergo LGN-dependent planar divisions to maintain self-renewability during mammalian neurogenesis. Nat. Cell Biol. 10, 93–101 (2008).
Sir, J. H. et al. Loss of centrioles causes chromosomal instability in vertebrate somatic cells. J. Cell Biol. 203, 747–756 (2013).
Pilaz, L. J. et al. Prolonged mitosis of neural progenitors alters cell fate in the developing brain. Neuron 89, 83–99 (2016).
Bury, L. et al. Plk4 and Aurora A cooperate in the initiation of acentriolar spindle assembly in mammalian oocytes. J. Cell Biol. 216, 3571–3590 (2017).
Kazazian, K. et al. Plk4 promotes cancer invasion and metastasis through Arp2/3 complex regulation of the actin cytoskeleton. Cancer Res. 77, 434–447 (2017).
Mason, J. M. et al. Functional characterization of CFI-400945, a Polo-like kinase 4 inhibitor, as a potential anticancer agent. Cancer Cell 26, 163–176 (2014).
Holland, A. J. & Cleveland, D. W. Polo-like kinase 4 inhibition: a strategy for cancer therapy? Cancer Cell 26, 151–153 (2014).
Meraldi, P., Lukas, J., Fry, A. M., Bartek, J. & Nigg, E. A. Centrosome duplication in mammalian somatic cells requires E2F and Cdk2-cyclin A. Nat. Cell Biol. 1, 88–93 (1999).
Knockleby, J. & Lee, H. Same partners, different dance: involvement of DNA replication proteins in centrosome regulation. Cell Cycle 9, 4487–4491 (2010).
Meraldi, P. & Nigg, E. A. Centrosome cohesion is regulated by a balance of kinase and phosphatase activities. J. Cell Sci. 114, 3749–3757 (2001).
Tanenbaum, M. E. et al. Kif15 cooperates with eg5 to promote bipolar spindle assembly. Curr. Biol. 19, 1703–1711 (2009).
Blangy, A. et al. Phosphorylation by p34cdc2 regulates spindle association of human Eg5, a kinesin-related motor essential for bipolar spindle formation in vivo. Cell 83, 1159–1169 (1995).
Oliver, T. G. et al. Caspase-2-mediated cleavage of Mdm2 creates a p53-induced positive feedback loop. Mol. Cell 43, 57–71 (2011).
Acknowledgements
The authors thank their laboratory members for helpful discussions and apologize to colleagues whose work could not be cited due to space limitations. Work in the authors' laboratories was supported by grants from the Swiss National Science Foundation (310030B-149641) to E.A.N. and a R01 research grant from the US National Institutes of Health (GM 114119), an American Cancer Society Scholar Grant (129742-RSG-16-156-01-CCG) and a March of Dimes Research Grant (1-FY17-698) to A.J.H.
Author information
Authors and Affiliations
Contributions
Both E.A.N. and A.J.H. researched data for the article, made substantial contributions to the discussion of content, wrote the article, and reviewed and edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Procentriole
-
A newly constructed centriole that is unable to duplicate.
- A tubules
-
A typical microtubule triplet is composed of A, B and C tubules, with the innermost A tubule being built from 13 protofilaments.
- Molecular rulers
-
Molecules of defined size that can be used to set distances between other structures.
- Axoneme
-
The nine-fold symmetrical microtubule-based structure at the centre of cilia and flagella.
- WD40 protein
-
A structural motif of approximately 40 amino acids, often terminating in a tryptophan-aspartic acid (WD) dipeptide.
- Planarians
-
Flatworms used as a model system to study regeneration.
- Hippo pathway
-
A signalling pathway that controls organ size in animals by restraining cell proliferation and promoting apoptosis.
- PIDDosome
-
A protein complex composed of death domain-containing protein CRADD (also known as RAIDD) and p53-induced death domain-containing protein 1 (PIDD1) that is implicated in the activation of caspase 2.
- Cytokinesis failure
-
Failure to physically separate the two daughter cells after chromosome segregation is completed.
- Neuroblasts
-
Dividing neuronal precursor cells.
- Aneuploidy
-
The presence of an abnormal chromosome number that is not a multiple of the haploid chromosome complement.
- Merotelic attachments
-
Spindle microtubule–chromosome attachments in which one kinetochore binds microtubules emanating from two centrosomes located on opposite sides of the mitotic spindle.
- Micronuclei
-
Small nuclei that are separate from the cell nucleus and that contain one or a few chromosomes or chromosome fragments.
- RAC1
-
A small GTPase member of the RAS superfamily with diverse cellular functions.
- Organoids
-
An in vitro culture system that mimics the micro-anatomy of an organ.
- Hypomorphic mutations
-
Mutations that cause a partial loss of gene function.
Rights and permissions
About this article
Cite this article
Nigg, E., Holland, A. Once and only once: mechanisms of centriole duplication and their deregulation in disease. Nat Rev Mol Cell Biol 19, 297–312 (2018). https://doi.org/10.1038/nrm.2017.127
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm.2017.127
- Springer Nature Limited
This article is cited by
-
Amplified centrosomes—more than just a threat
EMBO Reports (2024)
-
Molecular basis promoting centriole triplet microtubule assembly
Nature Communications (2024)
-
Spermatocytes have the capacity to segregate chromosomes despite centriole duplication failure
EMBO Reports (2024)
-
The two sides of chromosomal instability: drivers and brakes in cancer
Signal Transduction and Targeted Therapy (2024)
-
Male meiotic spindle poles are stabilized by TACC3 and cKAP5/chTOG differently from female meiotic or somatic mitotic spindles in mice
Scientific Reports (2024)